Abstract
Sustained proliferation is a hallmark of tumor cells. Cancer-associated alternative splicing (AS) events can provide proliferative advantages in tumors, suggesting that identifying aberrant RNA splicing events linked to proliferation in ovarian cancer (OC) may reveal novel therapeutic targets. In this study, we found that small nuclear ribonucleoprotein polypeptide E (SNRPE) was overexpressed in OC, particularly in the proliferative subtype, and indicated worse clinical prognosis. SNRPE knockdown significantly slowed tumor cell proliferation, inducing G1 phase cell cycle arrest and apoptosis. RNA sequencing analysis identified CTP synthase 1 (CTPS1), the rate-limiting factor in the conversion of UTP to CTP, as a critical downstream effector of SNRPE. Mechanistically, SNRPE deficiency led to the retention of intron 15 in CTPS1 mRNA, triggering the degradation of unspliced transcripts through the nonsense-mediated mRNA decay (NMD) pathway and reducing the level of functional CTPS1. Notably, CTPS1 knockdown significantly suppressed the tumor progression driven by SNRPE overexpression. Given that the loss of CTPS2 was prevalent in OC, OC cell proliferation could be more effectively controlled by the suppression of CTPS1. We propose a strategy to regulate CTPS1 expression by modulating its efficient splicing through SNRPE. Consequently, the SNRPE-CTPS1 axis may represent a potential therapeutic target for OC patients.
Similar content being viewed by others
Introduction
Ovarian cancer (OC) ranks first in terms of mortality rate among gynecological malignancies, with approximately 324,398 new cases and 206,839 deaths reported globally in 2022 [1]. More than 70% of OC patients present with advanced-stage disease at diagnosis because of obscure symptoms at onset, rapid progression, and the absence of reliable early detection methods. Despite the development of targeted therapies, primarily antiangiogenic agents and PARP inhibitors, improvements in the prognosis of OC are limited because of its propensity for extensive abdominal metastasis, tumor recurrence, and drug resistance. The overall 5-year survival rate of patients with OC is only 30–50% [2].
Sustained proliferation is a hallmark of tumor cells and represents a pivotal process in the development and progression of cancer [3]. Targeting mechanisms involved in uncontrolled cancer cell proliferation represents an efficient therapeutic strategy [4]. Several targeted compounds against tumor cell proliferation have been developed in recent years [5]. Notably, recent studies have revealed that OC patients with a proliferative subtype have shorter survival but may derive relatively greater therapeutic benefits from anti-angiogenic treatments [6, 7]. Consequently, identifying key molecular drivers underlying OC pathogenesis and progression, particularly those associated with proliferation, elucidating their mechanistic roles, and exploring novel targeted therapeutic strategies are of paramount importance for improving clinical outcomes in OC patients.
Alternative splicing (AS) is a key posttranscriptional regulatory mechanism through which eukaryotic cells increase their transcriptomic and proteomic diversity [8]. Aberrant splicing events are common in cancer and are associated with mutations or altered expression of components related to the splicing machinery [9]. Advances in therapies targeting RNA splicing, particularly small molecule splicing modulators and antisense oligonucleotides (ASOs), have shown preclinical efficacy in correcting pathogenic splicing events in cancer [10]. The dysregulation of AS mediated by spliceosome components and splicing factors has also been explored in OC. For instance, SNRPB promotes OC cell proliferation and invasion through the regulation of aberrant exon skipping of POLA1 and BRCA2 [11]. The splicing factor BUD31 drives OC progression by regulating the expression of anti-apoptotic BCL2L12 [12]. Moreover, the splicing factor WBP11 promotes the malignant progression of OC by mediating the intron retention (IR) of MCM7 [13]. However, understanding of the expression profile of critical spliceosome components in OC is superficial, and investigations of RNA splicing-based therapies in OC remain limited.
Genes encoding spliceosomal core components, which form a heteroheptameric Sm complex, have been shown to be overexpressed in malignant diseases. Among these genes, SNRPE was found to be among the most differentially expressed genes (DEGs) in lung adenocarcinoma, breast cancer, and OC, influencing the cell viability of breast, lung, and melanoma cancer cell lines by modulating cell autophagy via the mTOR pathway [14]. In hepatocellular carcinoma (HCC), SNRPE has been identified as an oncofetal splicing factor that is significantly associated with poor prognosis and drives HCC tumorigenesis through the regulation of FGFR4 pre-mRNA splicing [15]. Additionally, upregulation of SNRPE expression is related to poor prognosis in patients with breast cancer, and it could activate natural killer (NK) cell-mediated antitumor immunity by inducing tumor cell pyroptosis [16]. However, research on SNRPE in OC remains superficial. Using bioinformatics analysis, we preliminarily revealed that SNRPE expression was significantly elevated in OC patients, particularly in the proliferative subtype, and was correlated with poor survival. These findings indicated that SNRPE might serve as a driver of OC progression through its potential influence on tumor cell proliferation, warranting further validation and in-depth investigation into its precise regulatory mechanisms.
Here, we explored the expression levels and prognostic implications of SNRPE in OC on the basis of fresh-frozen tissue and tissue microarray (TMA) samples. A comprehensive functional evaluation of the effects of SNRPE on OC was conducted both in vitro and in vivo. Additionally, RNA sequencing revealed SNRPE-regulated AS events, through which key proliferation-associated downstream targets of SNRPE were identified and the regulatory mechanisms were further elucidated. In conclusion, our study demonstrates the oncogenic role of SNRPE in OC, offering in-depth understanding of its potential as a prognostic marker as well as a novel interventional target for OC patients.
Materials and methods
Bioinformatics analysis
We utilized the TCGA database (http://www.cbioportal.org/) to obtain gene expression profiles of Sm proteins in OC and its subtypes. Pan-cancer SNRPE expression patterns were obtained using TCGA data from the GEPIA3 database (https://gepia3.bioinfoliu.com/). Differential SNRPE expression across FIGO stages and histological grades was analyzed using the CSIOVDB database (http://csiovdb.mc.ntu.edu.tw/CSIOVDB.html). Kaplan-Meier plotter (https://kmplot.com/analysis/) was used to analyze the relationship between SNRPE expression and OC survival outcomes. Heatmap data visualization was conducted with TB tools [17]. Functional annotation, including Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses, was performed using the DAVID bioinformatics platform [18]. RNA-seq data were visualized using Sashimi plots generated by the Integrative Genomics Viewer (IGV) (https://igv.org/), while AS events were analyzed with rMATS software [19]. Transcriptomic information for target genes was retrieved from the Ensembl Genome Browser (https://www.ensembl.org/).
Patients and tissue samples
We collected fresh-frozen OC tissues, normal ovarian tissues, and fallopian tube (FT) tissues from patients at Qilu Hospital for quantitative real-time polymerase chain reaction (qPCR) and Western blot analyses. TMAs derived from the Department of Obstetrics and Gynecology of Qilu Hospital included normal FT and OC samples were used for Immunohistochemistry (IHC) staining and survival analysis. All the participants provided informed consent. The study was approved by the Ethics Committee of Qilu Hospital, Shandong University (Approval No. KYLL-202412-050).
RNA isolation and qPCR
RNA was isolated using the RNA-easy Isolation Reagent (Vazyme, China). Reverse transcription was performed using an All-in-One qRT SuperMix Kit (Vazyme, China). qPCR was conducted on a QuantStudio 3 system (Applied Biosystems, USA) using 2× SYBR qPCR Master Mix (Vazyme, China), with 18S serving as the internal reference. The relative mRNA expression levels of target molecules were calculated using the 2−ΔΔCT method.
Western blotting and antibodies
Proteins were extracted using RIPA lysis buffer supplemented with PMSF. After quantification, the proteins were separated by SDS‒PAGE and transferred onto PVDF membranes. The membrane was blocked and incubated with primary antibodies at 4 °C overnight. The next day, the membrane was incubated with secondary antibodies at room temperature for 1 h. The protein bands were visualized using Enhanced Chemiluminescence (ECL) Reagent (Merck Millipore, Germany), with β-actin serving as the internal control protein. Band intensities were quantified using ImageJ. The primary antibodies used in this study were as follows: SNRPE (Invitrogen, PA5-78173, 1:6000), CTPS1 (Proteintech, 15914-1-AP, 1:2000), β-actin (Sigma‒Aldrich, A5441, 1:6000), and Flag (Sigma‒Aldrich, F1804, 1:1000).
