Abstract
Mantle cell lymphoma (MCL) is considered one of the most aggressive B-cell lymphoid neoplasms. The transcription factor SOX11 is aberrantly expressed in conventional aggressive MCL, while it is not or weakly expressed in the leukemic non-nodal MCL subtype with a predominantly indolent clinical evolution. SOX11 is a key driver of MCL through the regulation of several oncogenic mechanisms, suggesting that it may be interacting with different protein complexes to exert its multiple actions. Using proteomic strategies, we characterized the SOX11-interactome and validated its physical interaction with SMARCA4, the catalytic subunit of the SWI/SNF chromatin-remodeling complex. SMARCA4 expression is directly regulated by SOX11, and its upregulation significantly associates with worse outcomes of patients. Integration of global DNA-binding and transcriptomic profiles revealed that SOX11 and SMARCA4 share binding sites enriched in open chromatin and active promoters and regulate common key oncogenic pathways crucial for MCL progression and aggressiveness. The SMARCA4-specific PROTAC-degrader AU-15330 significantly reduced SOX11 binding to specific regulatory regions and diminished the activation of BCR-, NIK-, and BCL2-signaling pathways. Moreover, SMARCA4 degradation significantly reduced proliferation and induced apoptosis of SOX11-positive MCL cells, highlighting AU-15330 as a promising therapeutic approach for patients who may relapse from current target therapies in MCL.

Similar content being viewed by others
Introduction
Mantle cell lymphoma (MCL) is considered one of the most aggressive B-cell lymphoid neoplasms, with very short responses to treatment, frequent relapses, and poor outcome of the patients. Remarkably, a subset of patients with predominantly leukemic non-nodal clinical disease (nnMCL) has an indolent evolution without the requirement of treatment for a long time [1]. SRY-related HMG-box gene 11 (SOX11) is aberrantly expressed in conventional MCL (cMCL), and negative or very weakly expressed in the nnMCL subtype [2, 3]. SOX11 contributes to MCL aggressive behavior by directly regulating critical targets and oncogenic pathways responsible for blocking late B-cell differentiation, promoting tumor angiogenesis, migration, tumor-stromal protective interactions, immune evasion mechanisms, and stem cell features [4,5,6,7,8,9]. SOX11 also promotes the transformation and aberrant expansion of B1a cells, in part through the activation of the B-cell receptor (BCR), in an Eμ-SOX11-eGFP transgenic mouse model [10].
SOX11 is a member of the SOX (Sex determining Region Y-related HMG-box) family of transcription factors characterized by binding at the minor groove of the DNA through its High Mobility Group (HMG) domain, inducing bending that brings regulatory regions of the DNA into proximity and enables gene activation by interaction of bound transcription factors [11, 12]. Moreover, SOX proteins need to interact with other proteins to establish a specific and stable binding to regulatory regions to modulate the transcription of its specific targets [13, 14]. Recently, SOX11 has been shown to have pioneering activity, initiating chromatin opening and facilitating subsequent transcriptional activity [15]. Although SOX11 participates in many oncogenic pathways, its mechanism of action is still unknown. Recent findings open the possibility that SOX11 interacts with different factors to participate in the specific activation of distinct pathogenic functions in MCL [15].
Here, we have characterized the interactome map of SOX11 in MCL, validated the physical interaction of SOX11 with SMARCA4, the central ATPase catalytic subunit of the SWItch/Sucrose NonFermentable (SWI/SNF) chromatin-remodeling complex, and recognized common DNA-binding sites and direct target genes regulated by the SOX11:SMARCA4 complex driving distinct oncogenic pathways in MCL.
Materials and Methods
MCL cell lines and primary samples
Five SOX11-positive MCL cell lines, Z138 (ATCC CRL-3001), Granta-519 (DSMZ ACC-342), JEKO-1 (ATCC CRL-3006), MINO (ATCC CRL-3000), HBL2, and one negative, JVM2 (ATCC CRL-3002) cell line, were used for western blot (WB), In situ-fluorescence Proximity Labeling Assay (fPLA) and/or in vitro experiments. JVM2TRE-SOX11:BioID2 and JVM2TRE-BioID2 stably transduced cell lines were generated to perform BioID2-based proximity labeling proteomic (BioID2PL) experiments. Z138CT, Z138SOX11KO, Z138SOX11Resc [9], JVM2TRE, JVM2TRE-SOX11FL:HA, and/or Granta-519 were used for WB, Cleavage Under Targets & Release Using Nuclease (CUT&RUN), RNA-sequencing (RNA-seq), and in vitro experiments. Somatic mutations in SMARCA4, ARID1A, or SMARCC1 genes have not been detected in any of the MCL cell lines or primary cell samples used in this work [16]. HEK293T (ATCC CRL-3216) cell line was used for lentivirus production.
MCL primary tumors cryopreserved at the Hospital Clínic/FCRB Biobank were used two for in situ-fPLA and CUT&RUN experiments. See more details on cell culture and generation of stable transduced MCL cell lines in Supplementary Methods.
BioID2-proximal labeling (PL) experiments and mass spectrometry (MS) analysis
For MS experiments, three biological replicates of JVM2TRE-SOX11:BioID2 and JVM2TRE-BioID2 cell lines were incubated with 0.1 μg/mL doxycycline and 50 μM biotin for 24 h. Cells were collected, and nuclear protein extraction was performed. Nuclear extracts were incubated overnight at 4 °C with streptavidin-coated magnetic beads (Pierce). After elution, biotinylated proteins by SOX11:BioID2 or BioID2 (used as control for unspecific binding) were digested with trypsin and subjected to LC-MS/MS analysis. See more details on protein extraction, purification, MS experiments and analysis in Supplementary Methods.
In situ-fluorescents Proximal Labeling Analyisis (fPLA)
Specific human anti-SOX11, anti-SMARCA4, anti-ARID1A, and anti-SMARCC1 antibodies (Table S1) were used for in situ-fPLA experiments, performed according to the manufacturer’s instructions (Merk) in one million SOX11-positive (Z138, Granta-519, JVM2TRE-SOX11FL:HA, JVM2TRE-SOX11ΔHMG:HA, and JVM2TRE-SOX11ΔTA:HA) and SOX11-negative JVM2TRE MCL cell lines.
Analysis of SMARCA4 expression in MCL primary tumors
SMARCA4 expression in MCL primary tumors ( > 80% purified tumor cells) was investigated in three cohorts with gene expression profile (GEPs) information from microarray data of 54 leukemic purified MCL primary tumors (GSE79196) [17], lymph node or peripheral blood samples of 39 MCL primary tumors (EGAD00010001842) [18], and RNA-seq data of 16 purified tumor cells from MCL samples (10 SOX11-positive and 6 SOX11-negative) [9]. For survival analysis, we used data from 40 leukemic purified MCL primary tumors (GSE79196) [17]. The maximally selected rank (Maxstat) statistics defined SMARCA4 high and low values (cut-off =6.82 expression units).
SMARCA4 expression in MCL was analyzed by RT-qPCR using Fast SYBR Green Master (Applied Biosystems) using specific primers (Table S2).
