Abstract
Pancreatic cancer is a highly aggressive tumor with a poor prognosis. The ubiquitin-proteasome system is crucial in maintaining protein homeostasis and regulating cellular senescence, thereby influencing tumorigenesis and progression; however, the specific mechanisms underlying this process remain unclear. Ubiquitin protein ligase E3A (UBE3A) is associated with various tumors; however, its potential role in pancreatic cancer warrants further investigation. The effects of UBE3A on cellular senescence were evaluated through in vitro and in vivo experiments, including SA-β-gal staining, EdU assays, detection of senescence-associated markers, and the establishment of subcutaneous xenograft and liver metastasis models. Mechanistic studies utilized Ub-MS, IP-MS, ChIP-seq, and RNA-seq to explore downstream pathways, followed by validation through ubiquitination assays, ChIP-qPCR, and dual-luciferase reporter assays. Our results showed that UBE3A was significantly upregulated in pancreatic cancer tissues and correlated with poor patient outcomes. Functional studies have demonstrated that UBE3A inhibits cellular senescence in pancreatic cancer cells, thereby promoting tumor proliferation and metastasis. Mechanistically, the histone variant macroH2A1 (mH2A1) recruits EZH2 to mediate H3K27me3 modification, repressing Telomerase reverse transcriptase (TERT) transcription. UBE3A interacts with mH2A1 through its N-terminal domain, leading to K48-linked polyubiquitination at the K167 residue, which accelerates mH2A1 degradation and upregulates TERT, enhancing the anti-senescence capacity of pancreatic cancer cells. Inhibition of UBE3A combined with the senolytic agent ABT‑263 induced apoptosis and inhibited tumor growth. The UBE3A/mH2A1/TERT axis enhances the anti-senescence capacity of pancreatic cancer cells and drives malignant progression, suggesting that UBE3A may serve as a novel therapeutic target for pancreatic cancer.
Similar content being viewed by others
Introduction
Pancreatic cancer is characterized by poor prognosis and high malignancy, with a 5-year survival rate of less than 10% [1]. Most patients are diagnosed with pancreatic cancer at an advanced stage, eliminating the possibility of surgical treatment; however, operative resection is the primary therapeutic modality. Chemotherapy, radiotherapy, and targeted therapies are currently available as pancreatic cancer treatment approaches; however, pancreatic cancer is markedly resistant to all these therapeutic modalities. Therefore, the identification of new and effective treatment strategies for pancreatic cancer is paramount.
Initially, senescence was described as the cessation of diploid cell proliferation caused by telomere shortening [2]. With advancing research, multiple senescence mechanisms have been identified across various cell types, including cancer cells [3]. Tumor cell senescence can occur early in oncogenesis. In early PanIN lesions, KRAS activation induces oncogene-induced senescence via the upregulation of p16, p53, and p21, thereby restraining tumor progression [4]. Anticancer therapies frequently induce cancer cell senescence through genotoxic or oxidative stress, leading to cell-cycle arrest and a senescence-associated secretory phenotype (SASP). Therefore, the induction of tumor senescence is considered a viable antitumor strategy [5]. However, persistent senescent cells can promote inflammation and suppress immune function, making their clearance critical for limiting tumor progression and preventing adverse effects. Notably, senescent cells undergo chromatin structural remodeling that alters transcriptional regulation and exposes exploitable vulnerabilities, enabling selective targeting by senolytic agents [6]. For example, small-molecule inhibitors targeting upregulated anti-apoptotic BCL-2 family proteins are a significant research focus [7]. Several studies have shown that Navitoclax (ABT-263), an inhibitor of BCL-2, BCL-XL, and BCL-W, can eliminate various senescent cancer cells by reactivating apoptotic pathways [8]. Thus, combining the induction of tumor cell senescence with senolytic agents may suppress tumor growth and remove deleterious senescent cancer cells, offering a promising avenue for more effective cancer therapies.
As cells senesce, the balance between protein synthesis and degradation is disrupted, resulting in the accumulation of abnormally folded proteins within the cell [9]. This perturbation impairs tumor cell function and heightens intracellular stress responses, inhibiting proliferation and survival. The ubiquitin–proteasome system (UPS), primarily responsible for regulating protein degradation, is a central mechanism for maintaining intracellular protein stability [10]. By modulating the UPS function, tumor cells can preserve proteostasis and cope with endogenous and environmental stresses [11]. Such regulation helps to alleviate oxidative and metabolic stress, slows the progression of cellular senescence in tumor cells, and may further drive tumor progression toward more aggressive phenotypes. Therefore, elucidating the molecular mechanisms by which the UPS regulates tumor cell senescence, identifying key anti‑senescence targets, and developing corresponding intervention strategies could open new avenues for enhancing cancer therapy.
Ubiquitin protein ligase E3A (UBE3A), also known as E6-associated protein (E6AP), belongs to the HECT protein family [12]. UBE3A accepts ubiquitin from E2 ubiquitin-conjugating enzymes via the formation of thioester linkages and transfers it to substrate proteins, marking them for degradation [13]. The N-terminus of UBE3A contains a Zn2+-binding AZUL domain responsible for substrate recruitment. Moreover, its C-terminus comprises a catalytic HECT domain, which is closely associated with Angelman syndrome, a neurodevelopmental disorder [14]. Additionally, UBE3A expression gradually decreases in the human cerebral cortex during aging [15]. Downregulation of UBE3A has also been linked to increased skin wrinkling in mammals, suggesting its potential as an anti-senescence agent [16]. Interestingly, the role of UBE3A in tumorigenesis has garnered significant attention [17,18,19]; however, its specific function in pancreatic cancer remains unclear. Therefore, exploring the role of UBE3A in pancreatic cancer and its potential anti-senescence mechanisms is critically important.
In addition to canonical histone components, histone variants exhibit unique genomic distributions. These variants can be incorporated into nucleosome subunits in a DNA replication-independent manner, replacing the existing nucleosome components and regulating gene transcription. Among them, the histone variant macroH2A1 (mH2A1), a member of the H2A family, replaces H2A in certain nucleosomes and acts as a transcriptional repression signal during gene transcription regulation [20]. Studies have demonstrated [21, 22] that mH2A1 interacts with EZH2 and EP300 within the genome, modulating downstream gene expression through epigenetic remodeling; thus, mH2A1 has been implicated in cellular senescence. Indeed, mH2A1 is involved in SASP [23] and senescence-associated heterochromatin foci (SAHF) formation [20]. Although previous studies have indicated that mH2A1 is downregulated in various cancers [24,25,26,27] and suppresses tumor growth, its specific role in pancreatic cancer has not yet been reported, highlighting the need for further investigation.
Telomerase plays a key role in regulating cellular senescence, with its activity directly determining a cell’s proliferative capacity [28]. In normal somatic cells, telomeres progressively shorten with each successive cell division, ultimately leading to replicative senescence. In contrast, telomerase reactivation and telomere length maintenance confer replicative immortality on cancer cells [29]. During the initiation and progression of pancreatic cancer, the mechanisms underlying telomerase function exhibit distinct characteristics. Although telomerase reactivation in most malignancies is commonly attributed to mutations in the telomerase reverse transcriptase (TERT) promoter, pancreatic cancer is distinct: restoration of telomerase activity in this tumor type is primarily associated with epigenetic alterations at the TERT locus [30]. Studies indicate that targeted inhibition of TERT promoter activity can effectively slow the proliferation and dissemination of pancreatic cancer cells [31]. Despite several regulatory mechanisms having been described, the whole regulatory network controlling TERT in pancreatic cancer remains incompletely understood.
In this study, we found that UBE3A is essential for antagonizing senescence in pancreatic cancer, and this function depends on its E3 ubiquitin ligase activity. We identified mH2A1 and TERT as downstream targets through mass spectrometry and high-throughput sequencing analyses. We demonstrate that UBE3A directly promotes K48-linked ubiquitination of mH2A1 at lysine 167 (K167), leading to its proteasome-mediated degradation. Degradation of mH2A1 reduces EZH2 binding and H3K27me3 levels at a TERT enhancer region, thereby facilitating TERT transcription. The UBE3A–mH2A1–TERT axis is critical in pancreatic cancer cell resistance to senescence. Notably, inhibition of UBE3A in combination with ABT-263 markedly suppresses pancreatic tumor growth. These results suggest that targeting UBE3A may represent a potential therapeutic strategy for pancreatic cancer.
Materials and methods
Bioinformatics Analysis
We performed weighted gene coexpression network analysis (WGCNA) to construct a weighted coexpression network using the GSE16515, GSE15471, and GSE28735 datasets from the Gene Expression Omnibus (GEO) [32]. After identifying modules that were significantly associated with tumors, they were compared with 466 senescence-related genes from the CellAge database. This process identified ten genes that were differentially expressed in relation to pancreatic cancer prognosis. Univariate Cox and LASSO regression analyses were conducted using R’s survival and glmnet packages (version 4.2.1). Prognostic correlations of UBE3A expression were evaluated using RNA-seq and clinical information from The Cancer Genome Atlas (TCGA) [33] and Genotype-Tissue Expression Project (GTEx) databases. Functional enrichment analyses of KEGG and GO pathways were performed using the clusterProfiler package in R.
Rigid‑body protein docking analysis
PDB files of the target proteins were retrieved from UniProtKB; biologically relevant chains were retained, and non-protein heteroatoms were removed. Rigid‑body protein–protein docking was performed using GRAMM with default parameters; both partners were submitted simultaneously, and the top 10 poses were collected and ranked by docking score. Binding energy (reference threshold ≈ −4.0 kcal·mol−1) was used as the primary selection criterion. The highest‑scoring pose was further evaluated for buried surface area, hydrogen bonds, and key interacting residues. Binding free energies and interface contacts were calculated using PDBePISA, and the results were visualized in PyMOL 3.1.
Human Samples
Tumor and adjacent tissue samples were collected from 60 patients with pancreatic cancer who underwent surgical resection at the Affiliated Hospitals of Guizhou Medical University. Among these patients, 35 were male and 25 were female, with an average age of 56.75 ± 8.6 years. The inclusion criteria were as follows: 1) a postoperative pathological diagnosis of pancreatic cancer; 2) no prior treatment before admission; and 3) patients who were conscious and able to communicate effectively. The exclusion criteria included: 1) patients with other tumors or severe infections; 2) patients with a history of treated pancreatic cancer or recurrence after treatment; 3) patients who voluntarily requested to withdraw from the study after enrollment; and 4) patients who were confused and unable to communicate normally. There was no attrition or dropout among the 60 patients whose clinical data were collected. Ethical approval for collecting samples and clinical data was granted by the Ethics Committee of Guizhou Medical University (Ethics Review No. 073 of 2019). All patients provided informed consent before participating in the study, ensuring that they understood the nature and purpose of the research. To minimize bias and enhance the study’s objectivity, blinding procedures were implemented during sample collection and data analysis. Personnel collecting tumor and adjacent tissue samples were blinded to the specific research objectives, ensuring impartiality in the handling of samples. Additionally, data analysts were blinded to patient group assignments during gene expression analyses and clinical data evaluations, allowing an unbiased interpretation of the results. The sample size was determined based on the results of the a priori effect size analysis.
Cell Lines
The pancreatic cancer cell lines AsPC-1, BxPC-3, MIA PaCa-2, PANC-1, and SW1990, along with the human pancreatic ductal epithelial cell lines HPDE6-C7 and tool cell HEK293T, were obtained from the American Type Culture Collection. These cells were cultured in DMEM and RPMI 1640, supplemented with antibiotics and fetal bovine serum, at 37 °C in a humidified incubator with 5% CO2. All cell lines were routinely authenticated and checked for contamination using short tandem repeat (STR) profiling and mycoplasma testing.