IHC staining analysis
The tissue sections were deparaffinized using xylene and rehydrated with graded ethanol solutions. Antigen retrieval was conducted with EDTA (pH 9.0) or citrate buffer (pH 6.0) according to the instructions. IHC staining was performed using a two-step IHC Kit (Zhongshan Biotechnology, China). The blocking of endogenous peroxidase activity, incubation with antibodies and color development were performed according to the instructions. After counterstaining with hematoxylin, microscopic evaluation was conducted to determine the histochemistry scores (H-scores) on the basis of the staining intensity and the percentage of positive cells. The antibody used for IHC was SNRPE (Invitrogen, PA5-78173, 1:500).
Cell culture
The HEY cell line was obtained from the laboratory of Dr. Liu. The SKOV3 and HEK293T cell lines were procured from the Chinese Academy of Sciences (Shanghai, China). The OVCAR8 cell line was provided by Dr. Ma’s laboratory. HEY, SKOV3, and HEK293T cells were cultured in DMEM, and OVCAR8 cells in RPMI 1640, all supplemented with 10% FBS. All the cells were cultured at 37 °C with 5% CO2.
Transient transfection, plasmid construction, and lentivirus infection
The siRNAs targeting SNRPE or CTPS1 mRNAs were synthesized by GenePharma (Shanghai, China). The specific sequences of the siRNAs are detailed in Tables S1 and S2. The PLKO.1 and PLKO.1-shSNRPE vectors for stable transfection were synthesized by Genepharma (Shanghai, China). The open reading frames (ORFs) encoding SNRPE, CTPS1_L, and CTPS1_S were inserted into the PCMV vector to construct overexpression plasmids. The plasmids were transfected into HEK293T cells to generate lentiviral particles. OC cells were infected with the lentiviruses, followed by selection with puromycin (2 μg/mL) for 5–7 days to establish stable OC cell lines. All transfection experiments were performed using Lipofectamine 2000 (Invitrogen, USA).
MTT assay
After the cells were plated in 96-well plates (500–1000 cells per well) and allowed to adhere to the surface, 20 μL of MTT reagent (5 mg/mL) was added to each well. Following the culture of the cells at 37°C for 4 hours, 100 μL of DMSO was added to dissolve the formazan crystals after the culture medium was removed. The absorbance was measured at 490 nm. Continuous culture and determination were performed for 5–7 consecutive days.
Colony formation assay
Cells were dispersed into 6-well plates (1000–1500 cells per well) and cultured for 10–14 days. The colonies were fixed with methanol, stained with gentian violet, and counted using ImageJ software.
EdU assay
Cells were seeded in 96-well plates (0.5–1 × 10⁴ cells per well) and cultured at 37 °C overnight. The EdU-594 Kit (Beyotime, China) was used for EdU immunofluorescence staining. Images were captured under a fluorescence microscope, and the cells were counted using ImageJ software. The EdU-positive cell ratio (%) = (EdU-positive cells/DAPI-positive cells) × 100%.
Cell migration and invasion assays
In the migration assay, after the Transwell chambers (Corning, USA) were placed in 24-well plates, 200 μL of serum-free cell suspension containing 0.5–1 × 10⁵ cells was seeded into the upper compartment. In the invasion assay, in addition to the aforementioned procedures, 60 μL of Matrigel (Corning, USA) diluted in serum-free medium at a ratio of 1:6 was precoated in the upper chamber. After incubation at 37 °C for 8–72 hours, the migrated or invaded cells in the lower chamber were fixed and stained. Three random fields per well were captured under microscope and counted.
Cell cycle and cell apoptosis assays
Cell cycle assay was performed using the Cell Cycle Staining Kit (MultiSciences Biotech, China) according to the manufacturer’s instructions, and cell apoptosis assay was conducted using the Annexin V-FITC/7-AAD Apoptosis Detection Kit (Goonie Biotech, China). The treated cells were detected using Flow Cytometer (Beckman Colter, USA). The data were analyzed using FlowJo software.
Xenograft models
Four-week-old female BALB/c nude mice (Gem Pharmatech, China) were used to establish xenograft models via subcutaneous injection of HEY cells (5×106 cells in 200 μL of PBS). For functional studies, negative control (PLKO.1-Ctrl) and stable SNRPE-knockdown (sh-SNRPE#1 and sh-SNRPE#2) HEY cells were used. In rescue experiments, HEY cells with negative control (PCMV-Ctrl), stable SNRPE-overexpressing (SNRPE), and stable CTPS1 knockdown following SNRPE-overexpression (SNRPE + sh-CTPS1) were utilized. All the mice were maintained under SPF conditions. Tumor growth was monitored every 3–4 days, and tumor volume was estimated using the formula: length × width²/2. On the basis of tumor progression, the mice were euthanized after 3–4 weeks. Excised tumors were photographed and weighed. The animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of Shandong University (Approval No. 25018).
RNA sequencing analysis
Total RNA was extracted from SKOV3 cells transiently transfected with either si-SNRPE or si-NC after 48 h using TRIzol reagent (Invitrogen, USA). RNA sequencing was conducted by Bioprofile Biotechnology Co., Ltd. (Shanghai, China). DEGs were identified on the basis of the criteria of |log2-fold change | ≥1 and padj < 0.05.
RT‒PCR splicing assay
Reverse transcription polymerase chain reaction (RT‒PCR) was conducted using 2×Taq Master Mix (Vazyme, China). The final PCR products were separated by agarose gel electrophoresis and scanned using a chemiluminescence gel imaging system (General Electric Company, USA). GAPDH was used as the internal reference.
RIP assay
RNA immunoprecipitation (RIP) assays were performed in SKOV3 cells overexpressing Flag-tagged SNRPE using a Magna RIP Kit (Millipore, Germany). Immunoprecipitation was conducted using an anti-Flag antibody (Sigma‒Aldrich, F1804), with IgG serving as the negative control. The expression levels of CTPS1 and FGFR4 mRNA in the immunoprecipitated complexes were analyzed by qPCR.
Statistical analysis
GraphPad Prism 10 was used for statistical analyses. Unpaired t-tests or ANOVA was used to assess differences between groups. The results are expressed as the mean ± SD of three independent experiments. Survival curves were generated using the Kaplan–Meier method, and differences in survival between groups were analyzed by the log-rank test. P < 0.05 was considered to indicate statistical significance (*P < 0.05, ** P < 0.01).
Results
SNRPE is highly expressed in OC and predicts poor prognosis
To explore the potential role of the core components of the spliceosome in the pathogenesis and progression of OC, we evaluated the differential expression of the genes encoding 7 core Sm proteins utilizing transcriptomic data sourced from the TCGA and illustrated the results with a heatmap (Fig. 1A). Compared with those in normal ovarian tissues, the expression levels of SNRPE and SNRPB mRNAs in OC tissues were significantly elevated (Fig. 1B and Fig. S1A). SNRPB has been identified as an important oncogenic driver that promotes OC progression [11]. We also analyzed the differential expression of SNRPE mRNA across various cancer types using TCGA data from the GEPIA3 database. The results confirmed that SNRPE mRNA expression was upregulated in the majority of malignant tumors (Fig. 1C), indicating its potential tumorigenic function. Few studies have investigated the function and underlying mechanism of SNRPE in carcinogenesis. Therefore, we focused our subsequent study on the impact of SNRPE in OC.
