Abstract
The scaffolding protein NEDD9 coordinates signaling downstream of integrins by interacting with focal adhesion kinase (FAK) and thereby promotes cell migration. NEDD9 expression is altered in a number of clinical conditions such as cancer, but its role in innate immunity against infections remains elusive. Transcriptome analysis of Salmonella Typhimurium (ST)-infected murine macrophages showed downregulation of NEDD9 and genes belonging to its signaling network. Bacterial infections induced host-mediated lysosomal degradation of NEDD9 in macrophages and PBMCs isolated from patients suffering from bloodstream infection. However, ST induced translocation of NEDD9 from the cytoplasm to ST-containing phagosomes and prevented their phagolysosome-mediated clearance by FAK/AKT activation, reflecting a bacterial evasion mechanism. Complete loss of NEDD9 significantly reduced bacterial burden and enhanced inflammation upon ST infection both in vitro and in vivo. Mechanistically, we show that NEDD9 activates the FAK-AKT pathway allowing phosphorylation of FAK and AKT to impair phagolysosomal-mediated clearance of bacteria. Our study has thus identified NEDD9 as a critical regulator of lysosomal function in macrophages and a potential host-directed therapeutic target to treat bacterial infections.
Classification: Biological Sciences, Microbiology

Macrophages downregulate NEDD9 to counteract ST mediated FAK-AKT activation. Upon infection with ST NEDD9 is translocated from the cytosol to ST-containing phagosomes. Loss of NEDD9 results in enhanced lysosomal capacities supporting bacterial clearance. Strikingly, ST recruits and activates FAK and AKT to suppress endosome-lysosome fusion, thereby bypassing lysosome-mediated pathogen clearance. Created in BioRender. Robinson, N. (2021) BioRender.com/n17r483.
Similar content being viewed by others
Introduction
Bloodstream infections (BSI) caused by bacteria often lead to severe clinical disorders in patients, potentially resulting in life-threating conditions such as sepsis, septic shock, and organ dysfunction [1, 2]. Sepsis remains the cause of approximately six million deaths worldwide, and is frequently underdiagnosed at an early stage when it is still reversible [3]. Moreover, the global emergence of multidrug-resistant pathogens is causing a rise in antibiotic-resistant infections, contributing to high morbidity and mortality of septic patients [4]. Thus, the WHO recently identified management of sepsis as a global health priority in order to improve prevention, diagnosis and clinical management [3]. Better understanding of the molecular mechanisms of BSI to establish alternative therapeutic strategies is essential for better management of bacterially-induced sepsis.
Macrophages are key cells that mediate an effective innate response against pathogens in the bloodstream. Invading pathogens are detected by pattern recognition receptors and degraded within the phagosomes after fusion with lysosomes [5]. Pro-inflammatory responses against pathogens often involve enhanced phagosome processing. Conversely, intracellular pathogens such as Salmonella can impair phagolysosomal degradation as a primary strategy to evade degradation by macrophages and reduce production of immunogenic peptides [6,7,8,9]. For example, the gastroenteritis causing non-typhoid Salmonella (NTS) strain Salmonella enterica serovar Typhimurium (S. Typhimurium, subsequently referred to as ST) actively suppresses phagolysosomal degradation by recruiting focal adhesion kinase (FAK) to ST-containing vacuoles [10]. Importantly, ST actively invades the bloodstream, particularly in immune-compromised patients, which can cause life-threatening conditions [11]. Recently, the numbers of invasive NTS associated infections have been on the rise, not only in low- and middle- income countries but also in high-income regions such as Australia [12] and Europe [13]. One reason is that treating infections caused by antibiotic-resistant Salmonella isolates is increasingly challenging due to a lack of new antibiotics.
NEDD9 (Neural Precursor Cell Expressed, Developmentally Down-Regulated 9) - also known as CAS-L (CRK-associated substrate-related protein) or HEF1 (Human enhancer of filamentation 1) - is a member of the CAS family. The CAS family is a group of non-catalytic scaffolding proteins regulating normal and pathological cellular signaling, thus modifying cell proliferation, adhesion, migration, and differentiation. They are characterized by protein interaction domains, including Src homology 3 (SH3) domain and a substrate domain, which allow them to interact with a wide range of signaling molecules and cytoskeletal components [14]. NEDD9 is frequently overexpressed and predictive for poor clinical outcome in numerous types of tumors [15,16,17]. Furthermore, NEDD9 enhances immune cell functions including chemokine-induced lymphocyte migration [18,19,20,21] and inflammatory responses in rheumatoid arthritis [22]. Upon activation of various cell surface receptors - as best described for integrins - NEDD9 forms complexes with FAK and other kinases to modify intracellular signaling pathways, promoting cell proliferation, migration, cytoskeletal remodeling and adhesion [16, 23, 24]. Nedd9 knockout mice are both viable and fertile, without a shortened lifespan. Nonetheless, differences in the distribution and quantity of immune cells have been observed in these mice, including an increased number of macrophages in the peripheral blood and spleen [16, 21]. However, whether NEDD9 plays a role in general or specifically during antibacterial host defenses has not yet been investigated.
In this study, we demonstrate a novel, functional role of NEDD9 in macrophage defense against bacterial pathogens. Initially, we identified NEDD9 and known NEDD9 interaction partners being significantly downregulated in murine macrophages upon ST infection. We analyzed murine bone-marrow derived macrophages (mBMDMs) from Nedd9wt/wt and Nedd9-/- mice as well as human macrophages, and found NEDD9 to be significantly and consistently downregulated and degraded by lysosomes upon infection with various bacteria, including ST. Genetic loss of NEDD9 strongly improved both bacterial clearance and secretion of pro-inflammatory cytokines, in vitro and in vivo. Furthermore, ST infection induced translocation of NEDD9 to ST-containing phagosomes, leading to activation of FAK-AKT. Conversely, loss of NEDD9 inhibited FAK-AKT-activation and enhanced phagolysosomal clearance of ST. Overall, our data suggest that downregulation of NEDD9 is a critical host defense mechanism of macrophages against bacterial infections, which is bypassed by ST.
Results
NEDD9 is downregulated in murine macrophages upon bacterial infection
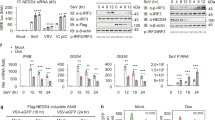
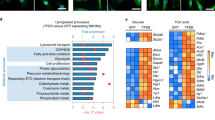
To assess the importance of NEDD9 in innate immune responses to bacterial infections, we reanalyzed a previously performed transcriptomics data analysis of wildtype mBMDMs 2 hours post ST infection and compared them to uninfected control cells [25]. We found that Nedd9 mRNA was significantly downregulated (approx. 95%). Nedd9 mRNA downregulation was accompanied by significant downregulation of known Nedd9 direct interactors such as Fak (also known as Ptk2), Ptk2b and Crkl as well as crucial Nedd9 signaling partners including Mapk1 and Dock1 (Fig. 1A, B) [16, 17, 26, 27]. Nedd9 downregulation could be confirmed in qRT-PCR from macrophage lysates infected with ST at 4 h timepoint (Fig. 1C). These results suggest a role for NEDD9 in host cell autonomous immune responses towards bacterial infection. In order to further elucidate the role of NEDD9 upon bacterial infection of macrophages, we infected mBMDMs with ST and analyzed NEDD9 protein levels by Western Blot analysis and immunofluorescence microscopy at timpoints 0.25, 0.5, 1, 4 and 24 hours. Consistent with our transcriptome and RNA data, we found NEDD9 significantly decreased at 4 hours post ST infection (Fig. 1D–F, Supplementary Fig. S1A) and almost completely depleted at 24 hours post infection (Fig. 1D,E).
