Abstract
The molecular mechanisms that link stress and biological rhythms still remain unclear. The habenula (Hb) is a key brain region involved in regulating diverse types of emotion-related behaviours while the suprachiasmatic nucleus (SCN) is the body’s central clock. To investigate the effects of chronic social stress on transcription patterns, we performed gene expression analysis in the Hb and SCN of stress-naïve and stress-exposed mice. Our analysis revealed a large number of differentially expressed genes and enrichment of synaptic and cell signalling pathways between resilient and stress-naïve mice at zeitgeber 16 (ZT16) in both the Hb and SCN. This transcriptomic signature was nighttime-specific and observed only in stress-resilient mice. In contrast, there were relatively few differences between the stress-susceptible and stress-naïve groups across time points. Our results reinforce the functional link between circadian gene expression patterns and differential responses to stress, thereby highlighting the importance of temporal expression patterns in homoeostatic stress responses.
Similar content being viewed by others
Introduction
Mood disorders are complex heterogeneous disorders that affect cognition, emotional processing and basic physiological functioning [1]. There is growing evidence of a tight connection between circadian rhythms and mood regulation [2]. A majority of depressed patients show more severe symptoms in the morning than in the evening [3]. Diurnal processes including diurnal rhythms in secretion of growth hormones and melatonin, daily core body temperature, and sleep/wake cycles are disrupted in some depressed patients [4, 5]. These internal rhythmic processes are maintained on a 24-h cycle by multiple peripheral clocks throughout the body which are synchronised to a single central principal clock in the mammalian hypothalamus—the suprachiasmatic nucleus (SCN) [6]. The SCN itself has been shown to affect depressive and anxious-like responses in mice [7, 8]. Furthermore, the SCN projects to several hypothalamic structures that regulate diurnal processes disrupted in depression, including the paraventricular nucleus, subparaventricular zone, and dorsomedial hypothalamus [4, 9]. Overall, direct and indirect SCN projections to mood-regulating brain areas are likely responsible for the diurnal rhythmicity of symptoms observed in depressed patients.
One such mood-regulating brain area that is thought to receive inputs from the SCN is the habenular complex (Hb), which is composed predominantly of lateral (LHb) and medial habenula (MHb) subregions [10]. The Hb relays information between the forebrain emotion-processing limbic system and the midbrain depression-associated aminergic centres [11]. The Hb is heavily involved in mediating stress responses [12, 13] and has also been associated with circadian modulation of depression [14]. Elevated diurnal firing in LHb neurons projecting to the serotonergic dorsal raphe nucleus (DRN) is associated with social avoidance in chronically stressed mice [15], while passive coping, a behaviour mirroring apathy and resignation in humans, has been associated with increased activity in numerous circuits, including: LHb-DRN projections [16], medial prefrontal cortex (mPFC) projections to LHb [17], and LHb projections to the ventral tegmental area (VTA) [18]. The pharmacological inhibition of the LHb rescues depressive-like behaviour [19, 20]. In contrast, MHb lesion impairs voluntary exercise and induces despair-like and anhedonic behavioural phenotypes in mice, without affecting the circadian period and sleep architecture [21,22,23]. Despite the apparent importance of Hb in mood regulation, the effects of stress on molecular pathways in the Hb remain underexplored.
Both LHb and MHb exhibit an independent endogenous molecular clock that is synchronised with the SCN clock [10, 24,25,26] which drives the daily variation in their firing [14, 24, 25]. Hb receives input from the SCN indirectly through the dorsomedial and lateral hypothalamic areas [27,28,29]. Some evidence suggests that SCN innervates the LHb directly [14, 26, 30]. Additionally, the Hb complex encompasses the caudal perihabenular region (PHb), which receives direct projections from the retina and has been shown to drive circadian-gated light modulation of a subset of depressive-like behaviours in mice, independently of the SCN [31,32,33,34]. Although the PHb does not respond to four-day social defeat stress [31], PHb-dependent light modulation of behaviour occurs through its projections to nucleus accumbens (NAc), mPFC and thalamic reticular nucleus – areas associated with mood, stress-coping and sleep regulation [31, 34, 35]. Hence, the source of depression-associated diurnal rhythm disruption might be due to perturbations of circadian rhythms at the level of the SCN, the Hb or both.
In order to investigate the relationship between mood regulation and diurnal rhythm disruption, we exposed mice to chronic social defeat stress (CSDS). CSDS has two important advantages over other models. Firstly, CSDS is a robust and widely used chronic stress model that induces depressive-like and PTSD-like symptoms in a subset of exposed rodents [36, 37]. Experimental mice exhibiting social avoidance are deemed as ‘stress-susceptible’, while those that do not are ‘stress-resilient’ [38]. These two subpopulations show different behavioural, physiological and molecular profiles, allowing for investigations of the underlying mechanisms leading to stress susceptibility or resilience [36]. CSDS in rodents disrupts core body temperature, locomotor activity rhythms, sleep architecture, and other basic physiological functioning, with some biological processes being disrupted in all stress-exposed mice and others only in the stress-susceptible phenotype [39,40,41,42,43]. As a social form of stress, it has higher translational value for modelling human mood disorders [37]. The other important reason for using the CSDS paradigm is that this model utilises a predictable stress exposure with standardised protocol and timing, unlike chronic mild stress (CMS). Hence, it is more suitable for investigation of time-dependent biological processes.
Few studies have explored the effect of CMS on core clock gene expression [44, 45], but the differences in core clock gene expression patterns in stress-resilient and stress-susceptible mice have not been investigated. CSDS was found to increase the amplitude of Per2 diurnal expression in the SCN [43]. Moreover, LHb cells are responsive to intermittent social defeat [31]. Specifically, LHb cells projecting to the DRN in stress-susceptible mice exhibit blunted diurnal activity compared to stress-resilient and stress-naïve mice [15]. Despite these earlier findings suggesting a significant interaction between diurnal rhythms and chronic stress, the temporal transcriptomic profiles of the SCN and Hb in the stress-resilient and stress-susceptible mice remain unexplored.
In this study, we investigated the effects of CSDS on the SCN and Hb transcriptome across the light/dark cycle. Mice exposed to CSDS were sampled at four time points: ZT4 (mid-inactive phase), ZT10 (transition to active phase), ZT16 (mid-active phase), and ZT22 (transition to inactive phase). We hypothesised that the greatest changes would be observed in the stress-susceptible mice in the Hb during the light phase. We expected relatively few differences between the resilient and stress-naïve mice, especially in the SCN. However, we found the largest transcriptomic changes to occur at ZT16 in resilient relative to stress-naïve mice, in both SCN and Hb. Our observations highlight the importance of establishing transcriptomic profiles across several time points to more accurately describe the homoeostatic changes in response to stress. We speculate that homoeostatic plasticity in the SCN may be important for modulating resilience to stress.
Material and Methods
Ethics approval
All experimental procedures were conducted in accordance with the National Institute of Health Guide for Care and Use of Laboratory Animals and approved by the New York University Abu Dhabi Animal Care and Use Committee (IACUC Protocol: 19-0004).
Animals and housing
Male C57BL/6J mice (8–10 weeks old; Jackson Laboratory) were housed in groups of 5 upon arrival for at least a week before the start of the chronic social defeat stress (CSDS) paradigm. CD1 male mice (retired breeders; Charles River Laboratories) were housed individually under standard housing conditions. All mice were housed with ad libitum access to food and water at 20 ± 1 °C and 50 ± 10% humidity under a 12-h light/dark cycle (7:00 am / ZT0: lights on; 7:00 pm /ZT12: lights off).
Chronic social defeat stress
CSDS was performed as described previously [38]. Experimental C57 mice were exposed to 10 min of physical stress followed by 24 h of sensory stress in the cage of an unfamiliar CD1 aggressor mouse (Fig. S1A). CSDS was repeated daily for 15 days between ZT8 and ZT9 with a novel CD1. The timing of the daily defeat sessions was selected to complement a long record of published studies that conducted CSDS during the light phase [36, 46,47,48], but failed to report or follow consistent stress-exposure timing across days. The ZT8-ZT9 was specifically selected since mice exhibit low activity levels [15], as well as lower levels of sleep pressure [49, 50]. Control mice were housed in pairs in a separate room and moved each day at the time of defeat session. All the mice were single-housed following the end of CSDS until sacrifice. Social avoidance towards an unfamiliar non-aggressive CD1 mouse was measured in a two-stage social interaction (SI) test, as previously reported [38]. The test was conducted between ZT7 and ZT10. TopScan video tracking system (CleverSys. Inc.) was used to automatically track mouse movement.
