Abstract
Anti-N-methyl-D-aspartate receptor (NMDAR) encephalitis (NMDARE) is a devastating autoimmune disease associated with the presence of autoantibodies targeting NMDAR1 in the cerebrospinal fluid (CSF) and serum of patients. Besides the critical roles of anti-NMDAR1 autoantibodies, studies have implicated other factors such as brain inflammation in NMDARE. To comprehensively uncover the molecular mechanisms underlying NMDARE, here we performed multi-omics analyses based on human forebrain organoids (hFOs). The transcriptomic and metabolomic analyses showed that hFOs exposed to either monoclonal anti-NMDAR1 IgG antibodies or purified patient CSF-derived IgG antibodies alone led to NMDAR hypofunction that caused a reduction of glutamate content and neuroactivities. Interestingly, hFOs exposed to either patient CSF or IgG-depleted patient CSF led to neuronal hyperexcitability rather than hypo neuroactivities. The following proteomic analysis and electrophysiological assays identified that the activated interleukin (IL)-17 signaling pathway in patient CSF accounted for the neuronal hyperexcitability. Neutralizing IL-17 alleviated the neuronal hyperexcitability in hFOs and seizure-like behaviors in mice exposed to CSF from NMDARE patients. Together, this study indicated that the anti-NMDAR1 antibodies and IL-17 signaling pathway shape NMDARE. Inactivating the IL-17 signaling pathway could be a potential therapeutic strategy for NMDARE treatment.
Similar content being viewed by others
Introduction
Anti-N-methyl-D-aspartate receptor (NMDAR) encephalitis (NMDARE) is an autoimmune disease characterized by the presence of autoantibodies targeting the NR1 subunit of NMDAR (NMDAR1) [1]. NMDARE shows complex and diverse clinical features, commonly including psychiatric symptoms, epileptic seizures, cognitive decline, and behavioral abnormalities [2,3,4]. Despite an increasing number of studies have emerged since its discovery in 2007 [1], the pathogenesis of NMDARE is not fully understood.
NMDARE is closely associated with the presence of autoreactive antibodies against NMDAR1 in the cerebrospinal fluid (CSF) and serum of patients [5]. Both in vitro and in vivo studies have shown that patient-derived anti-NMDAR1 antibodies (NMDAR1-Ab) lead to internalization and reduction of cell surface NMDAR1, resulting in NMDAR hypofunction such as decreased NMDAR-mediated synaptic currents [6,7,8]. Since NMDAR plays important roles in glutamatergic synaptic transmission, learning, and behavior [9], NMDAR hypofunction is thought to account for the psychiatric and behavioral symptoms in NMDARE. Though these core clinical features of NMDARE could be explained by NMDAR hypofunction [10,11,12], some clinical manifestations such as epileptic seizures are not well explained by NMDAR hypofunction. Though several studies reported that NMDAR1-Ab alone could lead to seizures [13,14,15,16], other studies showed that antibodies targeting NMDAR1 caused hypo neuroactivities rather than seizures [6,7,8, 12, 17]. Whether NMDAR1-Ab themselves cause all NMDARE symptoms including seizures needs further validation.
In addition to NMDAR1-Ab, brain inflammation may also play important roles in NMDARE18. Studies showed that molecules including cytokines and chemokines related to brain-blood-barrier damage and brain inflammation were significantly elevated in CSF of NMDARE patients [18,19,20,21,22,23]. Some studies proposed that both NMDAR1-Ab and brain inflammation contribute to the NMDARE phenotypes [24,25,26]. These studies provided new disease insights, but more evidence is needed to deepen our understanding of NMDARE.
Our current understanding of the molecular mechanisms of NMDARE is mainly based on in vitro 2D cell cultures and animal models. These models are limited to fully recapitulate the complex cellular interactions [27, 28] and human immune response specificity [29] involved in NMDARE, respectively. A human cell-based model that recapitulates key features of the human brain is urgently needed for the study of NMDARE.
Human brain organoids, the recently developed pluripotent stem cell-derived 3D cell cultures, have become attractive models that recapitulate complex cellular architecture and physiological function of the human brain [30, 31]. Brain organoids have been used to model viral encephalitis caused by herpes simplex virus 1 infection [32], providing potential improvement for the current therapeutic strategies. Given the merits of brain organoids, leveraging this model to study NMDARE facilitates uncovering the disease pathogenesis and improving the current therapeutic strategies.
Here we performed transcriptomic and metabolomic analysis based on human forebrain organoids (hFOs), showing that both monoclonal anti-NMDAR1 IgG antibodies and purified patient CSF-derived IgG antibodies led to NMDAR hypofunction that caused a reduction of glutamate content and neuroactivities in hFOs. The following proteomic analysis and animal experiments identified that the activated interleukin-17A (IL-17) signaling pathway in patient CSF mediated the seizure phenotypes in NMDARE. Together, this study indicated that the IL-17 signaling pathway and NMDAR1-Ab shape NMDARE. Neutralizing IL-17 may be a therapeutic strategy for NMDARE.
Materials and methods
CSF sample collection
All NMDARE patients in this study met the clinical diagnostic criteria for anti-NMDARE revised in 2016 [33]. Patient CSF samples were collected before initiating immunotherapy. CSF samples from non-affected controls were collected from individuals negative for the detection of brain inflammation and NMDAR1-Ab. The study was approved by the ethics committee of the First Affiliated Hospital of University of South China. Informed consent documents were signed by all the participants in this study. The demographic and clinical data of all participants are presented in Supplementary Table 1. The collected CSF samples were centrifuged at 1000 × g for 10 min at 4 °C. The supernatants were collected and stored at −80 °C for downstream analysis.
IgG purification and depletion from CSF
IgG antibodies were purified from CSF using the NAbTM Protein G Spin Kit (Thermo Fisher, 89979) according to the manufacturer’s instructions. In brief, set the centrifuge to 1000 × g and equilibrate the NAb spin column and buffers to room temperature in advance. Place the column in a 15 mL collection tube and centrifuge for 1 min to remove the storage solution. Discard the flow-through. Equilibrate the column by adding 2 mL Binding Buffer. Centrifuge for 1 min and discard the flow-through. After that, the column was ready for IgG purification or depletion.
Prepare CSF samples for purification by diluting them to 2 mL in the Binding Buffer. Add the CSF samples to the column and mix end-over-end for 10 min at room temperature. Transfer the column to a new 15 mL collection tube and centrifuge for 1 min. The flow-through solution was collected as IgG-depleted CSF samples. IgG antibodies were retained in the column. After the elution and neutralization steps, the purified IgG was obtained and concentrated with a freeze dryer (SCIENTZ, 10 N/A). The concentration of purified IgG was calibrated to 1 mg/mL with double-distilled water.
Cell culture
The human induced pluripotent stem cells (hiPSCs) derived from one healthy female donor (U2F) were obtained from the Cellapy Technology (Beijing, China). The pluripotency and karyotype of U2F hiPSCs were confirmed by immunofluorescence and karyotype analysis as shown in our previous study [34]. The hiPSCs were cultured in Matrigel (Corning, 354277)-coated plate supplemented with the mTeSR Plus medium (STEMCELL Technologies, 05825). The HEK 293 T cells were cultured in high-glucose DMEM (Gibco, C11995500BT) supplemented with 10% fetal bovine serum (Gibco, A3161001C) and 1% penicillin/streptomycin (Gibco, 10378016).
Cell-based assay
We performed a cell-based assay in HEK 293 T cells to confirm the presence of NMDAR1-Ab in CSF of NMDARE patients. The pcDNA3.1 plasmid containing NMDAR1 coding sequence was transfected into HEK 293 T cells using Lipofectamine 3000 transfection reagent (Thermo Fisher, L3000001). 48 h post-transfection, cells were fixed in 4% paraformaldehyde and permeabilized with 0.1% Triton X-100. Subsequently, cells were incubated with patient CSF (1:10) or rabbit anti-NMDAR1 primary antibody (Abcam, ab109182, 1:100) which was used as a positive control. Following this, cells were incubated with appropriate secondary antibodies FITC Anti-Human IgG antibody (Abcam, ab81051, 1:500) or Alexa Fluor 594-conjugated goat anti-rabbit IgG (Thermo Fisher, A11012, 1:500). After incubation with the DAPI nuclei staining solution (10 μg/ml, Invitrogen, D1306), fluorescent figures were taken using a microscope (Zeiss, Axio Vert A1).
