Abstract
Anorexia nervosa (AN) has extensive genetic correlations with other psychiatric disorders, and genetic risk for different psychiatric disorders was associated with distinct clinical courses in AN. Uncovering associations between transdiagnostic psychiatric genetic liability and AN outcomes can facilitate its personalized treatment. In this study, we investigated the associations between transdiagnostic psychiatric genetic liability and outcomes of AN. Genomic structural equation models were fitted to genome-wide association data for AN and psychiatric disorders with high genetic correlations with AN (obsessive-compulsive symptoms [OCS], major depressive disorder [MDD], schizophrenia, and anxiety disorders) to extract one shared and five trait-specific genetic components. Next, we calculated the polygenic risk scores (PRS) for these components, including PRSshared, PRSAN-specific, PRSOCS-specific, PRSMDD-specific, PRSSCZ-specific and PRSANX-specific, which index the shared genetic liability to all five psychiatric traits, and genetic liability specific to AN, OCS, MDD, SCZ and ANX, respectively. We then tested associations between these PRSs and clinical outcomes reported between 1997 and 2018 among AN cases from the Anorexia Nervosa Genetics Initiative (ANGI), linked to Swedish National Registers. The clinical outcomes included cumulative disease burden (i.e., number of diagnoses, medication prescriptions, and inpatient days), risks of psychiatric comorbidities, and AN symptomatology. Among 4028 included AN cases, the mean (SD) birth year was 1985 (9), and 3947 (98.0%) were female. Within AN, +1 SD increase of PRSshared was associated with 9–39% excess risk of disease burden and psychiatric comorbidity, whereas the associations between PRSAN-specific and most clinical outcomes were statistically non-significant. +1 SD increase of PRSMDD-specific was associated with 3–29% increased risk of AN disease burden. Our findings show that shared psychiatric liability is associated with more adverse AN outcomes, whereas AN-specific liability is not a good indicator for its clinical course. This study provides a novel perspective on factors influencing heterogeneity in AN clinical course.
Similar content being viewed by others
Introduction
Anorexia nervosa (AN) is a serious and often chronic psychiatric disorder with significant morbidity and mortality [1]. AN is characterized by very low body weight and an intense fear of gaining weight, with a lifetime prevalence of approximately 1% and a full recovery rate of less than 50% [2, 3]. It is a moderately heritable disorder with a twin-based heritability of 50–70% [4]. The latest genome-wide association study (GWAS) identified 8 genomic loci for AN and showed that 17% of variance in liability to AN is attributable to common genetic variants [5].
AN is highly comorbid with other psychiatric disorders [1]. It commonly co-occurs with obsessive-compulsive disorder (OCD) (14%), major depressive disorder (MDD) (73%) and anxiety disorders (ANX) (75%) and is associated with 6-fold greater risk of having schizophrenia (SCZ) [6,7,8,9]. Moreover, some clinical features of AN, such as perfectionism and obsessive exercise, have obsessive and compulsive characteristics [8]. Similarly, distorted perceptions of body shape in AN resemble delusional thinking in SCZ [10].
Phenotypic co-aggregation between AN and other psychiatric disorders can be partially explained by genetic factors, as indicated by their extensive genetic overlap. Notable genetic correlations (rg) based on common variants have been estimated at AN vs. OCD = 0.45, AN vs. MDD = 0.28, AN vs. SCZ = 0.25, and AN vs. ANX = 0.25 [5]. This strong but incomplete overlap indicates the existence of pleiotropic variants influencing multiple disorders as well as variants specific to each disorder.
The clinical course of AN is heterogeneous, with some cases being relatively brief but nearly a third giving way to a severe and enduring profile [1]. Interestingly, we have previously shown that the variance in AN course can be partially explained by genetic liability to other psychiatric disorders: genetic liability to SCZ was associated with distinct phenotypes in AN [11]. Furthermore, the transdiagnostic polygenic risk score (PRS) for AN and OCD demonstrated good performance in predicting the risk of AN [12].
Genomic structural equation modelling (gSEM) is a method to capture the multivariate genetic architecture of genetically correlated traits based on GWAS summary statistics. gSEM is capable of isolating a pleiotropic component representing broad genetic liability in addition to components representing liability specific to each trait. gSEM has been employed to distinguish shared and disorder-specific liability between BIP, MDD, and SCZ; autism spectrum disorder (ASD) and attention-deficit hyperactivity disorder (ADHD); as well as MDD and ANX [13,14,15,16]. Individual single nucleotide polymorphisms (SNPs) can be integrated into the gSEM model so that effects of a specific SNP on the shared liability component and trait-specific components can be estimated, allowing the construction of more predictive and valid PRSs [16]. These gSEM-derived PRSs indexing shared and non-shared genetic liabilities to psychiatric disorders can be further utilized to characterize the clinical heterogeneity of a specific disorder [13].