Tissue sampling and region isolation
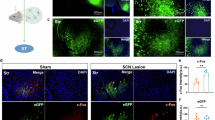
Mice were anaesthetised using isoflurane and sacrificed 48–72 h after the SI test by cervical dislocation. Sampling was done at four time points (ZT4, ZT10, ZT16, ZT22) across the light-dark cycle (Fig. 1A, B). Extractions were carried out on a cold sterile surface, and the extracted brains were flash-frozen by immersion into −60 °C isopentane (Sigma Aldrich M3263) for 20 s. The brains were kept on dry ice at -80°C until further processing. The flash-frozen brain samples were then sliced on a Leica CM1950 cryostat (250 µm thickness) (Fig. S1B). The regions were isolated with reference to the Mouse Brain Atlas [51]. For the SCN and Hb, 2–3 bilateral punches per region per biological replicate were combined in a single sample (Fig. S1B). There was no pooling across biological replicates.
A Timeline for investigating the effects of stress on temporal changes in gene expression in stress-naïve and stress-exposed animals. B Sampling time points and emerging comparison levels from pooling the sampling time points.
RNA isolation, library preparation and RNA-sequencing
RNA was isolated from the punches using the Qiagen RNeasy Micro Kit, as per manufacturer’s instructions. Isolated RNA was quantified and checked for quality on an Agilent Bioanalyzer instrument (Agilent Technologies) (Fig. S1C). Samples with RIN values above 7 were used for the library preparation. All the libraries were prepared together using the NEBNext® Ultra™ II RNA Library Prep Kit (New England BioLabs Inc.) as per the manufacturer’s protocol within one sitting. Sequencing was done on NovaSeq S1 flow cell (200 cycles) following Illumina’s recommendations.
Differential expression and pathway enrichment analysis
Count preprocessing
Raw FASTQ sequenced reads were first assessed for quality using FastQC (v0.11.5) [52]. Sequencing depth did not meaningfully differ between the phenotypes and timepoints, averaging 15.79 and 15.56 of TPM-normalised counts for Hb and SCN respectively (Supplementary Table 1). The reads were then passed through Trimmomatic (v0.36) for quality trimming [53] and processed with Fastp to remove poly-G tails and Novaseq-specific artefacts [54]. The reads were aligned to the mouse reference genome GRCm38.82 using HISAT2 with the default parameters [55]. The resulting SAM alignments were then converted to BAM format and coordinate-sorted using SAMtools (v1.3.1) [56]. Raw counts were generated by passing the sorted alignment files through HTSeq-count (v0.6.1p1) [57]. Concurrently, the sorted alignments were processed in Stringtie (v1.3.0) for transcriptome quantification [58]. Finally, Qualimap (v2.2.2) was used to generate RNAseq-specific QC metrics for each sample [59]. All of the methods were executed using predefined YAML workflows in the BioSAILs workflow management system [60].
Differential expression
Differential expression testing was performed in R package DESeq2 (v1.40.2) [61]. The p-values were adjusted using the Benjamini–Hochberg procedure. To obtain a better estimate of expression changes in low count genes, we performed log2-fold-change (LFC) shrinkage using the apeglm method [62]. We used an adjusted p-value of 0.05 (5% FDR) and a log2-fold-change (LFC) of 0.5 to identify differentially expressed genes (DEGs). Samples within the same behavioural phenotype from ZT4 and ZT10 time points were computationally pooled together to provide an estimate of the ‘daytime’ expression, while ZT16 and ZT22 time point samples were pooled to provide an estimate of the ‘nighttime’ expression for each phenotype (Fig. 1B). Pooling all the samples within the same behavioural phenotype provided an estimate of time-invariant ‘global’ expression corresponding to average expression throughout the day. Differential expression between the behavioural phenotypes was carried out at each of these temporal resolution levels.
SCN and Hb samples were assessed for presence of respective canonical markers from published scRNA-seq studies [63,64,65]. The sequenced SCN samples specifically expressed known SCN markers, while Hb strongly expressed both the LHb and MHb markers, suggesting sampling accuracy (Fig. S2). However, we did not manage to reliably sample the PHb (Fig. S2A). In order to assess whether technical biases were driving the observed differential expression patterns, the raw counts were normalised and stabilised for variance using DESeq2 with a design formula including region, time point and phenotype as covariates. The output was then used to compute sample similarity across all the genes (Fig. S3A), as well as the top 2000 most variable genes (Fig. S3B), which were then clustered based on their Eucledian distance. Beyond the sample region clustering, these heatmaps do not suggest any other specific grouping that would indicate a bias. Furthermore, we performed principle component analysis (PCA) on all the samples (data not shown). These results were replicated using surrogate variable analysis (SVA) (data not shown) [66]. First principal component represented the regional identity of the samples and accounted for 82% of data variability (Fig. S2B).
Pathway enrichment
Enrichment analysis was conducted in the Ingenuity Pathway Analysis (IPA, v.23.0, Qiagen) on all genes that had passed the 5% FDR significance criterion and was conducted only in those comparisons where the number of DEGs was higher than 20. Pathways were considered significantly enriched if –log(B-H p-value) was greater than 1.3. Based on the known roles of different genes and their relationship in the functionally-predefined pathways, IPA calculates z score which represents the predicted direction of change of the pathway (activation: z > 0; inhibition: z < 0). The analysis included only the pathways associated with the mouse nervous system, neurons, astrocytes, other tissues and unspecified cell types within the IPA framework.
Gene ontology
We visualised the associated DEGs and the top five IPA pathways from the resilient-to-control comparisons at ZT16 for both regions using gene-concept networks. Gene-concept networks represent the relative coordinates of the enriched pathways based on the number and differential expression of the associated DEGs. Therefore, it is possible to identify overlapping DEGs that have the largest contribution to the enrichment of the top five activated pathways.
In silico TF motif enrichment
To explore which transcription factors (TF) might be driving the differential expression at specific time points, we used FIMO [67] (fimo --oc {gene.fa} --verbosity 1 --bgfile --nrdb-- --thresh 1.0E-4 H12CORE_meme_format.meme {gene}) with the Mouse TF motif list from HOCOMOCO [68] to generate TF binding motif enrichment within 20 kb upstream of DEG promoters. The FASTA sequences were retrieved using BED2FASTA [67]. FIMO results were then filtered for hits corresponding to q ≤ 0.05 and for TF that were expressed above the median of normalised and transformed expression value. TF hits were then sum-aggregated by TF families. Given the unbalanced number of DEGs by conditions, aggregated TF hits were normalised by the number of hit genes per condition. Finally, the delta (Δ) between upregulated and downregulated hits was computed for each TF family. Positive delta values correspond to enriched binding of the TF family in the promoter of upregulated DEGs, and conversely negative values correspond to binding in the promoter of downregulated DEGs.
Detection of rhythmic genes
Log-normalised count values were used to determine the rhythmic expression profile of core clock genes across all three behavioural phenotypes using the non-parametric JTK method for identification of expression oscillations in genome-scale datasets [69]. JTK method was run in the R-implemented version of the ECHO (v.4.0) tool [70].
Results
CSDS induces social avoidance in a subset of stress-exposed mice
To investigate the effects of chronic stress on SCN and Hb transcriptome, we exposed mice to 15 days of CSDS. Following CSDS, mice were classified as stress-resilient or stress-susceptible using SI test (Fig. 1A, S4A–C). Stress-naïve and stress-resilient mice exhibited significantly larger social interaction values than the stress-susceptible mice (Fig. S4A–C). Resilient and susceptible mice exhibited decreased movement and velocity relative to the stress-naïve mice (Fig. S4D, E).
Stress-resiliency is associated with large gene upregulation in the SCN and Hb at ZT16
In order to investigate differential expression between the behavioural phenotypes at different time points, we sampled SCN and Hb at ZT4, ZT10, ZT16 and ZT22 time points 48–72 h following the SI test (Fig. 1A, B). Computationally merging ZT4 and ZT10 provided a daytime expression estimate, while merging ZT16 and ZT22 provided nighttime expression estimate (Fig. 1B). Combining all the time points together provided computational estimate of time-independent expression patterns (Fig. 1B). Comparing the phenotypes revealed that, in both the SCN and Hb, the highest number of significant genes was observed between resilient and stress-naïve mice at all temporal resolution comparison levels (Supplementary Tables 2, 3).