Generation of hFOs
The U2F hiPSCs were used to generate hFOs based on our published methods [34] with some modifications. Briefly, on day 0, hiPSCs were dissociated into single-cell suspension using Accutase solution (Sigma-Aldrich, A6964). 1 × 104 cells per well were seeded into the 96-well round-bottom ultra-low attachment plate (Corning, 7007) and cultured in the embryoid body formation medium for 5 days. On day 5, half of the medium was replaced with the Neural Induction Medium. On day 7, the medium was completely replaced with the Neural Induction Medium. On day 11, hFOs were cultured in the Organoid Differentiation Medium. From day 30, hFOs were cultured in the Organoid Maturation Medium. The Organoid Differentiation and Maturation Media were changed every 4–5 days. hFOs were characterized by the immunostaining analysis.
Immunostaining analysis
hFOs were fixed with 4% paraformaldehyde in PBS for 1 h at room temperature. hFOs were then incubated in 30% sucrose solution overnight at 4 °C. After embedding in O.C.T Compound (Tissue-Tek, 4583), hFOs were sectioned at a thickness of 16 μm using a cryostat (Leica, CM1950). The sections were permeabilized with 0.1% Triton X-100 in PBS for 30 min at room temperature. For staining of cell surface NMDAR1, hFOs sections were not treated with 0.1% Triton X-100. After washing with PBS three times, sections were blocked with 5% bovine serum albumin (Solarbio, A8020) for 1 h at 37 °C. Sections were then incubated with primary antibodies (e.g., Mouse Anti-NMDAR1, Abcam, ab134308) overnight at 4 °C. After washing with PBS three times, sections were incubated with secondary antibodies for 1 h and DAPI solution for 5 min at room temperature. Images were taken using a fluorescent microscope. The primary and secondary antibodies used in this study are shown in Supplementary Table 2.
Bulk organoid RNA sequencing (RNA-seq) and data analysis
For bulk organoid RNA-seq, two hFOs were randomly pooled together as one mixed sample in the NMDAR1-Ab exposure or control group. At least three mixed samples in each group were used for total RNA extraction with the miRNeasy Mini Kit (Qiagen, 217004). After assessing their quality, RNA samples were used for polyA+ selection RNA-seq library construction. The constructed libraries were then sequenced on the BGI DNBseq platform (150 bp, paired-end).
Raw RNA-seq reads were filtered to get clean reads using FastQC (v0.20.0). The clean reads were then aligned to the human genome hg38 using STAR (v2.7.6a). Gene expression quantification was conducted using RSEM (v1.3.0). The filterByExpr function in the edgeR package (v3.36.0) was used to filter out low-expression genes. The sva function in the SVA package (v3.42.0) was used to estimate the batch effect and other artifacts. Differential gene expression analysis between the NMDAR1-Ab exposure and control group was performed using the DEseq2 package (v1.34.0). P values were adjusted using the Benjamini–Hochberg method.
Protein-protein interaction (PPI) analysis
We used the downregulated differentially expressed genes (DEGs) in NMDAR1-Ab-exposed hFOs to construct a high-confidence (interaction score > 0.7) PPI network based on the STRING database (v11.5) (http://www.string-db.org/). Active interaction sources included text mining, experiments, databases, coexpression, neighborhood, gene fusion, and co-occurrence. The constructed PPI network was visualized using Cytoscape (v3.9.1). CytoHubba [35], a Cytoscape plugin, was used to explore hub nodes in the PPI network.
Real-time quantitative PCR (RT-qPCR) analysis
Total RNA extracted from two mixed hFOs was used to generate complementary DNA using the HiScript III RT SuperMix for qPCR (+gDNA wiper) (Vazyme, R323-01). RT-qPCR assay was performed using the ChamQ SYBR qPCR Master Mix (Vazyme, Q711-02) on the Real-Time PCR System (Thermo Fisher, QuantStudio 3). GAPDH was used as the internal reference gene. At least three technical replicates were used in the RT-qPCR analysis. Sequences of RT-qPCR primers are provided in Supplementary Table 3.
Metabolomic analysis of bulk hFOs
For the metabolomic analysis, three hFOs (day 110) from the NMDAR1-Ab exposure or control group were randomly pooled together as one mixed sample. Five mixed samples in each group were then used for targeted metabolomic analysis on the 600MRM platform (Biotree, China). Briefly, hFOs or quality control samples were lysed in 1000 μL acetonitrile-methanol-H2O (2:2:1, containing isotope internal standards). After sonication, samples were incubated at −40 °C for 2 h. The lysate was centrifuged (13000 g, 4 °C, 15 min) to get the supernatant. 800 μL supernatant of each sample was transferred to a new tube and concentrated with a freeze dryer. 160 μL of 60% acetonitrile was added to the dried samples, followed by centrifugation at 13000 g, 4 °C, 15 min. The supernatant of each sample was collected for LC-MS/MS analysis on the SCIEX QTRAP 6500+ system. Typical ion source parameters were: IonSpray Voltage = +5000 V/-4500V, Curtain Gas = 35 psi, Temperature = 400 °C, Ion Source Gas 1 = 50 psi, Ion Source Gas 2 = 50 psi. The content of metabolites (nmol/g) was quantified based on the formula (C*V*1.25/m), where C represents the calculated concentration (μmol/L), V represents the final sample volume (160 μL), and m represents the sample weight (mg).
To filter out outline samples and identify differentially expressed metabolites, the Orthogonal projections to latent structures-discriminant analysis (OPLS-DA) model was used. The variable importance in projection (VIP) value of the first principal component in the OPLS-DA model was calculated. Metabolites with VIP > 1 and P < 0.05 (Two-tailed t-test) were considered differentially expressed between the NMDAR1-Ab exposure and control hFOs. Functional annotation for the differentially expressed metabolites was performed using the online tool MetaboAnalyst 5.0 (https://www.metaboanalyst.ca/).
Microelectrode array (MEA) assay
MEA assays were used to assess electrophysiological properties after NMDAR1-Ab or patient CSF exposure in hFOs. Briefly, hFOs (day 110) were plated on a Matrigel-coated 24-well MEA culture plate. After 7 days of culture, hFOs were exposed to NMDAR1-Ab or patient CSF or controls for 24 h. The neuroactivities of hFOs were recorded using the MEA system (Axion Biosystems, Master Edge). Each recording duration was 15 min. The MEA data including the number of spikes, burst frequency, and network synchronization were analyzed using the software Axis Navigator (v3.6.2.2).
Calcium imaging analysis
For calcium imaging analysis, hFOs were loaded with 5 μM Fluo-4 AM (Beyotime, S1060) for 30 min at 37 °C. Imaging was performed using a fluorescent microscope (Zessi, Axio Vert A1) at a frequency of 2 Hz for 3 min. The resulting images were then analyzed using ImageJ (v2.1.0) as described previously [30]. Regions of interest (ROIs) were manually selected and mean fluorescence was calculated for each time frame. For each ROI, the change in fluorescence was calculated as follows: (F-Fbasal)/Fbackground, where Fbasal was the lowest mean fluorescence value across all frames, and Fbackground was the average mean fluorescence across all frames. In a given time period, the number of spikes represented the intensity of the calcium transient activities.
Olink proteomic analysis
CSF samples from 6 NMDARE patients and 6 non-affected controls were used for proteomic analysis using the Olink® target 96 inflammation panel (Olink Proteomics AB, Sweden). The proximity extension assay technology used for the Olink protocol enabled 92 inflammation-related proteins to be analyzed simultaneously. In brief, each targeted protein was tagged by pairs of oligonucleotide-labeled antibody probes. When matching pairs of antibodies bind to the protein, the oligonucleotides hybridize and finally create a unique PCR target sequence, which was subsequently detected and quantified using a microfluidic real-time PCR instrument (Signature Q100, LC-Bio Technology CO., Ltd., China). The final assay read-out is presented in normalized protein expression values and subsequently log2-transformed for statistical analysis. The Welch’s two-tailed t-test was used to identify differentially expressed proteins between the NMDARE patients and non-affected controls with a P value cutoff of 0.05.
