Abstract
Type 2 diabetes mellitus (T2DM) and major depressive disorder (MDD) are prevalent chronic conditions that frequently co-occur, suggesting potential shared mechanisms. Neuroimaging meta-analyses were conducted to identify gray matter volume (GMV) alterations in T2DM and MDD, with further identification of the overlapping regions between the two diseases. Large-scale genome-wide association study (GWAS) summary statistics were utilized, and conjunctional false discovery rate (conjFDR) analysis was applied to identify shared genes. Transcriptome-neuroimaging association and expression-trait association analyses were performed to identify and prioritize shared genes contributing to these structural brain changes. Knockout mouse models were employed to investigate the functional roles of the prioritized genes. Neuroimaging meta-analysis revealed significant GMV reductions in T2DM and MDD patients compared to controls, with overlapping decreases, such as the bilateral median cingulate gyri, right insula and superior temporal gyrus. ConjFDR analysis identified 664 genes shared between T2DM and MDD, with nine prioritized genes linked to GMV alterations. Knockout models of these genes revealed neural and metabolic abnormalities, highlighting their potential roles in shared neurobiological and metabolic pathways underlying both disorders. This integrative analysis identifies shared neuroimaging and molecular mechanisms between T2DM and MDD, offering novel insights into their comorbidity and underlying biological pathways.
Similar content being viewed by others
Introduction
Type 2 diabetes mellitus (T2DM) and major depressive disorder (MDD) are two of the most prevalent chronic health conditions, both contributing significantly to the global disease burden [1, 2]. T2DM primarily affects glucose metabolism [3], whereas MDD is characterized by persistent mood disturbances and emotional dysregulation [4]. Despite these distinct clinical manifestations, accumulating evidence has revealed a robust bidirectional relationship between the two disorders. A recent large-scale epidemiological study has reported significant comorbidity [4], with depression increasing the risk of diabetes-related hospitalization by fivefold in a major prospective cohort [5]. Moreover, depression is associated with worse outcomes among individuals with T2DM, including more complications and higher mortality [6]. However, the underlying mechanisms linking T2DM and MDD remain poorly understood, particularly at the molecular and biological level. Elucidating these shared pathophysiological pathways is crucial, as the co-occurrence of T2DM and MDD not only exacerbates disease burden but also complicates clinical management and impairs treatment outcomes.
Exploring the structural brain changes associated with T2DM and MDD is key to uncovering the neurobiological links between these two diseases [7, 8]. Among the various neuroimaging markers available, gray matter volume (GMV) is particularly relevant due to its stable measurement ability [9] and critical role in supporting cognitive functions [10], which is typically impaired in these disease [11, 12]. Identifying the shared GMV changes between T2DM and MDD could provide important insights into their common neurobiological mechanisms. However, neuroimaging studies within each disease have reported mixed findings regarding GMV changes. For example, in T2DM, while some studies have observed significant GMV reductions in regions such as the temporal cortex [13, 14], others have reported no significant changes in any region [15, 16]. Similarly, in MDD, GMV reductions are frequently reported in the inferior temporal and fusiform regions [17, 18], yet other research has failed to detect significant changes in whole brain [19] or identified GMV abnormalities in other brain regions [20]. These inconsistencies likely arise due to variations in sample size, methodologies, and population characteristics. To resolve these discrepancies, neuroimaging meta-analysis serves as a powerful approach to synthesize findings across multiple studies, providing more robust and reliable results [21,22,23,24,25]. By focusing on GMV alterations, neuroimaging meta-analyses can offer a clearer understanding of the common brain structural changes associated with T2DM and MDD, laying the foundation for identifying shared neurobiological mechanisms.
Both T2DM and MDD are highly heritable conditions, with heritability estimates of over 25% for T2DM [26] and 30–50% for MDD [27], indicating that genetic factors play a significant role in their development and progression. Genome-wide association studies (GWAS) have identified multiple genetic variants associated with each disease [28, 29], and recent research has suggested a genetic correlation between T2DM and MDD [30]. This implies that shared genetic factors may contribute to the susceptibility of both diseases, potentially explaining their frequent co-occurrence. Despite these findings, the exact genetic mechanisms driving GMV changes in both diseases are still not well understood. Investigating the genetic factors underlying GMV alterations is critical for understanding the neurobiological pathways that connect T2DM and MDD. Transcriptome-neuroimaging association analysis integrates gene expression data with neuroimaging phenotypes [31,32,33], providing an effective method for exploring shared genetic mechanisms and identifying specific genes linked to GMV changes. By utilizing resources like the Allen Human Brain Atlas (AHBA) [34], researchers can map gene expression profiles in regions where GMV alterations are observed in T2DM and MDD, uncovering the genetic drivers that contribute to these structural brain changes and advancing our understanding of the common pathways involved in both diseases.
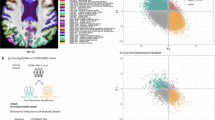
In this study, we aimed to investigate the shared molecular mechanisms underlying GMV changes in T2DM and MDD. We first conducted neuroimaging meta-analyses to identify GMV alterations specific to each disease and pinpoint the common changes occurring in both conditions. Following this, we utilized conjunctional false discovery rate (conjFDR) methods [35] to identify genes shared between the two diseases based on recent GWAS summary data for T2DM and MDD [28, 29]. Next, we performed transcriptome-neuroimaging association analyses with these shared genes, employing the AHBA to uncover the genetic drivers contributing to these structural brain changes. The identified shared genes were subsequently prioritized through expression-trait association analyses. Finally, knockout mouse models were employed to investigate the functional roles of the prioritized genes. The findings from this study not only enhance our understanding of the neurobiological mechanisms linking T2DM and MDD but also highlight potential genetic targets for future therapeutic interventions aimed at alleviating the cognitive and emotional impairments commonly associated with these diseases. An outline of the analytical framework is illustrated in Fig. 1.
A Neuroimaging meta-analyses were conducted to identify GMV alterations in T2DM and MDD, and to determine the overlapping regions of GMV reduction between the two conditions. B Shared genes were identified using conjFDR analysis applied to large-scale GWAS summary statistics for T2DM and MDD. Gene expression profiles were extracted from the AHBA database to construct the gene expression matrix. C Transcriptome-neuroimaging association analysis combined voxel-wise Z-values of GMV changes with the expression matrix of shared genes to identify genes associated with GMV alterations in each disorder and their overlap. D Expression-trait association analyses were used to prioritize shared genes, and functional validation was conducted through phenotypic profiling of knockout mouse models. AHBA allen human brain atlas, conjFDR conjunctional false discovery rate, GMV gray matter volume, HC healthy control, MDD major depression disorder, T2DM type 2 diabetes mellitus.
Materials and methods
Search strategy and selection criteria
A systematic search was conducted in the PubMed, Web of Science, and Embase databases to retrieve studies published prior to September 2024 using the following search terms: “voxel-based morphometry” or “VBM”, and “gray matter volume” or “grey matter volume” or “GMV”, and “major depressive disorder” or “major depression” or “MDD” or “type 2 diabetes mellitus” and “T2DM”. To identify additional studies, we also manually searched the reference lists of the included articles. Studies were included if they met all of the following criteria: (1) published in peer-reviewed English journals; (2) employed the voxel-based morphometry (VBM) approach to analyze whole-brain GMV in patients with MDD or T2DM compared to healthy controls; (3) provided peak coordinates of GMV alterations along with the corresponding statistical values in Montreal Neurological Institute (MNI) or Talairach space, or reported null findings; and (4) included participants aged between 18 and 65 years. Studies were excluded if they met any of the following criteria: (1) non-original research, such as meta-analyses, editorials, or review articles; or (2) included participants with comorbidities involving other mental illnesses. In cases where patient groups overlapped with those of another study, the study with the larger sample size was retained. If an article reported multiple independent patient samples, these were treated as separate datasets. Our meta-analysis was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines [36], and the detailed study selection procedures are summarized in Fig. 2.
