Abstract
Executive functions (EFs) are meta-cognitive abilities that orchestrate goal-directed behavior (i.e., set-shifting, working memory, and inhibitory control). Despite their strong genetic composition, the development of EFs is shaped by environmental exposures – as maternal distress, perinatal hypoxia, household dysfunction, neglection - which have varying degrees of impact depending on the duration or severity of the exposure, sensitive neurodevelopmental periods, and individual resiliency. Furthermore, they are negatively affected by aging and psychiatric, neurologic, and inflammatory diseases. MicroRNA dysregulation interferes with normal brain development and function and has been associated with various neuropsychiatric disorders; however, its effects on EFs remain unclear. Therefore, in this review we focus on the evidence regarding microRNA changes and their effects on EFs. We performed a systematic search from inception until October 2023 of four databases of human and animal studies. The results are presented narratively. Moreover, we conducted a bioinformatics analysis using experimental mRNA targets of the candidate microRNAs, as well as assessed the risk of bias of the included studies. We found 46 studies (23 in humans, 22 in animals, and one in both). The studies evaluated mild cognitive impairment, psychiatric disorders, and healthy aging. Gene mutations in MIR137 were associated with decreased EF performance, whereas MIR885 gene methylation was associated with increased executive functioning. Mutations in the genes of enzymes relevant for microRNA biosynthesis also impacted EFs. In the revised literature, the microRNAs that were consistently reported as dysregulated in two or more samples in relation to variations in EFs were miR-148a-3p for humans; miR-155, miR-30e, and miR-384-5p for rodents; and miR-132, miR-146a-5p, miR-148-3p, miR-181a-5p, miR-190b, miR-31, miR-501-3p, and miR-9-5p for both humans and rodents. The suggested regulatory pathways behind EFs included changes in neurogenesis, neurodevelopment, and synaptic plasticity/signaling. Changes in the various steps of microRNA biogenesis - such as mutations in the genes coding for microRNAs, reduced availability of relevant processing enzymes, or dysregulation of microRNA expression - are potentially associated with changes in EFs. Future research is required to better understand these associations in relation to developmental stage, diagnosis, disease severity, and the degree of the microRNA dysregulation.
Similar content being viewed by others
Introduction
Executive functions (EFs) are high-level cognitive processing skills that are relevant for self-regulation and goal-directed behaviors. [1] They support task initiation, performance, persistence, and completion and are therefore indispensable for occupational activities. [2] Some core abilities comprised under the umbrella term EFs are set-shifting (i.e., alternating between tasks), working memory (i.e., holding on to relevant information for relatively short periods of time), and inhibitory control (i.e., preventing the manifestation of an inappropriate cognitive process or behavior). [3] Although these abilities are separately identifiable, they share a top-level latent component named the common EF factor that explains most of the variability across them. [1]
EFs are presumed to be the result of the dynamic interplay between inherited factors and environmental influences, although they are also subject to independent influences of genes and environment separately. [4] The genetic architecture behind EFs is responsible for their high heritability and has been extensively studied. For example, twin studies estimated the heritability in children and adolescents to be as high as 0.77–1.00, whereby the latter value was obtained when researchers used the common EF factor. [5] Executive functioning involves activation of different neurotransmitters and circuits, including the serotoninergic, dopaminergic, and cholinergic systems, and changes in the genes related to the biosynthesis or transportation of these molecules are related to impairment of EFs. [6, 7] Recently, Hatoum et al. conducted the largest genome-wide association study (GWAS) on EFs to date. The study, which included more than 400 000 individuals in the UK biobank, found that genes related to inhibition (i.e., GABAergic processes) mediated EFs and also identified opposing genetic correlations between EFs and psychiatric disorders. [8]
Environmental adversities throughout the neurodevelopmental landscape contribute further to the shaping of EF. [9] The consequences of these exposures depend on various factors as their duration or severity, sensitive neurodevelopmental periods, and individual resiliency. Some examples include prenatal exposure to maternal distress, substance use or psychiatric disorders, perinatal hypoxia-ischemia, and adverse childhood experiences (ACEs). It is known that late childhood and adolescence are critical periods for the development of EFs, [10] therefore ACEs as physical abuse, sexual abuse, emotional or physical neglect during these periods of life can negatively impact EFs (mostly overall measures or working memory) into early and middle adulthood, and in the absence or presence of additional psychopathology (i.e., patients with bipolar disorder or schizophrenia). [9, 11] In adulthood, aging has mostly negative effects on EFs, although inhibitory control may increase with age. [12] EFs are also affected by chronic diseases, including cardiovascular and metabolic conditions. [13] Importantly, lower EF scores are found in most psychiatric diseases, including major depressive disorder (MDD), [14] bipolar disorder, [15] schizophrenia, [16] and post-traumatic stress disorder. [17] EFs are moldable skills and susceptible to improvement by cognitive remediation interventions, which are additionally effective for negative symptoms and overall functioning in patients with schizophrenia, in whom greater symptom severity and later onset-age do not prevent gains. [18, 19] Furthermore, meta-analyses showed that cognitive remediation is effective for improving EFs in adults with MDD, [20] and reducing symptoms severity and enhancing working memory in patients with anxiety disorders. [21] In summary, EFs are affected by genetic and environmental factors and are highly plastic and sensitive to rehabilitation.
Epigenetic modifications have a profound impact on the genome and increase or decrease the expression of genes. [22] Examples of such modifications include DNA methylation, histone modification, and regulation by non-coding RNAs, which include long non-coding RNAs and microRNAs. MicroRNAs are short non-coding sequences of around 22 nucleotides that were originally discovered in the genome of C. elegans three decades ago [23]; they participate in the silencing of mRNAs [24] and are assumed to be able to directly or indirectly modulate the expression of around a third of protein-coding genes. [25] The biosynthesis of microRNAs starts with the transcription of the corresponding gene. This primary microRNA is then processed by the nuclear enzymes DGCR8 and Drosha (which together form the microprocessor complex) to produce a precursor microRNA. The resulting sequence is translocated to the cytoplasm by exportin 5 and then further processed by the cytoplasmic enzyme DICER and by AGO proteins. The mature microRNA can exert effects in the cell or surrounding cells and can also reach distant tissues via the peripheral circulation, and the main overall effect of microRNAs is the regulation of gene transcription (Fig. 1A) [26,27,28,29]. Alterations in any of the relevant steps of microRNA biosynthesis can alter microRNA expression. In addition, microRNAs can target hundreds to thousands of mRNAs, being capable of regulating multiple genes and different individual biological mechanisms simultaneously leading to high functional pleiotropy of their effects, especially in complex systems as the nervous system. [30]
A Canonical pathway of microRNA biosynthesis. B Summary of the findings according to each step of the synthesis process. SNP, single nucleotide polymorphism; VNTR, variable number tandem repeat. Created in BioRender. Navarro Flores, A. (2026) https://BioRender.com/ztbg5cd. Partially adapted from Huang, E. [29], O’Carroll et al. [28], and Maffioletti et al. [27,28,29].
MicroRNAs are enriched in the brain in a tissue-specific manner. [31] They are expressed in various areas relevant for executive functioning, such as the frontal cortex and hippocampus, and are involved in crucial processes, such as neural genesis and differentiation, and synapse formation [32]. Therefore, dysregulation of microRNA expression due to changes in their biosynthesis or a mutation in the responsible gene can potentially induce various brain-related disorders [33]. For example, deletion of the region 22q11.2, which is critical for expression of the DiGeorge complex protein (a protein responsible for maturation of the microRNA), produces DiGeorge Syndrome; individuals with this deletion present with neurodevelopmental difficulties (learning disabilities) and a high prevalence of schizophrenia (25%) and other psychiatric disorders [34]. This example shows the importance of proper microRNAome expression for mental health. Moreover, many studies have reported dysregulated expression of microRNAs as potential biomarkers for certain psychiatric disorders [35,36,37], neurodegenerative disorders [33, 38], and chronic inflammatory conditions [39]. Establishing microRNAs as biomarkers for EFs would also be advantageous because microRNAs found peripherally (i.e., in blood samples) are informative of brain processes, highly stable in cell-free environments, and resistant to thaw-freeze cycles and could represent targets for future RNA-based therapies [40,41,42].
EFs are fundamental for regulating behavior, cognition, and mental health [43]. They decline not only in patients with psychiatric and neurologic disorders, but also in patients with chronic inflammatory diseases and during normal aging [44, 45]; while disturbances of EFs produce significant burden and impairment in activities of daily living and quality of life [46]. Understanding the role of microRNA expression represents an opportunity to better comprehend the epigenetic mechanisms involved. Furthermore, it may indicate targets that—with further research—could generate novel noninvasive biomarkers and potential therapeutic options. Here, we synthesize the evidence from published studies on the association between microRNA dysregulation and EFs in both human cohorts and animal models.
Methods
Literature search
This comprehensive literature review was performed according to the PRISMA guidelines [47]; a checklist of the items used is presented in the Supplement (eTable 1).
We searched the databases PubMed, WoS, and Scopus from inception until October 2023 by using the key terms “executive functions,” “working memory,” “Trail Making Test,” “microRNA,” and “miRNA” with Boolean connectors. The full search terms are presented in the Supplement (eTable 2). The inclusion criteria were as follows: 1) original article on observational (e.g., cross-sectional, case control, or cohort) or experimental study in humans or animals; 2) assessed EFs as a group of abilities or as individual skills such as working memory, set-shifting, and inhibitory control; and 3) assessed microRNA-related biology (e.g., microRNA expression, enzymes involved in microRNA synthesis, genes that regulate microRNA expression) and calculated correlations with executive functioning (i.e., explicitly stated that EFs were studied in general or named any individual EFs, including but not limited to working memory, set shifting, and inhibitory control). We excluded conference abstracts, studies on humans with cancer or animal models of cancer, studies in which the outcome of the microRNA analysis was not related to EFs (e.g., microRNA dysregulation in depression), and studies that analyzed only cognitive measures that were not EFs. No restrictions were placed on language, participant age, or publication time. One researcher (AN) screened the title and abstracts of the articles; in articles that appeared to be eligible for inclusion, this author read the full text, revised the inclusion criteria, and determined the final decision of inclusion. A second reviewer (UH) intervened in case of queries.
