Abstract
Background
Metastatic castration-resistant prostate cancer (mCRPC) remains a lethal disease with few biomarkers to inform treatment selection or patient prognosis. Methylation profiles of plasma circulating tumour DNA (ctDNA) accurately reflect tumour methylomes and may reveal novel biomarkers of mCRPC.
Methods
To establish a novel mCRPC-associated methylation signature for detection of ctDNA, we performed plasma methylome profiling on 27 mCRPC patients and 10 controls (cohort 1). Signature-based ctDNA detection was evaluated across prostate cancer (PC) disease stages using an internal cohort of 93 PC patients and 8 controls (cohort 2), and an external cohort of 115 PC patients (cohort 3).
Results
We established a 48-region methylation signature (cfMeCaP) capable of highly sensitive detection of ctDNA in mCRPC (100%, 84% and 95% in cohorts 1, 2 and 3, respectively). cfMeCaP methylation at mCRPC diagnosis was associated with poor progression-free survival (PFS) and overall survival in all three cohorts (p < 0.005), independent of routine clinical variables. Persistent serial detection of ctDNA using cfMeCaP was strongly associated with rapid mCRPC treatment failure (median PFS 4.4 vs. 65.5 months; p < 0.0001), while no detection predicted continued treatment response.
Conclusion
These results highlight cfMeCaP as a promising non-invasive biomarker for prognostication in mCRPC.
Similar content being viewed by others
Background
Metastatic castration resistant prostate cancer (mCRPC) presents a significant healthcare challenge worldwide due to high mortality rates [1, 2]. Standard treatment options, such as taxanes and androgen receptor signalling inhibitors (ARSI), provide only limited survival benefits owing to high levels of primary and acquired resistance [3, 4]. Currently, no method can accurately predict treatment resistance in mCRPC and routine tools to monitor therapy efficacy, i.e. prostate specific antigen (PSA) and imaging, remain suboptimal [3, 5]. Moreover, tumour tissue-based molecular analyses are challenging in this setting, as metastases are not routinely sampled. Hence, there is an urgent need for novel, minimally invasive biomarkers to guide personalised treatment. Plasma cell-free DNA (cfDNA) offers a promising strategy for biomarker discovery in mCRPC patients as a fraction of cfDNA may be tumour cell-derived (circulating tumour DNA; ctDNA) and has been shown to reflect the molecular alterations of matched tumour tissue [6].
Here, we employed a highly sensitive methodology, cell-free methylated DNA immunoprecipitation and high-throughput sequencing (cfMeDIP-seq) [7, 8], to characterise the methylome landscape of mCRPC using plasma cfDNA. By cfMeDIP-seq methylome profiling of plasma cfDNA samples from a cohort of 27 patients with mCRPC and 10 men without PC, we aimed to develop a mCRPC-associated cfDNA methylation signature and subsequently evaluate the signature in large independent cohorts, comprising more than 700 samples from 388 prostate cancer (PC) patients and 305 male controls. We established a novel 48-region mCRPC-associated cfDNA methylation signature (cfMeCaP) with high sensitivity and specificity for detection of methylated ctDNA (me-ctDNA) in plasma. We found high cfMeCaP methylation levels at mCRPC diagnosis to be associated with poor PSA progression-free survival (PSA-PFS) and poor overall survival (OS). Furthermore, during first-line ARSI treatment, longitudinal me-ctDNA detection analyses based on cfMeCaP could identify patients with a high risk of early treatment failure.
Materials and methods
Cohorts for cfMeDIP-seq analysis
Cohort 1 comprised of 27 patients with mCRPC (mCRPC) and 10 men without PC (non-PC control; NPCC), recruited at the Department of Urology, Aarhus University Hospital (AUH), Denmark (2016-2020).
Cohort 2 comprised of 43 patients with mCRPC (mCRPC), 40 with hormone sensitive PC (HSPC), 10 with localised PC (LPC) and 8 men without PC (NPCC), recruited at the Department of Urology, AUH (n = 79; 2006–2022) or at the Department of Oncology, Vejle Hospital, Denmark (n = 22; 2016–2020).
In both cohorts, metastases were verified by CT scan, bone scintigraphy or 18F-sodium fluoride PET (F-18Na/PET). Blood samples from all mCRPC patients (cohort 1 and cohort 2) were drawn immediately before the initiation of first-line mCRPC treatment (enzalutamide or abiraterone). LPC blood samples were drawn at the time of radical prostatectomy (RP). HSPC blood samples were collected immediately before initiating androgen deprivation therapy (ADT; n = 37) or during ADT (n = 3). All NPCC men had low serum PSA levels (<4 ng/ml), and prostate biopsies confirmed to be negative for PC. Clinical characteristics for both cohorts are shown in Table 1.
Cohort 3 (external validation) comprised 30 patients with localised PC (LPC) and 85 patients with mCRPC (mCRPC) recruited in USA and Canada [9]. Samples were collected at the time of RP (LPC patients) or at mCRPC baseline before initiation of first-line enzalutamide or abiraterone treatment (mCRPC patients). We downloaded cfMeDIP-seq data for patients in this cohort from EGAS00001005522. Additionally, we downloaded clinical follow-up data available for 72 out of the 85 mCRPC patients in this cohort [9, 10]. Clinical characteristics are shown in Table S1.
Methylation array data
We downloaded Infinium 450 K methylation array data from the Marmal-aid database [11] for 1057 blood cell samples, collected from men without cancer and of varying ages and ethnicities (referred to as Marmal-aid blood cell (MBC) dataset), as well as for 180 primary prostate tumour and 52 healthy prostate tissue samples (referred to as Marmal-aid tissue (MAT) dataset). Furthermore, we downloaded Infinium Methylation EPIC BeadChip data of blood cell samples from 245 healthy men (referred to as EPIC dataset) from GSE152026 [12]. Further details are available in the Supplementary Methods and Table S2.