CUT&RUN experiments and Analysis
CUT&RUN experiments were performed using Z138 (Z138CT), Z138SOX11KO [9], and Z138 cells treated with AU-15330 (0.5 μM) for 24 h (Z138AU-15330); JVM2TRE, JVM2TRE-SOX11FL:HA, and Granta-519. Three biological replicates were made for each of the conditions studied. The protocol was based on the EpiCypher® CUTANA™ CUT&RUN Protocol v2.0, incorporating some modifications. Chromatin was amplified following the instructions of the NEBNext® Ultra™ II DNA Library Prep Kit (Illumina). CUT&RUN libraries were amplified, pooled, and sequenced on a Nextseq500 to generate 75 bp paired-end reads. Quality control was performed using fastqc. Reads were aligned to the human genome (GRCh38) using BWA, sorted and quality-filtered using samtools. Peaks were called using MACS2, and only peaks with p-value < 0.01 and 100% overlap across all biological replicates were considered for downstream analyses. R was used for peak annotation with ChIPseeker. Motif enrichment analyses were carried out using SEA and MEME. Heatmaps were generated using DeepTools. See more detailed information on CUT&RUN experiments and analysis in Supplementary Methods.
RNA-sequencing experiments and analysis
RNA was extracted using the RNeasy Plus kit (QIAGEN). RNA-seq libraries were sequenced on a Nextseq2000. Sequencing reads were pseudo-aligned to reference the human genome GRCh38.p13 version with Kallisto. Differential expression was conducted using DESeq2 package.
Ethics approval and consent to participate
All methods were performed in accordance with the relevant guidelines and regulations. The study was approved by the Institutional Review Board of the Hospital Clínic (HCB/2022/0082) and in accordance with the Declaration of Helsinki. Informed consent was obtained from all patients.
Results
The SOX11-specific interactome
To identify endogenous proteins that dynamically interact with SOX11 in MCL cells, we performed a large-scale proteomic strategy using the BioID2PL technology followed by quantitative MS [19, 20] (Fig. S1A). We generated a stable transduced cell line ectopically expressing an improved BirA biotin ligase (BioID2) [21] fused to the SOX11 protein under the control of a tetracycline-inducible promoter (TRE) in the SOX11-negative JVM2 MCL cell line (JVM2TRE-SOX11:BioID2). The optimal expression and nuclear localization of the SOX11:BioID2 protein and its correct functionality were confirmed at 24 hours of Doxycycline (Dox)-induction, showing upregulation of already known SOX11-direct target genes, such as CD24 [4], PLXNB1 [22], PDGFA [6] and MSI2 [9] in JVM2TRE-SOX11:BioID2 compared to JVM2TRE-BioID2 MCL cell lines (Fig. S1B, C and S1D, respectively). Increased levels of biotinylated proteins in Dox-induced compared to -non-induced cells were purified from nuclear extracts using streptavidin beads, digested, and subjected to quantitative LC-MS/MS (Fig. S1E and S1F). MS analyses identified 92 proteins (log2fold change (FC) > 1.7 and adjusted p-value (adj.p-value)< 0.05) specifically enriched in three biological replicas of JVM2TRE-SOX11:BioID2 compared to JVM2TRE-BioID2 Dox-induced cells (SOX11-interactome) (Table S3). STRING protein network and Gene and ontology (GO) analyses revealed a highly interconnected interactome, mostly enriched in proteins involved in nucleosome assembly, chromatin reorganization and transcription regulation (Fig. 1A, B, respectively). Interestingly, several components of the SWI/SNF chromatin remodeling complex were identified among the highest confident SOX11-protein partners in MCL (SMARCA4 (adj.p-value = 10−25), ARID1A (adj.p-value = 10−14, and SMARCC1 (adj.p-value = 10−8) (Fig. 1C).
A STRING network analysis of the MCL SOX11-interactome, based on candidates with a log2FC > 1.7 and average of >2 spectral counts in the SOX11:BioID2 condition (Table S3). STRING database was used to create Protein-protein interacting networks (interactome). Edges represent putative interactions by STRING. Color clusters show different protein complexes within the SOX11-interactome. The mediator complex is represented in light blue, the integrator complex in green, the SWI/SNF complex in pink, the Histone acetyltransferase complex in dark blue, and DNA repair members in yellow. The interaction network was created using Cytoscape software 3.10.1. B GO functional analysis of the top biological processes (BP) enriched within the SOX11-interactome in MCL. C Volcano plot shows in the x-axis the log2-fold change enrichment and in the y-axis the -log10 adjusted p-value for each SOX11-protein partner identified by BioID2-based PL/MS approach in JVM2TRE-SOX11:BioID2 compared to JVM2TRE-BioID2 cell lines. Grey dots represent non-significant proteins; Black dots represent significantly enriched SOX11-specific interacting proteins identified by BioID2-based PL/MS experiments in JVM2TRE-SOX11:BioID2 compared to JVM2TRE-BioID2 cell lines (SOX11-interactome). Pink dots represent protein members of the SWI/SNF complex enriched in the SOX11-specific interactome (Table S3).
SOX11 physically interacts with SMARCA4
The nuclear co-localization and physical interaction of SOX11 and SMARCA4 were visualized by immunofluorescence and in situ-fPLA experiments in SOX11-positive MCL cell lines (Z138 and Granta-519, and the SOX11-negative MCL cell line JVM2 ectopically overexpressing the SOX11 full length (FL) protein (JVM2TRE-SOX11FL:HA)), but not in JVM2 control cells (Fig. 2A, B). Increased SOX11:SMARCA4 interaction observed in Z138 (TP53WT) compared to JVM2TRE-SOX11FL:HA (TP53WT) or Granta519 (TP53MUT) is independent of TP53 status and could be associated with higher SOX11 expression in Z138 compared to other MCL cell lines (Fig. S2A, B and S2I). The physical interaction of SOX11 and SMARCA4 was also observed in primary cMCL but not in nnMCL peripheral blood samples, highlighting the high sensitivity of this technique and confirming the absence or weak expression of SOX11 in nnMCL cases (Fig. 2C). On the contrary, physical interactions between SOX11 and ARID1A or SMARCC1 proteins were not observed in either Z138 or Granta-519 cell lines by in situ-fPLA (Fig. S2C, D, respectively). Co-immunoprecipitation (co-IP) experiments also verified SOX11-SMARCA4 interaction but no SOX11-ARID1A or -SMARCC1 interactions. SMARCA4 protein was detected, but not ARID1A or SMARCC1 proteins, in SOX11 IPs (Fig. 2D, E). While SOX11, ARID1A and SMARCC1 were detected in SMARCA4 IPs (Fig. 2F, G). These results suggest that SMARCA4 is the only subunit of an active SWI/SNF complex that physically interacts with SOX11.
