Abstract
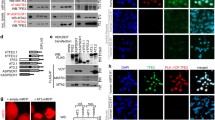
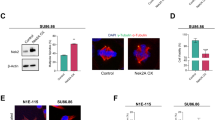
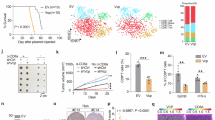
The proper orientation of centrosome and spindle is essential for genome stability; however, the mechanism that governs these processes remains elusive. Here, we demonstrated that polo-like kinase 1 (Plk1), a key mitotic kinase, phosphorylates residue Thr76 in VCP/p97 (an AAA-ATPase), at the centrosome from prophase to anaphase. This phosphorylation process recruits VCP to the centrosome and in this way, it regulates centrosome orientation. VCP exhibits strong co-localization with Eg5 (a mitotic kinesin motor), at the mitotic spindle, and the dephosphorylation of Thr76 in VCP is required for the enrichment of both VCP and Eg5 at the spindle, thus ensuring proper spindle architecture and chromosome segregation. We also showed that the phosphatase, PTEN, is responsible for the dephosphorylation of Thr76 in VCP; when PTEN was knocked down, the normal spread of VCP from the centrosome to the spindle was abolished. Cryo-EM structures of VCPT76A and VCPT76E, which represent dephosphorylated and phosphorylated states of VCP, respectively, revealed that the Thr76 phosphorylation modulates VCP by altering the inter-domain and inter-subunit interactions, and ultimately the nucleotide-binding pocket conformation. Interestingly, the tumor growth in nude mice implanted with VCPT76A-reconstituted cancer cells was significantly slower when compared with those implanted with VCPWT-reconstituted cancer cells. Collectively, our findings demonstrate that the phosphorylation and dephosphorylation switch of VCP regulates the architecture of centrosome and spindle for faithful chromosome segregation.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Data availability
Data supporting the present study are available from the corresponding author upon reasonable request.
References
Tanenbaum ME, Medema RH. Mechanisms of centrosome separation and bipolar spindle assembly. Dev Cell. 2010;19:797–806.
Nigg EA, Stearns T. The centrosome cycle: centriole biogenesis, duplication and inherent asymmetries. Nat Cell Biol. 2011;13:1154–60.
Nilsson J. Protein phosphatases in the regulation of mitosis. J Cell Biol. 2019;218:395–409.
Gelens L, Qian J, Bollen M, Saurin AT. The importance of kinase-phosphatase integration: lessons from mitosis. Trends Cell Biol. 2018;28:6–21.
Dominguez-Brauer C, Thu KL, Mason JM, Blaser H, Bray MR, Mak TW. Targeting mitosis in cancer: emerging strategies. Mol Cell. 2015;60:524–36.
Archambault V, Glover DM. Polo-like kinases: conservation and divergence in their functions and regulation. Nat Rev Mol Cell Biol. 2009;10:265–75.
Combes G, Alharbi I, Braga LG, Elowe S. Playing polo during mitosis: PLK1 takes the lead. Oncogene. 2017;36:4819–27.
Strebhardt K. Multifaceted polo-like kinases: drug targets and antitargets for cancer therapy. Nat Rev Drug Disco. 2010;9:643–60.
Gutteridge REA, Ndiaye MA, Liu X, Ahmad N. Plk1 inhibitors in cancer therapy: from laboratory to clinics. Mol Cancer Ther. 2016;15:1427–35.
Mann BJ, Wadsworth P. Kinesin-5 regulation and function in mitosis. Trends Cell Biol. 2019;29:66–79.
Bertran MT, Sdelci S, Regue L, Avruch J, Caelles C, Roig J. Nek9 is a Plk1-activated kinase that controls early centrosome separation through Nek6/7 and Eg5. EMBO J. 2011;30:2634–47.
Blangy A, Lane HA, d’Hérin P, Harper M, Kress M, Niggt EA. Phosphorylation by p34cdc2 regulates spindle association of human Eg5, a kinesin-related motor essential for bipolar spindle formation in vivo. Cell. 1995;83:1159–69.
Sawin KE, Mitchison TJ. Mutations in the kinesin-like protein Eg5 disrupting localization to the mitotic spindle. Proc Natl Acad Sci USA 1995;92:4289–93.
