Abstract
Temporomandibular joint (TMJ) osteoarthritis (OA) is a progressive degenerative disease in which the intricate mechanisms of the condyle cartilage damage are yet to be disclosed. A detailed characterization of cellular composition and signal networks involved in the pathogenesis is in need. This study, for the first time, identified chondrocytes heterogeneity and subchondral immune microenvironments based on single-cell transcriptomes of the mouse mandibular condyle, and provided insights into the cellular and signaling basis of the homeostasis of the condyle and pathogenesis of TMJOA by healthy-to-diseased comparisons. The differentially expressed gene pattern distinguished 8 chondrocyte subsets and 9 immune cell types in the condyle. Close interactions were found between these cell populations, with enhanced communication between chondrocytes and immune cell subsets in the condyles subjected to anterior premature contact (APC) model or unilateral anterior crossbite (UAC) model. The single-cell transcriptomic changes suggested predominant roles of neutrophils in immune-chondrocyte interactions in these occlusal disorder models. Neutrophils-derived TNF-α signaling was inferred to be the most greatly enhanced information flow in the condyle of the APC model, while increased Thbs1-Sdc4 signaling in mediating neutrophils-to-chondrocytes interactions was indicated in the UAC-model condyle, with predominant BSP signaling. Interfering in these signaling by drug administration demonstrated therapeutic potential for the APC- or UAC-induced TMJOA.
Similar content being viewed by others
Introduction
Temporomandibular joint (TMJ) osteoarthritis (OA) is a progressive degenerative disease that affects mandibular condylar cartilage and subchondral bone, causing pain, movement limitation and dysfunction of the joint.1,2,3 The mandibular condyle is distinct from articular hyaline cartilage in cellular components and extracellular matrix.1 TMJOA has highly complicated pathogenesis which is different from OA of other joints. Previous studies focused on the signals and pathological events in cartilage matrix degradation, chondrocyte metabolism and cell death, subchondral bone remodeling, and synovial fluid and tissue-mediated inflammation.1,2,3,4,5 However, the intricate mechanisms by which the degenerative changes in the cartilage and bone develop are far from being clearly elucidated.
It has been well established that inflammation dysregulates chondrocyte metabolism and causes matrix degradation.2,3 Outside stimuli trigger inflammatory responses in synovial tissue and cause tissue damage in the superficial layer of the condyle, leading to cartilage erosion and cracking. Recently, increasing evidence suggested the osteochondral junction as another site of importance where cartilage erosion takes place.6,7,8,9 Chondrocyte hypertrophy, abnormal endochondral osteogenesis and angiogenesis contribute to OA progression.6,7,8,9 Although with microstructural and histopathological changes observed, the molecular mechanisms are poorly understood. These mechanisms might involve complex cellular and signal networks. The lack of a detailed characterization of these networks has thusfar hampered understanding of the pathological mechanisms. This study was performed to bridge this research gap.
In this study we profiled single-cell transcriptomes of the mandibular condyles from mice of TMJ homeostasis or TMJOA induced by two types of occlusal disorder model. The single-cell transcriptional atlas reveals high heterogeneity of chondrocytes and complicated immune microenvironments in the condyle. Close interactions were inferred between chondrocyte sub-populations and multiple subsets of immune cells. The single-cell transcriptomic changes in occlusal disorder-induced TMJOA suggested predominant roles of neutrophils in immune-chondrocyte interactions that mediated inflammatory damage and deterioration of chondrocyte functions. Noteworthily, the dominant signaling that mediated the immune-chondrocyte interactions were different between the two types of occlusal disorder model. The functions of the signaling were in line with the TMJ histopathological changes, suggesting different immunoregulating mechanisms. We next tested the therapeutic potential of these mechanisms by interfering in these signaling using drug administration during model constructions. These mechanisms may provide new insights into the pathogenesis and potential therapeutic targets for TMJOA.
Results
Single-cell transcriptional profiling identified four principal cell types in mouse condyles
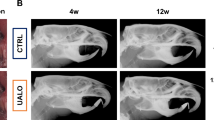
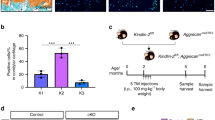
The anterior premature contact (APC) and unilateral anterior crossbite (UAC) models induced remarkable TMJOA-like lesions in the condyles (Fig. 1a–c). The histology demonstrated the rough surface, erosion of the cartilage by ossification regions, reduced thickness of the cartilage layer and disordered arrangements of cells in the cartilage (Fig. 1c). After unsupervised clustering of the 3 datasets combined (ctrl, APC, UAC), the captured cells were segregated into 14 distinct cell clusters (Fig. 1d), which were classified into chondrocytes, immune cells and endothelial cells/pericytes (Fig. 1e, Supplementary Fig. 1). Cells in clusters 0 ~ 4 and 6 ~ 10 highly expressed chondrocyte markers Sox9, Acan, and were identified as ‘chondrocytes’. Clusters 5 and 11 ~ 13 were annotated as ‘immune cells’ based on enrichment of Csf3r, Lyz2, Slpi, Retnlg and Cybb. Cluster 14 was identified as ‘endothelial cells/pericytes’ by expressing Pecam1, Cdh5, Rgs5 and Mcam. Cluster 14 was further classified into endothelial cells and pericytes (Supplementary Fig. 2). Compared with the control group, the condyles of the APC or UAC model had decreased proportions of chondrocytes and increased proportions of immune cells (Fig. 1f, g).