A In situ-fPLA showing SOX11 and SMARCA4 interaction (red spots) in SOX11-positive (Z138, Granta-519, and JVM2TRE-SOX11FL:HA), but not in the SOX11-negative (JVM2) MCL cell lines. SOX11-positive red fluorescent spots represent the interaction between SOX11 and SMARCA4. Cell nuclei were stained with DAPI (blue). B Quantification using the ImageJ software of in situ-fPLA red fluorescent spots shown in A. The signal was quantified as red fluorescent spots/cell nuclei (n = 300 nuclei). C In situ-fPLA showing the interaction of SOX11 and SMARCA4 (red spots) in SOX11-positive cMCL primary cells and not in SOX11-negative nnMCL case. Nuclear staining with DAPI (blue). D–G WB showing input and IP of SOX11, SMARCA4, ARID1A or SMARCC1 proteins co-immunoprecipitated upon specific D, E SOX11 pull-downs (IP SOX11) in JVM2TRE-SOX11:BioID2 (D) and Z138 (E) and (F, G) upon specific SMARCA4 pull-downs (IP SMARCA4) in (F) JVM2TRE-SOX11:BioID2 and (G) Z138. IgG antibody was used as a negative IP control (IP IgG). H Overlap of the histone core structures of the SOX11 and SWI/SNF complexes. In the model, SOX11 (red) is in a surface cavity next to the ATPase domain of SMARCA4 (green). Magnified view of the SOX11 binding to the SMARCA4/DNA surface cavity. SMARCA4 (green) and nucleosomal DNA (Brown) are shown as a surface representation. Figure prepared with the PyMOL Molecular Graphics System, Version 1.2r3pre (Schrödinger, LLC). I WB showing pulldown input and SMARCA4 pulldown upon immunoprecipitation (IP) with anti-HA antibody (α-HA) of HA-tagged SOX11 full-length (SOX11FL:HA) or SOX11 proteins with HMG or TA domains deleted (SOX11ΔHMG:HA or SOX11ΔTA:HA) expressed in JVM2 MCL cell line. All cell lines were induced with doxycycline (+). For each cell line, the first lane corresponds to its protein extract input (10% of total volume), and the second one the anti-HA IP (IP SOX11). JVM2TRE empty vector was used as a negative control. J In situ-fPLA of SMARCA4 and HA-tagged SOX11 full-length (SOX11FL:HA) or SOX11 proteins with HMG or TA domains deleted (SOX11ΔHMG:HA or SOX11ΔTA:HA) expressed in JVM2 MCL cell line, using an anti-SMARCA4 and an anti-HA antibodies. K Quantification using the ImageJ software of in situ-fPLA red fluorescent spots shown in (J). The signal was quantified as red fluorescent spots/cell nuclei (n = 500 nucleus). ∗p-value < .05; ∗∗p-value < .005, ∗∗∗p-value < .0005.
We generated a structural model, superimposing the histone cores of both SOX11- and SMARCA4-nucleosome complex three-dimensional structures [15, 23] (Fig. S2E, F, respectively), which showed that SOX11 binds to nucleosome-DNA near the location of the ATPase domain of SMARCA4 (Fig. 2H and Fig. S2G). In line, SMARCA4 protein levels in SOX11 co-IP and the number of in situ-fPLA spots significantly diminished in JVM2 cells expressing SOX11 proteins with the HMG or the transactivation domains (TA) deleted (SOX11ΔHMG:HA and SOX11ΔTA:HA, respectively), compared to full-length (SOX11FL:HA) expressing cells (Fig. 2I–K). Together, these results suggest that the physical interaction of SOX11 to SMARCA4 is through its HMG domain. However, the expression levels of SOX11ΔTA protein were very low, probably due to protein instability (Fig. S2H, I), which did not allow us to evaluate whether this domain is or is not involved in SOX11:SMARCA4 interaction.
SOX11:SMARCA4 genomic co-occupancy in MCL
SOX11- and SMARCA4-specific genome-wide binding sites were identified by CUT&RUN (C&R) experiments selecting SOX11 or SMARCA4 C&R-peaks in the SOX11/SMARCA4-positive MCL cell line Z138, that were not present in their respective SOX11-knockout (Z138SOX11KO) [9] or SMARCA4-negative Z138 cell line (Z138AU-15330). Z138AU-15330 cell line was obtained by treating Z138 MCL cell line with the proteolysis-targeting chimera (PROTAC) AU-15330, which efficiently degrades SMARCA4 [24] (Fig. 3A). Using three replicates, we identified 9319 SOX11-specific and 17574 SMARCA4-specific peaks, including 5751 and 9747 respectively annotated peaks (defined from -3000 bp upstream to +3000 bp downstream the transcription start sites (TSS)) in Z138 MCL cell lines (C&R-targets) (Fig. S3A, B and Table S4 and S5, respectively). Regulatory regions of already described SOX11-direct target genes, such as MSI [9], PDGFA [6], and BCL6 [5], were enriched in SOX11-peaks but not in IgG-peaks of control Z138 or in SOX11-peaks of Z138SOX11KO cell lines, demonstrating the specificity of SOX11-binding sites identified by our C&R experiments (Fig. S3C).
A WB showing SOX11 and SMARCA4 protein levels in Z138 control, Z138 MCL cell line treated with 0.5 μM of AU-15330 for 24 h (Z138AU-15330) and Z138SOX11-knockout (Z138SOX11KO). Actin was used as a loading control protein. B, C Heatmap profiles showing bound-genomic regions ( − 1 kb and +1 kb of the region’s center) of (B) the C&R read-intensity of SMARCA4 around SOX11-specific bound regions (C&R-peaks) and (C) C&R read-intensity of SOX11 around SMARCA4-specific bound regions (C&R-peaks) in Z138CT, Z138AU-15330 and Z138SOX11KO MCL cell lines (from left to right, respectively). D, E Heatmap showing tag density of H3K27ac, H3K4me3, H3K4me1, and DNase I cutting sites around (D) SOX11- or (E) SMARCA4-specific bound-genomic regions ( − 3 kb and +3 kb of the region’s center). F Stacked bar chart of the genomic distribution of SOX11-, SMARCA4-, and SOX11:SMARCA4 regions (C&R-peaks). For peak annotation, ChIPseeker was used. G Top enriched motifs were identified in the SOX11:SMARCA4-peaks using SEA (version 5.5.5) and MEME (version 5.5.5) in the Human TFs database (HOCOMOCO). H Dot chart of the more enriched transcription factor families. Each dot represents an individual transcription factor. The x-axis denotes the transcription factor families, and the y-axis indicates the significance level expressed as -log10(p-value).
Specific genomic regions bound by SMARCA4 were enriched in SOX11-peaks and vice versa (Fig. 3B, C, respectively). 58.84% of SOX11- and SMARCA4-specific C&R-peaks overlapped, sharing 8329 annotated peaks (SOX11:SMARCA4-peaks) (Table S6). SOX11 lost 56% of its binding sites in cells where SMARCA4 had been degraded. Interestingly, SMARCA4 lost 87.2% of its binding sites in SOX11KO cells (Fig. 3B, C and Fig. S3A, B). SOX11:SMARCA4-peaks (Table S6) were enriched in SOX11- and SMARCA4-specific peaks of other SOX11-positive MCL cell lines (Granta-519 and JVM2TRE-SOX11FL-HA), but not in SOX11-negative (JVM2TRE) or SMARCA4-negative (JVM2TRE-SOX11FL:HA treated with AU-15330) MCL cell lines (Fig. S3D–F, respectively). SOX11:SMARCA4-peaks were also enriched in SOX11- and SMARCA4-specific peaks of SOX11-positive cMCL cases, but not in peaks of SOX11-negative nnMCL cases, and diminished in SOX11-positive cases treated with AU-15330 (Fig. S4A, B). Our results confirm the genomic co-occupancy and dependency for binding of both proteins in several MCL cell lines and MCL cells from primary samples.
SOX11- and SMARCA4-specific peaks were enriched in the profiles of DNaseI hypersensitive sites (open chromatin) and active promoters (H3K27ac and H3K4me3), but not of active enhancers (H3K27ac and H3K4me1) [25] (Fig. 3D, E). Gene distribution, using HOMER annotation, also revealed that SOX11:SMARCA4-peaks were predominantly located in promoter regions (73.7%), at smaller numbers in distal intergenic regions (12.08%), introns (11.56%), and exons (2.65%) (Fig. 3F). DNA binding motif analysis showed that consensus motifs of the SP/KLF family, MYC-Associated Zing finger (MAZ), among other transcription factors, were strongly enriched within the SOX11:SMARCA4-peaks (Fig. 3G, H), suggesting they may be cofactors of the SOX11:SMARCA4 complex activity in MCL.
