Abstract
Several microRNAs play vital roles in placental development. miR-155 has been implicated in placental development and can directly interact with a variety of targets, including angiotensin type II receptor 1 (AT1R) (Agtr1) mRNA. The AT1R is pro-proliferative and promotes early placental development. We therefore tested the hypothesis that miR-155 downregulates Agtr1 mRNA expression and impairs placental development. Placentae and fetuses from wild-type C57Bl/6 mice (miR-155+/+, control) and C57Bl/6 mice with a null mutation in miR-155 (miR155-/-) were mated with males of the same genotype and analyzed on gestational day 18.5, when placental morphology and miR-155 and AGTR1 expression were assessed. Additionally, HTR8/SVneo cells were cultured with a miR-155 mimic to determine the effects on trophoblast proliferation, migration and invasion. miR-155-/- dams produced significantly heavier pups with unchanged placental weights and fetal-to-placental weight ratios. Placentae from miR-155-/- dams had significantly larger labyrinth zones and labyrinth-to-placental area ratios than controls, with altered stereological parameters. Placental Agtr1 mRNA and AGTR1 protein levels were significantly increased in miR-155-/- dams. Finally, in vitro treatment in human HTR-8/SVneo cells with the miR-155 mimic increased miR-155 expression, decreased AGTR1 mRNA levels and decreased the rates of trophoblast cell proliferation, migration and invasion. Thus, miR-155 is demonstrated to attenuate placental development in mice. We propose that this is at least partly due to its effects on the AT1R.

Similar content being viewed by others
Introduction
Placentation is a finely tuned process, relying on complex integrated genetic control. Post-transcriptional regulation by microRNAs (miRNAs) plays a role in this process [1]. miRNAs are short, non-coding RNA fragments that largely act by repressing mRNA translation. One miRNA that has been proposed to regulate placental development is miR-155 [2,3,4,5,6,7]. miR-155 is increased in placentae from pregnancies complicated by preeclampsia [8,9,10] and in vitro has been shown to inhibit trophoblast proliferation, migration, and invasion [6, 8]. Furthermore, pregnant mice with placental-specific miR-155 overexpression had significantly reduced fetal and placental weights, with decreased trophoblast invasion and insufficient remodeling of uterine spiral arteries [10]. Together, these studies provide strong evidence that miR-155 is a key regulator of placental development.
miRNAs can target multiple mRNA sequences, and miR-155 has been demonstrated to target several factors that contribute to placental development, including Cysteine-rich angiogenic inducer 61 (CYR61) [11], cyclin D1 (CCND1) [12], Forkhead box O-3 (FOXO3) [8], and Mothers against decapentaplegic homolog 2 (SMAD2) [6]. Outside of pregnancy, miR-155 has been shown to reduce the expression of the angiotensin II type 1 receptor gene (AGTR1) [13, 14], a known driver of placental development, and polymorphisms in the 3’-UTR of the AGTR1 mRNA are known to inhibit miR-155 binding [15]. Importantly, when human umbilical vein endothelial cells (HUVECs) isolated from healthy and preeclamptic pregnant women were transfected with a mature miR-155 plasmid, luciferase assays confirmed the direct effect of miR-155 on AGTR1 as a target gene [16]. However, their interactions in the placenta and in regulating placental development are yet to be investigated.
The AT1R is the receptor for angiotensin (Ang) II and, therefore, a key component of the reninangiotensin system (RAS). Although the circulating RAS is well known for its roles in cardiovascular control and salt and water homeostasis, tissue RASs also exist, including a placental RAS that plays an important role in placental development [17]. In tissues, Ang II acts via the AT1R to stimulate proliferation, migration [18], and angiogenesis [18, 19]. In humans, the expression of a number of the genes and proteins of the placental RAS, including the AT1R, are highest in early gestation [17], supporting its role in early placental development. The importance of adequate AT1R expression in placental development was established by Walther et al., where a null mutation in the gene encoding AT1R was associated with much smaller and more poorly vascularised placentae, which adversely impacted fetal development [20]. Therefore, the potential effect of miR-155 on AGTR1 mRNA expression and how this affects placentation requires investigation.
In this study, we utilized mice with a null mutation in miR-155 to examine the role of miR-155 in placental development by studying placental morphology and stereology and fetal development. The effect of genetic ablation of miR-155 on placental AT1R expression was also investigated as a key pathway in placental development. Given the importance of the placental renin-angiotensin system in controlling trophoblast proliferation [21, 22], we subsequently investigated the direct effects of miR-155 on trophoblast proliferation, migration and invasion, in addition to AT1R expression using HTR-8/SVneo cells.
Results
Placental and fetal weights from miR-155 +/+ (control) and miR-155 -/- mice
On pc day 18.5, pups from miR-155-/- dams had significantly greater fetal weights than pups from control dams (p = 0.003; Fig. 1A), but placental weights (Fig. 1B) and fetal to placental weight ratios (Fig. 1C) were not significantly different.
A Fetal and (B) placental weights, and C fetal to placental weight ratios in miR-155-/- and miR-155+/+ (control) groups. Data are represented as mean ± SEM. nmiR-155(+/+) = 18 litters with 4–12 fetuses/placentae each to a total of 96. nmiR-155(-/-) = 18 litters with 1–5 fetuses/placentae each to a total of 86. *indicates a significant difference from controls (p < 0.05) as analyzed using a linear mixed model with random intercept accounting for maternal familial correlation.
Numbers of embryos, viable embryos and resorptions were also assessed, however no significant differences were determined (Supplementary Fig. 2).
Placental morphology in control and miR-155 -/- mice
Placentae from miR-155-/- mice had a significantly larger labyrinth zone (p = 0.02; Fig. 2A) and labyrinth zone to placental area ratios (p = 0.04; Fig. 2B) compared with placentae from control mice. However, the area of the junctional zone (Fig. 2C), junctional zone to placental area ratio (Fig. 2D) and labyrinth zone to junctional zone ratio (Fig. 2E) were not different between control and miR-155-/- mice. This is shown in representative images of placentae from control (Fig. 2F) and miR-155-/- (Fig. 2G) mice.
A Labyrinth zone area, B labyrinth zone to placental area ratio, C junctional zone area, D junctional zone to placental area ratio and E labyrinth zone to junctional zone area in placentae miR-155-/- and control mice. F Labyrinth zone (LZ) compared with junctional zone (JZ) from the placenta of a control dam and G from the placenta of a miR-155-/- dam. H Magnified (40X objective) images depicting trophoblast cell morphology and spiral artery remodeling in the placenta of a control dam and I from the placenta of a miR-155-/- dam. Data are represented as mean ± SEM. nmiR-155(+/+) = 18 litters with 4–12 placentae each to a total of 96 (all placentae). nmiR-155(-/-) = 18 litters with 1–5 placentae each to a total of 86. * indicates a statistically significant difference from controls (p < 0.05).
