Abstract
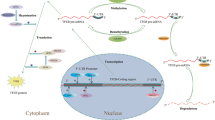
During autophagy, the coordinated actions of autophagosomes and lysosomes result in the controlled removal of damaged intracellular organelles and superfluous substrates. The evolutionary conservation of this process and its requirement for maintaining cellular homeostasis emphasizes the need to better dissect the pathways governing its molecular regulation. In our previously performed high-content screen, we assessed the effect of 1530 RNA-binding proteins on autophagy. Among the top regulators, we identified the eukaryotic translation initiation factor 4A-3 (eIF4A3). Here we show that depletion of eIF4A3 leads to a potent increase in autophagosome and lysosome biogenesis and an enhanced autophagic flux. This is mediated by the key autophagy transcription factor, TFEB, which becomes dephosphorylated and translocates from the cytoplasm to the nucleus where it elicits an integrated transcriptional response. We further identified an exon-skipping event in the transcript encoding for the direct TFEB kinase, GSK3B, which leads to a reduction in GSK3B expression and activity. Through analysis of TCGA data, we found a significant upregulation of eIF4A3 expression across several cancer types and confirmed the potential relevance of this newly identified signaling axis in human tumors. Hence, our data suggest a previously unrecognized role for eIF4A3 as a gatekeeper of autophagy through the control of TFEB activation, revealing a new mechanism for autophagy regulation.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Data availability
All datasets generated and analyzed in this study are deposited at the GitHub repository (https://github.com/ELELAB/eIF4A3_RNASEQ) or provided in the supplementary information files.
References
Mizushima N. A brief history of autophagy from cell biology to physiology and disease. Nat Cell Biol. 2018;20:521–7.
Levine B, Kroemer G. Biological functions of autophagy genes: a disease perspective. Cell. 2019;176:11–42.
Poillet-Perez L, White E. Role of tumor and host autophagy in cancer metabolism. Genes Dev. 2019;33:610–9.
Vanzo R, Bartkova J, Merchut-Maya JM, Hall A, Bouchal J, Dyrskjot L, et al. Autophagy role(s) in response to oncogenes and DNA replication stress. Cell Death Differ. 2019;27:1134–53.
Di Malta C, Cinque L, Settembre C. Transcriptional regulation of autophagy: mechanisms and diseases. Front Cell Dev Biol. 2019;7:114.
Fullgrabe J, Ghislat G, Cho DH, Rubinsztein DC. Transcriptional regulation of mammalian autophagy at a glance. J Cell Sci. 2016;129:3059–66.
Steingrimsson E, Copeland NG, Jenkins NA. Melanocytes and the microphthalmia transcription factor network. Annu Rev Genet. 2004;38:365–411.
Sardiello M, Palmieri M, di Ronza A, Medina DL, Valenza M, Gennarino VA, et al. A gene network regulating lysosomal biogenesis and function. Science. 2009;325:473–7.
Settembre C, Di Malta C, Polito VA, Garcia Arencibia M, Vetrini F, Erdin S, et al. TFEB links autophagy to lysosomal biogenesis. Science. 2011;332:1429–33.
Palmieri M, Impey S, Kang H, di Ronza A, Pelz C, Sardiello M, et al. Characterization of the CLEAR network reveals an integrated control of cellular clearance pathways. Hum Mol Genet. 2011;20:3852–66.
Puertollano R, Ferguson SM, Brugarolas J, Ballabio A. The complex relationship between TFEB transcription factor phosphorylation and subcellular localization. EMBO J. 2018;37:e98804.
Li Y, Xu M, Ding X, Yan C, Song Z, Chen L, et al. Protein kinase C controls lysosome biogenesis independently of mTORC1. Nat Cell Biol. 2016;18:1065–77.
Roczniak-Ferguson A, Petit CS, Froehlich F, Qian S, Ky J, Angarola B, et al. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci Signal. 2012;5:ra42.
Palmieri M, Pal R, Nelvagal HR, Lotfi P, Stinnett GR, Seymour ML, et al. mTORC1-independent TFEB activation via Akt inhibition promotes cellular clearance in neurodegenerative storage diseases. Nat Commun. 2017;8:14338.
Settembre C, Zoncu R, Medina DL, Vetrini F, Erdin S, Erdin S, et al. A lysosome-to-nucleus signalling mechanism senses and regulates the lysosome via mTOR and TFEB. EMBO J. 2012;31:1095–108.
Lubas M, Harder LM, Kumsta C, Tiessen I, Hansen M, Andersen JS, et al. eIF5A is required for autophagy by mediating ATG3 translation. EMBO Rep. 2018;19:e46072.