GO analysis revealed transcription regulation, apoptosis, cell division, and chromatin remodeling as the most significant biological processes (Fig. 4A), and VEGFA/VEGR2-, EGF/EGFR-, DNA repair, and the BCR-signaling as the most significantly enriched pathways within the SOX11:SMARCA4-peaks (in +/−3 Kb TSS) (Fig. 4B). The SOX11:SMARCA4 complex was also bound to regulatory regions of key proto-oncogenes (e.g., BLK, MAP3K14, MDM2, or CASTOR1) (Fig. 4C–F).
A, B GO functional annotation of the (A) biological processes (BP) and (B) pathways enriched in the SOX11:SMARCA4-peaks using DAVID software. C−F Genome view plot showing binding of SOX11 and SMARCA4 at regulatory regions of BLK and MAP3K14, MDM2, and CASTOR1 genes in Z138CT, Z138SOX11KO, and Z138AU-15330 cell lines. An IgG was used as an unspecific control of the technique. The graph represents one of the biological C&R triplicates. Antibodies and cell types for each track are shown on the left. All peaks obtained by C&R were called by MACS2 (p-value < 0.01).
Oncogenic transcriptional programs regulated by the SOX11:SMARCA4 complex in MCL
Z138 treated cell lines were treated with 0. 1 and 0.5 μM of AU-15330 for different hours (h). Treatments using 0.5 μM AU-15330 dose and time points that induce SMARCA4 degradation but not cell death (8 and 24 h) (Fig. S5A, B, respectively), were used to characterize the SMARCA4-specific GEP, comparing the RNA-seq transcriptomic profiles of Z138CT untreated (-) vs. Z138AU-15330 treated cells (adj.p-value < 0.05 and log2FC >|1 | ) (Fig. S5C−F and Table S7). Gene set enrichment analysis (GSEA) revealed that the BCR-signaling was one of the most significantly downregulated gene sets in Z138AU-15330 compared to the Z138CT cell line (Fig. S5G, H, respectively).
The SOX11-specific GEP was identified comparing RNA-seq transcriptomic profiles of SOX11-positive compared to SOX11-negative MCL cell lines (Z138CT vs. Z138SOX11KO, Z138SOX11Resc (Z138SOX11KO ectopically overexpressing Flag-SOX11) [9] vs. Z138SOX11KO (Fig. S6A) and JVM2TRE-SOX11FL:HA vs. JVM2TRE MCL cell lines (Fig. S2H)) (adj.p-value < 0.05 and log2FC >|1 | ), showing that SOX11 mostly activates transcription (Fig. S6B, C and S6D, E, and Tables S8 and S9, respectively). GSEA revealed that specific SOX11-GEPs were enriched in genes involved in the regulation of neural precursor cell proliferation, the Hippo signaling, and cell-cell signaling by WNT, among other pathways (Fig. S6F, G and S6H).
To identify the specific transcriptional program of the SOX11:SMARCA4 complex, we overlapped specific SMARCA4-GEPs (Table S7) and SOX11-GEPs (Table S8) with common SOX11:SMARCA4-peaks (Table S6) in Z138 MCL cell line, obtaining a total of 147 genes (SOX11:SMARCA4-direct targets) (Fig. 5A and Table S10). SOX11:SMARCA4-direct targets were significantly enriched in biological processes and pathways such as cell adhesion, apoptosis, cell migration, angiogenesis, B-cell differentiation and BCR-signaling, among others (Fig. 5B). SOX11:SMARCA4-direct targets were significantly upregulated in Z138CT compared to Z138SOX11KO or Z138AU-15330 cells, in JVM2TRESOX11FL:HA compared to JVM2TRE MCL cell lines, as well as in SOX11-positive cMCL compared to SOX11-negative nnMCL primary tumors (Fig. 5C).
A Overlap between SMARCA4-GEP (red), (Table S7; log2FC > |0.7 | & adjusted p-value < 0.05), SOX11-GEPs (green) (Table S8; log2FC > |0.5 | & adjusted p-value < 0.05), and SOX11:SMARCA4-Targets (blue) (Table S6). B GO of the Biological Processes (BP) (top) and pathway (bottom) enrichment analysis of the 147 SOX11:SMARCA4-direct target genes in MCL, found by the overlap in (A). The number of genes, fold enrichment, and –log10(p-value) for each BP and pathway are shown. C Dot plot showing at x-axis the NES, and at y-axis the 147 SOX11:SMARCA4-direct targets in Z138CT vs. Z138AU-15330, JVM2TRESOX11FL:HA vs. JVM2TRE (Dox-induced for 24 h and 120 h), Z138CT vs. Z138SOX11KO, Z138SOX11Resc vs. Z138SOX11KO MCL cell lines and cMCL vs nnMCL primary tumors. NES and FDR are shown, and statistical significance is assumed when FDR < 0.1. The size of the dots represents the number of enriched genes. D Genome view plot showing binding of SOX11 and SMARCA4 at regulatory regions of MSI2 and ROR1 genes in Z138, Z138SOX11KO, and Z138AU-15330 MCL cell line. The graph represents one of the biological C&R triplicates. Antibodies and cell types for each track are shown on the left. (E) Violin plots of MSI2 and ROR1 mRNA expression levels in Z138 control (CT) vs. AU-15330 treated cells during 8 h (AU8) or 24 h (AU24) and Z138SO11KO vs Z138CT and Z138SOX11Resc MCL cell lines, using the log2-transformed values. Datasets are indicated in the legend. Unpaired two-tailed Student’s t-test determined significances. p-values are shown in the graph. Statistical significance: ∗p-value < .05; ∗∗p-value < .005, ∗∗∗p-value < .0005, ∗∗∗∗p-value < .0001.
SOX11KO or SMARCA4-degradation led to the delocalization of both proteins from global genomic (Fig. 3B, C and Fig. S3A, B and S3D−F) and specific gene regulatory regions (e.g., MSI2 and ROR1) in MCL cell lines (Fig. 5D) and in primary tumors (Fig. S4A, B and S7A), subsequently impacting on the expression of their target genes (Fig. 5E and Fig. S7B, respectively), demonstrating that the SOX11:SMARCA4 complex is essential for directly activating gene transcription in MCL cell lines and primary tumors.
SMARCA4 is directly upregulated by SOX11 and associated with worse outcome of the patients
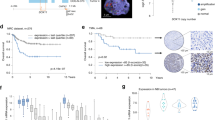
Using previously published gene expression and clinical data of three independent series (GSE79196 [17], EGAD00010001842 [18], and EGAD00001009422 [9]), we observed that SMARCA4 mRNA levels were significantly higher in SOX11-positive cMCL than in SOX11-negative nnMCL MCLs (Fig. 6A−C, respectively). Primary MCL of the blastoid variant, which has been associated with worse clinical outcome, displayed significantly higher SMARCA4 mRNA levels than classical and small cell variants (Fig. 6D).