Quality assessment
The quality of the included human and animal studies was evaluated separately. It was assessed by one author (AN), and discrepancies were solved by a second author (UH). For the human studies, we used the New Castle Ottawa tool [48] to evaluate case-control and cohort studies and adapted the tool for use with cross-sectional studies. For randomized controlled trials, we used the revised Cochrane risk-of-bias tool (RoB 2.0) [49]. The animal studies were assessed with the SYRCLE RoB tool [50], and the criteria were adapted to our research question in a similar way as used previously [51].The criteria are described in detail in the Supplement (eTables 3–6).
Synthesis of the evidence
The results are summarized as a narrative review, and the information on the human and animal studies is presented separately, both in the text and in the tables. Figures were created using Biorender [52].
Bioinformatics analysis
MicroRNAs were included if they were reported as being significantly dysregulated, the dysregulation correlated with impairment of EFs, and if they were found in experimental studies. We analyzed the human- and animal-derived candidates separately. Bioinformatics included all miRNAs independent of their brain expression status, [53] as peripheral miRNAs can influence brain function via direct crossing of the blood-brain-barrier or transport in extracellular vesicles. [40, 54]
The following four groups were created by using the direction of the microRNA regulation (i.e., up or down) and the associated effect on EFs (i.e., increased or decreased): Group 1, microRNA upregulated and EFs increased; Group 2, microRNA upregulated and EFs decreased; Group 3, microRNA downregulated and EFs increased; and Group 4, microRNA downregulated and EFs decreased.
Inclusion criteria
Context and tissue specificity
When analyzed in similar tissues and stages of development, the expression and biological effects of many microRNAs are conserved across mammals [55]; however, although rare, major differences can occur in expression and directionality of effect between species. [56] These differences could reflect species-specific microRNA profiles, especially in brain tissue and in certain environmental situations or disease models, and consequently warrant further investigation. [56, 57] Therefore, we analyzed human and animal studies separately and tolerated contradictory results between these two sample types.
Functional consistency
Functional consistency was the inclusion criterion for the studies that reported microRNA results within the same sample type (i.e., human or rodents). MicroRNA functional consistency is present when similar regulatory effects (in this case, increased or decreased EFs) are associated with regulation in the same direction (i.e., up or down). Consistency was also accepted when a correlation was reported in both logical directions, e.g., when upregulation of one microRNA was associated with higher EF performance and downregulation of the same microRNA was associated with lower EF performance. [58, 59] Moreover, if a correlation was found in one direction but not in the other (e.g., upregulation was associated with higher EF performance, but downregulation was not associated with changes in EFs) it was not assumed to inherently reflect inconsistency because unidirectional effects can be observed when functional threshold effects, [60, 61] molecular titration, [62] or redundant or compensatory roles of other regulatory molecules come into play. [63]
MicroRNAs that did not fulfill the above criteria within a sample type were considered inconsistent and excluded from the bioinformatics analysis. MicroRNAs that were reported as not having any human homology or were not annotated in the tool, as well as those from studies with models of alcohol abuse or negative behavioral results were also not included.
Network and gene ontology analysis
The targets of these microRNAs (coding genes) were identified using an in-house tool (created by DK & AF), as previously published [64]. The tool integrates exclusively experimentally validated targets collected from six different databases (NPInter [65], TarBase [66], TransmiR [67], RegNetwork [68], Rise [69], and STRING [70]). From those we selected the mRNA targets that were enriched in brain tissue (expression two-fold higher in brain) using the RNA Atlas [71], and gene ontology analyses were performed acknowledging the probabilistic nature of predictions. We included the biological processes section of the enrichGO function, by using a gene set enrichment analysis with false discovery rate correction (clusterProfiler package [72], R v.4.4.1 [73]).
A network analysis of these targets was performed with Cytoscape V.3.7.2. [74] We ranked nodes by degree centrality and, reported the top 10 hubs per group as an a priori decision to provide a concise yet representative subset of the most connected genes as an overview for biological interpretation.
Results
Overview of the literature search results
A total of 590 records were retrieved, 102 were further full-text assessed, and finally 46 studies were included. From them, 23 were studies only in humans, [75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97] 22 were studies only in animals, [98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119] and one was a study in both humans and animals [40]. Only animal studies in rodents were found, therefore the terms will be used interchangeably. A detailed description of the included human and animal studies is presented in Table 1 and Table 2, respectively. The selection process is summarized in eFigure 1, and a list of the excluded studies, together with the reasons for exclusion, is presented in eTable 7.
The included studies evaluated various disruptions along the microRNA biosynthesis process and their subsequent effects on EFs. Some studies used more than one method. Alterations in the genes coding for microRNAs were reported in nine human studies [77, 80, 82, 83, 86, 87, 91, 93, 96] (including gene mutations, variable number tandem repeats [VNTRs], and gene methylation) and in two animal studies (one overexpression [109] and one knockout [KO] transgenic mouse model [105]). The way in which reduced availability of the nuclear enzyme DGCR8 affects working memory was studied in four mice models of the 22q11.2 deletion syndrome (22q11.2 DS) [107, 113, 114, 118]; the manipulation of the cytoplasmic enzyme DICER was studied in a KO mouse model [106]; the expression of microRNAs was manipulated in 11 animal studies [40, 99,100,101, 103, 108,109,110,111,112, 115, 116] (by injection of mimics or microRNA inhibitors directly into the animal, e.g., by intracerebroventricular injection or peritoneal injection); and the differential expression of microRNAs was studied in 15 human studies [75, 76, 78, 79, 81, 84, 85, 88,89,90, 92, 94, 95, 97] (14 in blood, and one in saliva) and six animal studies [40, 98, 102, 104, 117, 119] (one in blood, one in the brainstem, three in the hippocampus, and three in the prefrontal cortex). These processes and the related findings are presented in Fig. 1.
Various animal models were included, and the models also used genetic engineering, electrophysiological measures, and immunohistochemistry methods. The most frequently used methods of analysis were GWASs (humans), and quantitative polymerase chain reaction (qPCR) and next-generation sequencing (both humans and animals). A summary of the common pipelines is presented in Fig. 2.
The methods used to assess the role of microRNA in executive functions (EFs) differ in various aspects. The most common were: (a) Regarding the selection or induction of the underlying pathology or age group being studied: a.1) In animal models: no genetic manipulation (examples are natural aging by using older animals or other behaviorally induced models, such as early life stress or social defeat); use of transgenic animals (with manipulation of genes such as DGCR8 or DICER that code for relevant enzymes related to the biosynthesis process); and administration of microRNAs inhibitors or mimics peripherally, e.g., by intranasal injection, or directly into the brain by stereotactic injections into the interventricular area or hippocampus; and a.2) In human studies: observational cohort, case-control, or cross-sectional studies without an intervention and experimental designs such as randomized controlled trials. b Regarding the methods used for assessing the phenotype, i.e., executive functioning: b.1) In animal models: behavioral assessments of working memory (other domains are still barely established) with the eight-arm water maze, Morris water maze, Novel Object Recognition test, Y water maze, or T water maze (these evaluations may require correction for visual and motor abilities); and b.2) In human studies: the domains of executive functions are broader, for instance the Trail Making Test can measure set shifting, the Card Sorting Test evaluates inhibitory control, and the digit symbol test assesses working memory. 3) Regarding the assessment of the genotype or the microRNA changes: 3a) sample collection, with examples of biomaterial ranging from saliva to brain-specific areas collected post mortem; 3b) measurement of microRNA expression either with tests for specific microRNAs that use polymerase chain reaction methods or with broader analysis by sequencing techniques; and (c) differential microRNA expression analysis by using pipelines in programming or statistical software, with possible further bioinformatics analysis, e.g., network analysis or gene set enrichment analysis. And (d) Regarding additional methods that can complement the research, for example: d.1) studies of cellular electrical or morphological changes after microRNA manipulation (with inhibitors or mimics) in cellular cultures derived from primary or human-derived induced pluripotent stem cells or organoids; and d.2) other omics techniques, such as proteome analysis or correlations with changes in structural or functional imaging markers. Created in BioRender. Navarro Flores, A. (2026) https://BioRender.com/ztbg5cd. Partially adapted from Faber, S. [164].
The disease models most commonly evaluated in the animal studies were cognitive impairment secondary to various injuries (n = 8) [100, 102,103,104,105, 111, 112, 119], Alzheimer disease (n = 4) [110, 115,116,117], 22q11.2 deletion syndrome (n = 4) [107, 113, 114, 118], and age-related mild cognitive impairment (MCI, n = 3) [40, 99, 109]. In the human studies, the conditions studied were both primary MCI and MCI secondary to other chronic conditions (n = 9) [40, 75, 76, 78, 85, 89, 94, 95, 97], schizophrenia (n = 6) [77, 82, 84, 86, 90, 93], cognitive aging (n = 3) [80, 83, 96], major depressive disorder (n = 2) [79, 81], and normal variations of cognition in healthy individuals (n = 2) [87, 91].
Quality assessment
In the human cohorts, items 2 (selection of the non-exposed cohort), 3 (ascertainment of the exposure), and 5 (comparability of cohorts) had a low mean risk of bias and items 1 (representativeness of the sample), 6 (outcome assessment), and 8 (follow-up), had a high mean risk of bias. In the case-control studies, items 1 (sample representativeness) and 4 (validated tool) had a low mean risk of bias; items 5 (comparability of samples) and 6 (outcome assessment) had a moderate mean risk of bias; and items 2 (sample size), 3 (characteristics of the non-respondents), and 7 (statistical test) had a high mean risk of bias. The only randomized controlled trial (RCT) included had a high risk of bias [97].
In animal studies, a low to moderate risk of bias was found only for the following items: similar baseline characteristics between groups, missing data, and other sources of bias, in this case financing from industry agencies. The rest of the items presented a moderate to high risk of bias.
A detailed depiction of the risk of bias assessment is presented in the Supplement (eTables 8–9, eFigures 2–4).
