Abstract
Salivary duct carcinoma (SDCa) is a rare cancer with high rate of metastases and poor survival despite aggressive multimodality treatment. This study analyzes the genetic changes in SDCa, their impact on cancer pathways, and evaluates whether molecular patterns can identify subgroups with distinct clinical characteristics and outcome. Clinicopathologic details and tissue samples from 66 patients (48 males, 18 females) treated between 1995 and 2018 were obtained from multiple institutions. Androgen receptor (AR) was assessed by immunohistochemistry, and the Illumina TruSight 170 gene panel was used for DNA sequencing. Male gender, lympho-vascular invasion, lymph node metastasis, and smoking were significant predictors of disease-free survival. AR was present in 79%. Frequently encountered alterations were mutations in TP53 (51%), PIK3CA (32%) and HRAS (22%), as well as amplifications of CDK4/6 (22%), ERBB2 (21%), MYC (16%), and deletions of CDKN2A (13%). TP53 mutation and MYC amplifications were associated with decreased disease-free survival. Analysis of cancer pathways revealed that the PI3K pathway was most commonly affected. Alterations in the cell cycle pathway were associated with impaired disease-free survival (HR 2.6, P = 0.038). Three subgroups based on AR and ERBB2 status were identified, which featured distinct molecular patterns and outcome. Among AR positive SDCa, HRAS mutations were restricted to AR positive tumors without ERBB2 amplification and HRAS mutations featured high co-occurrence with PIK3CA alterations, which seems specific to SDCa. AR negative SDCa were associated with poor disease-free survival in multivariate analysis (HR 4.5, P = 0.010) and none of these tumors exhibited ERBB2 amplification or HRAS mutations. AR and ERBB2 status in SDCa thus classifies tumors with distinct molecular profiles relevant to future targeted therapy. Furthermore, clinical factors such as smoking and molecular features such as MYC amplification may serve as markers of poor prognosis of SDCa.
Similar content being viewed by others
Introduction
Salivary duct carcinoma (SDCa) is a rare and aggressive primary salivary gland malignancy often presenting with facial nerve deficits and regional or distant metastasis. Men are more often affected and tend to have worse prognosis, but there are no other known predisposing factors [1]. The current multimodality therapy with radical surgical excision, adjuvant radiotherapy, and platinum-based chemotherapy achieves variable local and distant control and mortality remains high [1, 2]. Androgen receptor (AR) and human epidermal growth factor receptor 2 (HER2) expression have been observed in ~75–95% and 25–30% of SDCa, respectively, and small number of patients have been treated with androgen deprivation therapy and HER2-inhibitors. While initial response rates appear good, the development of resistance is common [3,4,5,6,7].
Targeted panel DNA sequencing has been previously performed in modest cohorts of SDCa from single institutions [3, 6, 8, 9]. However, genomic patterns that may unravel underlying risk factors have not been investigated. Furthermore, genetic patterns that may identify subgroups with different prognosis have also not been evaluated. Herein, we evaluate one of the largest multi-institutional cohorts of SDCa using a commercially available targeted gene panel and explore the prognostic utility of the genetic changes in SDCa.
Materials and methods
Samples were collected from pathology departments in Australia (Royal Prince Alfred Hospital, Sydney; ACT Pathology, Canberra; Liverpool Hospital, Liverpool) and Switzerland (Institute of Pathology, University of Bern, Bern). Approval was obtained from the local ethics committees. Clinical data were obtained by chart review. The cancer staging manual of the American Joint Committee on Cancer (8th Edition) was used for staging [10].
Histopathologic evaluation and targeted sequencing
SDCa was defined as per the World Health Organization Classification of Head and Neck Tumors (2017) [11]. All samples were reviewed and reclassified by two specialists in head and neck pathology (RG and MSD). In total, 66 SDCa were deemed suitable for analysis, including 48 males and 18 females.
The formalin fixed paraffin embedded blocks with highest tumor cellularity were selected. Immunohistochemistry for AR was performed as previously described [12]. Internationally validated criteria for interpretation of AR are lacking. Only strong intensity (intensity of 2–3+ as defined for estrogen and progesterone receptors in breast carcinoma) [13] in the majority of the nuclei (50% or more) was considered as positive [14, 15].
DNA was extracted using DNeasy (Qiagen, Venlo, Netherlands) and quantified by Qubit Fluorometric Quantitation (Thermo Fisher, Waltham, MA, USA). Library preparation and targeted capture were performed with TruSight Tumor 170 Kit (Illumina, San Diego, USA) according to the manufacturer’s protocol. Nine to 16 pooled samples were sequenced on NextSeq 500 (Illumina) using 2 × 150bp high-output flow cells. One sample failed DNA quality control, and two samples failed sequencing data quality control. Thus 63 cases were included for genomic analysis (47 males, 16 females). Median sequencing depth was 795× (interquartile range (IQR) 319–1,117).