Stereological analysis of placental tissue
Placental morphology of miR-155-/- and controls were evaluated using Merz grid stereological analysis (Table 1 and Supplementary Fig. 3). miR-155-/- placentae displayed a significant increase in trophoblast volume density (p < 0.0001) and trophoblast weight (p < 0.0001) when compared with the control placentae. Trophoblast surface density (p = 0.0005) and total surface area of trophoblasts (p < 0.0001) were also significantly increased in miR-155-/- placentae compared to controls. Interestingly, fetal capillary surface density was decreased in miR-155-/- placentae (p = 0.0001) compared to controls, as was mean barrier thickness, a pseudo-measure of syncytiotrophoblast barrier thickness (p = 0.005).
Effect of miR-155 deficiency on uteroplacental vascular function
Given the alterations to placenta stereology, we next sought to determine the impact of miR-155 deficiency on uterine and umbilical artery function in vivo. To achieve this, we undertook ultrasound biomicroscopy on pregnant miR-155-/- and control mice on day 17.5 postcoitum. Analysis of uterine artery function showed that there were no differences in the resistance index or the pulsatility index between control and miR-155-/- mice, both measures of uterine artery resistance to blood flow (Supplementary Fig. 4). Similarly, miR-155 deficiency did not impact the umbilical artery resistance index and pulsatility index, however, there was a reduction in end diastolic velocity (p = 0.036) were observed in the umbilical arteries of miR155-/- mice compared to those of control dams (Supplementary Fig. 5). Umbilical artery function was not correlated with fetal weight or fetal sex (data not shown).
miR-155 and AGTR1 mRNA, and AT1R protein expression, in control and miR-155 -/- mice
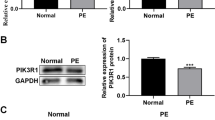
The expression of miR-155 was absent in placentae from miR-155-/- compared with control mice (p = 0.0003; Fig. 3A). miR-155 targets several different mRNA transcripts that encode for proteins that are associated with placental development, including Cyr61, Ccnd1, Foxo3, and Smad2. Therefore, qPCR was used to assess the relative abundance of these mRNAs in placentae from miR-155-/- compared with control mice, however no significant differences were observed between groups (data shown in Supplementary Fig. 6). Conversely, another known miR-155 target, AT1R, was altered in miR-155-/- mice placentae. Specifically, Agtr1 mRNA and AT1R protein levels were significantly increased in placentae from miR-155-/- compared with control mice (p = 0.01 and p = 0.004, respectively; Fig. 3B, C).
A Expression of miR-155, B Agtr1 mRNA and C AT1R protein levels in placentae from miR-155-/- and control mice. Data are represented as mean ± SEM. n = 4–9 placentae/dam for ten dams total. * indicates a statistically significant difference to controls (p < 0.05).
Representative full-length images of immunoblots are provided in Supplementary Fig. 7.
Effect of miR-155 on trophoblast proliferation in HTR-8/SVneo cells
In vitro experiments were employed to further investigate the effects of miR-155 on trophoblast cell function. HTR-8/SVneo cells treated with a miR-155 mimic had a significantly increased abundance of miR-155 (p = 0.003; Fig. 4A) and significantly decreased abundance of AGTR1 mRNA (p = 0.005; Fig. 4B) compared with the scrambled control. Treatment with 15, 30 and 60 pM of miR-155 mimic significantly decreased trophoblast proliferation compared with the control (p = 0.029, p = 0.005, p < 0.001, respectively; Fig. 4C). Proliferation was also significantly decreased at 15, 30 and 60 pM compared with 7.5 pM (p = 0.035, p = 0.047, p < 0.001, respectively). The cell index trajectory displays the gradient (slope) as an indication of cell activity.
A miR-155 and B AGTR1 mRNA expression in miR-155 mimic-treated and vehicle-treated HTR-8/SVneo cells. C The rate of proliferation, D migration and E invasion of HTR-8/SVneo cells transfected with different doses of a miR-155 mimic. The slopes of the lines indicate HTR8/SVneo cell proliferation, migration and invasion, respectively, over 72 h. Control (red), 7.5 pM mimic (blue), 15 pM mimic (green), 30 pM mimic (magenta), 60 pM mimic (cyan). Graphs D, E include an uncoated serum-free control (black); graph E includes an uncoated control (purple). Data are presented as mean ± SEM. n = 3 experiments, each in triplicate. * indicates a statistically significant difference to the vehicle control (p < 0.05). The same letter above the bars indicates that groups are not different from each other. A different letter above the bars indicates that groups are different (all p < 0.05).
As described in Supplementary Fig. 1, no significant change in proliferation was detected between the negative control (0 pM scrambled mimic + Lipofectamine) and all concentrations of the scrambled control, confirming that our results are miR-155 sequence-specific.
Effect of miR-155 on trophoblast migration in HTR-8/SVneo cells
Treatment with 15, 30 and 60 pM of miR-155 mimic significantly decreased trophoblast migration compared with the control (p = 0.001, p < 0.001, p < 0.001 respectively; Fig. 4D). Migration was also significantly decreased at 15, 30, and 60 pM compared with 7.5 pM (p = 0.013, p = 0.002, p < 0.001, respectively). Migration was significantly decreased at 60 pM compared with 15 and 30 pM (all p < 0.001). Migration was significantly decreased in SFC compared with all treatment conditions, i.e., 0, 7.5, 15, 30, and 60 pM mimic (all p < 0.001).
Effect of miR-155 on trophoblast invasion in HTR-8/SVneo cells
Treatment with 15, 30, and 60 pM of miR-155 mimic significantly decreased trophoblast invasion compared with the control (p = 0.048, p = 0.001, p < 0.001 respectively; Fig. 4E). Invasion was also significantly decreased at 30 and 60 pM compared with 7.5 pM (p = 0.011, p < 0.001, respectively). Invasion was significantly decreased at 30 pM compared with 15 pM (p = 0.038) and at 60 pM compared with 15 and 30 pM (all p < 0.001). Invasion was significantly decreased in USFC compared with 0, 7.5, 15, and 30 pM mimic and USFC (all p < 0.001) and was significantly increased in UC compared with all conditions, i.e., 0, 7.5, 15, 30, and 60 pM mimic (all p < 0.001).
Discussion
We have shown that miR-155 modulates placental cell function and development and that maternal and placental null mutation in miR-155 disrupts placental morphology by altering labyrinth and junctional zone development. We propose that this occurs, at least in part, due to the inhibitory action of miR-155 on Agtr1 mRNA expression in turn causing reduced AGTR1 synthesis.