Changes in microRNA-related genes
MIR137 variants: the most studied microRNA gene mutation impairs EFs
Gonzalez-Tirado et al. [91] performed a study in 230 healthy young Colombian adults (mean age, 21.2 years) and used polymerase chain reaction (PCR) to genotype the sample according to a functional VNTR for the MIR137 gene. They found that individuals who carried one copy of the 4-repeat allele showed better executive functioning, as measured by the Stroop test (Stroop facilitation scores). Liu et al. [87] performed a cross-sectional study in healthy participants in China (N = 290) and genotyped them for MIR137 rs1625579. On the basis of a functional magnetic resonance imaging experiment, they concluded that this single-nucleotide polymorphism (SNP) was associated with reduced connectivity in the dorsolateral prefrontal cortex and reduced working memory performance. Similarly, Ma et al. [86] conducted a case-control study of Chinese adults with schizophrenia (N = 1239; 611 cases and 628 controls) and evaluated working memory with the Digit Sequencing Task. They found that the T allele for the same SNP (rs1625579) was associated with lower working memory scores. Cosgrove et al. [93] performed a case-control study of 1000 Irish adults (808 patients with schizophrenia, 192 controls), most of them male (99%). The group studied the polygenic risk of the affected downstream genes that were targets of or were regulated by miR-137 and were relevant in cognition (including EFs) and found that the miR-137 polygenic score was inversely correlated with intelligence quotient, attention, and memory (i.e., episodic, visual, and working memory). Potkin et al. [82] combined data on gene differences identified in a GWAS with data from a working memory paradigm performed during functional magnetic resonance imaging [82]. The candidate genes found in the GWAS were submitted to an enrichment network analysis by using a gene set collection of experimental and putative targets, including microRNAs, curated by the authors. The authors found that the compromised genes regulated the expression of microRNA genes, including miR137, miR-448, and miR-218.
Negative results have also been reported for the MIR137 gene: Van Erp et al. [77] conducted a small case-control study in the USA in patients with schizophrenia (N = 111; 48 cases and 63 controls) and evaluated working memory with the Sternberg Item Recognition Paradigm during functional neuroimaging. They found higher hyperactivation of the dorsolateral prefrontal cortex and lower working memory scores in the patients than in the controls but no genotype-phenotype association.
Mutations in other genes related to microRNAs
SNPs in the MIR2113 gene were suggested to be related to general cognitive function. Andrews et al. studied this genetic marker in association with longitudinal cognitive performance, including working memory, across 12 years in a population of cognitively healthy older adults. They did not find an association with working memory, but they did find that the marker was related to an accelerated decline in episodic memory [96]. Similarly, Schroeder et al. studied working memory in older adults by using GWAS data. [80] The most significant SNP related to working memory performance was rs113948889, but additional SNPs were found to be related to other cognitive functions. The authors used a bioinformatic tool to predict which other proxy SNPs (in linkage disequilibrium with the ones found in their GWAS) were associated with microRNA-mRNA binding and found that rs1044950 (in linkage disequilibrium with rs113948889) was predicted to interfere with the binding of hsa-miR-138-5p and DCP1B. This relationship was confirmed with the luciferase assay in HEK293 cells. In postmortem human brains, both hsa-miR-138-5p and DCP1B were found in the areas of the hippocampus and frontal cortex, suggesting that an interaction may occur.
Effects of gene methylation on EFs
Some individuals maintain above average performance even as they age. Park et al. [83] performed a study in Korean older adults with above average performance in all cognitive domains, including EFs. They performed a methylation assay that included a microRNA gene site named MIR885 and found differentially methylated CpGs in the promoter regions of the MIR885 gene. The relative differentially methylated genes expression of this gene was two-fold higher in the group with successful cognitive aging than in the group with normal cognitive aging.
Reduced availability of nuclear microRNA processing enzymes
DiGeorge critical region 8 (DGCR8) is a subunit of the microprocessor complex that is required for the maturation of the longer primary microRNA into its short precursor form. [120] In 22q11.2 deletion syndrome, a neurodevelopmental disorder, both the DGCR8 and also other microRNA genes could be affected depending on the size of the deletion; consequently, an expression dysregulation is observed across the microRNAome. The phenotype includes cognitive impairment; a high risk for schizophrenia and other psychiatric disorders, such as anxiety and depression; and multiple physical malformations [121]. Fenelon et al. [114] engineered a mouse model of 22q11.2 DS by using a syntenic mutation. The DF16 +/− mouse was heterozygous to this mutation, and in the pyramidal neurons of the prefrontal cortex of this mouse, the authors found abnormal morphology that—during brief stimulation trains—was associated with short-term synaptic depression and a reduced initial phase of synaptic potentiation. These alterations in synaptic plasticity could explain the cognitive deficits in 22q11.2 DS and may provide a link between microRNA expression and higher order functions. In a later study, Fenelon et al. [113] found that mice with this mutation performed poorly in the Novel Object Recognition test of working memory. Another behavioral experiment was performed by Ouchi et al. [107] in a transgenic KO Dgcr8 mouse model: The group evaluated working memory with the Morris water maze and the Y maze and found reduced performance in these hippocampus-dependent tasks. The reduction in performance correlated with schizophrenia-related genes that were downregulated in this brain area and with a reduction in adult hippocampal stem/progenitor cells. Working memory performance improved after administration of insulin-like growth factor 2 in the hippocampus, a previously reported candidate for this function that was severely decreased in the mouse model [107].
Diamantopoulou et al. [118] also used the DF16 +/− mouse model along with a loss of function mutation in a gene with a major transcriptional effect due to microRNA dysregulation, Mirta22/Emc10. By adding this mutation, the authors rescued the schizophrenia-related phenotype of the mice, including impaired working memory performance [118].
Reduced availability of cytoplasmic microRNA processing enzymes
DICER is a cytoplasmic enzyme responsible for cleaving a microRNA into its mature form. Qiu et al. [106] generated a mouse model of DICER KO and found that postmitotic ablation of DICER in cortical vasoactive intestinal peptide-expressing interneurons (which are relevant for executive control) produced progressive cell loss in adulthood [106]. These mutant mice presented with a shortened life span and, paradoxically, performed better in the spatial working memory paradigm, even though learning and memory were not enhanced. Administration of valproate incremented the life span and reduced performance to levels similar to those in controls. The authors hypothesized that GABAergic circuits could be enhanced in and be responsible for this phenotype.
Differential microRNA expression correlates with EFs
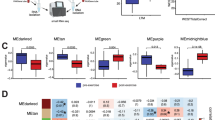
Studies reported dysregulation of microRNAs in both directions, i.e., upregulated and downregulated, and an association of microRNA dysregulation with increased and decreased EFs. A summary of the microRNA findings reported in at least two samples is presented below, and the full list of dysregulated microRNAs can be found in Table 3. Moreover, based on a recent report of microRNA expression in the human prefrontal cortex, [53] the reads per million of each candidate were included in this table to complement the interpretation of the results.
MicroRNAs consistently differentially expressed within sample types
In humans, miR-148a-3p was found to be upregulated and associated with reduced EFs in two different studies. [40, 85] Islam et al. [40] analyzed neurotypical participants from the PsyCourse Study [122] by assessing EFs with the Trail Making Test and Digit Symbol Test and working memory with the Digit Span Test. The authors performed a microRNAome sequencing analysis with total blood from these individuals and found that miR-148a-3p was significantly increased in participants who had MCI and were therefore at risk of developing dementia. In the other study, O’Meara et al. [85] measured EF performance in 31 patients with HIV who were at risk of cognitive decline. EFs were evaluated with the Trail Making Test parts A and B and the Controlled Oral Word Association Test. MicroRNA expression was measured by next-generation sequencing of plasma exosomes and compared between HIV patients with lower and higher EF performance, and the study found that miR-148a-3p was significantly upregulated in the former group.
In rodents, we found three microRNAs to be consistently dysregulated across samples: miR-155, miR-30e, and miR-384-5p. In a mouse model of traumatic brain injury, miR-155 was highly expressed in the microglia/microphages of the injured cortex. After injection of the inhibitor miR-155 antagomir and the consequent downregulation of miR-155 in the hippocampus, hippocampal expression of the pro-inflammatory biomarkers ITGAM, CD68, NOX2, p22phox, tumor necrosis factor alpha, and CCL2 was significantly reduced and the impairment of working memory was attenuated [111]. Similarly, inhibition of this microRNA in rats was associated with reduced expression of pro-inflammatory genes and rescued the impaired cognitive phenotype [116].
Xu et al. [100] found that rats overexpressing miRNA-30e in the central nervous system had cognitive impairments as measured with the Morris water maze. [100] Similarly, Chu et al. [119] showed in a mouse model of intoxication by polystyrene nano-plastics that upregulation of miRNA-30e was associated with reduced performance in the radial arm maze. [119] Finally, in a rat model of attention-deficit/hyperactivity disorder Xu et al. [101] found in two different samples that overexpression of miR-384—a highly conserved microRNA in humans and rodents—was associated with increased performance in the Morris water maze, whereas, in a different sample of rats, downregulation was associated with decreased performance in the same behavioral test. [101]
MicroRNAs consistently differentially expressed across sample types
The following microRNAs were reported as having functional consistency in at least one human and one animal sample: miR-132, miR-146a-5p, miR-148-3p, miR-181a-5p, miR-190b, miR-31, miR-501-3p, and miR-9-5p.
A study by Islam et al. [40] found that in addition to miE-148a-3p, the microRNAs miR-181a-5p and miR-146a-5p were also upregulated and associated with declined EFs in healthy human controls and mice models. In this study, mimics of the three microRNAs were injected into primary cultures of hippocampal neurons and immortalized microglia showing upregulation of genes related to neuronal plasticity, which is required for learning, and memory. Inhibitors of these microRNAs injected into the hippocampus of aged mice rescued the working memory performance up to levels comparable to those in young mice, indicating that these microRNAs may be a potential therapeutic option for MCI.
Qi et al. [81] assessed the effects on EFs of miR-132 expression in medication-naïve Chinese patients with depression. In these patients, miR-132 expression was upregulated and associated with reduced activation in the medial prefrontal cortex, an area relevant for executive performance. The authors found that grey matter thinning in prefrontal and hippocampal areas was associated with reduced scores in the EF test (Intra-Extra Dimensional Set Shift). This microRNA was also studied by a group examining the toxic effects on cognition produced by intrauterine fluoride exposure in mice [102]. The number of errors in the eight-arm maze test was higher in the exposed animals and in the highest dose group, and the hippocampal levels of miR-132 were higher in the exposed than in the non-exposed group. The levels of miR-124 and the DGCR8 gene were also incrementally increased [102].