Data processing
Paired-end short reads were aligned to the hs37d5 reference genome using BWA-MEM (v0.7.10-r789) [16]. NovoSort with default settings (Novocraft Technologies, Petaling Jaya, Malaysia) was used to flag duplicates and merge data from different sequencing lanes. Reads were realigned around insertions/deletions (indels) by GATK IndelRealigner (v3.3) [17]. Single nucleotide variants (SNVs) and indels were identified with VarDictJava (v1.4.6) [18]. Copy number variants (CNVs) were identified with CNVkit (v0.9.1) [19]. To minimize the effect of GC bias, Picard GcBiasSummaryMetrics (http://broadinstitute.github.io/picard/) was used to quantify the GC profile of each sample, and a pool of FFPE controls with similar GC profiles (as measured by the fourth quintile, GC_NC_60_79) were used as a reference. The number of reference controls pooled for each sample ranged from 15 to 42. Tumor purity was estimated using PureCN (v1.10.0) [20] in R (v3.6) [21]. Variant annotation was performed using COSMIC [22], ClinVar [23], dbSNP (v150) [24], 1000 Genomes [25], GnomAD [26] and CADD [27], using vcfanno [28] and Ensembl Variant Effect Predictor (v87) [29]. All analyses were performed on DNAnexus genomic analysis platform (www.dnanexus.com), using our in-house analysis pipeline, refynr2. Somatic variants were distinguished from germline variants using PureCN [20] which uses a statistical model based on the variant’s purity- and ploidy-adjusted variant allele frequency (VAF).
SNV, insertions/deletions and CNV were integrated following filtering of common sequencing artifacts. SNV were reported when deemed pathogenic or likely pathogenic in ClinVar [23], affected hotspots within COSMIC [22] ( ≥ 8 independent reports), or had CADD scores [27] of >15 and a VAF of ≥5%. All mutations were individually reviewed in the Integrative Genome Viewer (IGV) [30] using IGVNav [31]. Amplifications with a ploidy >5.5 were reported. To avoid false positives, long homozygous deletions were considered valid only when ploidy was ≤0.5 and the encompassed segment was <10 Mb long. Deletions in genes on the X chromosome and genes with <25 probes were excluded unless confirmed by manual review. CNVs were only called if they had a likely carcinogenic effect (amplifications for oncogenes, deletions for tumor suppressors). Tumor mutation burden (TMB) of ten samples with mean coverage <250X was not included in further analyses due to high false positive rate as per the manufacturer’s specifications. Missense mutations of TP53 were deemed pathogenic as per definition by the International Agency of Research on Cancer [32]. Loss of function mutations were defined as frameshift, splice site, or nonsense mutations.
Data from The Cancer Genome Atlas (TCGA) PanCancer cohort (n = 10,953) were analyzed using cBioPortal to facilitate comparison with other more common human malignancies [33]. Pathway analysis was performed using the Kyoto Encyclopedia of Genes and Genomes database [34], and curated according to publications based on TCGA data [35, 36]. Interplay of AR with the cell cycle pathway was established from known associations in prostate cancer [37]. To visualize results, R packages ComplexHeatmap [38] and Maftools [39] were implemented in RStudio (v1.1.456, R v3.5.1) [40].
Statistical analysis
Disease-free survival (DFS) was calculated from the date of surgery to the date of last follow-up, recurrence or death from any cause. Disease-specific survival (DSS) was calculated from the date of surgery to the date of death from SDCa. Five patients died from unrelated causes and were censored at the time of death. DFS and DSS were analyzed using univariable and multivariable Cox proportional hazards models. Survival curves were generated using the Kaplan–Meier method. Statistical analysis was performed using Stata version 12.0 SE (StataCorp LLC, 2011). All statistics were two-sided and p values < 0.05 were considered statistically significant.
Results
Clinical and pathological data
Demographic and clinical data are summarized in Table 1. Age did not differ significantly between female and male patients (mean 59.9 vs. 64.8 years, p = 0.16).
Histologically, the tumors typically resembled high-grade ductal carcinoma of the breast and were composed of expanded ducts with cribriform structures forming roman bridges and arches. Areas of central comedo necrosis were present (Fig. 1a). In all instances, the tumor cells showed apocrine morphology with cuboidal cells showing abundant eosinophilic cytoplasm and relatively round nuclei with distinctive nucleoli (Fig. 1b). Areas with apocrine snouts and decapitation secretions were also present (Fig. 1c). Occasional cases showed areas of intraductal carcinoma in the adjacent salivary gland parenchyma (Fig. 1d). Variant morphologic patterns such as micropapillary architecture (Fig. 1e) were also seen. Lympho-vascular invasion (LVI) and perineural invasion were seen in 67% and 59% of tumors, respectively. Lymph node metastasis (LNM) and distant metastases were present in 41 (62%) and 4 (6%) patients, respectively, at presentation. SDCa occurred as the malignant component of carcinoma ex pleomorphic adenoma in 7 (11%) cases. Some of these patients described a lump of nearly 20 years with sudden rapid enlargement. A sclerotic nodule or residual pleomorphic adenoma (Fig. 1f) was present in these cases.
a Areas resembling ductal carcinoma in situ with expanded ducts showing cribriform architecture with roman bridges and arches and central areas of necrosis (H&E ×40). b Polygonal cells with abundant eosinophilic cytoplasm and round nuclei with distinctive nucleoli (H&E ×400). c Apocrine appearing cells with apical snouts and decapitation secretion (H&E ×400). d The parotid gland adjacent to the invasive carcinoma showing intraductal carcinoma with peripheral myoepithelial cells and luminal apocrine cells with abundant cytoplasm and round nuclei with extensive nucleoli (H&E ×400). e SDCa with a dominant micropapillary pattern (H&E ×200). f SDCa arising as malignant transformation in pleomorphic adenoma. A relatively well demarcated sclerotic nodule is present. Dystrophic calcification is seen. Scattered tubular and spindle cell areas are seen within the sclerotic nodule with invasive carcinoma at the periphery (H&E ×100). g and h Immunohistochemistry for androgen receptor. Strong (intensity 2–3+) nuclear staining in the tumor cells (DAB ×40 and DAB ×200).