Given the consistently demonstrated symptomatic and genetic overlap between AN and other psychiatric disorders, we used gSEM to extract the shared and specific genetic components of AN and four psychiatric traits with high genetic correlations with AN (obsessive compulsive symptoms [OCS], MDD, SCZ and ANX). We calculated PRSs for these components and tested the associations between these PRSs and AN risk as well as a wide range of clinical outcomes to dissect the heterogeneous outcomes of AN.
Methods
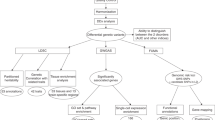
The work flow of the current study is presented in Fig. 1.
AN anorexia nervosa, SCZ schizophrenia, OCS obsessive-compulsive symptoms, OCD obsessive-compulsive disorder, MDD major depressive disorder, ANX anxiety disorders, gSEM genomic structural equation model, PRS polygenic risk scores, ASD autism spectrum disorder, ADHD attention-deficit hyperactivity disorder, SUD substance use disorder, BMI body mass index, CIA Clinical Impairment Assessment, CPRS Comprehensive Psychopathological Rating Scale, EDE-Q Eating Disorder Examination Questionnaire.
Source GWAS
The source GWAS summary statistics for AN (without ANGI-SE samples, Ncases = 4105, Ncontrols = 3793), OCS and SCZ were acquired from the Psychiatric Genomics Consortium (PGC), whereas the GWAS for MDD was derived from a meta-analysis of six European datasets, and the GWAS for ANX was based on the European subset of a multi-ancestry meta-analysis [5, 17,18,19,20]. All individuals in the component data sets were of European ancestry. Detailed information on source GWAS studies is available in Table S1.
Target genotype data
The target genotype data were acquired from the Swedish arm of the Anorexia Nervosa Genetics Initiative (ANGI-SE). ANGI is an international collaboration committed to collecting genotype and phenotype data from individuals with AN and controls to identify genetic and environmental risk factors of AN [21]. AN status was determined by a DSM-IV-based AN diagnosis or responses to the ED100K-V1 questionnaire [21]. Details regarding recruitment of participants in ANGI-SE are described in the Supplementary Methods.
In the current study, we included all ANGI-SE individuals with available linkage to Swedish National Registers [21]. The quality control (QC) and imputation of the genotype data were performed according to the RICOPILI pipeline, leaving a total of 4028 cases and 3846 controls [22]. Details regarding genotyping and QC are provided in the Supplementary Methods.
Clinical phenotypes
The data on clinical phenotypes in AN cases were derived from records in Swedish National registers from January 1, 1997 to December 31, 2018, using unique personal identification numbers to link data across registers [23]. We focused on five categories of phenotypes in the current study: (1) clinical diagnoses: general [somatic + psychiatric], somatic and psychiatric; (2) medication prescriptions: general, antipsychotics and antidepressants; (3) inpatient days: any and due to EDs ; (4) psychiatric comorbidities: OCD, SCZ, MDD, ANX, ASD, ADHD, and substance use disorder (SUD); (5) symptomatology of AN at first visit: BMI, Eating Disorder Examination Questionnaire (EDE-Q) score [24], depression subscale score from the Comprehensive Psychopathological Rating Scale (CPRS) [25], anxiety subscale score from the CPRS [25], and Clinical Impairment Assessment (CIA) questionnaire score [26, 27]. Number of clinical diagnoses, medication prescriptions, and inpatient days were defined as cumulative disease burden in the current study.
Data on diagnoses, inpatient days, medication prescriptions, and psychiatric comorbidities were available for all included AN cases, whereas information on AN symptomatology at first visit was only available for 1934 AN cases. Details about the Swedish National Registers that these phenotypes were derived from are described in the Supplementary Methods and Tables S2–S4.
Statistical analyses
GenomicSEM (gSEM)
We used gSEM to construct a common-factor model based on source GWAS datasets and extracted the loading of each SNP on the common factor. We refer to this as the “shared” component, indicating contribution of the SNP to general liability to the five disorders (Fig. S1A and Supplementary Methods). Next, we constructed five models to extract the loading of each SNP on the residual variance from each source GWAS after accounting for the common factor to represent contribution of the SNP to the liability specific to each disorder (Fig. S1B–F and Supplementary Methods). We refer to these as AN-specific, OCS-specific, SCZ-specific, MDD-specific, and ANX-specific effects. gSEM was performed with the R package “GenomicSEM” (“0.0.5”). Code and technical details are available online (https://github.com/GenomicSEM/GenomicSEM/) and in the Supplementary Methods.