Sawin KE, LeGuellec K, Philippe M, Mitchison TJ. Mitotic spindle organization by a plus-end-directed microtubule motor. Nature. 1992;359:540–3.
He J, Zhang Z, Ouyang M, Yang F, Hao H, Lamb KL, et al. PTEN regulates EG5 to control spindle architecture and chromosome congression during mitosis. Nat Commun. 2016;7:12355.
van Ree JH, Nam HJ, Jeganathan KB, Kanakkanthara A, van Deursen JM. Pten regulates spindle pole movement through Dlg1-mediated recruitment of Eg5 to centrosomes. Nat Cell Biol. 2016;18:814–21.
Muretta JM, Reddy BJN, Scarabelli G, Thompson AF, Jariwala S, Major J, et al. A posttranslational modification of the mitotic kinesin Eg5 that enhances its mechanochemical coupling and alters its mitotic function. Proc Natl Acad Sci USA. 2018;115:E1779–88.
van den Boom J, Meyer H. VCP/p97-mediated unfolding as a principle in protein homeostasis and signaling. Mol Cell. 2018;69:182–94.
Stach L, Freemont PS. The AAA+ ATPase p97, a cellular multitool. Biochem J. 2017;474:2953–76.
Bodnar N, Rapoport T. Toward an understanding of the Cdc48/p97 ATPase. F1000Res. 2017;6:1318.
Deng L, Wu RA, Sonneville R, Kochenova OV, Labib K, Pellman D, et al. Mitotic CDK promotes replisome disassembly, fork breakage, and complex DNA rearrangements. Mol Cell. 2019;73:915–29 e6.
Franz A, Orth M, Pirson PA, Sonneville R, Blow JJ, Gartner A, et al. CDC-48/p97 coordinates CDT-1 degradation with GINS chromatin dissociation to ensure faithful DNA replication. Mol Cell. 2011;44:85–96.
Wu RA, Semlow DR, Kamimae-Lanning AN, Kochenova OV, Chistol G, Hodskinson MR, et al. TRAIP is a master regulator of DNA interstrand crosslink repair. Nature. 2019;567:267–72.
Shi X, Zhu K, Ye Z, Yue J. VCP/p97 targets the nuclear export and degradation of p27(Kip1) during G1 to S phase transition. FASEB J. 2020.34:5193–207.
Raman M, Havens CG, Walter JC, Harper JW. A genome-wide screen identifies p97 as an essential regulator of DNA damage-dependent CDT1 destruction. Mol Cell. 2011;44:72–84.
Meerang M, Ritz D, Paliwal S, Garajova Z, Bosshard M, Mailand N, et al. The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks. Nat Cell Biol. 2011;13:1376–82.
Ramadan K, Bruderer R, Spiga FM, Popp O, Baur T, Gotta M, et al. Cdc48/p97 promotes reformation of the nucleus by extracting the kinase Aurora B from chromatin. Nature. 2007;450:1258–62.
Olmos Y, Hodgson L, Mantell J, Verkade P, Carlton JG. ESCRT-III controls nuclear envelope reformation. Nature. 2015;522:236–9.
Cao K, Nakajima R, Meyer HH, Zheng Y. The AAA-ATPase Cdc48/p97 regulates spindle disassembly at the end of mitosis. Cell. 2003;115:355–67.
Kress E, Schwager F, Holtackers R, Seiler J, Prodon F, Zanin E, et al. The UBXN-2/p37/p47 adaptors of CDC-48/p97 regulate mitosis by limiting the centrosomal recruitment of Aurora A. J Cell Biol. 2013;201:559–75.
Yu CC, Yang JC, Chang YC, Chuang JG, Lin CW, Wu MS, et al. VCP phosphorylation-dependent interaction partners prevent apoptosis in Helicobacter pylori-infected gastric epithelial cells. PLoS ONE. 2013;8:e55724.
Elia AE, Cantley LC, Yaffe MB. Proteomic screen finds pSer/pThr-binding domain localizing Plk1 to mitotic substrates. Science. 2003;299:1228–31.
Elia AE, Rellos P, Haire LF, Chao JW, Ivins FJ, Hoepker K, et al. The molecular basis for phosphodependent substrate targeting and regulation of Plks by the Polo-box domain. Cell. 2003;115:83–95.
Soderberg O, Gullberg M, Jarvius M, Ridderstrale K, Leuchowius KJ, Jarvius J, et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat Methods. 2006;3:995–1000.