In a sample of patients living with AIDS, miR-31-5p and miR-190b-5p were positively correlated with the scores on EF tests [95]. In line with those findings, a study of the cognitive effects of anesthetics in mice found that, after induction, the mice showed reduced alternation in the exploration of the arms in the Y maze and that miR-31 and miR-190b were significantly decreased in the medial prefrontal cortex and hippocampus [104].
Toyama et al. [103] studied the effects of vascular cognitive impairment in a model of bilateral common artery stenosis in mice and found that treatment with the microRNA inhibitor anti-miR-501-3p attenuated the spatial working memory deficits shown by the mice in the Y maze test [103]. Later, the same group studied community-dwelling Japanese older adults, some of whom presented comorbid diabetes mellitus or atherosclerosis, [78] and on the basis of the results hypothesized that vascular disruption contributes significantly to the burden of dementia. The authors evaluated the correlation between the serum expression of miR-501 measured by qPCR and the EF domain of the Montreal Cognitive Assessment and found that, regardless of the presence of a comorbid vascular condition, individuals with higher levels of miR-501 presented lower scores not only in EFs, but also in other cognitive domains, such as memory, attention, and visuospatial abilities.
Finally, for miR-9, Anderson et al. [97] found that after six months of an exercise intervention in older adults from the community, the levels of exosomal microRNA-9 were increased and associated with better performance in the Stroop A/C test [97]. Similarly, Gai et al. [112] showed that overexpression of miR-9-5p improved working memory in the hypoxia ischemia mouse model [112]. However, Malmevik et al. [108] inhibited the expression of three microRNAs in the hippocampi of mice including miR-9-5p by using a sponge design [108] and found no effects on the working memory performance of the mice in the Morris water maze test.
Bioinformatic assessment of candidate microRNAs
The network analysis showed few overlaps between the identified gene hubs, with nodes ranging from 120 (Mouse group 1) to 943 (Human group 1). Common gene hubs were genes that regulate transcription or the cell cycle, such as MYC and EP300 [123, 124]. A summary is presented in eTable 10. In the pathway analysis, upregulated microRNAs were related to processes involving neurogenesis, neurodevelopment, and synaptic plasticity/signaling, whereas downregulated microRNAs were associated with cell/neuron fate commitment, spinal cord development/neuron differentiation, and peripheral nervous/motor processes. The 10 most significant pathways found in each group are presented in Fig. 3. An extended version is presented in the Supplement (eTable 11).
Pathways involved in the association of microRNA dysregulation and changes in executive functions (EFs) in the following groups: (a) microRNAs upregulated and EFs increased in humans; (b) microRNAs upregulated and EFs decreased in humans; (c) microRNA downregulated and EFs increased in humans; (d) microRNAs upregulated and EFs increased in rodents; (e) microRNAs upregulated and EFs decreased in rodents; (f) microRNA downregulated and EFs increased in rodents; (g) microRNA downregulated and EFs decreased in rodents.
Discussion
Summary of the results
In this comprehensive review of the association between microRNA dysregulation and EFs in humans and animals, we found that alterations at the level of a microRNA or the related gene, loss of the microRNA biosynthesis machinery, and altered peripheral or tissue expression of microRNAs can either increase or decrease EFs. SNPs at the MIR137 gene were associated with reduced EFs. On the other hand, methylation in the promoter regions of the MIR885 gene was associated with successful cognitive aging. Loss of expression of the enzymes DGCR8 (22q11.2 DS) and DICER was associated with decreased working memory (a finding supported by electrophysiological findings). The consistently dysregulated microRNAs reported in two or more samples were as follows: in humans, miR-148a-3p; in rodents, miR-155, miR-30e, and miR-384-5p; and in both sample types, miR-132, miR-146a-5p, miR-148a-3p, miR-181a-5p, miR-190b, miR-31, miR-501-3p, and miR-9-5p. Gene ontology enrichment analysis showed that the targets of the dysregulated microRNAs were related to neurogenesis, neurodevelopment, and neuron fate commitment. Most of the human studies included patients with MCI or psychiatric conditions, and all of the studiesusing animal models evaluated only working memory. Our review thus identifies multiple potential targets on different levels that can be examined in future research, in particular in interventional studies aiming to improve higher-level cognition in humans.
Gene changes in EF performance
The rs113948889 SNP at the MIR137 gene was associated with reduced EFs. Gene mutations in microRNA areas had been reported. For example, in the mice genome 1700 SNPs were present in 702 pre-microRNAs and 609 SNPs in mature microRNAs and 841 SNPs could destabilize the secondary structure of the corresponding microRNA [125]. Also, a study in schizophrenia patients reported that cognitive impairment, including executive dysfunction, was longitudinally persistent and correlated with cortical structural changes, mostly in the frontal and temporal lobes [126]. In line with those results, a mega-analysis of GWASs on schizophrenia found that the candidate gene MIR137 was related to the pathology of the disease [127], and various authors replicated its association with the diagnosis of schizophrenia in independent studies [128]. VNTRs in this gene were found to be associated with morphological changes in the brain, such as a thicker left inferior temporal gyrus and deeper right postcentral sulcus, and with a cognitive deficit subtype of schizophrenia [129]. Other gene mutations may also be of interest and require future research.
Nuclear microRNA processing
The 22q11.2 deletion syndrome and the associated dysregulation by microRNAs may explain the associated high risk for schizophrenia, a psychiatric disorder strongly linked with deficient EFs [130]. The syndrome is characterized by a genetic mutation in the area closely related to the expression of the DGCR enzyme, which is needed for the biosynthesis of microRNAs. Different microRNAs have been reported as dysregulated in 22q11.2 DS, but a clear signature is still lacking [131]. Interestingly, patients with this syndrome have a high prevalence of various psychiatric disorders: As many as 70% of them have anxiety, depression, or attention-deficit/hyperactivity disorder and up to 25% have schizophrenia [34]. Impairment of EFs and other cognitive disabilities are also part of the phenotype [132]. These findings imply the potential role of microRNAs in the normal development of the brain and in maintaining optimal higher brain functioning.
DICER
In an inflammatory model resembling Parkinson disease in microglia and mice, one study showed that microglial DICER degradation is associated with increased inflammation and dopaminergic neuronal loss [133]. To this end, a physical exercise intervention in 3xTg mice (an Alzheimer disease model) showed that after 20 weeks, in the active group the levels of DICER and miR-29 were higher and the levels of the latter correlated with lower levels of one of its targets, Beta-secretase 1. This result might suggest that β−amyloid accumulation is related to lower levels of DICER [134]. Another experiment deleted the Dicer1 gene in the adult mouse forebrain and found downregulation of various microRNAs. After 12 weeks, the authors observed an increase in memory performance as a prodrome of neurodegeneration, suggesting the importance of microRNAs for cognitive processes in general [135].
Differential expression
The microRNAs that were reported as consistently dysregulated in two or more samples and had an inverse correlation with EFs were miR-155 and miR-30e in rodents and miR-132, miR-146a-5p, miR-148a-3p, miR-181a-5p, and miR-501-3p in both samples. MiR-384-5p presented a direct correlation with EFs in rodents, and miR-190b, miR-31, and miR-9-5p did so in both sample types.
A systematic review that included a meta-analysis of 147 different datasets on differential expression of microRNAs in Alzheimer disease patients found similar results regarding these microRNAs. [136] In line with our findings, the authors reported that across the studies, hsa-miR-501-3p was consistently upregulated in the brains of these patients. Additionally, miR-146a-5p was reported as upregulated in brain and downregulated in blood, which partially agrees with our results and highlights the tissue specificity of microRNA expression. Moreover, these findings suggest that because microRNA signatures are related to MCI, they could also serve as a predictor of an early stage of Alzheimer disease, justifying further research.
Another systematic review that evaluated microRNAs dysregulated in association with neurodegenerative disorders included 641 studies and identified similar microRNAs among the most frequently occurring candidates [33]. Like the present study, the review found that miR-146a-5p and miR-155-5p were upregulated in patients with neurogenerative disorders. Impairment of EFs is a common feature of neurodegenerative disorders [137], and the authors proposed that dysregulation of these microRNAs could affect shared pathways related to cognition, including an increase in proinflammatory markers, increased amyloid beta genesis, and altered neurogenesis and neural cell differentiation [33].
In addition to studying microRNAs, evaluating other classes of small non-coding RNAs could improve our understanding of the epigenetic mechanisms behind EFs. One such class is transfer RNA-derived small fragments (tRFs), which are produced through the cleavage of precursor or mature transfer RNAs, often in response to cellular stress. [138] This emerging class of small non-coding RNAs owes its diversity to the fact that tRFs can arise from precursor or mature transfer RNAs. [139] These fragments are not only diverse in origin, but also abundant in various biological fluids: They make up a large proportion of small RNAs in urine (84%), amniotic fluid (75%), and cerebrospinal fluid ( ~ 47%). [140] The roles of tRFs are similar to those of microRNAs in that they are capable of binding Argonaute proteins to silence the expression of mRNA. [141] Also noteworthy is that given their higher levels in neurons and their higher occurrence relative to microRNAs, tRFs likely play a broader role in regulating gene networks associated with EF-related processes. Further support for this hypothesis is provided by the finding that their dysregulation is linked to neurological disorders such as epilepsy, amyotrophic lateral sclerosis, and cognitive decline in Alzheimer disease. [142,143,144] The major role of EFs in healthy cognition and their sensitivity to brain disorders supports the hope that further research on tRFs could uncover new mechanisms and gene targets related to this phenotypic trait.
Bioinformatics analysis of microRNA targets
According to our exploratory bioinformatic analysis, the most relevant pathways through which microRNA regulation influences EFs include neurogenesis, nervous system development, and synapse regulation and signaling.