A Study selection process for T2DM. B Study selection process for MDD. GMV gray matter volume, HC healthy control, MDD major depressive disorder, N number of studies, T2DM type 2 diabetes mellitus.
The quality of the included studies was evaluated using a 10-point checklist, which assessed several key factors, including the demographic and clinical characteristics of the subjects, the methods used for image acquisition and analysis, and the quality of the reported results and conclusions (Supplementary Table 1). Each criterion was scored as 1, 0.5, or 0, depending on whether it was fully, partially, or not met (Supplementary Tables 2, 3). Studies scoring above 9 were considered eligible and were included in subsequent analyses. Additionally, the following data were extracted from the included studies: demographic information (e.g., sample size, mean age, gender), clinical characteristics (e.g., illness duration, medication status), and methodological details (e.g., field strength, statistical threshold, scanning parameters). The peak coordinates and relevant statistics (e.g., t-values or their equivalents) were also recorded.
TMUGH T2DM dataset
A T2DM dataset collected at Tianjin Medical University General Hospital (TMUGH) was included, comprising 156 right-handed individuals with T2DM and 148 healthy controls matched for age, sex, and handedness. Its inclusion in the meta-analysis increased the sample size and statistical power for detecting GMV alterations associated with T2DM. Detailed preprocessing and statistical procedures are provided in the Supplementary Materials.
Voxel-wise meta-analysis
A meta-analysis was performed using the Seed-based d Mapping with Permutation of Subject Images software (SDM-PSI, version 6.22; https://www.sdmproject.com/) to examine abnormal GMV changes in patients with T2DM and MDD. For each study, peak coordinates and corresponding statistical values (t-values, or P- and z-values converted to t-values via the SDM online converter) were utilized alongside the t-map from the TMUGH dataset to reconstruct full-brain effect-size images using default settings (full anisotropy = 1, full-width at half-maximum = 20 mm, voxel size = 2 mm, gray matter mask). Key steps included (1) generating multiple imputations for study and subject images, (2) performing group analyses on the imputed subject images within each study, and (3) conducting meta-analyses of the imputed study images using a random-effects model, with Rubin’s rules applied to integrate the final meta-analysis results [37]. Age was included as a covariate to control for potential confounding effects across both patient and control groups in the meta-analysis, as T2DM patients are typically older than those with MDD. To control for family-wise error, a subject-based nonparametric permutation test (1000 permutations) was conducted with threshold-free cluster enhancement (TFCE), setting significance at FWE-TFCE P < 0.05 with a minimum cluster extent of 100 voxels. Finally, heterogeneity across studies was assessed using Cochran’s Q test and I² statistics, and Egger’s test was employed to evaluate publication bias in significant findings [38].
To uncover shared neurobiological mechanisms, we performed SDM-PSI’s multimodal analysis [39] to identify common GMV changes across T2DM and MDD. Following the multimodal analysis, we conducted meta-regression analyses to examine how demographic and clinical factors moderated GMV changes within each condition. Specifically, for T2DM, we investigated the impact of variables such as the percentage of female participants, BMI, HbA1c levels, illness duration, and the proportion of patients on medication. For MDD, we assessed the percentage of female patients, onset age, illness duration, Hamilton Depression Rating Scale (HDRS) score, and the proportion of patients on medication. To assess the robustness of meta-analytic findings and explore potential sources of heterogeneity in MDD, we conducted subgroup analyses based on four consistently reported study-level variables: MRI field strength (1.5 vs. 3.0 T), scanner manufacturer (GE, Siemens, Philips), smoothing kernel size (8 mm vs. others), and medication status (medicated vs. unmedicated). These parameters were selected due to their frequent reporting and limited variability across included MDD datasets, enabling meaningful stratification. In contrast, subgroup analysis was not feasible for T2DM, given the smaller number of eligible studies (n = 15) and the limited availability of stratified metadata such as medication information.
Global genetic correlation between T2DM and MDD
GWAS summary statistics for T2DM were obtained from the DIAGRAM consortium, based on a large multi-ancestry meta-analysis involving 180,834 individuals with T2DM and 1,159,055 controls, partially derived from the UK Biobank [28]. For this study, we used the European ancestry subset, as it provided the largest sample size, comprising 80,154 cases and 853,816 controls. Depression summary statistics were obtained from a meta-analysis of six datasets, involving 371,184 individuals with depression and 978,703 controls, all of European ancestry [29]. To prevent sample overlap, which could introduce bias in the conjFDR analysis, participants from the UK Biobank were excluded from the depression dataset, and samples from 23andMe were removed due to access restrictions, resulting in a refined GWAS subset with 166,773 cases and 507,679 controls. Genome-wide genetic correlation between T2DM and MDD was estimated using linkage disequilibrium score regression (LDSC; https://github.com/bulik/ldsc). Analyses were restricted to HapMap3 SNPs, and LD scores were derived from the European reference panel of the 1000 Genomes Project.
Shared genes between T2DM and MDD
Conditional FDR (condFDR) and conjFDR analyses were used to enhance discovery for each trait and identify shared genetic variants between T2DM and MDD, respectively. CondFDR, based on an empirical Bayesian framework, increases the power of GWAS by identifying significant loci and quantifying genetic overlap across complex traits. ConjFDR extends condFDR by prioritizing SNPs associated with a primary trait (e.g., T2DM) through conditional adjustment for a secondary trait (e.g., MDD), thereby leveraging the combined power of two GWAS to enhance variant identification [40]. For more detailed methodological descriptions, please refer to the Supplementary Materials. We set the significance threshold for condFDR analysis at 0.01 and conjFDR analyses at 0.05, following prior studies [41,42,43]. To minimize bias from complex linkage disequilibrium (LD) patterns, SNPs within the MHC (chr6: 25,119,106-33,854,733), 8p23.1 (chr8: 7,200,000-12,500,000), and MAPT (chr17: 40,000,000-47,000,000) were excluded before fitting the model [44]. To visualize the cross-trait genetic enrichment of T2DM and MDD, quantile-quantile (Q-Q) plots were generated at four increasingly P-value thresholds (P < 1, P < 0.1, P < 0.01, and P < 0.001), where leftward deflection from the expected line indicated cross-trait enrichment as the strength of association in the secondary trait increased.
To identify genes shared between T2DM and MDD, we used FUMA (https://fuma.ctglab.nl/) with the following steps. Independent significant SNPs were selected based on the conjFDR < 0.05 and were required to be independent of each other, with r2 < 0.6 [40, 45, 46]. Candidate SNPs were defined as those with conjFDR < 0.10 and LD r2 ≥ 0.6 with at least one independent significant SNP. These candidate SNPs were then mapped to protein-coding genes using three complementary mapping approaches: positional mapping, linking SNPs to genes within a 10 kb physical distance; eQTL mapping, associating SNPs with genes affected by allelic variation in expression; and chromatin interaction mapping, which uses 3D genome structure to link SNPs to both nearby and distant genes. Genes identified by any of these mapping methods were considered shared genes between T2DM and MDD.