A Heat map of microarray analysis of murine bone marrow-derived macrophages (mBMDMs) infected with S. Typhimurium (ST) for 2 hours compared to uninfected control macrophages (UI). Gene expression of UI and infected mBMDMs as arbitrary unit (AU), red for high expression, blue for low expression. N = 3, biological replicates. B STRING Analysis of functional network of the NEDD9 signaling axis for significantly altered gene expression of the analysis described in (A). Genes are highlighted by coloring them according to their fold-change in expression. C Real-time PCR analysis of Nedd9 levels normalized to GAPDH by calculation of Δ CT values from mBMDMs infected with ST, unpaired t-test: p (UI vs. 4 h) = 0.0424, experiment n = 3 with at least 3 technical replicates. D Western blot analysis of mBMDMs uninfected (UI) and infected with ST at indicated time points followed by (E) densitometric analysis of NEDD9 protein level normalized to GAPDH, n = 5, biological replicates, one-way ANOVA: p (UI vs. 1 h) = 0.0162, p (UI vs. 4 h) = 0.0001, p (UI vs. 24 h) < 0.0001. F Quantification of immunofluorescent staining of intrazellular NEDD9 protein, UI n = 9, ST 4 h n = 11, biological replicates, unpaired t-test: p = 0.0019. A p-value > 0.05 equals not significant (ns), *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
Induction of macrophage defense mechanisms often depends on the degree of pathogen-mediated virulence [7]. To determine if these factors influenced control of NEDD9 expression, we next analyzed NEDD9 protein levels upon infection of mBMDM with ST Salmonella Pathogenicity Island SPI1 (ΔinvA) and SPI2 (ΔssaV) mutants, which have defects in the main type III secretion system (T3SS). NEDD9 levels were diminished despite the lack of SPI1 and SPI2 genes, suggesting that the decline in NEDD9 was independent of ST-specific virulence factors (Supplementary Fig. S1B). Next, we investigated if decreased NEDD9 is specific to ST. Therefore, we treated macrophages with ST derived LPS, which is a TLR-4 agonist as well as with Pam3CSK activating TLR-2. Interestingly, LPS alone was sufficient to completely decrease NEDD9 levels (Supplementary Fig. S1C) indicating that Nedd9 downregulation is specific for Gram-negative pathogens. This finding was further supported by western blot analysis of NEDD9 in response to Staphylococcus aureus (SA) and Shigella flexneri (SF) infection showing reduced NEDD9 levels only for the Gram-negative pathogen SF (Supplementary Fig. S1D–F).
Taken together, these data indicate that ST infection dynamically downregulates NEDD9 at both the mRNA and protein levels in a virulence independent manner.
NEDD9 translocates from cytoplasm to the phagolysosomal complex and inhibits lysosomal degradation upon ST infection
To understand the mechanism of NEDD9 downregulation in macrophages during bacterial infection, we analyzed whether the depletion of NEDD9 protein occurs by proteasomal degradation, as previously reported for other stimuli [28, 29]. To this end, we infected mBMDMs with ST and treated them with the proteasome inhibitor MG132 followed by Western blotting. However, proteasome inhibition did not prevent NEDD9 degradation (Supplementary Fig. S2A), implying a different degradation mechanism upon infection.
NEDD9 activity is often determined by relocalization from the cytosol to sites of defined cellular processes such as the mitotic spindle during mitosis [30,31,32] or the basal body in the process of ciliary disassembly [33]. In addition, proteins can also be degraded via the lysosomal pathway. Macrophages mainly eliminate invading pathogens by activating the phagolysosomal machinery. In our previous studies we have shown that host proteins such as SIRT-1 and AMPK traffic into lysosomes for degradation during host-pathogen interactions [7].
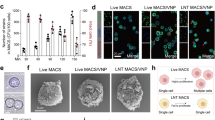
To assess whether NEDD9 subcellular localization is altered upon infection, we performed immunofluorescence staining of ST infected Nedd9wt/wt and Nedd9-/- mBMDMs as well as human macrophages derived from PBMCs of healthy donors. Untreated macrophages showed diffuse cytoplasmic expression of NEDD9. In contrast, NEDD9 clustered and colocalized with ST at 4 hours post infection (Fig. 2A) and with lysosomal-associated membrane protein 1 (LAMP-1) (Fig. 2B, Supplementary Fig. S2B, C). This suggests that ST infection triggers NEDD9 trafficking from the cytoplasm to the phagolysosome. Consistently, we detected NEDD9 in lysates of isolated ST-containing phagosomes 4 h post infection of Nedd9wt/wt mBMDMs (Fig. 2C). Notably, treatment with vacuolar ATPase inhibitor Concanamycin A significantly stabilized NEDD9 levels upon ST infection in mBMDMs (Fig. 2D, E). Interestingly, immunofluorescence staining using LysoTracker to track acidic organelles within macrophages upon ST infection revealed, besides colocalization with NEDD9 (Fig. 2A) a significantly increased intensity of LysoTracker in Nedd9-/- mBMDMs compared to Nedd9wt/wt mBMDMs indicating elevated lysosomal activity in the absence of NEDD9 (Fig. 2F, G). This data was further supported by a significant increase in lysosomal ß-galactosidase activity measured on C12FDG coated ST indicative of enhanced phagolysosomal processing (Fig. 2H). Collectively, these data suggest that in macrophages NEDD9 is degraded by the lysosomal machinery upon ST infection. Furthermore, it indicates that downregulation of NEDD9 facilitates phagolysosomal activity in macrophages upon bacterial infection.
A Representative confocal microscopy images of human macrophages uninfected (UI) and infected with ST for 4 hours and stained for ST (green) and NEDD9 (red). B Analysis of colocalization of NEDD9 with LAMP1 using Mander’s colocalization coefficient, n = 10, biological replicates, unpaired t-test: p = 0.0323. C Western blots of isolated phagosomes from Nedd9wt/wt and Nedd9-/- mBMDMs 4 h post ST infection and blotted for NEDD9, phagolysosomal marker Cathepsin D (Cath. D) and cytosolic marker GAPDH, n = 2, biological replicates. D Infection of mBMDMs with ST and parallel treatment with 100 nM concanamycin A (conc. A) followed by Western blotting and (E) densitometric analysis of NEDD9 protein level normalized to GAPDH and compared to uninfected cells (UI), n = 3, biological replicates, 2-way ANOVA: p (untreated UI vs. 4 h) = 0.0235, p (untreated UI vs. 24 h) = 0.0012, p (untreated UI vs. concanamycin A UI) = 0.0227, p (untreated 24 h vs. concanamycin A 24 h) = 0.0049. F Representative confocal microscopy images of Nedd9wt/wt and Nedd9-/- mBMDMs infected with ST for 4 hours and stained for ST (blue), NEDD9 (green) and lysotracker (red) and (G) integrated density analysis of LysoTracker. Nedd9wt/wt n = 8, Nedd9-/- n = 11, biological replicates, unpaired t-test: p = 0.0045. H Measurement of ß-Galactosidase activity by release of C12FDG in ST infected Nedd9wt/wt and Nedd9-/- mBMDMs treated with chloroquine (chloro) or non-treated (n.t.), n = 4, biological replicates, 2-way ANOVA: p (Nedd9wt/wt and Nedd9-/- ST 200 min.) = 0.0030, p (Nedd9-/- UT vs. Chloroquin ST 200 min.) < 0.0001, p (Nedd9wt/wt UT vs. Chloroquin ST 200 min.) = 0.042. *p ≤ 0.05, **p ≤ 0.01.
Nedd9 depletion enhances the inflammatory response of bacteria-infected macrophages and improves their bacterial clearance ex vivo
Next, we investigated how loss of Nedd9 regulates macrophage defense mechanisms upon ST infection. For this purpose, mBMDMs isolated from Nedd9wt/wt and Nedd9-/- mice were infected with ST and their inflammatory response was quantified by ELISA. Here, we found significantly higher expression of the pro-inflammatory cytokines IL-6 and TNF-α in response to ST infection in cell culture supernatants of Nedd9-deficient mBMDMs measured by ELISA (Fig. 3A, B). Interestingly, Nedd9-/- macrophages showed decreased NF-κB and p38 phosphorylation in Western blot analysis suggesting that NF-κb and p38 are regulated by other mechanisms, which are independent of the enhanced pro-inflammatory responses upon loss of Nedd9 (Fig. 3C–F). Strikingly, the enhanced pro-inflammatory response in Nedd9-/- macrophages was accompanied by significantly reduced bacterial burden of ST in Nedd9-/- compared to Nedd9wt/wt mBMDMs, shown as a higher fold reduction of bacterial burden, implying that loss of Nedd9 enhances the clearance of bacterial infection (Fig. 3G). Furthermore, reduced ST burden in Nedd9-/- mBMDMs could be rescued by lysosomal inhibition using concanamycin A (Fig. 3H). Taken together, our data suggest that NEDD9 downregulation during bacterial infection represents a cell-autonomous mechanism that enhances the antibacterial defenses against ST.