Construction of vectors for transfection
The siRNA used in this study was sourced from Sangon Biotech (China) and was transfected using the Lipofectamine 3000 Transfection Kit (Invitrogen, USA). The pGEX‑6 P‑1 vector was purchased from Sangon Biotech (China). Firefly luciferase vectors (pGL4.23 and pGL4.10) and Renilla luciferase vector (pGL4.74) were procured from Promega (USA). Plasmids encoding UBE3A, mH2A1 and H2A1 (GV657, GV658, GV141), lentiviral constructs for UBE3A overexpression and knockdown (GV298, GV341), lentiviral constructs for mH2A1 overexpression and knockdown (GV492, GV493), a TERT knockdown lentiviral construct (GV358), and an EZH2 lentiviral construct (GV298, GV341) were all generated by GeneChem (China). Experimental procedures were performed according to established protocols. See Table S3 for the referenced sequence information.
Antibodies and reagents
The antibodies and reagents used in this study are listed in Table S4.
Immunohistochemistry(IHC)
After deparaffinization and hydration, the prepared paraffin sections were sequentially subjected to blocking, removal of endogenous peroxidase, and antigen retrieval. The primary antibody was then added dropwise to the tissue sections, which were incubated overnight at 4 °C. The next day, the secondary antibody was applied dropwise to the tissue sections, followed by incubation at room temperature for 1 h. Finally, the tissue sections were stained with a DAB chromogenic solution and DAPI, and then visualized under an optical microscope. Immunohistochemical scoring was performed by two pathologists based on the staining intensity and percentage of positive cells. The staining intensity was graded on a four-point scale as follows: negative (0 points), weakly positive (1 point), positive (2 points), and strongly positive (3 points). The percentage of positive cells was scored as follows: less than 1% (0 points), 1-25% (1 point), 26-50% (2 points), 51-75% (3 points), and > 75% (4 points). The final score was the product of these two values. Scores below seven were classified as low expression, whereas scores above 7 were classified as high.
Multiplex Immunohistochemistry(mIHC)
The mIHC experiments were performed using a multiplex fluorescent IHC kit (Absin, China). Briefly, the tissue sections were deparaffinized, hydrated, subjected to antigen retrieval, and blocked. Primary and secondary antibodies were sequentially applied dropwise for subsequent incubation, followed by fluorescence staining. This process was repeated to label all the target genes. DAPI was subsequently used to stain the cell nuclei. Stained sections were observed and imaged using a fluorescence microscope (Nikon, Tokyo, Japan).
Immunofluorescence Colocalization Assay
The cells were fixed with 4% paraformaldehyde and permeabilized with 0.5% Triton X-100 for 20 min. After blocking, primary antibodies were added, and the samples were incubated overnight at 4 °C in the dark. Subsequently, fluorescently labeled secondary antibodies were added, and the samples were incubated at room temperature in the dark for 2 hours. Nuclear staining was performed, and the slides were observed and imaged using a fluorescence microscope.
Haematoxylin and Eosin(H&E)
H&E staining was performed using a Beyotime H&E staining kit (China). The tissue samples were first deparaffinized and hydrated. Then the sections were immersed in a hematoxylin solution for 10 minutes, followed by differentiation for a few seconds in a differentiation solution. The sections were then rinsed with running tap water for 10 min. An eosin stain was applied for 2 minutes, and the excess stain was removed with distilled water. Finally, the sections were dehydrated, cleared, and observed under a microscope.
Quantitative Real-Time PCR(qRT-PCR)
Total RNA was extracted from cells and tissues using an RNA extraction kit (ES Science, China) and reverse-transcribed to generate complementary DNA. PCRs were performed using TB Green® Premix Ex Taq™ reagent (Takara, Japan) on a CFX96 Real-Time PCR Detection System (Bio-Rad, USA). The data were analyzed using the 2−ΔΔCt method. Primers used for qRT-PCR are listed in Table S5.
Western blotting
After the proteins were successfully extracted from the tissues or cells, the protein concentrations in the samples were determined using a BCA protein assay kit (Solarbio, China). The samples were separated by electrophoresis and transferred onto PVDF membranes (Millipore, USA). The membranes were blocked with 5% nonfat milk and incubated overnight at 4 °C with primary antibodies. The following day, membranes were incubated with secondary antibodies at room temperature for 2 h. The color-developing solution was added dropwise to the PVDF membranes before analysis.
Senescence-Associated β-Galactosidase(SA-β-gal) Activity Assay
SA-β-gal activity was assessed using an SA-β-gal staining kit (Beyotime, China). The cells were first washed with PBS and fixed with β-galactosidase staining fixative. β-Galactosidase staining solution was added, and the cells were incubated overnight at 37 °C. SA-β-gal-positive cells were observed and quantified the following day under an optical microscope.
Cell Viability Assay
The CCK-8 reagent (Solarbio, China) was added to each well of the 96-well plates, and the absorbance of each well was measured using a microplate reader.
5-Ethynyl-2’-deoxyuridine(EdU) Assay
The experiment used the Click-iT EdU-555 Cell Proliferation Assay Kit (Servicebio, China). When the cells reached the logarithmic growth phase, the EdU stock solution was added, and the mixture was incubated for 2 h. After fixation and permeabilization, EdU and nuclear staining were performed. The proportion of EdU-positive cells was determined by fluorescence microscopy.
Cell Cycle Analysis
A cell cycle detection kit (Beyotime, China) was used for all the experiments. First, the cells were harvested by trypsinization to collect the cell pellet, which was then washed with pre-cooled PBS. After fixation, propidium iodide staining solution (0.5 mL) was added to each tube, followed by incubation at 37 °C in the dark for 30 min. Finally, the detection and analysis were performed using a flow cytometer.
Detection of Apoptosis
Apoptosis was detected using an Annexin V-FITC Apoptosis Detection Kit (Beyotime, China). A total of (5-10)×10⁴ cells were collected as pellets, to which 195 µL of Annexin V-FITC binding buffer, 5 µL of Annexin V-FITC, and 10 µL of PI staining solution were added in sequence with gentle mixing. The cells were incubated at room temperature in the dark for 20 min, after which flow cytometry was performed for detection and data analysis.
Enzyme-Linked Immunosorbent Assays(ELISAs)
ELISAs were performed using an ELISA kit (Jonlnbio, China). Briefly, 100 µL of the working standard solution or sample mixture was added to each well of a precoated microplate containing capture antibodies for IL-6, IL-8, CXCL1, CXCL6, and MMP3. The plates were then incubated at 37 °C for 60 min. After discarding the liquid, 300 µL of biotin-conjugated antibody working solution was added to each well and incubated at 37 °C for 60 min. Following three washes, 100 µL of HRP-conjugated secondary antibody solution was added to each well for incubation at 37 °C for 30 minutes. Subsequently, 50 µL of TMB substrate was added, and the plate was incubated at 37 °C in the dark for 15 min. Sample concentrations were calculated based on a standard curve.
Trypan blue staining
Pancreatic cancer cells were collected after trypsinization and resuspended in PBS, and the cell concentration was adjusted. The cell suspension was mixed with trypan blue solution, gently pipetted to mix, and incubated briefly at room temperature. An aliquot was applied to a hemocytometer and examined under light microscopy; cells stained uniformly blue were regarded as nonviable, while unstained, translucent cells were considered viable.
Coimmunoprecipitation(co-IP)
The cells were fully lysed with NP-40, and the protein supernatant was collected. Protein A + G magnetic beads (Beyotime, China) were added to the protein supernatant for pre-clearing, and the protein mixture was retained after discarding the beads. The fully conjugated magnetic bead‒antibody complex was added to the protein supernatant and incubated overnight at 4 °C. The beads were washed, separated using TBS, and retained the following day. After elution from the beads, the supernatant was collected for the subsequent experiments.
Immunoprecipitation–mass spectrometry (IP-MS)
IgG magnetic beads were used for pre-purification (pre-clearing) of pancreatic cancer cell samples, followed by immunoprecipitation and elution for mass spectrometry analysis. Mass spectrometry was performed on a Bruker timsTOF Pro instrument coupled to an UltiMate 3000 RSLCnano system. Peptides were separated on a C18 chromatography column, and data were acquired in the PASEF mode. The data were searched against the UniProt Human database (release 2023-06-19) using MaxQuant version 2.0.1.0 in conjunction with Andromeda. Enzyme specificity was set to Trypsin/P with up to two missed cleavages. Variable modifications included Oxidation, Acetylation, and Deamidation; fixed modification was Carbamidomethylation of Cys. The false discovery rate (FDR) was set to 1%. Downstream analyses and plotting were performed.
DIA-based ubiquitin proteomics analysis
UBE3A-overexpressing and vector-control pancreatic cancer cells were used for DIA ubiquitinome analysis. Proteins were extracted according to the APT SOP, and 15 μg of protein were used for SDS-PAGE to assess sample consistency. In-solution digestion with trypsin was performed using 10 mM DTT, 50 mM IAA, followed by acidification and desalting, at an enzyme-to-substrate ratio of 1:25 and incubation overnight at 37 °C. The digests were desalted on C18 and lyophilized. Ubiquitin remnant motif–based enrichment of K-ε-GG peptides was performed, and the enriched peptides were resuspended in 0.1% formic acid. DIA data were acquired on a Vanquish Neo HPLC system for chromatographic separation and analyzed on the Thermo Astral high-resolution mass spectrometer. MS1 scan range: 380–980 m/z; resolution: 240k; AGC target: 500%; maximum IT: 5 ms. MS2: 300 DIA windows, isolation window 2 m/z, HCD energy 25 eV, AGC target 300%, maximum IT 3 ms. Data were processed with Spectronaut using the following parameters: enzyme = Trypsin; maximum missed cleavages = 2; fixed modification = Carbamidomethyl (C); dynamic modifications = Oxidation (M), Acetyl (Protein N-term); identification of GlyGly (K) ubiquitin remnants; FDR < 1%.
Silver Staining
After electrophoresis, the gels were fixed for 20 min. The samples were washed with 30% ethanol and double-distilled water. The gel was sequentially treated with silver staining sensitizing solution, silver solution, and silver staining developer (Beyotime, China) at room temperature with gentle agitation, according to the manufacturer’s protocol, until the desired bands appeared. The reaction was terminated, and the gel was washed with double-distilled water. Finally, the gel was preserved, recorded, and photographed.
Coomassie Brilliant Blue Staining
After electrophoresis, the gel was washed five times with double-distilled water 5 times (5 min each time). A preheated Coomassie Brilliant Blue staining solution (Beyotime, China) was added, and the gel was placed on a horizontal shaker and agitated for 15 min to 1 h, until clear protein bands appeared. The excess background stain was removed by washing the gel with double-distilled water. The gel was then preserved and photographed.
GST Pull-Down Assay
Plasmids (UBE3A‑pGEX‑6p‑1, mH2A1‑pGEX‑6p‑1, and empty vector) were transformed into E. coli were inoculated into LB broth containing ampicillin and grown to an OD600 of ~0.5–0.6. At this point, protein expression was induced by adding IPTG, and the cells were then harvested. Bacterial pellets were lysed with lysis buffer containing lysozyme, and the lysates were centrifuged; the clarified supernatants were incubated with glutathione (GSH) affinity agarose magnetic beads, extensively washed, and the GST fusion proteins were recovered by elution. Myc‑mH2A1 or His‑EZH2 was expressed in HEK‑293 T cells; cell lysates were centrifuged, and the supernatants were bound to Myc‑ or His‑affinity resin, washed multiple times, and finally eluted by boiling in SDS‑PAGE sample buffer to obtain the tagged proteins. For the GST pull‑down assay, GST fusion proteins were first immobilized on glutathione magnetic beads, washed, incubated with Myc‑mH2A1 or His‑EZH2, and subjected to further washes. The beads were then treated with an elution buffer and boiled to recover the bound material. After adding the SDS-PAGE sample buffer, the samples were analyzed using a Western blot.