Shyu YJ, Hiatt SM, Duren HM, Ellis RE, Kerppola TK, Hu CD. Visualization of protein interactions in living Caenorhabditis elegans using bimolecular fluorescence complementation analysis. Nat Protoc. 2008;3:588–96.
Nakajima H, Toyoshima-Morimoto F, Taniguchi E, Nishida E. Identification of a consensus motif for Plk (Polo-like kinase) phosphorylation reveals Myt1 as a Plk1 substrate. J Biol Chem. 2003;278:25277–80.
Peters JM, Walsh MJ, Franke WW. An abundant and ubiquitous homo‐oligomeric ring‐shaped ATPase particle related to the putative vesicle fusion proteins Sec18p and NSF. EMBO J. 1990;9:1757–67.
Macůrek L, Lindqvist A, Lim D, Lampson MA, Klompmaker R, Freire R, et al. Polo-like kinase-1 is activated by aurora A to promote checkpoint recovery. Nature. 2008;455:119–23.
Seki A, Coppinger JA, Jang C-Y, Yates JR, Fang G. Bora and the kinase Aurora a cooperatively activate the kinase Plk1 and control mitotic entry. Science. 2008;320:1655–8.
Lee K, Rhee K. PLK1 phosphorylation of pericentrin initiates centrosome maturation at the onset of mitosis. J Cell Biol. 2011;195:1093–101.
van Vugt MA, van de Weerdt BC, Vader G, Janssen H, Calafat J, Klompmaker R, et al. Polo-like kinase-1 is required for bipolar spindle formation but is dispensable for anaphase promoting complex/Cdc20 activation and initiation of cytokinesis. J Biol Chem. 2004;279:36841–54.
Liu X, Erikson RL. Activation of Cdc2/cyclin B and inhibition of centrosome amplification in cells depleted of Plk1 by siRNA. Proc Natl Acad Sci USA. 2002;99:8672–6.
Li Y, Benezra R. Identification of a human mitotic checkpoint gene: hsMAD2. Science. 1996;274:246–8.
Kim DI, Jensen SC, Noble KA, Kc B, Roux KH, Motamedchaboki K, et al. An improved smaller biotin ligase for BioID proximity labeling. Mol Biol Cell. 2016;27:1188–96.
Anderson DJ, Le Moigne R, Djakovic S, Kumar B, Rice J, Wong S, et al. Targeting the AAA ATPase p97 as an approach to treat cancer through disruption of protein homeostasis. Cancer Cell. 2015;28:653–65.
Erdodi F, Rokolya A, Di Salvo J, Barany M, Barany K. Effect of okadaic acid on phosphorylation-dephosphorylation of myosin light chain in aortic smooth muscle homogenate. Biochem Biophys Res Commun. 1988;153:156–61.
Zhang XC, Piccini A, Myers MP, Van Aelst L, Tonks NK. Functional analysis of the protein phosphatase activity of PTEN. Biochem J. 2012;444:457–64.
Banerjee S, Bartesaghi A, Merk A, Rao P, Bulfer SL, Yan Y, et al. 2.3 A resolution cryo-EM structure of human p97 and mechanism of allosteric inhibition. Science. 2016;351:871–5.
Tang WK, Li D, Li CC, Esser L, Dai R, Guo L, et al. A novel ATP-dependent conformation in p97 N-D1 fragment revealed by crystal structures of disease-related mutants. EMBO J. 2010;29:2217–29.
Tang WK, Xia D. Altered intersubunit communication is the molecular basis for functional defects of pathogenic p97 mutants. J Biol Chem. 2013;288:36624–35.
Liu-Chittenden Y, Jain M, Gaskins K, Wang S, Merino MJ, Kotian S, et al. RARRES2 functions as a tumor suppressor by promoting beta-catenin phosphorylation/degradation and inhibiting p38 phosphorylation in adrenocortical carcinoma. Oncogene. 2017;36:3541–52.
Kim JY, Xue K, Cao M, Wang Q, Liu JY, Leader A, et al. Chemerin suppresses ovarian follicular development and its potential involvement in follicular arrest in rats treated chronically with dihydrotestosterone. Endocrinology. 2013;154:2912–23.