Infection of Nedd9wt/wt and Nedd9-/- mBMDMs with ST. A IL-6, unpaired t-test: p = 0.0289 and (B) TNF-α, Mann-Whitney test: p = 0.0286 in supernatants of mBMDMs 6 h post ST infection, n = 4, biological replicates. C Western blot analysis of NFК-B Ser536 phosphorylation 1 and 4 hours post infection with ST and (D) densitometric analysis of pNF-КB Ser536 normalized to GAPDH, n = 4, biological replicates, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 1 h) = 0.0005, p (Nedd9wt/wt UI vs. ST 4 h) = 0.0002, p (Nedd9wt/wt ST 1 h vs. Nedd9-/- ST 1 h) = 0.0024, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0103. E Western blot analysis of phosphorylated p38 MAPK Tyr180/Tyr182 1 h and 4 h post infection with ST and (F) densitometric analysis of phosphorylated p38 MAPK Tyr180/Tyr182 normalized to GAPDH, n = 2, biological replicates, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 1 h) = 0.0090, p (Nedd9wt/wt ST 1 h vs. Nedd9-/- ST 1 h) = 0.0043. G In vitro bacterial burden (CFU) in Nedd9wt/wt vs. Nedd9-/- mBMDMs plotted as fold reduction of 24 h to uptake (0 h), n = 3, experiments with at least 3 replicates, Mann-Whitney test: p = 0.0286. H CFU of Nedd9-/- mBMDMs treated with 100 nM Concanamycin A (Conc. A) and controls plotted as fold reduction of 24 h to uptake (0 h), n = 2, experiments with 3 replicates each, unpaired t-test: p = 0.0235. p> 0.05 equals not significant (ns), *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001.
Nedd9 knockout boosts inflammatory response upon ST infection and facilitates bacterial clearance in vivo
Given the elevated pro-inflammatory response and reduced bacterial load in Nedd9-/- mBMDMs in vitro, we investigated the impact of Nedd9 loss on bacterial clearance in vivo by challenging Nedd9wt/wt and Nedd9-/- mice intraperitoneally with ST. In line with the in vitro data, Nedd9 loss significantly enhanced the pro-inflammatory responses upon ST infection, as demonstrated by elevated cytokine IL-6 in spleens of ST-infected Nedd9-/- compared to Nedd9wt/wt mice (Fig. 4A) while loss of Nedd9 did not affect body or spleen weight upon infection (Supplementary Fig. S3A, B). In concordance with our in vitro data, Nedd9 loss significantly reduced ST burden in the spleens of mice compared to wildtype animals 4 days post infection (Fig. 4B). In line with a stronger pro-inflammatory response and lower ST burden, histopathological analysis of Nedd9-/- spleens revealed greater infiltration of lymphoid and myeloid cells as evidenced by follicles of significantly larger size and less defined, confluent shape and characterized by immune cells that spread more extensively (Fig. 4C–F). Consistently, flow cytometric analysis of spleens from Nedd9-/- compared to Nedd9wt/wt mice indicated significantly enhanced recruitment of CD11b+ myeloid cells, including macrophages (Fig. 4G). Thus, these results provide first evidence that NEDD9 downregulation is an important host innate immune defense against bacterial infection both in vitro and in vivo.
Infection of Nedd9wt/wt and Nedd9-/- mice with ST for 4 days. A IL-6 levels in spleens, n = 6 animals/group, Nedd9wt/wt vs. Nedd9-/-, unpaired t-test: p = 0.0127. B In vivo bacterial burden expressed as colony forming units (CFU) in spleens of Nedd9wt/wt and Nedd9-/- mice per gram of spleen tissue, Nedd9wt/wt n = 13, Nedd9-/- n = 20, Mann-Whitney test: p = 0.0072. C Representative HE-stained sections of spleens. Scale bars represent 1000 µm at 25x magnification and 200 µm at 100x magnification. D Analysis of mean follicle size per spleen area (µm), Nedd9wt/wt n = 8, Nedd9-/- n = 13, unpaired t-test: p = 0.0119. E Number of follicles > 50 µm, Nedd9wt/wt n = 18, Nedd9-/- n = 26, unpaired t-test: p = 0.0021. F Number of follicles ≤ 50 µm, Nedd9wt/wt n = 9, Nedd9-/- n = 13, unpaired t-test: p < 0.0001. G Quantification of infiltrating CD11b+ cells (macrophages) as % of CD45+ cells in spleen tissue by flow cytometry, Nedd9wt/wt n = 6, Nedd9-/- n = 7, unpaired t-test: p = 0.0222. *p ≤ 0.05, **p ≤ 0.01, ****p ≤ 0.0001.
NEDD9 promotes ST-mediated FAK-AKT activation and helps ST evade lysosomal degradation
A well described adaptation of ST to evade host defense mechanisms is to trigger FAK signaling by recruiting it to and activating it at the surface of ST-containing vacuoles [10]. Subsequently, FAK activates AKT, which inhibits autophagy and protects ST from lysosomal degradation [10]. We and others have previously shown that inhibition of the FAK effector kinase AKT increases lysosomal functions, resulting in enhanced ability of macrophages to eliminate bacterial pathogens [7, 8, 10]. NEDD9 is a direct interaction partner and substrate of FAK, being recruited by FAK to focal adhesions to regulate cellular adhesion and migration [34, 35]. Given that ST infection causes NEDD9 to accumulate at the site of ST-containing vacuoles and that NEDD9 expression aids ST in evading macrophage-mediated killing, we investigated whether NEDD9 plays a role in this adaptive mechanism of ST to escape host defenses. Therefore, we performed immunoblot analysis of Nedd9wt/wt and Nedd9-/- macrophages 1 hour and 4 hours post ST infection. While NEDD9 wildtype macrophages showed a tendency towards increased levels of total FAK and phosphorylated FAK Tyr397 (pFAK) after ST infection, loss of NEDD9 resulted in significantly decreased FAK and pFAK levels upon infection compared to wildtype controls (Fig. 5A–C). In line with our results and NEDD9 being well known to mediate FAK signaling, pFAK markedly colocalized with NEDD9 upon ST infection (Fig. 5D, Supplemental Fig. 4A). As expected according to the work of Owens et al. ST infection induced AKT phosphorylation at Thr308 and Ser473 in Nedd9wt/wt macrophages slightly after 1 hour and prominently after 4 hours (Fig. 5E–G). In contrast, NEDD9 loss prevented this sharp increase in AKT phosphorylation in macrophages, particularly at 4 hours post ST infection, and resulted in significantly less AKT activation compared to wildtype controls (Fig. 5E–G). Accordingly, phosphorylation of AKT substrates were decreased in ST-infected Nedd9-/- macrophages compared to wildtype controls, with significant changes in phosphorylated mTOR (pMTOR) as well as NDRG-1 (pNDRG-1) and a trend towards reduced expression of S6 kinase (pS6K) (Fig. 5 H, I, Supplemental Fig. 4B–D). Taken together, this indicates that NEDD9 is required for ST-induced FAK-AKT signaling. Consistent with this finding, treatment of Nedd9-/- mBMDMs with the AKT activator SC-79 significantly enhanced ST burden at 4 hours (Fig. 5J, Supplemental Fig. 4E) again indicating that downregulation of the FAK-AKT axis enhances lysosomal functions. Since FAK and AKT signaling have been previously shown to regulate autophagy, we analyzed the conversion of LC-3 I-to II, a specific marker of autophagy, which we found to be significantly increased in Nedd9-/- macrophages upon infection compared to wildtype controls (Fig. 5K, Supplemental Fig. 4F). Further, we analyzed p62 levels, a known receptor protein for the cargo destined for autophagic degradation in Nedd9wt/wt and Nedd9-/- macrophages. Here, we observed decreased levels of p62 in Nedd9-/- macrophages indicating enhanced lysosome-dependent autophagic flux upon infection (Fig. 5L, Supplemental Fig. 4F). This data is consistent with enhanced lysosomal activity observed by increasing cleavage of C12 FDG (Fig. 2H). Collectively, our data show that NEDD9 is necessary for functional ST-mediated FAK-AKT signaling and subsequent reduction in lysosomal function. Thus, NEDD9 downregulation is a plausible mechanism of the macrophage to prevent the host evasion mechanism of ST.