To further investigate the genomic loci shared between T2DM and MDD and better understand the biological roles of candidate SNPs, we performed the following steps. First, we identified lead SNPs among the independent significant SNPs as those in LD with other significant SNPs at r2 < 0.1. For each lead SNP, a locus was established by including its associated independent SNPs and their related candidate SNP. Loci within 250 kb of each other were merged to form a single genomic locus, and LD information for this analysis was obtained from the European ancestry reference panel of the 1000 Genomes Project. Second, all candidate SNPs were annotated with combined annotation-dependent depletion (CADD) scores, RegulomeDB scores, and chromatin states. CADD scores assess the potential deleteriousness of each variant by integrating multiple annotations, such as evolutionary conservation and predicted functional impacts, with higher scores suggesting a greater likelihood of pathogenicity [47]. A CADD score threshold of 12.37 was applied to identify variants with potential pathogenic effects. RegulomeDB scores measure the regulatory potential of SNPs, with values ranging from 1–7; lower scores (e.g., 1a, 1b) represent stronger regulatory evidence, helping to pinpoint SNPs likely involved in gene regulation [48]. Chromatin states classify genomic regions based on epigenetic markers, identifying functional elements such as promoters, enhancers, and heterochromatic regions [49, 50].
Transcriptome-neuroimaging association analysis
Gene expression data were sourced from the AHBA database, a comprehensive repository that provides spatially mapped gene expression data across the human brain, derived from postmortem samples of six neurotypical adult donors [34] (Supplementary Table 4). This dataset encompasses over 20,000 genes, measured with approximately 58,000 probes across 3702 tissue samples from various brain regions. The abagen toolbox (https://www.github.com/netneurolab/abagen) [51] was used to extract and preprocess the gene expression data, following several key steps [52, 53]. First, gene probes were reannotated, and those that were unreliable or unmatched to genes were discarded [54]. An intensity-based filter was then applied, removing probes whose intensity was below background noise in at least 50% of tissue samples. The MNI coordinates of the tissue samples were aligned with the anatomical images using the alleninf package (https://github.com/chrisgorgo/alleninf). For each gene, the probe exhibiting the highest differential stability was selected as its representative. Gene expression values were subsequently normalized using a scaled robust sigmoid function, which was applied across genes for each sample and donor and across samples for each gene and donor. Additional normalization was performed within each distinct brain structure (cortex, subcortical/brain stem, and cerebellum) to adjust for differences in gene expression patterns among these regions. After preprocessing, the shared genes identified through conjFDR analysis for T2DM and MDD were extracted from the resulting gene expression matrix, and used for subsequent transcriptome-neuroimaging association analysis.
Using the unthresholded voxel-wise meta-analytic z-map representing GMV differences between patients with T2DM/MDD and healthy controls, we calculated the mean z-value within a 6 mm radius sphere centered at each tissue sample’s MNI coordinate to represent the GMV difference for that sample. Within each disease group, cross-sample correlations between GMV differences and gene expression profiles were assessed using Pearson’s correlation to identify genes whose expression levels correlated with GMV alterations in T2DM and MDD, respectively. To control for multiple comparisons, the Benjamini-Hochberg FDR method was applied (FDR P < 0.05). Genes associated with GMV alterations in both T2DM and MDD were identified by intersecting the results from the two separate cross-spatial correlation analyses, highlighting potential shared genetic factors underlying GMV changes in both conditions. To assess the significance of these shared genes, we conducted a permutation test (1000 iterations) using the Brain Surrogate Maps with Autocorrelated Spatial Heterogeneity (BrainSMASH) method [55], with significance set at Pperm < 0.05.
Prioritizing shared genes using expression-trait association analyses
To further refine the shared genes identified through the transcriptome-neuroimaging association analysis, we employed expression-trait association analyses to prioritize genes potentially relevant to GMV alterations in both T2DM and MDD. We used S-PrediXcan [56], a method that estimates genetically regulated gene expression in specific tissues and correlates it with complex traits, to conduct single-tissue expression-trait association analyses. This approach integrates GWAS summary data with pre-computed eQTL prediction models derived from individual brain tissues available in the Genotype-Tissue Expression (GTEx v8) dataset [57], including the amygdala, anterior cingulate cortex (BA24), caudate, cerebellar hemisphere, cerebellum, cortex, frontal cortex (BA9), hippocampus, hypothalamus, nucleus accumbens, putamen, cervical spinal cord C1, and substantia nigra. Next, we employed S-MultiXcan [58], an extension of S-PrediXcan, to combine results across tissues, increasing statistical power and identifying shared genetic effects across brain regions. Genes lacking significant brain eQTL associations were excluded, and significant expression-trait associations were identified using a Benjamini-Hochberg FDR-corrected threshold of P < 0.05.
Abnormal phenotypes in knockout mouse models of prioritized shared genes
To explore the functional relevance of shared genes between T2DM and MDD, we leveraged the Mouse Genome Informatics (MGI) database (https://www.informatics.jax.org/), a comprehensive international resource dedicated to laboratory mice. MGI database provides valuable information on gene mutations, knockout models, and associated phenotypic markers, enabling us to investigate the phenotypic consequences of gene loss in mice. Genes retrieved from in the MGI database were subsequently queried for human homologs using the Ensembl BioMart platform (https://mart.ensembl.org/) to ensure cross-species validity and translational relevance of our findings. Our analysis primarily focused on phenotypic changes observed in the nervous system and metabolic system, particularly those related to brain structure, function, or fasting circulating glucose level. By examining the abnormal behavioral and neurobiological phenotypes of the shared genes in knockout or mutant mouse model, we aimed to explore how the loss of these genes may affect behavior, such as activity levels, anxiety, or cognitive function, which could be relevant to both T2DM and MDD.
Results
Included studies in meta-analysis
Our systematic search and selection process, as illustrated in Fig. 2, initially identified 211 records for T2DM and 1348 records for MDD through database searches. After removing duplicates, 157 records for T2DM and 868 for MDD were screened by title and abstract, which led to the exclusion of 107 T2DM and 693 MDD records, leaving 50 T2DM and 175 MDD articles for eligibility assessment. After applying rigorous inclusion and exclusion criteria for full-text screening, 15 studies on T2DM and 50 on MDD were finally selected for meta-analysis, with all selected studies receiving a quality score above 9 (Supplementary Tables 2, 3). Among these, 2 T2DM studies [59, 60] and 6 MDD studies [18, 20, 61,62,63,64] included different patient subgroups, which we treated as independent datasets. Additionally, we incorporated our own TMUGH T2DM dataset, comprising 156 T2DM patients and 148 healthy controls. In total, our meta-analysis included 18 datasets for T2DM (860 patients and 725 controls) and 57 datasets for MDD (3867 cases and 3810 controls). Detailed demographic information, clinical characteristics, and methodological specifics for each dataset are provided in Supplementary Tables 5, 6.
GMV differences in meta-analysis
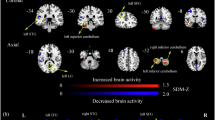
Compared to healthy controls, T2DM patients exhibited significant GMV reductions, primarily in the bilateral insula, bilateral median cingulate/paracingulate gyri (MCC), bilateral medial orbital part of superior frontal gyrus (SFGmO), and right superior temporal gyrus (STG). Additional reductions were observed in the crus I of left cerebellum, lobule IV/V of right cerebellum, left inferior occipital gyrus (IOG), inferior parietal gyrus (IPG), and left triangular and opercular parts of inferior frontal gyrus (IFGtriang/IFGoperc) (FWE-TFCE P < 0.05, Fig. 3A and Supplementary Table 7). In individuals with MDD, significant GMV decreases were observed predominantly in the bilateral SFGmO, bilateral IFGtriang, left IFGoperc, bilateral insula, left middle temporal gyrus (MTG), right middle frontal gyrus (MFG), right STG, bilateral MCC and the lobule VIII of right cerebellar hemisphere (FWE-TFCE P < 0.05, Fig. 3B and Supplementary Table 8). No regions with increased GMV were identified in either T2DM or MDD groups. The consistency of these findings was evaluated using Cochran’s Q test and I² statistics, both of which indicated no significant between-study heterogeneity, suggesting consistent GMV alterations across the studies. Furthermore, Egger’s test confirmed the absence of publication bias for the significant clusters (Supplementary Tables 7, 8).