Animals
Healthy C57BL/6 male mice (aged 8–10 weeks, weight 22–28 g) were bought from the Tianqin Biotechnology (Changsha, China) and habituated for one week before surgery. The room was maintained at a controlled temperature (21 ± 1 °C) and humidity (55 ± 10%) with 12 h light/12 h dark cycles. Food and water were available ad libitum. During all the animal experiments, mice were randomly allocated into different groups. The order of treatments and measurements, or animal/cage location were randomized to minimize potential confounders. All procedures were conducted in accordance with the Regulations for the Administration of Affairs Concerning Experimental Animals (China) and were approved by the Animal Care and Use Committee of the University of South China.
Intracerebroventricular infusion of human CSF in mice
To generate a mouse model of epileptic seizures, intracerebroventricular infusion of human CSF was performed using the micro-osmotic pumps (0.25 μL/h, model 1002, Alzet) and Brain Infusion Kit 3 (0008851, Alzet). The surgical procedures were based on the published methods [10] with some modifications. Briefly, the osmotic pumps were pre-loaded with 100 μL mixed patient CSF or mixed non-affected control CSF or 100 μL patient CSF premixed with 100 μg anti-IL-17 monoclonal antibodies (AIN457, Selleck, A2025). The pumps were then connected to the brain infusion cannula and vinyl catheter tubes in the Brain Infusion Kit 3. Mice were anaesthetized with 1.5% isoflurane and placed on a stereotaxic frame. A hole was drilled through the skull according to the following location: 0.02 mm anterior and 1.00 mm lateral from bregma, depth 0.22 mm. The infusion cannula was then inserted into one side of the lateral ventricles. Holes were secured with dental cement and pumps were subcutaneously implanted on the back of the mice. After the surgery, mice were immediately placed on a warmed heat pad until complete recovery.
Evaluation of seizure stages
Mice were observed continuously for at least 5 min to evaluate their seizure-like phenotypes after pentylenetetrazol (PTZ) injection. The severity of seizure was evaluated using the Racine scale [36]. A higher Racine scale score indicated a more severe seizure stage. Scoring of the Racine scale refers to the following standards: 0, normal behavior; 1, immobile, staring; 2, head nodding, shivering, facial clonus; 3, partial-body clonus, stiff and upright tail; 4, whole-body tonic-clonus, loss of postural control, falling; 5, non-intermittent tonic–clonus, supine, violent jumping and running; 6, death. Before evaluation of seizure stages, mice died for unknown reasons were excluded. Investigators were blinded to the group allocation. Data were processed in a blindness manner.
Statistical analysis
Data were presented as mean ± SEM unless indicated elsewhere. Data normality and variance were assessed using GraphPad Prism 9.0. Sample size was determined based on preliminary data and previous studies. No statistical methods were used to estimate sample size. Differences between two groups with normally distributed data were assessed using a two-tailed t-test. The Kruskal-Wallis test was used for multiple comparisons in non-normally distributed data. Two-way ANOVA was used for the statistical analysis of the MEA data. The hypergeometric test was used to assess the enrichment between NMDAR1-Ab exposure-induced DEGs and brain disorder-associated gene sets. P values in the differential gene expression analysis were adjusted using the Benjamini-Hochberg method to adjust for multiple testing. Statistical significance was defined as a P value less than 0.05 in all the statistical analyses.
Results
NMDAR1-Ab exposure reduces cell surface NMDAR1 intensity in hFOs
To initiate this study, we investigated the functional effects of NMDAR1-Ab exposure using hFOs. hFOs were generated from hiPSCs derived from one healthy donor based on our published methods [34] (Fig. 1a). The generated hFOs were featured with forebrain identity (FOXG1 + ) and contained ventricle-like structures marked by neural progenitors (SOX2 + ), intermediate zone marked by intermediate progenitors (TBR2 + ), and cortical-like regions containing neurons (MAP2 + ) (Fig. 1b) at early differentiation stage (< 40 days). In addition to neural lineage cell types (PAX6+ and MAP2 + ), cells that expressed astrocyte marker (GFAP + ) or oligodendrocyte marker (OLIG3 + ) were also presented in hFOs (day 110) (Fig. 1c), indicating that the hFOs contained major cell types emerging in the developing human brain. Importantly, the immunostaining confirmed the widespread expression of NMDAR1 in hFOs (Fig. 1c).
a Schematic diagram for hFOs culture. SB, SB431542; LDN, LDN193189; b Immunostaining characterization for representative control hFOs (day 30). Ventricular zone markers, SOX2 and ki67; intermediate zone marker, TBR2; cortical plate marker, MAP2; forebrain-specific marker, FOXG1. Scale bars, 50 μm and 25 μm. c Immunostaining characterization for major brain cell types in representative control hFOs (day 110). Neural stem cell marker, PAX6; neuron marker, MAP2; astrocyte marker, GAFP; oligodendrocyte marker, OLIG3; Scale bar, 25 μm. (d-g) Density of cell surface NMDAR1 and PSD95 in hFOs after exposure to monoclonal NMDAR1-Ab (d, e) or IgG purified from CSF of NMDARE patients (f, g). Synaptic marker, PSD95. At least 3 independent hFOs per group were used for the immunostaining analysis. Data are shown as Mean ± SEM. Scale bar, 20 μm. Two-tailed t-test, **P < 0.01; NS, non-significance.
A previous study used both commercial antibody targeting the N-terminal domain of NMDAR1 and human NMDAR1-Ab to construct the NMDARE model [37]. Here, hFOs on day 110 were exposed to the commercial monoclonal antibody targeting the N-terminal domain of NMDAR1 (Abcam, ab134308,10 μg/mL) or control IgG (CTRL-IgG) for 24 h, according to the concentration of anti-NMDAR1 IgG used in previous studies [8, 38]. The immunostaining analysis showed that NMDAR1-Ab exposure did not affect the total density of neurons (PSD95 + ) but led to an almost complete absence of cell surface NMDAR1 (NMDAR1 + ) in hFOs (Fig. 1d, e). To validate these findings in NMDARE patients, we collected CSF samples from 6 patients diagnosed with NMDARE and 6 non-affected controls (Supplementary Table 1). The cell-based assay confirmed the presence of anti-NMDAR1 IgG antibodies in all patient CSF samples (Methods and Supplementary Fig. 1). We then purified IgG antibodies from mixed CSF samples from the NMDARE patients (P-CSF-IgG) or non-affected controls (C-CSF-IgG). hFOs exposed to patient CSF-derived IgG antibodies also showed a nearly complete absence of cell surface NMDAR1 without changing the total density of neurons in hFOs (Fig. 1f, g). To confirm the apparent lack of cell surface NMDAR1 is not due to antibody competition in immunostaining, we directly incubate the sections of hFOs exposed to NMDAR1-Ab or control IgG with corresponding secondary antibodies without permeabilization. As shown in Supplementary Fig. 2, no fluorescence signal was detected in hFOs exposed to control IgG, and very few fluorescence signals for cell surface NMDAR1 were detected in hFOs exposed to either monoclonal NMDAR1-Ab or P-CSF-IgG. Notably, apparent fluorescence signals were detected in sections of hFOs exposed to either NMDAR1-Ab or P-CSF-IgG after permeabilization (Supplementary Fig. 2). These results indicated that the lack of cell surface NMDAR1 in immunostaining is not due to antibody competition but the consequence of antibody-mediated NMDAR1 internalization.