A Differential expression of genes encoding the 7 core Sm proteins between normal ovary (n = 8) and OC tissues (n = 585) from TCGA-OV-AffyU133a transcriptomic data, as visualized in a heatmap. B Differential expression analysis of SNRPE and SNRPB mRNAs between normal ovary (n = 8) and OC tissues (n = 585) from TCGA-OV-AffyU133a dataset. C Differential expression analysis of SNRPE mRNA across various cancer types using TCGA data from the GEPIA3 database. D Differential SNRPE mRNA expression levels across OC subtypes in TCGA database, including differentiated (n = 68), immunoreactive (n = 84), mesenchymal (n = 68), and proliferative (n = 79) subtypes. E Differential expression of genes encoding the 7 core Sm proteins in the proliferative subtype of OC based on TCGA database, as visualized in a heatmap. F, G Analysis of SNRPE mRNA expression levels according to the FIGO stage and histological grade of OC using the data from the CSIOVDB database. H Kaplan-Meier survival analysis assessing the impact of SNRPE on the OS (high-expression group, n = 958; low-expression group, n = 477) in OC patients. I qPCR analysis of SNRPE mRNA levels in OC tissues (n = 16) compared with normal FT tissues (n = 10) and normal ovary tissues (n = 10). J, K Western blot analysis of SNRPE protein levels in OC tissues (n = 9) compared with normal FT tissues (n = 8) and normal ovary tissues (n = 8). L Representative images of IHC staining for SNRPE in normal FT and OC based on TMAs. Scale bars: 50 μm. M Comparison of H-Scores of SNRPE from IHC in normal FT (n = 40) and OC (n = 101) samples based on TMAs. N The effect of SNRPE expression on the overall survival of OC patients analyzed by Kaplan-Meier analysis based on TMAs. The median expression according to the H-Scores was used to define high-expression group and low-expression group. The P values were obtained by Log-rank test (H, N), unpaired t-test (B, C, M) or one-way ANOVA (D, F, G, I, K). *P < 0.05, **P < 0.01.
Analysis across different molecular subtypes of OC in the TCGA database revealed that compared with differentiated, immunoreactive, and mesenchymal subtypes, the proliferative subtype exhibited significantly higher levels of SNRPE mRNA expression (Fig. 1D and Fig. S1B). Further comparison of the genes encoding 7 core Sm proteins in the proliferative subtype of OC revealed that SNRPE had the greatest proportion of highly expressed mRNAs (Fig. 1E and Fig. S1C). These results support a potential role for SNRPE in promoting cell proliferation in OC. Additional analysis of data from the CSIOVDB database [20] revealed that compared with patients with stage I disease, patients with FIGO stage III–IV disease had higher SNRPE mRNA expression (Fig. 1F). Compared with that in patients with histological grade 1 (G1) tumors, SNRPE mRNA expression was elevated in patients with grade 2 (G2) and grade 3 (G3) tumors (Fig. 1G). Furthermore, survival analysis using the Kaplan‒Meier plotter database revealed that high SNRPE mRNA expression correlated with poor overall survival (OS) in OC patients (Fig. 1H).
To validate these findings, we assessed the differential expression of SNRPE in fresh-frozen OC, normal ovary, and FT tissues. Both qPCR and Western blot analyses demonstrated that SNRPE expression levels were significantly elevated in OC tissues (Fig. 1I–K). Moreover, immunohistochemical analysis of TMAs from Qilu Hospital revealed that the H-scores of SNRPE were significantly greater in OC tissues than those in normal FT tissues (Fig. 1L, M). Additionally, survival analysis based on our TMA also revealed that high SNRPE expression was associated with poor OS (Fig. 1N).
Collectively, the above data indicate that SNRPE is highly expressed in OC, is correlated with unfavorable clinical prognosis, and plays a potential role in tumor proliferation. These findings revealed the value of SNRPE as a prognostic biomarker and its potential role in OC progression.
SNRPE deficiency suppresses the malignant biological behavior of OC cells
To investigate the biological functions of SNRPE, two independent siRNAs specifically targeting SNRPE mRNA were synthesized and used to suppress SNRPE expression in OC cells via transient transfection. The knockdown efficiency of SNRPE was verified using qPCR and Western blot (Fig. 2A, B), and the proliferative capacity of OC cells was assessed. The results of the MTT and colony formation assays demonstrated that the downregulation of SNRPE expression significantly inhibited the growth rate (Fig. 2C) and colony-forming ability of HEY, SKOV3, and OVCAR8 cells (Fig. 2D and Fig. S2A). EdU assays also revealed a marked reduction in the proportion of EdU-positive cells following SNRPE knockdown, indicating that SNRPE silencing significantly impaired DNA replication activity in OC cells (Fig. 2E, F; Fig. S2B, E, F). Furthermore, flow cytometry analysis demonstrated that SNRPE knockdown induced G1 phase cell cycle arrest (Fig. 2G, H and Fig. S2G, H) and triggered apoptosis (Fig. 2I, J and Fig. S2I, J) in OC cells. Moreover, we evaluated the effect of SNRPE expression on OC cell metastatic potential using Transwell assays. We found that SNRPE silencing significantly suppressed the migratory and invasive capacities of OC cells (Fig. S2C, D, K, L).
A, B The knockdown efficiency of SNRPE in HEY, SKOV3, and OVCAR8 cells was verified by qPCR and Western blot. C The effects of SNRPE knockdown on the growth rate of HEY, SKOV3, and OVCAR8 cells were examined by MTT assays. D The effects of SNRPE knockdown on the colony formation ability of HEY, SKOV3, and OVCAR8 cells were assessed by colony formation assays. E, F The proportion of proliferative cells in HEY and SKOV3 cells after SNRPE silencing were evaluated by EdU incorporation assays. Scale bars: 100 μm. G, H The proportions of G1, S or G2 phase cells among NC and SNRPE-silenced HEY cells were detected via cell cycle assays and flow cytometry. I, J The proportions of apoptotic NC and SNRPE-silenced HEY cells were determined via apoptosis assays and flow cytometry. K Photographs of subcutaneous xenograft tumors in nude mice in PLKO.1-Ctrl group and sh-SNRPE groups (n = 5 mice per group). L, M Comparison of tumor volume and weight between PLKO.1-Ctrl group and sh-SNRPE groups. The P values were obtained by one-way ANOVA. *P < 0.05, **P < 0.01.
Next, we investigated the role of SNRPE in OC tumor progression in vivo via subcutaneous injection of HEY cells, which were transfected with SNRPE-specific shRNA or negative control plasmids, into nude mice. Compared with those in the control group, both the tumor volume and weight in the sh-SNRPE groups were significantly lower (Fig. 2K–M), which indicated that SNRPE knockdown exerted potent antitumor effects on OC cells in vivo, consistent with the in vitro results.
These findings indicate that SNRPE knockdown significantly suppresses the proliferative capacity and metastatic potential of OC cells, providing compelling evidence for the regulatory role of SNRPE in OC progression.
SNRPE-targeted ASO inhibits the malignant progression of OC cells
ASOs are short, synthetic nucleic acid sequences that bind specifically to target mRNAs to modulate gene expression. To examine the potential of ASOs in OC, we synthesized ASOs specifically targeting SNRPE and assessed their effects on the malignant biological behaviors of OC cells.
HEY, SKOV3, and OVCAR8 cells were transfected with SNRPE‑ASOs, and the knockdown efficiency was verified by qPCR and Western blot analyses (Fig. 3A, B). ASO#2, which achieved the most effective suppression of SNRPE among the three OC cell lines, was selected for subsequent functional assays. MTT and colony formation assays demonstrated that SNRPE‑ASO treatment significantly inhibited cell proliferation and colony‑forming ability (Fig. 3C, D). Furthermore, Transwell assays revealed that SNRPE‑ASO markedly reduced the migratory and invasive capacities of OC cells (Fig. 3E–G).
A, B The impact of SNRPE-ASOs on the mRNA and protein expression levels of SNRPE in HEY, SKOV3, and OVCAR8 cells was assessed by qPCR and Western blot. C The effects of SNRPE-ASO#2 on the colony formation ability of HEY, SKOV3, and OVCAR8 cells were assessed by colony formation assays. D The effects of SNRPE-ASO#2 on the proliferative ability of HEY, SKOV3, and OVCAR8 cells were examined by MTT assays. E–G The effects of SNRPE-ASO#2 on the migratory and invasive abilities of HEY, SKOV3, and OVCAR8 cells were assessed by Transwell assays. The P values were obtained by one-way ANOVA (A) or unpaired t-test (C, D, G). *P < 0.05, **P < 0.01.
In summary, SNRPE-ASOs effectively suppressed the malignant biological behaviors of OC cells, consistent with the inhibitory effects observed in siRNA-mediated SNRPE knockdown experiments. These findings provide compelling evidence that SNRPE plays a critical role in the progression of OC.