A GMV reductions in T2DM patients compared to healthy controls. B GMV reductions in MDD patients compared to healthy controls. C Shared GMV reductions observed in both T2DM and MDD. For overlapping regions, the SDM-Z value with the smaller absolute magnitude was used as the representative value. The color bar indicates SDM-Z values, where darker colors correspond to larger decreases in GMV. Statistical maps are thresholded using FWE-TFCE-corrected P < 0.05 and cluster extent ≥ 100 voxels. GMV gray matter volume, IFG inferior frontal gyrus, INS insula, IOG inferior occipital gyrus, L left, MCC median cingulate/paracingulate gyri, MDD major depressive disorder, MFG median frontal gyrus, MTG middle temporal gyrus, ORBsup orbital part of the superior frontal gyrus, R right, SDM seed-based d mapping, SFG superior frontal gyrus, STG superior temporal gyrus.
The multimodal analysis revealed shared GMV reductions in T2DM and MDD patients, specifically in the bilateral MCC/SFGmO, left IFGtriang/IFGoperc, and right insula/STG (Fig. 3C, Supplementary Table 9). These results point to potential common neurobiological mechanisms underlying both conditions. In T2DM group, meta-regression analysis showed that GMV in the left IFGtriang was negatively correlated with BMI, GMV in the bilateral MCC was negatively correlated with HbA1c levels, and GMV in the lobule IV-V of left cerebellum was negatively associated with disease duration. In contrast, for MDD, meta-regression analysis indicated that GMV in the left MCC was negatively associated with HDRS scores. Detailed meta-regression results are provided in the Supplementary Table 10. Subgroup analyses in MDD revealed that the core regions exhibiting gray matter volume reductions—such as the bilateral IFG, left MTG, and right insula—were broadly consistent across strata defined by field strength, scanner manufacturer, smoothing kernel size, and medication status. These convergent patterns support the stability of the primary meta-analytic findings across varying methodological conditions.
Genetic correlation and shared genes between T2DM and MDD
LDSC revealed a significant positive genetic correlation between T2DM and MDD, with an rg of 0.219 and a P value of 2.05 × 10−22, indicating substantial polygenic overlap. As shown in Fig. 4A, the conditional Q-Q plot for T2DM exhibited a leftward shift with decreasing P-values for MDD, and conversely, the Q-Q plot for MDD displayed a leftward shift with decreasing P-values for T2DM, confirming the polygenic overlap between the two traits. Using conjFDR, we identified 128 independent significant SNPs and 3635 candidate SNPs. Among the candidate SNPs, the majority were located in intronic (1830, 50.3%) and intergenic (1158, 31.9%) regions, with 26 (0.7%) SNPs located within ncRNA exons and 449 (12.3%) SNPs in intronic region. Additionally, approximately 4.68% (170/3635) of the candidate SNPs were indicated as potentially pathogenic, based on their CADD scores exceeding 12.37. Around 3.49% (127/3635) of the candidate SNPs exhibited a higher likelihood of regulatory functionality, as indicated by RegulomeDB scores ranging from 1b–2c, based on evidence from eQTLs and chromatin marks. Furthermore, 83.99% (3054/3635) of the candidate SNPs were located in regions with a predominantly open chromatin configuration, characterized by a chromatin state score of less than 8. The information of candidate SNPs is shown in Fig. 5A. Through three mapping methods (positional mapping, eQTL mapping, and chromatin interaction mapping), we identified 664 protein-coding genes associated with candidate SNPs within loci shared between T2DM and MDD, as shown in Supplementary Table 11.
A Conditional Q-Q plots showing the polygenic overlap between T2DM and MDD. The left panel illustrates the nominal -log10(P) versus empirical -log10(q) values for T2DM conditioned on MDD, while the right panel shows the reverse (MDD conditioned on T2DM). B Manhattan plot depicting -log10(conjFDR) values for SNPs across chromosomes. The dotted horizontal line represents the threshold for significant shared associations (conjFDR < 0.05). Independent lead SNPs are highlighted in red. conjFDR conjunctional false discovery rate, CHR chromosome, Q-Q quantile-quantile, T2DM type 2 diabetes mellitus, MDD major depression disorder.
A Distribution of functional categories, RegulomeDB scores, and minimum chromatin states for candidate SNPs in shared loci associated with T2DM and MDD. B Distribution of functional categories, RegulomeDB scores, and minimum chromatin states for lead SNPs in shared loci associated with T2DM and MDD. In panels A and B, the y-axis represents the functional categories used to classify candidate SNPs and lead SNPs separately, while the x-axis indicates log10-transformed number of candidate SNPs and lead SNPs within each category. C Heatmap summarizing the expression-trait association results of nine shared genes. Colors represent log10-transformed P-values, with darker shades indicating significant associations. ACC anterior cingulate cortex, FDR false discovery rate, CSC cervical spinal cord C1, MDD major depression disorder, NA not available, NAc nucleus accumbens, ncRNA_in intron-derived non-coding RNA, ncRNA_ex exon-derived non-coding RNA, T2DM type 2 diabetes mellitus, UTR3 3′ untranslated region, UTR5 5′ untranslated region.
Among the 128 independent SNPs, 80 lead SNPs were identified (Fig. 4B). Of these, 55% (44/80) were found in intronic regions, and 15% (12/80) were intronic SNPs identified as non-synonymous mutations. Four of 80 lead SNPs (rs325485, rs6953951, rs10837055, rs813043) had CADD greater than 12.37, indicating potential pathogenicity. Two SNPs (rs11629241 and rs8066513) exhibited a higher likelihood of regulatory function, as indicated by a RegulomeDB score of 2b. Additionally, 90% (72/80) of lead SNPs were located in regions characterized by a predominantly open chromatin configuration, with chromatin state scores of less than 8. Supplementary Table 12 and Fig. 5B provide detailed information on lead SNPs. Additionally, we discovered 73 distinct genomic loci, with the most significant locus at chr17:65788157-66098979 (lead SNP: rs8072225, conjFDR = 6.355 × 10−7). Detailed information on these loci is provided in Supplementary Table 13.
Identification of genes associated with GMV alterations in T2DM and MDD
After preprocessing the gene expression data from AHBA, 3235 samples and 15,633 genes were retained. The shared genes identified through conjFDR analysis were then extracted, resulting in a final gene expression matrix containing 3235 samples and 528 genes. Subsequently, transcriptome-neuroimaging association analysis was conducted to explore the relationship between gene expression and GMV alterations in both MDD and T2DM. These analyses identified spatial correlations between gene expression and GMV alterations, with 189 genes linked to T2DM, 286 genes associated with MDD, and 154 genes shared between both diseases (Pperm < 0.05). The details of result are shown in Supplementary Tables 14–16. Based on these associations, the prioritizing expression-trait association analysis identified nine genes (FADS3, GIGYF2, KPNA2, LONRF2, MON1A, NEGR1, NICN1, PLXNB1, and SPPL3) as high-confidence genes associated with GMV alterations in both T2DM and MDD (Supplementary Table 17 and Fig. 5C).