Generation of PRSs
We generated PRSs using PRSice software (version 2.3.5) [28]. The target dataset was the genotype data from individuals in ANGI-SE. The base summary statistic datasets were the shared and trait-specific effects from gSEM. We clumped the SNPs at r2 < 0.1 within 250 kb and aggregated their effects at different p-value thresholds (5e-8, 1e-6, 1e-4, 0.001,0.01, 0.05, 0.1, 0.2, 0.5, 1) using the PRS-PCA method [29]. The final gSEM-derived PRSs utilized in association analyses were the standardized first components derived from principal component analyses (PCA) of PRSs at all thresholds. We performed PCA and standardization with R functions “princomp()” and “scale()”, respectively. Details for PRSs calculation are presented in Supplementary Methods.
Association analyses for gSEM PRSs with AN status and clinical phenotypes
To investigate the association between gSEM-derived PRSs and AN status, we conducted logistic regression analyses with binary AN status as the outcome in the 4028 AN cases and 3846 controls. Each gSEM-derived PRS was tested as an exposure variable separately. Odds ratios (ORs) represented the risk estimates +1 standard deviation (SD) increase of PRS.
To investigate the impact of gSEM-derived PRSs on the cumulative disease burden of AN cases, we conducted quasi-Poisson regression analyses with number of unique diagnoses (general, somatic, and psychiatric), number of unique medication prescriptions (any, antipsychotics, and antidepressants), and number of inpatient days (any and due to eating disorders) recorded from January 1, 1997 to December 31, 2018 as outcomes among 4028 AN cases. Each gSEM-derived PRS was tested as an exposure variable for association with these eight outcomes separately. We included a log-time offset term in each model to adjust for differences in follow-up time between individuals. Incidence rate ratios (IRRs) represented the risk estimates +1 SD increase of gSEM-derived PRS.
To investigate the impact of gSEM-derived PRSs on risks of psychiatric comorbidities in AN cases, we conducted Cox regression (survival) analyses with age as the underlying timescale and onset of OCD, SCZ, MDD, ANX, ASD, ADHD, and SUD as outcomes among 4028 AN cases. Each gSEM-derived PRS was tested as an exposure variable for association with these seven outcomes separately. Individuals were followed from January 1, 1997 until onset of disorder, death or December 31, 2018, whichever came first. Hazard ratios (HRs) represented the risk estimates +1 SD increase of gSEM-derived PRS.
To investigate the impact of gSEM-derived PRSs on the symptomatology of AN at first visit in AN cases, we performed linear regression analyses with BMI, EDE-Q scores, CIA scores, CPRS-depression and CPRS-anxiety scores as outcomes among 4028 AN cases. Each gSEM-derived PRS was tested as an exposure variable for association with these five outcomes separately. Regression coefficients represented change in symptoms +1 SD increase of gSEM-derived PRS.
All association analyses were performed in R version 4.2.3. In all regression models, we adjusted for birth year, sex, and the first 10 ancestry-informative principal components. The significance level in all association analyses above was set at two-sided P < 0.05. To correct for multiple tests, we also employed a Bonferroni-corrected significance level of two-sided P < 0.05/6 = 8.33 × 10−3 for association tests with AN status and two-sided P < 0.05/120 = 4.17 × 10−4 for association tests with clinical outcomes of AN. Details regarding the regression models are presented in the Supplementary Methods and Table S5.
Results
Descriptive characteristics of the study population
Of the 4028 AN cases in ANGI-SE, the mean (SD) birth year was 1985 (9), and 3947 (98.0%) were female, whereas among the 3846 controls, the mean (SD) birth year was 1978 (10), and 3776 (98.2%) were female. The descriptive characteristics of the 4028 individuals with AN are presented in Table S6.
Association between gSEM-derived PRSs and AN status
Only PRSAN-specific and PRSshared were associated with statistically significant increased risk of AN, whereas other gSEM-derived PRSs showed either statistically non-significant or decreased risk of AN after Bonferroni correction (Table 1). In contrast, all unmodified source GWAS PRSs were associated with an elevated risk of AN (Table S7).
Association between gSEM-derived PRSs and cumulative disease burden
PRSshared was associated with a higher risk for cumulative disease burden (Fig. 2A; Table S8): +1 SD increase in PRSshared was associated with receiving 12% (IRR, 1.12 95%CI, 1.09–1.15; P = 1.82 × 10−18) more general diagnoses, 16% (IRR, 1.16; 95%CI, 1.12–1.21; P = 5.46 × 10−16) more psychiatric diagnoses and 9% more somatic diagnoses (IRR, 1.09; 95%CI, 1.06–1.13; P = 7.01 × 10−8) at P < 0.05. For medication prescriptions, +1 SD increase in PRSshared was associated with 12% (IRR, 1.12; 95%CI, 1.09–1.15; P = 1.80 × 10−18) more general prescriptions, 27% (IRR, 1.27; 95%CI. 1.17–1.38; P = 8.06 × 10−9) more antipsychotic prescriptions, and 15% (IRR, 1.15; 95%CI, 1.11–1.20; P = 3.78 × 10−15) more antidepressant prescriptions at P < 0.05. PRSshared was also associated with having more inpatient days due to any illness (IRR, 1.19; 95%CI, 1.08–1.32; P = 5.79 × 10−4) at P < 0.05. After Bonferroni correction, all these associations remained significant except for the association with inpatient days.