A−C Violin plots showing SMARCA4 mRNA expression levels, calculated by the average of all microarray probes of SMARCA4, in (A) 30 SOX11-positive and 24 SOX11-negative MCL primary tumors (GSE79196) and (B) in 26 SOX11-positive cMCL and 13 SOX11-negative nnMCL primary tumors (EGAD00010001842), and (C) in 10 SOX11-positive and 6 SOX11-negative MCL primary tumors using the (log2-transformed values) of RNA-seq data (EGAD000010094222). Unpaired two-tailed Student’s t-test determined significances. p-values are shown in the graph. D Violin plot showing SMARCA4 mRNA levels in MCL primary tumors with small cells, classical or blastoid cytologic variant (EGAD00010001842). E–G Correlation between SOX11 and SMARCA4 mRNA expression levels in (E) GSE79196, (F) EGAD00010001842, and (G) EGAD000010094222. H, I Correlation between SOX11 and SMARCA4 mRNA expression levels considering only SOX11-positive primary tumors in (H) GSE79196 and (I) EGAD00010001842. The graph shows the Pearson Correlation coefficient (r), p-value, and the number of primary tumors analyzed (n). J, K Kaplan Meier curve and Cox regression analysis showing the association of SMARCA4 mRNA expression and OS, using previously published GEP and clinical data from (J) 40 SOX11-positive and SOX11-negative MCL primary samples (GSE79196) and (K) considering only SOX11-positive MCL primary tumors of the GSE79196 cohort. The maximally selected rank statistics (Maxstat) defined SMARCA4 high and low values (cut-off = 6.82 expression units). The hazard ratio (HR) with a 95% confidence interval (CI) and the Cox regression p-value are shown.
SMARCA4 expression positively correlated with SOX11 mRNA (Fig. 6E–G), also considering only SOX11-positive primary tumors (Fig. 6H, I). A significant correlation between SMARCA4 protein and mRNA levels (Pearson r = 0.88, p-value = 0.046, n = 3) and a significant positive correlation between SMARCA4 and SOX11 protein levels (Pearson r = 0.84, p-value = 0.037, n = 3) were observed in MCL cell lines (Fig. S8A and S8B, C, respectively). SMARCA4 mRNA and protein levels were reduced upon SOX11KO in Z138, and its levels were recovered upon SOX11 ectopic overexpression in Z138SOX11Resc, reaching similar levels as in the control Z138 MCL cell line and significantly increased in JVM2TRE-SOX11FL:HA compared to JVM2TRE cells (Fig. S8D, E and S2H). Together, our results support the direct involvement of SOX11 in the transcriptional activation of SMARCA4.
We observed that high SMARCA4 mRNA levels were significantly associated with shorter overall survival (OS) of patients (Fig. 6J), also when considering only SOX11-positive MCL tumors (Fig. 6K). Bivariate COX regression analyses showed that SMARCA4 had an overall survival prognostic value even when adjusting for SOX11 expression and other parameters of poor prognosis, such as high copy number alterations ( ≥ 6 CNA) and TP53 (17p13.1) alterations (Table S11), suggesting that SMARCA4 overexpression may be involved in the progression of this disease.
AU-15330 overcomes Ibrutinib resistance in MCL
SMARCA4 degradation led to the downregulation of mRNA and protein levels of several BCR-related genes in MCL cell lines (Fig. 7A, B and Fig. S9A and S9C, D). AU-15330 treatments also significantly reduced the phosphorylation of key BCR-signaling components (SYK and BTK) in Z138 (Fig. 7B and Fig. S9B) and in MINO MCL cell line (Fig. S9C–E), suggesting that it is regulating the BCR-signaling in MCL. To validate the decrease of the BCR activity upon AU-15330 treatments, we measured calcium (Ca2+) flux upon BCR stimulation with anti-IgM in different MCL cell lines. As expected, the ibrutinib sensitive (IS), MINO, but not Ibrutinib resistant (IR) MCL cell lines (Z138 or JVM2) showed Ca2+ flux reduction after Ibrutinib exposures, while upon AU-15330 exposure the Ca2+ flux reduction was observed in both, MINO (IS) and Z138 (IR) SOX11-positive, but not in SOX11-negative JVM2 (IR) MCL cell line (Fig. S9F). These results suggest that AU-15330 may overcome Ibrutinib resistance induced by the SOX11:SMARCA4 complex in MCL.
A Heatmap illustrating the scaled expression (Z-score) of BCR signaling pathway-related genes in RNA-seq GEPs of Z138CT and Z138AU-15330 MCL cell lines after 8 h and 24 h AU-15330 treatments (CT8 and CT24 vs AU8 and AU24, respectively) (n = 3). The relative value for each gene is depicted by color intensity, with red indicating upregulated and blue indicating downregulated genes. B WB experiments showing the protein levels of SMARCA4, SOX11, and total and phosphorylated (p) forms of BCR-related proteins (SYK and BTK) in Z138CT and Z138SOX11KO treated with DMSO (-) or AU-15330 (+) for 24 h, starved for 90 min, with (+) or without IgM (-) stimulation for 15 min. Actin protein was used as a loading control. C GSEA analysis of the gene set NIK-signature, comparing Z138CT vs. Z138AU-15330 cells after 8 h of treatments (Z138AU (8 h)). NES and FDR are shown, and statistical significance is assumed when FDR < 0.1. D Heatmap illustrating the scaled expression (Z-score) of NIK-signature in RNA-seq GEPs of Z138CT vs Z138AU-15330 MCL cell lines after 8 h and 24 h of treatment (CT8 and CT24 vs AU8 and AU24) (n = 3). The relative value for each gene is depicted by color intensity, with red indicating upregulated and blue indicating downregulated genes. E Bar plots showing cell proliferation (%) and (F) cell apoptosis (% of apoptotic cells; Annexin-V+ and Propidium-Iodide+ cells) of SOX11-positive/SMARCA4-high IR (Z138) and IS (MINO), and SOX11-negative/SMARCA4-low IR (JVM2) MCL cell lines, treated with AU-15330 (0.5 µM), Ibrutinib (1 µM), or the combination of AU-15330 plus Ibrutinib for 72 h, relative to untreated control cells (n = 3). G Bar plots showing BCL2 mRNA expression levels in SOX11-positive (Z138, MINO, JVM2TRESOX11FL:HA) and SOX11-negative (JVM2TRE) MCL cell lines, treated with DMSO or with AU-15330 (0.5 µM) for 24 h (n = 3), levels are relative to each control (DMSO treated) cells. Unpaired two-tails Student’s t-test determined significance. H WB experiments showing SMARCA4 and BCL2 protein levels in Z138, MINO, and JVM2 MCL cell lines treated with DMSO (-) or with 0.5 µM AU-15330 for 24 and 72 h. Tubulin protein was used as a loading control. I RT-qPCR analysis of SMARCA4, BLK, CD79A, and SYK mRNA levels in Z138SMARCA4KO or Z138AU-15330 cell lines relative to their non-targeting Z138 control cells and +DMSO, respectively. J, K Bar plots showing cell proliferation (%) (J) and cell apoptosis (% of apoptotic cells; Annexin-V+ and Propidium-Iodide+ cells) (K) of Z138SMARCA4KO or Z138AU-15330 cells relative to their non-targeting control cells and +DMSO, respectively. Data are shown as mean ± SEM. Statistical significance: ∗p-value < 0.05; ∗∗p-value < 0.005, ∗∗∗p-value < 0.0005.
Ibrutinib resistance in Z138 is caused by the constitutive activation of the alternative NF-κB-signaling [26], an essential proliferation pathway in MCL. Remarkably, we observed a significant decrease in the NF-κB-inducing kinase (NIK) signature in Z138AU-15330 compared to Z138CT cells (Fig. 7C, D). In line, Ibrutinib exposures decreased cell proliferation only in MINO cell line, while AU-15330 significantly reduced cell proliferation and increased apoptosis of both (IS) MINO and (IR) Z138 SOX11-positive, but not of the (IR) SOX11-negative JVM2 cell line (Fig. 7E, F).