Single-cell transcriptional profiling of the cells in the mandibular condyle. a The time frame and model construction. b A Schematic of the mouse mandibular condyle tissue harvested for single-cell transcriptomic analyses. c Saffron-O and fast green staining of the condyles of the healthy mice (ctrl) and mice subjected to the APC or UAC model. Scale bar: 50 μm. d Cell clusters from scRNA-seq analysis visualized by UMAP. e Feature plot showing the expression of chondrocytes markers, immune cells markers and endothelial cells/ pericytes markers. f Bar plot showing the proportion of cell types in ctrl, APC, and UAC group, respectively. g Visualization of cell clusters in scRNA-seq analysis of ctrl, APC, and UAC by UMAP
Chondrocyte heterogeneity in healthy and TMJOA condyles
Chondrocytes were the main cellular components in the mandibular condyle, constituting 91.36% of the total identified cells and 98.65% of the non-immune cells during homeostasis (Fig. 1f). The chondrocytes were heterogeneous in terms of transcriptional expressions and biological functions. Single-cell RNA sequencing (ScRNA-seq) of the 3 datasets combined (ctrl, APC, UAC) identified 16 cell clusters with distinct gene expression profiles (Fig. 2 & Supplementary Figs. 3, 4) and functions (Supplementary Fig. 5). Cells in cluster 7 and 9 were classified as ‘mature chondrocytes’ based on enrichment of chondrocyte markers Col9a1, Col11a2, Scrg1. Custer 0 and 2, also showing enrichment of these chondrocyte markers, was labeled ‘Col10a1high hypertrophic chondrocytes (HTC)’ due to high expressions of HTC markers Col10a1 and enrichment for GO terms ‘biomineralization’, ‘ossification’ and ‘osteoblast differentiation’. These clusters also abundantly expressed Ibsp. Cluster 4, 5 and 11 highly expressed pro-inflammatory markers Nos2, and was thus labeled ‘inflammation-associated (IA) chondrocytes’. These clusters also abundantly expressed Cxcl1, Cxcl2, Cx3cl1 and TGF-β signaling gene Inhba. GO enrichment analysis suggested functional features of leukocyte migration and activation, cytokine- and chemokine-mediated signaling for this cluster. Cells in cluster 6, 13, 14 and 15 were annotated as ‘Mmp13high chondrocytes’ featured by high expressions of Mmp13. These clusters also abundantly expressed Ifitm3 and Lum. Cluster 1 was labeled ‘reparative chondrocytes’ based on remarkably increased Atf3, Hspa1a and Hspa1b, and enrichment for GO terms ‘response to unfolded protein’ and ‘cellular response to topologically incorrect protein’, ‘negative regulation of inflammatory response’ and ‘negative regulation of leukocyte activation’. Clusters 3, 8 and 16 highly expressed fibroblast/fibrosis markers S100a4 (also known as fibroblast-specific protein 1, FSP1) and Abi3bp, thus were incorporated and annotated as ‘fibrotic chondrocytes’. These clusters also abundantly expressed Thbs4 and Mfap4. The enriched biological functions of this cluster were closely related to extracellular matrix organization, cell-matrix adhesion, regeneration and wound healing. Cluster 10 was identified as ‘mineralization-associated chondrocytes’ due to highly specialized functional features of biomineralization suggested by GO enrichment analysis. These cells highly expressed Ifitm5 and osteocalcin encoding genes Bglap and Bglap2. Cells in cluster 12 abundantly expressed stem-cell-related markers Stmn1, Birc5 and cell cycle genes Cdk1, Top2a, Cenpa and Mki67, thus was annotated as ‘chondrocyte progenitors’, with enriched biological processes related to mitosis and cell cycle.
The differentially expressed gene pattern distinguished 8 chondrocyte populations in the mandibular condyles in homeostasis and TMJOA. a Chondrocytes subtypes from scRNA-seq analysis visualized by UMAP. b Dot plot showing the expression of selected markers of various chondrocyte subtypes. c Multiplexed immunofluorescence staining of the healthy condyle for detection of progenitor cell markers BIRC5 (green), STMN1 (red) and DHFR (purple). Scale bar: 50 μm for the low-power field, 20 μm for the high-power field. d Co-immunofluorescence staining of the healthy condyle for detection of the fibrotic markers ABI3BP (green) and S100A4 (red). Scale bar: 50 μm
During homeostasis, the chondrocyte progenitors (Fig. 2c) and fibrotic chondrocytes (Fig. 2d) were found to reside in the superficial zone in the Condyle. The NOS2high IA chondrocytes resided below the superficial layer (Fig. 3a). The Col10a1high, Mmp13high, and Bglaphigh chondrocytes were distinct clusters, among which the cells highly expressing COL10 resided in the deep layer adjacent to the osteochondral junction, while the MMP13high OCNhigh chondrocytes were above the deep layer (Fig. 3b).
Comparison of cell composition and pseudotime developmental trajectory of the chondrocytes between healthy and TMJOA condyles. a Co-immunofluorescence staining of the condyle for the ‘fibrotic chondrocytes’ marker ABI3BP (green) and the ‘IA chondrocytes’ marker NOS2 (red). Scale bar: 50 μm for the left panels, 20 μm for the right panels. b Co-immunofluorescence staining of the condyle for Collagen-X (COL10, green), MMP13 (red) and osteocalcin (OCN, purple). Scale bar: 50 μm for the low-power field, 20 μm for the high-power field. c Bar plot showing the proportion of chondrocytes subtypes in Ctrl, APC, and UAC groups, respectively. Visualization of cell clusters in scRNA-seq analysis of ctrl, APC, and UAC chondrocytes subtypes by UMAP. d Pseudo-time analysis of chondrocytes subtypes using Monocle3 of Ctrl, APC, and UAC groups
Different TMJOA models induced different changes in frequency, composition and transcriptional phenotypes of chondrocyte and immune cell populations
ScRNA-seq revealed the transcriptomic changes in the condyles of occlusal disorder-induced TMJOA (Fig. 3). All the identified cell clusters were present in healthy and TMJOA condyles. The APC and UAC models induced reduced frequency of chondrocytes (Fig. 3c). Compared with the healthy condyle, the condyles of the APC model had increased proportions of Mmp13high chondrocytes, Col10a1high HTCs, and reduced proportions of IA chondrocytes, while the UAC-model condyle had reduced proportions of Col10a1high HTCs, IA chondrocytes, and increased proportions of Mmp13high chondrocytes, fibrotic chondrocytes and reparative chondrocytes (Fig. 3c). The proportion of chondrocyte progenitors witnessed a slight increase in the APC model and the UAC model compared to the control, which accounted for 4.07% and 3.67% in the APC model and the UAC model, respectively, and 3.48% in the control. This was in line with the immunofluorescence (Supplementary Fig. 6) that the TMJOA condyles showed regionally increased BIRC5high STMN1high cells though with regional decrease of these markers, indicating simultaneous occurrence of destruction and repair. The pseudotime analysis suggested disturbed developmental trajectory of chondrocytes in the APC and UAC models (Fig. 3d), consistent with the immunofluorescence (Fig. 3a, b) that showed disarranged and compacted chondrocytes losing cluster-dependent phenotypic characteristics during TMJOA.