Results are derived from quasi-Possion regression models based on 4028 AN cases with number of unique clinical diagnoses (any, psychiatric, somatic), prescriptions (any, antipsychotics, antidepressants) and inpatient days (any, due to EDs) as outcomes. For each of the eight outcomes, we constructed six models with PRSshared (A), PRSAN-specific (B), PRSOCS-specific (C), PRSMDD-specific (D), PRSSCZ-specific (E) and PRSANX-specific (F) as exposure variable, respectively. Incidence rate ratios indicate the risk estimates for +1 SD increase of PRS. Sex, birth year and first 10 ancestry-informative principal components were adjusted for in all models. The points represent incidence rate ratio estimates, and the error bars indicate 95% confidence intervals. Blue points represent effect estimates for clinical diagnoses, red points represent effects for medication prescriptions and yellow points represent effects for inpatient days. “*” represents association that remained significant after Bonferroni correction. “#” represents trending association at P < 0.05 but was not significant after Bonferroni correction. PRS polygenic risk scores, EDs eating disorders, IRR incidence rate ratio, 95%CI 95% confidence interval, AN anorexia nervosa, SCZ schizophrenia, OCS obsessive-compulsive symptoms, MDD major depressive disorder, ANX anxiety disorders.
PRSAN-specific was not statistically significantly associated with most cumulative disease burden outcomes at P < 0.05 (Fig. 2B; Table S8), except for a 17% increased risk of inpatient days due to EDs (IRR, 1.17; 95%CI, 1.04–1.32; P = 9.75 × 10−3). However, it became non-significant after Bonferroni correction. No statistically significant association was detected between PRSOCS-specific and cumulative disease burden at P < 0.05 (Fig. 2C; Table S8).
PRSMDD-specific was associated with generally elevated risk for cumulative disease burden (Fig. 2D; Table S8): +1 SD increase in PRSMDD-specific was associated with receiving 3% more general diagnoses (IRR, 1.03; 95%CI, 1.01–1.06; P = 7.58 × 10−3) and 7% more psychiatric diagnoses (IRR, 1.07; 95%CI, 1.03–1.11; P = 1.87 × 10−4), 6% more general prescriptions (IRR, 1.06; 95%CI, 1.03–1.08; P = 6.85 × 10−6), 14% more antipsychotic prescriptions (IRR, 1.14; 95%CI, 1.06–1.24; P = 5.43 × 10−4) and 9% more antidepressant prescriptions (IRR, 1.09; 95%CI, 1.05–1.13; P = 6.63 × 10−7) as well as 15% more inpatient days (IRR, 1.15; 95%CI, 1.04–1.26; P = 5.31 × 10−3) at P < 0.05. Only the associations with psychiatric diagnoses, any prescriptions and antidepressant prescriptions survived the Bonferroni correction.
PRSSCZ-specific and PRSANX-specific were generally associated with a lower risk for cumulative disease burden, but neither was statistically significant after Bonferroni correction (Fig. 2E, F; Table S8).
Association between gSEM-derived PRSs and risk of psychiatric comorbidities
Increased PRSshared was associated with increased risk of all psychiatric comorbidities at P < 0.05, with effect sizes ranging from +14–+58% excess risk (Fig. 3A; Table S9). Increased PRSMDD-specific was associated with elevated risks for ADHD, ANX, ASD, MDD and SUD at P < 0.05 (Fig. 3D; Table S9). Most associations did not withstand Bonferroni correction, except for the associations for PRSshared with ADHD, ANX, MDD and SUD as well as the one between PRSMDD-specific and ANX (Fig. 3; Table S9).