NMDAR1-Ab exposure in hFOs affects expression of genes enriched in synaptic transmission and glutamatergic pathways
We next performed RNA-seq analysis to investigate the functional impacts of NMDAR1-Ab exposure in hFOs. hFOs on day 110 were exposed to 10 μg/mL monoclonal NMDAR1-Ab or control IgG for 24 h, and were subsequently used for RNA-seq. Differential gene expression analysis between the NMDAR1-Ab-exposed and control hFOs identified a total of 2958 DEGs [False Discovery Rate (FDR) < 0.05]. Among the 2958 DEGs, 1646 genes were upregulated and 1312 genes were downregulated in NMDAR1-Ab-exposed hFOs (Fig. 2a and Supplementary data 1).
a Heatmap and volcano plots for RNA-seq data of NMDAR1-Ab exposure and control hFOs. Outlier samples were excluded based on the hierarchical cluster analysis. (b, c) GO and KEGG pathway analysis for upregulated (b) and downregulated (c) DEGs in NMDAR1-Ab-exposed hFOs, respectively. GO terms are shown with red bars, and KEGG pathways are shown with blue bars. d PPI network analysis for the downregulated DEGs in NMDAR1-Ab-exposed hFOs. The top 10 hub nodes with the most connections are highlighted in red. (e, f) RT-qPCR analysis for GRIN1 and GRIA2 (The hub nodes in the PPI network) in hFOs after exposure to monoclonal NMDAR1-Ab (e) or IgG purified from CSF of NMDARE patients (f). N ≥ 3 technical replicates per group. Data are shown as Mean ± SD. Two-tailed t-test, *P < 0.05, ***P < 0.001.
By performing functional annotation for the DEGs using WebGestalt [39], we found that the 1646 upregulated DEGs were enriched for biological functions including generation of precursor metabolites and energy, ribonucleoprotein complex biogenesis, and RNA splicing (FDR < 2.20E-16). The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis implicated the 1646 upregulated DEGs in pathways including thermogenesis, Huntington’s disease, Alzheimer’s disease, Parkinson’s disease, and oxidative phosphorylation (FDR < 2.20E-16) (Fig. 2b). For the 1312 downregulated DEGs, Gene Ontology (GO) enrichment analysis revealed their significant enrichment in axon development (FDR < 2.20E-16), regulation of trans-synaptic signaling (FDR < 2.20E-16), synaptic vesicle cycle (FDR = 4.20E-16), and calcium ion regulated exocytosis (FDR = 4.87E-13). The KEGG pathway analysis implicated the 1312 downregulated DEGs in nicotine addiction (FDR = 7.74E-10), morphine addiction (FDR = 9.52E-8), axon guidance (FDR = 1.53E-06), GABAergic synapse (FDR = 5.26E-06), and glutamatergic synapse (FDR = 6.60E-04) (Fig. 2c). These results highlighted that NMDAR1-Ab exposure downregulated sets of genes enriched in multiple neuronal functions, including the glutamatergic synapse.
We focused on the downregulated DEGs and used them to construct a PPI network based on the STRING database [40]. We then used CytoHubba [35] to explore key regulators in the PPI network that may mediate the effects of NMDAR1-Ab exposure. Nine of the twelve algorithms available in CytoHubba identified GRIN1 (NMDAR1) and GRIA2 as the top two hub nodes in the PPI network (Fig. 2d). To be noted, GRIN1 and GRIA2 are two genes encoding the NR1 subunit of NMDAR and AMPA-selective glutamate receptor 2 [41], respectively. Mutations in these two genes have been reported to be associated with neurodevelopmental disorders [42, 43]. Using RT-qPCR analysis, we confirmed the decreased expression of GRIN1 and GRIA2 in NMDAR1-Ab-exposed hFOs as compared to controls (Fig. 2e).
To validate the RNA-seq results, we treated the hFOs with IgG antibodies (10 μg/mL) purified from CSF of NMDARE patients or non-affected controls for 24 h. The RT-qPCR analysis showed that hFOs exposed to patient CSF-derived IgG antibodies also led to a decreased expression of GRIN1 and GRIA2 (Fig. 2f). These results together with the RNA-seq findings indicated that NMDAR1-Ab exposure functioned through affecting the glutamatergic synapse.
Genes affected by NMDAR1-Ab exposure are enriched for neuropsychiatric disorders
To explore the connection of NMDAR1-Ab exposure to human disease, we utilized the 1312 downregulated DEGs to perform the gene-disease association analysis using DisGeNET [44]. The association analysis displayed that the 1312 downregulated DEGs were significantly enriched with brain disorders including schizophrenia, absence epilepsy, autism spectrum disorders, intellectual disability, and bipolar disorder (FDR < 2.99E-05) (Supplementary Fig. 3 and Supplementary data 2). Notably, both GRIN1 and GRIA2 are associated with various neuropsychiatric disorders (Supplementary data 2). FUMA-GWAS [45] analysis also showed an enrichment of the 1312 downregulated DEGs with genes genetically associated with brain disorders, including schizophrenia (FDR = 3.42E-04), cognitive ability (FDR = 3.77E-04), epilepsy (FDR = 7.84E-03), and general factor of neuroticism (FDR = 9.37E-03) (Supplementary Fig. 3 and Supplementary data 2). These results implicated that NMDAR1-Ab exposure accounted for the neuropsychiatric symptoms presented in patients with NMDARE.
Metabolomic analysis confirms the decreased glutamate levels in NMDAR1-Ab-exposed hFOs
Since RNA-seq analysis revealed the effects of NMDAR1-Ab exposure on glutamatergic synapse, we performed metabolomic analysis to examine whether NMDAR1-Ab exposure decreased glutamate levels in hFOs. After exposure to monoclonal NMDAR1-Ab (10 μg/mL) or control IgG for 24 h, hFOs (day 110) were used for metabolomic analysis. Of the 206 metabolites detected in hFOs, a total of 12 metabolites were differentially expressed between the NMDAR1-Ab exposure and control hFOs (Fig. 3a and Supplementary data 3). Among the 12 differential metabolites, 4 metabolites were upregulated and 8 metabolites including L-glutamic acid were downregulated in NMDAR1-Ab-exposed hFOs, respectively. KEGG pathway analysis showed that these 12 differential metabolites were associated with pathways including ascorbate and aldarate metabolism, arginine biosynthesis, and glutathione metabolism (P < 0.05) (Fig. 3b). Notably, hFOs exposed to purified patient CSF-derived IgG antibodies also led to a reduction of L-glutamic acid content as measured by the enzyme-linked immunosorbent assay (ELISA) (Fig. 3c). These results suggested that NMDAR1-Ab exposure decreased cell surface NMDAR1 and led to the subsequent reduction of glutamate levels in hFOs.
a Scatter plot and volcano plot for the NMDAR1-Ab exposure and control hFOs. The orthogonal projections to latent structures-discriminant analysis (OPLS-DA) was used to cluster the hFOs and identify differentially expressed metabolites. VIP, variable importance in projection of the first principal component in the OPLS-DA model. b KEGG pathway analysis for the differentially expressed metabolites. c ELISA was used to measure glutamate levels in hFOs after exposure to IgG purified from CSF of NMDARE patients or non-affected controls. Data are shown as Mean ± SEM. N = 3 independent hFOs per group. Two-tailed t-test, *P < 0.05. d RT-qPCR analysis for GLUD1, GLUL, and GLS in hFOs after exposure to NMDAR1-Ab or control IgG. N ≥ 3 technical replicates per group. Data are shown as Mean ± SD. Two-tailed t-test, **P < 0.01; NS, non-significance.
To be noted, the RNA-seq and RT-qPCR analysis displayed that hFOs exposed to NMDAR1-Ab upregulated expression of GLUD1 and GLUL, two genes that encode the enzymes that respectively convert glutamate into α-ketoglutaric acid and glutamine. While the expression of GLS, a gene that encodes glutaminase to catalyze the production of glutamate, was unchanged upon NMDAR1-Ab exposure (Fig. 3d). These results suggested that enhanced expression of GLUD1 and GLUL may exacerbate the conversion of glutamate to downstream metabolites that caused a reduction of glutamate content after NMDAR1-Ab exposure in hFOs.