In Vivo Ubiquitination Assay
Pancreatic cancer cells from each treatment group were transfected with an HA‑Ub plasmid. Forty‑eight hours after transfection, cells were treated with MG132 (20 μM) for 6 hours and then harvested. Cells were lysed in NP‑40 lysis buffer, and mH2A1‑containing protein complexes were immunoprecipitated using anti‑mH2A1 antibody‑conjugated magnetic beads. The beads were washed and eluted to recover the mH2A1 complexes, which were analyzed by Western blotting and probed with an anti‑HA antibody to assess the cellular ubiquitination of mH2A1.
In vitro ubiquitination assay
An in vitro ubiquitination reaction was assembled in 40 mM Tris-HCl (pH 7.6), 2.5 mM Mg‑ATP, and 1 mM DTT. The reaction contained the E1 ubiquitin‑activating enzyme UBE1 (50 nM), the E2 ubiquitin‑conjugating enzyme UbcH7 (500 nM), HA‑ubiquitin (HA‑Ub, 200 μM), and the purified proteins to be tested (Myc‑mH2A1, Myc‑mH2A1 K167, and GST‑UBE3A). Reactions were incubated at 30 °C with shaking at 220 rpm for 30 minutes. After incubation, Myc-tagged proteins were immunoprecipitated using Myc affinity resin. The beads were washed and eluted to recover Myc‑mH2A1 complexes. The eluates were analyzed by Western blot to assess the ubiquitination status of mH2A1.
RNA-seq
The study included four biological replicates (mH2A1‑overexpressing PANC-1 cells versus control cells). Total RNA was extracted using TRIzol reagent and assessed for concentration and integrity using a NanoDrop spectrophotometer. Three micrograms of RNA were used as input for library preparation. mRNA was enriched using poly-T oligo-attached magnetic beads and fragmented in an Illumina proprietary fragmentation buffer containing divalent cations at an elevated temperature. First‑strand cDNA was synthesized with random primers and SuperScript II, followed by second‑strand synthesis using DNA Polymerase I and RNase H. Fragment ends were repaired using exonuclease/polymerase activities, enzymes were removed, and 3′ ends were adenylated before ligation of Illumina paired‑end adapters. Libraries were size‑selected for 400–500 bp fragments using AMPure XP, enriched by 15 cycles of PCR with Illumina PCR primers, purified with AMPure XP, and quantified on an Agilent 2100 Bioanalyzer using the High Sensitivity DNA assay. Libraries were sequenced on an Illumina NovaSeq 6000 platform. Raw sequencing reads underwent quality control and filtering to generate clean data, which were then aligned to the reference genome. Gene expression was quantified from the alignments, and differential expression analysis and subsequent downstream analyses were performed.
ChIP‒qPCR
The interactions between the proteins and DNA were analyzed using the SimpleChIP Enzymatic Chromatin IP Kit (CST, USA). Briefly, cells were fixed with formaldehyde, and the DNA was digested into fragments of an appropriate length using a micrococcal nuclease and sonicated. Once the DNA length and concentration met the required standards, the samples were incubated overnight with rotation at 4°C using a primary antibody. After elution, crosslink reversal, and DNA purification, PCR amplification was performed using specific primers. Primers used for ChIP-PCR are listed in Table S5.
ChIP‒seq
Immunoprecipitation in PANC-1 cells was performed using a ChIP-grade antibody against mH2A1. The workflow comprises: (1) formaldehyde fixation, quenching, cell lysis and nuclear isolation; (2) extraction of DNA–protein complexes and sonication to yield DNA fragments of 200–1000 bp; (3) immunoprecipitation, washing, elution, reversal of crosslinks and DNA purification; (4) quality assessment of the immunoprecipitated DNA and evaluation of IP efficiency by protein detection, followed by ChIP-seq library preparation. Library construction involves end repair and A-tailing, adapter ligation, magnetic bead cleanup, PCR amplification, quality control (using agarose gel electrophoresis or instrument-based QC), and Qubit quantification. After QC passes, the libraries are sequenced on the MGISEQ-T7 platform. Raw reads are filtered to obtain clean data, then aligned to the reference genome. Subsequently, genome-wide peak calling, binding site distribution analysis, and motif analysis were performed.
Dual-Luciferase Reporter Assay
The constructed reporter plasmid and the Renilla luciferase internal control plasmid were co-transfected into pancreatic cancer cells from each treatment group. A Dual-Luciferase Reporter Gene Assay Kit (Beyotime, China) was used for the analysis. After removing the cell culture medium, cell lysis buffer was added, and the supernatant was collected after thorough lysis. For each sample, 100 μL of firefly luciferase assay reagent was added. After mixing, the relative light unit (RLU) values were measured using a multimode microplate reader. Subsequently, 100 μL of Renilla luciferase assay reagent was added to the same samples, and after mixing again, RLU values were measured a second time. Activation of the target reporter gene in various samples was assessed by comparing the ratio of the firefly luciferase RLU value to the Renilla luciferase RLU value.
Animal studies
The Animal Ethics Committee of Guizhou Medical University approved the project (No. 19000672). Female BALB/c nude mice aged 6-8 weeks were procured from Collective Pharmaceuticals and maintained under controlled environmental conditions (temperature: 22 ± 1°C; relative humidity: 50 ± 1%; 12-hour light/dark cycle). Based on the effect size and variability reported in previous literature, we estimated that five mice per group would be necessary to ensure sufficient statistical power to detect significant changes in tumor growth and metastasis. To reduce bias and enhance the credibility of the experimental results, this study employed a blind methodology. Throughout the experiment, all tumor volume measurements, excisions, and weight recordings were conducted by independent researchers unaware of the group assignments to ensure the objectivity of the results. Additionally, during the data analysis phase, the analysts remained blind to the group information to maintain objectivity in the analytical process. To establish a subcutaneous tumor model, an appropriate volume of pancreatic cancer cells was injected into the right axilla of each mouse, and the tumor volume was measured at regular intervals. The mice were randomly assigned to different experimental groups to minimize bias. After seven weeks, the mice were euthanized, and the subcutaneous tumors were excised, weighed, and photographed. The mice were randomly assigned to groups for the liver metastasis model. After anesthetization, pancreatic cancer cells were carefully injected into the superficial layer of the spleen, ensuring no bleeding or leakage occurred at the injection site. The growth of the primary splenic tumor and liver metastases was monitored using bioluminescence imaging with an IVIS® Lumina III system. All animal experiments, including euthanasia protocols, adhered strictly to the animal management guidelines and standards set forth by Guizhou Medical University.
Statistical analysis
SPSS (USA) and GraphPad Prism (USA) were used for data processing and analysis. The associations between clinicopathological characteristics and UBE3A expression were evaluated using the chi-squared test. Comparisons between two groups were conducted using a two-tailed Student’s t-test, whereas comparisons among multiple groups were performed using analysis of variance (ANOVA). The assumption of homogeneity of variances was assessed using Levene’s test. If Levene’s test indicated equal variances (p > 0.05), standard t-tests and ANOVA were used; if not (p ≤ 0.05), Welch’s t-test and, for multiple comparisons, Welch’s ANOVA or appropriate post hoc tests (e.g., Games-Howell) were applied. Survival analysis was performed using the Kaplan-Meier method, and differences between groups were evaluated using the log-rank test. The Cox proportional hazards model was used for multivariate analysis. Statistical significance was set at p < 0.05.
Results
Screening Oncogenic Factors Associated with Cellular Senescence
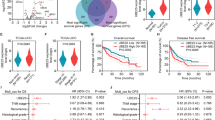
We used WGCNA and LASSO regression to identify aging-related oncogenes in pancreatic cancer. Initially, we integrated and corrected batch effects in the GSE28735, GSE15471, and GSE16515 datasets to generate a merged gene expression matrix. A scale-free coexpression network was subsequently constructed using WGCNA, with the optimal soft threshold power (β) set to seven with a scale-free topology fit index of 0.85 (Supplementary Fig. S1A-B). Based on the correlation with the “tumor” trait, we identified 16 candidate gene modules (Supplementary Fig. S1C). Among these modules, the “magenta” module was highly correlated with “tumor” (r = 0.85, P = 1.0e-170) (Supplementary Fig. S1D). We then cross-referenced the 77 genes in the “magenta” module with 466 senescence-associated genes listed in the CellAge database (Supplementary Fig. S1E), identifying ten key genes that are risk factors for pancreatic cancer prognosis (Supplementary Fig. S1F). These ten oncogenic factors were included in the LASSO model (Supplementary Fig. S1G-H) to establish a prognostic risk score as follows: risk score (-0.0258)×AGR2 + (0.1412)×FERMT1 + (0.1758)×UBE3A + (0.0894)×LGALS3 + (0.1574)×KRT19.
Notably, UBE3A exhibited the highest coefficient weights in the model. Further analysis using TCGA, GTEx, and UALCAN databases revealed that UBE3A was highly expressed in pancreatic cancer tissues (Supplementary Fig. S2A-B). Patients with high UBE3A expression showed significantly reduced overall survival (OS), disease-specific survival (DSS), and progression-free interval (PFI) (Supplementary Fig. S2C-E). GO and KEGG pathway analyses revealed that UBE3A was associated with pathways related to cellular senescence and telomerase regulation (Supplementary Fig. S2F-G). These findings prompted us to investigate the molecular mechanisms by which UBE3A regulates senescence in pancreatic cancer cells.
Validation of UBE3A Expression Levels in Pancreatic Cancer
To investigate UBE3A expression, we collected 60 paired pancreatic cancer tissue samples. qRT-PCR, Western blotting, and IHC analyses consistently demonstrated that UBE3A expression was significantly upregulated in pancreatic cancer tissues (Supplementary Fig. S3A-C). In addition, compared with that in adjacent tissues, the UBE3A score in tumor tissues increased, as shown by IHC (Supplementary Fig. S3D). High UBE3A expression was associated with reduced OS in pancreatic cancer patients (Supplementary Fig. S3E). We further explored the relationship between UBE3A expression and clinicopathological features of patients with pancreatic cancer. High UBE3A expression was significantly correlated with advanced TNM stage, larger tumor size, distant metastasis, perineural invasion, and lower tumor differentiation (Table S1). Cox regression analysis of clinicopathological features and prognosis revealed that UBE3A is an independent risk factor for poor outcomes in patients with pancreatic cancer (Table S2). Additionally, UBE3A expression was markedly increased in pancreatic cancer cell lines (Supplementary Fig. S3F-G). Based on these findings, we selected MIA PaCa-2 cells, which endogenously express low levels of UBE3A, and PANC-1 cells, which endogenously express high levels of UBE3A, for subsequent experiments.
Validation of the role of UBE3A in the regulation of cellular senescence
To modulate UBE3A expression in pancreatic cancer, we designed three siRNAs, among which si-UBE3A#2 and si-UBE3A#3 exhibited the most effective knockdown efficiency (Supplementary Fig. S4A, B). Lentiviral vectors for UBE3A overexpression and knockdown were constructed and successfully used to infect MIA PaCa-2 and PANC-1 cells (Supplementary Fig. S4C–F).