The immune cells were clustered into 12 populations, which were further annotated as 9 distinct cell types (Fig. 4 & Supplementary Figs. 7, 8). The occlusal disorder models induced remarkably increased frequency of immune cells (Fig. 1f). The cell composition of the immune cell clusters indicated an expansion of neutrophils, especially Stage II neutrophil subcluster in TMJOA, in contrast of remarkable shrinkage in monocytes, macrophages and B cells (Fig. 4d). Among the top 10 DEGs in the neutrophils, we observed significantly upregulated Abcg1 and downregulated Ftl1 genes in the condyle of the APC model, indicating dysregulated lipid homeostasis and cellular iron metabolism; while in the UAC model, increased expressions of Hspala/b,Il1r2 and Thbs1 in neutrophils were noted, indicating enhanced repair and tissue remodeling (Supplementary Fig. 9).
Single-cell transcriptomic atlas of the immune microenvironment of the TMJ condyle during homeostasis and TMJOA. a Immune cells subtypes from scRNA-seq analysis visualized by UMAP. b H & E staining showing the histology of the condyles. Scale bar: 50 μm. c Dot plot showing the expression of selected markers of various immune cells subtypes. d Bar plot showing the proportion of immune cells subtypes in ctrl, APC, and UAC, respectively. Visualization of cell clusters in scRNA-seq analysis of ctrl, APC, and UAC immune cells subtypes by UMAP
Transcriptionally inferred immune-chondrocyte interactions were enhanced and differentially regulated between the APC and UAC models
Transcriptionally inferred interactions were found between chondrocytes and immune cell subsets,and was enhanced in condyles of the occlusal disorder models (Fig. 5a), with distinct crosstalk signals (Fig. 5b, c). Compared with the healthy control, TNF signaling pathway was inferred to be the most greatly enhanced information flow in the condyle of the APC model (Fig. 5b), while BSP was indicated as the most increased signaling in the UAC model (Fig. 5c). Immunohistochemical staining confirmed high expression of TNF-α in chondrocytes and immune cells in the condyle of the APC model but not as high in the UAC model (Supplementary Fig. 10a). The immune cells at the osteochondral junction also expressed increased levels of IL-6 and IL-10 (Supplementary Fig. 10a).
Transcriptionally inferred cell-cell interactions and signaling flows in the APC- or UAC-exposed condyles compared with the healthy control. a Circle plots showing differential number of interactions and interaction strength compared the APC with the ctrl group, or compared the UAC group with the ctrl group. Red colored edges indicate increased signaling in the APC or UAC group and blue colored edges indicate decreased signaling in the ctrl group. b Bar plots showing the overall information flow of each signaling pathway in the ctrl group and the APC group. c Bar plots showing the overall information flow of each signaling pathway in the ctrl group and the UAC group
Single-cell transcriptome analysis suggested that the outgoing TNF signaling, while derived mainly from macrophages during homeostasis, greatly increased and predominantly derived from Stage II neutrophils in the condyle of the APC model, with enhanced potential interactions with other immune cells and chondrocytes (Fig. 6a, b, Supplementary Fig. 10b). The immunofluorescence showed significantly increased production of TNF-α in the chondrocytes, MPO+ neutrophils and CD68+ macrophages in the APC model (Fig. 6c, d). The outgoing BSP signaling, which increased greatly in the UAC model, derived mainly from Col10a1high HTCs, mineralization-associated chondrocytes and mature chondrocytes (Supplementary Fig. 10c). The signaling analysis suggested dominant roles of Thbs1-Sdc4 and Col2a1 signaling in mediating the Stage I/II neutrophils-to-chondrocyte interactions, with much stronger output of these signals from neutrophils towards chondrocytes in the UAC model than in healthy control (Fig. 7a, b). The Col2a1 signaling was enhanced more among chondrocytes than in neutrophils-to-chondrocyte interactions (Fig. 7a). The immunofluorescence (Fig. 7c, d) showed widespread BSP-positive areas from the cartilage to subchondral bone in the condyle of the UAC model, with high immunoreactivity in the bone and osteochondral junction areas. These areas co-localized with expressions of the endothelial marker CD31.