Identification of SNRPE downstream targets in OC cells
To investigate the molecular mechanism responsible for the oncogenic role of SNRPE in OC, we conducted RNA sequencing to identify DEGs in SKOV3 cells after transient knockdown of SNRPE. In total, 1157 statistically significant DEGs (|log2FC | ≥1, padj < 0.05) were screened, among which 528 genes were upregulated and 629 genes were downregulated following SNRPE inhibition (Fig. 4A, B).
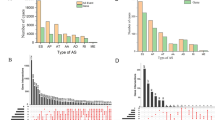
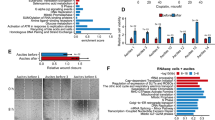
A, B Heatmap and volcano plot based on RNA sequencing data showing DEGs following SNRPE knockdown in SKOV3 cells. C, D Bubble plots illustrating the results of the GO and KEGG pathway analyses of the downregulated DEGs after SNRPE silencing in SKOV3 cells. E Venn diagram illustrating the overlapping genes between downregulated DEGs and IR-related genes after SNRPE knockdown in SKOV3 cells. F KEGG pathway analysis of genes related to differential IR events after SNRPE knockdown in SKOV3 cells. G Heatmap of RNA-seq expression data showing the expression patterns of four genes involved in pyrimidine metabolism following SNRPE knockdown in SKOV3 cells. H Sashimi plot of CTPS1 generated from RNA-seq BAM data, illustrating the splicing patterns following SNRPE knockdown in SKOV3 cells. I Correlation analysis of the relative mRNA expression levels of SNRPE and CTPS1 in fresh-frozen OC tissues (n = 16) from the Qilu cohort. J, K qPCR and Western blot analyses of the effect of SNRPE knockdown on CTPS1 expression levels in HEY, SKOV3 and OVCAR8 cells. The P values were obtained by one-way ANOVA. *P < 0.05, **P < 0.01.
To better understand the biological implications of these DEGs, we conducted GO and KEGG pathway analyses using the DAVID database. The results revealed that the downregulated DEGs were involved primarily in biological processes, such as homologous recombination, DNA metabolic process, cell cycle process, DNA repair, and chromosome organization (Fig. 4C), and pathways, such as the cell cycle, DNA replication, and metabolic pathways were significantly enriched (Fig. 4D).
Given that SNRPE is a core component of the spliceosome, we hypothesized that its dysregulation might lead to extensive AS abnormalities in OC cells. To obtain a global view of splicing alterations induced by SNRPE silencing, we analyzed RNA‑seq data to quantify all major categories of AS events, including exon skipping (SE), alternative 5′ splice sites (A5SS), alternative 3′ splice sites (A3SS), mutually exclusive exons (MXE), and IR.
Using rMATS with thresholds of |IncLevelDifference | > 0.1 and FDR < 0.05, we detected 7,945 significant AS events: SE (5,028), IR (1,101), MXE (844), A3SS (538), and A5SS (434) (Fig. S3A). For each AS category, we performed intersection analyses between genes exhibiting splicing alterations and DEGs in SNRPE‑silenced SKOV3 cells to identify candidates that were both aberrantly spliced and downregulated at the mRNA level (Fig. S3B–E).
We subsequently subjected these candidate genes to KEGG pathway enrichment analysis, which revealed distinct functional associations for different AS types. SE events were enriched in cancer‑related pathways, including metabolic signaling (metabolic pathways, carbon metabolism, pyruvate metabolism), cell cycle regulation, DNA replication, DNA repair (Fanconi anemia, base excision repair, nucleotide excision repair, homologous recombination), and biosynthesis of cofactors (Fig. S3F). MXE events were mainly linked to DNA repair pathways, particularly Fanconi anemia and homologous recombination. A3SS events were associated with metabolic pathways (fructose and mannose metabolism, porphyrin metabolism), biosynthesis of cofactors, and DNA repair processes (Fig. S3G). A5SS events showed enrichment in N‑glycan biosynthesis and general metabolic pathways.
Because IR is increasingly recognized as a critical AS pattern involved in tumor pathogenesis [21], we subsequently focused on IR events regulated by SNRPE in OC cells. By intersecting the 1699 downregulated DEGs (log₂FC ≤ −0.58, padj < 0.05) with the 1101 genes showing differential IR events, we identified 188 overlapping genes (Fig. 4E). KEGG pathway analysis revealed that these genes were enriched in metabolic pathways, biosynthesis of cofactors, the Fanconi anemia pathway, porphyrin metabolism, pyrimidine metabolism, and nucleotide excision repair (Fig. 4F).
Among the identified pathways, pyrimidine metabolism drew our early attention because nucleotide metabolism is a well‑established core process sustaining rapid cancer cell proliferation, and its dysregulation often reflects specific metabolic dependencies. As an essential pathway for DNA replication, RNA synthesis, and cellular bioenergetics, aberrant pyrimidine metabolism is a hallmark of cancer and supports uncontrolled tumor cell growth [22]. OC cells require elevated pyrimidine pools to support rapid proliferation and efficient DNA repair. Our previous work demonstrated that SNRPE expression is upregulated in proliferative OC subtypes and that modulating SNRPE levels significantly impacts OC cell proliferation. These findings led us to investigate the mechanisms by which SNRPE regulates pyrimidine metabolism.
Using RNA-seq analysis, we identified four genes (CAD, CTPS1, UCK1, and NME3) involved in pyrimidine metabolism that were significantly downregulated following SNRPE knockdown (Fig. 4G). To explore the IR events of these genes, we conducted Sashimi plot analysis, and the results demonstrated that only intron 15 of CTPS1 mRNA was significantly retained after SNRPE inhibition (Fig. 4H). We also detected SNRPE and CTPS1 mRNA expression levels in fresh-frozen OC tissues through qPCR and performed Pearson correlation analysis. The results revealed a significant positive correlation between SNRPE and CTPS1 mRNA expression levels in OC (Pearson’s r = 0.7290) (Fig. 4I). These findings supported the hypothesis that SNRPE might play a regulatory role in CTPS1. To validate this hypothesis, CTPS1 expression was assessed in OC cells following SNRPE knockdown. Both mRNA and protein levels of CTPS1 were markedly reduced upon SNRPE depletion (Fig. 4J, K).
In conclusion, we identified CTPS1 as an important downstream target of SNRPE, and the malignant behavior of OC cells driven by SNRPE may be partly mediated through its regulation of CTPS1.
CTPS1 expression is elevated in OC, and CTPS1 knockdown inhibits the malignant phenotypes of OC cells
To explore the role of CTPS1 in OC, we detected the mRNA and protein expression levels of CTPS1 in fresh-frozen OC, normal ovary, and FT tissues. The results demonstrated that CTPS1 expression was significantly elevated in OC tissues (Fig. 5A–C). Next, we knocked down CTPS1 in OC cell lines using siRNAs, validated the knockdown efficiency (Fig. 5D–E), and performed functional experiments. A growth curve analysis via MTT assays revealed that depletion of CTPS1 significantly hindered the proliferative capacity of OC cells (Fig. 5F). Colony formation assays also confirmed that silencing CTPS1 expression significantly weakened the clonogenic potential of OC cells (Fig. 5G and Fig. S4A). EdU incorporation assays demonstrated that silencing CTPS1 markedly decreased the proportion of OC cells with active DNA replication (Fig. 5H, I and Fig. S4B, E, F). Additionally, flow cytometry analysis confirmed that silencing CTPS1 induced cell cycle arrest at G1 phase (Fig. 5J,K and Fig. S4G, H) and triggered apoptosis (Fig. 5L, M and Fig. S4I, J) in OC cells. Furthermore, Transwell assays demonstrated that CTPS1 deficiency significantly suppressed the migratory and invasive abilities of OC cells (Fig. S4C, D, K, L).