AR protein expression was detected in 79% of tumors and did not significantly differ between genders (p = 0.18; Fig. 1g, h). The morphologic spectrum of AR negative cases is shown in Fig. 2. All tumors were composed of polygonal cells with abundant eosinophilic cytoplasm and round nuclei with distinctive nucleoli in keeping with apocrine/oncocytoid appearance (Fig. 2a). The tumors showed cribriform architecture (Fig. 2b, c) and occasional nests showed comedo necrosis (Fig. 2b). Intraductal component was present in some cases (Fig. 2d). Malignant transformation of pleomorphic adenoma was also seen (Fig. 2e, f).
a Expanded ductular structures lined by polygonal cells showing abundant eosinophilic cytoplasm and round nuclei with distinctive nucleoli. Apical snouts (arrow) are seen (H&E ×400). b and c Cribriform structures lined by oncocytoid cells. Areas of cribriform necrosis are present (H&E ×100 and ×200). d Intraductal component in the adjacent parotid gland parenchyma (H&E ×40). e Carcinoma arising on a background of pleomorphic adenoma. Sclerotic nodule with residual pleomorphic adenoma with infiltrative nests at the periphery (H&E ×40). f The malignant component shows apocrine morphology in the same tumor shown in e) (H&E ×100).
No patients were treated with androgen deprivation therapy. Two patients have received trastuzumab in adjuvant setting. One of these patients developed lung metastasis at 3 years of follow-up and the other remains disease free at 4 years of follow-up. However, these numbers are too limited for further analyses.
Somatic mutations
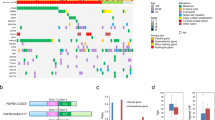
TMB was 15.8 mutations/Mb (IQR 6.8–24.8). Mutations were predominantly missense mutations (73%), and C→T transitions were most common (58%; Fig. 3a and b). The most frequently mutated gene was TP53 (51%; Fig. 3c). Of 35 mutations in 32 patients, 19 implied loss of function, while 16 were pathogenic missense mutations. There was a trend for higher rates of TP53 mutations amongst smokers (57% vs. 32%, p = 0.11), particularly loss of function mutations (43% vs. 11%, p = 0.019).
a Frequency and types of somatic mutations. b Fraction and absolute number of the six possible single nucleotide variants across the cohort. c Alterations in genes that were affected in at least 5% of patients. The bar chart on top shows the number and type of somatic mutation in each sample. Relevant clinical and histo-pathological data are shown for each patient. Genes are ordered according to their associated pathway and the frequency of events. Pathway names are shown on the left (RTK, receptor tyrosine kinase; Ras, Ras-Raf-MEK-ERK pathway).
PIK3CA mutations were detected in 20 (32%) and amplifications in 2 (3%) samples. Mutations concentrated in known hotspots around p.E545, p.E542, and p.H1047 (Fig. 4a). Mutant HRAS was present in 14 (22%) and amplified in 2 (3%) samples (Fig. 3c). Missense mutations occurred at two hotspots (G13 and Q61; Fig. 4a). Four samples also had concurrent splice site alterations or frameshift deletions. PIK3CA alterations co-occurred in 13 of 14 HRAS mutant samples (93%), compared with 14% in HRAS wild type tumors (p < 0.001; Fig. 4b, c). HRAS mutant SDCa featured fewer TP53 mutations than HRAS wild type samples (14% vs. 61%, p = 0.002; Fig. 4b, c). None of the seven SDCa arising from carcinoma ex pleomorphic adenoma featured HRAS mutations (p = 0.33; Fig. 3c).
a Pattern of co-occurring mutations in HRAS and PIK3CA. The colored dots mark the location and frequency of each mutation. Lines link concurrent mutations. Numbers in circles show the number of occurrences of each mutation pair. b Co-occurrence of mutations of the 25 most frequently mutated genes throughout the cohort. Significance levels are calculated by pairwise Fisher’s exact test. c Detailed pattern of oncogenic events (mutations and copy number variations) of TP53, PIK3CA, HRAS, and ERBB2.
Nine samples (14%) featured somatic mutations in several BRCA associated genes (BRCA2, 2 mutations, 1 deletion; BAP1, 4 mutations; BARD1, 2 mutations; BRIP1, 2 mutations; RAD51, 2 mutations). None of the HRAS mutant samples showed somatic mutations in BRCA associated genes.
Copy number variations
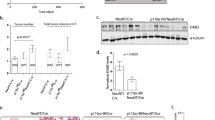
The most frequent CNVs were amplifications of ERBB2 (21%), MYC (16%), CDK4 (17%), CREBBP (14%), PIK3R1 (14%), as well as deletions in CDKN2A (13%) and RB1 (11%) (Fig. 3c). Of the 13 ERBB2 amplified samples, five (38%) had concomitant amplifications of CDK4 (p = 0.04), and eleven samples (85%) showed co-occurring TP53 mutations, while the rate of TP53 mutations was only 42% in samples without ERBB2 amplification (p = 0.011). None of the ERBB2 amplified samples had HRAS mutations (Fig. 4c).
Comparison with the TCGA PanCancer cohort
Assessment of the prevalence of HRAS mutations in the TCGA PanCancer cohort showed higher rates of HRAS mutations in SDCa than in any other human malignancy [11]. Moreover, co-occurrence of PIK3CA mutations was present in only 23% of other HRAS mutant tumors in the TCGA PanCancer cohort compared with 93% of SDCa (Figure S1 in the supplement) [33]. The rate of ERBB2 amplification in SDCa (21%) exceeded that of all other cancer types. The high rate of TP53 mutations in ERBB2 amplified SDCa (85%) was reproduced across the TCGA PanCancer cohort (62% vs. 34% in ERBB2- tumors, p < 0.001; Figure S2) [33]. Compared with the TCGA PanCancer cohort, SDCa also exhibited high rates of MYC amplifications (Figure S3).