A Western blot analysis of Nedd9wt/wt and Nedd9-/- mBMDMs infected with ST and (B) densitometric analysis of FAK protein level normalized to GAPDH, n = 3, 2-way ANOVA: p (Nedd9-/- UI vs. Nedd9-/- ST 4 h) = 0.0430, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0166 and (C) phosphorylated FAK protein level normalized to GAPDH 1 and 4 h post ST infection, n = 3, 2-way ANOVA: p (Nedd9wt/wt ST 1 h vs. Nedd9-/- ST 1 h) = 0.0219, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0090. D Infection of Nedd9wt/wt and Nedd9-/- mBMDMs with ST (1 h, 4 h, 24 h post infection, uninfected) and quantification of pFAK and NEDD9 colocalization, n = 5, biological replicates, unpaired t-test: p = 0.0028. E Infection of Nedd9wt/wt and Nedd9-/- mBMDMs with ST and Western blot analysis followed by densitometric analysis, n = 3, of (F) phosphorylated AKT Thr308 normalized to GAPDH, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 4 h) = 0.0079, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0199 and (G) phosphorylated AKT Ser473 normalized to GAPDH at indicated time points, n = 3, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 4 h) < 0.0001, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0003. Densitometric analysis of Western blots showing (H) phosphorylated mTOR Ser2448, n = 3, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 1 h) = 0.0401, p (Nedd9wt/wt UI vs. ST 4 h) < 0.0001, p (Nedd9-/- UI vs. ST 4 h) = 0.0036, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0029 and (I) phosphorylated NDRG1 normalized to GAPDH in Nedd9wt/wt and Nedd9-/- mBMDMs 1 and 4 h post ST infection, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 1 h) = 0.0067, p (Nedd9wt/wt UI vs. ST 4 h) = 0.0112, p (Nedd9wt/wt ST 1 h vs. Nedd9-/- ST 1 h) = 0.0038, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0040. J Bacterial burden (CFU/mL) of ST infected Nedd9-/- mBMDMs untreated and treated for 4 h with 1 µM AKT activator SC-79 in vitro, n = 4, unpaired t-test: p = 0.0418. Densitometric analysis of Western blots showing (K) LC3-I to LC3-II conversion normalized to ß-Actin in Nedd9wt/wt and Nedd9-/- mBMDMs 1 h and 4 h post ST infection, n = 3, 2-way ANOVA: p (Nedd9-/- UI vs. Nedd9-/- ST 4 h) = 0.0203, p (Nedd9wt/wt ST 1 h vs. Nedd9-/- ST 1 h) = 0.0128, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) = 0.0006 and (L) p62, n = 3, 2-way ANOVA: p (Nedd9wt/wt UI vs. ST 4 h) < 0.0001, p (Nedd9-/- UI vs. ST 4 h) = 0.0012, p (Nedd9wt/wt ST 1 h vs. Nedd9-/- ST 1 h) = 0.0179, p (Nedd9wt/wt ST 4 h vs. Nedd9-/- ST 4 h) < 0.0001. p > 0.05 equals not significant (ns), *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
NEDD9 is downregulated in human macrophages and patients with bacterial bloodstream infections
Finally, to determine whether NEDD9 downregulation and subsequent increased bacterial clearance is clinically significant, we infected human macrophages derived from healthy donors with ST and analyzed NEDD9 levels by Western Blot analysis. NEDD9 expression was strongly decreased in human macrophages 4 hours post ST infection (Fig. 6A). Next, we analyzed NEDD9 protein expression in PBMCs from patients suffering from various bacterial BSI and healthy controls (Supplemental Table S1). Consistent with our previous data, we observed that NEDD9 protein levels were significantly downregulated in patients with bacterial BSI compared to healthy controls (Fig. 6B, C). Next, we investigated the functional implications of NEDD9 loss in primary human macrophages by depleting NEDD9 using NEDD9-specific small interfering RNA followed by ST infection (Fig. 6D). Consistent with our previous data, NEDD9 knockdown resulted in a significantly lower bacterial burden (Fig. 6E) and increased levels of proinflammatory cytokine secretion (Fig. 6F). In conclusion, our results suggest that downregulation of NEDD9 is a host defense mechanism that could serve as a potential host-directed target to treat bacterial infection (graphical abstract).
A Infection of primary human macrophages with ST followed by Western blot analysis showing NEDD9 protein level 4 h post infection compared to uninfected (UI) cells, n = 2. B Representative Western Blot analysis and (C) statistical analysis of NEDD9 protein expression normalized to GAPDH in PBMCs from patients suffering from bacterial BSI, n = 9, versus healthy control donors (HC), n = 8, unpaired t-test: p = 0.0309. D Primary human macrophages were transfected with non-target siRNA (siNT) or siRNA targeting NEDD9 (siN9) followed by infection with ST for 1 h and 4 h. Western blot analysis shows NEDD9 protein expression and (E) assessment of in vitro bacterial burden (CFU/mL), n = 4, 2-way ANOVA: p (siNT 1 h vs. siN9 1 h) = 0.0006, p (siNT 4 h vs. siN9 4 h) = 0.0272. F IL-6 quantification using ELISA supernatant 4 h post-infection with ST, unpaired t-test: p = 0.0130, n = 3. *p ≤ 0.05, ***p ≤ 0.001.
Discussion
Although mostly studied for its role in cancer, NEDD9 has been shown to alter lymphocyte trafficking in secondary lymphoid organs, suggesting a role for NEDD9 in regulating immune responses [20]. However, neither the relevance of NEDD9 in cells of the innate immune system nor the role of NEDD9 in the response to infection has previously been systematically investigated. In this study, we observed that NEDD9 is downregulated transcriptionally and post-translationally in macrophages upon ST infection. In addition, treatment of macrophages with LPS – the most abundant antigen of Gram-negative bacteria – was sufficient to reduce NEDD9 levels.
To our knowledge, regulation of NEDD9 in the context of bacterial infection has not been reported previously. Only in the context of viral infection with human T-cell leukemia/lymphoma virus and interaction with oncogenic TAX protein NEDD9 was found to be upregulated and linked to oncogenesis [36]. Indeed, in many human cancer entities NEDD9 upregulation is associated with tumor aggressiveness, metastasis and resistance to chemotherapy [17, 37, 38]. However, in a mouse model for CML, loss of NEDD9 suppressed disease progression emphasizing its cell-specific function [20]. NEDD9 levels were decreased in macrophages in a virulence-independent manner by ST and can even be triggered by LPS stimulation but not by TLR-2 agonist or Gram- positive bacteria such as SA. Therefore, we propose that Nedd9 downregulation is a macrophage response specifically to Gram-negative bacterial infections.
In most cancer cells, NEDD9 promotes biological events such as migration, invasion, epithelial mesenchymal transition (EMT) and chemotaxis. Thereby, NEDD9 is primarily located in the cytosol but translocates to different sites in the cell in a context-specific manner, including focal adhesions to increase extra-cellular matrix binding [39] or the centrosome and basal body during the cell cycle to induce mitotic spindle formation [40] or ciliary disassembly [33]. In our study, we observed that ST infection led to a change in the localization of NEDD9 from the cytosol to bacteria-containing phagosomes and eventually degraded in lysosomes. NEDD9 is very dynamically regulated in its function as a scaffold protein, both transcriptionally and post-translationally. In this context, NEDD9 has been shown to be degraded by various stimuli, e.g. at the end of mitosis by proteasomal degradation [29]. In contrast, in our study, inhibition of the proteasomes did not alter the degradation of NEDD9 in macrophages after infection. Instead, inhibition of the lysosomal pathway resulted in stabilization of NEDD9 and increased bacterial burden in Nedd9-/- mBMDMs after ST infection. The fact that NEDD9 colocalized with bacteria-containing phagosomes including lysosomal marker LAMP1, as well as with pFAK upon ST infection and was downregulated via lysosomal degradation suggests a regulatory function in macrophage defense against bacteria. Moreover, phagolysosomal functions were significantly enhanced upon loss of NEDD9 resulting in reduced bacterial burden.