Sample processing and library preparation
The blood samples from patients in cohorts 1 and 2 were processed according to standard protocols [13], with plasma separated from cellular components by centrifugation (3000 × g for 10 min at 20 °C) within 2 h of blood draw. cfDNA was extracted from 2 to 4 mL of plasma using the QIAamp Circulating Nucleic acid kit (Qiagen) following the manufacturer’s instructions. cfDNA concentration and quality were evaluated by multiplex droplet digital PCR (ddPCR), as described previously [13], using the QX200 AutoDG ddPCR System (Bio-Rad). Samples were split into immunoprecipitated (IP) for cfMeDIP-seq and input control (IC) for low-pass whole genome sequencing (LPWGS). Libraries were prepared in accordance with the previously published protocol [7], with slight adjustments in reaction volumes to accommodate a lower input cfDNA concentration. Further details are provided in the Supplementary Methods.
Sequencing and initial data processing
Indexed cfMeDIP-seq libraries were paired-end sequenced (2 × 150bp) on an Illumina® Novaseq 6000 instrument (S-prime, S1, or S2 flow cell). Sequencing generated a median of 57.9 (14.3–113) million reads per IP library and 19.6 (3.8–68.5) million reads per IC library, corresponding to median coverages of 1.47x (0.27–4.27x) for IP libraries and 0.53x (0.08–3.37x) for IC libraries (Table S3). Raw Fastq files were demultiplexed (bcl2fastq, v2.20.0.422), and adaptor sequences were trimmed (Cutadapt, v1.16). Sequence reads were aligned and mapped to the hg19 reference genome (BWA MEM, v0.7.15), and PCR and optical duplicates were removed from each library (Samblaster, v0.1.24). The final bam files were realigned (GATK, v3.8.1.0), and Picard tools (v2.0.1) were used to calculate basic alignment statistics.
Bioinformatics processing of sequencing data
IC sample bam files were used as input for ichorCNA [14] to produce copy number profiles (bin size=10 kb) and estimate ctDNA fraction (ctDNA%). Samples with a ctDNA fraction above 3.0% were considered as ctDNA positive (Table S3). In silico quality control (QC) was evaluated for each IP and IC sample using the MEDIPS R package (v1.42.0) [15]. All samples passed initial QC, as they demonstrated efficient capture of methylated cfDNA fragments (median 2.8-fold enrichment in IP relative to IC samples) as well as sufficient sequencing depth (median depth of 1.47x for IP and 0.53x for IC samples; Table S3). Whole-methylome profiles were generated from IP sample bam files using the QSEA R package (v1.16.0) [16]. For each sample, read coverage was estimated per 300 base pair (bp) window across the genome. For normalisation of coverage data, copy number profiles of corresponding IC samples were imported into the QSEA workflow. For analysis, coverage per 300 bp window was converted into normalised reads per million mapped reads (nrpm).
Methylome landscape analysis
Differential methylation between mCRPC and NPCC samples in cohort 1 was analysed using QSEA, and 300 bp windows with a false-discovery rate (FDR) corrected p-value < 0.05, and absolute log2 fold change (log2FC) > 1.5 (hyper) or <−1.5 (hypo) were nominated as differentially methylated regions (DMRs). The genome-wide distribution of DMRs was visualised using the RIdeogram R package (v0.2.2). DMRs were annotated in relation to CpG and genic annotations using Bioconductor’s Annotatr package (v1.16.0) [17].
mCRPC signature creation using methylation data
We used DMRs identified in the methylome landscape analysis for the mCRPC signature creation. To improve the PC-specificity of the DMRs, we first filtered the DMRs using the MBC data to contain only DMRs that were commonly unmethylated in male blood cells, defined as: mean beta of the DMR < 0.05, and <1 × standard deviation.
We then used the cohort 1 methylation values to select DMRs that were highly hypermethylated in mCRPC patients (>1.2 nrpm), while showing very low methylation signal in NPCC (<0.05 nrpm). The resulting DMRs formed the methylation signature, with mean methylation levels (nrpm) of this signature reported hereafter.
For detection of me-ctDNA using the methylation signature, a fixed cutoff (me-ctDNA detection cutoff) was set based on cohort 1 NPCC, as: mean signature methylation (NPCC) + 2 × standard deviation (NPCC).
Me-ctDNA detection in a sample was then defined as: mean signature methylation (sample) > me-ctDNA detection cutoff.
Longitudinal me-ctDNA analyses during 1st-line mCRPC treatment
The changes in me-ctDNA during enzalutamide treatment were assessed using longitudinally collected plasma samples (at weeks 12 and 24) available for 22 mCRPC patients in cohort 2 (see Supplementary Methods). Based on the longitudinal me-ctDNA detection status, patients were divided into 2 groups: (1) Persistent: me-ctDNA detectable at baseline and in at least one of the two follow-up samples, (2) Non-persistent: me-ctDNA undetectable at baseline OR detectable at baseline but undetectable in both follow-up samples. Me-ctDNA dynamics during treatment were then evaluated in association to the duration of therapy response, assessed by serum PSA measurements and routine scans (bone scintigraphy or F-18Na/PET), respectively.
Statistical data analysis
All statistical analyses were conducted in R (v4.0.2), with two-sided p-values < 0.05 considered statistically significant. When appropriate, p-values were adjusted to correct for multiple testing using the Benjamini-Hochberg method. Mann-Whitney, Kruskal-Wallis, and Spearman’s correlation tests were used to compare groups. Exact McNemar’s test was used to compare sensitivity between ctDNA detection strategies. Code for analyses conducted in R is publicly available on GitHub at https://github.com/kkt8000/cfMeCaP-code.
Survival analyses, including Kaplan-Meier analyses and uni- and multivariate Cox regression analyses, were performed using the survminer (v0.4.9), survival (v3.1-12), and forestmodel (v0.6.2) R packages. PSA-PFS and OS from the time of treatment initiation were used as endpoints for survival analyses. Signature methylation levels were either included in the analyses as continuous variables, or patients were dichotomised into two groups (low and high signature methylation) based on the median methylation level of the specific cohort. For cox regression analyses, variables violating the proportional hazards assumptions were excluded. Further details are available in the supplementary methods.