Commonly affected pathways
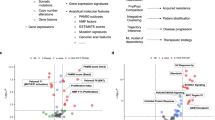
The PI3K pathway was most frequently involved (71% of all samples), followed by the p53 (57%) and cell cycle pathways (51%; Fig. 5). The majority (83%) of patients featured alterations in more than one pathway (Fig. 6a). Significant associations were found between receptor tyrosine kinase (RTK) and cell cycle pathways (p = 0.032), RTK and p53 pathways (p < 0.001), cell cycle and p53 pathways (p = 0.004), and alterations in the Ras pathway featured co-alterations in the PI3K pathway in 92% of the patients (p = 0.005; Fig. 6b).
The frequency of gene alterations in salivary duct carcinomas are summarized into six major pathways: RTK, Ras, Notch, PI3K, cell cycle, and p53. The frequency of alterations consistent with their expected mode of action in each gene is given within each gene’s box (left: somatic mutations; right: copy number variations). The percentage in each pathway represents the rate at which the pathway is affected across the entire cohort. Genes shaded in gray are not part of the sequencing panel. AR status was assessed by means of immunohistochemistry.
a illustrates the number of affected pathways for each patient (numbers above brackets). The number of affected pathways was significantly higher in tumors with lymph node metastasis (LNM; median 3 vs. 2 in without LNM, Mann-Whitney U Test p = 0.036) and lympho-vascular invasion (LVI; median 4 vs. 2 without LVI, p = 0.003). b Interaction map showing co-occurrence of oncogenic events between the different pathways. Colors indicate the rate of co-occurrence of the different pathways when the pathway in the left column is affected (green, high; pink, low). Significant associations are marked (Fisher’s exact test). c Frequency of concurrent pathway alterations for AR and ERBB2 positive tumors, AR positive and ERBB2 negative tumors, as well as AR and ERBB2 negative tumors. d Frequency of alterations in selected genes in three groups defined in C). Significant associations in c) and d) are marked (χ2-test).
Molecular patterns associated with AR and ERBB2 status
Of the 63 samples, 13 (21%) were AR positive and featured ERBB2 amplification (AR + /ERBB2+), 34 (54%) were AR + /ERBB2− and 12 (19%) were AR-/ERBB2−, while none were AR-/ERBB+. Three of seven SDCa arising from pleomorphic adenoma were AR + /ERBB+, two were AR + /ERBB2−, and two were AR-/ERBB2− (p = 0.224). AR status could not be determined in four cases (6%) due to lack of tissue. TMB was not significantly different in AR- tumors compared with AR + tumors (median 22.5 vs 15.8, p = 0.728).
AR + /ERBB2 + tumors were significantly associated with TP53 mutations (85% vs. 38% in AR + /ERBB2− vs. 50% in AR-/ERBB2−, p = 0.017) and alterations in the p53 pathway (100% vs. 41% vs. 58%, p = 0.001; Fig. 6c, d). There were no HRAS mutations in the AR + /ERBB2 + group, while one sample showed HRAS amplification.
AR + /ERBB2− tumors featured high rates of PIK3CA (50%, p = 0.005) and HRAS (41%, p = 0.009) mutations. In fact, all HRAS mutations clustered in this group. None of AR-/ERBB2- SDCa had HRAS or PIK3CA mutations (p = 0.009 and p = 0.005, respectively), although the PI3K pathway was frequently affected (Fig. 6c, d).
The cell cycle pathway was commonly affected in AR + /ERBB+ and AR-/ERBB2− SDCa, while significantly less in AR + /ERBB2− tumors (69%, 75%, and 35%, respectively, p = 0.02).
Survival analysis
Table 2 summarizes the associations of clinicopathological variables and frequent genomic alterations with DFS in univariate analysis. Significant predictors of impaired DFS on univariable analysis were male gender (p = 0.003), smoking history (p = 0.014), LVI (p = 0.024), and LNM (p = 0.013). The effect of smoking was dose dependent with heavy smokers (≥50 pack years) having a 720% increased risk of death or recurrence compared with non-smokers (p < 0.001; Fig. 7a). This association in heavy smokers remained significant for DSS (HR 6.0, 95%CI 1.52–24.11, p = 0.011). In multivariable analysis, both heavy smoking (HR 5.0, 95%CI 1.59–15.82, p = 0.006) and male gender (HR 4.1, 95%CI 1.12–15.18, p = 0.033) were significant predictors of DFS, independent of age (Fig. 7a, b).
a DFS of patients according to their smoking history. Moderate smokers had consumption of 1–50 pack years. Heavy smokers had consumption of >50 pack years. b DFS based on gender c) DFS according to TP53 mutations status. Loss of function mutations is defined as frameshift, splice site, or nonsense mutations. d Association of MYC amplification and DFS. e Association pathogenic alterations (mutations or copy number variations) in genes of the cell cycle pathway (CCND1, CCND2, CCND3, CCNE1, CDK4, CDK6, CDKN2A, RB1), and DFS. f Association of DFS with subgroups of SDCa based on ERBB2 and AR status, adjusted for effects of gender and the cell cycle pathway.