A qPCR analysis of CTPS1 mRNA expression levels in OC tissues (n = 16) compared with normal FT tissues (n = 10) and normal ovary tissues (n = 10). B, C Western blot analysis of CTPS1 protein levels in OC tissues (n = 9) compared with normal FT tissues (n = 8) and normal ovary tissues (n = 8). D, E The knockdown efficiency of CTPS1 in HEY, SKOV3, and OVCAR8 cells was verified by qPCR and Western blot assays. F The effects of CTPS1 knockdown on the growth rate of HEY, SKOV3, and OVCAR8 cells were evaluated by MTT assays. G The effects of CTPS1 knockdown on the colony formation ability of HEY, SKOV3, and OVCAR8 cells were assessed by colony formation assays. H–I The proportion of proliferative cells in HEY and SKOV3 cells after CTPS1 silencing was evaluated by EdU incorporation assays. Scale bars: 100 μm. J,K The proportions of G1, S or G2 phase cells among NC and CTPS1-silenced HEY cells were detected via cell cycle assays and flow cytometry. L,MThe proportions of apoptotic cells among the NC and CTPS1-silenced HEY cells were determined via apoptosis assays and flow cytometry. The P values were obtained by one-way ANOVA. *P < 0.05, **P < 0.01.
Collectively, these findings suggest that CTPS1 is highly expressed in OC tissues and plays a crucial role in promoting the proliferative and metastatic capacities of OC cells in vitro.
SNRPE regulates intron 15 retention of CTPS1 pre-mRNA through NMD in OC cells
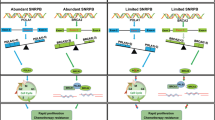
We hypothesized that SNRPE affects the expression of distinct CTPS1 transcripts in OC, which might impair the normal degradation of certain CTPS1 mRNAs and promote the generation of specific functional CTPS1 proteins. Thus, the mRNA transcript data of CTPS1 were retrieved from the Ensembl database (Fig. 6A). CTPS1-217 is the predominant transcript encoding the functional full-length CTPS1 protein (591 amino acids). CTPS1-221 retains intron 15 (where intron 15 refers to the 15th intron in CTPS1-217), which contains a premature termination codon (PTC). This intron-retained mRNA can either be recognized and degraded via the NMD pathway or encode a truncated CTPS1 protein (445 amino acids). As mentioned previously, the Sashimi plot visualization results revealed that the intron 15-retained CTPS1 transcript was increased after SNRPE silencing. These findings suggested that SNRPE might facilitate the splicing of intron 15 on CTPS1 pre-mRNA, leading to the production of functional CTPS1 protein and thereby promoting oncogenic processes in OC.
A Information on the CTPS1-217 (ENST00000650070.2) and CTPS1-221 (ENST00000696108.1) transcripts retrieved from the Ensembl database. B A zoomed view indicating the position of the retained intron 15 in CTPS1 and the products of the normal and abnormal splicing of CTPS1. The schematic also illustrates the locations of the qPCR and RT‒PCR primers. C qPCR analysis of differential expression levels between CTPS1-total and CTPS1-IR in OC tissues (n = 16). D qPCR analysis of CTPS1-IR mRNA levels in OC tissues (n = 16) compared with normal FT tissues (n = 10) and normal ovary tissues (n = 10). E The relative proportion of intron 15-retained CTPS1 transcript to total CTPS1 transcripts after SNRPE inhibition in OC cells through qPCR. F RT‒PCR and agarose gel electrophoresis analysis of the CTPS1 splicing isoforms in OC cells after SNRPE knockdown. G qPCR analysis of the mRNA stability of distinct CTPS1 transcripts in SKOV3 cells treated with actinomycin D. H qPCR analysis of the mRNA stability of total CTPS1 transcripts in SKOV3 cells with SNRPE knockdown or negative control cells. I The expression level of CTPS1 mRNA in SKOV3 cells with SNRPE knockdown or negative control cells that were cotransfected with si-UPF1. J The expression level of CTPS1 mRNA in SKOV3 cells with SNRPE knockdown or negative control cells treated with CHX. K RIP‒qPCR was performed to assess the binding of SNRPE to CTPS1 mRNA in SNRPE-overexpressing SKOV3 cells using an anti-Flag antibody. FGFR4 served as a positive control, and nonspecific IgG was used as a negative control. L The effects of CTPS1_L and CTPS1_S on the proliferation of HEY cells were evaluated by MTT assay. M The effects of CTPS1_L and CTPS1_S on the colony formation ability of HEY cells, as assessed by colony formation assay. The P values were obtained by one-way ANOVA (D, E, L, M) or unpaired t-test (C, G–K). *P < 0.05, **P < 0.01.
We designed specific primers targeting intron 15 of CTPS1 for qPCR to detect the CTPS1-221 transcript (Fig. 6B). Additionally, primers located on exon 9 and exon 10 of CTPS1 were designed for qPCR to detect total CTPS1 transcripts. First, the expression levels of various CTPS1 transcripts in tissues were evaluated by qPCR. The results demonstrated that in OC tissues, the mRNA expression of the intron 15-retained CTPS1 transcript (CTPS1-IR) was significantly lower than that of the total CTPS1 transcripts (Fig. 6C). However, no significant differences in CTPS1-IR mRNA expression were observed among FT, normal ovary, and OC tissues (Fig. 6D). Next, we used qPCR to assess the effect of SNRPE knockdown on CTPS1 mRNA expression levels in OC cell lines. The results revealed a significant increase in the proportion of CTPS1-IR transcripts relative to total CTPS1 transcripts following SNRPE knockdown in OC cells (Fig. 6E). Moreover, we designed specific primers on exon 15 and exon 16 of CTPS1 for RT‒PCR (Fig. 6B). The results of agarose gel electrophoresis demonstrated that SNRPE depletion in OC cells resulted in elevated expression of CTPS1-IR (172 bp), accompanied by reduced expression of normally spliced CTPS1 (93 bp) (Fig. 6F). Quantitative analysis also confirmed a marked increase in the proportion of CTPS1-IR (Fig. S5A). These findings collectively suggest that SNRPE plays a crucial role in facilitating the splicing of intron 15 on CTPS1 pre-mRNA.
To elucidate the molecular mechanism through which SNRPE regulates CTPS1, we performed an RNA stability assay. OC cells were treated with actinomycin D to inhibit transcription, and the expression levels of CTPS1 transcripts were dynamically detected at 0, 2, 4, and 8 h. The results demonstrated significantly reduced stability of the CTPS1-IR transcript (Fig. 6G and Fig. S5B). Additional analysis of the total CTPS1 transcripts also revealed a markedly decreased stability following SNRPE knockdown (Fig. 6H). These results indicated that SNRPE inhibition led to an increase in CTPS1-IR transcripts, which were less stable, ultimately resulting in a reduction in total CTPS1 expression.
The Ensembl database indicated that the intron 15-retained CTPS1 transcript could be degraded via the NMD pathway—a highly conserved cellular surveillance mechanism that identifies and degrades mRNAs containing PTCs to prevent the synthesis of truncated, potentially harmful proteins. To investigate whether SNRPE regulated CTPS1 expression through NMD in OC, we knocked down the expression of UPF1, a key component of the NMD pathway [23]. The results demonstrated that CTPS1 mRNA expression was significantly upregulated in SNRPE-silenced SKOV3 cells (Fig. 6I). Similarly, when NMD was impaired by cycloheximide (CHX) treatment in SNRPE-silenced SKOV3 cells, CTPS1 mRNA levels markedly increased (Fig. 6J). These data indicated that SNRPE depletion induced IR in CTPS1 pre-mRNA, with the resulting transcripts being susceptible to degradation by the NMD pathway.
Because SNRPE is a spliceosomal core protein, we next examined whether its depletion altered the levels of Sm proteins and spliceosomal snRNAs. qPCR analysis of SKOV3 and HEY cells revealed that SNRPE knockdown reduced the levels of multiple Sm-class snRNAs, including U1, U2, U4, U11, and U12, with U4 snRNA exhibiting the most pronounced decrease, but the expression of other Sm protein genes showed no significant difference (Fig. S5C). Given the essential role of SNRPE as a core component of the heptameric Sm ring in stabilizing Sm-class snRNAs, we propose that its depletion disrupts Sm ring assembly and leads to reduced U4 snRNA levels. U4 is required to pair with U6 and subsequently assemble with U5 into the U4/U6•U5 tri-snRNP, a critical pre-catalytic spliceosome subunit necessary for accurate 5′ splice site recognition [24]. Impaired tri-snRNP formation could plausibly explain the increased intron 15 retention observed in CTPS1 pre-mRNA and the consequent reduction in CTPS1 protein expression following SNRPE knockdown.