The impact of the miR-155 null mutation was readily observed when placental morphology was analyzed. In miR-155-/- placentae, there was a significant increase in the labyrinth zone area, as well as the labyrinth zone to total placenta area ratio. The labyrinth zone is closest to the fetus and is considered to be the functional zone of the placenta where nutrient, gas and waste exchange occurs [23]. This increase in labyrinth zone size suggests that there was increased trophoblast differentiation, resulting in increased substrate transfer capacity of the placenta and, in turn, an increased ability to accelerate growth and sustain a larger fetus [24]. Placentae from miR-155-/- mice had higher trophoblast component weight in conjunction with increased trophoblast surface area when assessed by stereology, although placental weight did not significantly change between miR-155-/- and control mice. Compared to placentae from control mice, placentae from miR-155-/- mice also had higher trophoblast volume density and higher trophoblast surface density, indicating increased numbers of trophoblasts and that a greater proportion of the placenta area is covered by trophoblasts, respectively. Placentae from miR-155-/- mice also had increased mean barrier thickness (an indication of syncytiotrophoblast barrier thickness) compared with controls. These results are in accordance with a previous study by Wang et al. [10], who showed that miR-155 overexpressing placentae exhibited a smaller labyrinth layer, and a disorganized arrangement of trophoblast cells [10]. Wang et al. also demonstrated however that the miR-155 overexpressing placentae displayed decreased placental vascular networks. This contrasts with the current study, where placentae from miR-155-/- showed decreased fetal capillary surface density. Thus, further investigation into the role of miR-155 in placental vascularization is warranted.
Indeed, while miR-155 deficiency did not affect uterine artery function, it did have some impact on umbilical artery function. Umbilical artery Doppler velocimetry assesses resistance to fetal blood flow within the placenta. Various conditions can compromise the placental vasculature, leading to decreased, absent, or reversed end-diastolic velocity in the umbilical artery [54]. We observed that maternal miR-155 deficiency was associated with decreased umbilical end-diastolic velocity compared to controls, however, since this change was not accompanied by fetal growth restriction or alterations in the resistance or pulsatility index, the clinical significance of this finding may be limited [53, 54].
Together, these changes in placental structure are consistent with our observation of larger fetuses in miR-155-/- compared with controls, as well as literature indicating a positive correlation between placental volume and fetal growth [25]. Indeed, at the same gestational age, miR-155-/- mice did have larger fetuses than controls, without any concomitant alteration to other fetal parameters. Given that we established the inhibitory action of miR-155 on AT1R expression, these findings correlate with the findings of Walther et al., where placental AT1R knockout resulted in smaller placentae and fetuses [20]. Our results are further corroborated by the findings of Wang et al., who observed that placenta specific miR-155 overexpression in pregnant dams produced smaller and significantly fewer pups [10].
While our data point to an association between miR-155 regulation of AT1R expression in influencing placental development, it is important to acknowledge that the effects of miR-155 deficiency on placental development may also be exacerbated by altered maternal immune parameters. We have previously shown profound changes to the maternal immune environment in pregnant miR-155 deficient female mice [26]. Maternal miR-155 was shown to be required for expansion of regulatory T cells to mediate robust pregnancy tolerance and support fetal development in mice [26]. Additionally, Treg cells contribute to the control of maternal vascular function during pregnancy [55, 56] further highlighting the potential link between miR-155 deficiency, immune cells and placental function. However, miR-155 driven immune alterations likely play a less essential role in the current study as pregnancies were generated by syngeneic matings, which are less dependent on maternal immune tolerance than the allogeneic mating model used in our previous work [26]. This may explain why in the current study, miR-155 deficiency leads to an increase in fetal weight, despite the fact that compromised Treg cell number and function in pregnancy is associated with pregnancy complications such as fetal growth restriction and preeclampsia [57, 58, 59].
Adequate trophoblast proliferation is essential for placental growth. We showed that treatment with a miR-155 mimic decreased HTR-8/SVneo cell proliferation in a dose-dependent manner, consistent with previous studies reporting that miR-155 decreases cell proliferation [5, 27]. Recent studies have confirmed that transfection of HTR-8/SVneo cells with a miR-155 mimic also results in reduced trophoblast migration and invasion [6]; likewise, inhibition of miR-155 expression in HTR-8/SVneo and JEG3 cell lines resulted in increased proliferation, migration and invasion [28]. This is coherent with our findings that miR-155 deficient placentae are more efficient through larger labyrinth zone area and increased trophoblast component weight and volume density, although we did not detect a significant difference in placental weight between the two groups. Furthermore, we demonstrated that miR-155 mimic treatment significantly reduced the migratory and invasive capacity of HTR-8/SVneo cells in a dose-dependent manner. As these processes are critical for extravillous trophoblast differentiation and successful uterine implantation [29], our findings suggest that miR-155 may impair placental development by disrupting these key pathways.
The actions of miR-155 are complex, as it is known to regulate many targets in addition to the gene encoding AT1R. Within the placenta, studies have associated miR-155 expression with downregulation of cysteine-rich protein 61 (CYR61), which is an important angiogenic regulating factor during pregnancy [11]. Importantly, CYR61 is essential for placental vascular integrity [30] and induces expression of vascular endothelial growth factor (VEGF) [31] and is shown to be decreased in preeclampsia [32]. Indeed, in a rat model of preeclampsia, miR-155 was increased, and VEGF decreased compared with controls [5]. miR-155 has also been implicated in the suppression of the PTEN 30-untranslated region, leading to reduced AP-1/NF-kB pathway activity [7]. miR-155 downregulates cyclin D1 [12], a critical regulator of cell cycle progression and transcriptional co-regulator, along with Smad2 [6], a signal transducer that interacts with the TGF-β receptors, and FOXO3 [8], which is suspected to function as a trigger for cellular apoptosis. Despite these reported interactions between miR-155 and targets that are likely involved in placental development and function, we found no evidence that the mRNA expression of these targets was altered in the placentae of miR-155-/- mice (Supplementary Fig. 6). We acknowledge that miR-155 may regulate not only these targets, but also other targets, post-transcriptionally, and that the fetal and placental phenotypes observed are likely the result of combined effects on multiple pathways. However, the substantial upregulation of AT1R expression in miR-155-/- placentae supports our hypothesis that miR-155 is at least partly acting through the AT1R to induce its effects.
Notably, miR-155 directly interacts with the 3’-UTR of AGTR1 mRNA [33], and the interaction between miR-155 and AGTR1 has been established in patient-derived HUVECs from preeclamptic and healthy pregnancies [16]. Furthermore, the silent polymorphism +1166A/C in the 3’-UTR of AGTR1 has been associated with hypertension and cardiovascular complications, with expression of the mutant allele resulting in decreased miR-155 expression, likely due to difficulty binding to the mRNA. AT1R protein expression was significantly increased and positively correlated with systolic and diastolic blood pressure in pregnancy [34]. The current study provides robust evidence that miR-155 negatively regulates placental development, and proposes a novel mechanism involving its interaction with AT1R. Specifically, we observed ~2-fold increased Agtr1 mRNA expression and almost 3-fold increased AT1R protein in miR-155-/- placentae, in the absence of changes at the mRNA level for other targets or miR-155, indicating an association between miR-155 and AT1R expression as well as placental growth. Further evidence for the miR-155/AT1R axis exerting control over placental cell growth was seen using treatment with a miR-155 mimic, which reduced AGTR1 mRNA expression by approximately half, also had a dose-dependent negative effect on HTR-8/SVneo cell proliferation.