AR status was not a significant predictor of DFS in univariable analysis, however after adjusting for the effect of gender, age and LNM, AR positivity was associated with an 84% improvement in DFS (HR 0.16, 95%CI 0.05–0.53, p = 0.003). Gender also remained significant (HR 5.15, 95% CI 1.63–16.22, p = 0.005). There was weak evidence for an interaction between AR status and gender (p = 0.09). Males with a negative AR status had 22.7 times the risk of recurrence or metastases compared with a female with negative AR status (p = 0.005).
Of the frequently mutated genes, TP53 mutation was associated with a significantly impaired DFS (p = 0.039), but only loss of function remained significant when separated from missense mutations (p = 0.017; Table 2; Fig. 7c). Patients with MYC amplification had poor DFS compared with patients without (p = 0.049; Fig. 7d). Mutations of PIK3CA (p = 0.41), HRAS (p = 0.69), and amplifications of ERBB2 (p = 0.67) were not associated with DFS. On a pathway level, aberrations in the cell cycle pathway were associated with reduced DFS (HR 2.8, 95%CI 1.36–5.75, p = 0.005; Fig. 7e).
None of the subgroups associated with AR/ERBB2 status showed any association with DFS. However, after adjusting for the effect of gender and involvement of the cell cycle pathway, AR-/ERBB2- SDCa had a 357% increased risk of recurrence or death (HR 4.6, 95%CI 1.45–14.43, p = 0.010) and a 239% increased risk of death due to SDCa (HR 3.4, 95% CI 1.00–11.49, p = 0.049) compared with AR + /ERBB2 + SDCa (Table S1, Fig. 7f).
Discussion
This multi-institutional study revealed subgroups of SDCa, including one subgroup (AR + /ERBB2-) featuring co-occurring HRAS and PIK3CA mutations. Whilst PI3K and p53 are the most commonly involved pathways, the cell cycle pathway appears to play an important prognostic role, especially in AR negative SDCa. This study also highlights the prognostic role of gender, LNM, LVI, and smoking.
Similar to other published series [3, 4, 9, 41, 42], our cohort demonstrates male predominance, high prevalence of smoking, and a predilection for the parotid gland. While there is some debate in the literature regarding poor prognosis in men [1, 4], our data show that males have impaired DFS. This is significantly worse if tumors lack AR expression.
The prevalence of smoking in our cohort was high (55%) compared with the general population (14–35% between 1980 and 2016 in Australia [43], 26–33% between 2001 and 2016 in Switzerland [44]). Association between smoking and SDCa has not been previously explored and most genomic SDCa studies do not report smoking status, but prevalence was high in Dalin et al. [3] (70%) and Morris et al. [8] (64%). Our data suggest that smoking is associated with impaired survival in a dose dependent manner (Fig. 7a), which may be attributable to the higher rate of TP53 loss of function mutations in smokers (43% vs. 11%, p = 0.019). These results also suggest a role of smoking in SDCa carcinogenesis, since TP53 mutations are associated with smoking [45].
MYC amplification was significantly associated with DFS in this cohort. Amplifications of MYC are associated with poor prognosis and resistance to chemotherapy in several malignancies [46], and its inhibition promises a therapeutic benefit. However, targeting MYC is difficult because of the protein’s structure, and direct MYC inhibition has not been achieved, although there have been recent advances [47].
AR negative SDCa was associated with impaired DFS in males only. Boon et al. [4] reported no association with outcome, but did not account for gender when assessing AR. Recent studies have demonstrated the benefit of AR blockade in SDCa in adjuvant and palliative settings [48, 49], but no patients in our historical cohort have received this treatment. Similarly, combined AR and ERBB2 status only had prognostic significance after adjusting for gender and cell cycle pathway involvement.
Subgroups of SDCa based on AR and ERBB2 status
AR + /ERBB2 + SDCa is amenable to androgen blockade and HER2 inhibition, both of which have demonstrated some efficacy in SDCa [48,49,50,51]. Apart from a remarkably high rate of co-alterations in the p53 pathway and TP53 mutations, AR + /ERBB2 + tumors exhibited frequent alterations in the cell cycle pathway (69%, P = 0.02; Fig. 4d, e) which are known to induce resistance to AR blockade and HER2-inhibitors [37, 52]. Furthermore, 31% of AR + /ERBB2 + SDCa demonstrated ERBB3 amplification and 38% demonstrated PIK3CA mutations. Both ERBB3 amplification and PIK3CA mutations have been described to confer resistance to HER2-inhibitors [52]. Therefore, AR and HER2 status alone are insufficient to predict the response to androgen deprivation and HER2-inhibitors, and targeted panel testing of genes such as ERBB3 and PIK3CA may help to identify patients with a high risk of resistance who would benefit from treatments downstream of the signaling cascades, such as CDK4/6-inhibitors, potentially in combination.