Acceleration of phagolysosomal maturation can be mediated through many different factors [41]. Secretion of cytokines is a well-established mechanism by which phagolysosomal maturation can be enhanced and ultimately support bacterial clearance [42]. NF-κb is a broad regulator not only of inflammation but also of NEDD9 transcription according to an in silico analysis of the NF-κb signaling pathway [17]. However, the interference of NEDD9 and NF-κb has not yet been further investigated. In line with the in silico analysis we found that loss of NEDD9 was associated with decreased phosphorylation of NF-κb and decreased phosphorylation of p38, whereas secretion of pro-inflammatory cytokines such as IL-6 and TNF-alpha was increased upon infection, suggesting NF-κb and p38 independent regulation of cytokine release by other pathways. This finding is consistent with previous studies showing increased secretion of pro-inflammatory cytokines and enhanced lysosomal functions despite reduced NF-κb phosphorylation, which may alternatively be mediated by mitochondrial metabolism [43]. Indeed, loss of NEDD9 not only resulted in increased release of pro-inflammatory cytokines but also in enhanced bacterial clearance, both in vitro and in vivo.
To evade host defense, we and others have shown Gram-negative bacteria such as ST develop an adaption mechanism by recruiting FAK and AKT at the site of ST-containing vacuoles to inhibit autophagy, which subsequently prevents their lysosomal degradation [7, 8]. FAK is known to be directly activated upon ST infection and thereby promotes bacterial survival via the AKT-mTOR signaling pathway [10]. This process is dependent on FAK kinase activity and negatively regulates autophagy via interaction with scaffolding protein p130CAS, a paralog of NEDD9 [10, 44]. We show that NEDD9 translocates to phagolysosomes upon infection where it colocalizes with FAK (Fig. 5D, E). NEDD9 is known to form a complex with kinases, in particular with FAK, to mediate their activity and regulate cellular processes such as migration, proliferation and adhesion [16, 23, 24]. As NEDD9 is recruited to ST-containing vacuoles we analyzed if NEDD9 is involved in the ST induced FAK-AKT pathway to inhibit lysosomal degradation. Indeed, our results show, that loss of NEDD9 leads to decreased FAK expression and phosphorylation in ST-infected macrophages. NEDD9-dependent FAK signaling has been previously reported in the context of cancer [16]. This data indicates that the presence of NEDD9 at ST-containing vacuoles is essential for FAK upregulation and activation upon infection, an ST survival mechanism to hinder their lysosomal degradation in macrophages.
Downstream of FAK, ST infection is known to induce phosphorylation of AKT in macrophages thereby bypassing immune defense mechanisms such as autophagy [7]. AKT is recognized as a general regulator of lysosomal function associated with bacterial infections and beyond, and is also a known downstream target of NEDD9 [8, 10]. In line with this, we observed phosphorylation of AKT and AKT substrates such as mTOR and NDRG-1 to be increased in Nedd9wt/wt mBMDMs but decreased upon Nedd9 loss, which suggests that AKT activation is dependent on NEDD9. Accordingly, bacterial load was reduced in Nedd9-/- compared to Nedd9wt/wt macrophages confirming previous findings that inhibition of AKT enhances bacterial clearance in macrophages. This data also corroborates with previous findings that flavonoids suppressing NEDD9 lead to lysosome-dependent downregulation of AKT [45, 46].
In addition, loss of Nedd9 resulted in significantly increased LysoTracker staining and increased cathepsin D expression in phagosomes isolated from Nedd9-/- cells as well as decreased p62 protein level, indicative for increased autophagic flux. In line with this, measuring lysosomal ß-galactosidase activity on C12FDG confirmed lysosomal function is enhanced in Nedd9-/- BMDMs upon infection. Thus, Nedd9 loss enhances phagolysosomal activity in macrophages and thereby counteracts the ST evasion mechanism of FAK-AKT activation to inhibit lysosomal function. In accordance with this, previous studies have shown that NEDD9 actively suppresses autophagy in lung cancer cells, and that autophagy is enhanced in NEDD9-deficient cells [47].
In summary, our study suggests that the downregulation of NEDD9 in macrophages serves as a host defense mechanism against Gram-negative bacterial infections. Conversely, it also indicates that ST and other Gram-negative pathogens exploit NEDD9 as a scaffold to recruit FAK to ST-containing vacuoles and to activate FAK and AKT, which in turn inhibits phagolysosomal degradation of the pathogens. However, as a host defense mechanism, macrophages degrade NEDD9 to prevent its hijacking by ST. This degradation restores proper phagolysosomal function, thereby enhancing bacterial clearance. Supporting this mechanism, we observed consistent downregulation of NEDD9 in PBMCs from patients with BSI, in line with our in vitro and in vivo data. Nonetheless, further studies are needed to elucidate the kinetics of NEDD9 downregulation and to explore its potential as a diagnostic marker or therapeutic target. In conclusion, our study reveals for the first time a functional link between NEDD9 and macrophage-mediated antibacterial defense (see graphical abstract), thereby expanding the current understanding of NEDD9 in immune responses. This findings suggest that targeting the NEDD9 pathway may represent a promosing host-directed strategy for the treatment of bacterial infections.
Material and Methods
Study approval
All animal procedures were conducted in accordance with the institutional guidelines on animal welfare and were approved by the North Rhine-Westphalian State Agency for Nature, Environment, and Consumer Protection (Landesamt für Natur, Umwelt and Verbraucherschutz [LANUV] Nordrhein-Westfalen; File no: 84-02.05.40.14.082 and 84-02.04.2015.A443) and the University of Cologne, Germany.
Human subjects
The study involving human blood samples from healthy donors was approved by the Local Ethics Commission of the University Hospital of Cologne, Germany (File no: 08-160). Written consent was received from participants prior to inclusion in the study. All clinical investigations were conducted according to the Declaration of Helsinki principles.
Animals
Mice were bred under standard conditions with ventilation system
Nedd9 knockout mice (Nedd9-/-) (Seo et al., 2006) and wild type mice of the same background C57Bl/6/J (Nedd9wt/wt) were held under specific pathogen free conditions in a conventional animal facility. For infection animals at the age of 8–12 weeks with a balanced mix of male and female animals were used. Animal randomization or blinding was not needed as WT and knockout mice were tested.
Pathogens
The Salmonella enterica Serovar Typhimurium strain (SL1344) was commercially purchased. The ST SPI1 mutant ΔssaV and SPI2 mutant ΔinvA were kindly provided by Ivan Dikic. Other pathogens used as controls in this study Shigella flexneri (ATCC 12022), Pseudomonas aeruginosa (ATCC 27853), Escherichia coli (ATCC 25922), Staphylococcus aureus (ATCC29213) or Listeria monocytogenes (EGD-e) were kindly provided by Georg Plum and are used as control strains in the routine patient diagnostics at the Institute of medical Microbiology, Immunology and Hygiene of the University Hospital Cologne. These strains are registered in the American Type Culture Collection (ATCC).
Bacteria preparation
Bacteria from a single colony was inoculated into 5 mL BHI medium and incubated overnight at 37 °C with constant agitation. Next, 1 mL bacterial suspension was transferred into 50 mL BHI and grown until the OD600 reached 1. The bacteria concentration was estimated by plating serial dilutions on BHI agar plates.
Cell culture and bacterial infection
Bone marrow cells were differentiated into macrophages for 7–10 days in RPMI medium supplemented with 20% L929 supernatant and 10% FBS. Mycoplasma test were regularly performed. mBMDMs were infected with specified bacteria at a MOI of 10. After infection, mBMDMs were incubated with the bacteria for 10 min at room temperature (RT) and then 30 min at 37 °C. This incubation time allowed bacteria to be internalized by macrophages. Cells were then washed with RPMI medium and incubated in medium supplemented with 50 µg/ml gentamicin. After 2 hours, the gentamicin concentration was reduced to 10 µg/ml. mBMDMs were treated with 10 µM AKT-Inhibitor VIII (Enzo lifesciences), 100 ng/ml ST LPS (Sigma), 100 nM Concanamycin A (Sigma), 50 µM MG132 (Selleckchem), 1 µM AKT-Activator SC79 (Selleckchem) dissolved in medium for 1 hour prior to infection. Human PBMCs were isolated from 30 mL EDTA blood using Lymphoprep (Stem Cells) and the monocytes were differentiated in RPMI containing 10% FCS and 50ng/mL M-CSF.