It is worthy to note that in rodents AT1R exists as two isoforms, Agtr1a and Agtr1b, which share a high degree of sequence similarity. Although Agtr1a is the predominant isoform expressed in the placenta [35], the primers used in our qPCR assay do not distinguish between the two isoforms. Therefore, the expression data presented here likely reflect combined levels of Agtr1a and Agtr1b. Future studies could employ isoform-specific approaches to further delineate the contribution of each receptor subtype to placental development. Furthermore, luciferase reporter assays or mutation of the miR-155 seed sequence binding site within Agtr1a and Agtr1b mRNA isoforms would provide definitive mechanistic confirmation, instead of the associations reported here which indicate that miR-155 may directly and/or indirectly regulate AGTR1 expression and contribute to placental development and pathology. However, these experiments are beyond the scope of the current study.
This study confirms the importance of adequate Ang II/AT1R signaling in placentation and reinforces the findings of Delforce et al., showing that specific inhibition of the AT1R by losartan inhibits expression of the angiogenic/proliferative trophoblast phenotype [36]. This phenotype is highly correlated with the expression of pro-angiogenic factors and cell viability [36]. The binding of Ang II to the AT1R has also been implicated in cytotrophoblast secretion of many hormones essential for healthy pregnancy, including placental lactogen, human chorionic gonadotropin and pregnancy-specific glycoprotein [37], as well as estradiol [38]. As mentioned previously, AT1R knockout in the placenta severely impedes placental growth and vasculogenesis [20]. Even intrauterine growth restriction is associated with reduced placental AGTR1 mRNA and AT1R protein [39], which is consistent with our results as miR-155-/- placentae expressed significantly higher levels of AT1R mRNA and protein, and produced larger fetuses.
Clinically, our findings have potential translational relevance, as miR-155 dysregulation has been associated with several human pregnancy complications. Significantly lower levels of miR-155 are recorded in first-trimester peripheral blood from women who later experienced recurrent pregnancy loss, suggesting that early miR-155 dysregulation may compromise pregnancy maintenance [40]. In contrast, elevated placental miR-155 expression has been observed in patients with preeclampsia, where it was shown to directly repress AGTR1 expression [10]. Other studies have similarly reported upregulation of miR-155 in preeclamptic placentae ([4, 28, 41,42,43]), and miR-155 has also been implicated in the pathogenesis of intrauterine growth restriction through its impact on trophoblast proliferation and invasion [6] and as a circulating biomarker of small for gestational age pregnancies [44]. These findings support a model in which precise regulation of miR-155 is essential for normal placental development and function, and implicate miR-155 as a potential biomarker or therapeutic target in pregnancy disorders. Indeed, Wang et al. [10] showed that antagomir-155 could eliminate the preeclampsia-like manifestations in pregnant mice in which miR-155 is overexpressed in the placenta [10]. Notably, this study was intended as mechanistic proof-of-concept, and while miR-155 may represent a useful biomarker, its direct therapeutic targeting would require caution due to the risks of disrupting renin–angiotensin signaling in pregnancy [45].
Indeed, the differential expression of miR-155 in maternal blood from women experiencing pregnancy complications raises the possibility that it may function as a cell-free or exosomal signal during pregnancy. Indeed, several studies have proposed miR-155 as a potential noninvasive biomarker for various pregnancy complications ([2, 4, 10]). While our study focused on placental expression, future investigations could explore whether miR-155 is released into the maternal circulation in response to placental stress or dysfunction, and whether it contributes to intercellular communication between the placenta and maternal tissues. This highlights the broader physiological relevance of miR-155 beyond its local effects within the placenta.
Overall, we have shown that culturing trophoblast cells with a miR-155 mimic decreased both AGTR1 mRNA and AT1R protein and impaired trophoblast proliferation, migration and invasion. This was supported by our finding that placentae from miR-155-/- dams had increased levels of Agtr1 mRNA and AT1R protein accompanied by increased fetal weights and an increase in trophoblasts and labyrinth zone area. The study therefore provides compelling evidence that miR-155 impacts placental morphology and functional capacity, resulting in altered fetal growth. We conclude that the effects of miR-155 on placental development and fetal growth are at least partly due to its regulation of AT1R expression.
Methods
Animal work
Bic/miR-155 (miR-155-/-) mice on a C57Bl/6 background [26] generated by Prof Klaus Rajewsky (CBR Institute for Biomedical Research, Harvard Medical School) [46] were purchased from The Jackson Laboratory (Bar Harbor, ME, USA, Stock No: 007745) and bred in-house at the University of Adelaide. C57Bl/6J (B6; miR-155+/+) wildtype control female mice were purchased from Animal Resource Centre, Perth. All mice were co-housed in specific pathogen-free conditions at the University of Adelaide Medical School Animal House on a 12-h light-dark cycle and were administered food and water ad libitum.
Experimental females were 8–12 weeks, and males were 10 weeks to 12 months in age.
For mating experiments, miR-155+/+ and miR-155-/- females were mated with the same genotype males, and 1–2 adult female mice were caged with one male for mating. Mice were checked for a vaginal plug daily (checked between 8 am and 10 am), and females on the day of plug detection were designated post coitum (pc) day 0.5. Mated females were housed in groups of 1–4 females per cage. No randomization or blinding was used.
The number of mice per experiment is listed in the corresponding Figure legend.
Ultrasound
On pc day 18.5, both uterine and umbilical artery function were assessed using an MX550D transducer probe on a Vevo 3100 ultrasound biomicroscope (VisualSonics®, ON, Canada), as described previously [55]. Briefly, mice were anaesthetised with isoflurane (5% induction, 1.5% maintenance, in medical air) and Doppler waveforms were acquired from uterine arteries near the lateral inferior margin of the utero-cervical junction and from the umbilical arteries of 3–4 fetuses per dam. During the ultrasound, the location of each fetus was marked on the skin of the abdomen of the anesthetised dam using a permanent marker. Immediately following the ultrasound, at postmortem, the maternal abdominal wall was carefully opened without disturbing the uterus, and corresponding marks were made on the uterine surface to identify the studied fetuses. These fetuses were dissected and weighed first to ensure accurate identification before processing the remaining fetuses. Each waveform was analyzed in triplicate, following previously established methods. Peak systolic velocity (PSV), end-diastolic velocity (EDV), time-averaged velocity (TAV) and heart rate averages were obtained from a minimum of three consecutive cardiac cycles. Resistance index (RI = (PSV − EDV)/PSV) and pulsatility index (PI = (PSV − EDV)/TAV) were calculated. Following Doppler imaging, dams were humanely killed by cervical dislocation. Each viable fetus was dissected from the amniotic sac and umbilical cord, and the fetuses and placentae were weighed. Placental tissues were either cryo-frozen or fixed in 10% formalin and washed in phosphate-buffered saline before embedding in paraffin.
RNA extraction and DNase treatment
Total RNA was extracted from crushed cryo-frozen placental tissue using TRIzol Reagent
(Thermo Fisher Scientific, Waltham, MA, USA), according to the manufacturer’s instructions.