Single-cell transcriptome analysis highlight predominant role of neutrophil-derived TNF signaling in mediating immune-chondrocyte interaction in the APC-exposed condyles. a Circle plot of TNF signaling network. b Bubble plot of the communication probability of all the significant ligand-receptor pairs that contributed to increasing signaling sent from Stage II neutrophils to each cell population in chondrocytes. c Co-immunofluorescence staining of the condyle for Myeloperoxidase (MPO, green) and TNF-α (red), or for CD68 (green) and TNF-α (red). Co-localization of MPO and TNF-α indicates production of TNF-α from neutrophils. Scale bar: 50 μm for the low-power field, 10 μm for the high-power field. d Semi-quantitation of the expression levels of MPO, CD68 and TNF-α. *P < 0.05. **P < 0.01. ***P < 0.001. ****P < 0.000 1
Enhanced chondrocyte-derived BSP signaling accompanied by increased angiogenic marker expression and neutrophil-derived THBS signaling in the UAC-exposed condyles. a Circle plot of THBS and COLLAGEN signaling networks in the ctrl group and the UAC group. b Bubble plot of the communication probability of all the significant ligand-receptor pairs that contributed to increasing signaling sent from Stage I neutrophils/ Stage II neutrophils to each cell population in chondrocytes. c Co-immunofluorescence staining of the condyle for BSP (green) and CD31 (red). Scale bar: 50 μm for the low-power field, 10 μm for the high-power field. d Semi-quantitation of the expression levels of BSP and CD31. **P < 0.01. ***P < 0.001
Signal-targeted drug administration alleviated cartilage erosion in TMJOA induced by different occlusal disorders
Compared with the healthy condylar cartilage with chondrocyte layers of well-arranged hierarchy and good integrity, the APC- or UAC-induced lesions presented with significantly reduced thickness of the cartilage layer and aggravated severity of OA assessed by Modified Mankin scores (P < 0.05, Fig. 8). Administration of TNF-α inhibitor Etanercept significantly ameliorated the cartilage damage induced by the APC model (P < 0.05, Fig. 8a, b), while the protective effect was not so pronounced in the mice subjected to the UAC model (Fig. 8c, d). The CXCR2/CXCR1 inhibitor Navarixin attenuated OA induced by the UAC model (P < 0.05, Fig. 8c, d), but had no statistical significance in affecting the condyle lesion caused by the APC model (På 0.05, Fig. 8a, b).
Signal-targeted therapeutic effect on TMJOA induced by different occlusal disorders. a Representative Masson staining, Saffron-O and fast green staining showing histological changes of the condyle subjected to the APC model, with or without administration of Etanercept or Navarixin. Scale bar: 50 μm. b Quantitative assessment of the histological changes evaluated by the thickness of cartilage and Modified Markins score. *P < 0.05. c Representative images showing histological changes of the condyle subjected to the UAC model, in the presence or absence of Etanercept or Navarixin treatment. Scale bar: 50 μm. d Statistical analyses of the thickness of cartilage and Modified Markins score. *P < 0.05
Discussion
This study is the first to identify transcriptome heterogeneity of chondrocytes in mouse TMJ condyle. A previous study on TMJ condyle of human embryos annotated 15 cell clusters including proliferating cells, chondrocytes, HTC, osteoblasts, leukocytes and multiple types of mesenchymal cells.10 Another study reported a single-cell transcriptomic atlas of human TMJOA condyle but without healthy control.11 It identified immune cells including macrophages, neutrophils, natural killer cells, T cells and B cells; and segregated chondrocytes into five clusters including Birc5 high cartilage progenitor cells.11 The other four clusters expressed genes involved in immunity, angiogenesis, neurogenesis and ossification but were unnamed. Thus far the data of healthy adult TMJ condyle is still lacking. This study, by presenting healthy-to-diseased comparisons, provided insights into the homeostasis of TMJ condyle and pathogenesis of TMJOA at single-cell resolution.
The cartilage of the condyle is conventionally divided into proliferative, mature and hypertrophic layers, among which the proliferative zone functions as a cell reservoir, while the hypertrophic zone has mineralizing features with expressions of MMP13, RUNX2, type I and X Collagens.12,13,14,15,16 Notably, the present study provided evidence that Mmp13high chondrocytes, Col1a1high chondrocytes and Col10a1high chondrocytes were distinct clusters. The chondrocytes positive for MMP13 or osteocalcin were found to reside mainly in the proliferative rather than the hypertrophic zone. The Collagen-Xhigh chondrocytes, displaying hypertrophic morphology and residing in the deep layer, were partially but not all positive for MMP13 or osteocalcin. In line with the developmental sequence indicated by the spatial relations, pseudotime analysis suggested that Mmp13high chondrocytes, Col1a1high Bglaphigh Ifitm5high mineralization-associated chondrocytes were at stages preceding mature chondrocytes, while Col1a1high chondrocytes were at the end of the trajectory. These findings suggested that in TMJ condyle chondrocytes possess mineralizing features early in proliferating stages. Collagen-X, rather than MMP13 or osteocalcin, could serve as a marker for HTC.
Consistent with the single-cell atlas of cartilage of other joints,17 this study identified STMN1high BIRC5high DHFRhigh chondrocyte progenitors in the superficial layer of the condyle. These cells have similar characteristics to and could also be termed ‘proliferating chondrocytes’. Progenitor cells with different markers but similar functions have also been reported in other studies on the TMJ.18,19,20,21 An ‘Mef2chigh Pth1rhigh pre-HTC cluster was previously identified in knee cartilage but lacks counterparts in the TMJ condyle.22,23 As revealed by our results, the enrichment of Mef2c and Pth1r was found in clusters of mature chondrocytes and HTCs. It is likely that the pre-hypertrophic phenotype was transitional and could not be clearly separated. Analogously, we didn’t annotate a ‘pre-fibrotic chondrocytes’ subtype which was previously reported.22 A S100A4high ABI3BPhigh ‘fibrotic chondrocytes’ population was annotated and found to reside in the superficial zone of the condyle cartilage, following the chondrocyte progenitors along the pseudotime trajectory. A NOS2 high chondrocyte population characterized by pro-inflammatory phenotypes was identified closely below the superficial layer, following the fibrotic chondrocytes along the pseudotime trajectory. Noteworthily, the orderly arranged spatial relations and pseudotime trajectory of the chondrocyte clusters during homeostasis were remarkably disturbed by occlusal disorder-induced TMJOA. The diseased condyles showed disordered pseudotime trajectory and disorganized cells compacted in the eroded cartilage layer. These results highlighted the distinct transcriptomic profile of TMJ condyle chondrocytes relative to other joints, and suggested that chondrocyte developmental trajectory might be implicated in TMJOA pathogenesis.