Adult neurogenesis was first observed in rodent models and was later confirmed in the adult human hippocampus (dentate gyrus) to be responsible for lifelong synaptic plasticity. [145,146,147] Alterations in neurogenesis that occur during aging or in neuropsychiatric conditions can impact executive functioning, hence producing impaired cognitive flexibility and working memory [148]. This process is susceptible to environmental modifications such as enrichment [149], but epigenetic regulations also play an important role. For instance, increased levels of miR-132, a candidate that was identified also in this review, improved memory deficits in a mouse model of Alzheimer disease by rescuing hippocampal neurogenesis. [150] On the other hand, hippocampal upregulation of miR-146-5p was associated with higher cognitive impairment, whereas its downregulation increased hippocampal neurogenesis and rescued the phenotype. [151, 152] In the case of the miR-17-92 cluster, inhibition of its expression in a transgenic mouse model led to reduced hippocampal neurogenesis and a phenotype of cognitive impairment [153], which was later observed to be associated with symptoms of anxiety and depression [154].
Regulation by microRNAs also takes place during neurodevelopment, including forebrain development, neuronal differentiation and maturation, and synaptogenesis [155]. One example is miR-124, which regulates neuronal development by targeting the Notch pathway. [156] Further study of the involvement of these and other mechanisms in the regulation of EFs by microRNA candidates is granted.
Synaptic plasticity in the prefrontal cortex (an area relevant to EFs) is mediated by dopamine signaling and explains to some extent differences in the performance of rodents in cognitive tasks [157]. MicroRNAs regulate synaptic plasticity by silencing relevant plasticity mediators and dendritic proteins needed for signal transmission [158]. For example, inhibition of microRNA 135a-5p expression in mice produces synaptic dysfunction and memory impairment [157]. In the case of major depressive disorder, several microRNAs interfere with synaptic signaling and protein-synthesis dependent synaptic plasticity [159]. These mechanisms could potentially explain cognitive deficits in major depressive disorder and other psychiatric disorders.
Conclusions and future perspectives
Alterations of microRNA biosynthesis at the gene level, loss of the microRNA biosynthesis machinery, and altered peripheral or tissue-specific expression of microRNAs affect EFs. These changes mostly reduce EFs, but some improvement was also observed. A total of eight candidates were found consistently across sample types (i.e., in both human and rodent studies). The targets of the significantly dysregulated microRNAs were related to common pathways such as neurogenesis, neurodevelopment, and synaptic plasticity. Although it was not the focus of this review, isoforms of miRNAs (isomiRs) which are variants of the canonical from of different sequences or length, improve the specificity of target regulation and have been explored to improve the precision of miRNA-based biomarkers, [160] showing promising results in animal models of epilepsy. [161]
Most of the human studies were in patients with MCI or psychiatric conditions, and all of the animal studies evaluated only working memory. These microRNA signatures require further confirmation as potential biomarkers in human studies with larger sample sizes and in controlled and blinded interventions. If confirmed as such, these signatures could not only be easily implemented in clinical practice to classify at-risk patients and provide additional cognitive training, but they could also be considered as therapeutic targets to enhance EFs or prevent their decline. Therefore, research on this topic is necessary to improve the quality of life of patients with psychiatric, neurologic, and inflammatory disorders and of healthy aging individuals. Additionally, longitudinal studies could provide a broader perspective of the relationship between microRNA expression and EFs, both of which are dynamic constructs, to identify predictive signatures that correlate with the various trajectories of EFs.
Limitations
The high heterogeneity resulting from the various methodological approaches used in the studies (i.e., selection of different diagnoses, use of different microRNA processing methods and bioinformatic pipelines, and differences in statistical analysis methods) prevented us from evaluating the effects related to the commonly dysregulated microRNAs by a more detailed analysis, such as a pooled meta-analysis.
Most of the included studies had a moderate to high risk of bias. Human cohorts were deficient in the representativeness of the sample chosen, blinding of the personnel involved in data processing and analysis, and the longitudinal assessment of the outcomes.
This review identified mostly cross-sectional human studies assessing miRNA expression. Only two studies used a longitudinal approach, but no repeated measures of microRNA expression in humans related to executive functions were found. Cross-sectional designs can establish associations between miRNA expression and executive function phenotypes but cannot infer causality due to temporal ambiguity, reverse causation, and confounding. Longitudinal studies tracking miRNA dynamics over time, paired with cognitive assessments, are needed to elucidate directional relationships and rule out reverse causality (e.g., cognitive decline influencing miRNA expression).
In most of the case-control studies, sample sizes were reduced, non-responders were not clearly described, assessors were not blinded, and the comparability of the sample with other studies was decreased because of the type of statistical analyses performed. Animal studies lacked randomization of housing, blinding of outcome assessment, and a registered or publicly available protocol, all of which could limit the reliability and validity of the results.
Our bioinformatics pipeline presented limitations which are common to miRNA research. The in-house tool aggregated experimentally validated targets from NPInter (ncRNA-protein/RNA), TarBase (miRNA-mRNA), TransmiR (transcription factor-miRNA), RegNetwork (regulatory networks), Rise (regulatory interaction search), and STRING (protein interactions). While validated targets provide higher confidence, they present limited coverage with an overrepresentation of studies in the field of cancer. [162] These limitations propagate to the pathway enrichment analysis, where Gene Ontology over-representation can reflect biological processes biased towards that field. [163] We therefore interpret networks and pathways as hypothesis-generating at the module level, not as confirmed direct 3′UTR/promoter interactions. Definitive validation requires future testing with functional assays (e.g., 3′UTR luciferase reporters with mutagenesis, CLIP). The above-mentioned limitations should be taken into consideration when interpreting the results of this review.
References
Friedman NP, Miyake A. Unity and diversity of executive functions: individual differences as a window on cognitive structure. Cortex. 2017;86:186–204.
Jurado MB, Rosselli M. The elusive nature of executive functions: a review of our current understanding. Neuropsychology Rev. 2007;17:213–33.
Diamond A. Executive functions. Annu Rev Psychol. 2013;64:135–68.
Hatoum AS. The etiology of executive functioning is nature and nurture. Biol Psychiatry. 2023;94:e5–6.
Li JJ, Roberts DK Genetic influences on executive functions across the life span. In: Wiebe S, Karbach J (eds). Executive Function: Development Across the Lifespan. Routledge2017.
Logue SF, Gould TJ. The neural and genetic basis of executive function: attention, cognitive flexibility, and response inhibition. Pharmacology, biochemistry, Behav. 2014;123:45–54.
Barnes JJ, Dean AJ, Nandam LS, O’Connell RG, Bellgrove MA. The molecular genetics of executive function: role of monoamine system genes. Biol Psychiatry. 2011;69:e127–43.
Hatoum AS, Morrison CL, Mitchell EC, Lam M, Benca-Bachman CE, Reineberg AE, et al. Genome-wide association study shows that executive functioning is influenced by GABAergic processes and is a neurocognitive genetic correlate of psychiatric disorders. Biol Psychiatry. 2023;93:59–70.
Miguel PM, Meaney MJ, Silveira PP. New research perspectives on the interplay between genes and environment on executive function development. Biol Psychiatry. 2023;94:131–41.
Tervo-Clemmens B, Calabro FJ, Parr AC, Fedor J, Foran W, Luna B. A canonical trajectory of executive function maturation from adolescence to adulthood. Nat Commun. 2023;14:6922.
Lund JI, Boles K, Radford A, Toombs E, Mushquash CJ. A systematic review of childhood adversity and executive functions outcomes among adults. Arch Clin neuropsychology. 2022;37:1118–32.
Veríssimo J, Verhaeghen P, Goldman N, Weinstein M, Ullman MT. Evidence that ageing yields improvements as well as declines across attention and executive functions. Nat Hum Behav. 2022;6:97–110.
Lobo JD, Goodman ZT, Schmaus JA, Uddin LQ, McIntosh RC. Association of cardiometabolic health factors with age-related executive function and episodic memory. Neuropsychology, development, cognition Sect B, Aging, neuropsychology cognition. 2022;29:746–60.
Pan Z, Park C, Brietzke E, Zuckerman H, Rong C, Mansur RB, et al. Cognitive impairment in major depressive disorder. CNS Spectr. 2019;24:22–29.
Bourne C, Aydemir Ö, Balanzá-Martínez V, Bora E, Brissos S, Cavanagh JTO, et al. Neuropsychological testing of cognitive impairment in euthymic bipolar disorder: an individual patient data meta-analysis. Acta Psychiatrica Scandinavica. 2013;128:149–62.
Navarro-Flores A, Heilbronner M, Rafiee H, Wendel B, Papiol S, Adorjan K, et al. Polygenic scores for executive functioning as predictors of performance improvements after repeated testing in major psychiatric disorders. Sci Rep. 2026;16:9199.
Nyvold O, Nygaard E, Augusti EM, Tamnes CK. Unity or diversity of executive functioning in children and adolescents with post-traumatic stress symptoms? a systematic review and meta-analysis. Child neuropsychology. 2022;28:374–93.
Ganesan K, Steinbeis N. Development and plasticity of executive functions: a value-based account. Curr Opin Psychol. 2022;44:215–9.
Taylor R, Cella M, Wykes T. Cognitive remediation is an evidence-based psychological therapy: isn’t it time it was treated like one? Behav Modif. 2025;49:502–26.
Mokhtari S, Mokhtari A, Bakizadeh F, Moradi A, Shalbafan M. Cognitive rehabilitation for improving cognitive functions and reducing the severity of depressive symptoms in adult patients with Major Depressive Disorder: a systematic review and meta-analysis of randomized controlled clinical trials. BMC psychiatry. 2023;23:77.
Mokhtari S, Mokhtari A, Saeed F, Aghayan SN, Boroon M, Shalbafan M. Cognitive rehabilitation in anxiety disorders: a systematic review and meta-analysis of randomized controlled trials. Sci Rep. 2025;15:38420.
Zhang L, Lu Q, Chang C. Epigenetics in health and disease. Adv Exp Med Biol. 2020;1253:3–55.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54.
Mohr AM, Mott JL. Overview of microRNA biology. Semin liver Dis. 2015;35:3–11.
Selbach M, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N. Widespread changes in protein synthesis induced by microRNAs. Nature. 2008;455:58–63.
Ha M, Kim VN. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol. 2014;15:509–24.
Maffioletti E, Tardito D, Gennarelli M, Bocchio-Chiavetto L. Micro spies from the brain to the periphery: new clues from studies on microRNAs in neuropsychiatric disorders. Front Cell Neurosci. 2014;8:75.