Tissues (~0.15 g) were homogenized with 1.5 mL of TRIzol in a Precelleys24 homogenizer (5000 RPM, 2 × 30 s, then 1 × 20 s, Thermo Fisher Scientific). DNase I treatment (Qiagen GMBH, Hilden, Germany) was performed on all samples. The integrity of the total RNA and miRNAs were examined by gel electrophoresis and quantified using the Nanodrop 2000 (Thermo Fisher Scientific, data not shown [47]). Samples were used for further analysis if the 260:280 and 260:230 nm ratios were 1.8–2.1 and 2.0–2.2, respectively.
miRNA analysis
Expression of miR-155 was measured by qPCR [21]. Total RNA (5 ng) samples underwent reverse transcription to cDNA (TaqMan miRNA Reverse Transcription Kit and probes, Assay ID #002571 for human miR-155, #002571 for mouse miR-155, #001094 for RNU44 and # 002491 for miR-20a; Thermo Fisher Scientific) according to the manufacturer’s instructions. Samples then underwent quantitative PCR (qPCR) using TaqMan Universal PCR master mix, according to the manufacturer’s instructions (Thermo Fisher Scientific). Data was generated using a 7500 Real-Time PCR System (Thermo Fisher Scientific). The expression of miR-155 was determined by calculating 2−ΔΔCT using RNU44 (a highly conserved small nucleolar RNA in the growth arrest-specific 5 transcript) as the human reference gene and miR-20a as the mouse reference gene.
Quantitative reverse transcriptase polymerase chain reaction (qPCR)
All RNA samples underwent reverse transcription to generate cDNA according to the manufacturer’s instructions (Superscript III First-Strand Synthesis for RT, Thermo Fisher Scientific). Total RNA was spiked with a known amount of Alien RNA (Stratagene; 107 copies per µg of total RNA). qPCR was performed in an Applied Biosystems 7500 Real-Time PCR System using SYBR Green for detection. Each reaction mixture contained 5 μL of SYBR Green PCR master mix (Applied Biosystems), primers, 10 ng cDNA, and water to 10 μL. Messenger RNA abundance was calculated as described previously, using the 2–ΔΔCT method and expressed relative to Alien mRNA and a calibrator cDNA (prepared from pooled mouse kidney tissue) [17]. Primer sequences for Agtr1 (primers capture expression of both Agtr1a and Agtr1b isoforms) [48]; Foxo3 F: CTCTCAGGCTCCTCACTGTA, R: ATGAGTTCACTACGGATGAT; Ccnd1 F: GAACAGACAAGCACATTAATAGA, R: GCTTCAGTTCATGAGTCTTATTCC; Cyr61 F: CGAGTTACCAATGACAACCCAG, R: TGCAGCACCGGCCATCTA, Smad2 F: GAGTGTGGATTGTTACCTTTG, R: CTCAACTCTCTGGTAGTGGTA.
Protein extraction
Protein was isolated from tissues using a radioimmunoprecipitation assay buffer (RIPA, 50 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, 1% Triton X–100, 1% sodium dodecyl sulfate, SDS) supplemented with a Pierce Halt™ complete protease inhibitor cocktail tablet (Thermo Fisher Scientific), in a ratio of 250 μL buffer to 200 mg of tissue. Samples were then homogenized using the Precelleys24 homogenizer (5000 RPM, 2 × 30 s, then 1 × 20 s, Thermo Fisher Scientific, Waltham, MA, USA) every 10 min across a 30 min period and cooled on ice in between. Protein levels were measured using the Pierce BCA Protein assay kit (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Immunoblotting
Immunoblotting was performed as described previously [48, 49]. Briefly, samples were loaded into Bis-Tris methane 4–12% gels in duplicate before electrophoresis (Thermo Fisher Scientific). Proteins were then transferred to a polyvinylidene difluoride (PVDF) membrane (Thermo Fisher Scientific) using the wet sandwich method immersed in Tris Buffered Saline + Tween20 (TBST) transfer buffer. The PVDF membrane was then completely dried and re-activated before immunodetection. The membrane was rocked in a blocking solution (5% BSA, 5% skim milk in Tris buffered saline (TBS)) for 2 h on a rocker at 22 °C. The primary antibody solution was then added (1:1000 dilution, Abcam, Cambridge, United Kingdom, #ab18801), and samples were incubated at 4 °C overnight. The secondary anti-rabbit antibody solution was added (1:5000 dilution, Millipore, Burlington, MA, USA; #12-348) and incubated at 22 °C on a rocker for 1 h. Membranes were rinsed before signal detection using an Amersham ECL detection kit and Amersham Imager 600 (both GE Healthcare, Chicago, IL, USA). Membranes were then dried and stripped using 0.2 M NaOH and, using a rabbit polyclonal anti-β-actin antibody (1:5000 dilution, Abcam; ab8227), were probed to detect β-actin for normalization. The ratio of the protein of interest to β-actin was averaged for duplicate lanes, and differences between blots were corrected using an internal control (a pooled mouse kidney sample). ImageJ software was used for assessment.
Morphometric analysis
The mid-sagittal placental labyrinth and junctional zones were morphometrically analyzed, as previously described [50]. Briefly, tissue sections were stained with haemotoxylin and eosin and mounted before imaging. QuPath software [51] was used to measure the cross-sectional areas of the labyrinth, junctional zone and total placenta. From these measurements, ratios of labyrinth to junctional zone, labyrinth to total placenta and junctional zone to total placenta were calculated.
Placental stereological analysis
Immunofluorescent dual labeling was performed as previously described [50]. Briefly, placental tissues were incubated with both mouse anti-cytokeratin (1:50 dilution, Agilent Technologies, Glostrup, Denmark; #M7018) and rabbit anti-vimentin (1:250 dilution, Abcam; #ab92547) primary antibodies overnight at 4 °C in 1% BSA-PBS. Following PBS-T washes, sides were incubated with both goat anti-mouse IgG H&L Alexa Fluor® 594 (1:500 dilution, Abcam; #ab150116) and goat anti-rabbit IgG H&L Alexa Fluor® 488 (1:500 dilution, Abcam; #ab150077) secondary antibodies, for 1 h at RT in 1% BSA-PBS. DAPIProLong Gold Antifade solution (Thermo Fisher Scientific) was applied before imaging at 20× magnification using an Axio Imager M2 microscope (Carl Zeiss AG, Jena, Germany) and analyzed using Zen imaging (Carl Zeiss AG) and ImageJ software (National Institutes of Health).
Immunofluorescent dual labeling was performed following established and widely accepted methodologies for quantifying placental architecture and blood space composition, as described in multiple prior studies ([23, 24, 50, 52]). To determine the proportion of placental trophoblasts, fetal capillaries, and maternal blood space, placental tissue dual labeled for cytokeratin and vimentin (as described above) underwent stereological analysis. The proportion of each component was quantified using point counting on an isotropic L-36 Merz grid using ImageJ imaging software. To ensure objectivity, uniform random sampling was employed to collect eight fields of view at 20x magnification per placental tissue.