In the SCN (Fig. 2A, B), at global (average gene expression across all 4 time points) and daytime (average gene expression at ZT4 and ZT10) comparisons only a few genes were differentially expressed between any of the groups. Comparison of individual gene expression patterns at ZT4 and ZT10 time points did not reveal substantial differences between the phenotypes. However, at nighttime (average gene expression at ZT16 and ZT22) comparisons, we found 338 DEGs in resilient relative to stress-naïve mice, of which a large majority (~80%) were upregulated (Fig. 2A, C). The nighttime differences in the SCN predominantly arose from the differential expression observed at ZT16 where we found 758 DEGs in resilient relative to stress-naïve mice, of which ~60% were upregulated (Fig. 2B, E). At ZT22, we observed 34 upregulated DEGs in resilient relative to susceptible mice. However, other phenotypic comparisons at ZT22 did not reveal any major differential expression patterns. These results suggest that the largest changes in stress-induced SCN gene expression occur in resilient mice during nighttime at ZT16.
A Number of DEGs at pooled time points. At the global comparison level, there were 16, 5 and 7 DEGs in resilient-to-control, susceptible-to-control and resilient-to-susceptible comparisons, respectively. Daytime comparison revealed 2, 3 and 1 DEGs for resilient-to-control, susceptible-to-control and susceptible-to-resilient comparisons, respectively. Nighttime comparison revealed 338, 11 and 10 DEGs for resilient-to-control, susceptible-to-control and susceptible-to-resilient comparisons, respectively. B Number of DEGs at specific time points. At ZT4, there were 0, 3 and 7 DEGs for resilient-to-control, susceptible-to-control and resilient-to-susceptible comparisons. At ZT10, there were no DEGs for any phenotypic comparison. At ZT16, there were 758, 17 and 23 DEGs for resilient-to-control, susceptible-to-control and resilient-to-susceptible comparisons, respectively. At ZT22, there were 0, 0 and 34 DEGs for resilient-to-control, susceptible-to-control and resilient-to-susceptible, respectively. C Nighttime comparison: Volcano plot showing significant DEGs (FDR < 0.05) in resilient mice relative to controls at nighttime comparison level in SCN. D ZT16 comparison: Volcano plot showing significant DEGs (FDR < 0.05) in resilient mice relative to controls at ZT16 in SCN. Red and blue points represent significantly upregulated and downregulated genes, respectively.
In the Hb (Fig. 3A, B), differential expression patterns between the phenotypes were somewhat similar to those observed in the SCN. Global comparisons in the Hb revealed 68 upregulated DEGs in the resilient relative to stress-naïve mice (Fig. 3A, C). Gene expression patterns did not differ between any of the three behavioural phenotypes during the daytime. Specifically, ZT4 and ZT10 time point comparisons did not yield any discernible differences in gene expression patterns (Fig. 3B). However, nighttime comparison revealed 55 upregulated DEGs in resilient relative to stress-naïve mice (Fig. 3A, D). We found that the large differences in global and nighttime gene expression profiles are again mainly due to the large numbers of upregulated genes at ZT16 in stress-resilient relative to control and susceptible mice (Fig. 3A, B). Specifically, 143 DEGs were identified in the resilient relative to stress-naïve mice at ZT16, with almost all genes being upregulated (Fig. 3B, E). We observed 78 upregulated DEGs in resilient mice compared to stress-susceptible mice in the Hb, unlike in the SCN (Fig. 3B). Only small differences were observed between the susceptible and control mice at ZT16 in the Hb. No robust differences were observed at ZT22 between any of the behavioural phenotypes in the Hb (Fig. 3B). Hence, stress-induced differences in Hb gene expression patterns in the resilient mice, both at global and nighttime analysis, are driven by changes at the ZT16 time point, similar to the SCN.
A Number of DEGs at pooled time points. At the global comparison level, there were 69, 0 and 1 DEGs for resilient-to-control, susceptible-to-control and resilient-to-susceptible comparisons, respectively. At the daytime comparison level, there were 0, 0 and 2 DEGs for resilient-to-control, susceptible-to-control and susceptible-to-resilient comparisons, respectively. At the nighttime comparison level, there were 55, 0 and 0 DEGs for resilient-to-control, susceptible-to-control and susceptible-to-resilient comparisons, respectively. B Number of DEGs at specific sampled time points. At ZT4, there were 1, 0 and 0 DEGs for resilient-to-control, susceptible-to-control and resilient-to-susceptible comparisons. At ZT10, there were no DEGs for resilient-to-control and resilient-to-susceptible comparisons, and only 1 DEG in susceptible-to-control comparison. At ZT16, there were 143, 2 and 80 DEGs for resilient-to-control, susceptible-to-control and resilient-to-susceptible comparisons, respectively. At ZT22, there were no DEGs in any of the phenotypic comparisons. C Global comparison: Volcano plot showing significant DEGs (FDR < 0.05) in resilient mice relative to controls at global comparison level in Hb. DEGs were labelled based on |log2FC| > 1.0 and biological relevance. D Nighttime comparison: Volcano plot showing significant DEGs (FDR < 0.05) in resilient mice relative to controls at nighttime comparison level in Hb. E ZT16 comparison: Volcano plot showing significant DEGs (FDR < 0.05) in resilient mice relative to controls at ZT16 time point in Hb. Red and blue points represent significantly upregulated and downregulated genes, respectively.
Given the strong differential expression at ZT16 between resilient and stress-naïve mice, we examined in sillico possible TF candidates among DEGs that could drive these differences (Fig. S5). We found a greater number of possible TFs associated with downregulated than upregulated DEGs in both SCN and Hb. The two brain regions showed enrichment of 35 common TF families. Differential expression in the SCN and Hb was associated with 8 and 39 region-specific TF families, respectively.
Overall, these results show robust transcriptomic differences in resilient relative to stress-naïve mice in both the SCN and Hb. The largest observed differences occur at ZT16 in both regions, with SCN showing larger transcriptional response than the Hb in terms of DEG numbers and associated LFC. Overall, susceptible mice did not show significant differential expression patterns in most comparisons relative to either the resilient or control mice. Taken together, the results suggest resiliency-associated transcriptional signature to be temporally-restricted to the middle of the active phase.
Stress resilience is associated with enrichment of signalling and plasticity pathways in SCN at ZT16
To better understand the molecular pathways and functions of DEGs between the resilient and stress-naïve mice, we performed pathway enrichment analysis in the SCN at nighttime and at ZT16 and found a number of enriched pathways (Supplementary Table 4, Fig. 4). At ZT16, the enriched pathways included an expanded list of those observed from the nighttime comparison (Supplementary Table 4, Fig. 4A, B). All the enriched pathways in the resilient mice can be divided into two broad functional groups: signalling and plasticity-related pathways (Fig. 4A). Furthermore, at ZT16, ‘circadian rhythm signalling’ pathway was significantly enriched in the resilient mice. Both signalling and plasticity-related pathways in the resilient mice were enriched by common DEGs (subset shown in Supplementary Table 5). Many upregulated DEGs were associated with most enriched pathways (Fig. 4C). These results suggest that stress resilience is strongly associated with upregulation of relatively few genes in SCN at ZT16 which subsequently have broad impact on signalling and plasticity regulation, possibly through their role as TFs.
The activation pattern of enriched biological pathways in the SCN. All lists are sorted by adjusted p-value. Red denotes activation (z > 0), blue denotes inhibition (z < 0), white denotes no net effect (z = 0), and grey denotes uncertain activation pattern. A Enriched signalling pathways. B Enriched plasticity-associated pathways. C Gene-concept networks of the 5 most enriched IPA pathways at ZT16 between resilient and control phenotypes in the SCN.
Stress resilience is associated with enrichment of synaptogenesis and calcium signalling pathways in the Hb at ZT16
In the Hb, we explored pathway enrichment at global, nighttime and ZT16 comparison levels between the resilient and stress-naïve mice (Fig. 5). Fewer pathways were as highly enriched and activated in the Hb of the resilient mice at ZT16 compared to the SCN (Supplementary Table 6, Fig. 5A, B). The most enriched pathways in this comparison were associated with mechanisms of synaptic plasticity (Supplementary Table 6, Fig. 5B). Similar to the SCN, ‘circadian rhythm signalling’ pathway was enriched in the resilient mice at global, nighttime and ZT16 comparisons. Relatively few DEGs were associated with a high number of enriched pathways in the Hb of resilient mice (Supplementary Table 5). Furthermore, the most enriched pathways were associated with a handful of genes (Fig. 5C). These results suggest that stress resilience is associated with upregulation of plasticity-associated genes in the Hb at ZT16. Although both the SCN and Hb exhibited robust changes specifically at ZT16, the SCN shows larger and functionally more diverse transcriptomic response to stress than the Hb.