O’Carroll D, Schaefer A. General principals of miRNA biogenesis and regulation in the brain. Neuropsychopharmacology. 2013;38:39–54.
Huang E. miRNA Processing Mechanisms in the Brain. Brain Sci. 2022;12:1366.
Gurtan AM, Sharp PA. The role of miRNAs in regulating gene expression networks. J Mol Biol. 2013;425:3582–600.
Sun E, Shi Y. MicroRNAs: small molecules with big roles in neurodevelopment and diseases. Exp Neurol. 2015;268:46–53.
Chen W, Qin C. General hallmarks of microRNAs in brain evolution and development. RNA Biol. 2015;12:701–8.
Juźwik CA, S. Drake S, Zhang Y, Paradis-Isler N, Sylvester A, Amar-Zifkin A, et al. microRNA dysregulation in neurodegenerative diseases: a systematic review. Prog Neurobiol. 2019;182:101664.
Vorstman JA, Breetvelt EJ, Duijff SN, Eliez S, Schneider M, Jalbrzikowski M, et al. Cognitive decline preceding the onset of psychosis in patients with 22q11.2 deletion syndrome. JAMA psychiatry. 2015;72:377–85.
Luoni A, Riva MA. MicroRNAs and psychiatric disorders: from aetiology to treatment. Pharmacology therapeutics. 2016;167:13–27.
Mendes-Silva AP, Fujimura PT, Silva J, Teixeira AL, Vieira EM, Guedes PHG, et al. Brain-enriched MicroRNA-184 is downregulated in older adults with major depressive disorder: a translational study. J Psychiatr Res. 2019;111:110–20.
Navarro-Flores A, Heilbronner U. Opportunities to advance microRNA research in psychiatry. Eur Neuropsychopharmacol. 2025;92:26–28.
Mendes-Silva AP, Pereira KS, Tolentino-Araujo GT, Nicolau Ede S, Silva-Ferreira CM, Teixeira AL, et al. Shared biologic pathways between alzheimer disease and major depression: a systematic review of MicroRNA expression studies. Am J geriatric psychiatry. 2016;24:903–12.
King KP, Humiston T, Gowey MA, Murdaugh DL, Dutton GR, Lansing AH. A biobehavioural and social-structural model of inflammation and executive function in pediatric chronic health conditions. Health Psychol Rev. 2022;18:24–40.
Islam MR, Kaurani L, Berulava T, Heilbronner U, Budde M, Centeno TP, et al. A microRNA signature that correlates with cognition and is a target against cognitive decline. EMBO Mol Med. 2021;13:e13659.
Rao P, Benito E, Fischer A. MicroRNAs as biomarkers for CNS disease. Front Mol Neurosci. 2013;6:39.
Salta E, De Strooper B. Noncoding RNAs in neurodegeneration. Nat Rev Neurosci. 2017;18:627–40.
Halse M, Steinsbekk S, Hammar Å, Wichstrøm L. Longitudinal relations between impaired executive function and symptoms of psychiatric disorders in childhood. J child Psychol psychiatry, allied Discip. 2022;63:1574–82.
McIntosh EC, Tureson K, Rotblatt LJ, Singer EJ, Thames AD. HIV, vascular risk factors, and cognition in the combination antiretroviral therapy era: a systematic review and meta-analysis. J Int Neuropsychological Soc. 2021;27:365–81.
Girotti M, Adler SM, Bulin SE, Fucich EA, Paredes D, Morilak DA. Prefrontal cortex executive processes affected by stress in health and disease. Prog Neuro-psychopharmacol Biol psychiatry. 2018;85:161–79.
Wollesen B, Wildbredt A, van Schooten KS, Lim ML, Delbaere K. The effects of cognitive-motor training interventions on executive functions in older people: a systematic review and meta-analysis. Eur Rev aging Phys Act. 2020;17:9.
Moher D, Liberati A, Tetzlaff J, Altman DG. The PG. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA Statement. PLOS Med. 2009;6:e1000097.
Wells GA, Shea B, O’Connell D, Peterson J, Welch V, Losos M et al. The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. 2000.
Eldridge S, Campbell M, Campbell M, Drahota DA, Giraudeau B, Higgins J et al. Revised Cochrane risk of bias tool for randomized trials (RoB 2.0). 2016.
Hooijmans CR, Rovers MM, de Vries RB, Leenaars M, Ritskes-Hoitinga M, Langendam MW. SYRCLE’s risk of bias tool for animal studies. BMC Med Res Methodol. 2014;14:43.
Fabian-Jessing BK, Vallentin MF, Secher N, Hansen FB, Dezfulian C, Granfeldt A, et al. Animal models of cardiac arrest: A systematic review of bias and reporting. Resuscitation. 2018;125:16–21.
BioRender B. BioRender. Com2024.
Vattathil SM, Gerasimov ES, Canon SM, Lori A, Tan SSM, Kim PJ, et al. Mapping the microRNA landscape in the older adult brain and its genetic contribution to neuropsychiatric conditions. Nat aging. 2025;5:306–19.
Matsumoto J, Stewart T, Banks WA, Zhang J. The transport mechanism of extracellular vesicles at the blood-brain barrier. Curr Pharm Des. 2017;23:6206–14.
Sempere LF, Freemantle S, Pitha-Rowe I, Moss E, Dmitrovsky E, Ambros V. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 2004;5:R13.
Mor E, Shomron N. Species-specific microRNA regulation influences phenotypic variability: perspectives on species-specific microRNA regulation. BioEssays. 2013;35:881–8.
Hu HY, Guo S, Xi J, Yan Z, Fu N, Zhang X, et al. MicroRNA expression and regulation in human, chimpanzee, and macaque brains. PLoS Genet. 2011;7:e1002327.
Li X, Wang Q, Zheng Y, Lv S, Ning S, Sun J, et al. Prioritizing human cancer microRNAs based on genes’ functional consistency between microRNA and cancer. Nucleic acids Res. 2011;39:e153.
Gong X, Wu R, Wang H, Guo X, Wang D, Gu Y, et al. Evaluating the consistency of differential expression of microRNA detected in human cancers. Mol cancer therapeutics. 2011;10:752–60.
Flynt AS, Lai EC. Biological principles of microRNA-mediated regulation: shared themes amid diversity. Nat Rev Genet. 2008;9:831–42.
Lee T, Wang N, Houel S, Couts K, Old W, Ahn N. Dosage and temporal thresholds in microRNA proteomics. Mol Cell Proteom. 2015;14:289–302.
Ebert MS Molecular titration by microRNAs and target mimic inhibitors. Massachusetts Institute of Technology2010.
Fischer S, Handrick R, Aschrafi A, Otte K. Unveiling the principle of microRNA-mediated redundancy in cellular pathway regulation. RNA Biol. 2015;12:238–47.
Epple R, Krüger D, Berulava T, Brehm G, Ninov M, Islam R, et al. The coding and small non-coding hippocampal synaptic RNAome. Mol Neurobiol. 2021;58:2940–53.
Teng X, Chen X, Xue H, Tang Y, Zhang P, Kang Q, et al. NPInter v4.0: an integrated database of ncRNA interactions. Nucleic acids Res. 2020;48:D160–d165.
Karagkouni D, Paraskevopoulou MD, Chatzopoulos S, Vlachos IS, Tastsoglou S, Kanellos I, et al. DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA-gene interactions. Nucleic acids Res. 2018;46:D239–d245.
Tong Z, Cui Q, Wang J, Zhou Y. TransmiR v2.0: an updated transcription factor-microRNA regulation database. Nucleic acids Res. 2019;47:D253–d258.
Liu ZP, Wu C, Miao H, Wu H. RegNetwork: an integrated database of transcriptional and post-transcriptional regulatory networks in human and mouse. Database. 2015;2015:bav095.
El Fatimy R, Li S, Chen Z, Mushannen T, Gongala S, Wei Z, et al. MicroRNA-132 provides neuroprotection for tauopathies via multiple signaling pathways. Acta neuropathologica. 2018;136:537–55.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic acids Res. 2019;47:D607–d613.
Lorenzi L, Chiu H-S, Avila Cobos F, Gross S, Volders P-J, Cannoodt R, et al. The RNA Atlas expands the catalog of human non-coding RNAs. Nat Biotechnol. 2021;39:1453–65.
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation. 2021;2:100141.
Team RC. RA language and environment for statistical computing, R Foundation for Statistical. 2020.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Zhang H, Yang S, Zhu W, Niu T, Wang J, Yang M, et al. Exosomal miR-let-7c-5p is involved in the cognitive function of type 2 diabetes mellitus patients by interleukin 10: a cross-sectional study. J diabetes. 2023;15:978–86.
Wang Z, Li XN, Yang SN, Wang Y, Gao KJ, Han B, et al. Exosomal miR-320e through wnt2targeted inhibition of the Wnt/β-catenin pathway allevisate cerebral small vessel disease and cognitive impairment. World J psychiatry. 2023;13:630–44.
van Erp TG, Guella I, Vawter MP, Turner J, Brown GG, McCarthy G, et al. Schizophrenia miR-137 locus risk genotype is associated with dorsolateral prefrontal cortex hyperactivation. Biol Psychiatry. 2014;75:398–405.
Toyama K, Spin JM, Tsao PS, Maruyama K, Osawa H, Mogi M, et al. Serum microRNA-501-3p is a potential diagnostic tool for detecting mild cognitive impairment: Ehime genome study. J neurochemistry. 2023;166:960–71.
Sun N, Yang C, He X, Liu Z, Liu S, Li X, et al. Impact of expression and genetic variation of microRNA-34b/c on cognitive dysfunction in patients with major depressive disorder. Neuropsychiatric Dis Treat. 2020;16:1543–54.
Schröeder J, Ansaloni S, Schilling M, Liu T, Radke J, Jaedicke M, et al. MicroRNA-138 is a potential regulator of memory performance in humans. Front Hum Neurosci. 2014;8:501.
Qi S, Yang X, Zhao L, Calhoun VD, Perrone-Bizzozero N, Liu S, et al. MicroRNA132 associated multimodal neuroimaging patterns in unmedicated major depressive disorder. Brain. 2018;141:916–26.