Results
The cfDNA methylome landscape of mCRPC investigated by cfMeDIP-seq
For cfDNA methylome profiling in mCRPC patients, we performed cfMeDIP-seq on plasma samples from 27 mCRPC patients and 10 NPCC men in cohort 1 (Fig. 1a). Clinical characteristics are summarised in Table 1. Plasma ctDNA fractions, as estimated using ichorCNA, ranged from 22.3% to 72.4% in mCRPC patients, while no ctDNA was detected in NPCCs (Fig. S1A and Table S3).
a cfMeDIP-seq workflow, including generation of whole-genome methylation profiles from immunoprecipitated (IP) samples and copy-number profiles from input control (IC) samples. b DMR analyses of cohort 1. Red dots represent regions being hypermethylated in mCRPC patients relative to NPCCs (hyperDMRs: adj. p < 0.05, log fold change > 1.5). Blue dots represent regions being hypomethylated in mCRPC patients relative to NPCCs (hypoDMRs: adj. p < 0.05, log fold change < −1.5). c CpG annotation of cohort 1 hyperDMRs and hypoDMRs, respectively. d Gene relation annotation of cohort 1 hyperDMRs and hypoDMRs, respectively. The percentage of hyperDMRs or hypoDMRs, respectively, associated with a given annotation is shown on top of bars. Elements in (a) were created in BioRender: Sørensen, K. (2025) https://BioRender.com/khrfkru.
In cohort 1, we identified 38,841 differentially methylated genomic regions (DMRs) in cfDNA from mCRPC patients compared to NPCCs. Of these, 20,654 regions were hypermethylated (hyperDMR) and 18,187 were hypomethylated (hypoDMR) in mCRPC (Fig. 1b and Table S4). DMRs were widely distributed across the human genome (Fig. S1B). The majority of hyperDMRs mapped to CpG-rich genomic elements (CpG islands and shores, 12,348/20,654; Fig. 1c) and/or to genic elements (15,305/20,654; Fig. 1d), while most hypoDMRs mapped to CpG-poor shelves and open sea regions (17,233/18,187; Fig. 1c) and less than half to genic elements (6802/18,187; Fig. 1d). These findings align with the known PC methylome landscape, which is characterised by aberrant hypermethylation of CpG rich elements as well as overall global loss of DNA methylation, as identified in prior tumour tissue-based studies [18,19,20]. Moreover, several hyperDMRs identified in our analysis were associated to genes previously described as common targets for aberrant hypermethylation in PC tumours (e.g. GSTP1, RASSF1, CCDC181, and HAPLN3 [20,21,22,23]; Table S4), further supporting the validity of our cfDNA-based findings.
Development of mCRPC-specific cfDNA methylation signature; cfMeCaP
Next, based on the 20,654 hyperDMRs identified in mCRPC patients in cohort 1, we sought to establish a signature for detection of PC-associated methylated ctDNA (me-ctDNA) in plasma. To improve specificity for PC, we employed two stringent filtration steps (Fig. 2a; see also ‘Materials and methods’). First, we limited our analysis to hyperDMRs that were unmethylated in a large set of blood cell samples from cancer-free men, thereby excluding a total of 18,357 hyperDMRs. Second, we selected hyperDMRs that were unmethylated in cohort 1 NPCCs while highly methylated in cohort 1 mCRPC, thereby excluding an additional 2249 hyperDMRs (Fig. 2a). This identified 48 mCRPC-associated hyperDMRs, distributed across the whole genome (Fig. S2A and Table S5). We refer to this novel genome-wide methylation signature as the ‘cell-free DNA Methylation signature for Cancer of the Prostate’ (cfMeCaP; Fig. 2a).
a Schematic overview of the creation of cfMeCaP. First, cohort 1 hyperDMRs were filtered against a large reference methylome of blood cell samples (n = 1057) to retain only regions that were also unmethylated in blood cells. Next, based on cohort 1 methylation levels, hyperDMRs that were highly methylated in mCRPC patients (>1.2 nrpm) while unmethylated in NPCCs (<0.05 nrpm) were retained. This filtration identified 48 genomic regions (the cfMeCaP signature). cfMeCaP methylation levels were tested in 4 different datasets to evaluate the signature’s association to PC. b Mean cfMeCaP methylation of cohort 1 (cfMeDIP-seq data). c Mean cfMeCaP methylation of 245 healthy men from GSE152026 (EPIC data). d Mean cfMeCaP methylation of 52 healthy prostate samples and 180 primary prostate tumour samples from the Marmal-aid database (Marmal-aid tissue; MAT, 450 K data). e Mean cfMeCaP methylation of cohort 2 (cfMeDIP-seq data). f Mean cfMeCaP methylation of cohort 3 (cfMeDIP-seq data). Nrpm= Normalised reads per million, gDNA genomic DNA. Elements in (a) were created in BioRender: Sørensen, K. (2025) https://BioRender.com/l90k0wj.
As expected, in cohort 1, cfMeCaP showed significantly (p < 0.001) higher methylation levels in mCRPC patients relative to NPCCs (Fig. 2b). In further support of its PC-specificity, cfMeCaP methylation levels were extremely low (median β = 0.06) in blood cell samples from an external cohort of 245 healthy men (EPIC dataset, Fig. 2c), while they were significantly higher in an external set of 180 prostate tumour tissue (median β = 0.42) compared to 52 healthy prostate tissue samples (median β = 0.15) (p < 0.001; MAT dataset, Fig. 2d).
cfMeCaP is consistently methylated across PC cohorts
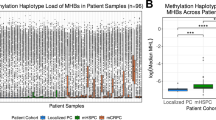
For independent validation, we first tested cfMeCaP in cohort 2 (Fig. 2a and Table 1), comprising plasma samples from 43 mCRPC patients and 8 men without cancer (NPCCs), alongside 40 HSPC and 10 LPC patients to investigate if cfMeCaP signals were detectable in earlier-stage PC. In cohort 2, we observed significantly higher cfMeCaP levels in mCRPC patients (median: 0.24 nrpm) compared to NPCCs (median: 0.02 nrpm; p < 0.001; Fig. 2e). Moreover, no significant difference in cfMeCaP methylation levels was observed between LPC (median 0.02 nrpm), HSPC (median: 0.03 nrpm), and NPCC men (median 0.02 nrpm) in this cohort (p > 0.05; Fig. 2e). Consistent with this, in an external validation cohort (cohort 3, Table S1) of plasma samples from 85 mCRPC patients and 30 LPC patients, we observed significantly higher cfMeCaP levels in mCRPC patients (median: 0.56 nrpm) compared to LPC patients (median: 0.02 nrpm; p < 0.001; Fig. 2f). Together, these results confirmed the consistency of cfMeCaP across multiple PC cohorts.