NIK transcriptionally activates the BCL2 pathway, facilitating resistance to apoptosis during tumor development [27]. Accordingly, we observed significantly lower BCL2 mRNA and protein levels in AU-15330 treated compared to non-treated cells of only SOX11-positive cell lines, but not in the SOX11-negative JVM2TRE MCL cell line (Fig. 7G, H). BCL2 mRNA levels decreased upon SOX11KO in Z138 or increased in JVM2TRE-SOX11FL:HA compared to JVM2TRE cells, although those changes were not statistically significant (Fig. S9G). Together, our results suggest that BCL2 is regulated by SMARCA4 only in SOX11-positive MCL cells, probably through the direct binding to MAP3K14 (encoding for NIK protein) regulatory regions (Fig. 4D) and transcriptional activation by the SOX11:SMARCA4 complex.
AU-15330 is a dual degrader of SMARCA4 and SMARCA2 [24]. Compared to SMARCA4, SMARCA2 is weakly expressed in MCL cell lines and primary samples (Fig. S10A, B, respectively). However, to fully confirm that phenotypic changes observed after AU-15330 treatments were through SMARCA4 degradation, we knocked out SMARCA4 in Z138 cells using the CRISPR-Cas9 genome editing. A pool of KO cells (SMARCA4KO), showing about 50% SMARCA4 protein reduction, but not SMARCA2 protein levels (Fig. S10C), displayed a significant decrease in the expression of several BCR-related genes and cell proliferation (Fig. 7I, J). Additionally, apoptosis significantly increased compared to control cells, reaching similar phenotypes to the ones observed after AU-15330 exposures (Fig. 7K). Together, our results highlight AU-15330, through specific degradation of SMARCA4, as promising target therapy to overcome Ibrutinib resistance in MCL.
Discussion
Here, using a large-scale and highly sensitive approach, capable of detecting also weak and/or transient protein interactions with our bait protein (SOX11:BioID2), we have identified the specific interactome map of SOX11. Several subunits of the SWI/SNF chromatin remodeling complex were identified as the most confident SOX11-binding proteins.
SWI/SNF complexes use the energy of ATP hydrolysis to mobilize nucleosomes and modulate chromatin architecture [28, 29]. They play a crucial role in transcriptional regulation, mainly by facilitating transcription through remodeling of nucleosomes to provide access to the transcription machinery and transcription factors [30]. Three major subfamilies of SWI/SNF complexes have been described in mammals: The canonical BAF (cBAF), the polybromo-associated BAF (PBAF), and the noncanonical BAF (ncBAF or GBAF). BAF complexes are composed of 8-12 subunits, containing common subunits (e.g., one or two mutually exclusive catalytic ATPase subunits, SMARCA4 and/or SMARCA2) [30] and complex-specific subunits [31]. We have identified SMARCA4 and SMARCC1 (common subunits of all three BAF complexes) and specific subunits of the cBAF complex (ARID1A and ARID1B) within the SOX11-interactome. However, no specific subunits of the gBAF or pBAF complexes were identified, suggesting that SMARCA4 interacts with SOX11 as part of an active cBAF complex, which mostly binds enhancer regions [32].
Detection of SOX11 signals varies depending on the technique and antibody used. BioID2-PL/MS is the most sensitive detecting interactions in live cells, including weak and transient ones. In situ fPLA technique needs cell fixation that could lose detecting some transient interactions. Co-IP experiments use several mechanical manipulations and stringent buffers that could break weak protein-protein interactions; However, if it is detected it would mean that there is a strong interaction between the proteins analyzed. BioID2-PL identified SOX11 interactions with cBAF complex proteins (SMARCA4, ARID1A, SMARCC1), but only SMARCA4 interaction was confirmed by in situ fPLA and co-IP/WB experiments, suggesting a strong interaction between SOX11 and SMARCA4, but weak or transitory with ARID1A and SMARCC1, in MCL cell lines and primary tumors. Together, our results suggest that SMARCA4 is physically interacting with SOX11, ARID1A and SMARCC1 to support SOX11-specific transcriptional program. However, we cannot discard that SMARCA4 may also interact with ARID1A and SMARCC1, as part of the cBAF complex, supporting other physiologic activities of MCL cells. Our results also support that SOX11 may interact with other protein complexes to exert its multifunctional activity in MCL cells.
Our three structural model and the loss of interaction between SMARCA4 and the SOX11ΔHMG protein suggest that SOX11 and SMARCA4 might be physically interacting through their respective HMG and ATPase domains within the nucleosome. However, further experiments need to be performed to confirm whether SOX11 TA and the SMARCA4 ATPase domain are indeed involved.
We have performed the first comprehensive analysis of genome-wide SOX11-specific binding sites in MCL. SOX11 and SMARCA4 share more than 50% of their specific C&R-peaks in preferentially open chromatin and active promoter regions. SOX11 lost 56% of its binding sites in MCL cell lines and primary tumors treated with AU15330, demonstrating co-occupancy and its dependence on SMARCA4 as a cofactor to mediate its oncogenic transcriptional activity. Intriguingly, SMARCA4 lost 87.2% of its binding sites in SOX11KO cells, suggesting that SMARCA4 needs to interact with SOX11 to bind specific genomic regions in MCL. Together, these results suggest that SOX11 binds the SMARCA4 ATPase catalytic domain of the active cBAF complex, enabling chromatin accessibility to recruit the SOX11:SMARCA4 complex at active promoter regions and modulate transcription in MCL cells. 44% of SOX11-binding sites are not sheared with SMARCA4, suggesting that SOX11 also binds to DNA together with other proteins as cofactors, which is consistent with multiple roles of SOX11 in MCL pathogenesis. ELF1 and FOXK1 proteins were identified within the 92 proteins of the SOX11-interactome, and their specific binding motifs were enriched within the SOX11:SMARCA4-peaks, suggesting that those proteins could be participating in the SOX11:SMARCA4 complex in MCL.
The role of SMARCA4 in cancer remains controversial. Several studies suggest that SMARCA4 acts as either a tumor suppressor or oncogene in a cell-type and cancer context-specific manner. Recently, several evidence highlighted the capacity of SMARCA4 to promote tumorigenesis, often through overexpression, deregulating key oncogenic pathways for the development of breast, gastric, and prostate cancers [32,33,34,35]. Like in adrenergic neuroblastoma [36], we have observed that SMARCA4 upregulation is directly regulated by SOX11 in MCL. High levels of SMARCA4 were significantly associated with higher expression in SOX11-positive cMCL and blastoid variants and lower OS of patients, suggesting its involvement in the aggressive behavior of MCL. Our results suggest that SMARCA4 expression is essential for normal B cells; however, its upregulation, induced by SOX11, contributes to the aggressive MCL phenotype by promoting SOX11-mediated oncogenic transcriptional programs. Further studies, analyzing SMARCA4 expression in in situ mantle cell lesions, are needed to explore whether SMARCA4 activity could be also an early event in MCL pathogenesis.