Potkin SG, Macciardi F, Guffanti G, Fallon JH, Wang Q, Turner JA, et al. Identifying gene regulatory networks in schizophrenia. NeuroImage. 2010;53:839–47.
Park J, Lee Y, Won CW. CEND1 and miR885 methylation changes associated with successful cognitive aging in community-dwelling older adults. Exp Gerontology. 2022;160:111704.
Pan SJ, Feng W, Li YL, Huang JC, Chen S, Cui YM, et al. The microRNA-195-BDNF pathway and cognitive deficits in schizophrenia patients with minimal antipsychotic medication exposure. Transl Psychiatry. 2021;11:117.
OʼMeara T, Kong Y, Chiarella J, Price RW, Chaudhury R, Liu X, et al. Exosomal MicroRNAs Associate With Neuropsychological Performance in Individuals With HIV Infection on Antiretroviral Therapy. J acquired immune deficiency syndromes. 2019;82:514–22.
Ma G, Yin J, Fu J, Luo X, Zhou H, Tao H, et al. Association of a miRNA-137 polymorphism with schizophrenia in a Southern Chinese Han population. BioMed Res Int. 2014;2014:751267.
Liu B, Zhang X, Hou B, Li J, Qiu C, Qin W, et al. The impact of MIR137 on dorsolateral prefrontal-hippocampal functional connectivity in healthy subjects. Neuropsychopharmacology. 2014;39:2153–60.
Ignacio C, Hicks SD, Burke P, Lewis L, Szombathyne-Meszaros Z, Middleton FA. Alterations in serum microRNA in humans with alcohol use disorders impact cell proliferation and cell death pathways and predict structural and functional changes in brain. BMC Neurosci. 2015;16:55.
Han L, Tang Y, Bai X, Liang X, Fan Y, Shen Y, et al. Association of the serum microRNA-29 family with cognitive impairment in Parkinson’s disease. Aging. 2020;12:13518–28.
Gou MZ, Pan SJ, Tong JH, Zhou YF, Han JR, Xie T, et al. Effects of microRNA-181b-5p on cognitive deficits in first-episode patients with schizophrenia: Mediated by BCL-2. J Psychiatr Res. 2021;136:358–65.
González-Giraldo Y, González-Reyes RE, Forero DA. A functional variant in MIR137, a candidate gene for schizophrenia, affects Stroop test performance in young adults. Psychiatry Res. 2016;236:202–5.
Dypås LB, Duale N, Olsen AK, Bustamante M, Maitre L, Escaramis G, et al. Blood miRNA levels associated with ADHD traits in children across six European birth cohorts. BMC psychiatry. 2023;23:696.
Cosgrove D, Harold D, Mothersill O, Anney R, Hill MJ, Bray NJ, et al. MiR-137-derived polygenic risk: effects on cognitive performance in patients with schizophrenia and controls. Transl Psychiatry. 2017;7:e1012.
Bijkerk R, Kallenberg MH, Zijlstra LE, van den Berg BM, de Bresser J, Hammer S, et al. Circulating angiopoietin-2 and angiogenic microRNAs associate with cerebral small vessel disease and cognitive decline in older patients reaching end-stage renal disease. Nephrology, dialysis, Transplant. 2022;37:498–506.
Aparicio JM, Xu YX, Li YL, Colantuoni C, Dastgheyb R, Williams DW, et al. Plasma microRNAs are associated with domain-specific cognitive function in people with HIV. Aids. 2021;35:1795–804.
Andrews SJ, Das D, Anstey KJ, Easteal S. Association of AKAP6 and MIR2113 with cognitive performance in a population-based sample of older adults. Genes, Brain Behav. 2017;16:472–8.
Anderson-Hanley C, Barcelos NM, Zimmerman EA, Gillen RW, Dunnam M, Cohen BD, et al. The aerobic and cognitive exercise study (ACES) for community-dwelling older adults with or at-risk for mild cognitive impairment (MCI): neuropsychological, neurobiological and neuroimaging outcomes of a randomized clinical trial. Front aging Neurosci. 2018;10:76.
Zhu X, Hollinger KR, Huang Y, Borjabad A, Kim BH, Arab T, et al. Neutral sphingomyelinase 2 inhibition attenuates extracellular vesicle release and improves neurobehavioral deficits in murine HIV. Neurobiol Dis. 2022;169:105734.
Zhang Y, Deng W, Wang W, Song A, Mukama O, Deng S, et al. MicroRNA-206 down-regulated human umbilical cord mesenchymal stem cells alleviate cognitive decline in D-galactose-induced aging mice. Cell death discovery. 2022;8:304.
Xu Y, Liu Z, Song X, Zhang K, Li X, Li J, et al. Cerebralcare Granule® attenuates cognitive impairment in rats continuously overexpressing microRNA-30e. Mol Med Rep. 2015;12:8032–40.
Xu Q, Ou J, Zhang Q, Tang R, Wang J, Hong Q, et al. Effects of aberrant miR-384-5p expression on learning and memory in a rat model of attention deficit hyperactivity disorder. Front Neurol. 2019;10:1414.
Wang J, Zhang Y, Guo Z, Li R, Xue X, Sun Z, et al. Effects of perinatal fluoride exposure on the expressions of miR-124 and miR-132 in hippocampus of mouse pups. Chemosphere. 2018;197:117–22.
Toyama K, Spin JM, Deng AC, Huang TT, Wei K, Wagenhäuser MU, et al. MicroRNA-Mediated therapy modulating blood-brain barrier disruption improves vascular cognitive impairment. Arteriosclerosis, thrombosis, Vasc Biol. 2018;38:1392–406.
Shan LG, Ma D, Zhang CS, Xiong W, Zhang Y. miRNAs may regulate GABAergic transmission associated genes in aged rats with anesthetics-induced recognition and working memory dysfunction. Brain Res. 2017;1670:191–200.
Sarkar S, Engler-Chiurazzi EB, Cavendish JZ, Povroznik JM, Russell AE, Quintana DD, et al. Over-expression of miR-34a induces rapid cognitive impairment and Alzheimer’s disease-like pathology. Brain Res. 2019;1721:146327.
Qiu F, Mao X, Liu P, Wu J, Zhang Y, Sun D, et al. microRNA Deficiency in VIP+ Interneurons leads to cortical circuit dysfunction. Cereb cortex. 2020;30:2229–49.
Ouchi Y, Banno Y, Shimizu Y, Ando S, Hasegawa H, Adachi K, et al. Reduced Adult hippocampal neurogenesis and working memory deficits in the Dgcr8- Deficient mouse model of 22q11.2 deletion-associated schizophrenia can be rescued by IGF2. J Neurosci. 2013;33:9408–19.
Malmevik J, Petri R, Knauff P, Brattås PL, Åkerblom M, Jakobsson J. Distinct cognitive effects and underlying transcriptome changes upon inhibition of individual miRNAs in hippocampal neurons. Sci Rep. 2016;6:19879.
Liu K, Yin Y, Le Y, Ouyang W, Pan A, Huang J, et al. Age-related Loss of miR-124 causes cognitive deficits via derepressing RyR3 expression. Aging Dis. 2022;13:1455–70.
Koh HS, Lee S, Lee HJ, Min JW, Iwatsubo T, Teunissen CE, et al. Targeting MicroRNA-485-3p blocks alzheimer’s disease progression. Int J Mol Sci. 2021;22:13136.
Henry RJ, Doran SJ, Barrett JP, Meadows VE, Sabirzhanov B, Stoica BA, et al. Inhibition of miR-155 limits neuroinflammation and improves functional recovery after experimental traumatic brain injury in mice. Neurotherapeutics. 2019;16:216–30.
Gai CC, Xing XH, Song Y, Zhao YJ, Jiang ZG, Cheng YH, et al. Up-Regulation of miR-9-5p inhibits hypoxia-ischemia brain damage through the DDIT4-Mediated autophagy pathways in neonatal mice. Drug Des Dev Ther. 2023;17:1175–89.
Fénelon K, Xu B, Lai CS, Mukai J, Markx S, Stark KL, et al. The pattern of cortical dysfunction in a mouse model of a schizophrenia-related microdeletion. J Neurosci. 2013;33:14825–39.
Fénelon K, Mukai J, Xu B, Hsu PK, Drew LJ, Karayiorgou M, et al. Deficiency of Dgcr8, a gene disrupted by the 22q11.2 microdeletion, results in altered short-term plasticity in the prefrontal cortex. Proc Natl Acad Sci USA. 2011;108:4447–52.
Berchenko OG, Levicheva NO, Bevzyuk DO, Sokolik VV. The effect of miR-101 on the memory of rats with a model of Alzheimer’s disease. Regulatory Mechanisms Biosyst. 2020;11:354–9.
Liu D, Zhao D, Zhao Y, Wang Y, Zhao Y, Wen C. Inhibition of microRNA-155 Alleviates cognitive impairment in alzheimer’s disease and involvement of neuroinflammation. Curr Alzheimer Res. 2019;16:473–82.
Kawakita R, Takata T, Nonaka W, Hamada Y, Iwama H, Kobara H, et al. Age‑related brainstem degeneration through microRNA modulation in mice. Mol Med Rep. 2023;28:146.
Diamantopoulou A, Sun Z, Mukai J, Xu B, Fenelon K, Karayiorgou M, et al. Loss-of-function mutation in Mirta22/Emc10 rescues specific schizophrenia-related phenotypes in a mouse model of the 22q11.2 deletion. Proc Natl Acad Sci USA. 2017;114:E6127–e6136.
Chu C, Zhang Y, Liu Q, Pang Y, Niu Y, Zhang R. Identification of ceRNA network to explain the mechanism of cognitive dysfunctions induced by PS NPs in mice. Ecotoxicol Environ Saf. 2022;241:113785.
Wang Y, Medvid R, Melton C, Jaenisch R, Blelloch R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat Genet. 2007;39:380–5.
McDonald-McGinn DM, Sullivan KE, Marino B, Philip N, Swillen A, Vorstman JA, et al. 22q11.2 deletion syndrome. Nat Rev Dis primers. 2015;1:15071.