The SA-β-gal assay revealed that in pancreatic cancer cells cultured for 50 consecutive passages, the ratio of SA-β-gal-positive cells (Fig. 1A-B) was substantially reduced when UBE3A was overexpressed. Moreover, the UBE3A knockdown increased the SA-β-gal-positive cell ratio (Supplementary Fig. S5A, B). The CCK-8 assay results demonstrated that UBE3A overexpression resulted in a faster proliferation rate (Fig. 1C), whereas UBE3A knockdown reduced the proliferation rate of pancreatic cancer cells (Supplementary Fig. S5C). Similarly, there was a significant increase in the proportion of EdU-positive cells among the UBE3A-overexpressing pancreatic cancer cells (Fig. 1D, E). In contrast, the EdU-positive cell rate decreased in the UBE3A-knockdown cells (Supplementary Fig. S5D, E). Flow cytometric analysis revealed that UBE3A overexpression accelerated cell cycle progression in pancreatic cancer cells (Fig. 1F, G), whereas UBE3A knockdown induced cell cycle arrest at the G0/G1 phase (Supplementary Fig. S5F, G). Furthermore, UBE3A overexpression reduced the levels of SAHF markers (Fig. 1H, I); conversely, UBE3A knockdown in pancreatic cancer cells led to opposite trends in SAHF marker levels (Supplementary Fig. S5H, I). We conducted ELISAs to evaluate the SASP. UBE3A overexpression significantly decreased the secretion of SASP factors, including IL-6, IL-8, CXCL1, CXCL6, and MMP3 in the culture supernatants of pancreatic cancer cells (Fig. 1J), whereas UBE3A knockdown increased the secretion of these SASP markers (Supplementary Fig. S5J).
A, B SA-β-gal staining was used to assess the senescence level of MIA PaCa-2 cells. Experiments were performed in three independent biological replicates (n = 3). For each replicate, three fields were imaged and counted under a 20× objective, and the mean of these three fields was taken as the value for that replicate. The number of SA-β-gal-positive cells and the total number of cells were recorded, and the percentage of positive cells was calculated as (positive cells / total cells) × 100%. C Cell proliferation was assessed using the CCK-8 kit. Experiments were performed in three independent biological replicates (n = 3); each treatment was assayed in three technical replicates for each biological replicate. Absorbance was measured at 450 nm using a microplate reader, and values were background-corrected by subtracting the absorbance of wells containing medium only. For each biological replicate, the mean of the technical replicates was used as the replicate value. D, E EdU staining was used to assess cellular DNA synthesis. Experiments were performed in three independent biological replicates (n = 3). Three randomly selected fields were imaged and counted for each biological replicate under a fluorescence microscope at 40× magnification. The mean of these three fields was taken as the value for that replicate. The EdU-positive rate was calculated as (EdU-positive cells / total cells) × 100%. F, G Flow cytometry was used to assess cell-cycle distribution. Experiments were performed in three independent biological replicates (n = 3). For each biological replicate, cells were fixed, stained with propidium iodide, and analyzed by flow cytometry. A minimum of 10,000 gated single-cell events were acquired per replicate, with debris and doublets excluded. Cell-cycle phase distributions (G0/G1, S, and G2/M) were calculated using cell-cycle analysis software. H, I Western blotting was used to assess the expression of SAHF markers in pancreatic cancer cells across experimental groups. Experiments were performed in three independent biological replicates (n = 3). Representative blots are shown; band intensities were quantified by densitometry, normalized to α-tubulin. J ELISAs were used to quantify SASP-related factors in cell culture supernatants. Each group was analyzed in three independent biological replicates (n = 3); within each biological replicate, samples were assayed in technical triplicates according to the manufacturer’s instructions. Absolute concentrations were calculated from standard curves. **P < 0.01, ***P < 0.001, ****P < 0.0001. P-values were calculated using a paired, two-tailed Student’s t-test. Final data are presented as mean ± SEM of three independent experiments.
UBE3A promotes pancreatic cancer growth and metastasis
We first established subcutaneous tumors in nude mice to determine whether UBE3A affects the growth of pancreatic cancer cells in vivo. The results revealed that UBE3A overexpression led to a significant increase in both tumor volume and weight (Fig. 2A–C). In contrast, UBE3A knockdown markedly reduced the tumor volume and weight (Fig. 2D–F). Furthermore, a liver metastasis model was constructed by injecting pancreatic cancer cells overexpressing UBE3A or UBE3A knockdown into the subcapsular region of the spleen of nude mice. In this model, UBE3A overexpression significantly increased the number of liver metastatic lesions, whereas UBE3A knockdown reduced or eliminated liver metastases (Fig. 2G–I). This observation was further corroborated by H&E staining of the liver metastatic lesions (Fig. 2J, K).
A–F Pancreatic cancer cell suspensions were injected subcutaneously into the right axilla of nude mice. Animals were assigned to Vector, UBE3A, sh-Ctrl, sh-UBE3A#2, and sh-UBE3A#3 groups; tumor volume and weight were recorded and compared. G–I Primary splenic tumors and hepatic metastatic lesions were measured and analyzed for volume and fluorescence intensity. J, K Representative H&E-stained images of liver metastases. Quantitative fluorescence was measured using a small animal in vivo imaging system. Tumor volume was measured with calipers, and excised tumors were weighed using an electronic balance. Data are presented as mean ± SEM. Statistical comparisons were performed using two-tailed Student’s t-tests; significance is indicated as ***P < 0.001 and ****P < 0.0001.
UBE3A regulates the expression of the mH2A1 protein
To further elucidate the regulatory role of UBE3A in pancreatic cancer, we performed ubiquitinome profiling on a paired set of PANC‑1 cell samples (UBE3A‑overexpressing and vector control) to identify UBE3A‑mediated ubiquitination events. Ub-MS analysis revealed 435 differentially expressed ubiquitinated peptides corresponding to 355 ubiquitinated proteins, with the majority located in the nucleus and the remainder in the cytoplasm (Figure S6A, B). In the UBE3A overexpression group, 220 ubiquitinated peptides were upregulated, and 215 were downregulated (Figure S6A). Notably, more than 57.72% of proteins were ubiquitinated at two or more sites, with an average ubiquitination frequency of 0.77% per protein (Figure S6C, D). Enrichment analysis indicated these proteins were closely associated with proteasomal metabolism, chromatin remodeling, nucleosome assembly, aging regulation, and telomere maintenance (Figure S6E). KEGG pathway analysis revealed that UBE3A-mediated ubiquitination is critical in modulating cancer-associated transcriptional dysregulation (Figure S6F). Identified ubiquitination substrates include the histone H2A domain (Figure S6G). In addition, we performed immunoprecipitation coupled with mass spectrometry (IP‑MS) in PANC-1 cells using an anti-UBE3A antibody to identify UBE3A-interacting proteins and potential substrates. Using IP‑MS, we identified 85 UBE3A‑binding peptides corresponding to 62 candidate interacting proteins, including H4, RO40, LMNA, mH2A1, and PARP1 (Figure S6H; Figure S6K). The peptides ranged in length from 7 to 22 amino acids, with 14-residue peptides being the most prevalent (Figure S6I). The IP-MS dataset was of high quality, displaying virtually no missed-cleavage peptides (Figure S6J). By comparing these candidate proteins with genes related to telomere regulation, we identified mH2A1 as a possible effector molecule (Figure S6L). In addition, we identified multiple ubiquitination sites on mH2A1, including K116, K123, K167, and K304; ubiquitination at K167 was the most statistically significant, with a Significance B value of 0.026 (Fig. S6M).
Immunofluorescence colocalization assays revealed that UBE3A and mH2A1 share similar subcellular localization, predominantly in the nucleus (Fig. 3A–C). Silver staining analysis revealed a distinct protein band at approximately 40 kDa in the IP group compared to the IgG control group, likely due to the molecular weight of mH2A1 (Fig. 3D). Subsequent co-IP experiments confirmed that UBE3A interacts with mH2A1 in pancreatic cancer cells (Fig. 3E, F). Additionally, GST pull-down assays demonstrated that UBE3A is directly bound to mH2A1 in vitro (Fig. 3G). To explore the structural basis of this interaction, we constructed two UBE3A recombinant proteins: Flag-UBE3A ΔN (amino acids 401-875) and Flag-UBE3A ΔC (amino acids 1-400) (Fig. 3H). Co-IP experiments revealed that Myc-mH2A1 coprecipitated with full-length Flag-UBE3A (FL) and Flag-UBE3A ΔC, but not with Flag-UBE3A ΔN. These findings indicated that the N-terminal domain of UBE3A is critical for its interaction with mH2A1 (Fig. 3I). Similarly, we generated four recombinant mH2A1 proteins, Myc-mH2A1 (amino acids 1-120), Myc-mH2A1 (amino acids 1-181), Myc-mH2A1 (amino acids 182-369), and Myc-mH2A1 (amino acids 121-369) (Fig. 3J). Co-IP experiments demonstrated that Flag-UBE3A coprecipitated with full-length Myc-mH2A1, the 121-369 fragment, and the 182-369 fragment but not with the 1-120 or 1-181 fragments. These results indicate that the region of mH2A1 containing amino acid residues 182-369 is essential for its interaction with UBE3A (Fig. 3K). We further performed rigid docking on the GRAMM platform; the top-ranked conformation had a binding energy of −4.9 kcal·mol−1 and formed multiple hydrogen bonds at the interface (Fig. 3L–N). UBE3A THR-348 and mH2A1 PRO-266 are likely the primary interaction sites, as they reside within the experimentally validated interface (Fig. 3H–K) and form short hydrogen bonds (~2.5 Å).
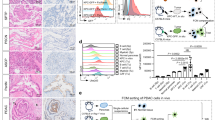
A–C Subcellular colocalization analysis. UBE3A (Red) and mH2A1 (Green) distribution was examined by immunofluorescence in PANC‑1 and MIA PaCa‑2 cells; two‑channel fluorescence intensities and overlap were extracted and quantified using ImageJ, and statistical analyses were performed in GraphPad Prism. Representative images are shown (experiments were independently repeated three times). D Silver staining following anti‑UBE3A immunoprecipitation was performed to validate precipitation efficiency; samples included Input, IgG control, and IP. The experiment was independently repeated three times, presenting a representative result. E, F Reciprocal Co‑IP experiments in two pancreatic cancer cell lines: forward Co‑IP using a UBE3A antibody to detect mH2A1 enrichment, and reverse Co‑IP using an mH2A1 antibody to detect UBE3A enrichment. All Co‑IP assays were independently repeated three times, and representative immunoblots are shown. G GST and GST‑UBE3A were expressed in Escherichia coli and affinity‑purified, while Myc‑mH2A1 was expressed in HEK293T cells and obtained using a purification kit. Recombinant protein purity was confirmed by Coomassie Brilliant Blue staining, followed by GST pull‑down assays to evaluate enrichment of Myc‑mH2A1 by GST/GST‑UBE3A. The experiment was independently repeated three times. H, I Full‑length Flag‑UBE3A (FL), N‑terminal deletion mutant (ΔN), and C-terminal deletion mutant (ΔC) were cotransfected with Myc‑mH2A1 in pancreatic cancer cells, and Co‑IP assays were performed to assess the contribution of UBE3A domains to the interaction. Experiments were independently repeated three times. J, K Full‑length Myc‑mH2A1 (FL) and fragments (1–120, 1–181, 182–369, 121–369) were cotransfected with Flag‑UBE3A in pancreatic cancer cells, and Co‑IP assays were conducted to map the mH2A1 interaction region. Experiments were independently repeated three times. L–N Protein–protein rigid docking was performed with GRAMM using UniProtKB‑derived structures; of the top 10 poses, the highest‑ranked conformation showed a binding energy of −4.9 kcal·mol−1, and PDBePISA/PyMOL analysis revealed a stable interface with multiple hydrogen bonds formed by key residues at several contact sites. O, P mH2A1 expression was evaluated at the mRNA level by qRT‑PCR and at the protein level by Western blot in Vector, UBE3A overexpression, sh‑Ctrl, sh‑UBE3A#2, and sh‑UBE3A#3 groups. All experiments included three biological replicates; data are presented as mean ± SEM. Group comparisons in panel O were performed using an independent‑sample, two‑sided Student’s t‑test. Q IHC was used to detect mH2A1 expression and distribution in 60 pancreatic cancer specimens and paired adjacent noncancerous tissues. R IHC scores for tumor versus adjacent tissue are compared. S–V To assess spatial expression patterns, Adjacent tissue sections were stained concurrently for UBE3A and mH2A1. W Spearman rank correlation analysis was performed on the IHC scores from the 60 cases to evaluate the relationship between UBE3A and mH2A1; the correlation coefficient (r) and p-value are reported. X Samples were dichotomized by an IHC score threshold into UBE3A high and low expression groups, and the percentages of mH2A1 high/low expression were compared between groups. For each specimen, three paraffin‑embedded sections were randomly selected as independent replicates for IHC. Tumor and adjacent non‑tumorous tissues were compared during IHC scoring. For each paraffin section, three microscopic fields within the target area were randomly chosen and scored independently. The arithmetic mean of these three field scores was taken as the final score for that specimen. Scoring was performed independently by two experienced pathologists under blinded conditions. Group differences in proportions were assessed by the chi‑square (χ2) test; the differences were statistically significant (**P < 0.01).