The activation pattern of enriched biological pathways in the Hb. All lists are sorted by adjusted p-value. Red denotes activation (z > 0), blue denotes inhibition (z < 0), white denotes no net effect (z = 0), and grey denotes uncertain activation pattern. A Enriched signalling pathways. B Enriched plasticity-associated pathways. C Gene-concept networks of 5 most enriched IPA pathways at ZT16 between resilient and control phenotypes in the Hb.
Stress induces arrhythmicity of Per1 and Arntl in the SCN of stress exposed mice
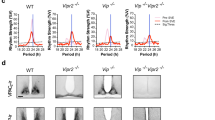
Given the enrichment of ‘circadian rhythms signalling’ pathway, we explored whether chronic stress affects the rhythmicity of core clock genes in the SCN and Hb. To this end, we performed JTK analysis and found rhythmic expression of the core clock genes (Per1, Per2, Per3, Cry1, and Arntl) in the SCN (padj <0.05) of stress-naïve mice (Fig. 6A, B, Supplementary Table 7). Clock was arrhythmic in both brain regions across all the phenotypes. In the SCN, Per1 was arrhythmic specifically in the stress-susceptible mice. Arntl was arrhythmic in both stress-resilient and stress-susceptible mice. In contrast Cry2 was rhythmic only in the SCN of resilient mice. In the Hb, the expression of core clock genes was more uniform across the phenotypes. With the exception of Cry1 and Clock, all genes were rhythmic in stress-resilient and stress-susceptible mice. Moreover, Per1, Per3 and Cry1 were arrhythmic in stress-naïve mice, but rhythmic in resilient and susceptible mice (Fig. 6A, B). Beyond core clock genes, large differences in numbers of phenotype-specific rhythmic genes were observed in both the SCN and Hb (Fig. S6).
A Temporal expression of core clock genes in the SCN and Hb of control, resilient and susceptible mice. All the clock genes show previously reported temporal expression in the SCN of control mice. B Dotplot showing rhythmic expression of clock genes in SCN and Hb. SCN: Per1 is arhythmic stress-susceptible mice. Arntl is arrhythmic in both resilient and susceptible mice. Cry2 is rhythmic in resilient mice. Hb: Per1, Per3 and Cry1 are arrhythmic in the stress-naïve mice, while being rhythmic in both the resilient and susceptible mice.
Discussion
Stress resilience and homoeostatic response
The close relationship between diurnal rhythms and mood disorders warranted a systemic investigation of transcriptomic changes in the principal clock (SCN) and mood-regulating brain regions. We focused on the Hb because it is a semi-autonomous oscillator with a strong role in stress regulation and mood disorders [14]. We found that chronic stress elicited a robust change in the gene expression in the SCN and to a lesser degree in the Hb (Figs. 2–3). This transcriptomic signature was specific to resilient mice and was strongest at ZT16. In contrast, comparing susceptible and stress-naïve mice did not reveal strong differential expression patterns. Large differential upregulation in resilient mice was associated with enrichment and predicted activation of signalling and plasticity-related pathways in both regions (Figs. 4–5). By computationally pooling the samples and decreasing the temporal resolution, we demonstrated that such strong transcriptomic signatures are lost.
Resilience is an active homoeostatic process and involves transcriptional changes in a diffuse network of interconnected brain regions [71]. The current understanding is that chronic stress imposes an adaptive challenge that requires behavioural flexibility and physiological homoeostatic buffering [72]. The inability to elicit such a response leads to failure in stress-coping and strongly contributes to associated stress-induced pathologies [72, 73]. Resilient mice do not show behavioural deficits likely because of the large transcriptomic changes such as those observed in our study. Pioneering studies found that resilient mice exhibit large and unique upregulation of neurotrophic factors and stress-responsive genes in the prefrontal cortex [36]. Others found strong gene upregulation in resilient mice relative to stress-naïve and susceptible mice in midbrain regions such as the VTA and NAc [74, 75]. The physiological consequences of large transcriptomic changes in resilient mice are also likely responsible for observed differences in circuit-specific neural activity between resilient and susceptible mice. For example, VTA dopaminergic (DA) cells in susceptible, but not resilient, mice exhibit elevated firing rate [76]. The pathophysiological elevated firing is counteracted in resilient mice by compensatory mechanisms that are absent in susceptible mice [77]. We speculate that the relatively negligable differences between resilient and susceptible mice were observed because chronic stress induces similar transcriptional changes in both phenotypes. However, larger differential expression in resilient mice would be sufficient in driving necessary homoeostatic response for acquisition of the resilient phenotype. Susceptible mice do not adapt to the same degree as resilient mice and are transcriptomically found in an ‘intermediate’ state [36, 77], explaining negligible differences in relation to both resilient and stress-naïve mice.
Our findings that large transcriptomic changes occur in resilient mice at a specific night time point (ZT16) add a significant temporal dimension to the current understanding of the molecular mechanisms that drive stress-resilience. Our data, along with previously published physiological and circuit studies, suggest that temporal patterns of regional activity may play a crucial role in driving a particular behavioural phenotype [15]. It provides further evidence of a tight association between large transcriptional changes and successful stress-coping. Future studies with sampling performed over several day/night cycles following chronic stress would allow for temporal stability estimation of this transcriptomic signature.
SCN transcriptomic profile
We observed significant upregulation of Adora2a in the SCN of resilient mice at ZT16. Adenosine is known to regulate Per1 and Per2 expression via A1/A2A receptor activation which is hypothesised to integrate light and sleep signalling to track homoeostatic sleep pressure and regulate sleep architecture - both of which were previously found to be differentially impacted in resilient and susceptible mice [39, 41, 78]. Upregulation of Adora2a in the SCN of resilient mice at ZT16 may be a subcomponent of adaptive homoeostatic response that maintains consolidated sleep [39, 41]. Interestingly, the upregulation of A2A receptors in the forebrain is associated with depressive-like behaviour and decreased synaptic plasticity following unpredictable CMS [79].
Resilient mice also displayed strong upregulation of Ntrk1 and Ngfr in the SCN at ZT16. NGF binds to Ntrk1 and Ngfr [80, 81]. NGF levels are significantly lower in depressed patients [82] while increasing its levels has antidepressant properties [83, 84]. Furthermore, BDNF, which has functional roles in synaptic plasticity, stress response, and antidepressant action in several regions, also binds to Ngfr [85, 86]. Ngfr is specifically synthesised in cholinergic neurons which strongly modulate SCN neuronal excitability and circadian rhythmicity at night [87,88,89].
We also observed high upregulation of cholinergic muscarinic receptors Chrm 1-5. These receptors have a functional role in fear and emotional responsiveness [90, 91]. Cholinergic signalling in the SCN can induce phase shifts in locomotor rhythms [92]. However, at present it is unclear how increased muscarinic receptor expression at ZT16 in the SCN drives resilience. A recent study demonstrated that the stress-associated neuropeptide CRH increases cholinergic interneuron firing that causes potentiation of dopamine transmission [93, 94]. Besides cholinergic receptors, Drd2 and Crh were also upregulated in the SCN of resilient mice at ZT16. Dopaminergic input from VTA to SCN modulates circadian entrainment through activation of excitatory Drd1 receptors [95]. The putative homoeostatic role of inhibitory Drd2 in circadian regulation in resilient mice remains unknown. Our observation that Crh is upregulated in resilient mice adds to the recent findings that CRH-expressing neurons are also present in the SCN [96]. CRH increases gating of BDNF signalling [97]. Thus, increased CRH expression at ZT16 may be responsible for putative BDNF-induced antidepressant-like effects in stress-resilient mice.
The resilient phenotype has also been associated with increased cAMP and GPCR-mediated signalling [74, 98]. These plasticity-related pathways were also enriched in our dataset and predicted to be activated in the SCN of resilient mice. However, the functional consequences of these changes are unclear since plasticity-related transcription factor Creb3l1 was downregulated, while many others, such as Gbx1, were strongly upregulated in the resilient mice. Previously, however, Creb downregulation has been associated with stress resilience [99, 100]. Overall, few studies have examined stress effects on SCN physiology and those predominantly focused on the expression of clock genes [7, 101,102,103]. Our observations highlight potentially novel molecular mechanisms in the SCN that promote resilience to stress, warranting further investigation.