Linder P, Jankowsky E. From unwinding to clamping - the DEAD box RNA helicase family. Nat Rev Mol Cell Biol. 2011;12:505–16.
Chan CC, Dostie J, Diem MD, Feng W, Mann M, Rappsilber J, et al. eIF4A3 is a novel component of the exon junction complex. RNA. 2004;10:200–9.
Le Hir H, Gatfield D, Izaurralde E, Moore MJ. The exon-exon junction complex provides a binding platform for factors involved in mRNA export and nonsense-mediated mRNA decay. EMBO J. 2001;20:4987–97.
Wang Z, Murigneux V, Le Hir H. Transcriptome-wide modulation of splicing by the exon junction complex. Genome Biol. 2014;15:551.
Choe J, Ryu I, Park OH, Park J, Cho H, Yoo JS, et al. eIF4AIII enhances translation of nuclear cap-binding complex-bound mRNAs by promoting disruption of secondary structures in 5’UTR. Proc Natl Acad Sci USA. 2014;111:E4577–4586.
Nott A, Le Hir H, Moore MJ. Splicing enhances translation in mammalian cells: an additional function of the exon junction complex. Genes Dev. 2004;18:210–22.
Blazquez L, Emmett W, Faraway R, Pineda JMB, Bajew S, Gohr A, et al. Exon junction complex shapes the transcriptome by repressing recursive splicing. Mol Cell. 2018;72:496.e9–509.e9.
Boehm V, Britto-Borges T, Steckelberg AL, Singh KK, Gerbracht JV, Gueney E, et al. Exon junction complexes suppress spurious splice sites to safeguard transcriptome integrity. Mol Cell. 2018;72:482.e7–95.e7.
Michelle L, Cloutier A, Toutant J, Shkreta L, Thibault P, Durand M, et al. Proteins associated with the exon junction complex also control the alternative splicing of apoptotic regulators. Mol Cell Biol. 2012;32:954–67.
Mazloomian A, Araki S, Ohori M, El-Naggar AM, Yap D, Bashashati A, et al. Pharmacological systems analysis defines EIF4A3 functions in cell-cycle and RNA stress granule formation. Commun Biol. 2019;2:165.
Ryu I, Won YS, Ha H, Kim E, Park Y, Kim MK, et al. eIF4A3 phosphorylation by CDKs affects NMD during the cell cycle. Cell Rep. 2019;26:2126.e9–39.e9.
Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A, et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy. 2016;12:1–222.
Holdgaard SG, Cianfanelli V, Pupo E, Lambrughi M, Lubas M, Nielsen JC, et al. Selective autophagy maintains centrosome integrity and accurate mitosis by turnover of centriolar satellites. Nat Commun. 2019;10:4176.
Tiessen I, Abildgaard MH, Lubas M, Gylling HM, Steinhauer C, Pietras EJ, et al. A high-throughput screen identifies the long non-coding RNA DRAIC as a regulator of autophagy. Oncogene. 2019;38:5127–41.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN, et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci USA. 2014;111:E5593–5601.
Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science. 2002;298:1912–34.
Abildgaard MH, Brynjolfsdottir SH, Frankel LB. The Autophagy-RNA Interplay: Degradation and Beyond. Trends Biochem Sci. 2020;45:845–57.
Gatica D, Hu G, Liu X, Zhang N, Williamson PR, Klionsky DJ. The Pat1-Lsm complex stabilizes ATG mRNA during nitrogen starvation-induced autophagy. Mol Cell. 2019;73:314.e4–24.4.
Tang HW, Hu Y, Chen CL, Xia B, Zirin J, Yuan M, et al. The TORC1-regulated CPA complex rewires an RNA processing network to drive autophagy and metabolic reprogramming. Cell Metab. 2018;27:1040.e8–54.e8.
Xu G, Li T, Chen J, Li C, Zhao H, Yao C, et al. Autosomal dominant retinitis pigmentosa-associated gene PRPF8 is essential for hypoxia-induced mitophagy through regulating ULK1 mRNA splicing. Autophagy. 2018;14:1818–30.
Zhang H, Alsaleh G, Feltham J, Sun Y, Napolitano G, Riffelmacher T, et al. Polyamines control eIF5A hypusination, TFEB translation, and autophagy to reverse B cell senescence. Mol Cell. 2019;76:110.e9–25.e9.
Medina DL, Di Paola S, Peluso I, Armani A, De Stefani D, Venditti R, et al. Lysosomal calcium signalling regulates autophagy through calcineurin and TFEB. Nat Cell Biol. 2015;17:288–99.