ST phagosome isolation
ST was grown in BHI broth until the OD600 reached 1, and then biotinylated with EZ-link NHS-Biotin reagent (Thermo Fisher Scientific). After washing, biotinylated bacteria were incubated with siMAG Streptavidin ferrofluid (Chemicell) at 37 °C. Biotinylated ST bound to Streptavidin ferrofluid was then separated using a magnet and the bacteria were quantified using BHI agar plates. Subsequently, mBMDMs were infected with the biotinylated ST bound to the Streptavidin ferrofluid at a MOI of 10. At each time point, ST-containing phagosomes were isolated using equilibration and lysis buffer, as previously described for bead phagosome isolation.
In vitro bacterial burden and ELISA
After 24 hours post-infection, mBMDMs were lysed with 1% Triton X-100, 0.01% SDS in PBS. Several dilutions of the lysate were plated on BHI plates and incubated overnight at 37 °C. The next day, ST CFUs were enumerated. Supernatants were collected and analyzed for IL-6 and TNF-α secretion using ELISA kits (R&D, DY406-05, DY410-05), according to the manufacturer’s instructions.
Estimation of bacterial burden in vivo
Mice were infected (i.p. injection) with 100 CFU ST. After 4 days of infection, mice were euthanized. The spleens were isolated, weighted and homogenized using a gentleMACS Dissociator (Miltenyi Biotec, Bergisch Gladbach, Germany) in sterile PBS. Extracts of the homogenized spleens were plated on BHI Agar plates. After 24 h incubation at 37 °C, the bacterial colonies were enumerated. The number of colonies was normalized to per gram of tissue. Spleen extracts were also used for ELISA (described above).
Tissue histology
Spleen and liver samples were obtained from in vivo experiments and cross-sectioned in equal-sized slices 2 mm in width. The respective slices were snap-frozen in liquid nitrogen covered with Tissue-Tek® and stored at −80 °C until further use. Cryostat sections were cut to 3 µm and attached to adhesive glass slides (Dako, Hamburg, Germany), fixed with 4% phosphate-buffered formalin for 30 sec, and stained with H&E. After rinsing in water, the slides were dehydrated via an ascending ethanol gradient, immersed in xylene and mounted on glass coverslips. The tissue sections were imaged using a Hitachi HV-F202 UXGA 3CCD camera attached to Leica DM5500B microscope. Pictures were generated using the DISKUS program (Hilgers Technisches Büro, Königswinter, Germany).
Western blotting
mBMDMs were lysed in RIPA buffer supplemented with a 1X protease/phosphatase inhibitor cocktail (Thermo Fisher Scientific). Protein concentrations were estimated using Pierce® BCA Protein Assay Kit (Thermo Fisher Scientific), according to the manufacturer’s instructions. Equal amounts of protein were separated on 10% or 12% SDS-PAGE gels or on 4–20% Mini-PROTEAN® TGX™ Precast Protein Gels (Biorad #4561094) using a broad range, color-coded prestained protein marker (#14208 Cell Signaling). The proteins were then transferred onto a PVDF membrane and probed with the following antibodies: HEF1/Nedd9 (#4044 Cell Signaling), phospho-NF-κB p65 Ser536 (#3033 Cell Signaling), NF-κb p65 (#4764 Cell Signaling), Lamp-1 (sc-17768 Santa Cruz Biotechnology), phospho-Akt Ser473 (#4060 Cell Signaling), phospho-Akt Thr 308 (#2965 Cell Signaling), Akt (#4691 Cell Signaling), phospho-FAK Tyr397 (#3283 Cell Signaling), FAK (#3285S Cell signaling), phospho-mTOR Ser2448 (#4561094 Cell Signaling), phospho-p70 S6 Kinase Thr389 (#9234#9205 Cell Signaling), phospho-NDRG-1 (#3217 Cell Signaling), p62 (#5114 Cell Signaling), LC-3 (L7543 Sigma), phospho-P38 (Cell Signaling), Cathepsin D (#2284 Cell Signaling), β-actin (A2228 Sigma-Aldrich) and GAPDH (#2118S Cell Signaling). GAPDH and ß-Actin was used as a loading control. After incubation with secondary HRP-conjugated antibodies (R&D), the blots were developed using ECL reagent (GE Healthcare). Densitometric analysis of Western Blots was performed using lab image 1D software (Kapelan Bio-Imaging GmbH, Leipzig, Germany).
Immunofluorescence staining and confocal microscopy
mBMDMs and human macrophages grown on glass coverslips were infected with ST and fixed with 4% formaldehyde or methanol in PBS for 15 min at RT. When indicated, the cells were treated with LysoTracker before fixation at the indicated time points. The samples were then permeabilized with 0.3% Triton X-100 in PBS for 5 min and then incubated with Image-iT® R FX (#I36933, Invitrogen) and blocked with 5% (w/v) normal goat serum (Life Technologies). The cells were then incubated overnight with specific primary antibodies NEDD99 (#4044S, cell signaling), phospho-FAK Tyr397 (#3283 Cell Signaling), Lamp-1 (sc-17768 Santa Cruz Biotechnology) and ST LPS (MA1-83451, Thermo Fisher Scientific) at 4 °C. Alternatively, PE-labeled NEDD9 antibody was used (Novus NB100-1699PE). After washing with 0.03% Triton in PBS, the cells were incubated with either Alexa Fluor 594-conjugated goat anti-rabbit or anti-mouse secondary antibody or Alexa Fluor 488-conjugated goat anti-rabbit or anti-mouse (Life Technologies) for 1 hour at RT protected from light. The cover-slips were then mounted using ProLong® Gold antifade DAPI (#P36953, Life Technologies) to stain the nuclei. The slides were imaged using a 63X objective under a confocal microscope (Leica SP8, Leica Microsystems).
Colocalization was analyzed using Colocalization Analysis plugins (ImageJ software). First, the Colocalization Test plugin with Fay randomization method was performed to calculate Pearson’s correlation coefficient for the two channels in each selected ROI (25 × 25 pixels). This value was compared with what would be expected for random overlap. The observed correlation was considered significant if it was greater than 95% of the correlations between channel 1 and a number of randomized channel 2 images. All ROIs with Pearson’s coefficient p value of ≥ 0.95 were further analyzed by the Colocalization Threshold plugin to calculate thresholded Mander’s coefficients [tM1 colocalization value for channel 1 (red); tM2 colocalization value for channel 2 (green or blue)] and to generate scatterplots with linear regression line and thresholds.
NEDD9 siRNA knockdown in human macrophages
Human macrophages were transfected with either 50 nM non-targeting siRNA (#SR-CL000-005, Eurogentec) or siRNA for NEDD9 (Dharmacon) using the transfection reagent Lipofectamine 3000 (Life Technologies), according to the manufacturer’s instructions.
RNA isolation, reverse transcription and qPCR
RNA was isolated using RNeasy® Mini Kit followed by reverse transcription using GoScript™ Reverse Transcription System according to the manufacturer’s protocol. Quantitative real-time PCR for Nedd9 was run on a 7500 Fast cycler system (Applied Biosystems, Foster City, California, United States) Nedd9 mRNA expression was determined using TaqMan™ Fast Advanced Master Mix. GAPDH was used as a reference gene to normalize NEDD9 expression.
Gene expression array
Total RNA was extracted from uninfected or ST-infected (2 hours) Nedd9-/- mBMDMs using the RNeasy Mini Kit (Qiagen), and subsequently treated with RNase-free DNase I for 30 min at 37 °C to remove residual DNA. Library preparation was carried out according to Illumina’s protocol for ‘Preparing Samples for Sequencing of mRNA’ (Part #1004898, Rev. A). Briefly, mRNA was isolated from total RNA with oligo(dT) magnetic beads. Upon purification and chemical fragmentation, cleaved RNA was copied into first strand cDNA using reverse transcriptase and random primers. Second strand cDNA was synthesized using DNA Polymerase I and RNase H. Following end repair, addition of a single A base, and adaptor ligation, cDNA was size-selected on agarose gels and products were further purified and enriched by PCR amplification to create the final cDNA library. Library integrity was analyzed using an Agilent Technologies 2100 Bioanalyzer and libraries were subsequently sequenced on an Illumina HiSeq 2000 platform (BGI Genomics, China) (GSE84375) [25].