Hb transcriptomic profile
Adcy1 and Adcy9 were upregulated in the Hb of resilient relative to stress-naïve mice at ZT16. Adenylyl cyclases play a pivotal role in both cAMP-mediated and the ERK½-CREB signalling cascade which are known to modulate neuronal physiology [104,105,106]. Overexpression of Adcy1 in the forebrain prevents BDNF downregulation in stressed mice, thereby promoting stress resiliency [107]. Adcy1 forebrain overexpression also induces stronger long-term potentiation (LTP) [106]. We also observed ZT16-specific upregulation of other plasticity-modulating genes in resilient mice, such as Grin2a, Grin2b, Cacna1h, Camk2b, and Ppp3ca which correlate with large predicted activation of ‘synaptogenesis’ and ‘LTP signalling’ pathways. Resilient mice were found to exhibit larger hippocampal LTP relative to susceptible mice because of higher expression of Grin2b and Camk2 [108]. Upregulation of plasticity-related genes may be a part of a larger homoeostatic response associated with stress resiliency [109, 110].
We also observed upregulation of Gad1, Gabra5, Kcnq3 and Npy1r genes in stress-resilient mice. Sub-chronic mild stress decreases long-term depression (LTD) of GABAergic neurons in the LHb [111]. Furthermore, male mice with Kcnq3 mutation that inhibits GABA binding exhibit reduced self-grooming, behaviour indicative of apathy [37, 112]. Neuropeptide Y action through its Y1 receptor has been associated with stress resilience and anxiolysis in several brain regions, including the Hb [113, 114]. In the LHb, Npy1r-expressing neurons play a critical role in chronic stress adaptation by driving palatable food intake and reducing anxious-like behaviour [115]. Thus, it is possible that elevated Npy1r expression at ZT16 in the Hb drives increased food intake in resilient mice to compensate for lower levels of adipose tissue and serum leptin [116]. Our data suggests that Hb might play an important role in promoting stress-resilience, beyond its known role in promoting depressive-like phenotype.
Clock genes
Since our analysis revealed a strong effect of stress on transcriptional profiles in the SCN and Hb at a specific time point, we examined the rhythmic expression of clock genes in both of these regions. We found that the resilient and susceptible mice exhibit similar rhythmic expression profile of the core clock genes in the SCN with the exception of Per1 and Cry2 which were differentially modulated by CSDS. Chronic stress induced arrhythmicity of Arntl in the SCN of both resilient and susceptible mice which might imply that putative homoeostatic adaptations in resilient mice do not protect against stress-induced disruption in rhythmic expression of this specific clock gene. Despite previous observation that knockdown of Arntl leads to despairful-, anxious- and helpless-like behaviour following CMS, our results suggest that Arntl arrhythmicity in the SCN does not directly impact mood regulation [7]. Interestingly, Cry2 was rhythmically expressed in the stress-resilient mice, unlike stress-naïve and stress-susceptible mice, potentially pointing to a compensatory mechanism involving induced rhythmic expression of Cry2. Using in vitro bioluminescence methods, one study found that Per2 expression amplitude was increased in the SCN of mice that underwent nighttime CSDS, but not daytime CSDS [43]. Although inconclusive due to low temporal resolution, our data also suggests amplitude blunting. Others have shown that exposure to CSDS during night induces differences in locomotor activity and tissue-specific Per2 expression profile [117, 118], but also that resting-phase CSDS more strongly supresses locomotor activity [101]. In contrast no difference in Per2 expression was observed in mice kept in constant dim red light where clock rhythms are free running [119]. These observations imply that phase-specific CSDS exposure modulates rhythmic behavioural and gene expression profiles. Future studies will investigate whether the transcriptomic profile of mice exposed to chronic stress at night induces similar transcriptomic profile to the one observed in resilient mice in our study. Finally, rhythmic expression profile of SCN of stress-naïve mice in our study match previously published, more spatially-resolved findings [64].
In our study, Per1, Per3, and Cry1 were arrhythmic in the Hb of stress-naïve mice, but not resilient and susceptible mice. Although unexpected, these results can be attributed to the subdivisional differences at the level of MHb and LHb [10, 24,25,26]. Clock gene expression in the MHb has yet to be systematically investigated in mice, though it has been reported previously in hamsters and rats [120, 121]. Despite both the LHb and MHb showing diurnal variation in firing [24, 25], the LHb displays more robust rhythms with higher amplitude than the MHb [10]. In the present study we did not differentiate between the two subregions and the reduced spatial resolution might have contributed to the observed arrhythmicity of clock genes in stress-naïve mice. Moreover, different stressors have been previously shown to differentially impact neuronal dynamics [37]. Thus, the differential effect of CSDS and CMS on the transcriptomic landscape may be explained by the utilised stressor type. Future studies using higher sampling resolution would be required to fully describe effects on the phase of core clock gene expression.
Limitations
There are several limitations to our study. An increasing number of studies report stronger perturbations following nighttime, rather than daytime, defeat [42, 43, 117, 118, 122]. Our daytime defeat may miss potential molecular differences between resilient and susceptible mice that might be evident in mice exposed to nighttime stress. Furthermore, bulk RNA-seq approach offers limited spatial resolution where neither the subregions nor neuronal subpopulations can be clearly differentiated. It has been previously shown that cell types and subpopulations can widely differ in rhythmic profiles of core clock genes [64]. Even though PHb does not respond to short-duration CSDS [31], it was not reliably sampled in our study. Lastly, sampling at additional time points to increase the temporal resolution would have allowed for more robust temporal gene expression profiling. Future studies will elucidate subregion-specific effects using techniques of greater spatial resolution, such as scRNA-seq, and with larger number of time points for increased temporal resolution.
Concluding remarks
The SCN and Hb are heterogeneous regions with many functionally and molecularly distinct subpopulations. Previous transcriptomic studies have only examined specific subpopulations of the Hb for purposes of circuit manipulations [18, 35, 123]. Likewise, studies linking SCN and stress were limited to investigating a limited predetermined list of genes [7, 43]. To the best of our knowledge, no study has systematically examined the SCN or Hb bulk transcriptome in the context of murine chronic stress models across multiple time points throughout the diurnal cycle. Our study is the first to show that the largest transcriptomic changes in the SCN and Hb of resilient mice occur at night with a peak at specific time point. This highlights the importance of sampling over a 24-hour cycle to accurately determine dynamic changes in gene expression following stress exposure, since failing to do so conceals strong transcriptomic signatures as demonstrated with our computational pooling of day and night time points. Most studies fail to consider diurnal changes in expression profiles. Our time-dependent transcriptomic analysis suggests novel potential molecular mechanisms for driving resilience to stress that would not be evident with daytime-limited sampling. Such findings further support time-dependent medication treatment administration, which might enhance its efficacy. Crucially, our results highlight the link between temporal expression patterns and homoeostatic stress responses that remains to be investigated further.
Data availability
Behavioural and sequencing data is avaiable through the G-Node Infrastructure (GIN) repository (https://doi.org/10.12751/g-node.jj57l9). Any additional information will be made available by the corresponding author upon reasonable request.
References
Maletic V, Robinson M, Oakes T, Iyengar S, Ball SG, Russell J. Neurobiology of depression: an integrated view of key findings. Int J Clin Pract. 2007;61:2030–40.
Li JZ, Bunney BG, Meng F, Hagenauer MH, Walsh DM, Vawter MP, et al. Circadian patterns of gene expression in the human brain and disruption in major depressive disorder. Proc Natl Acad Sci USA 2013;110:9950–5.
Joyce PR, Porter RJ, Mulder RT, Luty SE, McKenzie JM, Miller AL, et al. Reversed diurnal variation in depression: associations with a differential antidepressant response, tryptophan:large neutral amino acid ratio and serotonin transporter polymorphisms. Psychol Med. 2005;35:511–7.
Germain A, Kupfer DJ. Circadian rhythm disturbances in depression. Hum Psychopharmacol Clin Exp. 2008;23:571–85.
Walker WH, Walton JC, DeVries AC, Nelson RJ. Circadian rhythm disruption and mental health. Transl Psychiatry. 2020;10:28.
Takahashi JS. Transcriptional architecture of the mammalian circadian clock. Nat Rev Genet. 2017;18:164–79.
Landgraf D, Long JE, Proulx CD, Barandas R, Malinow R, Welsh DK. Genetic disruption of circadian rhythms in the suprachiasmatic nucleus causes helplessness, behavioral despair, and anxiety-like behavior in mice. Biol Psychiatry. 2016;80:827–35.