Results are derived from quasi-Possion regression models based on 4028 AN cases with ADHD, ANX, ASD, MDD, OCD, SCZ and SUD as the outcome variables. For each of the seven outcomes, we constructed six models with PRSshared (A), PRSAN-specific (B), PRSOCS-specific (C), PRSMDD-specific (D), PRSSCZ-specific (E) and PRSANX-specific (F) as exposure variable, respectively. Incidence rate ratios indicate the risk estimates for +1 SD increase of PRS. Sex, birth year and first 10 ancestry-informative principal components were adjusted for in all models. The points represent hazard ratio estimates, and the error bars indicate 95% confidence intervals. “*” represents association that remained significant after Bonferroni correction. “#” represents trending association at P < 0.05 but was not significant after Bonferroni correction. PRS polygenic risk scores, AN anorexia nervosa, OCS obsessive-compulsive symptoms, OCD obsessive-compulsive disorder, SCZ schizophrenia, MDD major depressive disorder, ANX anxiety disorders, ASD autism spectrum disorder, ADHD attention-deficit hyperactivity disorder, SUD substance use disorder.
Association between gSEM-derived PRSs and AN symptomatology at first clinical visit
PRSshared was associated with more severe ED symptoms (regression coefficient, 0.07; 95%CI, 0.01–0.14; P = 0.03), greater clinical impairment (regression coefficient, 1.13; 95%CI, 0.45–1.82; P = 1.25 × 10−3), greater self-reported anxiety (regression coefficient, 0.52; 95%CI, 0.27–0.77; P = 4.43 × 10−5) and depression (regression coefficient, 0.46; 95%CI, 0.17–0.74; P = 1.71 × 10−3) at P < 0.05 (Fig. 4A; Table S10). Statistically significant associations were also observed between PRSAN-specific and lower BMI (regression coefficient, −0.41; 95%CI, −0.58–−0.24; P = 1.78 × 10−6) at P < 0.05 (Fig. 4B; Table S10). After correction, only the associations for PRSshared with self-reported anxiety symptoms and PRSAN-specific with lower BMI remained significant.
Results are derived from linear regression models with BMI, EDE-Q score, CPRS-depression score, CPRS-anxiety score and CIA score as the outcome variables. The analyses were based on 1934 ANGI-SE AN cases with available data on symptomatology in Stepwise Quality Register. For each of the five outcomes, we constructed six models with PRSshared (A), PRSAN-specific (B), PRSOCS-specific (C), PRSMDD-specific (D), PRSSCZ-specific (E) and PRSANX-specific (F) as the exposure variable, respectively. Regression coefficients indicate the risk estimates for +1 SD increase of PRS. Sex, birth year and first 10 ancestry-informative principal components were adjusted for in all models. The points represent regression coefficients, and the error bars indicate 95% confidence intervals. “*” represents association that remained significant after Bonferroni correction. “#” represents trending association at P < 0.05 but was not significant after Bonferroni correction. PRS polygenic risk scores, AN anorexia nervosa, OCS obsessive-compulsive symptoms, SCZ schizophrenia, MDD major depressive disorder, ANX anxiety disorders, BMI body mass index, CIA Clinical Impairment Assessment, CPRS Comprehensive Psychopathological Rating Scale, CPRS-dep CPRS-depression score, CPRS-anx CPRS-anxiety score, EDE-Q Eating Disorder Examination Questionnaire.
Discussion
This study is the first to leverage transdiagnostic psychiatric genetic liability to dissect the clinical heterogeneity of AN so far. We found that shared psychiatric genetic liability was a consistent predictor of disease burden, risk of psychiatric comorbidity and clinical impairment within AN, whereas AN-specific liability was not a good indicator of its clinical course. These findings provide a novel perspective on the heterogeneous etiology and clinical course of AN.
We observed that only AN-specific liability and shared liability were associated with an elevated AN risk. However, all other gSEM-derived trait-specific PRSs were associated with either a statistically non-significant or reduced risk of AN. This offers a completely different perspective on AN etiology compared to the consistently positive relationships between unmodified single-disorder PRSs and AN risk. Since the single-disorder PRSs can capture both shared and trait-specific genetic factors, their associations with increased AN risk might be attributed to the shared psychiatric genetic component. These findings suggest that both general psychiatric and AN-specific genetic factors are underlying the onset of AN.
Shared psychiatric liability was associated with greater cumulative disease burden, elevated risks of psychiatric comorbidities and more severe symptoms and clinical impairment in AN. The findings are consistent with previous studies revealing a positive association between family history of SCZ and AN disease burden, as well as between OCD-AN shared liability and AN symptoms [11, 12]. According to the “p factor” theory, there are shared psychopathological mechanisms underlying multiple psychiatric disorders which could be captured by a common latent factor [30]. The positive association between shared liability and psychiatric disease burden in AN offers biological evidence for the “p-factor” in this population. Furthermore, we shed new light on the genetic underpinnings of this “p-factor” by revealing the relationship between shared liability and other clinical outcomes (i.e. more somatic diagnoses, more medication prescriptions, more inpatient days, more severe symptoms and clinical impairment) in AN.