Mouse knockout models of identified novel genes
To investigate the functional relevance of the prioritized shared genes between T2DM and MDD, we utilized the MGI resource to explore knockout mouse models and their associated phenotypic modifications. For example, among the prioritized genes, GIGYF2 and NEGR1 exhibited various abnormal phenotypes related to the neural system, including abnormal neuron number, motor neuron degeneration, and decreased brain weight, highlighting the potential roles these genes play in maintaining neural integrity and function. Additionally, three of the prioritized genes (GIGYF2, LONRF2, and MON1A) were linked to metabolic abnormalities such as increased fasting circulating glucose levels and impaired glucose tolerance, which align with known disturbances in glucose metabolism observed in T2DM. These metabolic alterations suggest that these genes may regulate both neural functions and glucose homeostasis. Furthermore, all these four genes were found to be direct homologs in humans, suggesting their potential importance in shared neurobiological and metabolic pathways underlying both T2DM and MDD. Detailed information of all prioritized shared genes, as well as their phenotypic effects in knockout mouse models, is provided in Supplementary Table 18.
Discussion
To our knowledge, this study is the first to investigate shared neuroimaging and molecular mechanisms between T2DM and MDD. We identified significant commonalities in brain structural changes, with GMV reductions observed in regions such as the bilateral MCC/ORBsup, left IFGtriang/IFGoperc, and right insula/STG, which are associated with emotional regulation, cognitive functions, and metabolic processes. Complementing these neuroimaging findings, genetic analyses identified shared genes between T2DM and MDD. Transcriptome analysis linked these genetic factors to brain structural alterations, offering insights into pathways through which genetic risk may drive both neural and metabolic dysfunction. Together, these findings deepen our understanding of the shared biological basis of T2DM and MDD and provide a foundation for future research on comorbidity mechanisms.
In this study, shared reductions in GMV were observed in T2DM and MDD patients, highlighting potential neurobiological links between these conditions. The insula plays a critical role in interoceptive awareness, integrating physiological signals to create a sense of bodily state, and in emotional regulation [65, 66], where it supports adaptive responses to stress and emotional stimuli. Dysfunction in this region leads to poor emotional regulation, disrupted physiological states, and destroyed working memory, manifesting as heightened affective symptoms and disruptions in somatic perception commonly seen in T2DM and MDD [67, 68]. Similarly, the STG, involved in auditory processing and emotional perception [69, 70], contributes to interpreting social and emotional cues [71]. Structural changes in this region could lead to impaired auditory processing and difficulties in social interactions, with MDD patients often exhibiting reduced ability in sensory processing and integration [72] and T2DM patients experiencing related deficits in communication and perception [73]. The IFG, particularly the triangular and opercular parts, supports higher-order cognitive functions, such as language processing, inhibitory control, and goal-directed behavior [74, 75]. Thus, reductions in IFG GMV may lead to cognitive rigidity and impairments in verbal reasoning. These effects may be further exacerbated by metabolic dysfunction in T2DM [76]. The MCC and SFGmO, which are involved in decision-making, emotional regulation, and stress processing, also show reductions in GMV [77, 78], indicating deficits in managing cognitive and affective responses. The MCC supports attentional control and conflict monitoring [79], crucial for coping with stress and maintaining emotional resilience [80], while the SFGmO facilitates decision-making and emotional evaluation [78], with dysfunction in these regions linked to maladaptive stress responses and impaired behavioral regulation in both conditions. Together, these findings suggest shared structural changes that underpin the cognitive, emotional, and physiological disruptions characteristic of T2DM and MDD.
The conjFDR analysis was used to detect the shared genetic architecture between MDD and T2DM. Stratified conditional Q-Q plots revealed SNP enrichment between MDD and T2DM, highlighting substantial polygenic overlap. Additionally, a previous study revealed there exists genetic correlation between T2DM and MDD [30]. In the present study, we revealed significant genetic overlap between T2DM and MDD, identifying 73 shared loci. Among these, the most significant shared locus, (chr17:65788157-66098979, lead SNP: rs8072225) was located within the intronic region of the BPTF gene, which encodes a transcription factor involved in chromatin remodeling and has been linked to neurodevelopmental disorders [81, 82] and linked with depression [29]. This finding highlights the potential role of transcriptional regulation in common genetic basis of T2DM and MDD. Additionally, most of the shared candidate SNPs were found in intronic (50.3%) and intergenic (31.9%) regions, with a smaller proportion (4.7%) classified as potentially pathogenic based on high CADD scores. Many of these SNPs exhibited regulatory functionality, supported by RegulomeDB scores and chromatin state annotations, indicating their likely influence on gene expression [48, 50]. By integrating positional, eQTL, and chromatin interaction mapping, we identified 664 shared protein-coding genes, further underscoring the molecular connections between these two conditions. These results suggest that regulatory regions and transcriptional control play crucial roles in mediating the genetic links between T2DM and MDD, offering new insights into the molecular mechanisms underlying their comorbidity.
Building upon these findings, transcriptome–neuroimaging and expression–trait association analyses were conducted to identify and prioritize shared genes contributing to GMV changes in T2DM and MDD. Among the nine prioritized genes, several are involved in both neural and metabolic pathways. NEGR1 is associated with adult hippocampal neurogenesis, affective regulation [83], and systemic metabolism improvement [84]. FADS3 is linked to lipid metabolism and colocalizes with genetic signals for psychiatric disorders [85]. GIGYF2, a synapse-enriched transcript with specific neuronal localization [86], may regulate insulin-related signaling [87]. LONRF2 and MON1A are involved in neuronal energy metabolism; LONRF2 contributes to motor neuron survival and is implicated in neurodegeneration with metabolic disruption [88], while MON1A plays a role in autophagosome maturation, with dysfunction implicated in both neurodegeneration and altered energy regulation [89]. PLXNB1 has been associated with mood disorders through its effects on neurite outgrowth and brain connectivity [90], although its metabolic role remains to be clarified. NICN1 is involved in HbA1c-related metabolic signaling [91], while SPPL3 is functionally implicated in protein modification, cellular metabolism, and immune-inflammatory responses, and has been linked to MDD risk [92].
The MGI database suggests that mouse gene knockout phenotypes resemble the clinical manifestations of human diseases. The mouse model exhibited abnormality in the nervous system, including neurodegeneration and reduced brain weight, which align with neuropathological changes reported in human MDD [93, 94] and T2DM cases [95, 96]. Additionally, the mice exhibited abnormal fasting circulating glucose levels, which is a hallmark feature of T2DM patients and also a recognized risk factor for MDD [97]. The shared genes represent promising targets for multi-mechanistic interventions addressing T2DM-MDD comorbidity. Their dual involvement in neurobiological and metabolic processes suggests that interventions simultaneously enhancing neuroplasticity while improving metabolic function may yield synergistic benefits. Notably, targeting convergent pathways such as neuroinflammation (SPPL3), autophagy (MON1A), and neuronal energy metabolism (LONRF2) may vulnerabilities shared by both disorders. Future research should explore combinatorial approaches, while considering patient stratification based on these molecular signatures.
Despite the findings of this study, several limitations should be acknowledged. First, the datasets used in the meta-analysis were cross-sectional, limiting the ability to infer causal relationships between GMV alterations and disease phenotypes in T2DM and MDD. Second, while the subgroup analyses provided additional insight into potential sources of variability, some subgroup comparisons were based on a limited number of datasets. Therefore, these findings should be interpreted with caution and considered exploratory in nature. Third, the meta-analysis included an unbalanced number of datasets for T2DM (18 datasets) and MDD (57 datasets), which may impact the statistical power of the results. The disparity in sample sizes could lead to less reliable comparisons of GMV alterations between the two conditions. Fourth, while age was included as a covariate to control for age differences between T2DM and MDD patients, this adjustment may not entirely remove the impact of age on GMV alterations. Fifth, although most genes have identifiable orthologs in the MGI database, caution should be exercised when interpreting these results due to inherent interspecies differences between mice and humans. Future studies utilizing parallel experiments in non-human primates or organoid models are needed to address this gap. Finally, the inclusion of studies with varying methodologies, imaging protocols, and quality could introduce heterogeneity in the results, potentially affecting the consistency and reliability of the findings. Although heterogeneity tests indicated no significant differences, subtle methodological variations may still influence the results and their interpretation.