Budde M, Anderson-Schmidt H, Gade K, Reich-Erkelenz D, Adorjan K, Kalman JL, et al. A longitudinal approach to biological psychiatric research: The PsyCourse study. Am J Med Genet Part B, Neuropsychiatric Genet. 2019;180:89–102.
Celikkaya B, Durak T, Farooqi AA, Inci K, Tokgun PE, Tokgun O. The effects of MYC on exosomes derived from cancer cells in the context of breast cancer. Chem Biol drug Des. 2023;102:65–75.
van Voorden AJ, Keijser R, Veenboer GJM, Lopes Cardozo SA, Diek D, Vlaardingerbroek JA, et al. EP300 facilitates human trophoblast stem cell differentiation. Proc Natl Acad Sci USA. 2023;120:e2217405120.
Omariba G, Xu F, Wang M, Li K, Zhou Y, Xiao J. Genome-Wide Analysis of MicroRNA-related single nucleotide polymorphisms (SNPs) in mouse genome. Sci Rep. 2020;10:5789.
Heilbronner U, Samara M, Leucht S, Falkai P, Schulze TG. The longitudinal course of schizophrenia across the lifespan: clinical, cognitive, and neurobiological aspects. Harv Rev psychiatry. 2016;24:118–28.
Consortium SPG-WASG. Genome-wide association study identifies five new schizophrenia loci. Nat Genet. 2011;43:969–76.
Sakamoto K, Crowley JJ. A comprehensive review of the genetic and biological evidence supports a role for MicroRNA-137 in the etiology of schizophrenia. Am J Med Genet Part B, Neuropsychiatric Genet. 2018;177:242–56.
Mahmoudi E, Atkins JR, Quidé Y, Reay WR, Cairns HM, Fitzsimmons C, et al. The MIR137 VNTR rs58335419 Is associated with cognitive impairment in schizophrenia and altered cortical morphology. Schizophrenia Bull. 2021;47:495–504.
Forstner AJ, Degenhardt F, Schratt G, Nöthen MM. MicroRNAs as the cause of schizophrenia in 22q11.2 deletion carriers, and possible implications for idiopathic disease: a mini-review. Front Mol Neurosci. 2013;6:47.
de la Morena MT, Eitson JL, Dozmorov IM, Belkaya S, Hoover AR, Anguiano E, et al. Signature MicroRNA expression patterns identified in humans with 22q11.2 deletion/DiGeorge syndrome. Clin immunology. 2013;147:11–22.
Biswas AB, Furniss F. Cognitive phenotype and psychiatric disorder in 22q11.2 deletion syndrome: a review. Res developmental disabilities. 2016;53-54:242–57.
Wang Q, He Q, Chen Y, Shao W, Yuan C, Wang Y. JNK-mediated microglial DICER degradation potentiates inflammatory responses to induce dopaminergic neuron loss. J neuroinflammation. 2018;15:184.
Dungan CM, Valentino T, Vechetti IJ Jr., Zdunek CJ, Murphy MP, Lin AL, et al. Exercise-mediated alteration of hippocampal Dicer mRNA and miRNAs is associated with lower BACE1 gene expression and Aβ(1-42) in female 3xTg-AD mice. J Neurophysiol. 2020;124:1571–7.
Konopka W, Kiryk A, Novak M, Herwerth M, Parkitna JR, Wawrzyniak M, et al. MicroRNA loss enhances learning and memory in mice. J Neurosci. 2010;30:14835–42.
Takousis P, Sadlon A, Schulz J, Wohlers I, Dobricic V, Middleton L, et al. Differential expression of microRNAs in Alzheimer’s disease brain, blood, and cerebrospinal fluid. Alzheimer’s & dementia : the journal of the Alzheimer’s. Association. 2019;15:1468–77.
Moreira HS, Costa AS, Castro SL, Lima CF, Vicente SG. Assessing executive dysfunction in neurodegenerative disorders: a critical review of brief neuropsychological tools. Front aging Neurosci. 2017;9:369.
Patil AH, Halushka MK. miRge3.0: a comprehensive microRNA and tRF sequencing analysis pipeline. NAR genomics Bioinforma. 2021;3:lqab068.
Pliatsika V, Loher P, Magee R, Telonis AG, Londin E, Shigematsu M, et al. MINTbase v2.0: a comprehensive database for tRNA-derived fragments that includes nuclear and mitochondrial fragments from all the cancer genome atlas projects. Nucleic acids Res. 2017;46:D152–59.
Godoy PM, Bhakta NR, Barczak AJ, Cakmak H, Fisher S, MacKenzie TC, et al. Large differences in small RNA composition between human biofluids. Cell Rep. 2018;25:1346–58.
Chen Q, Zhang X, Shi J, Yan M, Zhou T. Origins and evolving functionalities of tRNA-derived small RNAs. Trends biochemical Sci. 2021;46:790–804.
Winek K, Soreq H. Emerging roles of transfer RNA fragments in the CNS. Brain. 2025;148:2631–45.
Mathew BA, Katta M, Ludhiadch A, Singh P, Munshi A. Role of tRNA-Derived fragments in neurological disorders: a review. Mol Neurobiol. 2023;60:655–71.
Fagan SG, Helm M, Prehn JHM. tRNA-derived fragments: A new class of non-coding RNA with key roles in nervous system function and dysfunction. Prog Neurobiol. 2021;205:102118.
Eriksson PS, Perfilieva E, Björk-Eriksson T, Alborn AM, Nordborg C, Peterson DA, et al. Neurogenesis in the adult human hippocampus. Nat Med. 1998;4:1313–7.
Knoth R, Singec I, Ditter M, Pantazis G, Capetian P, Meyer RP, et al. Murine features of neurogenesis in the human hippocampus across the lifespan from 0–100 years. PLoS one. 2010;5:e8809.
Roy NS, Wang S, Jiang L, Kang J, Benraiss A, Harrison-Restelli C, et al. In vitro neurogenesis by progenitor cells isolated from the adult human hippocampus. Nat Med. 2000;6:271–7.
Toda T, Parylak SL, Linker SB, Gage FH. The role of adult hippocampal neurogenesis in brain health and disease. Mol Psychiatry. 2019;24:67–87.
Grońska-Pęski M, Gonçalves JT, Hébert JM. Enriched environment promotes adult hippocampal neurogenesis through FGFRs. J Neurosci. 2021;41:2899–910.
Walgrave H, Balusu S, Snoeck S, Vanden Eynden E, Craessaerts K, Thrupp N, et al. Restoring miR-132 expression rescues adult hippocampal neurogenesis and memory deficits in Alzheimer’s disease. Cell stem Cell. 2021;28:1805–1821.e1808.
Deng LJ, Wu D, Yang XF, Li T. miR-146a-5p Modulates Adult Hippocampal Neurogenesis Deficits Through Klf4/p-Stat3 Signaling in APP/PS1 Mice. Neuroscience. 2023;526:314–25.
Fan C, Li Y, Lan T, Wang W, Long Y, Yu SY. Microglia secrete miR-146a-5p-containing exosomes to regulate neurogenesis in depression. Mol Ther. 2022;30:1300–14.
Pan WL, Chopp M, Fan B, Zhang R, Wang X, Hu J, et al. Ablation of the microRNA-17-92 cluster in neural stem cells diminishes adult hippocampal neurogenesis and cognitive function. FASEB J. 2019;33:5257–67.
Jin J, Kim SN, Liu X, Zhang H, Zhang C, Seo JS, et al. miR-17-92 Cluster regulates adult hippocampal neurogenesis, anxiety, and depression. Cell Rep. 2016;16:1653–63.
Liu Q, Zhang L, Li H. New insights: MicroRNA function in CNS development and psychiatric diseases. Curr Pharmacology Rep. 2018;4:132–44.
Gu X, Xu X, Jia C, Wang J, Zhang J, Gao Q, et al. Molecular mechanisms involved in the regulation of neurodevelopment by miR-124. Mol Neurobiol. 2023;60:3569–83.
Zheng K, Hu F, Zhou Y, Zhang J, Zheng J, Lai C, et al. miR-135a-5p mediates memory and synaptic impairments via the Rock2/Adducin1 signaling pathway in a mouse model of Alzheimer’s disease. Nat Commun. 2021;12:1903.
Smalheiser NR, Lugli G. microRNA regulation of synaptic plasticity. Neuromolecular Med. 2009;11:133–40.
Rahmani S, Kadkhoda S, Ghafouri-Fard S. Synaptic plasticity and depression: the role of miRNAs dysregulation. Mol Biol Rep. 2022;49:9759–65.
McAllan AL, Pillman KA, Gearing LJ, Gantier MP. IsomiR stoichiometry changes as disease biomarkers. Mol Ther Nucleic acids. 2025;36:102578.
van Vliet EA, Scheper M, Mills JD, Puhakka N, Szydlowska K, Ferracin M, et al. Circulating microRNAs and isomiRs as biomarkers for the initial insult and epileptogenesis in four experimental epilepsy models: the EPITARGET study. Epilepsia. 2024;65:3406–20.
Mockly S, Seitz H. Inconsistencies and limitations of current MicroRNA target identification methods. Methods Mol Biol. 2019;1970:291–314.
Godard P, van Eyll J. Pathway analysis from lists of microRNAs: common pitfalls and alternative strategy. Nucleic acids Res. 2015;43:3490–7.
Faber S Mouse Behavioral Tests: Learning & Memory. 2023.
Funding
Urs Heilbronner was supported by European Union’s Horizon 2020 Research and Innovation Program (PSY-PGx, grant agreement No 945151) and the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation, project number 514201724).The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results. Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
ANF: Conceptualization, methodology, evidence synthesis, bioinformatics analysis, writing – original draft, writing – review & editing, and approval of the manuscript. DMK: bioinformatics analysis & supervision, writing – review & editing, and approval of the manuscript. LK: Supervision, writing – review & editing, and approval of the manuscript. AF: Supervision, writing – review & editing, and approval of the manuscript. TGS: Supervision, writing – review & editing, and approval of the manuscript. UH: Conceptualization, supervision, writing – review & editing, and approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Navarro-Flores, A., Krüger, D.M., Kaurani, L. et al. The role of microRNAs in executive functions: a comprehensive review and bioinformatics analysis of human and animal studies. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03623-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03623-2