Then we validated its regulatory role in pancreatic cancer cells to investigate the potential mechanism by which UBE3A regulates mH2A1 expression. Western blotting and qRT-PCR indicated that UBE3A can affect mH2A1 protein expression only (Fig. 3O, P). IHC further demonstrated that mH2A1 expression was higher in the adjacent tissues than in the pancreatic cancer tissues (Fig. 3Q, R). Notably, mH2A1 was consistently expressed at lower levels in pancreatic cancer tissues than in UBE3A tissues(Fig. 3S–V), with IHC scores showing a negative correlation (Fig. 3W, X). These findings suggest that UBE3A may regulate mH2A1 via posttranslational modifications.
The UBE3A-Mediated Ubiquitination and Degradation of mH2A1 Drives Pancreatic Cancer Cell Resistance to Senescence
Proteasome inhibition with MG132 increased mH2A1 protein levels, whereas the lysosomal inhibitor NH4Cl had minimal effect (Fig. 4A, B). In cycloheximide (CHX) chase assays, MG132 delayed mH2A1 degradation in pancreatic cancer cells (Fig. 4C, D). Transfection with increasing amounts of wild-type UBE3A (UBE3A-WT) reduced both endogenous and exogenous mH2A1 levels in a dose-dependent manner, whereas the catalytically inactive mutant UBE3A-C833A did not (Fig. 4E–G). Consistent with a proteasome-dependent mechanism, MG132-treated pancreatic cancer cells exhibited no significant differences in mH2A1 levels whether UBE3A was overexpressed or knocked down (Fig. 4H, I). CHX chase experiments showed that UBE3A overexpression shortened the mH2A1 half-life, which was restored by UBE3A knockdown (Fig. 4J, K, Fig. 4S, T).
A Treatment with 20 μM MG132 caused a time‑dependent increase in mH2A1 protein levels in pancreatic cancer cells. B Treatment with 2.5 mM NH4Cl did not alter mH2A1 protein levels over the indicated time course. C, D CHX (40 μg/ml) chase experiments shortened the half-life of mH2A1 protein, and this degradation was inhibited by co-treatment with MG132, indicating proteasome-dependent turnover. E, F Increasing doses of transfected Flag‑UBE3A plasmid produced a dose‑dependent decrease in endogenous and exogenous Myc‑mH2A1 protein levels. G Gradual transfection of the catalytically inactive mutant UBE3A‑C833A did not significantly alter mH2A1 expression in pancreatic cancer cells, indicating that UBE3A catalytic activity is required for mH2A1 regulation. H, I UBE3A modulates the effect of CHX on mH2A1 protein half‑life. Following MG132 treatment, UBE3A no longer regulates mH2A1 protein levels, consistent with a proteasome‑dependent mechanism. J, K Under CHX treatment, UBE3A overexpression further shortened mH2A1 protein half‑life, whereas UBE3A knockdown delayed mH2A1 degradation. L, M Stable UBE3A‑overexpressing or UBE3A‑knockdown pancreatic cancer cells were transfected with HA‑Ub, endogenous mH2A1 was immunoprecipitated, and mH2A1 ubiquitination was assessed by Western blot. N UBE3A regulation of exogenous Myc‑mH2A1 ubiquitination was dose‑dependent and time‑dependent in in vivo ubiquitination assays. O Protein purification was assessed by Coomassie staining of GST and GST‑UBE3A. An in vitro reconstituted ubiquitination assay was then performed by incubating E1 (UBE1), E2 (UbCH7), HA‑Ub, and purified proteins (GST or GST‑UBE3A together with Myc‑mH2A1‑WT or Myc‑mH2A1‑K167A) for 30 min; Myc‑mH2A1 was subsequently immunoprecipitated with anti‑Myc affinity resin, and its ubiquitination was detected by Western blot. P pancreatic cancer cells were co‑transfected with combinations of UBE3A‑WT, UBE3A‑C833A, mH2A1‑WT, mH2A1‑K167A, and ubiquitin plasmids; Myc‑mH2A1 was immunoprecipitated to evaluate in vivo ubiquitination. Q, R The impact of UBE3A on mH2A1 ubiquitination was evaluated in pancreatic cancer cells transfected with ubiquitin mutants favoring distinct linkage types (K6, K11, K27, K29, K33, K48, K63). S, T Evaluate the effect of modulating UBE3A expression on the degradation rate of mH2A1 protein. All experiments were independently repeated three times; representative immunoblots are shown. Quantitative data are presented as mean ± SEM. Two‑group comparisons for panels (S, T) were analyzed by a two‑tailed unpaired Student’s t‑test (***P < 0.001; ****P < 0.0001).
The regulatory role of UBE3A in the ubiquitination of mH2A1 was further investigated. In UBE3A-knockdown PANC-1 cells, the level of ubiquitinated mH2A1 was significantly reduced, whereas in UBE3A-overexpressing MIA PaCa-2 cells, the level of ubiquitinated mH2A1 was markedly increased (Fig. 4L, M). Furthermore, the level of mH2A1 ubiquitination was positively correlated with both the expression level of UBE3A and the duration of its activity (Fig. 4N). To validate these findings further, we constructed the Myc-mH2A1 protein and its K167A mutant to evaluate ubiquitination in vitro. The results showed that GST-UBE3A effectively catalyzed the ubiquitination of Myc-mH2A1 but failed to ubiquitinate Myc-mH2A1 K167A (Fig. 4O). In vivo, the mH2A1 K167A mutant or the UBE3A C833A mutant did not induce mH2A1 ubiquitination (Fig. 4P). Additionally, in pancreatic cancer cells transfected with plasmids expressing different HA-Ub chains, we observed that UBE3A promoted the polyubiquitination of mH2A1, specifically on K48-linked chains (Fig. 4Q, R), which supports the notion that UBE3A mediates the proteasomal degradation of mH2A1.
A, B RNA sequencing (RNA‑seq) was performed on pancreatic cancer cells transfected with mH2A1 plasmids (Vector vs. mH2A1; four biological replicates, n = 4). Total RNA had an RIN value of ≥ 8.0 and concentrations sufficient for library preparation. Libraries were sequenced on an Illumina platform using paired‑end 150 bp reads, yielding approximately 4–6 × 10^7 reads per sample with a mean GC content of ~46%. Sequencing quality was high (> 98% of bases ≥ Q30), and the overall mapping rate to the GRCh38 reference genome exceeded 98%. Heatmap (A) shows the global expression patterns of differentially expressed genes upon mH2A1 overexpression; volcano plot (B) displays significance versus fold change (filter criteria: |Log2 FC | ≥ 1.5, P < 0.05). C, D ChIP‑seq was performed in pancreatic cancer cells to identify mH2A1 binding sites (one pair, n = 1). This study was conducted on the MGISEQ-T7 sequencing platform to perform ChIP-seq sequencing. A MGISEQ-T7 paired-end (PE) library with an insert size of approximately 300 bp was constructed and sequenced. Raw sequencing data were subjected to quality control and analyses of the ChIP-seq data. The total number of raw reads exceeded 80 million, with Q20 and Q30 values of > 99% and ≥ 97%, respectively, and an average GC content of ~45%. The effective rate was 99.9%. The heatmap depicts the distribution of mH2A1 peaks surrounding transcription start sites (TSS). E Distribution of mH2A1 binding sites across genomic functional regions. Identified binding sites were annotated to promoters, 5′ UTRs, exons, introns, 3′ UTRs, and intergenic regions; the chart displays the relative abundance of each category as a percentage. F The intersection of RNA‑seq and ChIP‑seq results revealed potential mH2A1 interaction targets. G IGV genome browser visualization of 17 putative mH2A1 binding sites at the TERT locus. The sites indicated by the blue boxes correspond to individual peaks and are significantly enriched relative to the input control (P < 0.05). H, I Multiplex immunofluorescence staining of UBE3A, mH2A1, and TERT in pancreatic tumor and adjacent normal tissues. Fluorescence intensities for the yellow (UBE3A), green (mH2A1), and red (TERT) channels were quantified using Image J. Each group included three independent biological replicates (n = 3); for each replicate, three random fields were acquired, and the mean of the three fields was used for analysis. Statistical comparisons were performed using a paired two‑tailed Student’s t‑test. J–M In two pancreatic cancer cell lines stably overexpressing or knocking down UBE3A, qRT‑PCR and Western blot were used to assess expression changes of UBE3A, mH2A1, and TERT. All experiments were independently repeated thrice (biological replicates, n = 3). Western blotting band intensities were quantified by densitometry and normalized to α‑Tubulin. Statistical comparisons were performed using a paired two‑tailed Student’s t‑test. All quantitative data are presented as mean ± SEM. Significance is indicated as **P < 0.01, ***P < 0.001, ****P < 0.0001.
Rescue experiments demonstrated that mH2A1 overexpression significantly promoted the senescent phenotype of pancreatic cancer cells and abolished the anti-senescence effect of UBE3A (Supplementary Fig. S7A–I). Notably, across multiple assays—including SA-β-gal staining, CCK-8 proliferation assay, SAHF marker analysis, EdU incorporation, and flow cytometry—only UBE3A‑WT exerted a significant regulatory effect on the senescence phenotype, whereas UBE3A‑C833A and UBE3A‑ΔN showed no observable effects (Supplementary Figs. S8A–I and S9A–I). Furthermore, the senescence‑promoting effect of mH2A1‑WT was reversed by UBE3A‑WT but not by the catalytically inactive UBE3A‑C833A mutant (Supplementary Fig. S8A–I and S9A–I). Moreover, mice bearing UBE3A-WT tumors exhibited significantly increased subcutaneous tumor volume and weight (Figure S10A–C) and pronounced hepatic metastasis (Figure S10D–G). In contrast, mice bearing tumors overexpressing mH2A1 showed markedly reduced subcutaneous tumor volume and weight (Figure S10A–C), and concurrent overexpression of UBE3A in mH2A1-overexpressing tumors reversed these effects. Notably, the catalytically inactive UBE3A-C833A mutant neither promoted tumor growth nor metastasis. These results indicate that UBE3A-mediated ubiquitination of mH2A1 is required for UBE3A’s tumor-promoting activity.