This study reveals shared neuroimaging and molecular signatures between T2DM and MDD, highlighting potential common pathways contributing to both disorders. By integrating genetic, transcriptomic, and neuroimaging data, we identified significant brain structural changes and underlying genetic mechanisms shared between T2DM and MDD. The prioritization of shared genes and their functional relevance in knockout mouse models suggests that both metabolic and neurobiological mechanisms may contribute to these conditions. These findings not only enhance our understanding of the shared genetic markers and brain structural alterations associated with T2DM and MDD but also provide a foundation for future research into shared therapeutic targets.
Data availability
All datasets used in this study are publicly accessible. The GWAS summary statistics of MDD were obtained from the iPSYCH consortium at https://ipsych.dk/en/research/downloads/; the GWAS summary statistics of T2DM were downloaded from the DIAGRAM consortium (http://diagram-consortium.org/downloads.html). The gene expression data were retrieved from the AHBA website (https://human.brain-map.org/static/download). The gene expression prediction models used for TWAS are available at https://zenodo.org/records/3519321#.XfEKwtF7m90.
Code availability
The code used in the present study is shared on public repositories:
abagen toolbox (https://github.com/rmarkello/abagen),
ConjFDR and condFDR (https://github.com/precimed/pleiofdr),
MetaXcan (https://github.com/hakyimlab/MetaXcan).
References
Marx W, Penninx B, Solmi M, Furukawa TA, Firth J, Carvalho AF, et al. Major depressive disorder. Nat Rev Dis Primers. 2023;9:44.
Ahmad E, Lim S, Lamptey R, Webb DR, Davies MJ. Type 2 diabetes. Lancet. 2022;400:1803–20.
Mason IC, Qian J, Adler GK, Scheer F. Impact of circadian disruption on glucose metabolism: implications for type 2 diabetes. Diabetologia. 2020;63:462–72.
Berk M, Köhler-Forsberg O, Turner M, Penninx B, Wrobel A, Firth J, et al. Comorbidity between major depressive disorder and physical diseases: a comprehensive review of epidemiology, mechanisms and management. World Psychiatry. 2023;22:366–87.
Frank P, Batty GD, Pentti J, Jokela M, Poole L, Ervasti J, et al. Association between depression and physical conditions requiring hospitalization. JAMA Psychiatry. 2023;80:690–9.
Possidente C, Fanelli G, Serretti A, Fabbri C. Clinical insights into the cross-link between mood disorders and type 2 diabetes: a review of longitudinal studies and Mendelian randomisation analyses. Neurosci Biobehav Rev. 2023;152:105298.
Zhao H, Wang F, Luo GH, Lei H, Peng F, Ren QP, et al. Assessment of structural brain changes in patients with type 2 diabetes mellitus using the MRI-based brain atrophy and lesion index. Neural Regen Res. 2022;17:618–24.
Zhang FF, Peng W, Sweeney JA, Jia ZY, Gong QY. Brain structure alterations in depression: psychoradiological evidence. CNS Neurosci Ther. 2018;24:994–1003.
Winkler AM, Kochunov P, Blangero J, Almasy L, Zilles K, Fox PT, et al. Cortical thickness or grey matter volume? The importance of selecting the phenotype for imaging genetics studies. Neuroimage. 2010;53:1135–46.
Kharabian Masouleh S, Eickhoff SB, Hoffstaedter F, Genon S. Empirical examination of the replicability of associations between brain structure and psychological variables. Elife. 2019;8:e43464.
MacKenzie LE, Uher R, Pavlova B. Cognitive performance in first-degree relatives of individuals with vs without major depressive disorder: a meta-analysis. JAMA Psychiatry. 2019;76:297–305.
Strachan MW, Reynolds RM, Marioni RE, Price JF. Cognitive function, dementia and type 2 diabetes mellitus in the elderly. Nat Rev Endocrinol. 2011;7:108–14.
Wang C, Fu K, Liu H, Xing F, Zhang S. Brain structural changes and their correlation with vascular disease in type 2 diabetes mellitus patients: a voxel-based morphometric study. Neural Regen Res. 2014;9:1548–56.
Wang Y, Yan Y, Wei J, Yang X, Wang M, Zhao L, et al. Down-regulated miR-16-2 in peripheral blood is positively correlated with decreased bilateral insula volume in patients with major depressive disorder. J Affect Disord. 2023;338:137–43.
Fang F, Lai MY, Huang JJ, Kang M, Ma MM, Li KA, et al. Compensatory hippocampal connectivity in young adults with early-stage type 2 diabetes. J Clin Endocrinol Metab. 2019;104:3025–38.
Chen J, Zhang J, Liu X, Wang X, Xu X, Li H, et al. Abnormal subcortical nuclei shapes in patients with type 2 diabetes mellitus. Eur Radiol. 2017;27:4247–56.
Yang Y, Li X, Cui Y, Liu K, Qu H, Lu Y, et al. Reduced gray matter volume in orbitofrontal cortex across schizophrenia, major depressive disorder, and bipolar disorder: a comparative imaging study. Front Neurosci. 2022;16:919272.
Arnone D, McKie S, Elliott R, Juhasz G, Thomas EJ, Downey D, et al. State-dependent changes in hippocampal grey matter in depression. Mol Psychiatry. 2013;18:1265–72.
Jia Z, Huang X, Wu Q, Zhang T, Lui S, Zhang J, et al. High-field magnetic resonance imaging of suicidality in patients with major depressive disorder. Am J Psychiatry. 2010;167:1381–90.
Peng H, Wu K, Li J, Qi H, Guo S, Chi M, et al. Increased suicide attempts in young depressed patients with abnormal temporal-parietal-limbic gray matter volume. J Affect Disord. 2014;165:69–73.
Müller VI, Cieslik EC, Laird AR, Fox PT, Radua J, Mataix-Cols D, et al. Ten simple rules for neuroimaging meta-analysis. Neurosci Biobehav Rev. 2018;84:151–61.
Tahmasian M, Sepehry AA, Samea F, Khodadadifar T, Soltaninejad Z, Javaheripour N, et al. Practical recommendations to conduct a neuroimaging meta-analysis for neuropsychiatric disorders. Hum Brain Mapp. 2019;40:5142–54.
Zhang Z, Zhang Y, Wang H, Lei M, Jiang Y, Xiong D, et al. Resting-state network alterations in depression: a comprehensive meta-analysis of functional connectivity. Psychol Med. 2025;55:e63.
Ma J, Xue K, Wang X, Cai M, Wang X, Li J, et al. Gray matter volume abnormalities in vascular cognitive impairment and their association with gene expression profiles. Meta-Radiology. 2023;1:100035.
Cai M, Wang R, Liu M, Du X, Xue K, Ji Y, et al. Disrupted local functional connectivity in schizophrenia: an updated and extended meta-analysis. Schizophrenia. 2022;8:93.
Prasad RB, Groop L. Genetics of type 2 diabetes-pitfalls and possibilities. Genes. 2015;6:87–123.
Kendall KM, Van Assche E, Andlauer TFM, Choi KW, Luykx JJ, Schulte EC, et al. The genetic basis of major depression. Psychol Med. 2021;51:2217–30.
Mahajan A, Spracklen CN, Zhang W, Ng MCY, Petty LE, Kitajima H, et al. Multi-ancestry genetic study of type 2 diabetes highlights the power of diverse populations for discovery and translation. Nat Genet. 2022;54:560–72.