AR + /ERBB2− SDCa is characterized by frequent PI3K pathway alterations and HRAS mutations, while TP53 mutations and cell cycle pathway alterations were less common than in the other groups (Fig. 6d, e). All HRAS mutated SDCa were AR + /ERBB2-, and they featured a high rate of concurrent PIK3CA alterations (93%; Fig. 4b, c). Other studies also demonstrate co-occurrence of HRAS and PIK3CA mutations in SDCa [3, 41, 53,54,55], while this co-occurrence is present in only 23% of HRAS mutated tumors in the TCGA PanCancer cohort (Figure S1) [33, 56]. Co-occurring HRAS and PIK3CA alterations were thus specific to AR + /ERBB2- SDCa. However, Gargano et al. recently described this co-mutation in HER2 positive SDCa as well [55]. Unlike our study, they used HER2 immunohistochemistry and not ERBB2 amplification as the determinant of HER2/ERBB2 status, and the two methods can occasionally be discordant [57]. In the study of Dalin et al. [3], co-occurring PIK3CA mutations were limited to HRAS p.G13R mutations. Similarly, HRAS p.G13R mutations were associated with PIK3CA mutations around H1047 in our study, but we also observed co-occurring HRAS p.Q61R and PIK3CA p.E542 mutations (Fig. 4a). Interestingly, and in line with other studies [53, 55], HRAS mutations seem to be exclusive to SDCa arising de novo, as none of the SDCa arising from carcinoma ex pleomorphic adenoma were affected by HRAS mutations. The structure of HRAS makes it a difficult target, and efforts to develop drugs directed directly at HRAS have been unsuccessful. The inhibition of proteins downstream in the Ras-Raf-MEK-ERK pathway has therefore received growing attention, and may offer a therapeutic approach for HRAS-mutant SDCa in the future [58].
All AR negative SDCa were also ERBB2 negative, which has also been described in other studies [3, 59]. Interestingly, although alterations in the PI3K pathway were common, PIK3CA mutations did not occur in this subgroup (Fig. 6c, d). Lack of AR expression always raises concerns that this group may not represent SDCa despite the presence of apocrine/polygonal/oncocytoid morphology and PI3K pathway alterations. The possibility that some of these tumors may represent adenocarcinoma, not otherwise specified, cannot be entirely excluded. Other differential diagnoses such as high-grade transformation in a pre-existing salivary gland neoplasm with obliteration of the low-grade component also needs to be considered. Indeed, one case has demonstrated NR4A3 rearrangement, a change recently described in acinic cell carcinoma [60]. It is important to note that AR-/ERBB2- SDCa had the worst prognosis and the fact that they are not amenable to androgen blockade and HER2-inhibition underlines that these are oncologically challenging tumors. These tumors also had the highest rates of alterations in the cell cycle pathway (75%; Fig. 6c), which was the only pathway significantly associated with impaired DFS (Fig. 7e). These findings highlight the need to recognize high-grade primary salivary gland carcinoma with apocrine/polygonal/oncocytoid morphology or carcinoma ex pleomorphic adenoma that demonstrate AR-/ERBB2- profile as a distinct molecular entity with a prognosis more dismal than AR + SDCa and with limited therapeutic options.
Most molecular hallmark alterations in SDCa characterized in this study (AR, ERBB2, HRAS, MYC and PI3K pathway involvement) impact cell cycle progression via direct or indirect upregulation of CCND1–3 and CDK4/6, even when the genes within the cell cycle pathway are not altered (Fig. 5). Therefore, although AR-/ERBB2- and AR + /ERBB2 + SDCa more commonly featured alterations in the cell cycle pathway than AR + /ERBB2- SDCa (Fig. 6c), this pathway may offer a treatment option for all three subgroups of SDCa and warrants further research.
In conclusion, this study describes several clinical and molecular features that may serve as markers of poor prognosis of SDCa. Smoking, which has not previously been associated with SDCa, seems to negatively impact outcome. Importantly, three subgroups of SDCa based on AR and ERBB2 status were identified, which exhibit distinct molecular profiles relevant to future targeted therapy.
References
Osborn V, Givi B, Lee A, Sheth N, Roden D, Schwartz D, et al. Characterization, treatment and outcomes of salivary ductal carcinoma using the National Cancer Database. Oral Oncol. 2017;71:41–46.
Guzzo M, Locati LD, Prott FJ, Gatta G, McGurk M, Licitra L, et al. Major and minor salivary gland tumors. Crit Rev Oncol Hematol. 2010;74:134–48.
Dalin MG, Desrichard A, Katabi N, Makarov V, Walsh LA, Lee K-W, et al. Comprehensive molecular characterization of salivary duct carcinoma reveals actionable targets and similarity to apocrine breast cancer. Clin Cancer Res. 2016;22:4623–33.
Boon E, Bel M, van Boxtel W, van der Graaf WTA, van Es RJJ, Eerenstein SEJ, et al. A clinicopathological study and prognostic factor analysis of 177 salivary duct carcinoma patients from The Netherlands. Int J Cancer. 2018;143:758–66.
Soper MS, Iganej S, Thompson LDR. Definitive treatment of androgen receptor-positive salivary duct carcinoma with androgen deprivation therapy and external beam radiotherapy. Head Neck. 2014;36:E4–E7.
Khoo TK, Yu B, Smith JA, Clarke AJ, Luk PP, Selinger CI, et al. Somatic mutations in salivary duct carcinoma and potential therapeutic targets. Oncotarget. 2017;8:75893–903.
Park JC, Ma TM, Rooper L, Hembrough T, Foss RD, Schmitt NC, et al. Exceptional responses to pertuzumab, trastuzumab, and docetaxel in human epidermal growth factor receptor-2 high expressing salivary duct carcinomas. Head Neck. 2018;40:E100–E106.
Morris LGT, Chandramohan R, West L, Zehir A, Chakravarty D, Pfister DG, et al. The molecular landscape of recurrent and metastatic head and neck cancers. JAMA Oncol. 2017;3:244–55.
Ross JS, Gay LM, Wang K, Vergilio JA, Suh J, Ramkissoon S, et al. Comprehensive genomic profiles of metastatic and relapsed salivary gland carcinomas are associated with tumor type and reveal new routes to targeted therapies. Ann Oncol. 2017;28:2539–46.