Moreover, RIP assays were performed using anti‑Flag antibodies in SKOV3 cells overexpressing Flag‑tagged SNRPE. FGFR4, a transcript previously shown to associate with SNRPE [15], was included as a positive control to confirm that the assay could detect known SNRPE‑binding targets. IgG was used as a negative control. As expected, CTPS1 mRNA was significantly enriched in the SNRPE IP (Fig. 6K), indicating that CTPS1 mRNA bound to the SNRPE protein in OC cells. These results suggested that SNRPE maintained the stability and high expression level of CTPS1 by regulating the efficient splicing of intron 15, thereby promoting OC progression.
To investigate the difference in biological functions between the two CTPS1 transcripts in OC cells, we constructed two plasmids—CTPS1_L (overexpressing normally spliced CTPS1) and CTPS1_S (overexpressing intron 15-retained CTPS1) —which were transfected into OC cells with endogenous CTPS1 knockdown by siRNAs. MTT and colony formation assays demonstrated that compared with the control treatment, CTPS1_L significantly increased the proliferation capacity of OC cells, whereas CTPS1_S resulted in relatively minimal differences (Fig. 6L, M). These findings supported the stronger oncogenic effects of normally spliced CTPS1.
CTPS1 mediates the malignant phenotypes of SNRPE in OC cells
To validate the role of CTPS1 as a key downstream effector of SNRPE in OC progression, rescue experiments were performed. We established stable SNRPE-overexpressing OC cells and validated the overexpression efficiency by qPCR and Western blot (Fig. 7A, B). CTPS1 was subsequently knocked down through transient siRNA transfection in these SNRPE-overexpressing cells. The proliferative capacity of OC cells was assessed by MTT and colony formation assays. SNRPE overexpression increased the growth rate and colony-forming capacity of OC cells, whereas CTPS1 depletion significantly attenuated these abilities (Fig. 7C, D and Fig. S6A). Furthermore, Transwell assays revealed that SNRPE overexpression increased the migratory and invasive abilities of OC cells, whereas CTPS1 knockdown markedly suppressed this metastatic potential (Fig. 7E, F). To evaluate tumorigenic potential in vivo, HEY cell lines stably overexpressing SNRPE were transfected with CTPS1-specific shRNA plasmids to achieve stable knockdown of CTPS1 expression. Three groups of cells— (1) HEY cells with stable SNRPE overexpression plus CTPS1 knockdown (SNRPE + sh-CTPS1 group), (2) HEY cells with stable SNRPE overexpression alone (SNRPE group), and (3) control HEY cells (PCMV-Ctrl group) —were subsequently subcutaneously injected into nude mice to establish a xenograft model (n = 5 per group). SNRPE-overexpressing OC cells significantly increased in tumor volume and weight. Conversely, CTPS1 knockdown markedly attenuated the tumor-promoting effects of SNRPE in vivo (Fig. 7G–I).
A, B The overexpression levels of SNRPE mRNA and protein (Flag-tagged) in HEY and SKOV3 cells were verified by qPCR and Western blot. C, D MTT and colony formation assays demonstrated that CTPS1 depletion significantly attenuated the enhanced proliferative and colony-forming capacities induced by SNRPE overexpression in HEY and SKOV3 cells. E, F Transwell assays demonstrated that CTPS1 knockdown markedly suppressed the enhanced migratory and invasive potential induced by SNRPE overexpression in HEY and SKOV3 cells. G Xenograft assays demonstrated that CTPS1 knockdown markedly weakened the enhanced tumorigenic capacity of OC cells induced by SNRPE overexpression in vivo. H, I Comparison of tumor volume and weight among the three groups shown in Fig. 7G. The P values were obtained by unpaired t-test (A) or one-way ANOVA (C, F, H, I). *P < 0.05, **P < 0.01.
In addition, to investigate whether CTPS1 overexpression can rescue or reverse the inhibitory effects of SNRPE knockdown on tumor progression, we performed rescue experiments in HEY cells in which CTPS1 was ectopically overexpressed in the context of SNRPE knockdown. MTT and colony formation assays demonstrated that CTPS1 overexpression partially restored cell proliferation and alleviated the suppression of proliferation and clonogenic capacity induced by SNRPE depletion (Fig. S6B, C). Transwell assays revealed that the impaired migratory and invasive abilities due to SNRPE depletion could be partially restored by CTPS1 overexpression (Fig. S6D).
Collectively, these findings indicate that CTPS1 is a critical downstream effector through which SNRPE exerts its tumor‑promoting role in OC.
Discussion
This study represents the comprehensive investigation of SNRPE expression levels, clinical significance, biological functions, and regulatory mechanisms in OC. We found that SNRPE was highly expressed in OC and was associated with poor clinical prognosis. SNRPE knockdown significantly suppressed cell proliferation, induced G1 phase arrest and apoptosis, and impaired the metastatic potential of OC cells. Mechanistically, SNRPE regulated the efficient splicing of intron 15 in CTPS1 pre-mRNA, maintained high expression of CTPS1 by reducing NMD pathway-mediated degradation, and thereby promoted OC progression. CTPS1 depletion significantly suppressed the effect of SNRPE on tumor progression. Overall, we identified SNRPE as a critical driver that promotes the malignant progression of OC through the regulation of CTPS1 pre-mRNA AS, suggesting its potential as a novel therapeutic target.
SNRPE is one of the seven Sm proteins [25]. Sm proteins bind to uridine-rich small nuclear RNAs to form small nuclear ribonucleoprotein particles, which collectively constitute the spliceosome complex involved in the splicing process of pre-mRNAs [26]. Dysregulation of Sm proteins is associated with aberrant splicing, leading to unexpected mRNA variants from a single gene, and may play an oncogenic role in multiple malignancies [27]. Our study revealed aberrantly high SNRPE expression in OC tissues and its positive correlation with poor prognosis. The upregulated expression of SNRPE has also been observed in breast cancer, HCC, lung cancer, and prostate cancer, typically correlating with poor survival [14,15,16, 28, 29]. These findings are consistent with our observations in OC, suggesting that SNRPE may serve as an oncogenic driver. Notably, SNRPE expression was significantly higher in the proliferative subtype than in the other OC subtypes, suggesting that SNRPE might regulate the proliferation process of OC cells. Both in vitro and in vivo experiments further demonstrated that SNRPE promoted the rapid growth of OC cells. Additionally, SNRPE inhibition induced G1 phase arrest and apoptosis in OC cells. However, autophagy has been observed in SNRPE-knockdown breast cancer cells [14], whereas in HCC cells, SNRPE inhibition leads to G2/M phase arrest and apoptosis [15]. These discrepancies may arise from tissue-specific variations in the frequency and patterns of AS, which could result in functional diversification of SNRPE across different cancer types [15].
RNA sequencing analysis revealed CTPS1 as a key downstream effector of SNRPE. CTP synthetase serves a rate-limiting role in de novo CTP synthesis and is required for the formation of RNA, DNA, and phospholipids [30], which are essential for tumor cell proliferation. There are two isoforms of CTPS: CTPS1 is the main isoform that participates in cell proliferation, whereas CTPS2 plays a compensatory role when CTPS1 is deficient [31]. In rapidly proliferating cells, particularly in cancers and immune-related diseases, both CTPS activity and CTP nucleotide levels frequently present upregulation [32]. In triple-negative breast cancer, CTPS1 expression is significantly upregulated and correlated with poor survival outcomes, and it is activated by YBX1 to promote tumor progression [33]. High expression levels of CTPS1 were linked to advanced disease and poor outcomes in multiple myeloma; in addition, a selective inhibitor of CTPS1 effectively inhibited cell proliferation and led to cell death through apoptosis [34]. Here, we demonstrated elevated expression of CTPS1 and a positive correlation between CTPS1 and SNRPE expression in OC tissues. Silencing CTPS1 effectively attenuated the malignant biological potential of OC cells. Notably, consistent with SNRPE inhibition, CTPS1 knockdown induced cell cycle arrest at G1 phase and apoptosis. Furthermore, CTPS1 knockdown in SNRPE-overexpressing OC cells significantly attenuated the tumor-promoting effects of SNRPE, further indicating that CTPS1 serves as a crucial downstream effector that mediates SNRPE-driven OC progression. Notably, a recent study revealed that OC tumors with CTPS2 deficiency are significantly sensitive to CTPS1 inhibitors (STP-B), and that CTPS2 deficiency occurs in up to 24% of OC cases [35]. These results identify CTPS1 as a promising therapeutic target for OC, meriting further investigation.