The Single-cell transcriptome analysis revealed potentially enhanced BSP signaling in the condyle of the UAC model compared with homeostasis. Previous studies supported the implication of BSP as a key mediator of angiogenesis in bone formation, fracture healing and the pathology of OA, in line with our findings that the BSP-immunoreactive areas co-localized with angiogenesis marker CD31.24,25,26 Neo-angiogenesis at the osteochondral junction was an important pathogenic feature of TMJOA.3 New blood vessels invade the cartilage, promoting chondrocyte hypertrophy and matrix mineralization. Endochondral angiogenesis, coupled with neoneurogenesis, exacerbates pain and promotes cartilage ossification.27,28,29
In this study, the histological examinations and scRNA-seq analysis implicated angiogenesis as one of the major pathogenic factors for UAC-induced TMJOA. Considering the pro-angiogenic effect of Thbs1-Sdc4 signaling,30,31,32 the increased immune cell infiltration subsequent to blood vessel invasion might exacerbate cartilage erosion by promoting a positive feedback loop. Enhanced Thbs1-Sdc4 signaling was found to be the dominant change in the UAC-perturbed chondrocyte-immune cell interaction, highlighting the crucial role of angiogenesis in UAC-induced pathogenesis. Thus, we tested the therapeutic effect of Navarixin administration on UAC-model mice, attempting to block this pathogenic process by targeting neutrophil infiltration. As a result, condyle cartilage damage was significantly attenuated by administration of the CXCR2/CXCR1 inhibitor Navarixin, though far from fully recovering to normal. The limitation in the therapeutic effect might result from multiple possible mechanisms. First, production of proangiogenic factors from chondrocytes3; second, weak resistance of cartilage to vessel formation due to cartilage matrix degradation and proteoglycan depletion2,3; third, local hypoxia microenvironment where hypoxia-inducible factor 1 (HIF-1) activates its target genes that promote neovascularization.2,33,34 There is hardly a solution that could inhibit angiogenesis by blocking all the three ways. There were multiple angiogenic factors such as HIF-1,2,33,34 high-mobility group box 1 (HMGB1),35 VEGF,28 Notch,36 Platelet-Derived Growth Factor BB(PDGF-BB),37 IL-6 and IL-1β.38,39 These signaling might contribute to forming of compensating mechanisms that promote neo-angiogenesis when CXCR2/CXCR1 signaling was suppressed. For instance, aberrantly activated Notch signaling could be mediated by degradative proteinases and inflammatory cytokines such as MMP13, IL-1β and IL-6.40,41
One of the reasons for the difficulty in treating TMJOA is that its pathogenesis involves diverse factors of genetic, mechanical and biochemical causes. The diverse etiologies may correspond to distinct pathogenic mechanisms. The animal models of TMJOA are currently established by creation of disordered occlusion, intra-articular injection of inflammatory mediators, genetic modification or surgical manipulations such as partial perforations of the discs.1,3 Despite the success in inducing OA-like lesions, these models are still insufficient in mimicking the complex clinical conditions where OA may result from a single or combined etiological factors. UAC and bite-raising models were used to mimic TMJOA based on the knowledge that excessive or unbalanced joint loading causes mechanical stimuli that induce and aggravate TMJOA.1,3
In this study, the difference in transcriptome-inferred immune-chondrocyte crosstalk mechanisms between the two models suggested that TMJOA pathogenesis might be highly context-dependent rather than represented by mechanisms unveiled through a specific model. Even the occlusal loading-induced TMJOA involved distinct pathogenesis depending on the types of occlusal disorder, and the pathogenesis-targeted therapies should be adjusted accordingly. However, the functional implications of this study should be interpreted with caution due to inherent limitations of the transcriptomic data. The scRNA-seq data inferred potential cell-cell communication mediated by ligand-receptor co-expression patterns, rather than providing direct evidence of protein-protein interactions necessary to validate functional binding events. Further studies are warranted to prioritize functional validation to translate observational findings into actionable therapeutic strategies.
Previous study on human TMJOA condyle11 identified similar immune cell types and chondrocyte clusters with different functional enrichment as compared to our study. However, limited comparisons could be made regarding pathogenic mechanisms due to lack of a healthy control and missing information on TMJOA etiology for the human sample.11 The results of this study could not be directly extrapolated to human diseases as murine and human TMJ differ anatomically and pathologically. However, this study demonstrated the highly intricate TMJOA pathogenesis that may vary between different etiological factors. The pathogenic mechanisms and targeted therapies for TMJOA may be more appropriately stated based on specific etiological conditions, rather than being universally applied. It would be of significance to extend further investigations into diseased-and-healthy comparisons in human tissues.
Materials and methods
Animal experiments
The animal experiments were conducted in compliance with the protocols approved by the Ethics Committee of West China Stomatology School of Sichuan University. To investigate the single-cell transcriptomes and histology of the condyle, 90 female C57BL/6 mice aged 4 weeks old (Ensiweier, Chengdu, China) were purchased from the same batch and were bred under identical conditions to minimize interindividual variability (for details see Supplementary data). The 90 mice were evenly allocated into three groups: healthy control, APC model and UAC model. The APC model was constructed by binding a resin ramp at the palatal side of the upper incisors (Fig. 1a). The UAC model was constructed by binding a pair of stainless steel tubes onto the left maxillary and mandibular incisors. The mice were forced to shift the mandible to the left to as the lower tube was longer than the incisor to prevent normal occlusion with the upper tooth (Fig. 1a). The APC and UAC models created two types of occlusal disorders that mimicked clinical scenarios that may happen during the development or restoration of dentition. The experimental staff checked the mice twice per day to ensure retainment of the resin ramps or steel tubes. The devices were fixed promptly in the cases of missing caused by scratching or clenching. The mice were sacrificed with CO2 euthanization at the endpoints of 4 weeks.42 For each group, the mandibles of 20 mice were harvested for single-cell transcriptomes and that of 10 mice were for harvested for histology, histochemistry and immunofluorescence.