Zhang H, Wang Y, Liu Z, Yao B, Dou C, Xu M, et al. Lymphocyte-specific protein 1 inhibits the growth of hepatocellular carcinoma by suppressing ERK1/2 phosphorylation. FEBS Open Bio. 2016;6:1227–37.
Irigoyen M, Pajares MJ, Agorreta J, Ponz-Sarvise M, Salvo E, Lozano MD, et al. TGFBI expression is associated with a better response to chemotherapy in NSCLC. Mol Cancer. 2010;9:130.
Smith CC, Lee KS, Li B, Laing JM, Hersl J, Shvartsbeyn M, et al. Restored expression of the atypical heat shock protein H11/HspB8 inhibits the growth of genetically diverse melanoma tumors through activation of novel TAK1-dependent death pathways. Cell Death Dis. 2012;3:e371.
Lewis AM, Varghese S, Xu H, Alexander HR. Interleukin-1 and cancer progression: the emerging role of interleukin-1 receptor antagonist as a novel therapeutic agent in cancer treatment. J Transl Med. 2006;4:48.
Alfaro C, Sanmamed MF, Rodríguez-Ruiz ME, Teijeira Á, Oñate C, González Á, et al. Interleukin-8 in cancer pathogenesis, treatment and follow-up. Cancer Treat Rev. 2017;60:24–31.
Masjedi A, Hashemi V, Hojjat-Farsangi M, Ghalamfarsa G, Azizi G, Yousefi M, et al. The significant role of interleukin-6 and its signaling pathway in the immunopathogenesis and treatment of breast cancer. Biomed Pharmacother. 2018;108:1415–24.
Kimonis VE, Fulchiero E, Vesa J, Watts G. VCP disease associated with myopathy, Paget disease of bone and frontotemporal dementia: review of a unique disorder. Biochim Biophys Acta. 2008;1782:744–8.
Dobrynin G, Popp O, Romer T, Bremer S, Schmitz MH, Gerlich DW, et al. Cdc48/p97–Ufd1–Npl4 antagonizes Aurora B during chromosome segregation in HeLa cells. J Cell Sci. 2011;124:1571–80.
Xu J, Shen C, Wang T, Quan J. Structural basis for the inhibition of Polo-like kinase 1. Nat Struct Mol Biol. 2013;20:1047.
Lowery DM, Lim D, Yaffe MB. Structure and function of Polo-like kinases. Oncogene. 2005;24:248–59.
Golan A, Yudkovsky Y, Hershko A. The cyclin-ubiquitin ligase activity of cyclosome/APC is jointly activated by protein kinases Cdk1-cyclin B and Plk. J Biol Chem. 2002;277:15552–7.
May KM, Reynolds N, Cullen CF, Yanagida M, Ohkura H. Polo boxes and Cut23 (Apc8) mediate an interaction between polo kinase and the anaphase-promoting complex for fission yeast mitosis. J Cell Biol. 2002;156:23–8.
Liu X, Zhou T, Kuriyama R, Erikson RL. Molecular interactions of Polo-like-kinase 1 with the mitotic kinesin-like protein CHO1/MKLP-1. J Cell Sci. 2004;117:3233–46.
Neef RD, Preisinger C, Sutcliffe J, Kopajtich R, Nigg EA, Mayer TU, et al. Phosphorylation of mitotic kinesin-like protein 2 by polo-like kinase 1 is required for cytokinesis. J Cell Biol. 2003;162:863–76.
Zhang Z, Hou SQ, He J, Gu T, Yin Y, Shen WH. PTEN regulates PLK1 and controls chromosomal stability during cell division. Cell Cycle. 2016;15:2476–85.
Tibarewal P, Zilidis G, Spinelli L, Schurch N, Maccario H, Gray A, et al. PTEN protein phosphatase activity correlates with control of gene expression and invasion, a tumor-suppressing phenotype, but not with AKT activity. Sci Signal. 2012;5:ra18.
Tamura M, Gu J, Matsumoto K, Aota S, Parsons R, Yamada KM. Inhibition of cell migration, spreading, and focal adhesions by tumor suppressor PTEN. Science. 1998;280:1614–7.
Gu T, Zhang Z, Wang J, Guo J, Shen WH, Yin Y. CREB is a novel nuclear target of PTEN phosphatase. Cancer Res. 2011;71:2821–5.