Als TD, Kurki MI, Grove J, Voloudakis G, Therrien K, Tasanko E, et al. Depression pathophysiology, risk prediction of recurrence and comorbid psychiatric disorders using genome-wide analyses. Nat Med. 2023;29:1832–44.
Bergstedt J, Pasman JA, Ma Z, Harder A, Yao S, Parker N, et al. Distinct biological signature and modifiable risk factors underlie the comorbidity between major depressive disorder and cardiovascular disease. Nat Cardiovasc Res. 2024;3:754–69.
Cai M, Ji Y, Zhao Q, Xue H, Sun Z, Wang H, et al. Homotopic functional connectivity disruptions in schizophrenia and their associated gene expression. Neuroimage. 2024;289:120551.
Wang H, Zhao Q, Zhang Y, Ma J, Lei M, Zhang Z, et al. Shared genetic architecture of cortical thickness alterations in major depressive disorder and schizophrenia. Prog Neuropsychopharmacol Biol Psychiatry. 2024;135:111121.
Xue K, Liang S, Yang B, Zhu D, Xie Y, Qin W, et al. Local dynamic spontaneous brain activity changes in first-episode, treatment-naïve patients with major depressive disorder and their associated gene expression profiles. Psychol Med. 2022;52:2052–61.
Hawrylycz MJ, Lein ES, Guillozet-Bongaarts AL, Shen EH, Ng L, Miller JA, et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature. 2012;489:391–9.
Ghaffar A, Nyholt DR. Integrating eQTL and GWAS data characterises established and identifies novel migraine risk loci. Hum Genet. 2023;142:1113–37.
Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. Open Med. 2009;3:e123–30.
Albajes-Eizagirre A, Solanes A, Vieta E, Radua J. Voxel-based meta-analysis via permutation of subject images (PSI): theory and implementation for SDM. Neuroimage. 2019;186:174–84.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–60.
Radua J, Mataix-Cols D, Phillips ML, El-Hage W, Kronhaus DM, Cardoner N, et al. A new meta-analytic method for neuroimaging studies that combines reported peak coordinates and statistical parametric maps. Eur Psychiatry. 2012;27:605–11.
Smeland OB, Frei O, Shadrin A, O’Connell K, Fan CC, Bahrami S, et al. Discovery of shared genomic loci using the conditional false discovery rate approach. Hum Genet. 2020;139:85–94.
Andreassen OA, Thompson WK, Schork AJ, Ripke S, Mattingsdal M, Kelsoe JR, et al. Improved detection of common variants associated with schizophrenia and bipolar disorder using pleiotropy-informed conditional false discovery rate. PLoS Genet. 2013;9:e1003455.
Liu M, Wang L, Zhang Y, Dong H, Wang C, Chen Y, et al. Investigating the shared genetic architecture between depression and subcortical volumes. Nat Commun. 2024;15:7647.
Zhao Q, Wang S, Xiong D, Liu M, Zhang Y, Zhao G, et al. Genome-wide analysis identifies novel shared loci between depression and white matter microstructure. Mol Psychiatry. 2025;30:3455–65.
Schwartzman A, Lin X. The effect of correlation in false discovery rate estimation. Biometrika. 2011;98:199–214.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Zhao Q, Xu J, Shi Z, Zhang Y, Du X, Zhai Y, et al. Genome-wide pleiotropy analysis reveals shared genetic associations between type 2 diabetes mellitus and subcortical brain volumes. Research. 2025;8:0688.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–5.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30.
Ernst J, Kellis M. ChromHMM: automating chromatin-state discovery and characterization. Nat Methods. 2012;9:215–6.
Markello RD, Arnatkeviciute A, Poline JB, Fulcher BD, Fornito A, Misic B. Standardizing workflows in imaging transcriptomics with the abagen toolbox. Elife. 2021;10:e72129.
Arnatkeviciute A, Fulcher BD, Fornito A. A practical guide to linking brain-wide gene expression and neuroimaging data. Neuroimage. 2019;189:353–67.
Xue K, Guo L, Zhu W, Liang S, Xu Q, Ma L, et al. Transcriptional signatures of the cortical morphometric similarity network gradient in first-episode, treatment-naive major depressive disorder. Neuropsychopharmacology. 2023;48:518–28.
Arloth J, Bader DM, Röh S, Altmann A. Re-annotator: annotation pipeline for microarray probe sequences. PLoS One. 2015;10:e0139516.
Burt JB, Helmer M, Shinn M, Anticevic A, Murray JD. Generative modeling of brain maps with spatial autocorrelation. Neuroimage. 2020;220:117038.
Barbeira AN, Pividori M, Zheng J, Wheeler HE, Nicolae DL, Im HK. Integrating predicted transcriptome from multiple tissues improves association detection. PLoS Genet. 2019;15:e1007889.
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020;369:1318–30.
Barbeira AN, Dickinson SP, Bonazzola R, Zheng J, Wheeler HE, Torres JM, et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat Commun. 2018;9:1825.
Gao Y, Sui C, Chen B, Xin H, Che Y, Zhang X, et al. Voxel-based morphometry reveals the correlation between gray matter volume and serum P-tau-181 in type 2 diabetes mellitus patients with different HbA1c levels. Front Neurosci. 2023;17:1202374.
Zhang Y, Zhang X, Zhang J, Liu C, Yuan Q, Yin X, et al. Gray matter volume abnormalities in type 2 diabetes mellitus with and without mild cognitive impairment. Neurosci Lett. 2014;562:1–6.
Lu S, Wu C, Jia L, Fang Z, Lu J, Mou T, et al. Increased plasma levels of IL-6 are associated with striatal structural atrophy in major depressive disorder patients with anhedonia. Front Psychiatry. 2022;13:1016735.
Salvadore G, Nugent AC, Lemaitre H, Luckenbaugh DA, Tinsley R, Cannon DM, et al. Prefrontal cortical abnormalities in currently depressed versus currently remitted patients with major depressive disorder. Neuroimage. 2011;54:2643–51.
Serra-Blasco M, Portella MJ, Gómez-Ansón B, de Diego-Adeliño J, Vives-Gilabert Y, Puigdemont D, et al. Effects of illness duration and treatment resistance on grey matter abnormalities in major depression. Br J Psychiatry. 2013;202:434–40.
Zhao S, Du Y, Zhang Y, Wang X, Xia Y, Sun H, et al. Gray matter reduction is associated with cognitive dysfunction in depressed patients comorbid with subclinical hypothyroidism. Front Aging Neurosci. 2023;15:1106792.
Tan Y, Yan R, Gao Y, Zhang M, Northoff G. Spatial-topographic nestedness of interoceptive regions within the networks of decision making and emotion regulation: combining ALE meta-analysis and MACM analysis. Neuroimage. 2022;260:119500.
Gogolla N. The insular cortex. Curr Biol. 2017;27:R580–r6.
Coccaro EF, Lazarus S, Joseph J, Wyne K, Drossos T, Phillipson L, et al. Emotional regulation and diabetes distress in adults with type 1 and type 2 diabetes. Diabetes Care. 2021;44:20–5.
Zhou H, Liu J, Wu Y, Huang Z, Wang W, Ma Y, et al. Unveiling the interoception impairment in various major depressive disorder stages. CNS Neurosci Ther. 2024;30:e14923.
Nourski KV. Auditory processing in the human cortex: an intracranial electrophysiology perspective. Laryngoscope Investig Otolaryngol. 2017;2:147–56.
Yi HG, Leonard MK, Chang EF. The encoding of speech sounds in the superior temporal gyrus. Neuron. 2019;102:1096–110.
Klasen M, von Marschall C, Isman G, Zvyagintsev M, Gur RC, Mathiak K. Prosody production networks are modulated by sensory cues and social context. Soc Cogn Affect Neurosci. 2018;13:418–29.