NMDAR1-Ab exposure leads to hypo neuroactivities in hFOs
The RNA-seq and metabolomic analyses indicated the inhibited glutamatergic function upon NMDAR1-Ab exposure, we used MEA assays to examine whether NMDAR1-Ab exposure affected neuroactivities. After exposure to monoclonal NMDAR1-Ab for 24 h, hFOs exhibited a decreased number of spikes and weighted mean firing rate when compared with controls (Fig. 4a). The hypo neuroactivities after NMDAR1-Ab exposure were further confirmed by the calcium imaging analysis (Fig. 4b). Intriguingly, the purified patient CSF-derived IgG antibodies also led to hypo neuroactivities in hFOs as detected by the MEA assays (Fig. 4c). Together, these results indicated that NMDAR1-Ab alone led to hypo neuroactivities rather than neuronal hyperexcitability observed in patients with NMDARE.
a MEA analysis for hFOs after exposure to monoclonal NMDAR1-Ab or control IgG. Data are shown as Mean ± SEM (N ≥ 3 hFOs per group). Number of spikes and weighted mean neuron firing rate at 0 h were normalized to 1. Two-way ANOVA, **P < 0.01, ****P < 0.0001. Scale bar, 200 μm. b Calcium imaging analysis for hFOs after exposure to monoclonal NMDAR1-Ab or control IgG. Regions of interest (ROIs), N = 24 from 3 independent hFOs per group. Data are shown as Mean ± SEM. Two-tailed t-test, ***P < 0.001. Scale bar, 100 μm. c MEA analysis for hFOs after exposure to IgG purified from CSF of NMDARE patients or non-affected controls. Number of spikes and weighted mean neuron firing rate at 0 h were normalized to 1. N ≥ 3 independent hFOs per group. Data are shown as Mean ± SEM. Two-way ANOVA, *P < 0.05, **P < 0.01; NS, non-significance. Scale bar, 200 μm.
hFOs exposed to either patient CSF or IgG-depleted patient CSF causes hyper neuroactivities
We next tested how patient CSF exposure could affect neuroactivities. To ensure hFOs were exposed to 10 μg/mL IgG antibodies, the CSF samples were diluted in hFOs culture medium according to their IgG concentrations determined by ELISA. hFOs (day 110) were then exposed to the diluted CSF from the NMDARE patients or non-affected controls for 24 h. The immunostaining analysis showed that patient CSF dramatically decreased the density of cell surface NMDAR1(Fig. 5a and Supplementary Fig. 4a) without changing the total density of neurons (PSD95 + ) in hFOs (Supplementary Fig. 4b). We then performed MEA and calcium imaging analyses, finding that hFOs exposed to the mixed patient CSF exhibited neuronal hyperexcitability when compared with non-affected controls (Fig. 5b). Given purified patient CSF-derived IgG antibodies led to hypo neuroactivities, these results indicated that the presence of substances in patient CSF rather than IgG antibodies accounted for the neuronal hyperexcitability. To confirm this, we removed IgG antibodies from the CSF and examined how IgG-depleted CSF supernatant affected neuroactivities. After confirming the complete removal of IgG from CSF samples by cell-based assay (Supplementary Fig. 5), hFOs were treated with IgG-depleted CSF supernatant for 24 h. As shown by the MEA assays, hFOs treated with IgG-depleted patient CSF exhibited neuronal hyperexcitability as compared to controls (Fig. 5c). Together, the above results verified that substance in patient CSF rather than NMDAR1-Ab accounted for the neuronal hyperexcitability.
a Density of PSD95 and cell surface NMDAR1 in hFOs after exposure to mixed CSF from NMDARE patients or non-affected controls. N ≥ 3 independent hFOs per group were used for the immunostaining analysis. Scale bar, 20 μm. b MEA and calcium imaging analysis for hFOs after exposure to mixed CSF from NMDARE patients or non-affected controls. Data are shown as Mean ± SEM. Number of spikes and weighted mean neuron firing rate at 0 h were normalized to 1. N ≥ 3 independent hFOs per group for MEA analysis. Two-way ANOVA for MEA analysis, *P < 0.05, **P < 0.01, ***P < 0.001; NS, non-significance. N = 24 ROIs from 3 independent hFOs in each group for calcium imaging analysis. Two-tailed t-test for calcium imaging analysis, ***P < 0.001; NS, non-significance. Scale bars, 200 μm and 100 μm. c MEA analysis for hFOs after exposure to mixed CSF from NMDARE patients or non-affected controls with IgG depletion. N ≥ 3 independent hFOs per group. Data are shown as Mean ± SEM. Number of spikes and weighted mean neuron firing rate at 0 h were normalized to 1. Two-way ANOVA, *P < 0.05. Scale bar, 200 μm.
Activated IL-17 signaling pathway in patient CSF accounts for the neuronal hyperexcitability
Since elevated brain inflammation usually emerges in NMDARE patients [18] and plays important roles in regulating neuroactivities [46], we proposed that elevated levels of proinflammatory factors in patient CSF took response for the neuronal hyperexcitability. We then performed Olink proteomic analysis using CSF samples from 6 NMDARE patients and 6 non-affected controls, which allowed sensitive high-throughput profiling of CSF proteins with low cross-reactivity and off-target [47]. Of the 92 proteins in the Olink® Target 96 inflammation panel, 14 differential proteins including the proinflammatory factors IL-6 and interferon-gamma (IFN-γ) were identified (Fig. 6a and Supplementary data 4). Notably, all the 14 differential proteins were upregulated in NMDARE patient CSF. The 14 upregulated proteins were enriched in positive regulation of response to external stimulus (FDR = 1.27E-04), T cell activation (FDR = 3.25E-04), and IL-17 signaling pathway (FDR = 1.86E-03) (Fig. 6b). Since IL-17 plays critical roles in driving inflammation [48], the activation of IL-17 signaling pathway may indicate inflammation in CSF of NMDARE patients. To be noted, a recent meta-analysis showed that the contents of proinflammatory factors including IL-17, IL-6, and IFN-γ were significantly higher in CSF from NMDARE patients than those in non-affected controls [23]. Given that IL-17 could induce the release of IL-6 and IFN-γ [49,50,51], activation of IL-17 signaling pathway may be the origin that caused neuronal hyperexcitability.
a Volcano plot for the differential proteins between CSF from NMDARE patients and non-affected controls. b GO and KEGG pathway analysis for the differential proteins. The top 10 terms are shown. c RT-qPCR analysis for ACT1 and TRAF6 after IL-17 exposure in hFOs. N ≥ 3 technical replicates per group. Data are shown as Mean ± SD. Two-tailed t-test, *P < 0.05, ***P < 0.001. d Calcium imaging analysis for hFOs after exposure to IL-17. N = 24 ROIs from 3 independent hFOs per group. Data are shown as Mean ± SEM. Two-tailed t-test, ***P < 0.001. Scale bar, 100 μm. e MEA analysis for hFOs (N ≥ 3 hFOs per group) after exposure to IL-17. f MEA analysis for hFOs (N ≥ 3 hFOs per group) after exposure to IgG-depleted control CSF, IgG-depleted patient CSF with or without IL-17 neutralizing antibody (AIN457). Isotype control IgG was used as control for AIN457. e, f Data are shown as Mean ± SEM. Number of spikes and weighted mean neuron firing rate at 0 h were normalized to 1. Two-way ANOVA, *P < 0.05; NS, non-significance. Scale bar, 200 μm.
To validate this hypothesis, we treated the hFOs with IL-17 for 24 h at 100 ng/mL, a concentration reported to induce neuronal hyperexcitability in mouse brain slice cultures [52]. RT-qPCR analysis confirmed the upregulation of two key activators of IL-17 signaling pathway (ACT1 and TRAF6) after IL-17 exposure in hFOs (Fig. 6c), indicating that the IL-17 signaling pathway was activated. The following MEA and calcium imaging analysis showed that hFOs exposed to IL-17 showed neuronal hyperexcitability when compared with controls (Fig. 6d, e). To further confirm the role of IL-17 in neuroactivities, hFOs were exposed to IgG-depleted patient CSF with or without anti-IL-17 monoclonal antibodies (AIN457, 100 μg) and IgG-depleted control CSF. The following MEA assay showed that neutralizing IL-17 alleviates neuronal hyperexcitability in hFOs exposed to IgG-depleted patient CSF when compared to controls (Fig. 6f). These results indicated that the activated IL-17 signaling pathway in patient CSF accounted for the neuronal hyperexcitability.