Amin MB, Edge SB, American Joint Committee on Cancer. AJCC Cancer Staging Manual, Eight Edition. New York: Springer International Publishing; 2017.
El-Nagger AK, Chan JK, Grandis JR, Takashi T, Slootweg PJ WHO Classification of Head and Neck Tumours. 4th ed. Lyon: International Agency for Research on Cancer, 2017.
Luk PP, Weston JD, Yu B, Selinger CI, Ekmejian R, Eviston TJ, et al. Salivary duct carcinoma: Clinicopathologic features, morphologic spectrum, and somatic mutations. Head Neck. 2016;38 Suppl 1:E1838–47.
Allison KH, Hammond MEH, Dowsett M, McKernin SE, Carey LA, Fitzgibbons PL, et al. Estrogen and progesterone receptor testing in breast cancer: ASCO/CAP guideline update. J Clin Oncol. 2020;38:1346–66.
Williams L, Thompson LDR, Seethala RR, Weinreb I, Assaad AM, Tuluc M, et al. Salivary duct carcinoma: the predominance of apocrine morphology, prevalence of histologic variants, and androgen receptor expression. Am J Surg Pathol. 2015;39:705–13.
Hammond MEH, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, et al. American society of clinical oncology/college of american pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010;28:2784–95.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Lai Z, Markovets A, Ahdesmaki M, Chapman B, Hofmann O, McEwen R, et al. VarDict: a novel and versatile variant caller for next-generation sequencing in cancer research. Nucleic Acids Res. 2016;44:e108–e108.
Talevich E, Shain AH, Botton T, Bastian BC. CNVkit: Genome-Wide Copy Number Detection and Visualization from Targeted DNA Sequencing. PLOS Comput Biol. 2016;12:e1004873.
Riester M, Singh AP, Brannon AR, Yu K, Campbell CD, Chiang DY, et al. PureCN: copy number calling and SNV classification using targeted short read sequencing. Source Code Biol Med. 2016;11:13.
R Development Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria.
Forbes SA, Beare D, Boutselakis H, Bamford S, Bindal N, Tate J, et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 2017;45:D777–D783.
Landrum MJ, Lee JM, Benson M, Brown G, Chao C, Chitipiralla S, et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2016;44:D862–8.
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 2001;29:308–11.
Consortium T. 1000 GP. A global reference for human genetic variation. Nature. 2015;526:68–74.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. Variation across 141,456 human exomes and genomes reveals the spectrum of loss-of-function intolerance across human protein-coding genes. bioRxiv. 2019:531210. https://doi.org/10.1101/531210.
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res 2019;47:D886–D894.
Pedersen BS, Layer RM, Quinlan AR. Vcfanno: fast, flexible annotation of genetic variants. Genome Biol. 2016;17:118.
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GRS, Thormann A, et al. The ensembl variant effect predictor. Genome Biol. 2016;17:122.
Robinson JT, Thorvaldsdóttir H, Wenger AM, Zehir A, Mesirov JP. Variant review with the integrative genomics viewer. Cancer Res. 2017;77:e31–e34.
Barnell EK, Ronning P, Campbell KM, Krysiak K, Ainscough BJ, Sheta LM, et al. Standard operating procedure for somatic variant refinement of sequencing data with paired tumor and normal samples. Genet Med. 2019;21:972–81.
Bouaoun L, Sonkin D, Ardin M, Hollstein M, Byrnes G, Zavadil J, et al. TP53 variations in human cancers: new lessons from the IARC TP53 database and genomics data. Hum Mutat. 2016;37:865–76.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Kanehisa M, Sato Y, Furumichi M, Morishima K, Tanabe M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 2019;47:D590–5.
Sanchez-Vega F, Mina M, Armenia J, Chatila WK, Luna A, La KC. et al. Oncogenic signaling pathways in The Cancer Genome Atlas. Cell. 2018;173:321–37.e10.
Zhang Y, Ng PK, Kucherlapati M, Chen F, Liu Y, Tsang YH, et al. A Pan-Cancer Proteogenomic Atlas of PI3K/AKT/mTOR Pathway Alterations. Cancer Cell. 2017;31:820–32.e3.
Balk SP, Knudsen KE. AR, the cell cycle, and prostate cancer. Nucl Recept Signal. 2008;6:e001.
Gu Z, Eils R, Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–9.
Mayakonda A, Koeffler HP Maftools: Efficient analysis, visualization and summarization of MAF files from large-scale cohort based cancer studies. bioRxiv. 2016:052662. https://doi.org/10.1101/052662.
RStudio Team. RStudio: Integrated development for R. RStudio Inc, Boston, MA. 2016.
Ku BM, Jung HA, Sun J-M, Ko YH, Jeong H-S, Son Y-I, et al. High-throughput profiling identifies clinically actionable mutations in salivary duct carcinoma. J Transl Med. 2014;12:299.
Sawabe M, Ito H, Takahara T, Oze I, Kawakita D, Yatabe Y, et al. Heterogeneous impact of smoking on major salivary gland cancer according to histopathological subtype: a case-control study. Cancer 2018;124:118–24.
Greenhalgh EM, Bayly M, Winstanley MH 1.3 Prevalence of smoking—adults. In Scollo, MM and Winstanley, MH, editors. Tobacco in Australia. Melbourne: Cancer Council Victoria.
Addiction Monitoring Switzerland. Tabak » Prävalenz. Bern: Federal Office of Public Health, 2019 [cited 26 Movember 2019]. Available from https://www.suchtmonitoring.ch/de/1/1-2.html?tabak-pravalenz-pravalenz-des-tabakkonsums.