Aberrant RNA splicing is recognized as a molecular characteristic that is present in almost all tumor types. Splicing dysregulation can drive tumorigenesis through multiple mechanisms, which contribute to malignant biological characteristics, such as enhanced cell proliferation, decreased apoptosis, and evasion of immune surveillance [36]. For instance, SRSF1-regulated AS of the RPS6KB1 gene is associated with persistent cell proliferation and tumor growth [37]. As a common mode of AS, IR also represents a widespread mechanism of tumor development [9]. Intron-retained mRNAs play an undeniable role in diverse diseases, including cancer [21, 38]. For example, SNRPD2 regulates the IR of DDX39A to promote HCC oncogenesis [27]. PABPN1 contributes to cell proliferation and DNA damage repair through the IR of SNHG19 in hepatoblastoma (HB) [39]. Intron-retained sequences typically contain PTCs, resulting in the removal of IR transcripts by surveillance mechanisms, including NMD, thereby preventing the production of potentially deleterious proteins [38]. In rare instances, IR transcripts may also undergo translation, generating small quantities of functionally relevant truncated protein isoforms through an in-frame stop codon within the retained intron [40]. Here, we demonstrate that SNRPE ensures the expression of functional CTPS1 by facilitating the efficient splicing of intron 15 in CTPS1 pre-mRNA. Conversely, depletion of SNRPE leads to the accumulation of CTPS1 transcripts that retain intron 15, which exhibit reduced stability because of PTC introduction and subsequent degradation via the NMD pathway, ultimately attenuating the malignant phenotype of OC cells. These findings further indicate that upregulated SNRPE in OC plays an oncogenic role by promoting the splicing of CTPS1 pre-mRNA.
At the molecular level, SNRPE is a core component of the heptameric Sm ring, which assembles around the Sm site of Sm-class snRNAs and is essential for their cytoplasmic maturation, stability, and import into the nucleus. In our study, qPCR analysis of SKOV3 and HEY cells revealed that SNRPE depletion selectively impaired snRNA biogenesis—particularly U4 snRNA—rather than broadly affecting Sm-protein transcription. U4 snRNA plays a pivotal role in pairing with U6 snRNA to form the U4/U6 di-snRNP, which subsequently associates with U5 snRNP to create the U4/U6•U5 tri-snRNP—a critical precatalytic spliceosomal subunit required for accurate 5′ splice-site recognition [24]. While we observed a substantial reduction in U4 snRNA, we explicitly acknowledge that this is an expression-level finding. Logically, such depletion is expected to hinder tri-snRNP formation and thereby compromise spliceosome integrity. This inference is strongly supported by prior reports demonstrating that deficiency or mutation of SNRPE disrupts Sm ring assembly and impairs pre-mRNA splicing in human diseases such as non-syndromic microcephaly [41]. Furthermore, destabilization of Sm-ring components leads to selective snRNA instability and mislocalization [42], ultimately arresting spliceosome progression at the precatalytic stage [43]. However, recognizing the preliminary nature of these mechanics, we categorize this proposed pathway as a “functional working model” rather than a definitive structural proof. A mechanistic pitfall in our current work is that we have not yet provided high-resolution structural insights or direct biochemical assays to visualize the specific failure of tri-snRNP assembly. Future studies will aim to directly evaluate tri-snRNP assembly and spliceosomal subcomplex dynamics to further substantiate this proposed mechanism.
Based on our findings, loss of SNRPE function led to intron 15 retention in CTPS1 pre‑mRNA, generating a PTC and activating NMD. The resulting reduction in CTPS1 protein levels is likely to impair the de novo nucleotide biosynthesis pathway, consequently depleting nucleotide pools, hindering DNA replication, inducing G1 phase arrest, and promoting apoptosis. This sequence of events provides a plausible mechanistic link between SNRPE depletion, spliceosome dysfunction, and tumor growth suppression.
Building on these mechanistic insights, it is crucial to evaluate the translational implications of modulating SNRPE in a therapeutic setting. While targeting such a central spliceosomal component could potentially suppress oncogenic splicing programs, its intrinsic role within the Sm ring raises significant concerns regarding specificity, global splicing disruption, and potential toxicity. To address these challenges, we propose a two‑tiered therapeutic strategy: (i) upstream modulation using safe, FDA‑approved small molecules to selectively attenuate SNRPE‑driven pathological splicing functions—drawing on the precedent of the SNRPD2–digitoxin study [27]—and (ii) downstream precision interference with aberrant CTPS1 intron 15 splicing via ASOs or splice‑modulating CRISPR tools to induce IR, activate NMD, and lower CTPS1 protein abundance. Such a strategy is designed to improve target specificity, enhance translational potential, and minimize the disruption of global splicing networks.
Despite the promising findings, several limitations in our study merit discussion. First, regarding the breadth of downstream targets, our study primarily focused on the interaction between SNRPE and CTPS1. While we confirmed that CTPS1 is a critical effector, SNRPE is a fundamental component of the core splicing machinery. Therefore, its silencing likely triggers a broad perturbation of the global splicing landscape rather than affecting a single gene. It is plausible that SNRPE depletion induces widespread AS events—particularly IR—in other gene sets critical for OC progression. Second, regarding mechanistic depth, while we identified CTPS1 via RNA-seq, we lacked high-resolution maps of SNRPE-RNA interactions. The specific motifs recognized by SNRPE in OC cells and the distinctions between direct binding versus indirect effects mediated by spliceosome assembly defects remain to be fully elucidated. As proposed regarding our future directions, studies integrating UV crosslinking and immunoprecipitation followed by sequencing (CLIP-seq) with transcriptomic analysis are needed. This multi-omics approach will be essential to identify alternate transcript targets, construct a comprehensive SNRPE-regulated splicing landscape, and determine whether SNRPE possesses non-canonical functions beyond its traditional role in the Sm ring.
In summary, SNRPE expression is upregulated in OC and correlates with adverse clinical outcomes. Functionally, SNRPE promotes OC progression by facilitating the precise splicing of intron 15 in CTPS1 pre‑mRNA. Inhibition of SNRPE and its downstream effector CTPS1 markedly reduced the proliferative and metastatic potential of OC cells, while inducing G1 phase arrest and apoptosis. Collectively, these findings deepen our understanding of SNRPE‑mediated oncogenic mechanisms in OC and underscore the therapeutic potential of targeting the SNRPE–CTPS1 axis as a novel precision strategy for OC treatment.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74:229–63.
Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. 2025;75:10–45.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Feitelson MA, Arzumanyan A, Kulathinal RJ, Blain SW, Holcombe RF, Mahajna J, et al. Sustained proliferation in cancer: mechanisms and novel therapeutic targets. Semin Cancer Biol. 2015;35:S25–S54.
Patra D, Bhavya K, Ramprasad P, Kalia M, Pal D. Anti-cancer drug molecules targeting cancer cell cycle and proliferation. Adv Protein Chem Struct Biol. 2023;135:343–95.
Konecny, GE, Wang, C, Hamidi, H, Winterhoff, B, Kalli, KR, Dering, J, et al. Prognostic and therapeutic relevance of molecular subtypes in high-grade serous ovarian cancer. J Natl Cancer Inst. 2014;106:249.
Kommoss S, Winterhoff B, Oberg AL, Konecny GE, Wang C, Riska SM, et al. Bevacizumab may differentially improve ovarian cancer outcome in patients with proliferative and mesenchymal molecular subtypes. Clin Cancer Res. 2017;23:3794–801.
Graveley BR. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 2001;17:100–7.
Bonnal SC, Lopez-Oreja I, Valcarcel J. Roles and mechanisms of alternative splicing in cancer - implications for care. Nat Rev Clin Oncol. 2020;17:457–74.
Anczukow O, Allain FH, Angarola BL, Black DL, Brooks AN, Cheng C, et al. Steering research on mRNA splicing in cancer towards clinical translation. Nat Rev Cancer. 2024;24:887–905.