Lu F, Cui Q, Huang X, Li L, Duan X, Chen H, et al. Anomalous intrinsic connectivity within and between visual and auditory networks in major depressive disorder. Prog Neuropsychopharmacol Biol Psychiatry. 2020;100:109889.
Hu XJ, Lau CC, Ruan RQ. Exploring auditory temporal resolution and dichotic listening skills among individuals with type 2 diabetes mellitus. Hear Res. 2024;450:109067.
Turker S, Kuhnke P, Eickhoff SB, Caspers S, Hartwigsen G. Cortical, subcortical, and cerebellar contributions to language processing: a meta-analytic review of 403 neuroimaging experiments. Psychol Bull. 2023;149:699–723.
Asplund CL, Todd JJ, Snyder AP, Marois R. A central role for the lateral prefrontal cortex in goal-directed and stimulus-driven attention. Nat Neurosci. 2010;13:507–12.
Amorim JA, Coppotelli G, Rolo AP, Palmeira CM, Ross JM, Sinclair DA. Mitochondrial and metabolic dysfunction in ageing and age-related diseases. Nat Rev Endocrinol. 2022;18:243–58.
Shackman AJ, Salomons TV, Slagter HA, Fox AS, Winter JJ, Davidson RJ. The integration of negative affect, pain and cognitive control in the cingulate cortex. Nat Rev Neurosci. 2011;12:154–67.
Manes F, Sahakian B, Clark L, Rogers R, Antoun N, Aitken M, et al. Decision-making processes following damage to the prefrontal cortex. Brain. 2002;125:624–39.
Zhang R, Geng X, Lee TMC. Large-scale functional neural network correlates of response inhibition: an fMRI meta-analysis. Brain Struct Funct. 2017;222:3973–90.
Sinha R, Lacadie CM, Constable RT, Seo D. Dynamic neural activity during stress signals resilient coping. Proc Natl Acad Sci USA. 2016;113:8837–42.
Richart L, Carrillo-de Santa Pau E, Río-Machín A, de Andrés MP, Cigudosa JC, Lobo VJS, et al. BPTF is required for c-MYC transcriptional activity and in vivo tumorigenesis. Nat Commun. 2016;7:10153.
Stankiewicz P, Khan TN, Szafranski P, Slattery L, Streff H, Vetrini F, et al. Haploinsufficiency of the chromatin remodeler BPTF causes syndromic developmental and speech delay, postnatal microcephaly, and dysmorphic features. Am J Hum Genet. 2017;101:503–15.
Noh K, Lee H, Choi TY, Joo Y, Kim SJ, Kim H, et al. Negr1 controls adult hippocampal neurogenesis and affective behaviors. Mol Psychiatry. 2019;24:1189–205.
Nigro P, Carbone N, Caputo T, Vamvini M, Columbus M, Hirshman M, et al. 8638 NEGR1 is a Mediator of Exercise-Induced Neuritogenesis in Subcutaneous White Adipose Tissue, Implications for Improved Systemic Metabolism. J Endocr Soc. 2024;8:bvae163.009.
Zhao Q, Baranova A, Liu D, Cao H, Zhang F. Bidirectional causal associations between plasma metabolites and bipolar disorder. Mol Psychiatry. 2025.
Yang J, Jin N, Zhang S, Tan Y, Chen Z, Huang X, et al. Genome-wide profiling and functional characterization of circular RNAs in neural development and injury: insights from a rat model research. Cell Mol Life Sci. 2025;82:135.
Zhao Y, Stankovic S, Koprulu M, Wheeler E, Day FR, Lango Allen H, et al. GIGYF1 loss of function is associated with clonal mosaicism and adverse metabolic health. Nat Commun. 2021;12:4178.
Li D, Johmura Y, Morimoto S, Doi M, Nakanishi K, Ozawa M, et al. LONRF2 is a protein quality control ubiquitin ligase whose deficiency causes late-onset neurological deficits. Nat Aging. 2023;3:1001–19.
Cai CZ, Zhuang XX, Zhu Q, Wu MY, Su H, Wang XJ, et al. Enhancing autophagy maturation with CCZ1-MON1A complex alleviates neuropathology and memory defects in Alzheimer disease models. Theranostics. 2022;12:1738–55.
Yang G, Ullah HMA, Parker E, Gorsi B, Libowitz M, Maguire C, et al. Neurite outgrowth deficits caused by rare PLXNB1 mutation in pediatric bipolar disorder. Mol Psychiatry. 2023;28:2525–39.
Taylor HJ, Hung YH, Narisu N, Erdos MR, Kanke M, Yan T, et al. Human pancreatic islet microRNAs implicated in diabetes and related traits by large-scale genetic analysis. Proc Natl Acad Sci USA. 2023;120:e2206797120.
Li X, Luo Z, Gu C, Hall LS, McIntosh AM, Zeng Y, et al. Common variants on 6q16.2, 12q24.31 and 16p13.3 are associated with major depressive disorder. Neuropsychopharmacology. 2018;43:2146–53.
Koolschijn PC, van Haren NE, Lensvelt-Mulders GJ, Hulshoff Pol HE, Kahn RS. Brain volume abnormalities in major depressive disorder: a meta-analysis of magnetic resonance imaging studies. Hum Brain Mapp. 2009;30:3719–35.
Wang H, He Y, Sun Z, Ren S, Liu M, Wang G, et al. Microglia in depression: an overview of microglia in the pathogenesis and treatment of depression. J Neuroinflammation. 2022;19:132.
Farhadi A, Vosough M, Zhang JS, Tahamtani Y, Shahpasand K. A possible neurodegeneration mechanism triggered by diabetes. Trends Endocrinol Metab. 2019;30:692–700.
Moran C, Beare R, Wang W, Callisaya M, Srikanth V. Type 2 diabetes mellitus, brain atrophy, and cognitive decline. Neurology. 2019;92:e823–e30.
Wang Q, Li Y, Ren H, Huang Q, Wang X, Zhou Y, et al. Metabolic characteristics, prevalence of anxiety and its influencing factors in first-episode and drug-naïve major depressive disorder patients with impaired fasting glucose. J Affect Disord. 2023;324:341–8.
Acknowledgements
Thanks to all the authors of the included studies for giving us the availability to use the data to perform the present meta-analysis.
Funding
This work was funded by the Tianjin Natural Science Foundation (19JCYBJC25100, 21JCYBJC01280, 23JCZXJC00120, 21ZXJBSY00120), the Beijing-Tianjin-Hebei Basic Research Collaboration Project (J230040), and Tianjin Key Medical Discipline Construction Project (TJYXZDXK-3-008C).
Author information
Authors and Affiliations
Contributions
JX, XD, and YZ (Ying Zhai): performed data curation, analyzed the data, and wrote the original draft. QZ (Qiyu Zhao) and YZ (Yang Zhang): validated the methodology and interpreted the data. HW and YW: verified the extracted data. YP: provided supervision and conceived the study concept. FL and QZ (Quan Zhang): secured funding, designed the study methodology, and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The T2DM data collection was approved by the Ethics Committee of Tianjin Medical University General Hospital (approval number: IRB2020-YX-009-01), and written informed consent was obtained from all participants. Additionally, written informed consent for the publication of identifiable images was obtained from all participants whose images are included in this article. All methods were performed in accordance with relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, J., Du, X., Zhai, Y. et al. Shared neuroimaging and molecular profiles in type 2 diabetes mellitus and major depressive disorder: an integrative analysis of genetic, transcriptomic, and neuroimaging data. Transl Psychiatry 15, 352 (2025). https://doi.org/10.1038/s41398-025-03585-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41398-025-03585-z