Neutralizing IL-17 alleviates seizure-like behaviors in mice exposed to CSF from NMDARE patients
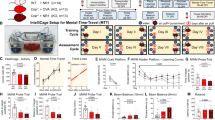
To further validate the effects of IL-17 signaling pathway on neuroactivities in vivo, we generated a mouse model intracerebroventricularly injected with CSF from NMDARE patients or non-affected controls using osmotic pumps (Methods and Fig. 7a). In the initial 7 days, mice injected with either control CSF or patient CSF did not develop any seizure-like behaviors. To induce a seizure-like phenotype in mice following the CSF injection, PTZ was intraperitoneally injected at a subthreshold dosage (40 mg/kg) on day 7. Under this PTZ dose, mice co-injected with PTZ and control CSF did not develop seizure-like phenotypes, while all of the mice co-injected with PTZ and CSF from NMDARE patients exhibited seizure-like behaviors in 5 min after PTZ injection (Supplementary Movie 1). These results suggested that mice exposed to CSF from NMDARE patients are more vulnerable to seizures than controls. Interestingly, the severity of seizure-like phenotypes indicated by scores of the Racine scale could be alleviated (lower Racine scores) in mice that were pretreated with anti-IL-17 monoclonal antibodies (AIN457, 100 μg) (Fig. 7b and Supplementary Movie 1). These results supported that the activated IL-17 signaling pathway accounted for the neuronal hyperexcitability.
a Schematic diagram for generating a mouse model of epileptic seizure. Created with BioRender.com. b score of the Racine scale (seizure stage) was used to indicate the severity of seizures. N ≥ 7 mice in each group. The Kruskal-Wallis test, *P < 0.05, ***P < 0.001; NS, non-significance.
Discussion
Discovered only a few years ago, NMDARE has emerged as one of the most common types of nonviral encephalitis. The lack of a clear understanding of the disease pathogenesis has led to challenges in precision diagnosis and treatment. The current therapeutic strategy for NMDARE relies on various forms of immunotherapy such as glucocorticoid therapy. However, this treatment lacks a precision target and may cause side effects. Patients with NMDARE usually take a long time to recover and may experience long-term sequelae and relapse after symptom onset [53, 54]. Discovering a newly effective therapeutic target will improve the treatment strategy and outcome.
By leveraging human brain organoids as a model, our study showed that NMDAR1-Ab exposure alone led to NMDAR hypofunction as indicated by the reduction of cell surface NMDAR1, GRIN1 expression, glutamate content, and synaptic transmission in hFOs. Given that activation of NMDAR requires extracellular glutamate, the reduction of GRIN1 expression and glutamate content after NMDAR1-Ab treatment implicates the existence of a feedback mechanism to maintain homeostasis in response to NMDAR hypofunction. The NMDAR hypofunction may account for the psychiatric symptoms in patients with NMDARE since expression of many neuropsychiatric disorder-related genes was altered after NMDAR1-Ab exposure. Though NMDAR1-Ab exposure did not cause a seizure-like electrophysiological phenotype in hFOs, expression of a large set of epilepsy-related genes was altered. These altered genes including STX1B, SCN1A, and RYR2 were significantly overlapped with those identified by the exome sequencing [55] and genome-wide association studies for epilepsy [56]. These results suggested that NMDAR1-Ab exposure may alter the threshold of epileptic seizures. Interestingly, as shown in several studies [16, 57], NMDAR1-Ab treatment in rats or mice caused NMDAR hypofunction but hyperexcitability of hippocampal circuits, highlighting the complexity of circuit disturbance that may lead to seizures. These studies may explain the relationship between NMDAR hypofunction and increased seizure susceptibility from the aspect of brain circuits. In addition to disturbed brain circuits, NMDAR1-Ab treatment was found to cause seizures and hippocampal inflammation in mice with increased levels of inflammatory markers [13], suggesting that activation of brain inflammation may also contribute to the seizure phenotype in NMDARE.
IL-17, a proinflammatory cytokine, was found to cause neuronal hyperexcitability in this study. A previous study also showed that IL-17 induced neuronal hyperexcitability in mouse brain slice cultures, and IL-17A receptor knock-out mice displayed significantly delayed seizure onset in a kainic acid-induced seizure model [52]. Our study also showed that neutralizing IL-17 alleviated seizure-like phenotypes in hFOs and a mouse model exposed to NMDARE CSF. Importantly, IL-17 levels in CSF were found to be higher in NMDARE patients in a meta-analysis [23]. Given higher levels of IL-17 in CSF were significantly correlated with poor prognosis and relapse of NMDARE patients [58, 59], evaluating IL-17 levels in CSF may benefit predicting the seizure probability and prognosis of the patients. More importantly, selectively blocking IL-17A receptor or neutralizing IL-17 to inactivate the IL-17 signaling pathway may be an effective therapeutic strategy for NMDARE.
IL-17 plays important roles in host defense response, inflammation modulation, and the maintenance of barrier integrity [60, 61]. IL-17 was shown to induce a disruption of the blood-brain barrier [62]. The activation of IL-17 signaling in CSF of NMDARE patients suggested a break of the blood-brain barrier. The loss of blood-brain barrier may facilitate the entry of NMDAR1-Ab or proinflammation factors such as IL-6 [63] into the brain microenvironment. Neutralizing IL-17 or its signaling has the potential to alleviate brain inflammation and psychiatric symptoms in NMDARE. In addition, we noticed that differential cytokines in CSF of NMDARE patients were enriched for T cell activation. Given that IL-17 was mainly produced from Th17 T cells, targeting the Th17 cell lineage may be an alternative solution for NMDARE treatment.
In summary, using hFOs-based multi-omics analyses, this study reveals the molecular mechanisms underlying NMDARE. Given the genetic variabilities across iPSC lines, we replicated key experiments in hFOs derived from other two normal iPSC lines (F1F and GD1405101) to improve the generalizability. Key findings including NMDAR internalization and hypofunction, and neuronal hypo activities caused by NMDAR1-Ab exposure were validated (Supplementary Fig. 6–8). The contribution of activated IL-17 signaling pathway to neuronal hyperexcitability was also validated in these two iPSC lines-derived hFOs (Supplementary Fig. 8).
These findings support the notion that the anti-NMDAR1 antibodies and IL-17 signaling pathway shape NMDARE. However, several limitations of this study should also be noted. First, electrophysiological experiments in this study are limited for in-depth analysis, more detailed electrophysiological recordings with respect to excitatory and inhibitory currents are needed to explain the origin of neuronal hyperexcitability caused by patient CSF or IL-17 treatment. Second, whether the activated IL-17 signaling pathway is the primary driver for neuronal hyperexcitability could not be determined since IL-6 and IFN-γ could both increase neuronal excitability [64, 65] and were also elevated in NMDARE patient CSF. The third, though CSF IL-17 levels can potentially predict the seizure probability and prognosis of NMDARE patients, more clinical details and correlations are needed to assess the accuracy. Moreover, whether IL-17 neutralization improves other NMDARE symptoms such as cognitive decline and behavioral abnormalities needs further investigations. The fourth, Olink proteomic analysis in this study only covered a panel that included 92 inflammation-associated proteins, other molecules that contribute to disease phenotypes may not be discovered. On the other hand, the sample size used for Olink proteomic analysis is small, more CSF samples could be collected for further study to strengthen our findings and discover more disease-associated molecules. Last, though major brain cell types existed in hFOs, the lack of microglia makes hFOs limited to fully resemble the immune response system in the human brain. Microglia-containing brain organoids or a brain organoid co-culture system containing T cells and B cells could be developed as more complex models to better mimic NMDARE immune responses in the future.
Data availability
RNA-seq data generated in this study have been deposited in the Gene Expression Omnibus under accession code GSE256148. Because the raw data of the MEA, metabolomic, and proteomic data are huge and presented in diverse native formats, these raw data are available on request from the corresponding authors. All the other data associated with this study are shown in the manuscript and supplementary materials.
References
Dalmau J, Tüzün E, Wu H-y, Masjuan J, Rossi JE, Voloschin A, et al. Paraneoplastic anti–N-methyl-D-aspartate receptor encephalitis associated with ovarian teratoma. Ann Neurol. 2007;61:25–36.