To examine the therapeutic effect of signal-targeted drug administration on APC- or UAC-induced murine TMJOA, 42 female C57BL/6 mice aged 4 weeks old (Ensiweier) were bred and evenly allocated into seven groups: vehicle control, APC model, APC model with Etanercept administration, APC model with Navarixin administration, UAC model, UAC model with Etanercept administration, UAC model with Navarixin administration. For Etanercept administration, the TNF-α inhibitor Etanercept (Medchemexpress LLC, NJ, USA, 5 mg/kg) was applied by intraperitoneal injection (i.p.) every other day from the onset of APC or UAC model. For Navarixin administration, the CXCR2/CXCR1 inhibitor Navarixin (Medchemexpress LLC, 30 mg/kg, i.p.) was administered every other day during the experimental time frame. For the vehicle control and the APC or UAC model, the mice received i.p. injection of 0.5% DMSO. At the endpoint, the mice were sacrificed under euthanization and their mandibles were harvested for histological examination.
Cell preparation
The articular cartilage tissue was dissected from the Condyles (as illustrated in Fig. 1b). Upon collection, the tissues underwent a thorough washing process in ice-cold PBS (Hyclone). Subsequently, dissociation was performed for 4 h using Collagenase Ⅱ (Sigma, 2 mg/mL) and Trypsin (Sigma, 0.20%), strictly adhering to the manufacturer’s instructions. Depending on the viscosity of the homogenate, an optional DNase Ⅰ (Sigma) treatment was applied. Fluorescence Cell Analyzer (Countstar® Rigel S2) with AO/PI reagent after removal erythrocytes (Solarbio) and then debris and dead cells removal was decided to be performed or not (Miltenyi). Finally fresh cells were washed twice in the RPMI1640 (Gibco) and then resuspended at 1 × 106 cells per mL in RPMI1640 and 2% FBS (Gibco).
Single cell library construction and sequencing
ScRNA-seq libraries were prepared using SeekOne® MM Single Cell 3’ library preparation kit (SeekGene). Briefly, a suitable quantity of cells was introduced into the flow channel of the SeekOne® MM chip, which featured 170 000 microwells. These cells were permitted to settle into the microwells under the influence of gravity. Following the removal of cells that failed to settle, an adequate volume of Cell Barcoded Magnetic Beads (CBBs) was dispensed into the flow channel. With the aid of a magnetic field, these CBBs were also allowed to settle into the microwells. Subsequently, any excess CBBs were washed away. The cells within the MM chip were lysed, thereby releasing RNA that was subsequently captured by the CBBs situated in the same microwell. All CBBs were then gathered, and reverse transcription was carried out at 37 °C for a duration of 30 min. This process enabled the labeling of cDNA with the cell barcode present on the beads. An Exonuclease I treatment was subsequently applied to eliminate any unused primer on the CBBs. Afterward, the barcoded cDNA on the CBBs was hybridized with a random primer. This primer had a reads 2 SeqPrimer sequence at its 5’ end and was capable of extending to form the second strand of DNA, with the cell barcode positioned at the 3’ end. The resulting second strand of DNA was then denatured off the CBBs, purified, and amplified through a PCR reaction. The amplified cDNA product was subsequently cleansed to eliminate any unwanted fragments and was ligated with full-length sequencing adapters and sample indices via indexed PCR. The indexed sequencing libraries were then purified using VAHTS DNA Clean Beads (Vazyme), quantified with a Qubit fluorometer (Thermo Fisher Scientific), and analyzed using a Bio-Fragment Analyzer (Bioptic Qsep400). Finally, these libraries were sequenced on an illumina NovaSeq 6000 platform, with a read length of PE150.
Quality control and scRNA-seq data pre-processing
The raw sequencing data was processed by Fastp firstly43 to trim primer sequences and low-quality bases. Subsequently, Seeksoul tools (Seekgene, V1.0.0) was used to process the resting data, demultiplex cellular barcodes and map reads to the mouse GRCm39 reference genome to generate a feature-barcode matrix, which was used for the downstream analyses. For each sample, the data output was analyzed using the Seurat package (version 4.3.0.1) within the R software environment (version 4.2.1). Low-quality cells were filtered by removing cells with expressed genes (> 7 500 and < 500) or the percent of mitochondrial genes > 10%. Additionally, cells identified as doublets were eliminated using the DoubletFinder package (version 2.0.4). A total of 20 496 unique genes were detected across 37636 cells, which were then selected for subsequent bioinformatics analysis.
Data integration, dimension reduction and cell clustering
The scRNA-seq data from the control, APC model and UAC model were analysed using “Seurat” package (version 4.3.0.1). After data pre-processing, the 2 000 most variable genes were selected based on variance stabilizing transformation (VST). Then we identified anchors for data integration using the “FindIntegrationAnchors” function. Next, these identified anchors were fed into the “IntegrateData” function, which facilitated the creation of a new integrated matrix that consolidates all three datasets. The data was then scaled, and principal component analysis (PCA) was performed to reduce its dimensionality. Batch effects were removed by leveraging the “harmony” reduction method in the “Seurat” package on the top 10 PCA components. Clustering analysis was then conducted on the integrated datasets using the “FindClusters” function from the “Seurat” package. Cells were clustered using the shared nearest neighbour (SNN) modularity optimisation-based clustering algorithm. The resulting clusters were visualized using the uniform manifold approximation and projection (UMAP) algorithm. Clustree was used to determine the resolution of the clustering, with resolution = 0.4 for the overall cells clustering. For sub-clustering analysis, a similar workflow was applied, encompassing the identification of variable genes, dimensionality reduction, and clustering. The resolutions of the chondrocytes subtypes, the immune cells subtypes and the endothelial cellsperpericytes were 1.1, 0.4, and 0.4, respectively.
scRNA-seq data analyses
Cell clusters annotation,ontology enrichment analysis, pseudotime trajectory analysis and Cell-cell communication analysis were performed as described in the Supplementary materials.