Vadnie CA, Petersen KA, Eberhardt LA, Hildebrand MA, Cerwensky AJ, Zhang H, et al. The suprachiasmatic nucleus regulates anxiety-like behavior in mice. Front Neurosci. 2022;15:765850.
Vujovic N, Gooley JJ, Jhou TC, Saper CB. Projections from the subparaventricular zone define four channels of output from the circadian timing system: Projections from the SPZ. J Comp Neurol. 2015;523:2714–37.
Baño-Otálora B, Piggins HD. Contributions of the lateral habenula to circadian timekeeping. Pharmacol Biochem Behav. 2017;162:46–54.
Yang Y, Wang H, Hu J, Hu H. Lateral habenula in the pathophysiology of depression. Curr Opin Neurobiol. 2018;48:90–6.
Baker PM, Mathis V, Lecourtier L, Simmons SC, Nugent FS, Hill S, et al. Lateral habenula beyond avoidance: roles in stress, memory, and decision-making with implications for psychiatric disorders. Front Syst Neurosci. 2022;16:826475.
Gold PW, Kadriu B. A major role for the lateral habenula in depressive illness: physiologic and molecular mechanisms. Front Psychiatry. 2019;10:320.
Young CJ, Lyons D, Piggins HD. Circadian influences on the habenula and their potential contribution to neuropsychiatric disorders. Front Behav Neurosci. 2022;15:815700.
Liu H, Rastogi A, Narain P, Xu Q, Sabanovic M, Alhammadi AD, et al. Blunted diurnal firing in lateral habenula projections to dorsal raphe nucleus and delayed photoentrainment in stress-susceptible mice. PLoS Biol. 2021;19:e3000709.
Coffey KR, Marx RE, Vo EK, Nair SG, Neumaier JF. Chemogenetic inhibition of lateral habenula projections to the dorsal raphe nucleus reduces passive coping and perseverative reward seeking in rats. Neuropsychopharmacology. 2020;45:1115–24.
Warden MR, Selimbeyoglu A, Mirzabekov JJ, Lo M, Thompson KR, Kim S-Y, et al. A prefrontal cortex–brainstem neuronal projection that controls response to behavioural challenge. Nature. 2012;492:428–32.
Cerniauskas I, Winterer J, De Jong JW, Lukacsovich D, Yang H, Khan F, et al. Chronic stress induces activity, synaptic, and transcriptional remodeling of the lateral habenula associated with deficits in motivated behaviors. Neuron. 2019;104:899–915.e8.
Yang L-M, Hu B, Xia Y-H, Zhang B-L, Zhao H. Lateral habenula lesions improve the behavioral response in depressed rats via increasing the serotonin level in dorsal raphe nucleus. Behav Brain Res. 2008;188:84–90.
Winter C, Vollmayr B, Djodari-Irani A, Klein J, Sartorius A. Pharmacological inhibition of the lateral habenula improves depressive-like behavior in an animal model of treatment resistant depression. Behav Brain Res. 2011;216:463–5.
Hsu Y-WA, Wang SD, Wang S, Morton G, Zariwala HA, De La Iglesia HO, et al. Role of the dorsal medial habenula in the regulation of voluntary activity, motor function, hedonic state, and primary reinforcement. J Neurosci. 2014;34:11366–84.
Hsu Y-WA, Morton G, Guy EG, Wang SD, Turner EE. Dorsal medial habenula regulation of mood-related behaviors and primary reinforcement by tachykinin-expressing habenula neurons. eneuro. 2016;3:ENEURO.0109–16.2016.
Hsu Y-WA, Gile JJ, Perez JG, Morton G, Ben-Hamo M, Turner EE, et al. The dorsal medial habenula minimally impacts circadian regulation of locomotor activity and sleep. J Biol Rhythms. 2017;32:444–55.
Sakhi K, Wegner S, Belle MDC, Howarth M, Delagrange P, Brown TM, et al. Intrinsic and extrinsic cues regulate the daily profile of mouse lateral habenula neuronal activity. J Physiol. 2014;592:5025–45.
Sakhi K, Belle MDC, Gossan N, Delagrange P, Piggins HD. Daily variation in the electrophysiological activity of mouse medial habenula neurones. J Physiol. 2014;592:587–603.
Zhang C, Truong KK, Zhou Q-Y. Efferent projections of prokineticin 2 expressing neurons in the mouse suprachiasmatic nucleus. PLoS ONE. 2009;4:e7151.
Morin LP. Neuroanatomy of the extended circadian rhythm system. Exp Neurol. 2013;243:4–20.
Poller WC, Madai VI, Bernard R, Laube G, Veh RW. A glutamatergic projection from the lateral hypothalamus targets VTA-projecting neurons in the lateral habenula of the rat. Brain Res. 2013;1507:45–60.
Stamatakis AM, Van Swieten M, Basiri ML, Blair GA, Kantak P, Stuber GD. Lateral hypothalamic area glutamatergic neurons and their projections to the lateral habenula regulate feeding and reward. J Neurosci. 2016;36:302–11.
Hu H, Cui Y, Yang Y. Circuits and functions of the lateral habenula in health and in disease. Nat Rev Neurosci. 2020;21:277–95.
Fernandez DC, Fogerson PM, Lazzerini Ospri L, Thomsen MB, Layne RM, Severin D, et al. Light affects mood and learning through distinct retina-brain pathways. Cell. 2018;175:71–84.e18.
Hattar S, Kumar M, Park A, Tong P, Tung J, Yau K-W, et al. Central projections of melanopsin-expressing retinal ganglion cells in the mouse. J Comp Neurol. 2006;497:326–49.
Morin LP, Studholme KM. Retinofugal projections in the mouse. J Compar Neurol. 2014;522:3733–53.
An K, Zhao H, Miao Y, Xu Q, Li Y-F, Ma Y-Q, et al. A circadian rhythm-gated subcortical pathway for nighttime-light-induced depressive-like behaviors in mice. Nat Neurosci. 2020;23:869–80.
Weil T, Daly KM, Yarur Castillo H, Thomsen MB, Wang H, Mercau ME, et al. Daily changes in light influence mood via inhibitory networks within the thalamic perihabenular nucleus. Sci Adv. 2022;8:eabn3567.
Krishnan V, Han M-H, Graham DL, Berton O, Renthal W, Russo SJ, et al. Molecular adaptations underlying susceptibility and resistance to social defeat in brain reward regions. Cell. 2007;131:391–404.
Petković A, Chaudhury D. Encore: Behavioural animal models of stress, depression and mood disorders. Front Behav Neurosci. 2022;16:931964.
Golden SA, Covington HE, Berton O, Russo SJ. A standardized protocol for repeated social defeat stress in mice. Nat Protoc. 2011;6:1183–91.
Radwan B, Jansen G, Chaudhury D. Abnormal sleep signals vulnerability to chronic social defeat stress. Front Neurosci. 2021;14:610655.
Haniffa S, Narain P, Hughes MA, Petković A, Šušić M, Mlambo V, et al. Chronic social stress blunts core body temperature and molecular rhythms of Rbm3 and Cirbp in mouse lateral habenula. Open Biol. 2023;13:220380.
Radwan B, Jansen G, Chaudhury D. Sleep-wake dynamics pre- and post-exposure to chronic social stress. iScience. 2021;24:103204.
Bartlang MS, Neumann ID, Slattery DA, Uschold-Schmidt N, Kraus D, Helfrich-Förster C, et al. Time matters: pathological effects of repeated psychosocial stress during the active, but not inactive, phase of male mice. J Endocrinol. 2012;215:425–37.
Bartlang MS, Savelyev SA, Johansson A-S, Reber SO, Helfrich-Förster C, Lundkvist GBS. Repeated psychosocial stress at night, but not day, affects the central molecular clock. Chronobiol Int. 2014;31:996–1007.
Christiansen S, Bouzinova E, Fahrenkrug J, Wiborg O. Altered expression pattern of clock genes in a rat model of depression. IJNPPY. 2016;19:pyw061.
Logan RW, Edgar N, Gillman AG, Hoffman D, Zhu X, McClung CA. Chronic stress induces brain region-specific alterations of molecular rhythms that correlate with depression-like behavior in mice. Biol Psychiatry. 2015;78:249–58.
Berton O, McClung CA, DiLeone RJ, Krishnan V, Renthal W, Russo SJ, et al. Essential role of BDNF in the mesolimbic dopamine pathway in social defeat stress. Science. 2006;311:864–8.