Quantification of hydrolase activity
1 × 108 ST in 100 µl PBS were labeled with 350 µg/ml 5-dodecanoylaminofluorescein-di-β-D-galactopyranoside (C12FDG, D2893, Thermo Fisher Scientific) in 0.1 M NaHCO3 buffer, pH 9.6 for 1 hour at 37 °C on a tube rotator. mBMDMs (50,000 cells/well) were infected at a MOI of 50 in a 96-well black flat bottom plate and then incubated at 37 °C in HBSS with Ca2+ and Mg2+ in a Tristar2 multimode plate reader LB 942 (Berthold Technologies). Unquenching of C12FDG fluorescence was measured at 120 s intervals using standard GFP filters. Phagosomal acidification, and thereby acid hydrolase activity, was inhibited by 5 µM chloroquine.
Bioinformatic analyses
Raw sequencing reads were filtered into clean reads and were aligned to murine reference sequences (ftp://ftp.ncbi.nih.gov/refseq/M_musculus/mRNA_Prot/) using SOAPaligner/SOAP2 [48]. Gene expression values were then calculated by the RPKM method [49] and differentially expressed genes (DEGs) among uninfected and S.Typhimurium-infected (2 hours) Nedd9-/- mBMDMs were identified. Up and downregulation of single genes was assessed by calculating the log2 of fold changes (log2[RPKM WT ST/RPKM WT]). Correction for false positive (type I errors) and false negative (type II) errors was performed based on the false discovery rate (FDR) method [50]. ‘FDR ≤ 0.001 and log2 ratio ≥ 1’ were set as the cutoff to determine the significance of DEGs.
Statistical Analysis
Statistical analysis and data evaluation were carried with GraphPad Prism version 10.2.3 for Windows, GraphPad Software, La Jolla, California, United States. Data are presented as mean ± SD if not otherwise indicated. Significance is indicated directly in the figures as *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Unpaired t-test was performed to compare two independent groups with normally distributed data. Data of two independent groups that are not normally distributed were compared by using Mann-Whitney test. One-way ANOVA was used to compare three or more independent groups for one single factor and Kruskal-Wallis test was used when assumptions of ANOVA were not met. Two-way ANOVA was used to determine the effect of two different independent factors.
Data availability
All data generated or analysed during this study are included in this published article.
Change history
11 August 2025
The original online version of this article was revised: The author Sebastian J. Theobald was initially written as Sebastian Theobald.
29 August 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41419-025-07919-z
References
Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA. 2016;315:801–10.
Cohen J, Vincent JL, Adhikari NK, Machado FR, Angus DC, Calandra T, et al. Sepsis: a roadmap for future research. Lancet Infect Dis. 2015;15:581–614.
WHO. Improving prevention, diagnosis and clinical management of sepsis. 2017; https://apps.who.int/gb/ebwha/pdf_files/WHA70/A70_13-en.pdf?ua=1,%20accessed%2027%20June%202020.
Laxminarayan R, Duse A, Wattal C, Zaidi AK, Wertheim HF, Sumpradit N, et al. Antibiotic resistance-the need for global solutions. Lancet Infect Dis. 2013;13:1057–98.
Ginhoux F, Guilliams M. Tissue-resident macrophage ontogeny and homeostasis. Immunity. 2016;44:439–49.
Robinson N, McComb S, Mulligan R, Dudani R, Krishnan L, Sad S. Type I interferon induces necroptosis in macrophages during infection with Salmonella enterica serovar Typhimurium. Nat Immunol. 2012;13:954–62.
Ganesan R, Hos NJ, Gutierrez S, Fischer J, Stepek JM, Daglidu E, et al. Salmonella Typhimurium disrupts Sirt1/AMPK checkpoint control of mTOR to impair autophagy. PLoS Pathog. 2017;13:e1006227.
Fischer J, Gutierrez S, Ganesan R, Calabrese C, Ranjan R, Cildir G, et al. Leptin signaling impairs macrophage defenses against Salmonella Typhimurium. Proc Natl Acad Sci USA. 2019;116:16551–60.
Hos NJ, Ganesan R, Gutierrez S, Hos D, Klimek J, Abdullah Z, et al. Type I interferon enhances necroptosis of Salmonella Typhimurium-infected macrophages by impairing antioxidative stress responses. J Cell Biol. 2017;216:4107–21.
Owen KA, Meyer CB, Bouton AH, Casanova JE. Activation of focal adhesion kinase by Salmonella suppresses autophagy via an Akt/mTOR signaling pathway and promotes bacterial survival in macrophages. PLoS Pathog. 2014;10:e1004159.
Gordon MA. Salmonella infections in immunocompromised adults. J Infect. 2008;56:413–22.
Parisi A, Crump JA, Stafford R, Glass K, Howden BP, Kirk MD. Increasing incidence of invasive nontyphoidal Salmonella infections in Queensland, Australia, 2007-2016. PLoS Negl Trop Dis. 2019;13:e0007187.
Mughini-Gras L, Pijnacker R, Duijster J, Heck M, Wit B, Veldman K, et al. Changing epidemiology of invasive non-typhoid Salmonella infection: a nationwide population-based registry study. Clin Microbiol Infect. 2020;26:941.e9–e14.
Tikhmyanova N, Little JL, Golemis EA. CAS proteins in normal and pathological cell growth control. Cell Mol Life Sci. 2010;67:1025–48.
Gabbasov R, Xiao F, Howe CG, Bickel LE, O’Brien SW, Benrubi D, et al. NEDD9 promotes oncogenic signaling, a stem/mesenchymal gene signature, and aggressive ovarian cancer growth in mice. Oncogene. 2018;37:4854–70.
Izumchenko E, Singh MK, Plotnikova OV, Tikhmyanova N, Little JL, Serebriiskii IG, et al. NEDD9 promotes oncogenic signaling in mammary tumor development. Cancer Res. 2009;69:7198–206.
Shagisultanova E, Gaponova AV, Gabbasov R, Nicolas E, Golemis EA. Preclinical and clinical studies of the NEDD9 scaffold protein in cancer and other diseases. Gene. 2015;567:1–11.
Ohashi Y, Iwata S, Kamiguchi K, Morimoto C. Tyrosine phosphorylation of Crk-associated substrate lymphocyte-type is a critical element in TCR- and beta 1 integrin-induced T lymphocyte migration. J Immunol. 1999;163:3727–34.
Gu JJ, Lavau CP, Pugacheva E, Soderblom EJ, Moseley MA, Pendergast AM. Abl family kinases modulate T cell-mediated inflammation and chemokine-induced migration through the adaptor HEF1 and the GTPase Rap1. Sci Signal. 2012;5:ra51.
Seo S, Nakamoto T, Takeshita M, Lu J, Sato T, Suzuki T, et al. Crk-associated substrate lymphocyte type regulates myeloid cell motility and suppresses the progression of leukemia induced by p210Bcr/Abl. Cancer Sci. 2011;102:2109–17.
Seo S, Asai T, Saito T, Suzuki T, Morishita Y, Nakamoto T, et al. Crk-associated substrate lymphocyte type is required for lymphocyte trafficking and marginal zone B cell maintenance. J Immunol. 2005;175:3492–501.
Katayose T, Iwata S, Oyaizu N, Hosono O, Yamada T, Dang NH, et al. The role of Cas-L/NEDD9 as a regulator of collagen-induced arthritis in a murine model. Biochem Biophys Res Commun. 2015;460:1069–75.
Sima N, Cheng X, Ye F, Ma D, Xie X, Lü W. The overexpression of scaffolding protein NEDD9 promotes migration and invasion in cervical cancer via tyrosine phosphorylated FAK and SRC. PLoS One. 2013;8:e74594.
Baquiran JB, Bradbury P, O’Neill GM. Tyrosine Y189 in the substrate domain of the adhesion docking protein NEDD9 is conserved with p130Cas Y253 and regulates NEDD9-mediated migration and focal adhesion dynamics. PLoS One. 2013;8:e69304.