Li Y, Chen Z, Peng J, Yuan C, Yan S, Yang N, et al. The splicing factor SNRPB promotes ovarian cancer progression through regulating aberrant exon skipping of POLA1 and BRCA2. Oncogene. 2023;42:2386–401.
Wang Z, Wang S, Qin J, Zhang X, Lu G, Liu H, et al. Splicing factor BUD31 promotes ovarian cancer progression through sustaining the expression of anti-apoptotic BCL2L12. Nat Commun. 2022;13:6246.
Wei Y, Chen Z, Li Y, Song K. The splicing factor WBP11 mediates MCM7 intron retention to promote the malignant progression of ovarian cancer. Oncogene. 2024;43:1565–78.
Quidville V, Alsafadi S, Goubar A, Commo F, Scott V, Pioche-Durieu C, et al. Targeting the deregulated spliceosome core machinery in cancer cells triggers mTOR blockade and autophagy. Cancer Res. 2013;73:2247–58.
Wu Q, Liao R, Miao C, Hasnat M, Li L, Sun L, et al. Oncofetal SNRPE promotes HCC tumorigenesis by regulating the FGFR4 expression through alternative splicing. Br J Cancer. 2024;131:77–89.
Ji Z, Wang Z, Guo X, Li J, Cai Y, Li K. Targeting SNRPE to induce pyroptosis enhances antitumor immunity in breast cancer. Int J Med Sci. 2025;22:2419–33.
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13:1194–202.
Sherman BT, Hao M, Qiu J, Jiao X, Baseler MW, Lane HC, et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022;50:W216–W221.
Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN, et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci USA. 2014;111:E5593–601.
Tan TZ, Yang H, Ye J, Low J, Choolani M, Tan DS, et al. CSIOVDB: a microarray gene expression database of epithelial ovarian cancer subtype. Oncotarget. 2015;6:43843–52.
Monteuuis G, Schmitz U, Petrova V, Kearney PS, Rasko JEJ. Holding on to Junk Bonds: Intron Retention in Cancer and Therapy. Cancer Res. 2021;81:779–89.
Siddiqui A, Ceppi P. A non-proliferative role of pyrimidine metabolism in cancer. Mol Metab. 2020;35:100962.
Kim YK, Maquat LE. UPFront and center in RNA decay: UPF1 in nonsense-mediated mRNA decay and beyond. RNA. 2019;25:407–22.
Nguyen TH, Galej WP, Bai XC, Savva CG, Newman AJ, Scheres SH, et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature. 2015;523:47–52.
Salgado-Garrido J, Bragado-Nilsson E, Kandels-Lewis S, Seraphin B. Sm and Sm-like proteins assemble in two related complexes of deep evolutionary origin. EMBO J. 1999;18:3451–62.
Zhou Z, Licklider LJ, Gygi SP, Reed R. Comprehensive proteomic analysis of the human spliceosome. Nature. 2002;419:182–5.
Chang C, Li L, Su L, Yang F, Zha Q, Sun M, et al. Intron retention of DDX39A driven by SNRPD2 is a crucial splicing axis for oncogenic MYC/spliceosome program in hepatocellular carcinoma. Adv Sci (Weinh). 2024;11:e2403387.
Tamura K, Furihata M, Tsunoda T, Ashida S, Takata R, Obara W, et al. Molecular features of hormone-refractory prostate cancer cells by genome-wide gene expression profiles. Cancer Res. 2007;67:5117–25.
Liu J, Gu L, Zhang D, Li W. Determining the prognostic value of spliceosome-related genes in hepatocellular carcinoma patients. Front Mol Biosci. 2022;9:759792.
Thangadurai S, Bajgiran M, Manickam S, Mohana-Kumaran N, Azzam G. CTP synthase: the hissing of the cellular serpent. Histochem Cell Biol. 2022;158:517–34.
Minet, N., Boschat, AC, Lane, R, Laughton, D, Beer, P, Asnagli, H, et al. Differential roles of CTP synthetases CTPS1 and CTPS2 in cell proliferation. Life Sci Alliance. 2023;6:e202302066.
Carcamo WC, Satoh M, Kasahara H, Terada N, Hamazaki T, Chan JY, et al. Induction of cytoplasmic rods and rings structures by inhibition of the CTP and GTP synthetic pathway in mammalian cells. PLoS One. 2011;6:e29690.
Lin Y, Zhang J, Li Y, Guo W, Chen L, Chen M, et al. CTPS1 promotes malignant progression of triple-negative breast cancer with transcriptional activation by YBX1. J Transl Med. 2022;20:17.
Pfeiffer C, Grandits AM, Asnagli H, Schneller A, Huber J, Zojer N, et al. CTPS1 is a novel therapeutic target in multiple myeloma which synergizes with inhibition of CHEK1, ATR or WEE1. Leukemia. 2024;38:181–92.
Beer PA, Thomson JP, Gourley C, Meynert A, Parker AE, Schwartz B, et al. Vulnerability to selective CTPS1 inhibition in cancers lacking CTPS2. Cancer Res. 2025;85:1620.
Bradley RK, Anczukow O. RNA splicing dysregulation and the hallmarks of cancer. Nat Rev Cancer. 2023;23:135–55.
Karni R, de Stanchina E, Lowe SW, Sinha R, Mu D, Krainer AR. The gene encoding the splicing factor SF2/ASF is a proto-oncogene. Nat Struct Mol Biol. 2007;14:185–93.
Wong JJ, Au AY, Ritchie W, Rasko JE. Intron retention in mRNA: no longer nonsense: known and putative roles of intron retention in normal and disease biology. Bioessays. 2016;38:41–9.
Zhen N, Zhu J, Mao S, Zhang Q, Gu S, Ma J, et al. Alternative splicing of lncRNAs from SNHG family alters snoRNA expression and induces chemoresistance in hepatoblastoma. Cell Mol Gastroenterol Hepatol. 2023;16:735–55.
Taylor R, Hamid F, Fielding T, Gordon PM, Maloney M, Makeyev EV, et al. Prematurely terminated intron-retaining mRNAs invade axons in SFPQ null-driven neurodegeneration and are a hallmark of ALS. Nat Commun. 2022;13:6994.
Chen T, Zhang B, Ziegenhals T, Prusty AB, Frohler S, Grimm C, et al. A missense mutation in SNRPE linked to non-syndromal microcephaly interferes with U snRNP assembly and pre-mRNA splicing. PLoS Genet. 2019;15:e1008460.
Becker D, Hirsch AG, Bender L, Lingner T, Salinas G, Krebber H. Nuclear Pre-snRNA export is an essential quality assurance mechanism for functional spliceosomes. Cell Rep. 2019;27:3199–3214.
Fu, X. and Hoskins, AA. Dynamics and evolutionary conservation of B complex protein recruitment during spliceosome activation. Nucleic Acids Res. 2025;53:606642.
Acknowledgements
This work was supported by the Tai-Shan Scholar Project of Shandong Province (No. ts20070743), the National High-Level Medical Talent Program (National Distinguished Physician), and Shandong Provincial Natural Science Foundation (Grant No. ZR2023MH183).
Author information
Authors and Affiliations
Contributions
Conception and design: Beihua Kong and Yingwei Li. Methodology: Beihua Kong and Yingwei Li. Acquisition of data: Yingying Pu. Analysis and interpretation of data: Yingying Pu and Yingwei Li. Technical or material support: Zhongshao Chen, Qianqian Gao, Yanling Liu and Ning Yang. Study supervision: Beihua Kong and Yingwei Li. Writing, review, and/or revision of the manuscript: Yingying Pu and Yingwei Li. Final approval: All authors.
Corresponding authors
Ethics declarations
Ethics
Informed consent was obtained from all participants. Ethical approval was granted by the Ethics Committee of Qilu Hospital, Shandong University (KYLL-202412-050). The animal study was approved by the Institutional Animal Care and Use Committee (IACUC) of Shandong University (25018). All methods were performed in accordance with the relevant guidelines and regulations.
Competing interests
The authors have declared no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pu, Y., Chen, Z., Gao, Q. et al. Spliceosomal component SNRPE drives cell proliferation by regulating CTP synthase 1 mRNA splicing in ovarian cancer. Oncogene (2026). https://doi.org/10.1038/s41388-026-03764-2
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41388-026-03764-2