AN-specific liability did not show statistically significant associations with most of the clinical features in AN. It is interesting that AN-specific liability was a predictor for AN status as expected, but not for its clinical outcomes, suggesting that distinct genetic mechanisms may underlie the onset and prognosis of AN. However, its associations with lower BMI and more inpatient days due to EDs suggest a potential role of AN-specific liability as a predictor for AN-specific symptoms and severity, which is consistent with a previous study revealing that AN PRS showed better performance in predicting ED severity compared to a cross-disorder PRS [12]. Given the low heritability of AN-specific genetic effects (Table S11), the analyses should be replicated when larger GWASs emerge.
MDD-specific liability was associated with greater disease burden within AN. Early-life depression has previously been found to be associated with an increased risk of somatic conditions, possibly due to shared inflammatory or metabolic mechanisms [31,32,33,34]. Moreover, genetic overlap has been observed between MDD and endocrine disorders, obesity, and inflammatory cytokines [33, 35,36,37,38]. The association between MDD-specific genetic risk and poor health status in AN might be mediated by these biological mechanisms. Our findings should be interpreted with caution given the low heritability of disorder-specific genetic effects and still growing sample sizes of source GWASs.
Limitations
Our study has several limitations. Firstly, the source GWAS for AN has relatively small sample size, and the GWAS for OCS is based on self-reported obsessive compulsive symptoms, so our analyses should be replicated when larger and diagnosis-based GWASs for AN and OCD respectively are available. Secondly, although our gSEM model fit was good and we showed that the gSEM-derived components had acceptable validity in our study population (Supplementary Results, Table S12), the results for ANX-specific liability should be interpreted with caution since the correlation between PRSANX-specific and PRSANX was not strong (Supplementary Results, Table S13 and Fig. S2). Moreover, data on AN symptoms were missing for a considerable proportion of participants and were measured by self-report questionnaires which might lead to underestimates due to subjective denial of symptoms [39]. Given the limited follow-up time, our data might not cover first diagnoses of some psychiatric comorbidities with an early onset age, such as ADHD and ASD. Finally, effects from rare variants and structural variations were not considered in the current study.
Conclusions
Our findings show that shared instead of AN-specific liability was a strong predictor for adverse outcomes of AN, suggesting that genetic risk profiles for AN diagnosis may be distinct from those for AN outcomes. We provide a novel perspective on the heterogenous clinical outcomes within AN by identifying clinically relevant genetic components.
Data availability
GWAS summary statistics for anorexia nervosa, obsessive-compulsive symptoms, schizophrenia, major depressive disorder and anxiety disorders are publicly available at the Psychiatric Genomics Consortium data downloads portal: https://pgc.unc.edu/for-researchers/download-results/. The individual genotype data for ANGI-SE participants are deposited in dbGaP (http://www.ncbi.nlm.nih.gov/gap) under accession number phs001541.v1.p1. However, their linked information from the Swedish National Registers cannot be shared publicly due to legal and ethical restrictions.
References
Treasure J, Duarte TA, Schmidt U. Eating disorders. Lancet. 2020;395:899–911.
Duncan L, Yilmaz Z, Gaspar H, Walters R, Goldstein J, Anttila V, et al. Significant locus and metabolic genetic correlations revealed in genome-wide association study of anorexia nervosa. Am J Psychiatry. 2017;174:850–8.
Steinhausen HC. The outcome of anorexia nervosa in the 20th century. Am J Psychiatry. 2002;159:1284–93.
Yilmaz Z, Hardaway JA, Bulik CM. Genetics and epigenetics of eating disorders. Adv Genom Genet. 2015;5:131–50.
Watson HJ, Yilmaz Z, Thornton LM, Hübel C, Coleman JRI, Gaspar HA, et al. Genome-wide association study identifies eight risk loci and implicates metabo-psychiatric origins for anorexia nervosa. Nat Genet. 2019;51:1207–14.
Fernandez-Aranda F, Pinheiro AP, Tozzi F, Thornton LM, Fichter MM, Halmi KA, et al. Symptom profile of major depressive disorder in women with eating disorders. Aust N Z J Psychiatry. 2007;41:24–31.
Godart NT, Flament MF, Perdereau F, Jeammet P. Comorbidity between eating disorders and anxiety disorders: a review. Int J Eat Disord. 2002;32:253–70.
Drakes DH, Fawcett EJ, Rose JP, Carter-Major JC, Fawcett JM. Comorbid obsessive-compulsive disorder in individuals with eating disorders: an epidemiological meta-analysis. J Psychiatr Res. 2021;141:176–91.
Zhang R, Larsen JT, Kuja-Halkola R, Thornton L, Yao S, Larsson H, et al. Familial co-aggregation of schizophrenia and eating disorders in Sweden and Denmark. Mol Psychiatry. 2021;26:5389–97.