The volume densities (Vd) of placental trophoblasts, fetal capillaries, and maternal blood space, defined as the proportion of the reference volume (i.e., placental tissue) occupied by each component, were calculated using the following formula:
Where Pa is the total number of points that have fallen on the component of interest, and PT is the total number of points applied to the image [24, 52].
The component weights (CW), representing the total mass of each placental component, the following formula was used:
The surface density (Sv) of placental trophoblasts, fetal capillaries, and maternal blood space, quantifying the amount of surface area per unit of reference volume, were calculated using the following formula:
Where La is the number of intercepts of the applied lines with the components of interest, and
LT is the total length of the lines applied to the section [24, 52].
The total surface area (ST), which represents the full surface area for each component within the placental labyrinth zone, was determined by the equation:
Total Surface Area; ST = Sv × placental weight × labyrinth proportion
The mean barrier thickness (BT), reflecting the average thickness of the syncytiotrophoblasts layer between the maternal and fetal circulations, was calculated using the following equation:
Where Vd and Sv are the trophoblast volume density and trophoblast surface density, respectively [24, 52].
To assess reproducibility, observations were repeated on sections randomly throughout the analysis. The variation observed was less than 5%.
Cell culture
HTR-8/SVneo cells (an immortalized first-trimester trophoblast cell line provided by Professor Charles Graham, Queens University, Ontario, free of mycoplasma) were cultured at 37 °C with 5% CO2 in RPMI1640 (Danaher Corporation, Washington, D.C., USA) supplemented with 10% heat-inactivated FCS (ISAFC Biosciences) and 1% L-glutamine (Thermo Fisher Scientific). Three separate cultures of HTR-8/SVneo cells were made, and from each of these three cultures, three sets of 2 × 105 cells from passages 10–20 were plated (n = 9). After 48 h, 30pM of the miRVana miR-155 mimic (Thermo Fisher Scientific) was transfected into each well, using Lipofectamine (Thermo Fisher Scientific) as the transfection vector. To confirm a sequence-specific response, other wells were treated with Lipofectamine alone (negative control) or with the addition of a scrambled control (where nucleotides are scrambled in a random order to confirm sequence order is specific) at concentrations of 7.5, 15, 30 and 60 pM; data supplied in Supplementary Fig. 1. Cells and culture media were individually collected and snap frozen in liquid nitrogen, then stored at −80 °C before RNA analysis.
Proliferation analysis
Fifty μL of cell culture medium was added to each well of an xCELLigence E-plate 16 (ACEA Biosciences Inc., San Diego, CA) and allowed to equilibrate at room temperature for 30 min. A background reading in the xCELLigence Real-Time Cell Analysis Multi Plate (RCTA MP) system was then carried out. 1 × 104 HTR-8/SVneo cells from passages 10–20 (cultured as above) were plated in each well with an additional 100 μL of incubation medium and again allowed to equilibrate for 30 min at room temperature. 24 h after cell plating, miRNA-155 miRVana mimic, scrambled mimics or vehicle (Lipofectamine) were added to each well at 0, 7.5, 15, 30 or 60 pM. As the proliferation plates have gold microelectrodes on the bottom of each well, the proliferation of cells impedes electrical conductance. Cell index was generated as a measure of the electrical resistance as cells proliferated. The cell index was measured every 30 min for 48 h in the xCELLigence RCTA MP system. After 48 h of incubation with treatment, data were collected and analyzed. The rate of proliferation was determined by measuring the slope of the line (cell index) over time.
Migration analysis
For cell migration analysis, an xCELLigence CIM-Plate 16 (ACEA Biosciences Inc., San Diego, CA) was used containing two chambers. 160 μL of incubation medium, with 10% FBS, was added to each well of the lower chamber, excepting a serum-free control (SFC). 50 μL of serum-free media was added to each well of the upper chamber, and the CIM-plate was equilibrated for 1 h at 37 °C, before a background measurement was taken using the RTCA software (Agilent, Santa Clara, CA, USA). A total of 3 × 104 HTR-8/SVneo cells were plated in each well of the upper chamber before equilibrating for 30 min at room temperature. CIM plates were placed in the RTCA DP cradle and cell index readings were taken every 30 min for 72 h using the RTCA software (Agilent, Santa Clara, CA, USA). 24 h after cell plating, miRNA-155 miRVana mimic, scrambled mimics or vehicle (Lipofectamine) were added to each well at 0, 7.5, 15, 30 or 60 pM.
Invasion analysis
For cell invasion analysis, an xCELLigence CIM-Plate 16 (ACEA Biosciences Inc., San Diego, CA) was used containing two chambers. 160 μL of incubation medium, with 10% FBS, was added to each well of the lower chamber. 20 μL of Matrigel (1:20 ratio with serum-free incubation medium) pre-coated each well of the upper chamber prior to addition of 50 μL of serum-free incubation medium. Controls included 2 wells without Matrigel coating (“uncoated controls”; UC) and 2 wells without Matrigel coating or serum in the lower chamber (“uncoated serum-free control”; USFC). The CIM-plate was then equilibrated for 1 h at 37 °C, before a background measurement was taken using the RTCA software (Agilent, Santa Clara, CA, USA). A total of 3 × 104 HTR-8/SVneo cells were plated in each well of the upper chamber before equilibrating for 30 min at room temperature. CIM plates were placed in the RTCA DP cradle and cell index readings were taken every 30 min for 72 h using the RTCA software (Agilent, Santa Clara, CA, USA). 24 h after cell plating, miRNA-155 miRVana mimic, scrambled mimics or vehicle (Lipofectamine) were added to each well at 0, 7.5, 15, 30, or 60 pM.
Statistical analyses
Statistical analysis was undertaken using SPSS Statistics. Where multiple placentae or pups per mother were assessed, a linear mixed model with random intercept accounting for maternal familial correlation was used. Where a single placenta from each mother was assessed, a MannWhitney test was used. Stereology was analyzed by GraphPad Prism 8. Stereological analysis was statistically validated using Two-way ANOVA and Šídák’s multiple comparisons test, with variation between measurements assessed via Kruskal–Wallis: Dunn’s multiple comparisons test. All graphs were generated using GraphPad Prism 8. Differences between groups were considered significant for p < 0.05.
Ethics
All experimental protocols were approved by the University of Adelaide Animal Ethics Committee (approval number: M-2014-023, M-2016-009), using methods in accordance with the Australian code for the care and use of animals for scientific purposes.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files. Should any raw data files be needed in another format they are available from the corresponding author upon reasonable request.
References
Mouillet J, Ouyang Y, Coyne C, Sadovsky Y. MicroRNAs in placental health and disease. Am J Obstet Gynaecol. 2015;213:163–72.
Azizi F, Gargari S, Shahmirzadi S, Dodange F, Amiri V, Mirfakhraie R. et al. Evaluation of Placental mir-155-5p and Long Non-coding RNA sONE expression in patients with severe preeclampsia. Int. J Mol Cell Med. 2017;6:22
Faraoni I, Antonet F, Cardone J, Bonmassar E. miR-155 gene: a typical multifunctional microRNA. Biochim Biophys Acta. 2009;1792:497–505.