A recent study has identified an activating SMARCA4 missense mutation that shows enhanced ATPases and chromatin remodeling activities of the SWF/SNF complex, leading to tumor progression in colorectal cancers [37]. SMARCA4 mutations identified in MCL are mostly missense mutations [18, 38,39,40,41]. We have recently identified an association between SMARCA4 mutations and SOX11 positivity in MCL and BL [18, 22], and most SMARCA4 missense mutations are located at the ATPase domain (aa1009-1314), where the physical interaction between SMARCA4 and SOX11 occurs. Previous works have reported that the acquisition of SMARCA4 mutations may contribute to disease progression in patients with ibrutinib-venetoclax-resistant MCL [39, 42]. Here, we have demonstrated that the SOX11:SMARCA4 interaction contributes to the oncogenic transcriptional program of SOX11, directly activating genes crucial for MCL progression, such as BCR-related genes. Together, our findings suggest that missense mutations at the SMARCA4 ATPase domain may be critical activating mutation that may increase the binding capability of the SOX11:SMARCA4 complex to specific regulatory regions, leading to a higher transcriptional activity of common direct targets, such as BCR-related genes, to accelerate the progression of MCL and its resistance to Ibrutinib. However, further investigations are required to demonstrate this hypothesis in MCL, and if so, what are possible molecular mechanisms.
AU-15330 induces specific degradation of SMARCA4, resulting in potent inhibition of tumor cell proliferation in prostate and other cancers [24]. In MCL, we have observed that AU-15330 significantly reduced SOX11 binding to specific genomic regions, including binding sites of genes related to the BCR signaling. Z138AU-15330 cells showed decreased gene expression and signaling activation of the BCR pathway compared to Z138CT cells. Moreover, AU-15330 treatments reduced calcium flux, NIK-signaling and BCL2 expression, cell proliferation, and induced apoptosis of both Ibrutinib-sensible and -resistant SOX11/SMARCA4-expressing cells but not in the JVM2 SOX11-negative/SMARCA4low MCL cell line. Similar effects were observed in SMARCA4KO cells, suggesting that AU-15330 treatments, by the specific degradation of SMARCA4 in MCL cells, could represent a promising therapeutic strategy for patients who relapse from Ibrutinib, Venetoclax, and other target therapies.
Overall, we have identified the specific SOX11-interactome map and demonstrated the physical interaction of SOX11 with SMARCA4 in MCL cells. SMARCA4 upregulation is directly regulated by SOX11 and associated with worse outcome of patients with MCL. SOX11 and SMARCA4 bind common regulatory regions of active promoters to directly regulate the expression of genes related to pathways crucial for MCL pathogenesis. Downregulation of SMARCA4, by AU-15330 treatments or SMARCA4KO, significantly reduces SOX11 binding to specific regulatory regions and diminished the activation of BCR-, NIK-, and BCL2-signaling pathways, as well as significantly reduces proliferation and induced apoptosis of Ibrutinib-sensible and -resistant SOX11-positive MCL cells. Further experiments will be required to understand how missense mutations at the SMARCA4 ATPase domain may alter the interaction between SOX11 and SMARCA4 and the binding capability of the SOX11:SMARCA4 complex to specific regulatory regions in MCL cells. Disruption of the SOX11:SMARCA4 complex could represent an innovative therapeutic strategy to overcome resistance and relapse to current target therapies, such as Ibrutinib, frequently seen in patients with MCL.
Data availability
The BioID-PL/MS data generated in this study have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD051143. CUT&RUN and RNA-seq data generated in this study have been deposited in the European Genome-phenome Archive and will be available at the time of acceptance of this manuscript.
References
Swerdlow SH, Campo E, Harris NL. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. Revised 4th Edition. (2017).
Fernàndez V, Salamero O, Espinet B, Solé F, Royo C, Navarro A. et al. Genomic and Gene Expression Profiling Defines Indolent Forms of Mantle Cell Lymphoma. Cancer Res. 2010;70:1408–18.
Royo C, Navarro A, Clot G, Salaverria I, Giné E, Jares P, et al. Non-nodal type of mantle cell lymphoma is a specific biological and clinical subgroup of the disease. Leukemia. 2012;26:1895–8.
Vegliante MC, Palomero J, Pérez-Galán P, Roué G, Castellano G, Navarro A, et al. SOX11 regulates PAX5 expression and blocks terminal B-cell differentiation in aggressive mantle cell lymphoma. Blood. 2013;121:2175–85.
Palomero J, Vegliante MC, Eguileor A, Rodríguez ML, Balsas P, Martínez D, et al. SOX11 defines two different subtypes of mantle cell lymphoma through transcriptional regulation of BCL6. Leukemia. 2016;30:1596–9.
Palomero J, Vegliante MC, Rodríguez ML, Eguileor Á, Castellano G, Planas-Rigol E, et al. SOX11 promotes tumor angiogenesis through transcriptional regulation of PDGFA in mantle cell lymphoma. Blood. 2014;124:2235–47.
Balsas P, Palomero J, Eguileor Á, Rodríguez ML, Vegliante MC, Planas-Rigol E, et al. SOX11 promotes tumor protective microenvironment interactions through CXCR4 and FAK regulation in mantle cell lymphoma. Blood. 2017;130:501–13.
Balsas P, Veloza L, Clot G, Sureda-Gómez M, Rodríguez M-L, Masaoutis C, et al. SOX11, CD70, and Treg cells configure the tumor immune microenvironment of aggressive mantle cell lymphoma. Blood. 2021;138:2202–15.
Sureda-Gómez M, Balsas P, Rodríguez M-L, Nadeu F, De Bolòs A, Eguileor Á, et al. Tumorigenic role of Musashi-2 in aggressive mantle cell lymphoma. Leukemia. 2023;37:408–21.
Kuo P-Y, Jatiani SS, Rahman AH, Edwards D, Jiang Z, Ahr K, et al. SOX11 augments BCR signaling to drive MCL-like tumor development. Blood. 2018;131:2247–55.
Hou L, Srivastava Y, Jauch R. Molecular basis for the genome engagement by Sox proteins. Semin Cell Dev Biol. 2017;63:2–12.
Grosschedl R, Giese K, Pagel J. HMG domain proteins: architectural elements in the assembly of nucleoprotein structures. Trends Genet. 1994;10:94–100.
Wilson M, Koopman P. Matching SOX: partner proteins and co-factors of the SOX family of transcriptional regulators. Curr Opin Genet Dev. 2002;12:441–6.
Wissmuller S. The high-mobility-group domain of Sox proteins interacts with DNA-binding domains of many transcription factors. Nucleic Acids Res. 2006;34:1735–44.
Dodonova SO, Zhu F, Dienemann C, Taipale J, Cramer P. Nucleosome-bound SOX2 and SOX11 structures elucidate pioneer factor function. Nature. 2020;580:669–72.
Beà S, Valdés-Mas R, Navarro A, Salaverria I, Martín-Garcia D, Jares P, et al. Landscape of somatic mutations and clonal evolution in mantle cell lymphoma. Proc Natl Acad Sci. 2013;110:18250–5.
Navarro A, Clot G, Martínez-Trillos A, Pinyol M, Jares P, González-Farré B, et al. Improved classification of leukemic B-cell lymphoproliferative disorders using a transcriptional and genetic classifier. Haematologica. 2017;102:e360–e363.
Nadeu F, Martin-Garcia D, Clot G, Díaz-Navarro A, Duran-Ferrer M, Navarro A, et al. Genomic and epigenomic insights into the origin, pathogenesis, and clinical behavior of mantle cell lymphoma subtypes. Blood. 2020;136:1419–32.
Roux KJ, Kim DI, Raida M, Burke B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J Cell Biol. 2012;196:801–10.