Lopez JP, Brivio E, Santambrogio A, De Donno C, Kos A, Peters M, et al. Single-cell molecular profiling of all three components of the HPA axis reveals adrenal ABCB1 as a regulator of stress adaptation. Sci Adv. 2021;7:eabe4497.
Morel C, Montgomery SE, Li L, Durand-de Cuttoli R, Teichman EM, Juarez B, et al. Midbrain projection to the basolateral amygdala encodes anxiety-like but not depression-like behaviors. Nat Commun. 2022;13:1532.
Radwan B, Yanez Touzet A, Hammami S, Chaudhury D. Prolonged exposure to social stress impairs homeostatic sleep regulation. Front Neurosci. 2021;15:633955.
Deboer T. Behavioral and electrophysiological correlates of sleep and sleep homeostasis. In: Meerlo P, Benca RM, Abel T, editors. Sleep, neuronal plasticity and brain function. Berlin: Springer; 2013. p. 1–24.
Paxinos G, Franklin KBJ. Paxinos and Franklin’s The mouse brain in stereotaxic coordinates. 5th ed. London: Elsevier; 2019.
Andrews S. FastQC: a quality control tool for high throughput sequence data [Online]. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Pertea M, Kim D, Pertea GM, Leek JT, Salzberg SL. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc. 2016;11:1650–67.
García-Alcalde F, Okonechnikov K, Carbonell J, Cruz LM, Götz S, Tarazona S, et al. Qualimap: evaluating next-generation sequencing alignment data. Bioinformatics. 2012;28:2678–9.
Rowe J, Drou N, Youssef A, Gunsalus KC. BioSAILs: versatile workflow management for high-throughput data analysis. Bioinformatics. 2019. https://doi.org/10.1101/509455.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Zhu A, Ibrahim JG, Love MI. Heavy-tailed prior distributions for sequence count data: removing the noise and preserving large differences. Bioinformatics. 2019;35:2084–92.
Wallace ML, Huang KW, Hochbaum D, Hyun M, Radeljic G, Sabatini BL. Anatomical and single-cell transcriptional profiling of the murine habenular complex. eLife. 2020;9:e51271.
Wen S, Ma D, Zhao M, Xie L, Wu Q, Gou L, et al. Spatiotemporal single-cell analysis of gene expression in the mouse suprachiasmatic nucleus. Nat Neurosci. 2020;23:456–67.
Hashikawa Y, Hashikawa K, Rossi MA, Basiri ML, Liu Y, Johnston NL, et al. Transcriptional and spatial resolution of cell types in the mammalian habenula. Neuron. 2020;106:743–58.e5.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28:882–3.
Bailey TL, Johnson J, Grant CE, Noble WS. The MEME Suite. Nucleic Acids Res. 2015;43:W39–49.
Vorontsov IE, Eliseeva IA, Zinkevich A, Nikonov M, Abramov S, Boytsov A, et al. HOCOMOCO in 2024: a rebuild of the curated collection of binding models for human and mouse transcription factors. Nucleic Acids Res. 2024;52:D154–63.
Hughes ME, Hogenesch JB, Kornacker K. JTK_CYCLE: An Efficient Nonparametric Algorithm for Detecting Rhythmic Components in Genome-Scale Data Sets. J Biol Rhythms. 2010;25:372–80.
De Los Santos H, Collins EJ, Mann C, Sagan AW, Jankowski MS, Bennett KP, et al. ECHO: an application for detection and analysis of oscillators identifies metabolic regulation on genome-wide circadian output. Bioinformatics. 2020;36:773–81.
Franklin TB, Saab BJ, Mansuy IM. Neural mechanisms of stress resilience and vulnerability. Neuron. 2012;75:747–61.
Pfau ML, Russo SJ. Peripheral and central mechanisms of stress resilience. Neurobiol Stress. 2015;1:66–79.
Wu G, Feder A, Cohen H, Kim JJ, Calderon S, Charney DS, et al. Understanding resilience. Front Behav Neurosci. 2013;7. https://doi.org/10.3389/fnbeh.2013.00010.
Bagot RC, Cates HM, Purushothaman I, Lorsch ZS, Walker DM, Wang J, et al. Circuit-wide transcriptional profiling reveals brain region-specific gene networks regulating depression susceptibility. Neuron. 2016;90:969–83.
Golden SA, Christoffel DJ, Heshmati M, Hodes GE, Magida J, Davis K, et al. Epigenetic regulation of RAC1 induces synaptic remodeling in stress disorders and depression. Nat Med. 2013;19:337–44.
Chaudhury D, Walsh JJ, Friedman AK, Juarez B, Ku SM, Koo JW, et al. Rapid regulation of depression-related behaviours by control of midbrain dopamine neurons. Nature. 2013;493:532–6.
Friedman AK, Walsh JJ, Juarez B, Ku SM, Chaudhury D, Wang J, et al. Enhancing depression mechanisms in midbrain dopamine neurons achieves homeostatic resilience. Science. 2014;344:313–9.
Jagannath A, Varga N, Dallmann R, Rando G, Gosselin P, Ebrahimjee F, et al. Adenosine integrates light and sleep signalling for the regulation of circadian timing in mice. Nat Commun. 2021;12:2113.
Van Calker D, Biber K, Domschke K, Serchov T. The role of adenosine receptors in mood and anxiety disorders. J Neurochem. 2019;151:11–27.
Bathina S, Das UN. Brain-derived neurotrophic factor and its clinical implications. Arch Med Sci. 2015;6:1164–78.
Miranda M, Morici JF, Zanoni MB, Bekinschtein P. Brain-derived neurotrophic factor: a key molecule for memory in the healthy and the pathological brain. Front Cell Neurosci. 2019;13:363.
Tseng P-T, Lin P-Y, Tu K-Y, Cheng Y-S, Wu C-K, Chen Y-W. Significantly lower nerve growth factor levels in patients with major depressive disorder than in healthy subjects: a meta-analysis and systematic review. Neuropsyciatr Dis Treat. 2015;11:925.
Mondal AC, Fatima M. Direct and indirect evidences of BDNF and NGF as key modulators in depression: role of antidepressants treatment. Int J Neurosci. 2019;129:283–96.
Overstreet DH, Fredericks K, Knapp D, Breese G, McMichael J. Nerve growth factor (NGF) has novel antidepressant-like properties in rats. Pharmacol Biochem Behav. 2010;94:553–60.
Chaudhury D, Liu H, Han M-H. Neuronal correlates of depression. Cell Mol Life Sci. 2015;72:4825–48.
Koo JW, Chaudhury D, Han M-H, Nestler EJ. Role of mesolimbic brain-derived neurotrophic factor in depression. Biol Psychiatry. 2019;86:738–48.
Bina KG, Rusak B. Muscarinic receptors mediate carbachol-induced phase shifts of circadian activity rhythms in Syrian hamsters. Brain Res. 1996;743:202–11.
Bina KG, Rusak B. Nerve growth factor phase shifts circadian activity rhythms in Syrian hamsters. Neurosci Lett. 1996;206:97–100.
Liu C, Gillette M. Cholinergic regulation of the suprachiasmatic nucleus circadian rhythm via a muscarinic mechanism at night. J Neurosci. 1996;16:744–51.
Navarria A, Wohleb ES, Voleti B, Ota KT, Dutheil S, Lepack AE, et al. Rapid antidepressant actions of scopolamine: role of medial prefrontal cortex and M1-subtype muscarinic acetylcholine receptors. Neurobiol Dis. 2015;82:254–61.
Santini E, Sepulveda-Orengo M, Porter JT. Muscarinic receptors modulate the intrinsic excitability of infralimbic neurons and consolidation of fear extinction. Neuropsychopharmacol. 2012;37:2047–56.
Hut RA, Van Der Zee EA. The cholinergic system, circadian rhythmicity, and time memory. Behav Brain Res. 2011;221:466–80.
Lemos JC, Shin JH, Alvarez VA. Striatal cholinergic interneurons are a novel target of corticotropin releasing factor. J Neurosci. 2019;39:5647–61.
Maksymetz J, Joffe ME, Moran SP, Stansley BJ, Li B, Temple K, et al. M1 muscarinic receptors modulate fear-related inputs to the prefrontal cortex: implications for novel treatments of posttraumatic stress disorder. Biol Psychiatry. 2019;85:989–1000.
Grippo RM, Purohit AM, Zhang Q, Zweifel LS, Güler AD. Direct midbrain dopamine input to the suprachiasmatic nucleus accelerates circadian entrainment. Curr Biol. 2017;27:2465–75.e3.