Gan L, Liu Z, Wei M, Chen Y, Yang X, Chen L, et al. MiR-210 and miR-155 as potential diagnostic markers for pre-eclampsia pregnancies. Medicine. 2017;96:7515.
Liu Q, Yang J. Expression and significance of miR155 and vascular endothelial growth factor in placenta of rats with preeclampsia. Int J Clin Exp Med. 2015;8:15731–7.
Wu JX, Shi M, Gong BM, Ji BW, Hu CC, Wang GC, et al. An miRNA-mRNA integrative analysis in human placentas and mice: role of the Smad2/miR-155-5p axis in the development of fetal growth restriction. Front Bioeng Biotechnol. 2023;11:1159805.
Xue P, Zheng M, Diao Z, Shen L, Liu M, Gong P, et al. miR-155* mediates suppressive effect of PTEN 30-untranslated region on AP-1/NF-kB pathway in HTR-8/SVneo cells. Placenta. 2013;34:650656.
Luo X, Pan C, Guo X, Gu C, Huang Y, Guo J, et al. Methylation mediated silencing of miR-155 suppresses the development of preeclampsia in vitro and in vivo by targeting FOXO3. Mediat Inflamm. 2022;2022:4250621.
Pineles B, Romero R, Montenegro D, Tarca A, Han Y, Kim Y, et al. Distinct subsets of microRNAs are expressed differentially in the human placentas of patients with preeclampsia. Am J Obstet Gynecol. 2007;196:261–6.
Wang Z, Liu D, Dai Y, Li R, Zheng Y, Zhao G, et al. Elevated placental microRNA-155 is a biomarker of a preeclamptic subtype. Hypertension. 2023;80:370–84.
Zhang Y, Diao Z, Su L. MicroRNA-155 contributes to preeclampsia by down-regulating CYR61. Am J Obstet Gynaecol. 2010;202:466–7.
Dai Y, Qiu Z, Diao Z, Shen L, Xue P, Sun H, et al. MicroRNA-155 inhibits proliferation and migration of human extravillous trophoblast derived HTR-8/SVneo cells via down-regulating cyclin D1. Placenta. 2012;33:824–9.
Martin MM, Lee EJ, Buckenberger JA, Schmittgen TD, Elton TS. MicroRNA-155 regulates human angiotensin II type 1 receptor expression in fibroblasts. J Biol Chem. 2006;281:1827718284.
Zhu N, Zhang D, Chen S, Liu X, Lin L, Huang X, et al. Endothelial enriched microRNAs regulate angiotensin II-induced endothelial inflammation and migration. Atherosclerosis. 2011;215:286–93.
Sethupathy P, Borel C, Gagnebin M, Grant GR, Deutsch S, Elton TS, et al. Human microRNA-155 on chromosome 21 differentially interacts with its polymorphic target in the AGTR1 3’ untranslated region: a mechanism for functional single-nucleotide polymorphisms related to phenotypes. Am J Hum Genet. 2007;81:405–13.
Cheng W, Liu T, Jiang F, Liu C, Zhao X, Gao Y, et al. microRNA-155 regulates angiotensin II type 1 receptor expression in umbilical vein endothelial cells from severely pre-eclamptic pregnant women. Int J Mol Med. 2011;27:393–9.
Pringle K, Tadros M, Callister R, Lumbers E. The expression and localization of the human placental prorenin/renin-angiotensin system throughout pregnancy: Roles in trophoblast invasion and angiogenesis?. Placenta. 2011;32:956–62.
Ishimatsua S, Itakurab A, Okadaa MKT, Iwaseb A, Kajiyamaa H, Inoa K, et al. Angiotensin II augmented migration and invasion of choriocarcinoma cells involves PI3K activation through the AT1 receptor. Placenta. 2006;27:587–91.
Herr D, Rodewald M, Fraser HM, Hack G, Konrad R, Kreienberg R, et al. Regulation of endothelial proliferation by the renin–angiotensin system in human umbilical vein endothelial cells. J Soc Reprod Fertiity. 2008;136:125–30.
Walther T, Jank A, Heringer-Walther S, Horn L, Stepan H. Angiotensin II type 1 receptor has impact on murine placentation. Placenta. 2008;29:905–9.
Arthurs A, Lumbers E, Pringle K. MicroRNA mimics that target the placental renin-angiotensin system inhibit trophoblast proliferation. Mol Hum Reprod. 2019;25:218–27.
Arthurs A, Lumbers ER, Delforce SJ, Mathe A, Morris BJ, Pringle KG. The role of oxygen in the regulation of miRNAs in control of the placental renin-angiotensin system. Mol Hum Reprod. 2019;25:206–17.
Coan P, Ferguson-Smith A, Burton G. Developmental dynamics of the definitive mouse placenta assessed by stereology. Biol Reprod. 2004;70:1806–13.
Roberts T, Sohlstrom A, Kind KL, Earl RA, Khong TY, Robinson JS, et al. Maternal food restriction reduces the exchange surface area and increases the barrier thickness of the placenta in the guinea-pig. Placenta. 2001;22:177–85.
Virupaxi R, Potturi B, Shirol V. Morphology of placenta and its relation with small for date babies in 950 Live births. Recent Res Sci Technol. 2011;3:123–26.
Schjenken JE, Moldenhauer LM, Zhang B, Care AS, Groome HM, Chan H-Y, et al. MicroRNA miR-155 is required for expansion of regulatory T cells to mediate robust pregnancy tolerance in mice. Mucosal Immunol. 2020;13:609–25.
Li X, Li C, Dong X, Gou W. MicroRNA-155 inhibits migration of trophoblast cells and contributes to the pathogenesis of severe preeclampsia by regulating endothelial nitric oxide synthase. Mol Med Rep. 2014;10:550–4.
Wu H-Y, Liu K, Zhang J-L. LINC00240/miR-155 axis regulates function of trophoblasts and M2 macrophage polarization via modulating oxidative stress-induced pyroptosis in preeclampsia. Mol Med. 2022;28:119.
Davies J, Pollheimer J, Yong H, Kokkinos M, Kalionis B, Knöfler M, et al. Epithelial– mesenchymal transition during extravillous trophoblast differentiation. Cell Adh Migr. 2016;10:310–21.
Mo F, Muntean AG, Chen CC, Stolz DB, Watkins SC, Lau LF. CYR61 (CCN1) is essential for placental development and vascular integrity. Mol Cell Biol. 2002;22:8709–20.
Zhou D, Herrick DJ, Rosenbloom J, Chaqour B. Cyr61 mediates the expression of VEGF, αvintegrin, and α-actin genes through cytoskeletally based mechanotransduction mechanisms in bladder smooth muscle cells. J Appl Physiol. 1985;98:2344–54.