Gutiérrez S, Fischer J, Ganesan R, Hos NJ, Cildir G, Wolke M, et al. Salmonella Typhimurium impairs glycolysis-mediated acidification of phagosomes to evade macrophage defense. PLoS Pathog. 2021;17:e1009943.
Cabodi S, del Pilar Camacho-Leal M, Di Stefano P, Defilippi P. Integrin signalling adaptors: not only figurants in the cancer story. Nat Rev Cancer. 2010;10:858–70.
Fashena SJ, Einarson MB, O’Neill GM, Patriotis C, Golemis EA. Dissection of HEF1-dependent functions in motility and transcriptional regulation. J Cell Sci. 2002;115:99–111.
Liu X, Elia AE, Law SF, Golemis EA, Farley J, Wang T. A novel ability of Smad3 to regulate proteasomal degradation of a Cas family member HEF1. EMBO J. 2000;19:6759–69.
Zheng M, McKeown-Longo PJ. Cell adhesion regulates Ser/Thr phosphorylation and proteasomal degradation of HEF1. J Cell Sci. 2006;119:96–103.
Pugacheva EN, Golemis EA. The focal adhesion scaffolding protein HEF1 regulates activation of the Aurora-A and Nek2 kinases at the centrosome. Nat Cell Biol. 2005;7:937–46.
Dadke D, Jarnik M, Pugacheva EN, Singh MK, Golemis EA. Deregulation of HEF1 impairs M-phase progression by disrupting the RhoA activation cycle. Mol Biol Cell. 2006;17:1204–17.
Ice RJ, McLaughlin SL, Livengood RH, Culp MV, Eddy ER, Ivanov AV, et al. NEDD9 depletion destabilizes Aurora A kinase and heightens the efficacy of Aurora A inhibitors: implications for treatment of metastatic solid tumors. Cancer Res. 2013;73:3168–80.
Pugacheva EN, Jablonski SA, Hartman TR, Henske EP, Golemis EA. HEF1-dependent Aurora A activation induces disassembly of the primary cilium. Cell. 2007;129:1351–63.
Minegishi M, Tachibana K, Sato T, Iwata S, Nojima Y, Morimoto C. Structure and function of Cas-L, a 105-kD Crk-associated substrate-related protein that is involved in beta 1 integrin-mediated signaling in lymphocytes. J Exp Med. 1996;184:1365–75.
Natarajan M, Stewart JE, Golemis EA, Pugacheva EN, Alexandropoulos K, Cox BD, et al. HEF1 is a necessary and specific downstream effector of FAK that promotes the migration of glioblastoma cells. Oncogene. 2006;25:1721–32.
Iwata S, Souta-Kuribara A, Yamakawa A, Sasaki T, Shimizu T, Hosono O, et al. HTLV-I Tax induces and associates with Crk-associated substrate lymphocyte type (Cas-L). Oncogene. 2005;24:1262–71.
Kong C, Wang C, Wang L, Ma M, Niu C, Sun X, et al. NEDD9 is a positive regulator of epithelial-mesenchymal transition and promotes invasion in aggressive breast cancer. PLoS One. 2011;6:e22666.
Rusyn L, Reinartz S, Nikiforov A, Mikhael N, Vom Stein A, Kohlhas V, et al. The scaffold protein NEDD9 is necessary for leukemia-cell migration and disease progression in a mouse model of chronic lymphocytic leukemia. Leukemia. 2022;36:1794–805.
Zhong J, Baquiran JB, Bonakdar N, Lees J, Ching YW, Pugacheva E, et al. NEDD9 stabilizes focal adhesions, increases binding to the extra-cellular matrix and differentially effects 2D versus 3D cell migration. PLoS One. 2012;7:e35058.
Pugacheva EN, Golemis EA. HEF1-aurora A interactions: points of dialog between the cell cycle and cell attachment signaling networks. Cell Cycle. 2006;5:384–91.
Pauwels AM, Trost M, Beyaert R, Hoffmann E. Patterns, Receptors, and Signals: Regulation of Phagosome Maturation. Trends Immunol. 2017;38:407–22.
Nguyen T, Robinson N, Allison SE, Coombes BK, Sad S, Krishnan L. IL-10 produced by trophoblast cells inhibits phagosome maturation leading to profound intracellular proliferation of Salmonella enterica Typhimurium. Placenta. 2013;34:765–74.
Mills EL, Kelly B, O’Neill LAJ. Mitochondria are the powerhouses of immunity. Nat Immunol. 2017;18:488–98.
Shi J, Casanova JE. Invasion of host cells by Salmonella typhimurium requires focal adhesion kinase and p130Cas. Mol Biol Cell. 2006;17:4698–708.
Zhou RT, He M, Yu Z, Liang Y, Nie Y, Tai S, et al. Baicalein inhibits pancreatic cancer cell proliferation and invasion via suppression of NEDD9 expression and its downstream Akt and ERK signaling pathways. Oncotarget. 2017;8:56351–63.
Dai J, Van Wie PG, Fai LY, Kim D, Wang L, Poyil P, et al. Downregulation of NEDD9 by apigenin suppresses migration, invasion, and metastasis of colorectal cancer cells. Toxicol Appl Pharmacol. 2016;311:106–12.
Deneka AY, Kopp MC, Nikonova AS, Gaponova AV, Kiseleva AA, Hensley HH, et al. Restrains Autophagy to Limit Growth of Early Stage Non-Small Cell Lung Cancer. Cancer Res. 2021;81:3717–26.
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics. 2009;25:1966–7.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;5:621–8.
Benjamini Y, Yekutieli D. The Annals of Statistics- The Control of the False Discovery Rate in Multiple Testing under Dependency, Vol. 29, pp. 1165-88.
Acknowledgements
We thank Monika Keiten-Schmitz and Elke Weber for technical support. This work was supported by funding from the German Center for Infectious Research (DZIF) (80185MLJFI to J.F. and TTU 08.928 to M.K.), the German Research Council (DFG, SE 2280/3-1 to T.S.-N.), the German Ministry of Science and Education (BMBF) as part of the e:Med initiative (01ZX1406 to T.S.-N.), the Exzellenz initiieren (E.I.) - Stiftung Kölner Krebsforschung (to T.S.-N.), the “Deutsche Jose-Carreras Leukaemiestiftung e.V.” (DJCLS 19 R/2021), the medical faculty of the University of Cologne (Gusyk program to J.F. and T.S.-N.; Köln Fortune, 252/2013 to T.S.-N) and the medical faculty of Münster (No. 2024_002). Jonel Trebicka was supported by the German Research Foundation (DFG) project ID 403224013 – SFB 1382 (A09), by the German Federal Ministry of Education and Research (BMBF) for the DEEP-HCC project and by the Hessian Ministry of Higher Education, Research and the Arts (HMWK) for the ENABLE and ACLF-I cluster projects. The MICROB-PREDICT (project ID 825694), DECISION (project ID 847949), GALAXY (project ID 668031), LIVERHOPE (project ID 731875), and IHMCSA (project ID 964590) projects have received funding from the European Union’s Horizon 2020 research and innovation program. The manuscript reflects only the authors’ views, and the European Commission is not responsible for any use that may be made of the information it contains. The funders had no influence on study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.F., T.S.-N., N.R.; Methodology: N.R., J.F.; Investigation: L.R., N.H., Z.H., C.C., P.M., J.W.U.F. J.F., J.S., L.L., S.J.T., P-H.N., M.H., A.G., H.N., F.K.; Writing – Original Draft: J.F., L.R.; Writing – Review & Editing: T.S.-N., J.F., N.H., J.W.U.F, J.S., N.R., J.T., F.K.; Funding Acquisition: T.S.-N.,J.F, Resources: J.F., T.S.-N., N.R., J.R., M.K., J.T., C.L. Supervision: J.F., T.S.-N., Patient Recruitment J.S., J.J.V., R.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests. Jonel Trebicka has received speaking and/or consulting fees from Versantis, Gore, Boehringer-Ingelheim, Falk, Grifols, Genfit and CSL Behring.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Gerry Melino
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fischer, J., Rusyn, L., Krus, F. et al. Macrophages downregulate NEDD9 to counteract S. Typhimurium- mediated FAK-AKT activation and lysosome inhibition. Cell Death Dis 16, 445 (2025). https://doi.org/10.1038/s41419-025-07634-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-025-07634-9