Wang Y, Hu P, Shan Q, Huang C, Huang Z, Chen P, et al. Single-cell morphological characterization of CRH neurons throughout the whole mouse brain. BMC Biol. 2021;19:47.
Walsh JJ, Friedman AK, Sun H, Heller EA, Ku SM, Juarez B, et al. Stress and CRF gate neural activation of BDNF in the mesolimbic reward pathway. Nat Neurosci. 2014;17:27–9.
Yang J, Sun J, Lu Y, An T, Lu W, Wang J-H. Revision to psychopharmacology mRNA and microRNA profiles are associated with stress susceptibility and resilience induced by psychological stress in the prefrontal cortex. Psychopharmacology. 2020;237:3067–93.
Pascual Cuadrado D, Todorov H, Lerner R, Islami L, Bindila L, Gerber S, et al. Long‐term molecular differences between resilient and susceptible mice after a single traumatic exposure. Br J Pharmacol. 2022;179:4161–80.
Manners MT, Brynildsen JK, Schechter M, Liu X, Eacret D, Blendy JA. CREB deletion increases resilience to stress and downregulates inflammatory gene expression in the hippocampus. Brain Behav Immunity. 2019;81:388–98.
Ota SM, Suchecki D, Meerlo P. Chronic social defeat stress suppresses locomotor activity but does not affect the free-running circadian period of the activity rhythm in mice. Neurobiol Sleep Circadian Rhythms. 2018;5:1–7.
Ota SM, Kong X, Hut R, Suchecki D, Meerlo P. The impact of stress and stress hormones on endogenous clocks and circadian rhythms. Front Neuroendocrinol. 2021;63:100931.
Tahara Y, Shiraishi T, Kikuchi Y, Haraguchi A, Kuriki D, Sasaki H, et al. Entrainment of the mouse circadian clock by sub-acute physical and psychological stress. Sci Rep. 2015;5:11417.
Ha S, Redmond L. ERK mediates activity dependent neuronal complexity via sustained activity and CREB-mediated signaling. Dev Neurobiol. 2008;68:1565–79.
Lee D. Global and local missions of cAMP signaling in neural plasticity, learning, and memory. Front Pharmacol. 2015;6:161.
Wang H, Ferguson GD, Pineda VV, Cundiff PE, Storm DR. Overexpression of type-1 adenylyl cyclase in mouse forebrain enhances recognition memory and LTP. Nat Neurosci. 2004;7:635–42.
Yang M, Ding Q, Zhang M, Moon C, Wang H. Forebrain overexpression of type 1 adenylyl cyclase promotes molecular stability and behavioral resilience to physical stress. Neurobiol Stress. 2020;13:100237.
Lee C-W, Fang Y-P, Chu M-C, Chung Y-J, Chi H, Tang C-W, et al. Differential mechanisms of synaptic plasticity for susceptibility and resilience to chronic social defeat stress in male mice. Biochem Biophys Res Commun. 2021;562:112–8.
Christoffel DJ, Golden SA, Russo SJ. Structural and synaptic plasticity in stress-related disorders. Rev. Neurosci. 2011;22:535–49.
Wang J, Hodes GE, Zhang H, Zhang S, Zhao W, Golden SA, et al. Epigenetic modulation of inflammation and synaptic plasticity promotes resilience against stress in mice. Nat Commun. 2018;9:477.
Lalive AL, Nuno‐Perez A, Tchenio A, Mameli M. Mild stress accumulation limits GABAergic synaptic plasticity in the lateral habenula. Eur J Neuroscience. 2022;55:377–87.
Chen KJ, Yoshimura R, Edmundo CA, Truong TM, Civelli O, Alachkar A, et al. Behavioral and neuro-functional consequences of eliminating the KCNQ3 GABA binding site in mice. Front Mol Neurosci. 2023;16:1192628.
Reichmann F, Holzer P. Neuropeptide Y: a stressful review. Neuropeptides. 2016;55:99–109.
Sweis BM, Veverka KK, Dhillon ES, Urban JH, Lucas LR. Individual differences in the effects of chronic stress on memory: behavioral and neurochemical correlates of resiliency. Neuroscience. 2013;246:142–59.
Ip CK, Zhang L, Farzi A, Qi Y, Clarke I, Reed F, et al. Amygdala NPY circuits promote the development of accelerated obesity under chronic stress conditions. Cell Metabolism. 2019;30:111–28.e6.
Chuang J-C, Cui H, Mason BL, Mahgoub M, Bookout AL, Yu HG, et al. Chronic social defeat stress disrupts regulation of lipid synthesis. J Lipid Res. 2010;51:1344–53.
Bartlang MS, Oster H, Helfrich-Förster C. Repeated psychosocial stress at night affects the circadian activity rhythm of male mice. J Biol Rhythms. 2015;30:228–41.
Kong X, Ota SM, Suchecki D, Lan A, Peereboom AI, Hut RA, et al. Chronic social defeat stress shifts peripheral circadian clocks in male mice in a tissue-specific and time-of-day dependent fashion. J Biol Rhythms. 2022;37:164–76.
Ota SM, Hut RA, Riede SJ, Crosby P, Suchecki D, Meerlo P. Social stress and glucocorticoids alter PERIOD2 rhythmicity in the liver, but not in the suprachiasmatic nucleus. Horm Behav. 2020;120:104683.
Shieh K-R. Distribution of the rhythm-related genes rPERIOD1, rPERIOD2, and rCLOCK, in the rat brain. Neuroscience. 2003;118:831–43.
Yamamoto S, Shigeyoshi Y, Ishida Y, Fukuyama T, Yamaguchi S, Yagita K, et al. Expression of thePer1 gene in the hamster: Brain atlas and circadian characteristics in the suprachiasmatic nucleus. J Comp Neurol. 2001;430:518–32.
Bush BJ, Donnay C, Andrews E-JA, Lewis-Sanders D, Gray CL, Qiao Z, et al. Non-rapid eye movement sleep determines resilience to social stress. eLife. 2022;11:e80206.
Cui Y, Yang Y, Ni Z, Dong Y, Cai G, Foncelle A, et al. Astroglial Kir4.1 in the lateral habenula drives neuronal bursts in depression. Nature. 2018;554:323–7.
Acknowledgements
The authors thank Professor Justin Blau and Professor Dan Ohtan Wang for critical reading of the manuscript draft. The authors have received funding from the following sources: NYUAD Start-Up Fund (DC), NYUAD Annual Research Fund (DC), NYUAD Research Enhancement Fund (DC), University Research Challenge Fund (DC), Al Jalila Research Foundation (AJF201638; DC), NYUAD Center for Brain and Health (NYUAD Research Institute CG012 Tamkeen grant; DC), NYUAD Center for Genomics and Systems Biology (NYUAD Research Institute ADHPG-CGSB Tamkeen grant; PN). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Figure 1 was created in Biorender (Petkovic, A. (2024) BioRender.com/g34m454).
Author information
Authors and Affiliations
Contributions
Conceptualisation: Priyam Narain, Aleksa Petković, Marko Šušić, Dipesh Chaudhury Data curation: Aleksa Petković, Marko Šušić, Priyam Narain Formal analysis: Marko Šušić, Aleksa Petković, Priyam Narain, Giuseppe Antonio-Saldi, Nizar Drou Visualisation: Marko Šušić, Aleksa Petković, Giuseppe Antonio-Saldi, Priyam Narain Validation: Aleksa Petković, Priyam Narain, Marko Šušić, Giuseppe Antonio-Saldi Funding acquisition: Dipesh Chaudhury Investigation: Aleksa Petković, Marko Šušić, Priyam Narain, Salma Haniffa, Marc Arnoux, Mariam Anwar Methodology: Priyam Narain, Aleksa Petković, Marko Šušić, Dipesh Chaudhury Project administration: Priyam Narain, Dipesh Chaudhury Resources: Dipesh Chaudhury Software: Marko Šušić, Giuseppe Antonio-Saldi, Aleksa Petković, Nizar Drou, Priyam Narain, Writing-original draft: Aleksa Petković, Priyam Narain, Dipesh Chaudhury Writing-review-editing: Aleksa Petković, Priyam Narain, Dipesh Chaudhury, Giuseppe Antonio-Saldi, Marko Šušić Supervision: Dipesh Chaudhury.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Narain, P., Petković, A., Šušić, M. et al. Nighttime-specific differential gene expression in suprachiasmatic nucleus and habenula is associated with resilience to chronic social stress. Transl Psychiatry 14, 407 (2024). https://doi.org/10.1038/s41398-024-03100-w
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41398-024-03100-w