Varnaitė R, MacNeill SA. Meet the neighbors: Mapping local protein interactomes by proximity‐dependent labeling with BioID. Proteomics. 2016;16:2503–18.
Kim DI, Jensen SC, Noble KA, KC B, Roux KH, Motamedchaboki K, et al. An improved smaller biotin ligase for BioID proximity labeling. Mol Biol Cell. 2016;27:1188–96.
Sureda-Gómez M, Iaccarino I, De Bolòs A, Meyer M, Balsas P, Richter J, et al. SOX11 expression is restricted to EBV-negative Burkitt lymphoma and is associated with molecular genetic features. Blood. 2024;144:187–200.
He S, Wu Z, Tian Y, Yu Z, Yu J, Wang X, et al. Structure of nucleosome-bound human BAF complex. Science. 2020;367:875–81.
Xiao L, Parolia A, Qiao Y, Bawa P, Eyunni S, Mannan R, et al. Targeting SWI/SNF ATPases in enhancer-addicted prostate cancer. Nature. 2022;601:434–9.
Queirós AC, Beekman R, Vilarrasa-Blasi R, Duran-Ferrer M, Clot G, Merkel A, et al. Decoding the DNA Methylome of Mantle Cell Lymphoma in the Light of the Entire B Cell Lineage. Cancer Cell. 2016;30:806–21.
Saba NS, Liu D, Herman SEM, Underbayev C, Tian X, Behrend D, et al. Pathogenic role of B-cell receptor signaling and canonical NF-κB activation in mantle cell lymphoma. Blood. 2016;128:82–92.
You Z, Ouyang H, Lopatin D, Polver PJ, Wang C-Y. Nuclear Factor-κB-inducible Death Effector Domain-containing Protein Suppresses Tumor Necrosis Factor-mediated Apoptosis by Inhibiting Caspase-8 Activity. J Biol Chem. 2001;276:26398–404.
Imbalzano AN, Kwon H, Green MR, Kingston RE. Facilitated binding of TATA-binding protein to nucleosomal DNA. Nature. 1994;370:481–5.
Kwon H, Imbalzano AN, Khavari PA, Kingston RE, Green MR. Nucleosome disruption and enhancement of activator binding by a human SW1/SNF complex. Nature. 1994;370:477–81.
Centore RC, Sandoval GJ, Soares LMM, Kadoch C, Chan HM. Mammalian SWI/SNF Chromatin Remodeling Complexes: Emerging Mechanisms and Therapeutic Strategies. Trends Genet. 2020;36:936–50.
Malone HA, Roberts CWM. Chromatin remodellers as therapeutic targets. Nat Rev Drug Discov. 2024;23:661–81.
Shi J, Whyte WA, Zepeda-Mendoza CJ, Milazzo JP, Shen C, Roe J-S, et al. Role of SWI/SNF in acute leukemia maintenance and enhancer-mediated Myc regulation. Genes Dev. 2013;27:2648–62.
Buscarlet M, Krasteva V, Ho L, Simon C, Hébert J, Wilhelm B, et al. Essential role of BRG, the ATPase subunit of BAF chromatin remodeling complexes, in leukemia maintenance. Blood. 2014;123:1720–8.
Jubierre L, Soriano A, Planells-Ferrer L, París-Coderch L, Tenbaum SP, Romero OA, et al. BRG1/SMARCA4 is essential for neuroblastoma cell viability through modulation of cell death and survival pathways. Oncogene. 2016;35:5179–90.
Guerrero-Martínez JA, Reyes JC. High expression of SMARCA4 or SMARCA2 is frequently associated with an opposite prognosis in cancer. Sci Rep. 2018;8:2043.
Decaesteker B, Louwagie A, Loontiens S, De Vloed F, Bekaert S-L, Roels J, et al. SOX11 regulates SWI/SNF complex components as member of the adrenergic neuroblastoma core regulatory circuitry. Nat Commun. 2023;14:1267.
Zeng X, Yao B, Liu J, Gong GW, Liu M, Li J, et al. The SMARCA4R1157W mutation facilitates chromatin remodeling and confers PRMT1/SMARCA4 inhibitors sensitivity in colorectal cancer. NPJ Precis Oncol. 2023;7:28.
Jain P, Wang M. High-risk MCL: recognition and treatment. Blood. 2025;145:683–95.
Zhao S, Kanagal-Shamanna R, Navsaria L, Ok CY, Zhang S, Nomie K, et al. Efficacy of venetoclax in high risk relapsed mantle cell lymphoma (MCL) - outcomes and mutation profile from venetoclax resistant MCL patients. Am J Hematol. 2020;95:623–9.
Jain P, Zhang S, Kanagal-Shamanna R, Ok CY, Nomie K, Gonzalez GN, et al. Genomic profiles and clinical outcomes of de novo blastoid/pleomorphic MCL are distinct from those of transformed MCL. Blood Adv. 2020;4:1038–50.
Jain P, Wang M. Mantle cell lymphoma: 2019 update on the diagnosis, pathogenesis, prognostication, and management. Am J Hematol. 2019;94:710–25.
Agarwal R, Chan YC, Tam CS, Hunter T, Vassiliadis D, Teh CE, et al. Dynamic molecular monitoring reveals that SWI–SNF mutations mediate resistance to ibrutinib plus venetoclax in mantle cell lymphoma. Nat Med. 2019;25:119–29.
Acknowledgements
This work was supported by the Ministerio de Ciencia e Innovación (Grant No. PID2021-124048OB-100 to VA and PID2021-123054OB-I00 to EC), co-financed by the Sub-Directorate General for Evaluation and the European Regional Development Fund (FEDER-“Una manera de hacer Europa”); Generalitat de Catalunya Suport Grups de Recerca (AGAUR: 2021-SGR-01293 to VA and 2021-SGR-01172 to EC); Fundació La Marató de TV3 (201901-30 to VA and JV) and CERCA Programme / Generalitat de Catalunya. ADB is supported by the Agencia de Gestió d’ajuts universitaris i de recerca (AGAUR-PERIS Salut; SLT017/20/000161). MCC is supported by the Agencia de Gestió d’ajuts universitaris i de recerca (AGAUR; investigador novell-FI2024), financed by the European Union, Next Generation EU. MSG was supported by the Agencia de Gestió d’ajuts universitaris i de recerca (AGAUR; investigador novell-FI2018; FI-1 00266). We thank Dr. Francesc Canals for his technical assistance in the analysis of samples by mass spectrometry. This work was developed at the Centro Esther Koplowitz (CEK), Barcelona, Spain. We are grateful to S. Ruiz for her logistics assistance.
Author information
Authors and Affiliations
Contributions
ADB performed all in vitro experiments, designed experiments, and analyzed data; MCC performed cell proliferation experiments, MSG, GC and CL analyzed RNA-seq data, MRL performed in vitro experiments; AM contributed to the execution of C&R experiments and SC, RA and JIMS to the interpretation of the C&R-seq data; whereas EGOGenomics company performed the C&R-seq computational analyses; DC and HP contributed to the execution of in vitro experiments, EC identified MCL tumors; DR performed structural remodeling, JV analyzed MS data and VA supervised and designed experiments, analyzed data and wrote the manuscript; and all authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
De Bolòs, A., Carreras-Caballé, M., Sureda-Gómez, M. et al. The SOX11:SMARCA4 complex is a driver of oncogenic transcriptional programs in mantle cell lymphoma. Blood Cancer J. 15, 127 (2025). https://doi.org/10.1038/s41408-025-01333-6
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41408-025-01333-6