Identification of Downstream Targets of mH2A1
To further research the molecular mechanisms by which mH2A1 regulates cellular senescence, transcriptomic sequencing was performed on four pairs of mH2A1-overexpressing cell lines and control pancreatic cancer cell lines. The results revealed 844 upregulated and 1640 downregulated genes following mH2A1 overexpression (p < 0.05, |log₂FC | > 1.5) (Fig. 5A, B). Moreover, we conducted ChIP-seq experiments using a ChIP-grade antibody against mH2A1, and the ChIP-seq signal profile identified 3,789 genes that are potentially involved in DNA-protein interactions with mH2A1. The IP sample reads were predominantly distributed on either side of each annotated gene’s transcriptional start site (TSS) (Fig. 5C, D). mH2A1 binding sites are broadly distributed across genomic functional elements, including exons, introns, intergenic regions, TSS, and regions proximal to transcription termination sites(Fig. 5E). After cross‑referencing the 1640 genes identified by RNA‑seq as negatively correlated with mH2A1 expression, the 3789 genes from ChIP‑seq that potentially interact with mH2A1, and a set of 173 genes involved in telomerase regulation, TERT may represent a tentative downstream candidate target of mH2A1 (Fig. 5F). Visualization of the ChIP-seq results using the IGV genome browser revealed 17 binding sites for mH2A1 within the TERT genomic region (Fig. 5G). Additionally, mIHC revealed that UBE3A and TERT were upregulated, whereas mH2A1 was downregulated in pancreatic cancer tissues compared to adjacent normal tissues (Fig. 5H, I). In pancreatic cancer cells, upregulation of UBE3A was associated with decreased mH2A1 expression and increased TERT expression. Conversely, UBE3A knockdown elevated mH2A1 levels and reduced TERT expression(Fig. 5J–M).
The UBE3A/mH2A1/TERT Axis Mediates Pancreatic Cancer Cell Resistance to Senescence
The transcriptional repressor, mH2A1, was found to have 17 binding sites within the TERT genomic region, with one site located upstream of the TSS and the remaining sites located downstream (Fig. 6A). Further, to investigate the specific binding interactions between mH2A1 and TERT, we designed primers targeting these binding site sequences and performed ChIP‒qPCR. The results revealed significant enrichment of mH2A1 at all 17 sites compared to the IgG control group, but only site 10 exhibited enrichment above the input background level (Fig. 6B; Supplementary Fig. S11A). These findings suggest that site 10 is the primary binding site for mH2A1 and TERT. Interestingly, site 10 was not located within the classical promoter region of TERT ( + 99 to −2000 bp). Studies have reported that mH2A1 modulates osteoclastogenesis in breast cancer by recruiting EZH2 to enhancer regions, thereby increasing local levels of H3K27me3 histone modifications and suppressing LOX gene expression [21]. We hypothesized that mH2A1 suppresses TERT transcription by modulating its enhancer activity. To test this hypothesis, we examined the enrichment of enhancer markers H3K27ac, H3K4me1, and H3K4me2 at site 10 in pancreatic cancer cells. All three markers were significantly enriched at site 10, suggesting they were near the TERT enhancer (Fig. 6C–E; Supplementary Fig. S11B–D). Furthermore, overexpression of mH2A1 significantly increased the enrichment of mH2A1, H3K27me3, and EZH2 at site 10, whereas mH2A1 knockdown had the opposite effect (Fig. 6F–H; Supplementary Fig. S11E–G). We further isolated Flag-H2A and Flag-mH2A1 nucleosomes from the pancreatic cancer cells. Compared to Flag-H2A nucleosomes, Flag-mH2A1 nucleosomes showed significant enrichment of EZH2 and H3K27me3 (Supplementary Fig. S12A). Co-IP and GST pull-down assays confirmed that mH2A1 directly interacted with EZH2 in vivo and in vitro (Supplementary Fig. S12C–E). The overexpression of EZH2 increased H3K27me3 levels and decreased TERT expression, whereas EZH2 knockdown reduced H3K27me3 levels and increased TERT expression (Supplementary Fig. S12B). These findings suggest that mH2A1 recruits EZH2 to mediate H3K27me3 modification at site 10, suppressing TERT enhancer activity and transcription. To validate this mechanism, we constructed luciferase reporter vectors containing site 10 using pGL4.23 along with the corresponding control vectors (Fig. 6I). Dual-luciferase reporter assays revealed that mH2A1 overexpression significantly increased the fluorescence intensity of the reporter vector containing site 10, but not of the pGL4.23 or pGL4.10 control vectors (Fig. 6J; Supplementary Fig. S11H). Consistent with this, mH2A1 overexpression reduced TERT expression in pancreatic cancer cells (Fig. 6K–M), whereas mH2A1 knockdown increased TERT expression (Supplementary Fig. S11I, J). Rescue experiments demonstrated that UBE3A overexpression increased TERT expression, which was reversed by mH2A1 overexpression (Supplementary Fig. S11K-L).
A A schematic showing the 17 mH2A1 binding sites across the TERT locus. B Specific primers were designed for the 17 binding sites above. ChIP-qPCR was performed to assess mH2A1 enrichment at each site (biological replicates n = 3; qPCR with technical triplicates; enrichment was calculated relative to input). C–E Enrichment of enhancer marks H3K27ac (C) H3K4me1 (D) and H3K4me2 (E) at binding site 10 was assessed by ChIP‑qPCR (biological replicates n = 3; qPCR with technical triplicates; enrichment relative to input). F–H mH2A1 upregulation significantly increased the enrichment of mH2A1 (F) H3K27me3 (G) and EZH2 (H) at binding site 10 as measured by ChIP‑qPCR (biological replicates n = 3; qPCR with technical triplicates; enrichment relative to input). I A schematic of the luciferase reporter constructs used (pGL4.10, pGL4.23, and pGL4.23+Site10). J pGL4.10, pGL4.23, and pGL4.23+Site10 luciferase constructs were co‑transfected with mH2A1 into PANC‑1 cells, and relative luciferase activity was measured by dual‑luciferase assay (Firefly luciferase normalized to Renilla). Each independent biological replicate included technical triplicates; all experiments were performed three times, with biological replicates (n = 3). K–M Effects of mH2A1 on TERT expression were evaluated by qRT‑PCR and Western blot (qRT‑PCR data were normalized to α‑Tubulin and calculated by the 2^-ΔΔCt method; Western blot band intensities were quantified by densitometry and normalized to α‑Tubulin; all experiments were repeated three independent times, biological replicates n = 3). N–P PANC‑1 cell lines expressing Vector, UBE3A‑WT, UBE3A‑C833A, UBE3A‑ΔN, and mH2A1‑WT were generated, and ChIP‑PCR was used to assess enrichment of mH2A1, EZH2, and H3K27me3 at TERT‑Site10 in each group (biological replicates n = 3; qPCR with technical triplicates; enrichment relative to input). Q The pGL4.23+Site10 reporter was co‑transfected with Vector, UBE3A‑WT, UBE3A‑C833A, UBE3A‑ΔN, mH2A1‑WT into PANC‑1 cells, and relative luciferase activity (Firefly/Renilla) was measured by dual‑luciferase assay. Each independent biological replicate included technical triplicates; biological replicates n = 3. R–T qRT-PCR and Western blot analyses of TERT expression were performed across the different treatment groups (qRT-PCR normalized to α-Tubulin; Western blot band densitometry normalized to α-Tubulin). All experiments were repeated three times, with biological replicates (n = 3). All quantitative data are presented as mean ± SEM. Statistical comparisons were performed using a paired two‑tailed Student’s t‑test. Significance is indicated as *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
ChIP‑PCR, dual‑luciferase reporter assays, qPCR, and immunoblotting demonstrated that UBE3A‑WT markedly reduced the enrichment of mH2A1, EZH2, and H3K27me3 at TERT‑Site10 (Fig. 6N–Q, Supplementary Fig. S11M–P) and transcriptionally upregulated TERT (Fig. 6R–T, Supplementary Fig. S11Q, R); by contrast, the catalytic mutant UBE3A‑C833A and the N‑terminal deletion mutant UBE3A‑ΔN lacked these effects. Further analyses indicated that mH2A1 promotes the enrichment of EZH2 and H3K27me3 at TERT‑Site10, and this promotion is abrogated by UBE3A‑WT but not by UBE3A‑C833A (Fig. 6N–Q, Supplementary Fig. S11M–P). TERT knockdown induced more pronounced senescent features in pancreatic cancer cells and reversed the anti-senescence effects of UBE3A (Supplementary Fig. S13A–I). These findings elucidate the molecular mechanism by which UBE3A regulates senescence in pancreatic cancer cells, highlighting the UBE3A/mH2A1/TERT axis as a critical pathway.
Targeting UBE3A combined with senolytic agents further suppresses pancreatic cancer growth
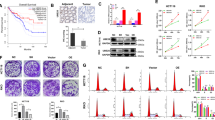
Excessive telomere elongation often leads to replicative senescence and the upregulation of antiapoptotic members of the BCL-2 family, rendering cells resistant to apoptotic signals. ABT-263 is a senolytic agent that targets the anti-apoptotic BCL-2 family and effectively eliminates senescent cells [34]. Therefore, we hypothesized that inhibiting UBE3A in combination with ABT-263 would result in a more potent anticancer effect. Consistent with this, our experiments showed that the anti-apoptotic proteins Bcl-2, Bcl-W, and Bcl-xL were significantly upregulated in senescent pancreatic cancer cells, and their expression increased further following UBE3A knockdown (Fig. 7A). We identified an optimal concentration of ABT‑263 (2.5 μM; Fig. 7B), consistent with previous results [34]. ABT‑263 alone does not induce death in non‑senescent PANC‑1 cells at this concentration. Annexin V/PI flow cytometry analysis revealed that neither UBE3A knockdown nor ABT-263 treatment significantly affected the apoptosis of pancreatic cancer cells. However, combining UBE3A knockdown with ABT-263 treatment markedly increased the percentage of apoptotic pancreatic cancer cells (Fig. 7C-D). Consistently, SA-β-gal staining demonstrated that UBE3A knockdown combined with ABT-263 significantly reduced the number of senescent pancreatic cancer cells (Fig. 7E, F).
A Western blot analysis of the impacts of UBE3A overexpression or knockdown on the expression of anti‑apoptotic BCL family proteins (including BCL‑2, BCL‑W, and BCL‑XL) in pancreatic cancer cells. Western blots were performed on independently prepared samples (biological replicates n = 3); band intensities were quantified by densitometry and normalized to α‑Tubulin. B Cell viability of PANC‑1 cells following treatment with varying concentrations of ABT‑263 was assessed by the trypan blue exclusion assay. Adherent cells were detached with trypsin; the resulting cell suspension was stained with trypan blue and counted using a hemocytometer to determine the percentage of viable cells. Experiments were performed with three independent biological replicates (n = 3). Data are presented as mean ± standard deviation. C, D In the senescent pancreatic cancer cell model, apoptosis was evaluated by flow cytometry using Annexin V‑FITC/PI following sh‑UBE3A and/or ABT‑263 treatment. Flow cytometry data were acquired from samples of three independent experiments, recording at least 10,000 events per sample. E, F Senescent cell positivity rates in PANC‑1 cells after sh‑UBE3A and/or ABT‑263 treatment were assessed by SA‑β‑gal staining. For each biological replicate, three random fields were imaged (20× objective), and the mean of the three fields was used as the value for that replicate. All cell‑based experiments were independently repeated three times (n = 3). G Schematic of the subcutaneous tumor model and experimental grouping: following establishment of PANC‑1 xenografts, animals were randomized into four groups: sh‑Ctrl, sh‑UBE3A, sh‑UBE3A + TA‑65, and sh‑UBE3A + ABT‑263. One week after tumor implantation, the sh‑UBE3A + TA‑65 group received TA‑65 by intraperitoneal injection at 25 mg/kg every 3 days for 4 weeks; the sh‑UBE3A + ABT‑263 group received ABT‑263 by intraperitoneal injection at 50 mg/kg every 3 days for 4 weeks. Calipers were used to measure tumor volumes during the treatment period, and tumors were subsequently excised and weighed at the endpoint. Each group comprised 5 mice (n = 5; one tumor per mouse). H–J Representative photographs of subcutaneous tumors from each group are shown, with comparative and statistical analyses of tumor volumes and weights (n = 5 per group). K–M At the study endpoint, animals were euthanized and tumors harvested for IHC to detect the proliferation markers PCNA and Ki67. IHC quantification was performed on 5 animals per group, with three random fields counted per animal. Data are presented as mean ± SEM. Statistical comparisons were made using two‑tailed Student’s t tests; significance is indicated as ***P < 0.001, ****P < 0.0001.