Morylowska-Topolska J, Ziemiński R, Molas A, Gajewski J, Flis M, Stelmach E, et al. Schizophrenia and anorexia nervosa - reciprocal relationships. A literature review. Psychiatr Pol. 2017;51:261–70.
Zhang R, Kuja-Halkola R, Birgegård A, Larsson H, Lichtenstein P, Bulik CM, et al. Association of family history of schizophrenia and clinical outcomes in individuals with eating disorders. Psychol Med. 2023;53:371–8.
Yilmaz Z, Schaumberg K, Halvorsen M, Goodman EL, Brosof LC, Crowley JJ, et al. Predicting eating disorder and anxiety symptoms using disorder-specific and transdiagnostic polygenic scores for anorexia nervosa and obsessive-compulsive disorder. Psychol Med. 2022;53:1–15.
Richards AL, Cardno A, Harold G, Craddock NJ, Di Florio A, Jones L, et al. Genetic liabilities differentiating bipolar disorder, schizophrenia, and major depressive disorder, and phenotypic heterogeneity in bipolar disorder. JAMA Psychiatry. 2022;79:1032–9.
Peyre H, Schoeler T, Liu C, Williams CM, Hoertel N, Havdahl A, et al. Combining multivariate genomic approaches to elucidate the comorbidity between autism spectrum disorder and attention deficit hyperactivity disorder. J Child Psychol Psychiatry. 2021;62:1285–96.
Thorp JG, Campos AI, Grotzinger AD, Gerring ZF, An J, Ong JS, et al. Symptom-level modelling unravels the shared genetic architecture of anxiety and depression. Nat Hum Behav. 2021;5:1432–42.
Grotzinger AD, Rhemtulla M, de Vlaming R, Ritchie SJ, Mallard TT, Hill WD, et al. Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat Hum Behav. 2019;3:513–25.
Trubetskoy V, Pardiñas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Als TD, Kurki MI, Grove J, Voloudakis G, Therrien K, Tasanko E, et al. Depression pathophysiology, risk prediction of recurrence and comorbid psychiatric disorders using genome-wide analyses. Nat Med. 2023;29:1832–44.
Friligkou E, Løkhammer S, Cabrera-Mendoza B, Shen J, He J, Deiana G, et al. Gene discovery and biological insights into anxiety disorders from a large-scale multi-ancestry genome-wide association study. Nat Genet. 2024;56:2036–45.
Strom NI, Burton CL, Iyegbe C, Silzer T, Antonyan L, Pool R, et al. Genome-wide association study of obsessive-compulsive symptoms including 33,943 individuals from the general population. Mol Psychiatry. 2024;29:2714–23.
Thornton LM, Munn-Chernoff MA, Baker JH, Juréus A, Parker R, Henders AK, et al. The Anorexia Nervosa Genetics Initiative (ANGI): Overview and methods. Contemp Clin Trials. 2018;74:61–9.
Lam M, Awasthi S, Watson HJ, Goldstein J, Panagiotaropoulou G, Trubetskoy V, et al. RICOPILI: Rapid Imputation for COnsortias PIpeLIne. Bioinformatics. 2020;36:930–3.
Ludvigsson JF, Almqvist C, Bonamy AK, Ljung R, Michaëlsson K, Neovius M, et al. Registers of the Swedish total population and their use in medical research. Eur J Epidemiol. 2016;31:125–36.
Fairburn CG, Beglin SJ. Assessment of eating disorders: interview or self-report questionnaire? Int J Eat Disord. 1994;16:363–70.
Svanborg P, Asberg M. A new self-rating scale for depression and anxiety states based on the comprehensive psychopathological rating scale. Acta Psychiatr Scand. 1994;89:21–8.
Bohn K, Doll HA, Cooper Z, O’Connor M, Palmer RL, Fairburn CG. The measurement of impairment due to eating disorder psychopathology. Behav Res Ther. 2008;46:1105–10.
Birgegård A, Björck C, Clinton D. Quality assurance of specialised treatment of eating disorders using large-scale Internet-based collection systems: methods, results and lessons learned from designing the Stepwise database. Eur Eat Disord Rev. 2010;18:251–9.
Choi SW, O’Reilly PF. PRSice-2: polygenic risk score software for biobank-scale data. Gigascience. 2019;8:giz082.
Coombes BJ, Ploner A, Bergen SE, Biernacka JM. A principal component approach to improve association testing with polygenic risk scores. Genet Epidemiol. 2020;44:676–86.
Caspi A, Houts RM, Belsky DW, Goldman-Mellor SJ, Harrington H, Israel S, et al. The p factor: one general psychopathology factor in the structure of psychiatric disorders? Clin Psychol Sci. 2014;2:119–37.