Ito M, Iwatani M, Kamada Y, Sogabe S, Nakao S, Tanaka T, et al. Discovery of selective ATP-competitive eIF4A3 inhibitors. Bioorg Med Chem. 2017;25:2200–9.
Iwatani-Yoshihara M, Ito M, Ishibashi Y, Oki H, Tanaka T, Morishita D, et al. Discovery and characterization of a eukaryotic initiation factor 4A-3-selective inhibitor that suppresses nonsense-mediated mRNA decay. ACS Chem Biol. 2017;12:1760–8.
Mizojiri R, Nakata D, Satoh Y, Morishita D, Shibata S, Iwatani-Yoshihara M, et al. Discovery of novel 5-(piperazine-1-carbonyl)pyridin-2(1H)-one derivatives as orally eIF4A3-selective inhibitors. ACS Med Chem Lett. 2017;8:1077–82.
Ito M, Tanaka T, Cary DR, Iwatani-Yoshihara M, Kamada Y, Kawamoto T, et al. Discovery of novel 1,4-diacylpiperazines as selective and cell-active eIF4A3 inhibitors. J Med Chem. 2017;60:3335–51.
Nezich CL, Wang C, Fogel AI, Youle RJ. MiT/TFE transcription factors are activated during mitophagy downstream of Parkin and Atg5. J Cell Biol. 2015;210:435–50.
Settembre C, De Cegli R, Mansueto G, Saha PK, Vetrini F, Visvikis O, et al. TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat Cell Biol. 2013;15:647–58.
Decressac M, Mattsson B, Weikop P, Lundblad M, Jakobsson J, Bjorklund A. TFEB-mediated autophagy rescues midbrain dopamine neurons from alpha-synuclein toxicity. Proc Natl Acad Sci USA. 2013;110:E1817–26.
Tsunemi T, Ashe TD, Morrison BE, Soriano KR, Au J, Roque RA, et al. PGC-1alpha rescues Huntington’s disease proteotoxicity by preventing oxidative stress and promoting TFEB function. Sci Transl Med. 2012;4:142ra197.
Spampanato C, Feeney E, Li L, Cardone M, Lim JA, Annunziata F, et al. Transcription factor EB (TFEB) is a new therapeutic target for Pompe disease. EMBO Mol Med. 2013;5:691–706.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44:W90–97. W1
Mounir M, Lucchetta M, Silva TC, Olsen C, Bontempi G, Chen X, et al. New functionalities in the TCGAbiolinks package for the study and integration of cancer data from GDC and GTEx. PLoS Comput Biol. 2019;15:e1006701.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Acknowledgements
We thank Vanda Turcanova for assistance with cloning and Jernej Ule for helpful advice regarding the splicing analysis. We thank Valentina Cianfanelli for providing the CMV-Sigma14-TFEB-GFP plasmid and Andrea Ballabio for providing the TFEB phospho Serine 138 antibody. In addition, thanks to Anders H. Lund for constructive feedback and fruitful discussions regarding this work.
Funding
The work for this study was supported from The Lundbeck Foundation (R272-2017-3872), The Novo Nordisk Foundation (NNF19OC0056107), The Danish Cancer Society (R209-A13011), The Danish Council for Independent Research (DFF-7016-00313), and The Danish National Research Foundation (project CARD, DNRF 125).
Author information
Authors and Affiliations
Contributions
LBF developed the study concept; DS, THK, MHA, ARL, and LBF performed experiments; LB, MT, EP, and THK analyzed the sequencing and TCGA data; LBF, EP, and JB provided supervision; LBF, DS, and THK wrote the manuscript; all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
The study did not require any ethical approval.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by D. Rubinsztein
Supplementary information
Rights and permissions
About this article
Cite this article
Sakellariou, D., Tiberti, M., Kleiber, T.H. et al. eIF4A3 regulates the TFEB-mediated transcriptional response via GSK3B to control autophagy. Cell Death Differ 28, 3344–3356 (2021). https://doi.org/10.1038/s41418-021-00822-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41418-021-00822-y
This article is cited by
-
Post-transcriptional activation of TGF-β signaling via SMAD mRNA stabilization by EIF4A3 promotes EMT and metastasis in gastric cancer
Cell Biology and Toxicology (2026)
-
Post-transcriptional dynamics and RNA homeostasis in autophagy and cancer
Cell Death & Differentiation (2025)
-
PRKAA2, MTOR, and TFEB in the regulation of lysosomal damage response and autophagy
Journal of Molecular Medicine (2024)