Zhang S, Huang WC, Li P, Guo H, Poh SB, Brady SW, et al. Combating trastuzumab resistance by targeting SRC, a common node downstream of multiple resistance pathways. Nat Med. 2011;17:461–9.
Shi Y, Wang J, Chandarlapaty S, Cross J, Thompson C, Rosen N, et al. PTEN is a protein tyrosine phosphatase for IRS1. Nat Struct Mol Biol. 2014;21:522–7.
Ben-David U, Amon A. Context is everything: aneuploidy in cancer. Nat Rev Genet. 2020;21:44–62.
Gordon DJ, Resio B, Pellman D. Causes and consequences of aneuploidy in cancer. Nat Rev Genet. 2012;13:189–203.
Sheltzer JM, Ko JH, Replogle JM, Habibe Burgos NC, Chung ES, Meehl CM, et al. Single-chromosome gains commonly function as tumor suppressors. Cancer Cell. 2017;31:240–55.
Wei W-J, Sun H-Y, Ting KY, Zhang L-H, Lee H-C, Li G-R, et al. Inhibition of cardiomyocytes differentiation of mouse embryonic stem cells by CD38/cADPR/Ca2+ signaling pathway. J Biol Chem. 2012;287:35599–611.
Wang Q, Guo W, Hao B, Shi X, Lu Y, Wong CW, et al. Mechanistic study of TRPM2-Ca2+-CAMK2-BECN1 signaling in oxidative stress-induced autophagy inhibition. Autophagy. 2016;12:1340–54.
Chin YR, Yuan X, Balk SP, Toker A. PTEN-deficient tumors depend on AKT2 for maintenance and survival. Cancer discovery. 2014;4:942–55.
Dunn KW, Kamocka MM, McDonald JH. A practical guide to evaluating colocalization in biological microscopy. Am J Physiol-Cell Physiol. 2011;300:C723–42.
Acknowledgements
We thank members of Yue laboratory for advice on the manuscript. We would like to thank the Cryo-EM center of Southern University of Science and Technology for Cryo-EM data collection and the HPC-Service Station in Cryo-EM center of Southern University of Science and Technology for data processing.
Funding
This work was supported by Hong Kong Research Grant Council grants (11101717 and 11103620 to JY), Ministry of Science and Technology of the People’s Republic of China (2019YFA0906000 to HZ), NSFC (21778045, 32070702, and 82161128014 to JY), ITF (MRP/064/21 and GHP/097/20GD to JY), Guangdong Science and Technology Program (2017B030301018 to HZ), Shenzhen Science and Technology Innovation Committee (SGDX20201103093201010, JSGG20200225150702770, and JCYJ20210324134007020 to JY, and ZDSYS20140509142721429 to HZ).
Author information
Authors and Affiliations
Contributions
KZ, SL, HZ, and JY designed the experiments; KZ, YC, XS, ZM, HZ, ZY, RW and JY performed the experiments; KZ, YC, ZY, YG, CL, LZ, SL, HZ, and JY analyzed the data; and KZ, YC, HZ and JY wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All animal experiments were performed following the university laboratory animal guidelines and with approval from the Animal Committee of City University of Hong Kong. Pathological sections of lung cancer patients were obtained from the Pathology Department of the Second Affiliated Hospital of Chongqing Medical University, with informed consent and approval from the Clinical Research Ethics Committee of the Second Affiliated Hospital of Chongqing Medical University. The patients were provided and signed the consent forms.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Piacentini
Supplementary information
Rights and permissions
About this article
Cite this article
Zhu, K., Cai, Y., Si, X. et al. The phosphorylation and dephosphorylation switch of VCP/p97 regulates the architecture of centrosome and spindle. Cell Death Differ 29, 2070–2088 (2022). https://doi.org/10.1038/s41418-022-01000-4
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41418-022-01000-4
This article is cited by
-
The nuclear exosome co-factor MTR4 shapes the transcriptome for meiotic initiation
Nature Communications (2025)
-
DUSP1 interacts with and dephosphorylates VCP to improve mitochondrial quality control against endotoxemia-induced myocardial dysfunction
Cellular and Molecular Life Sciences (2023)
-
Vacuolin-1 enhances RA-induced differentiation of human myeloblastic leukemia cells: evidence for involvement of a CD11b/FAK/LYN/SLP-76 axis subject to endosomal regulation that drives late differentiation steps
Cell & Bioscience (2022)