Abstract
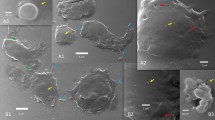
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus is highly contagious and causes lymphocytopenia, but the underlying mechanisms are poorly understood. We demonstrate here that heterotypic cell-in-cell structures with lymphocytes inside multinucleate syncytia are prevalent in the lung tissues of coronavirus disease 2019 (COVID-19) patients. These unique cellular structures are a direct result of SARS-CoV-2 infection, as the expression of the SARS-CoV-2 spike glycoprotein is sufficient to induce a rapid (~45.1 nm/s) membrane fusion to produce syncytium, which could readily internalize multiple lines of lymphocytes to form typical cell-in-cell structures, remarkably leading to the death of internalized cells. This membrane fusion is dictated by a bi-arginine motif within the polybasic S1/S2 cleavage site, which is frequently present in the surface glycoprotein of most highly contagious viruses. Moreover, candidate anti-viral drugs could efficiently inhibit spike glycoprotein processing, membrane fusion, and cell-in-cell formation. Together, we delineate a molecular and cellular rationale for SARS-CoV-2 pathogenesis and identify novel targets for COVID-19 therapy.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–3.
Wu F, Zhao S, Yu B, Chen Y-M, Wang W, Song Z-G, et al. A new coronavirus associated with human respiratory disease in China. Nature. 2020;https://doi.org/10.1038/s41586-020-2008-3.
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl J Med. 2020;382:727–33.
Li F. Structure, function, and evolution of coronavirus spike proteins. Annu Rev Virol. 2016;3:237–61.
Jiang X, Zhang Z, Wang C, Ren H, Gao L, Peng H, et al. Bimodular effects of D614G mutation on the spike glycoprotein of SARS-CoV-2 enhance protein processing, membrane fusion, and viral infectivity. Signal Transduct Target Ther. 2020;5:268–71.
Yurkovetskiy L, Wang X, Pascal KE, Tomkins-Tinch C, Nyalile TP, Wang Y, et al. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell. 2020;183:739–51.e8.
Wang C, Horby PW, Hayden FG, Gao GF. A novel coronavirus outbreak of global health concern. Lancet. 2020;395:470–3.
WHO. Coronavirus disease (COVID-2019) situation report-174. Situation reports. 2020;doi: 20200318-sitrep-174-covid-19:https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports.
Guan WJ, Ni ZY, Hu Y, Liang WH, Ou CQ, He JX, et al. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 2020; https://doi.org/10.1056/NEJMoa2002032.
Huang H, Chen Z, Sun Q. Mammalian cell competitions, cell-in-cell phenomena and their biomedical implications. Curr Mol Med. 2015;15:852–60.
Mackay HL, Muller PAJ. Biological relevance of cell-in-cell in cancers. Biochem Soc Trans. 2019;47:725–32.
Sun Q, Luo T, Ren Y, Florey O, Shirasawa S, Sasazuki T, et al. Competition between human cells by entosis. Cell Res. 2014;24:1299–310.
Rizzotto D, Villunger A. P53 clears aneuploid cells by entosis. Cell Death Differ. 2020; https://doi.org/10.1038/s41418-020-00659-x.
Liang J, Niu Z, Zhang B, Yu X, Zheng Y, Wang C, et al. p53-dependent elimination of aneuploid mitotic offspring by entosis. Cell Death Differ. 2021;28:799–813.
Zhang X, Niu Z, Qin H, Fan J, Wang M, Zhang B, et al. Subtype-based prognostic analysis of cell-in-cell structures in early breast cancer. Front Oncol. 2019;9:895.
Huang H, He M, Zhang Y, Zhang B, Niu Z, Zheng Y, et al. Identification and validation of heterotypic cell-in-cell structure as an adverse prognostic predictor for young patients of resectable pancreatic ductal adenocarcinoma. Signal Transduct Target Ther. 2020;5:246–8.
Fan J, Fang Q, Yang Y, Cui M, Zhao M, Qi J, et al. Role of heterotypic neutrophil-in-tumor structure in the prognosis of patients with Buccal Mucosa squamous cell carcinoma. Front Oncol. 2020;10:541878.
Wang M, Niu Z, Qin H, Ruan B, Zheng Y, Ning X, et al. Mechanical ring interfaces between adherens junction and contractile actomyosin to coordinate entotic cell-in-cell formation. Cell Rep. 2020;32:108071.
Sun Q, Cibas ES, Huang H, Hodgson L, Overholtzer M. Induction of entosis by epithelial cadherin expression. Cell Res. 2014;24:1288–98.
Niu Z, He M, Sun Q. Molecular mechanisms underlying cell-in-cell formation: core machineries and beyond. J Mol Cell Biol. 2021; https://doi.org/10.1093/jmcb/mjab015.
Ruan B, Niu Z, Jiang X, Li Z, Tai Y, Huang H, et al. High frequency of cell-in-cell formation in heterogeneous human breast cancer tissue in a patient with poor prognosis: a case report and literature review. Front Oncol. 2019;9:1444.
Huang H, Chen A, Wang T, Wang M, Ning X, He M, et al. Detecting cell-in-cell structures in human tumor samples by E-cadherin/CD68/CD45 triple staining. Oncotarget. 2015;6:20278–87.
Su Y, Ren H, Tang M, Zheng Y, Zhang B, Wang C, et al. Role and dynamics of vacuolar pH during cell-in-cell mediated death. Cell Death Dis. 2021;12:119.
Fais S, Overholtzer M. Cell-in-cell phenomena in cancer. Nat Rev Cancer. 2018;18:758–66.