cfMeCaP enables efficient detection of plasma me-ctDNA
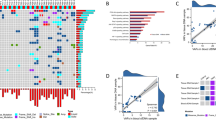
To further explore the potential of cfMeCaP as a tool to detect me-ctDNA in plasma from PC patients, we defined a cutoff (0.0289 nrpm) for me-ctDNA detection based on cfMeCaP methylation levels of NPCCs in cohort 1 (see ‘Materials and methods’) and used it to classify patients as me-ctDNA positive or negative. Accordingly, cfMeCaP detected me-ctDNA in all mCRPC patients in cohort 1, while none of the NPCC men were me-ctDNA positive (Fig. 3a). Testing this cutoff, cfMeCaP detected me-ctDNA in 84% of mCRPC patients in cohort 2 (Fig. 3b), whereas 95% of mCRPC patients were me-ctDNA positive in cohort 3 (Fig. 3c). Furthermore, cfMeCaP was able to detect me-ctDNA in 20% of LPC and 45% of HSPC patients in cohort 2 (Fig. 3b), as well as 20% of LPC patients in cohort 3 (Fig. 3c).
a Detection of methylated ctDNA (me-ctDNA) using the cfMeCaP signature in cohort 1. b cfMeCaP me-ctDNA detection in cohort 2 (internal validation). c cfMeCaP me-ctDNA detection in cohort 3 (external validation). d Detection of ctDNA, as estimated from copy number profiles using ichorCNA, in cohort 1. e IchorCNA copy number-based ctDNA detection in cohort 2. f Detection of ctDNA, as estimated from mutation calls from targeted sequencing data, in a subset of cohort 3 (72 mCRPC patients). g cfMeCaP me-ctDNA detection stratified by IchorCNA copy number-based ctDNA detection in patients from cohort 1 and 2 (n = 120). h cfMeCaP me-ctDNA detection stratified by mutation-based ctDNA detection in a subset of cohort 3 mCRPC patients (n = 72).
To compare cfMeCaP me-ctDNA detection with routine clinical markers, we assessed its relationship with PSA and metastatic characteristics for PC patients in cohorts 1–3. We observed no differences in serum PSA levels between me-ctDNA positive and negative PC patients in any of the cohorts (p > 0.05, Fig. S3A–C). Among mCRPC patients, where all individuals had metastatic disease, me-ctDNA detection appeared to vary by metastatic site, with the highest detection rates observed in those with visceral involvement (Fig. S3D–F). In HSPC, me-ctDNA positivity was enriched in patients with M1 compared to M0 disease (p < 0.001, McNemar’s tests, Fig. S3G).
cfMeCaP is superior to genomics-based ctDNA detection strategies
Next, we explored the performance of cfMeCaP in relation to a copy number-based (i.e. ichorCNA analysis of LPWGS data) and a mutation-based (i.e. deep targeted sequencing of 72 mCRPC driver genes) ctDNA detection strategy. While the copy number-based strategy, performed on cohort 1 and cohort 2 patients, achieved 100% sensitivity in cohort 1 (Fig. 3d), this strategy was inferior to cfMeCaP in cohort 2, with none of the LPC patients, and only 15% of the HSPC and 63% of the mCRPC patients in this cohort being identified as ctDNA positive (Fig. 3e and Table S6). Similarly, a mutation-based detection strategy, performed on a subset of mCRPC patients in cohort 3 (n = 72), detected ctDNA in only 76% of the mCRPC patients (Fig. 3f and Table S6). Furthermore, cfMeCaP detected me-ctDNA in all PC samples classified as ctDNA positive by either the copy number (cohort 1 + 2, n = 120) or the mutation-based (cohort 3, n = 72) strategies, and an additional 38% and 76% of samples classified as ctDNA negative by the copy number and mutation-based strategies, respectively (Fig. 3g, h). Together, our results support the superior sensitivity of the cfMeCaP-based me-ctDNA detection strategy.
Longitudinal cfMeCaP profiling identifies early treatment failure
To explore the potential of cfMeCaP in predicting early treatment failure in mCRPC patients, we combined cfMeCaP me-ctDNA detection with longitudinal plasma sampling at weeks 12 and 24, using 22 mCRPC patients from cohort 2 (Fig. 4a). Response to treatment was monitored using routine PSA measurements and scans every 12 weeks. In this cohort, all patients with detectable me-ctDNA at baseline and in at least one follow-up sample (i.e. Persistent group, n = 13) had either primary treatment resistance or had PSA and/or radiographic progression within 1 year of treatment initiation (Fig. 4b–d). In contrast, none of the patients with undetectable me-ctDNA at baseline or undetectable in both follow-up samples (i.e. Non-persistent group, n = 9) experienced PSA or radiographic progression within 1 year of treatment initiation (Fig. 4b–d). On median, patients in the Persistent group had PSA-PFS and radiographic PFS (rPFS) of 4.4 months and 5.4 months, respectively, compared to 65.5 months and 67.5 months for patients in the Non-persistent group. These results indicate that longitudinal me-ctDNA detection during 1st-line enzalutamide treatment is associated with early treatment failure.