Histological and histochemical examination
After being fixed in paraformaldehyde (4%) overnight at 4 °C, the mandibles of the mice were dehydrated and then embedded in paraffin. Sections of the Condyle were made in saggital direction at the thickness of 4 μm. H & E staining was conducted to observe the histology. Masson staining was performed using Masson Stain Kit (Powerful Biology, Wuhan, China). Saffron-O and fast green staining was performed using Saffron-O and Fast Green Stain Kit (Powerful Biology). By using Image J (v1.8.0, NIH), the thickness of cartilage was calculated as the mean value of the thickness measured at quarter points of the anterior, middle, and posterior region of the condyle. Each sample was assessed three times and the average thickness was used for statistical analysis. The severity of cartilage damage was evaluated by Modified Markins score.44,45 Histochemical staining and double/multiplexed immunofluorescence were conducted as described in the Supplementary materials.
Statistical analyses
Statistical analyses of scRNA-seq data were conducted using a combination of tools, including Seurat, clusterProfiler, Monocle3, and CellChat. Differential expression analysis was performed using the wilcoxon rank-sum test. All statistical analyses were carried out using R software (version 4.2.1). Immunochemical semiquantitative data were analyzed using GraphPad Prism software (GraphPad Software Inc, San Diego, CA, USA). Differences between groups were detected by one-way analysis of variance (ANOVA) followed by Sidak’s multiple comparison. Data were presented as mean ± standard deviation (SD). A P-value < 0.05 was considered to be statistically significant.
Data availability
The data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Lu, K. et al. Molecular signaling in temporomandibular joint osteoarthritis. J. Orthop. Transl. 32, 21–27 (2022).
Cardoneanu, A. et al. Temporomandibular joint osteoarthritis: Pathogenic mechanisms involving the cartilage and subchondral bone, and potential therapeutic strategies for joint regeneration. Int. J. Mol. Sci. 24, 171 (2022).
Li, B., Guan, G., Mei, L., Jiao, K. & Li, H. Pathological mechanism of chondrocytes and the surrounding environment during osteoarthritis of temporomandibular joint. J. Cell. Mol. Med. 25, 4902–4911 (2021).
Sillmann, Y. M. et al. Intra-articular injection of orthobiologics for temporomandibular joint osteoarthritis: A systematic review of randomized controlled trials. Int. J. Oral Maxillofac. Surg. S0901502725000086 https://doi.org/10.1016/j.ijom.2025.01.008 (2025).
Tang, Y. H., Van Bakelen, N. B., Gareb, B. & Spijkervet, F. K. L. Arthrocentesis versus conservative treatments for temporomandibular joint disorders: A systematic review with meta-analyses and trial sequential analyses. J. Cranio-Maxillo-fac. Surg. Off. Publ. Eur. Assoc. Cranio-Maxillo-fac. Surg. 53, 250–261 (2025).
Luo, X., Jiang, Y., Bi, R., Jiang, N. & Zhu, S. Inhibition of notch signaling pathway temporally postpones the cartilage degradation progress of temporomandibular joint arthritis in mice. J. Cranio-Maxillo-fac. Surg. Off. Publ. Eur. Assoc. Cranio-Maxillo-fac. Surg. 46, 1132–1138 (2018).
Qin, W. et al. Neurovascularization inhibiting dual responsive hydrogel for alleviating the progression of osteoarthritis. Nat. Commun. 16, 1390 (2025).
Qian, Y. et al. M2 macrophage-derived exosomal miR-26b-5p regulates macrophage polarization and chondrocyte hypertrophy by targeting TLR3 and COL10A1 to alleviate osteoarthritis. J. Nanobiotechnol. 22, 72 (2024).
Zhang, C. et al. IHH-GLI-1-HIF-2α signalling influences hypertrophic chondrocytes to exacerbate osteoarthritis progression. J. Orthop. Transl. 49, 207–217 (2024).
Zhu, Q. et al. Single-cell RNA sequencing analysis of the temporomandibular joint condyle in 3 and 4-month-old human embryos. Cell Biosci 13, 130 (2023).
Zhang, D. et al. Single-cell RNA sequencing reveals neurovascular-osteochondral network crosstalk during temporomandibular joint osteoarthritis: Pilot study in a human condylar cartilage. Heliyon 9, e20749 (2023).
Bibb, C. A., Pullinger, A. G. & Baldioceda, F. Serial variation in histological character of articular soft tissue in young human adult temporomandibular joint condyles. Arch. Oral Biol. 38, 343–352 (1993).
Orthodontic Treatment of Class III Malocclusion. (Bentham Science Publishers Ltd, Sharjah, United Arab Emirates, 2014).
Singh, M. & Detamore, M. S. Biomechanical properties of the mandibular condylar cartilage and their relevance to the TMJ disc. J. Biomech. 42, 405–417 (2009).
Wang, L., Lazebnik, M. & Detamore, M. S. Hyaline cartilage cells outperform mandibular condylar cartilage cells in a TMJ fibrocartilage tissue engineering application. Osteoarthritis Cartilage 17, 346–353 (2009).
Van Bellinghen, X. et al. Temporomandibular joint regenerative medicine. Int. J. Mol. Sci. 19, 446 (2018).
Gu, Y. et al. Single-cell RNA sequencing in osteoarthritis. Cell Prolif 56, e13517 (2023).
Bi, R. et al. Divergent chondro/osteogenic transduction laws of fibrocartilage stem cell drive temporomandibular joint osteoarthritis in growing mice. Int. J. Oral Sci. 15, 36 (2023).
Embree, M. C. et al. Exploiting endogenous fibrocartilage stem cells to regenerate cartilage and repair joint injury. Nat. Commun. 7, 13073 (2016).
Nathan, J. et al. Fibrocartilage stem cells engraft and self-organize into vascularized bone. J. Dent. Res. 97, 329–337 (2018).
Bi, R. et al. Identification of human temporomandibular joint fibrocartilage stem cells with distinct chondrogenic capacity. Osteoarthritis Cartilage 28, 842–852 (2020).