Dalmau J, Gleichman AJ, Hughes EG, Rossi JE, Peng X, Lai M, et al. Anti-NMDA-receptor encephalitis: case series and analysis of the effects of antibodies. Lancet Neurol. 2008;7:1091–8.
Irani SR, Bera K, Waters P, Zuliani L, Maxwell S, Zandi MS, et al. N-methyl-D-aspartate antibody encephalitis: temporal progression of clinical and paraclinical observations in a predominantly non-paraneoplastic disorder of both sexes. Brain. 2010;133:1655–67.
Finke C, Kopp UA, Prüss H, Dalmau J, Wandinger KP, Ploner CJ. Cognitive deficits following anti-NMDA receptor encephalitis. J Neurol Neurosurg Psychiatry. 2012;83:195–8.
Wagnon I, Hélie P, Bardou I, Regnauld C, Lesec L, Leprince J, et al. Autoimmune encephalitis mediated by B-cell response against N-methyl-d-aspartate receptor. Brain. 2020;143:2957–72.
Hughes EG, Peng X, Gleichman AJ, Lai M, Zhou L, Tsou R, et al. Cellular and synaptic mechanisms of anti-NMDA receptor encephalitis. J Neurosci. 2010;30:5866–75.
Castillo-Gómez E, Oliveira B, Tapken D, Bertrand S, Klein-Schmidt C, Pan H, et al. All naturally occurring autoantibodies against the NMDA receptor subunit NR1 have pathogenic potential irrespective of epitope and immunoglobulin class. Mol Psychiatry. 2017;22:1776–84.
Wenke NK, Kreye J, Andrzejak E, van Casteren A, Leubner J, Murgueitio MS, et al. N-methyl-D-aspartate receptor dysfunction by unmutated human antibodies against the NR1 subunit. Ann Neurol. 2019;85:771–6.
Moscato EH, Peng X, Jain A, Parsons TD, Dalmau J, Balice-Gordon RJ. Acute mechanisms underlying antibody effects in anti–N-methyl-D-aspartate receptor encephalitis. Ann Neurol. 2014;76:108–19.
Planagumà J, Leypoldt F, Mannara F, Gutiérrez-Cuesta J, Martín-García E, Aguilar E, et al. Human N-methyl D-aspartate receptor antibodies alter memory and behaviour in mice. Brain. 2015;138:94–109.
Jones BE, Tovar KR, Goehring A, Jalali-Yazdi F, Okada NJ, Gouaux E, et al. Autoimmune receptor encephalitis in mice induced by active immunization with conformationally stabilized holoreceptors. Sci Transl Med. 2019;11:eaaw0044.
Ding Y, Zhou Z, Chen J, Peng Y, Wang H, Qiu W, et al. Anti-NMDAR encephalitis induced in mice by active immunization with a peptide from the amino-terminal domain of the GluN1 subunit. J Neuroinflammation. 2021;18:53.
Taraschenko O, Fox HS, Zekeridou A, Pittock SJ, Eldridge E, Farukhuddin F, et al. Seizures and memory impairment induced by patient-derived anti-N-methyl-D-aspartate receptor antibodies in mice are attenuated by anakinra, an interleukin-1 receptor antagonist. Epilepsia. 2021;62:671–82.
Taraschenko O, Fox HS, Pittock SJ, Zekeridou A, Gafurova M, Eldridge E, et al. A mouse model of seizures in anti–N-methyl-d-aspartate receptor encephalitis. Epilepsia. 2019;60:452–63.
Taraschenko O, Fox HS, Eldridge E, Wang W, Dowd SW, Al-Saleem F, et al. Monoclonal antibodies from anti-nmda receptor encephalitis patient as a tool to study autoimmune seizures. Front Neurosci. 2021;15:710650.
Wright SK, Rosch RE, Wilson MA, Upadhya MA, Dhangar DR, Clarke-Bland C, et al. Multimodal electrophysiological analyses reveal that reduced synaptic excitatory neurotransmission underlies seizures in a model of NMDAR antibody-mediated encephalitis. Commun Biol. 2021;4:1106.
Wang X, Ma C, Liu CY, Li GJ, Zhao D, Han DF. Neuronal NMDAR Currents of the hippocampus and learning performance in autoimmune anti-NMDAR encephalitis and Involvement of TNF-α and IL-6. Front Neurol. 2019;10:684.
Liba Z, Kayserova J, Elisak M, Marusic P, Nohejlova H, Hanzalova J, et al. Anti-N-methyl-D-aspartate receptor encephalitis: the clinical course in light of the chemokine and cytokine levels in cerebrospinal fluid. J Neuroinflammation. 2016;13:55.
Kothur K, Wienholt L, Mohammad SS, Tantsis EM, Pillai S, Britton PN, et al. Utility of CSF cytokine/chemokines as markers of active intrathecal inflammation: comparison of demyelinating, anti-NMDAR and enteroviral encephalitis. PLoS One. 2016;11:e0161656.
Ai P, Zhang X, Xie Z, Liu G, Liu X, Pan S, et al. The HMGB1 is increased in CSF of patients with an Anti-NMDAR encephalitis. Acta Neurol Scand. 2018;137:277–82.
Ding Y, Yang C, Zhou Z, Peng Y, Chen J, Pan S, et al. Clinical significance of soluble adhesion molecules in anti-NMDAR encephalitis patients. Ann Clin Transl Neurol. 2019;6:945–53.
Li Q, Chen J, Yin M, Zhao J, Lu F, Wang Z, et al. High Level of Soluble CD146 in cerebrospinal fluid might be a biomarker of severity of Anti-N-Methyl-D-Aspartate receptor encephalitis. Front Immunol. 2021;12:680424.
Ma Y, Wang J, Guo S, Meng Z, Ren Y, Xie Y, et al. Cytokine/chemokine levels in the CSF and serum of anti-NMDAR encephalitis: a systematic review and meta-analysis. Front Immunol. 2022;13:1064007.
Wilke JBH, Hindermann M, Berghoff SA, Zihsler S, Arinrad S, Ronnenberg A, et al. Autoantibodies against NMDA receptor 1 modify rather than cause encephalitis. Mol Psychiatry. 2021;26:7746–59.
Pan H, Oliveira B, Saher G, Dere E, Tapken D, Mitjans M, et al. Uncoupling the widespread occurrence of anti-NMDAR1 autoantibodies from neuropsychiatric disease in a novel autoimmune model. Mol Psychiatry. 2019;24:1489–501.
Arinrad S, Wilke JBH, Seelbach A, Doeren J, Hindermann M, Butt UJ, et al. NMDAR1 autoantibodies amplify behavioral phenotypes of genetic white matter inflammation: a mild encephalitis model with neuropsychiatric relevance. Mol Psychiatry. 2022;27:4974–83.
Matute C, Palma A, Serrano-Regal MP, Maudes E, Barman S, Sánchez-Gómez MV, et al. N-Methyl-D-Aspartate receptor antibodies in autoimmune encephalopathy alter oligodendrocyte function. Ann Neurol. 2020;87:670–6.
Ismail FS, Faustmann PM. Astrocytes and their potential role in anti-NMDA receptor encephalitis. Med Hypotheses. 2020;139:109612.
Giannoccaro MP, Wright SK, Vincent A. In vivo mechanisms of antibody-mediated neurological disorders: animal models and potential implications. Front Neurol. 2019;10:1394.
Lancaster MA, Renner M, Martin CA, Wenzel D, Bicknell LS, Hurles ME, et al. Cerebral organoids model human brain development and microcephaly. Nature. 2013;501:373–9.
Uzquiano A, Kedaigle AJ, Pigoni M, Paulsen B, Adiconis X, Kim K, et al. Proper acquisition of cell class identity in organoids allows definition of fate specification programs of the human cerebral cortex. Cell. 2022;185:3770–88.e27.
Rybak-Wolf A, Wyler E, Pentimalli TM, Legnini I, Oliveras Martinez A, Glažar P, et al. Modelling viral encephalitis caused by herpes simplex virus 1 infection in cerebral organoids. Nat Microbiol. 2023;8:1252–66.