a Schematic overview of the longitudinal plasma sample collection in a subset of cohort 2 mCRPC patients receiving enzalutamide as first-line mCRPC treatment (n = 22). Patients were monitored for disease progression at 12 weeks and 24 weeks after treatment initiation using routine PSA measurements and scans. Plasma samples were collected at baseline, at the 12-week follow-up scan, and at the 24-week follow-up scan, and subjected to whole methylome profiling followed by cfMeCaP analyses. b cfMeCaP me-ctDNA detection analyses in cohort 2 longitudinal samples. Based on the longitudinal me-ctDNA detection status, patients were split into two me-ctDNA dynamics groups: the Non-persistent group (n = 9) and the Persistent group (n = 13). Days to follow-up events outside the range of the figure are indicated by colour coding: Yellow: Days to PSA progression, Red: Days to radiographic progression, Black: End of protocol due to death, Grey: Patient still responding – days to last follow-up. c cfMeCaP me-ctDNA dynamics groups stratified by the development of PSA progression within 1 year from enzalutamide initiation. d cfMeCaP me-ctDNA dynamics groups stratified by the development of radiographic progression within 1 year from enzalutamide initiation. Elements in (a) were created in BioRender: Sørensen, K. (2025) https://BioRender.com/3fq8x90.
cfMeCaP methylation at mCRPC baseline is associated with poor outcome
Finally, we explored the prognostic potential of cfMeCaP at mCRPC baseline. Given that most patients had detectable me-ctDNA at mCRPC baseline (100%, 84% and 95% in cohorts 1, 2, and 3, respectively), using me-ctDNA detectability for prognostic stratification would result in highly unbalanced groups, limiting statistical power (Fig. S4). Therefore, we instead focused on the cfMeCaP methylation levels. In cohort 1, high cfMeCaP methylation was associated with significantly shorter time to PSA-progression on first-line ARSI treatment in both Kaplan–Meier (p = 0.014; Fig. S5A) and univariate cox regression (p = 0.027; Fig. S6A) analyses. This association was validated in both cohort 2 (p < 0.015; Figs. S5B and S6B) and cohort 3 mCRPC patients (p < 0.002; Figs. S5C and S6C). In line with these findings, high cfMeCaP methylation levels were associated with significantly shorter OS for mCRPC patients in cohort 1 (p < 0.009; Figs. S5D and S6D), which was validated in cohort 2 (p < 0.002; Figs. S5E and S6E) and cohort 3 (p < 0.001; Figs. S5F and S6F). Furthermore, in multivariate Cox regression analyses, high cfMeCaP methylation remained a significant adverse predictor of PSA-PFS and OS, independent of previously described prognostic clinical features (i.e. serum PSA level and localisation of the metastases) in all three mCRPC cohorts (p < 0.05; Fig. 5a–f). Taken together, our results emphasise the promising clinical potential of using cfDNA-derived multi-marker methylation signatures for mCRPC prognostication.
a Multivariate cox-regression analysis in cohort 1 using PSA-PFS as endpoint. b Multivariate cox-regression analysis in cohort 2 using PSA-PFS as endpoint. c Multivariate cox-regression analysis in cohort 3 using PSA-PFS as endpoint. d Multivariate cox-regression analysis in cohort 1 using OS as endpoint. e Multivariate cox-regression analysis in cohort 2 using OS as endpoint. f Multivariate cox-regression analysis in cohort 3 using OS as endpoint. All parameters except “Metastases” are analysed as continuous variables. Grey p-values indicate non-significance (p > 0.05). HR hazard ratio, LN lymph node metastases.
Discussion
In the present study, we used methylome profiling of plasma cfDNA from patients with mCRPC and men without cancer to show that clinically relevant genome-wide methylation markers can be identified in cfDNA from liquid biopsies. As a significant proportion of cfDNA arise from blood cells [24], tumour-specific cell-free methylation signals may become diluted by the methylation signals from blood cells, as evident from e.g. the low sensitivity of certain methylation-based classifiers in predicting different cancer types [25,26,27]. One approach to increase tumour specificity of methylation signals is to exclude blood cell-derived methylation signals from cfDNA methylomes using parallel whole-methylome sequencing of matched blood cell DNA [28]. Although this approach may effectively remove patient-specific blood cell methylation signals, it comes with a substantial increase in analytical costs. Instead, in our analysis, we generated an in-silico reference methylome of blood cell DNA from a large cohort of cancer-free men, using publicly available methylation data, and subsequently restricted our analyses to regions that are unmethylated in blood cells. To reduce the potential influence of age-associated methylation changes in blood cells of older adult men, NPCCs in this study were selected to match the age range of the patients, and the blood cell reference methylome was derived predominantly from adult donors (median age 60 years).
The genome-wide methylation signature (cfMeCaP) established in this study consisted of 48 genomic regions that were significantly hypermethylated in plasma from mCRPC patients compared to NPCCs. While plasma cfMeCaP methylation in LPCs or HSPCs did not significantly differ from NPCCs, we showed that cfMeCaP was hypermethylated in primary PC tumours compared to healthy prostate tissue, supporting the biological validity of the signature. Consistent with this, multiple cfMeCaP regions were associated with gene families previously described as hypermethylated in primary PC tumours compared to healthy prostate tissue, including e.g. the LIM homeobox gene family (LHX6, LHX9 [29,30,31]), the ANTP homeobox gene family (HOXD1 [29, 31]), and the Oligodendrocyte Transcription Factor gene family (OLIG2, OLIG3 [29, 31, 32]). Additionally, one cfMeCaP region (cfMeCaP region 3) was annotated to the CCDC181 promoter, a region previously reported as part of a three-gene prognostic methylation signature in PC [20].
The cfMeCaP signature was capable of highly sensitive detection of me-ctDNA in mCRPC patients in all three cohorts (84–100% sensitivity). This is in accordance with previous studies, which in general have reported high sensitivity for plasma ctDNA detection as well as high ctDNA% in mCRPC patients, detected using different strategies such as mutations [33, 34] or copy-number variation [35, 36]. We also compared the performance of cfMeCaP to alternative ctDNA detection approaches. We found that cfMeCaP outperformed ichorCNA, a method based on copy-number variation estimated from low-coverage WGS data. While differences in sequencing depth (ichorCNA-based: ~0.5x, cfMeCaP-based: ~1.5x) may contribute to discrepancies in the performance of these strategies, the cfMeCaP strategy also outperformed a mutation-based ctDNA detection strategy, in which the ctDNA% was estimated from targeted sequencing data with a median sequencing depth of 688x [10]. Together, our findings highlight that a multi-methylation marker signature shows great potential in tumour-naïve ctDNA detection in mCRPC.