Sebastian, A. et al. Single-cell RNA-seq reveals transcriptomic heterogeneity and post-traumatic osteoarthritis-associated early molecular changes in mouse articular chondrocytes. Cells 10, 1462 (2021).
Ji, Q. et al. Single-cell RNA-seq analysis reveals the progression of human osteoarthritis. Ann. Rheum. Dis. 78, 100–110 (2019).
Pesesse, L. et al. Bone sialoprotein as a potential key factor implicated in the pathophysiology of osteoarthritis. Osteoarthritis Cartilage 22, 547–556 (2014).
Bellahcène, A. et al. Bone sialoprotein mediates human endothelial cell attachment and migration and promotes angiogenesis. Circ. Res. 86, 885–891 (2000).
Kriegel, A. et al. Bone sialoprotein immobilized in collagen type I enhances angiogenesis in vitro and in ovo. Polymers 15, 1007 (2023).
Babarina, A. V., Möllers, U., Bittner, K., Vischer, P. & Bruckner, P. Role of the subchondral vascular system in endochondral ossification: Endothelial cell-derived proteinases derepress late cartilage differentiation in vitro. Matrix Biol. J. Int. Soc. Matrix Biol. 20, 205–213 (2001).
Mapp, P. I. & Walsh, D. A. Mechanisms and targets of angiogenesis and nerve growth in osteoarthritis. Nat. Rev. Rheumatol. 8, 390–398 (2012).
Hu, Y., Chen, X., Wang, S., Jing, Y. & Su, J. Subchondral bone microenvironment in osteoarthritis and pain. Bone Res. 9, 20 (2021).
Ferrari do Outeiro-Bernstein, M. A. et al. A recombinant NH(2)-terminal heparin-binding domain of the adhesive glycoprotein, thrombospondin-1, promotes endothelial tube formation and cell survival: A possible role for syndecan-4 proteoglycan. Matrix Biol. J. Int. Soc. Matrix Biol. 21, 311–324 (2002).
Dias, J. V. et al. A motif within the N-terminal domain of TSP-1 specifically promotes the proangiogenic activity of endothelial colony-forming cells. Biochem. Pharmacol. 84, 1014–1023 (2012).
Nunes, S. S. et al. Syndecan-4 contributes to endothelial tubulogenesis through interactions with two motifs inside the pro-angiogenic N-terminal domain of thrombospondin-1. J. Cell. Physiol. 214, 828–837 (2008).
Zhang, J. et al. The role of HIF-1α in hypoxic metabolic reprogramming in osteoarthritis. Pharmacol. Res. 213, 107649 (2025).
Yudoh, K., Nakamura, H., Masuko-Hongo, K., Kato, T. & Nishioka, K. Catabolic stress induces expression of hypoxia-inducible factor (HIF)-1α in articular chondrocytes: Involvement of HIF-1α in the pathogenesis of osteoarthritis. Arthritis Res. Ther. 7, R904 (2005).
Feng, Y., Hu, S., Liu, L., Ke, J. & Long, X. HMGB1 contributes to osteoarthritis of temporomandibular joint by inducing synovial angiogenesis. J. Oral Rehabil. 48, 551–559 (2021).
Gao, W. et al. Notch signalling pathways mediate synovial angiogenesis in response to vascular endothelial growth factor and angiopoietin 2. Ann. Rheum. Dis. 72, 1080–1088 (2013).
Su, W. et al. Angiogenesis stimulated by elevated PDGF-BB in subchondral bone contributes to osteoarthritis development. JCI Insight 5, e135446 (2020).
Wang, Y. et al. TNF-α-induced LRG1 promotes angiogenesis and mesenchymal stem cell migration in the subchondral bone during osteoarthritis. Cell Death Dis 8, e2715–e2715 (2017).
Shrivastava, M., Battaglino, R. & Ye, L. A comprehensive review on biomarkers associated with painful temporomandibular disorders. Int. J. Oral Sci. 13, 23 (2021).
Yoon, K. & Gaiano, N. Notch signaling in the mammalian central nervous system: Insights from mouse mutants. Nat. Neurosci. 8, 709–715 (2005).
Wang, Q.-Y. et al. Osteochondral angiogenesis in rat mandibular condyles with osteoarthritis-like changes. Arch. Oral Biol. 57, 620–629 (2012).
Yang, F. et al. Abnormal mechanical stress induced chondrocyte senescence by YAP loss-mediated METTL3 upregulation. Oral Dis 30, 3308–3320 (2024).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinforma. Oxf. Engl. 34, i884–i890 (2018).
van der Sluijs, J. A. et al. The reliability of the mankin score for osteoarthritis. J. Orthop. Res. Off. Publ. Orthop. Res. Soc 10, 58–61 (1992).
Kuyinu, E. L., Narayanan, G., Nair, L. S. & Laurencin, C. T. Animal models of osteoarthritis: Classification, update, and measurement of outcomes. J. Orthop. Surg. 11, 19 (2016).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 81701006, 82271018, 82371002, 32171308) and the Nature Science Foundation of Sichuan Province (Grant No. 2022NSFC0695).
Author information
Authors and Affiliations
Contributions
K.W. performed the experiments and data collection, interpreted the results and drafted the manuscript; Y.L. performed data analysis and visualization, and drafted the manuscript. S.X. performed data visualization and manuscript editing. N.S. and P.W. participated in data analysis and interpretation. X.Y., Q.L. and X.H. contributed to conception and methodology, H.X. contributed to conception and design, methodology, financial support, manuscript reviewing and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, K., Li, Y., Xiang, S. et al. Single-cell transcriptional atlas reveals distinct immune-chondrocyte crosstalk mechanisms in temporomandibular joint osteoarthritis induced by different types of occlusal disorder. Int J Oral Sci 18, 22 (2026). https://doi.org/10.1038/s41368-025-00424-1
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41368-025-00424-1