Halvorsen AR, Silwal-Pandit L, Meza-Zepeda LA, Vodak D, Vu P, Sagerup C, et al. TP53 mutation spectrum in smokers and never smoking lung cancer patients. Front Genet. 2016;7:85.
Kalkat M, De Melo J, Hickman KA, Lourenco C, Redel C, Resetca D, et al. MYC deregulation in primary human cancers. Genes (Basel). 2017;8:151.
Chen H, Liu H, Qing G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal Transduct Target Ther. 2018;3:5.
van Boxtel W, van Herpen C. Improving survival in salivary duct cancer with adjuvant androgen deprivation therapy. Oncotarget. 2019;10:3833–4.
Boon E, van Boxtel W, Buter J, Baatenburg de Jong RJ, van Es RJJ, Bel M, et al. Androgen deprivation therapy for androgen receptor-positive advanced salivary duct carcinoma: a nationwide case series of 35 patients in The Netherlands. Head Neck. 2018;40:605–13.
Takahashi H, Tada Y, Saotome T, Akazawa K, Ojiri H, Fushimi C, et al. Phase II trial of trastuzumab and docetaxel in patients with human epidermal growth factor receptor 2–positive salivary duct carcinoma. J Clin Oncol. 2019;37:125–34.
van Boxtel W, Boon E, Weijs WLJ, van den Hoogen FJA, Flucke UE, van Herpen CML. Combination of docetaxel, trastuzumab and pertuzumab or treatment with trastuzumab-emtansine for metastatic salivary duct carcinoma. Oral Oncol. 2017;72:198–200.
Vernieri C, Milano M, Brambilla M, Mennitto A, Maggi C, Cona MS, et al. Resistance mechanisms to anti-HER2 therapies in HER2-positive breast cancer: current knowledge, new research directions and therapeutic perspectives. Crit Rev Oncol Hematol. 2019;139:53–66.
Wang K, Russell JS, McDermott JD, Elvin JA, Khaira D, Johnson A, et al. Profiling of 149 salivary duct carcinomas, carcinoma ex pleomorphic adenomas, and adenocarcinomas, not otherwise specified reveals actionable genomic alterations. Clin Cancer Res. 2016;22:6061–8.
Chiosea SI, Williams L, Griffith CC, Thompson LDR, Weinreb I, Bauman JE, et al. Molecular characterization of apocrine salivary duct carcinoma. Am J Surg Pathol. 2015;39:744–52.
Gargano SM, Senarathne W, Feldman R, Florento E, Stafford P, Swensen J, et al. Novel therapeutic targets in salivary duct carcinoma uncovered by comprehensive molecular profiling. Cancer Med. 2019;8:7322–9.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Ross DS, Zehir A, Cheng DT, Benayed R, Nafa K, Hechtman JF. et al. Next-generation assessment of human growth factor receptor 2 (ERBB2) amplification status: clinical validation in the context of a hybrid capture-based, comprehensive solid tumor genomic profiling assay. J Mol Diagnostics. 2017;19:244–54.
Vaseva AV, Yohe ME. Targeting RAS in pediatric cancer: is it becoming a reality? Curr Opin Pediatr. 2020;32:48–56.
Di Palma S, Simpson RHW, Marchiò C, Skálová A, Ungari M, Sandison A, et al. Salivary duct carcinomas can be classified into luminal androgen receptor-positive, HER2 and basal-like phenotypes. Histopathology. 2012;61:629–43.
Haller F, Skálová A, Ihrler S, Märkl B, Bieg M, Moskalev EA, et al. Nuclear NR4A3 immunostaining is a specific and sensitive novel marker for acinic cell carcinoma of the salivary glands. Am J Surg Pathol. 2019;43:1264–72.
Acknowledgements
The results of the comparison to other tumor types are based upon data generated by the TCGA Research Network: https://www.cancer.gov/tcga. This work was supported by a Cancer Institute NSW Early Career Fellowship (grant number 13/ECF/1–46) to [MJC], Cancer Institute NSW Career Development Fellowship (grant number 2019/ CDF004) to [MJC], NSW Health Early-Mid Career Fellowship, Cancer Council NSW (grant number RG 15–19) to [MJC], and philanthropic donations from the Kinghorn Foundation as well as the Neal Wald Trust. We would like to thank Mary Abbey, Senior Scientist, for preparing the tissues from Anatomical Pathology, ACT Pathology, The Canberra Hospital, and Trina Lum, NSW Pathology, Royal Prince Alfred Hospital, for immunohistochemistry.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Mueller, S.A., Gauthier, ME.A., Blackburn, J. et al. Molecular patterns in salivary duct carcinoma identify prognostic subgroups. Mod Pathol 33, 1896–1909 (2020). https://doi.org/10.1038/s41379-020-0576-2
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41379-020-0576-2
This article is cited by
-
Potentially actionable molecular alterations in particular related to poor oncologic outcomes in salivary gland carcinomas
BMC Cancer (2025)
-
LH–RH agonist monotherapy for androgen receptor-positive recurrent and/or metastatic salivary gland cancer: a retrospective study
International Journal of Clinical Oncology (2025)
-
Histologic characteristics and predictors of outcome in epithelial-myoepithelial carcinoma (EMC) of salivary gland: a multi-center study of 75 cases
Virchows Archiv (2025)
-
Speicheldrüsenkarzinome – ein aktueller Überblick
Die Pathologie (2023)
-
Speicheldrüsenkarzinome – ein aktueller Überblick
Die Onkologie (2023)