Using our cfMeCaP methylation signature, the majority of mCRPC patients had detectable levels of me-ctDNA at the time of mCRPC diagnosis. When assessed for its prognostic potential, we found high cfMeCaP methylation levels at mCRPC baseline to be an adverse predictor of both PSA-PFS and OS, independent of established clinical parameters such as serum PSA levels or the localisation of metastasis. Our results confirm and expand on previous studies that have also reported on the prognostic biomarker potential of ctDNA methylation in mCRPC [9, 37,38,39] as well as other cancer types, including head and neck squamous cell carcinoma (HNSCC) [28] and hepatocellular carcinoma [40]. Several studies have reported genomics-based ctDNA% as an adverse predictor of PSA-PFS and OS in mCRPC [33,34,35], suggesting that ctDNA methylation may primarily reflect plasma ctDNA levels. However, in our analysis, high cfMeCaP methylation remained a significant adverse predictor of both PSA-PFS and OS, independent of the ichorCNA-based and mutation-based ctDNA% in cohort 1 and 3 mCRPC patients, respectively (Fig. S7A–F). In accordance with this, a previous study reported a composite methylation score of 5 genomic regions to hold prognostic potential independent of ctDNA abundance in HNSCC patients [28]. These findings suggest that cfDNA methylation analysis may provide a refined prognostication of mCRPC patients, possibly due to its improved analytical sensitivity. To support clinical implementation, larger independent cohorts are needed to validate cfMeCaP as a prognostic biomarker for mCRPC, either as a continuous measure or to define cohort- or context-specific cutoffs if dichotomisation is desired.
Moreover, we showed that longitudinal testing of me-ctDNA during the initial months of mCRPC treatment could identify patients with rapid disease progression, for whom treatment intensification might provide additional clinical benefits. In line with these findings, a previous study showed significantly shorter rPFS in mCRPC patients with detectable ctDNA at baseline and after 9 weeks of treatment (enzalutamide w/o atezolizumab), compared to patients with undetectable ctDNA at both time points [34]. Extensive research efforts are currently directed towards demonstrating the clinical utility of ctDNA-guided monitoring of therapy response in multiple cancer types. For example in patients with advanced non-small-cell lung cancer undergoing systemic therapy, ctDNA-based molecular progression was detectable before radiographic progression [41]. The study concluded that a third of all therapy cycles were likely ineffective as they were administered after molecular progression. Such a conclusion might also have important significance for mCRPC treatment, as 20-40% of mCRPC patients are known to harbour primary ARSI resistance and are less likely to benefit from ARSI, the current mainstay for mCRPC treatment [42,43,44]. Noteworthily, we further showed that a me-ctDNA analysis at baseline and 12 + 24 weeks was able to identify patients who did not progress within 12 months of treatment initiation. For these patients, intensive monitoring of the disease might not be necessary, enabling healthcare resources to be reallocated where they are needed the most.
Although the cfMeCaP signature was developed entirely in the mCRPC setting, it enabled me-ctDNA detection in one in five LPC patients and approximately every other HSPC patient. While low ctDNA abundance is generally characteristic of earlier stages of PC, previous studies have associated detectable ctDNA with rapid disease progression in both LPC and HSPC [23, 45, 46]. Given the small sample size and limited clinical information of LPC/HSPC patients in our study, larger cohorts with longitudinal follow-up are now warranted to determine the prognostic utility of the cfMeCaP signature in early-stage and treatment-naïve PC.
Our study is, however, not without limitations. First, due to the inclusion of only a few NPCCs, available 450 K data were used to exclude methylation signals from normal blood. This restricted our signature to regions that are covered by the 450 K array. Second, by focusing the signature development on mCRPC patients, methylation signals from early-stage PC could potentially be missed, in turn limiting the sensitivity of cfMeCaP in these early stages. Third, since cfMeCaP relies on plasma ctDNA, tumours that do not shed detectable levels of ctDNA might be missed by this method. Fourth, additional prognostic clinical markers, such as LDH or neutrophil-to-lymphocyte ratio, were not available across all mCRPC cohorts and therefore could not be included in the survival analyses. Finally, while the longitudinal me-ctDNA dynamics show promising clinical potential, larger studies are needed to further clarify the association between me-ctDNA dynamics and response to therapy in patients with mCRPC.
Conclusions
In conclusion, this study established a tumour-naïve signature (cfMeCaP) for the detection of methylated ctDNA in PC. Compared to existing copy number/mutation-based methods, our cfMeCaP signature showed superior sensitivity in the detection of cancer across different stages of PC. High cfMeCaP at mCRPC baseline was significantly associated with poor PSA-PFS and OS, and cfMeCaP-based me-ctDNA dynamics across time-points proved to be an effective indicator of early treatment failure, strongly encouraging further clinical validation of these results.
Data availability
The data generated in this study are available through restricted access from GenomeDK (https://genome.au.dk/) under the accession number GDK000020 (https://genome.au.dk/library/GDK000020). All other data are available within the article or its Supplementary Materials. External datasets are available from their respective repositories, as described in the ‘Materials and methods’.
References
Khoshkar Y, Westerberg M, Adolfsson J, Bill-Axelson A, Olsson H, Eklund M, et al. Mortality in men with castration-resistant prostate cancer—A long-term follow-up of a population-based real-world cohort. BJUI Compass. 2021;3:173–83.
Le TK, Duong QH, Baylot V, Fargette C, Baboudjian M, Colleaux L, et al. Castration-resistant prostate cancer: from uncovered resistance mechanisms to current treatments. Cancers. 2023;15:5047.
Teo MY, Rathkopf DE, Kantoff P. Treatment of advanced prostate cancer. Annu Rev Med. 2019;70:479–99.
Chi K, Hotte SJ, Joshua AM, North S, Wyatt AW, Collins LL, et al. Treatment of mCRPC in the AR-axis-targeted therapy-resistant state. Ann Oncol. 2015;26:2044–56.