We further established a subcutaneous tumor xenograft model to validate the synergistic antitumor activity of UBE3A knockout combined with ABT-263 (Fig. 7G). In a subcutaneous tumor model, UBE3A knockdown slowed tumor growth; however, this inhibitory effect was reversed upon treatment with the telomerase activator TA-65 (Fig. 8H–J). Interestingly, in mice injected subcutaneously with UBE3A-knockdown PANC-1 cells, combined treatment with ABT-263 resulted in an even more significant suppression of tumor growth (Fig. 7H–J). Immunohistochemical analysis of tumor tissues revealed that UBE3A knockdown reduced the expression of proliferation markers Ki67 and PCNA. Furthermore, the knockout of UBE3A, combined with ABT-263, substantially reduces the expression levels of Ki-67 and PCNA. (Fig. 7K–M). These findings suggest that targeting UBE3A in combination with ABT-263 treatment suppresses pancreatic tumor growth more effectively.
This figure was created using the Figdraw 2.0 online platform (ID: WYWWR3bc23).
Discussion
Pancreatic cancer is known for its highly invasive and metastatic nature, and despite surgical intervention, patient outcomes remain poor. Targeted therapy is the cornerstone of cancer treatment; however, effective therapeutic options for pancreatic cancer are still limited. Inducing tumor cell senescence has emerged as a robust anticancer strategy because it inhibits tumor growth and enhances sensitivity to combination therapies. As critical components of the UPS, E3 ubiquitin ligases play essential regulatory roles in tumor cell senescence by modulating protein homeostasis. An in-depth exploration of these mechanisms may provide a crucial foundation for developing novel targeted therapeutic strategies.
To identify targets in regulating pancreatic cancer cell senescence, we culled publicly available databases and integrated clinical data from patients with pancreatic cancer. We analyzed the expression profile of UBE3A in pancreatic cancer cells. The results revealed that UBE3A is upregulated in pancreatic cancer, and its elevated expression is closely correlated with tumor size, pathological stage, and poor patient outcomes. The anti-senescence effects of the E3 ubiquitin ligase UBE3A have not been fully validated in tumor cells, despite the pro-tumorigenic roles of UBE3A being documented in various cancers. For example, UBE3A mediates the ubiquitination and degradation of p53, thereby promoting malignant progression in hepatocellular carcinoma [35]. In lung cancer [36], UBE3A cooperates with MCM6 to regulate tumor cell proliferation and migration. Additionally, UBE3A induces the ubiquitination and degradation of PBRM1, thereby modulating the sensitivity of renal cancer cells to CDK4/6 inhibitors [37]. In this study, we employed complementary in vitro and in vivo approaches to demonstrate that UBE3A suppresses senescence in pancreatic cancer cells, thereby promoting tumor growth and metastasis. This pro-tumorigenic effect depends on UBE3A’s catalytic cysteine residue, Cys833. Additionally, inhibiting UBE3A while administering the senolytic agent ABT-263 more significantly impeded the growth of subcutaneous tumors. This finding aligns with previous reports, suggesting that inducing pancreatic cancer cell senescence with senescence-inducing agents (e.g., etoposide, radiation, and gemcitabine) more effectively suppresses tumor growth when used in combination with ABT-263 [34, 38]. These insights provide critical evidence for the further exploration of the mechanistic role of UBE3A in pancreatic cancer.
The histone H2A variant, mH2A1, characterized by its unique macrodomain, forms specific complexes within chromatin and regulates various biological processes, including cellular senescence, energy metabolism, and stem cell properties [20]. Multiple studies have demonstrated the critical role of the UPS in regulating the stability of the mH2A1 protein. For example, following DNA damage, RNA168 catalyzes the ubiquitination of mH2A1, thereby facilitating the accumulation of downstream repair factors in damaged chromatin [39]. Additionally, the degradation and ubiquitination of mH2A1 in the nucleus can be induced by TRIM59, thereby enhancing STAT3 signaling and promoting tumorigenesis [40]. Moreover, the BRCA1/BARD1 complex regulates mH2A1 ubiquitination to encourage senescence in fibroblasts [41]. Ub-MS and IP-MS analyses initially established the direct ubiquitination of mH2A1 by UBE3A. Further ubiquitination assays demonstrated that UBE3A interacts with mH2A1 through its N-terminal domain and promotes polyubiquitination at the K167 residue of mH2A1 via K48-linked chains, thereby targeting it for degradation through the UPS. This process regulates the senescence of pancreatic cancer cells. The mH2A1-K167A mutation markedly reduces mH2A1 ubiquitination, thereby restoring protein stability and further supporting lysine 167 as a functional UBE3A modification site. Given prior reports that TRIM59 can also recognize and modify this residue [40], we hypothesize that distinct E3 ubiquitin ligases may exert competitive or cooperative regulation at the same lysine, resulting in divergent biological outcomes. For instance, Rad18 mediates monoubiquitination of PCNA at K164, whereas HLTF and SHPRH promote K63-linked polyubiquitination at the same site [42]; likewise, multiple E3s (e.g., MDM2 [43], Pirh2 [44], COP1 [45], ARF-BP1 [46]) catalyze K48-linked chains on the C-terminal lysine cluster of p53 to drive proteasomal degradation. To clarify the relationships among these enzymes and their distinct functional consequences, future studies should compare the chain-type specificities and spatiotemporal expression patterns of UBE3A and TRIM59, and validate their interplay using co-expression and competitive interaction assays.
Although several studies have explored the relationship between mH2A1 and cellular senescence, investigations into its precise mechanisms remain relatively limited. As a histone variant, mH2A1 can replace conventional H2A histones, altering chromatin structure. These structural changes often result in increased compactness in specific genomic regions, thereby suppressing the transcriptional activity of associated genes [47]. To investigate the molecular mechanisms underlying mH2A1-mediated senescence in pancreatic cancer cells, we integrated RNA-seq and ChIP-seq analyses, identifying TERT as the downstream effector of mH2A1. TERT, the catalytic subunit of Telomerase, plays a critical role in maintaining telomere length and cellular division. Telomeres progressively shorten during continuous cell division, eventually leading to replicative senescence. However, TERT is abnormally upregulated in many cancer cells, enabling telomere maintenance and allowing cancer cells to evade standard senescence signals, thereby facilitating unlimited proliferation [30]. Our findings revealed that TERT is regulated by mH2A1 and contains 17 potential regions within its DNA sequence with which it can interact with mH2A1. Further analysis identified binding site 10 ( + 20345 to +20556) as the primary site for the mH2A1-TERT interaction. Although this region is distal to the promoter, it is highly enriched with H3K27ac, H3K4me1, and H3K4me2, suggesting it functions as an enhancer region for TERT. Previous studies [21] indicated that nucleosomes formed by mH2A1 can recruit EZH2, which mediates H3K27me3 within enhancer regions, thereby suppressing gene transcription. Similarly, our study demonstrated that mH2A1 and its associated nucleosomes show increased enrichment of EZH2 and H3K27me3 compared to H2A nucleosomes. Elevated levels of EZH2 increase intracellular H3K27me3 levels, ultimately suppressing TERT expression.
Our results further indicate that UBE3A (C833) plays a crucial role in regulating TERT transcription. The UBE3A-C833A mutant appears to lose the ability to ubiquitinate mH2A1, resulting in increased mH2A1 expression and consequent enrichment of a complex comprising mH2A1, EZH2, and H3K27me3 at the TERT-Site10 locus, thereby repressing TERT expression at the transcriptional level. Notably, in the context of human papillomavirus (HPV) infection, UBE3A has also been reported as an essential regulator of TERT transcription: HPV16 E6 associates with UBE3A to form a functional E3 ubiquitin ligase complex that, on one hand, mediates polyubiquitination and proteasomal degradation of the transcriptional repressor NFX1-91, relieving repression of the TERT promoter; on the other hand, cooperates with c-Myc and the NFX1-123 isoform to activate TERT transcription and increase telomerase activity [48, 49]. In addition [50, 51], E6 can, through UBE3A-dependent ubiquitination activity, promote increased acetylation of histone H3 at the TERT promoter, further relieving transcriptional repression. These observations demonstrate that UBE3A exerts multi-layered regulation of TERT across different cellular contexts and molecular mechanisms.
Despite our findings highlighting the key role of UBE3A in cell senescence and tumor progression, and elucidating the underlying mechanism, several limitations remain. First, the precise molecular mechanisms by which UBE3A cooperates with ABT-263 to suppress tumor growth are not yet fully understood; further work is needed to dissect their roles within senescence-associated signaling and apoptotic pathways. Second, the potential cooperative or competitive modification at mH2A1-K167 by UBE3A and TRIM59, as well as the downstream molecular basis for gene expression regulation, senescence, and pro-tumorigenic effects, have not been elucidated; additional experiments are required to delineate their relationship and mechanisms. Third, although the study focuses on pancreatic cancer, it remains unknown whether UBE3A influences pancreatic cancer malignancy through other pathways in the context of HPV, necessitating investigations in HPV-related models. These limitations point to directions for future research.
In summary, our study highlights the crucial role of UBE3A in the senescence of pancreatic cancer cells. We elucidated the mechanism by which UBE3A ubiquitinates mH2A1 and further revealed that the UBE3A/mH2A1 axis promotes TERT expression by modulating EZH2-driven H3K27me3, conferring anti-senescence properties, and thereby promoting malignant progression in pancreatic cancer. Moreover, concomitant suppression of UBE3A expression and administration of the senolytic agent ABT-263 is a viable therapeutic strategy for further inhibiting tumor growth. These findings provide a novel perspective for developing targeted treatment strategies for pancreatic cancer.
Data availability
The original datasets for this study are available upon request from the corresponding author, and all data analyzed in this study are included in the article and its supplementary files.
Code availability
The source codes for this study are available from the corresponding author upon reasonable request.
References
Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA Cancer J Clin. 2024;74:12–49.
Thau H, Gerjol BP, Hahn K, von Gudenberg RW, Knoedler L, Stallcup K, et al. Senescence as a molecular target in skin aging and disease. Ageing Res Rev. 2025;105:102686.
McHugh D, Durán IA-O, Gil JA-O. Senescence as a therapeutic target in cancer and age-related diseases. Nat Rev Drug Discov. 2025;24:57–71.
Yang K, Li X, Xie KA-O. Senescence program and its reprogramming in pancreatic premalignancy. Cell Death Dis. 2023;14:528.