Graus F, Titulaer MJ, Balu R, Benseler S, Bien CG, Cellucci T, et al. A clinical approach to diagnosis of autoimmune encephalitis. Lancet Neurol. 2016;15:391–404.
Meng Q, Zhang W, Wang X, Jiao C, Xu S, Liu C, et al. Human forebrain organoids reveal connections between valproic acid exposure and autism risk. Transl Psychiatry. 2022;12:130.
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8:S11.
Lüttjohann A, Fabene PF, van Luijtelaar G. A revised Racine’s scale for PTZ-induced seizures in rats. Physiol Behav. 2009;98:579–86.
Taraschenko O, Fox HS, Pittock SJ, Zekeridou A, Gafurova M, Eldridge E, et al. A mouse model of seizures in anti-N-methyl-d-aspartate receptor encephalitis. Epilepsia. 2019;60:452–63.
Kreye J, Wenke NK, Chayka M, Leubner J, Murugan R, Maier N, et al. Human cerebrospinal fluid monoclonal N-methyl-D-aspartate receptor autoantibodies are sufficient for encephalitis pathogenesis. Brain. 2016;139:2641–52.
Zhang B, Kirov S, Snoddy J. WebGestalt: an integrated system for exploring gene sets in various biological contexts. Nucleic Acids Res. 2005;33:W741–8.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47:D607–d13.
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev. 2010;62:405–96.
Chen W, Shieh C, Swanger SA, Tankovic A, Au M, McGuire M, et al. GRIN1 mutation associated with intellectual disability alters NMDA receptor trafficking and function. J Hum Genet. 2017;62:589–97.
Salpietro V, Dixon CL, Guo H, Bello OD, Vandrovcova J, Efthymiou S, et al. AMPA receptor GluA2 subunit defects are a cause of neurodevelopmental disorders. Nat Commun. 2019;10:3094.
Piñero J, Ramírez-Anguita JM, Saüch-Pitarch J, Ronzano F, Centeno E, Sanz F, et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 2019;48:D845–55.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Vezzani A, Balosso S, Ravizza T. Neuroinflammatory pathways as treatment targets and biomarkers in epilepsy. Nat Rev Neurol. 2019;15:459–72.
Zhong W, Edfors F, Gummesson A, Bergström G, Fagerberg L, Uhlén M. Next generation plasma proteome profiling to monitor health and disease. Nat Commun. 2021;12:2493.
Li X, Bechara R, Zhao J, McGeachy MJ, Gaffen SL. IL-17 receptor-based signaling and implications for disease. Nat Immunol. 2019;20:1594–602.
Fossiez F, Djossou O, Chomarat P, Flores-Romo L, Ait-Yahia S, Maat C, et al. T cell interleukin-17 induces stromal cells to produce proinflammatory and hematopoietic cytokines. J Exp Med. 1996;183:2593–603.
Li L, Huang L, Vergis AL, Ye H, Bajwa A, Narayan V, et al. IL-17 produced by neutrophils regulates IFN-gamma-mediated neutrophil migration in mouse kidney ischemia-reperfusion injury. J Clin Invest. 2010;120:331–42.
Hwang S-Y, Kim J-Y, Kim K-W, Park M-K, Moon Y, Kim W-U, et al. IL-17 induces production of IL-6 and IL-8 in rheumatoid arthritis synovial fibroblasts via NF-κB- and PI3-kinase/Akt-dependent pathways. Arthritis Res Ther. 2004;6:R120.
Xu D, Robinson AP, Ishii T, Duncan DS, Alden TD, Goings GE, et al. Peripherally derived T regulatory and γδ T cells have opposing roles in the pathogenesis of intractable pediatric epilepsy. J Exp Med. 2018;215:1169–86.
Heine J, Kopp UA, Klag J, Ploner CJ, Prüss H, Finke C. Long-term cognitive outcome in Anti-N-Methyl-D-Aspartate Receptor Encephalitis. Ann Neurol. 2021;90:949–61.
Gong X, Chen C, Liu X, Lin J, Li A, Guo K, et al. Long-term functional outcomes and relapse of Anti-NMDA receptor encephalitis: a cohort study in western China. Neurol Neuroimmunol Neuroinflamm. 2021;8:D845–55.
Allen AS, Berkovic SF, Cossette P, Delanty N, Dlugos D, Eichler EE, et al. De novo mutations in epileptic encephalopathies. Nature. 2013;501:217–21.
Stevelink R, Campbell C, Chen S, Abou-Khalil B, Adesoji OM, Afawi Z, et al. GWAS meta-analysis of over 29,000 people with epilepsy identifies 26 risk loci and subtype-specific genetic architecture. Nat Genet. 2023;55:1471–82.
Ceanga M, Rahmati V, Haselmann H, Schmidl L, Hunter D, Brauer AK, et al. Human NMDAR autoantibodies disrupt excitatory-inhibitory balance, leading to hippocampal network hypersynchrony. Cell Rep. 2023;42:113166.
Peng Y, Liu B, Pei S, Zheng D, Wang Z, Ji T, et al. Higher CSF Levels of NLRP3 inflammasome is associated with poor prognosis of Anti-N-methyl-D-Aspartate receptor encephalitis. Front Immunol. 2019;10:905.
Zeng C, Chen L, Chen B, Cai Y, Li P, Yan L, et al. Th17 cells were recruited and accumulated in the cerebrospinal fluid and correlated with the poor prognosis of anti-NMDAR encephalitis. Acta Biochim Biophys Sin. 2018;50:1266–73.
Huangfu L, Li R, Huang Y, Wang S. The IL-17 family in diseases: from bench to bedside. Signal Transduct Target Ther. 2023;8:402.
Wang C, Zhang C-J, Martin BN, Bulek K, Kang Z, Zhao J, et al. IL-17 induced NOTCH1 activation in oligodendrocyte progenitor cells enhances proliferation and inflammatory gene expression. Nat Commun. 2017;8:15508.
Kebir H, Kreymborg K, Ifergan I, Dodelet-Devillers A, Cayrol R, Bernard M, et al. Human TH17 lymphocytes promote blood-brain barrier disruption and central nervous system inflammation. Nat Med. 2007;13:1173–5.
Veldhoen M. Interleukin 17 is a chief orchestrator of immunity. Nat Immunol. 2017;18:612–21.
Zhu PingJ, Huang W, Kalikulov D, Yoo JongW, Placzek AndonN, Stoica L, et al. Suppression of PKR Promotes Network Excitability and Enhanced Cognition by Interferon-γ-Mediated Disinhibition. Cell. 2011;147:1384–96.
Vezzani A, Viviani B. Neuromodulatory properties of inflammatory cytokines and their impact on neuronal excitability. Neuropharmacology. 2015;96:70–82.
Acknowledgements
We thank professor Ewen Tu for providing support and guidance for this study. This work was supported by grants from the National Natural Science Foundation of China (Nos. 82171504 and 81901359), the Natural Science Foundation of Hunan Province (No. 2022JJ20035), and the Science and Technology Innovation Program of Hunan Province (No. 2022RC1214).
Author information
Authors and Affiliations
Contributions
QM, BT, LZ, and JJ initiated the study and supervised the overall procedures. YUL and XW helped for the study design and data analysis. ZX, JW, and HMW did the cell and organoid culture, as well as the ELISA and MEA assays. HY performed the differential gene expression analysis. ZX, BW, and HY performed the metabolomic and Olink proteomic analysis. ZX and HY wrote the manuscript, HW and YX helped to revise the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All animal experiments were conducted in accordance with the Regulations for the Administration of Affairs Concerning Experimental Animals (China) and were approved by the Animal Care and Use Committee of the University of South China. All the participants signed the informed consent documents. Experimental procedures associated with human-derived samples were approved by the ethics committee of the First Affiliated Hospital of University of South China.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, Z., Yan, H., Wang, B. et al. The anti-NMDAR1 antibodies and IL-17 signaling pathway shape NMDAR encephalitis. Mol Psychiatry (2025). https://doi.org/10.1038/s41380-025-03434-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-025-03434-x