Wang S, He MF, Chen YH, Wang MY, Yu XM, Bai J, et al. Rapid reuptake of granzyme B leads to emperitosis: an apoptotic cell-in-cell death of immune killer cells inside tumor cells. Cell Death Dis. 2013;4:e856.
Luo W, Yu H, Gou J, Li X, Sun Y, Li J, et al. Clinical pathology of critical patient with novel coronavirus pneumonia (COVID-19). Preprint at https://www.preprints.org/manuscript/202002.0407/v1.
Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med. 2020; https://doi.org/10.1016/s2213-2600(20)30076-x.
Fox SE, Akmatbekov A, Harbert JL, Li G, Brown JQ, Vander Heide RS. Pulmonary and cardiac pathology in Covid-19: the first autopsy series from New Orleans. medRxiv. 2020; https://doi.org/10.1101/2020.04.06.20050575:2020.04.06.20050575.
Zhou L, Niu Z, Jiang X, Zhang Z, Zheng Y, Wang Z, et al. SARS-CoV-2 targets by the pscRNA profiling of ACE2. TMPRSS2 Furin Proteases iScience. 2020;23:1–15.
Boonstra S, Blijleven JS, Roos WH, Onck PR, van der Giessen E, van Oijen AM. Hemagglutinin-mediated membrane fusion: a biophysical perspective. Annu Rev Biophys. 2018;47:153–73.
Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. 2020;26:450–2.
Ledford H. Safety fears over drug hyped to treat the coronavirus spark global confusion. Nature 2020;582:18–9.
Liu J, Cao R, Xu M, Wang X, Zhang H, Hu H, et al. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Disco. 2020;6:16.
Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30:269–71.
Fantini J, Di Scala C, Chahinian H, Yahi N. Structural and molecular modelling studies reveal a new mechanism of action of chloroquine and hydroxychloroquine against SARS-CoV-2 infection. Int J Antimicrob Agents. 2020;55:105960.
Amin M, Abbas G. Docking study of Chloroquine and Hydroxychloroquine interaction with SARS-CoV-2 spike glycoprotein-An in silico insight into the comparative efficacy of repurposing antiviral drugs. J Biomol Struct Dyn. 2020; https://doi.org/10.1080/07391102.2020.1775703:1-11.
Braga L, Ali H, Secco I, Chiavacci E, Neves G, Goldhill D, et al. Drugs that inhibit TMEM16 proteins block SARS-CoV-2 Spike-induced syncytia. Nature. 2021; https://doi.org/10.1038/s41586-021-03491-6.
Wang C, Chen A, Ruan B, Niu Z, Su Y, Qin H, et al. PCDH7 inhibits the formation of homotypic cell-in-cell structure. Front Cell Dev Biol. 2020;8:1–12.
Acknowledgements
We thank Dr. Cheng Cao, Dr. Hui Zhong and Dr. Guan Yang at Beijing Institute of Biotechnology for reagents. Dr. Hongguang Ren and Dr. Junjie Yue at Beijing Institute of Biotechnology for assistance in bioinformatics.
Funding
This work was supported by the Key Science & Technology Project of Beijing Educational Committee and the Beijing Municipal Natural Science Foundation (KZ202110025029 to HH), Beijing Municipal Administration of Hospitals Incubating Program (PX2021033 to HH), the National Key Research & Development Program of China (2018YFA0900804 to YZ and ZC, 2019YFA0903801 to QS), and the National Natural Science Foundation of China (81872314 to ZC, 31970685 to QS).
Author information
Authors and Affiliations
Contributions
Concept and design: QS; Phenotype: QS and ZZ; Tissue staining, imaging and analysis: ZN and YQW; Human specimen and sections: XY, XB, DF, MP, YYW and LL; Virus induced fusion: HP and PZ; S protein induced fusion and imaging: ZZ and ZN; Cell-in-cell experiments: YZ, YS, YCZ, MH; Gene cloning: BZ; Expression detection: CW; Data analysis: YCZ, MT, WG, XJ, HR, LG; Bioinformatics: QS; Figures: QS, HH and YCZ; Data interpretation: QS, HH; Manuscript: QS, GM, HH, XW, HS; Funding: QS, HH, YZ and ZC. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics statement
The autopsy for COVID-19 death was carried out with informed consent under the approval of Ethics Committee of Wuhan Infectious Diseases Hospital (KY-2020-15.01) and Ethics Committee of the First Affiliated Hospital of Army Medical University (KY2020298).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Y. Shi
Supplementary information
41418_2021_782_MOESM4_ESM.tif (download TIF )
Fig. S4 Subtyping T lymphocytes internalized into syncytia, and effects of candidate compounds on CIC formation. Related to Figure 3.
Rights and permissions
About this article
Cite this article
Zhang, Z., Zheng, Y., Niu, Z. et al. SARS-CoV-2 spike protein dictates syncytium-mediated lymphocyte elimination. Cell Death Differ 28, 2765–2777 (2021). https://doi.org/10.1038/s41418-021-00782-3
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41418-021-00782-3
This article is cited by
-
The Effect of Antibodies in the Presence of Syncytia During Viral Infections
Bulletin of Mathematical Biology (2025)
-
Homotypic cell-in-cell structure as a novel prognostic predictor in non-small cell lung cancer and frequently localized at the invasive front
Scientific Reports (2024)
-
A Comprehensive Analysis of NRP1 in Malignancies Provide Therapeutic Implication for Treating Cancer Patients Infected with SARS-CoV-2
Biochemical Genetics (2024)
-
A high-throughput screening system for SARS-CoV-2 entry inhibition, syncytia formation and cell toxicity
Biological Procedures Online (2023)
-
Ultrastructural study confirms the formation of single and heterotypic syncytial cells in bronchoalveolar fluids of COVID-19 patients
Virology Journal (2023)