Gellhaus A, Schmidt M, Dunk C, Lye SJ, Kimmig R, Winterhager E. Decreased expression of the angiogenic regulators CYR61 (CCN1) and NOV (CCN3) in human placenta is associated with preeclampsia. Mol Hum Reprod. 2006;12:389–99.
Zheng L, Xu CC, Chen WD, Shen WL, Ruan CC, Zhu LM, et al. MicroRNA-155 regulates angiotensin II type 1 receptor expression and phenotypic differentiation in vascular adventitial fibroblasts. Biochem Biophys Res Commun. 2010;400:483–8.
Ceolotto G, Papparella I, Bortoluzzi A, Strapazzon G, Ragazzo F, Bratti P, et al. Interplay between miR-155, AT1R A1166C polymorphism, and AT1R expression in young untreated hypertensives. Am J Hypertens. 2011;24:241–6.
Gao H, Yallampalli U, Yallampalli C. Maternal protein restriction reduces expression of angiotensin I-converting enzyme 2 in rat placental labyrinth zone in late pregnancy. Biol Reprod. 2012;86:31–38.
Delforce S, Lumbers ER, Morosin S, Wang Y, Pringle KG. The Angiotensin II type 1 receptor mediates the effects of low oxygen on early placental angiogenesis. Placenta. 2019;75:54–61.
Kalenga M, de Gasparo M, Thomas K, De Hertogh R. Angiotensin II induces human placental lactogen and pregnancy-specific beta 1-glycoprotein secretion via an angiotensin AT1 receptor. Eur J Pharm. 1994;268:231–6.
Kalenga M, De Gasparo M, Thomas K, De Hertogh R. Angiotensin-II stimulates estradiol secretion from human placental explants through AT1 receptor activation. J Clin Endocrinol Metab. 1995;80:1233–7.
Li X, Shams M, Zhu J, Khalig A, Wilkes M, Whittle M, et al. Cellular localization of AT1 receptor mRNA and protein in normal placenta and its reduced expression in intrauterine growth restriction. Angiotensin II stimulates the release of vasorelaxants. J Clin Investig. 1998;101:442–54.
Winger E, Reed JL, Ji XF. First trimester PBMC MicroRNA predicts adverse pregnancy outcome. Am J Reprod Immunol. 2014;72:515–26.
Mayor-Lynn K, Toloubeydokhti T, Cruz AC, Chegini N. Expression profile of microRNAs and mRNAs in human placentas from pregnancies complicated by preeclampsia and preterm labor. Reprod Sci. 2011;18:46–56.
Ntsethe A, Mackraj I. An Investigation of exosome concentration and exosomal microRNA (miR155 and miR-222) expression in pregnant women with gestational hypertension and preeclampsia. Int J Women's Health. 2022;14:1681–9.
Wang Z, Zheng M, Duan H, Wang Y, Dai Y, Tang H. et al. Increased circulating miR-155 identifies a subtype of preeclamptic patients. BMC Pregnancy Childbirth. 2025;25:319
Hromadnikova I, Kotlabova K, Krofta L. First-trimester screening for fetal growth restriction and small-for-gestational-age pregnancies without preeclampsia using cardiovascular disease-associated microRNA biomarkers. Biomedicines. 2022;10:718.
National Institute for Health and Care Excellence (NICE). Hypertension in pregnancy: diagnosis and management. NICE Guideline 133 [NG133]. London; 2019. https://www.nice.org.uk/guidance/ng133/resources/hypertension-in-pregnancy-diagnosis-and-management-pdf-66141717671365.
Thai T, Calado D, Casola S, Ansel K, Xiao C, Xue Y, et al. Regulation of the germinal center response by microRNA-155. Science. 2007;316:604–8.
Arthurs AL, Mccullough D, Williamson JM, Jankovic-Karasoulos T, Smith MD, Roberts CT. Factors influencing RNA yield from placenta tissue. Placenta. 2023;140:80–83.
Wang Y, Lumbers E, Arthurs A, Corbisier de Meaultsart C, Mathe A, Avery-Kiejda K, et al. Regulation of the human placental (pro)renin receptor-prorenin-angiotensin system by microRNAs. Mol Hum Reprod. 2018;24:453–64.
Arthurs AL, Smith MD, Hintural MD, Breen J, McCullough D, Thornton FI, Leemaqz SY. et al. Placental inflammasomemRNA levels differ by mode of delivery and fetal sex. Front Immunol. 2022;13:807750
Schofield L, Kahl R, Rodrigues S, Fisher J, Endacott S, Delforce S. et al. Placental deficiency of the (pro) renin receptor ((P) RR) reduces placental development and functional capacity. Front Cell Dev Biol. 2023;11:1212898
Bankhead P, Loughrey MB, Fernández JA, Dombrowski Y, McArt DG, Dunne PD, et al. QuPath: open source software for digital pathology image analysis. Sci Rep. 2017;7:1–7.
Weibel ER, Kistler GS, Scherle WF. Practical stereological methods for morphometric cytology. J cell Biol. 1966;30:23–38.
Kingdom J. et al. Pathology and clinical implications of abnormal umbilical artery Doppler waveforms. Ultrasound Obstet Gynecol. 1997;9:271–286.
Bligard K. H. et al. Clinical significance of umbilical artery intermittent vs persistent absent end-diastolic velocity in growth restricted fetuses. Am J Obstet Gynecol. 2022;227:519. e511–e519.
Hosking S. L. et al. Treg cells promote decidual vascular remodeling and modulate uterine NK cells in pregnant mice. JCI Insight. 2024;10:e169836.
Care A. S. et al. Reduction in regulatory T cells in early pregnancy causes uterine artery dysfunction in mice. Hypertension. 2018;72:177–187.
Saito S. et al. What is the role of regulatory T cells in the success of implantation and early pregnancy? J Assist Reprod Genet. 2007;24:379–386.
Steinborn A. et al. Pregnancy-associated diseases are characterized by the composition of the systemic regulatory T cell (Treg) pool with distinct subsets of Tregs. Clin Exp Immunol. 2012;167:84–98.
Prins J. R. et al. Preeclampsia is associated with lower percentages of regulatory T cells in maternal blood. Hypertensionin Pregnancy. 2009;28:300–311.
Funding
ALA is supported by a Channel 7 Children's Foundation grant.
Author information
Authors and Affiliations
Contributions
ALA performed experimental work and formal analysis of data, original draft preparation, and review and editing. ERL conceptualized this work, provided supervision, interpretation of data, review and editing of the manuscript. LS and CL performed experimental work and formal analysis of data, and reviewed and edited the manuscript. PYC performed experimental work, and reviewed and edited the manuscript. ASC and JES performed experimental work, formal analysis of data and reviewed and edited the manuscript. SAR reviewed and edited the manuscript. KGP conceptualized this work, provided supervision, funding acquisition, interpretation of data, review and editing of the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arthurs, A.L., Lumbers, E.R., Schofield, L. et al. miR-155 suppresses angiotensin II type 1 receptor synthesis during placental morphogenesis. Cell Death Discov. 12, 49 (2026). https://doi.org/10.1038/s41420-025-02892-0
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41420-025-02892-0