Wang BA-O, Han J, Elisseeff JH, Demaria MA-O. The senescence-associated secretory phenotype and its physiological and pathological implications. Nat Rev Mol Cell Biol. 2024;25:958–78.
Bousset LA-O, Gil JA-O. Targeting senescence as an anticancer therapy. Mol Oncol. 2022;16:3855–80.
Czajkowski K, Herbet M, Murias M, Piątkowska-Chmiel I. Senolytics: charting a new course or enhancing existing anti-tumor therapies?. Cell Oncol (Dordr). 2025;48:351–71.
Rysanek D, Vasicova P, Kolla JN, Sedlak D, Andera L, Bartek J, et al. Synergism of BCL-2 family inhibitors facilitates selective elimination of senescent cells. Aging (Albany NY). 2022;14:6381–414.
Morimoto RI. Cell-Nonautonomous Regulation of Proteostasis in Aging and Disease. Cold Spring Harb Perspect Biol. 2020;12:a034074.
Rai MA-O, Curley MA-O, Coleman Z, Demontis FA-O. Contribution of proteases to the hallmarks of aging and to age-related neurodegeneration. Aging Cell. 2022;21:e13603.
Spano DA-O, Catara G. Targeting the Ubiquitin-Proteasome System and Recent Advances in Cancer Therapy. Cells. 2023;13:29.
Lopez SJ, Segal DJ, LaSalle JM. UBE3A: An E3 Ubiquitin Ligase With Genome-Wide Impact in Neurodevelopmental Disease. Front Mol Neurosci. 2018;11:476.
LaSalle JM, Reiter LT, Chamberlain SJ. Epigenetic regulation of UBE3A and roles in human neurodevelopmental disorders. Epigenomics. 2015;7:1213–28.
Owais A, Mishra RK, Kiyokawa HA-O. The HECT E3 Ligase E6AP/UBE3A as a Therapeutic Target in Cancer and Neurological Disorders. Cancers (Basel). 2020;12:2108.
Williams K, Irwin DA, Jones DG, Murphy KM. Dramatic Loss of Ube3A Expression during Aging of the Mammalian Cortex. Front Aging Neurosci. 2010;2:18.
Jeon HY, Jeong YW, Kim YW, Jeong YI, Hossein SM, Yang H, et al. Senescence is accelerated through donor cell specificity in cloned pigs. Int J Mol Med. 2012;30:383–91.
Zheng Z, Zhang B, Yu H, Li S, Song N, Jin X, et al. UBE3A activates the NOTCH pathway and promotes esophageal cancer progression by degradation of ZNF185. Int J Biol Sci. 2021;17:3024–35.
Zhang N, Shen J, Gou L, Cao M, Ding W, Luo P, et al. UBE3A deletion enhances the efficiency of immunotherapy in non-small-cell lung cancer. Bioengineered. 2022;13:11577–92.
Song L, Liu S, Zeng S, Zhang L, Li X. miR-375 Modulates Radiosensitivity of HR-HPV-Positive Cervical Cancer Cells by Targeting UBE3A through the p53 Pathway. Med Sci Monit. 2015;21:2210–7.
Lo Re O, Vinciguerra M. Histone MacroH2A1: A Chromatin Point of Intersection between Fasting, Senescence and Cellular Regeneration. Genes (Basel). 2017;8:367.
Kim J, Shin Y, Lee S, Kim M, Punj V, Lu JF, et al. Regulation of Breast Cancer-Induced Osteoclastogenesis by MacroH2A1.2 Involving EZH2-Mediated H3K27me3. Cell Rep. 2018;24:224–37.
Doyen CM, An W, Angelov D, Bondarenko V, Mietton F, Studitsky VM, et al. Mechanism of polymerase II transcription repression by the histone variant macroH2A. Mol Cell Biol. 2006;26:1156–64.
Chen H, Ruiz PD, McKimpson WM, Novikov L, Kitsis RN, Gamble MJ. MacroH2A1 and ATM Play Opposing Roles in Paracrine Senescence and the Senescence-Associated Secretory Phenotype. Mol Cell. 2015;59:719–31.
Giallongo S, Di Rosa M, Caltabiano R, Longhitano L, Reibaldi M, Distefano A, et al. Loss of macroH2A1 decreases mitochondrial metabolism and reduces the aggressiveness of uveal melanoma cells. Aging (Albany NY). 2020;12:9745–60.
Lo Re O, Mazza T, Giallongo S, Sanna P, Rappa F, Vinh Luong T, et al. Loss of histone macroH2A1 in hepatocellular carcinoma cells promotes paracrine-mediated chemoresistance and CD4(+)CD25(+)FoxP3(+) regulatory T cells activation. Theranostics. 2020;10:910–24.
Park SJ, Shim JW, Park HS, Eum DY, Park MT, Mi Yi J, et al. MacroH2A1 downregulation enhances the stem-like properties of bladder cancer cells by transactivation of Lin28B. Oncogene. 2016;35:1292–301.
Xu D, Li CF, Zhang X, Gong Z, Chan CH, Lee SW, et al. Skp2-macroH2A1-CDK8 axis orchestrates G2/M transition and tumorigenesis. Nat Commun. 2015;6:6641.
Assalve G, Lunetti P, Rocca MS, Cosci I, Di Nisio A, Ferlin A, et al. Exploring the Link Between Telomeres and Mitochondria: Mechanisms and Implications in Different Cell Types. Int J Mol Sci. 2025;26:993.
Lanna A. Unexpected links between cancer and telomere state. Semin Cancer Biol. 2025;110:46–55.
Liu M, Zhang Y, Jian Y, Gu L, Zhang D, Zhou H, et al. The regulations of telomerase reverse transcriptase (TERT) in cancer. Cell Death Dis. 2024;15:90.
Shou S, Li Y, Chen J, Zhang X, Zhang C, Jiang X, et al. Understanding, diagnosing, and treating pancreatic cancer from the perspective of telomeres and telomerase. Cancer Gene Ther. 2024;31:1292–305.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res. 2013;41:D991–995.
Tomczak K, Czerwińska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (Pozn). 2015;19:A68–77.
Jaber S, Warnier M, Leers C, Vernier M, Goehrig D, Médard JJ, et al. Targeting chemoresistant senescent pancreatic cancer cells improves conventional treatment efficacy. Mol Biomed. 2023;4:4.
Zhang X, Bian S, Ni Y, Zhou L, Yang C, Zhang C, et al. Minichromosome maintenance protein family member 6 mediates hepatocellular carcinoma progression by recruiting UBE3A to induce P53 ubiquitination. Int J Biol Macromol. 2023;248:125854.
Luo Y, Yang Y, Yang C, Li C, Hu R, Geng W, et al. UBE3A and MCM6 synergistically regulate the proliferation and migration of lung adenocarcinoma cells. FEBS Open Bio. 2023;13:1756–71.
Liu W, Zhang B, Zhang D, Guo F, Ye K, Zhu L, et al. The RBPJ/DAPK3/UBE3A signaling axis induces PBRM1 degradation to modulate the sensitivity of renal cell carcinoma to CDK4/6 inhibitors. Cell Death Dis. 2022;13:295.
Revskij D, Woitas A, Kolle B, Umstatter C, Zechner D, Khan FM, et al. Effects of triggers of senescence and senolysis in murine pancreatic cancer cells. Hepatobiliary Pancreat Dis Int. 2024;23:628–37.
Kelliher JL, West KL, Gong Q, Leung JWC. Histone H2A variants alpha1-extension helix directs RNF168-mediated ubiquitination. Nat Commun. 2020;11:2462.
Sang Y, Li Y, Zhang Y, Alvarez AA, Yu B, Zhang W, et al. CDK5-dependent phosphorylation and nuclear translocation of TRIM59 promotes macroH2A1 ubiquitination and tumorigenicity. Nat Commun. 2019;10:4013.
Kim BJ, Chan DW, Jung SY, Chen Y, Qin J, Wang Y. The Histone Variant MacroH2A1 Is a BRCA1 Ubiquitin Ligase Substrate. Cell Rep. 2017;19:1758–66.
Hoege C, Pfander B, Moldovan GL, Pyrowolakis G, Jentsch S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature. 2002;419:135–41.
Huang Y, Che X, Wang PW, Qu X. p53/MDM2 signaling pathway in aging, senescence and tumorigenesis. Semin Cancer Biol. 2024;101:44–57.
Leng RP, Lin Y, Ma W, Wu H, Lemmers B, Chung S, et al. Pirh2, a p53-induced ubiquitin-protein ligase, promotes p53 degradation. Cell. 2003;112:779–91.
Su Y, Luo Y, Zhang P, Lin H, Pu W, Zhang H, et al. Glucose-induced CRL4(COP1)-p53 axis amplifies glycometabolism to drive tumorigenesis. Mol Cell. 2023;83:2316–2331.e2317.
Zhang X, Berger FG, Yang J, Lu X. USP4 inhibits p53 through deubiquitinating and stabilizing ARF-BP1. Embo j. 2011;30:2177–89.
Guberovic I, Farkas M, Corujo D, Buschbeck M. Evolution, structure and function of divergent macroH2A1 splice isoforms. Semin Cell Dev Biol. 2023;135:43–49.
Liu X, Yuan H, Fu B, Disbrow GL, Apolinario T, Tomaic V, et al. The E6AP ubiquitin ligase is required for transactivation of the hTERT promoter by the human papillomavirus E6 oncoprotein. J Biol Chem. 2005;280:10807–16.
Gewin L, Myers H, Kiyono T, Galloway DA. Identification of a novel telomerase repressor that interacts with the human papillomavirus type-16 E6/E6-AP complex. Genes Dev. 2004;18:2269–82.
James MA, Lee JH, Klingelhutz AJ. HPV16-E6 associated hTERT promoter acetylation is E6AP dependent, increased in later passage cells and enhanced by loss of p300. Int J Cancer. 2006;119:1878–85.
Sekaric P, Cherry JJ, Androphy EJ. Binding of human papillomavirus type 16 E6 to E6AP is not required for activation of hTERT. J Virol. 2008;82:71–76.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 82460507, 82560464, 82560475), Guizhou Provincial Department of Science and Technology Key Project (ZK[2024]066), Guizhou Provincial Health Commission Project (gzwkj2024-373), Guizhou Provincial Scientific Foundation for Basic Research (Qiankehe Basic-ZK[2024]GNRH230), and Guizhou Medical University Affiliated Hospital 2024 NSFC Cultivation Program Regional Funding Project (gyfynsfc[2024]-54).
Author information
Authors and Affiliations
Contributions
LR and YP conceived the project. LR, RL, and XF designed the experiments, analyzed the data, and drafted the manuscript. SC, PL, and SL performed the in vitro cell experiments and in vivo animal studies. XS and CZ assisted with data collation and analysis. XS, XW, and YP provided critical revisions to the manuscript. All authors have approved the final version of the manuscript and are accountable for their contributions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Human Ethics Committee of the Affiliated Hospital of Guizhou Medical University (2019 Ethical Review No. 073) and the Animal Ethics Committee of Guizhou Medical University (No.19000672). All participants were informed of the study’s purpose and provided written informed consent. All methods were performed in accordance with the relevant guidelines and regulations.
Consent for publication
All co-authors have consented to the version of the manuscript for publication.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by: Professor Boris Zhivotovsky
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ren, L., Lu, R., Fei, X. et al. UBE3A-mediated mH2A1 Ubiquitination activates TERT transcription to promote senescence resistance in pancreatic cancer. Cell Death Dis 17, 274 (2026). https://doi.org/10.1038/s41419-026-08480-z
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41419-026-08480-z