Tian S, Lei Z, Gong Z, Sun Z, Xu D, Piao M. Clinical implication of prognostic and predictive biomarkers for castration-resistant prostate cancer: a systematic review. Cancer Cell Int. 2020;20:409.
Wyatt AW, Annala M, Aggarwal R, Beja K, Feng F, Youngren J, et al. Concordance of circulating tumor DNA and matched metastatic tissue biopsy in prostate cancer. J Natl Cancer Inst. 2017;109:djx118.
Shen SY, Burgener JM, Bratman SV, De Carvalho DD. Preparation of cfMeDIP-seq libraries for methylome profiling of plasma cell-free DNA. Nat Protoc. 2019;14:2749–80.
Shen SY, Singhania R, Fehringer G, Chakravarthy A, Roehrl MHA, Chadwick D, et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature. 2018;563:579–83.
Chen S, Petricca J, Ye W, Guan J, Zeng Y, Cheng N, et al. The cell-free DNA methylome captures distinctions between localized and metastatic prostate tumors. Nat Commun. 2022;13:6467.
Annala M, Vandekerkhove G, Khalaf D, Taavitsainen S, Beja K, Warner EW, et al. Circulating tumor DNA genomics correlate with resistance to abiraterone and enzalutamide in prostate cancer. Cancer Discov. 2018;8:444–57.
Lowe R, Rakyan VK. Marmal-aid - a database for Infinium HumanMethylation450. BMC Bioinform. 2013;14:359.
Hannon E, Dempster EL, Mansell G, Burrage J, Bass N, Bohlken MM, et al. DNA methylation meta-analysis reveals cellular alterations in psychosis and markers of treatment-resistant schizophrenia. eLife. 2021;10:e58430.
Thomas R, Lone VS, Rune T, Heidi T, Søren V, Iver N, et al. Analysis of circulating tumour DNA to monitor disease burden following colorectal cancer surgery. Gut. 2016;65:625.
Adalsteinsson VA, Ha G, Freeman SS, Choudhury AD, Stover DG, Parsons HA, et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat Commun. 2017;8:1324.
Lienhard M, Grimm C, Morkel M, Herwig R, Chavez L. MEDIPS: genome-wide differential coverage analysis of sequencing data derived from DNA enrichment experiments. Bioinformatics. 2014;30:284–6.
Lienhard M, Grasse S, Rolff J, Frese S, Schirmer U, Becker M, et al. QSEA—modelling of genome-wide DNA methylation from sequencing enrichment experiments. Nucleic Acids Res. 2017;45:e44.
Cavalcante RG, Sartor MA. annotatr: genomic regions in context. Bioinformatics. 2017;33:2381–3.
Strand SH, Switnicki M, Moller M, Haldrup C, Storebjerg TM, Hedegaard J, et al. RHCG and TCAF1 promoter hypermethylation predicts biochemical recurrence in prostate cancer patients treated by radical prostatectomy. Oncotarget. 2017;8:5774–88.
Zhao SG, Chen WS, Li H, Foye A, Zhang M, Sjöström M, et al. The DNA methylation landscape of advanced prostate cancer. Nat Genet. 2020;52:778–89.
Haldrup C, Mundbjerg K, Vestergaard EM, Lamy P, Wild P, Schulz WA, et al. DNA methylation signatures for prediction of biochemical recurrence after radical prostatectomy of clinically localized prostate cancer. J Clin Oncol. 2013;31:3250–8.
Lee WH, Morton RA, Epstein JI, Brooks JD, Campbell PA, Bova GS, et al. Cytidine methylation of regulatory sequences near the pi-class glutathione S-transferase gene accompanies human prostatic carcinogenesis. Proc Natl Acad Sci USA. 1994;91:11733–7.
Liu L, Yoon J-H, Dammann R, Pfeifer GP. Frequent hypermethylation of the RASSF1A gene in prostate cancer. Oncogene. 2002;21:6835–40.
Bjerre MT, Nørgaard M, Larsen OH, Jensen SØ, Strand SH, Østergren P, et al. Epigenetic analysis of circulating tumor dna in localized and metastatic prostate cancer: evaluation of clinical biomarker potential. Cells. 2020;9:1362.
Moss J, Magenheim J, Neiman D, Zemmour H, Loyfer N, Korach A, et al. Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease. Nat Commun. 2018;9:5068.
Liu MC, Oxnard GR, Klein EA, Swanton C, Seiden MV, Liu MC, et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann Oncol. 2020;31:745–59.
Bronkhorst AJ, Ungerer V, Holdenrieder S. The emerging role of cell-free DNA as a molecular marker for cancer management. Biomol Detect Quantif. 2019;17:100087.
Sharma M, Verma RK, Kumar S, Kumar V. Computational challenges in detection of cancer using cell-free DNA methylation. Comput Struct Biotechnol J. 2022;20:26–39.
Burgener JM, Zou J, Zhao Z, Zheng Y, Shen SY, Huang SH, et al. Tumor-naïve multimodal profiling of circulating tumor dna in head and neck squamous cell carcinoma. Clin Cancer Res. 2021;27:4230–44.
Kim JH, Dhanasekaran SM, Prensner JR, Cao X, Robinson D, Kalyana-Sundaram S, et al. Deep sequencing reveals distinct patterns of DNA methylation in prostate cancer. Genome Res. 2011;21:1028–41.
Kron K, Trudel D, Pethe V, Briollais L, Fleshner N, van der Kwast T, et al. Altered DNA methylation landscapes of polycomb-repressed loci are associated with prostate cancer progression and ERG oncogene expression in prostate cancer. Clin Cancer Res. 2013;19:3450–61.
Massie CE, Mills IG, Lynch AG. The importance of DNA methylation in prostate cancer development. J Steroid Biochem Mol Biol. 2017;166:1–15.
Aryee MJ, Liu W, Engelmann JC, Nuhn P, Gurel M, Haffner MC, et al. DNA methylation alterations exhibit intraindividual stability and interindividual heterogeneity in prostate cancer metastases. Sci Transl Med. 2013;5:169ra10–ra10.