Leone M, Kuja-Halkola R, Leval A, D’Onofrio BM, Larsson H, Lichtenstein P, et al. Association of youth depression with subsequent somatic diseases and premature death. JAMA Psychiatry. 2021;78:302–10.
Hu X, Pang H, Liu J, Wang Y, Lou Y, Zhao Y. A network medicine-based approach to explore the relationship between depression and inflammation. Front Psychiatry. 2023;14:1184188.
Leone M, Kuja-Halkola R, Leval A, Butwicka A, Skov J, Zhang R, et al. Genetic and environmental contribution to the co-occurrence of endocrine-metabolic disorders and depression: a nationwide Swedish Study of Siblings. Am J Psychiatry. 2022;179:824–32.
Chu K, Cadar D, Iob E, Frank P. Excess body weight and specific types of depressive symptoms: Is there a mediating role of systemic low-grade inflammation? Brain Behav Immun. 2023;108:233–44.
Maina JG, Balkhiyarova Z, Nouwen A, Pupko I, Ulrich A, Boissel M, et al. Bidirectional Mendelian randomization and multiphenotype GWAS show causality and shared pathophysiology between depression and type 2 diabetes. Diabetes Care. 2023;46:1707–14.
Hughes AM, Sanderson E, Morris T, Ayorech Z, Tesli M, Ask H, et al. Body mass index and childhood symptoms of depression, anxiety, and attention-deficit hyperactivity disorder: a within-family Mendelian randomization study. Elife. 2022;11:e74320.
Liao SF, Su CY, Su MH, Chen CY, Chen CY, Lin YF, et al. Associations of polygenic risks, depression, and obesity-related traits in Taiwan Biobank. J Affect Disord. 2023;320:397–403.
Draganov M, Arranz MJ, Salazar J, de Diego-Adeliño J, Gallego-Fabrega C, Jubero M, et al. Association study of polymorphisms within inflammatory genes and methylation status in treatment response in major depression. Eur Psychiatry. 2019;60:7–13.
Vandereycken W. Denial of illness in anorexia nervosa—a conceptual review: part 1 diagnostic significance and assessment. Eur Eat Disord Rev. 2006;14:341–51.
Funding
This study was funded by the Chinese Scholarship Council (CSC202206010089) and Vetenskapsrådet 2021-03126 (PI: Sarah E. Bergen). Cynthia M. Bulik is supported by NIMH (R56MH129437; R01MH120170; R01MH124871; R01MH119084; R01MH118278; R01MH124871); Swedish Research Council (Vetenskapsrådet, award: 538-2013-8864); Lundbeck Foundation (Grant no. R276-2018-4581). The Anorexia Nervosa Genetics Initiative (ANGI) was an initiative of the Klarman Family Foundation. We acknowledge and thank Stina Borg for her support on ANGI data extraction, as well as Ruyue Zhang for her analytic support and data curation. The computations and data handling were enabled by resources provided by the National Academic Infrastructure for Supercomputing in Sweden (NAISS) and the Swedish National Infrastructure for Computing (SNIC) at Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX) partially funded by the Swedish Research Council through grant agreements no. 2022-06725 and no. 2018-05973. The funding organizations had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication. Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Consortia
Contributions
Z-AL participated in the conceptual generation of the study and acquisition of data, conducted statistical analyses, and drafted the original manuscript. AP contributed substantially to the study design and statistical analyses and provided critical comments on the manuscript. AB helped with the study design and provided comments on the manuscript. ML oversaw parts of the ANGI-SE data collection and commented on the manuscript. CMB leads the ANGI study and was responsible for its funding and the overall data collection. She also offered comments on the manuscript. SEB was responsible for acquiring funding, conceptualization and supervision of this study, helped with the statistical analyses and provided critical comments during the manuscript preparation. The manuscript was circulated among members of Eating Disorders Working Group of the Psychiatric Genomics Consortium for comments.
Corresponding author
Ethics declarations
Competing interests
Cynthia M. Bulik reports: Lundbeckfonden (grant recipient); Pearson (author, royalty recipient); Equip Health Inc. (Stakeholder Advisory Board). Other authors declare no conflict of interest.
Ethics approval and consent to participate
All methods were performed in accordance with the relevant guidelines and regulations. The ANGI-SE study was approved by the regional Ethical Review Board in Stockholm (DNR 2013/112-31/2). Informed consent was obtained from all participants.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, ZA., Ploner, A., Birgegård, A. et al. Leveraging transdiagnostic genetic liability to psychiatric disorders to dissect clinical outcomes of anorexia nervosa. Mol Psychiatry 31, 1475–1484 (2026). https://doi.org/10.1038/s41380-025-03264-x
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41380-025-03264-x