Fonseca NM, Maurice-Dror C, Herberts C, Tu W, Fan W, Murtha AJ, et al. Prediction of plasma ctDNA fraction and prognostic implications of liquid biopsy in advanced prostate cancer. Nat Commun. 2024;15:1828.
Sweeney CJ, Petry R, Xu C, Childress M, He J, Fabrizio D, et al. Circulating tumor DNA assessment for treatment monitoring adds value to PSA in metastatic castration-resistant prostate cancer. Clin Cancer Res. 2024;30:4115–22.
Nørgaard M, Bjerre MT, Fredsøe J, Vang S, Jensen JB, De Laere B, et al. Prognostic value of low-pass whole genome sequencing of circulating tumor DNA in metastatic castration-resistant prostate cancer. Clin Chem. 2023;69:386–98.
Sumanasuriya S, Seed G, Parr H, Christova R, Pope L, Bertan C, et al. Elucidating prostate cancer behaviour during treatment via low-pass whole-genome sequencing of circulating tumour DNA. Eur Urol. 2021;80:243–53.
Wu A, Cremaschi P, Wetterskog D, Conteduca V, Franceschini GM, Kleftogiannis D, et al. Genome-wide plasma DNA methylation features of metastatic prostate cancer. J Clin Investig. 2020;130:1991–2000.
Kluge K, Lotz V, Einspieler H, Haberl D, Spielvogel C, Amereller D, et al. Imaging and outcome correlates of ctDNA methylation markers in prostate cancer: a comparative, cross-sectional [⁶⁸Ga]Ga-PSMA-11 PET/CT study. Clin Epigenetics. 2025;17:36.
Franceschini GM, Quaini O, Mizuno K, Orlando F, Ciani Y, Ku S-Y, et al. Noninvasive detection of neuroendocrine prostate cancer through targeted cell-free DNA methylation. Cancer Discov. 2024;14:424–45.
Xu R-h, Wei W, Krawczyk M, Wang W, Luo H, Flagg K, et al. Circulating tumour DNA methylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nat Mater. 2017;16:1155–61.
Frank MS, Andersen CSA, Ahlborn LB, Pallisgaard N, Bodtger U, Gehl J. Circulating Tumor DNA monitoring reveals molecular progression before radiologic progression in a real-life cohort of patients with advanced non–small cell lung cancer. Cancer Res Commun. 2022;2:1174–87.
Scher HI, Beer TM, Higano CS, Anand A, Taplin M-E, Efstathiou E, et al. Antitumour activity of MDV3100 in castration-resistant prostate cancer: a phase 1–2 study. Lancet. 2010;375:1437–46.
Scher HI, Fizazi K, Saad F, Taplin M-E, Sternberg CN, Miller K, et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N Engl J Med. 2012;367:1187–97.
Ryan CJ, Smith MR, Bono JSD, Molina A, Logothetis CJ, Souza PD, et al. Abiraterone in metastatic prostate cancer without previous chemotherapy. N Engl J Med. 2013;368:138–48.
Hennigan ST, Trostel SY, Terrigino NT, Voznesensky OS, Schaefer RJ, Whitlock NC, et al. Low abundance of circulating tumor DNA in localized prostate cancer. JCO Precis Oncol. 2019;3:1–13.
Lau E, McCoy P, Reeves F, Chow K, Clarkson M, Kwan EM, et al. Detection of ctDNA in plasma of patients with clinically localised prostate cancer is associated with rapid disease progression. Genome Med. 2020;12:72.
Acknowledgements
The authors would like to thank all staff involved in the patient recruitment and sample collection at the Departments of Urology at Aarhus University Hospital, as well as the laboratory technicians, clinical academics, and bioinformaticians at the Department of Molecular Medicine for their invaluable help during the project. We acknowledge Aarhus Genome Data Center for providing access to the GenomeDK high-performance computing facility.
Funding
This project is supported by the Danish Cancer Society, the Novo Nordisk Foundation, the Central Denmark Region Health Fund, the Graduate School of Health – Aarhus University, and the Danish Cancer Research Foundation. Open access funding provided by Aarhus Universitet.
Author information
Authors and Affiliations
Contributions
KK, MN, PL, RE, and KDS designed the study. TFH, PJSO, AZ and MB provided study materials. KK and LI processed and analysed the data. KK, LI, PVS, and MN interpreted the results. KK, LI, PVS, and KDS wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
RE has received honoraria from GU-ASCO, Janssen, University of Chicago, Dana Farber Cancer Institute USA as a speaker, as well as educational honorarium from Bayer and Ipsen. RE is a member of an external expert committee to AstraZeneca UK, the Active Surveillance Movember Committee, and the SAB of Our Future Health. RE undertakes private practice as a sole trader at The Royal Marsden NHS Foundation Trust and 90 Sloane Street SW1X 9PQ and 280 Kings Road SW3 4NX, London, UK.
Ethical approval and consent to participate
All research forming the basis of this study was conducted in accordance with relevant rules, regulations, and guidelines and conform to the principles of the Declaration of Helsinki. Written informed consent was obtained from all participants prior to their donation of plasma samples for a research biobank (approved by The Central Denmark Region Committees on Health Research Ethics [jr. nr. 2000-0299], The Southern Denmark Region Committees on Health Research Ethics [S-20160029], and The Danish Data Protection Agency [jr. nr. 2013-41-2041, jr. nr. 2007-58-0010, and jr. nr. 17/34479]). The present study was approved by the Danish National Committee on Health Research Ethics [jr. nr. 1302791 and jr. nr. 1603542], who waived the requirement for patient consent, as (1) the new analyses were considered to not pose any health risk or other significant disadvantages to the participants and (2) an expert committee was established to deal with potential (although unlikely) incidental genetic findings in this new study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kondrup, K., Iisager, L., Salachan, P.V. et al. Genome-wide methylome profiling of cell-free DNA enables prognostication of patients with castration-resistant prostate cancer. Br J Cancer (2026). https://doi.org/10.1038/s41416-026-03432-y
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41416-026-03432-y