Abstract
RIPK3 is a key regulator of necroptosis, but the specific roles of its kinase-dependent and -independent functions in disease pathogenesis remain poorly understood. Here, we generated and characterized RIPK3 D143N kinase-dead knock-in mice, a novel kinase-inactive model that selectively disrupts RIPK3 kinase activity without inducing spontaneous apoptosis. Unlike previously reported kinase-inactive Ripk3D161N/D161N mice, which exhibit embryonic lethality by triggering apoptosis, Ripk3D143N/D143N mice are viable and fertile, demonstrating that RIPK3 kinase activity is dispensable for development. The RIPK3 D143N mutation effectively blocks necroptosis induced by multiple stimuli and fully rescues embryonic lethality of caspase-8-deficient mice. Notably, Ripk3D143N/D143N mice were significantly less protected from TNF-driven inflammatory disease than RIPK3-deficient mice, revealing a critical kinase-independent role for RIPK3. This scaffold function drives inflammation and tissue damage through JAK-STAT1 activation, as pharmacological inhibition of JAK1/2 effectively reduces disease pathogenesis. Thus, our findings establish Ripk3D143N/D143N mice as a valuable model for dissecting the kinase and scaffold functions of RIPK3, and highlights the therapeutic potential of targeting its scaffold function in inflammatory diseases.

Schematic diagram showing the kinase-dependent and kinase-independent function in cell death and inflammation. A novel kinase-inactive RIPK3 mouse model (Ripk3D143N/D143N) defines the kinase-dependent and kinase-independent role of RIPK3 in cell death and inflammation.
Similar content being viewed by others
Introduction
Apoptosis is a well-characterized form of programmed cell death mediated by caspase activation [1, 2]. The extrinsic apoptotic pathway is initiated by death receptor signaling, leading to caspase-8 activation, while the intrinsic pathway involves mitochondrial signaling and caspase-9 activation [3, 4]. Both pathways further activate the executioner caspase-3 and caspase-7 to mediate apoptotic cell death [5]. When caspase-8 activity is compromised, cells undergo necroptosis, which is a form of regulated necrosis tightly regulated by RIP kinases RIPK1 and RIPK3 [5,6,7,8,9,10,11].
Necroptosis plays critical roles in inflammatory diseases [12, 13], neurodegenerative diseases [14,15,16], and host defense against pathogens [17, 18]. In TNF-induced necroptosis, inhibition or genetic deletion of caspase-8 (Casp-8) or FADD impairs apoptotic signaling and induces RIPK1 phosphorylation [19]. Activated RIPK1 then recruits and activates RIPK3 through RIP homotypic interaction motif (RHIM) domain-mediated interaction, forming the necrosome complex [20,21,22]. In addition to TNF signaling, toll-like receptor3/4 (TLR3/4) [23, 24], Z-DNA-binding protein1 (ZBP1/DAI) [25, 26], and pathogen sensors [18, 21] can also activate RIPK3, which functions as a central regulator of necroptosis. Activated RIPK3 phosphorylates mixed lineage kinase domain-like pseudokinase (MLKL), triggering its oligomerization and plasma membrane translocation to mediate necrosis [27,28,29]. Necroptosis causes plasma membrane breakdown, leading to the release of intracellular contents and DAMPs that amplify inflammatory responses [30].
Although the kinase activity of RIPK3 is essential for necroptosis, evidence suggests a kinase-independent function of RIPK3 in regulating apoptosis. However, RIPK3 kinase inhibitors (e.g. GSK'872) directly induce apoptosis, and the D161N kinase-dead mutation causes embryonic lethality in mice [31,32,33,34]. This occurs because kinase site alterations promote conformational changes in RIPK3 that expose its RHIM domain, facilitating recruitment of the RIPK1-FADD-caspase-8 complex to activate apoptosis [35]. Notably, this RIPK3/RIPK1-dependent apoptosis pathway differs from classical TNF-induced RIPK1-mediated apoptosis and is resistant to RIPK1 kinase inhibition [36, 37]. Additionally, both RIPK3-dependent necroptosis (kinase-dependent) and RIPK3-mediated apoptosis (kinase-independent) contribute to the pathogenesis of influenza A virus (IAV) infection [38, 39], highlighting the intricate functions of RIPK3 in modulating cell death and disease via kinase-dependent and kinase-independent mechanisms.
While RIPK3-dependent necroptosis is a driver of inflammation, emerging evidence indicates that RIPK3 also promotes pro-inflammatory cytokine production through cell death-independent mechanisms [6]. Genetic ablation of RIPK3 markedly ameliorates inflammation and tissue damage in diverse inflammatory disorders, including TNF-induced systemic inflammatory response syndrome (SIRS) [13, 40], graft-versus-host disease (GVHD) [12, 41], amyotrophic lateral sclerosis(ALS) [42], sepsis [43], kidney and heart ischemia-reperfusion injury [44] and acute pancreatitis [20]. Both RIPK3-mediated necroptosis (kinase-dependent) and apoptosis (kinase-independent) contribute to the pathogenesis of IAV infection [38, 45,46,47], highlighting the intricate functions of RIPK3 in modulating cell death and disease via kinase-dependent and kinase-independent mechanisms. Although these findings demonstrate the critical roles of RIPK3 in pathogenesis, the pathogenic contributions of its kinase-dependent and kinase-independent activities remain poorly defined.
In this study, we establish a novel kinase-inactive RIPK3 mouse model (Ripk3D143N/D143N) to define the precise role of RIPK3 kinase activity in cell death and inflammation. The viable and fertile RIPK3 D143N mice completely rescue caspase-8-deficiency-induced embryonic lethality. Notably, using this model, we demonstrate critical contributions of RIPK3 scaffold functions in TNF-induced SIRS through promoting JAK-STAT1 signaling. Pharmacological JAK1/2 inhibition ameliorated hypothermia and pathological damage in cecum. Our study reveals the scaffold function of RIPK3 in driving inflammatory disorder and provides new insights for therapeutic strategies targeting RIPK3 scaffold beyond its kinase activity.
Results
The kinase-inactive Ripk3 D143N/D143N mice develop normally
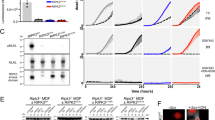
Current RIPK3 kinase-inactive models, such as the embryonic-lethal D161N mutation and the poorly expressed K51A variant [32, 37], have limitations, prompting the need for new models. Structural modeling of the murine RIPK3 kinase domain revealed close spatial proximity between D143 and D161 residues (Fig. 1A). To examine the impact of the kinase-dead D143N mutant on apoptosis, which has been reported not to spontaneously induce cell death in 3T3-SA cells [37]. Ripk3-/- mouse embryonic fibroblasts (MEFs) were transduced with wild-type (WT), D143N, or D161N RIPK3. Notably, only RIPK3 D161N-expressing MEFs exhibited significant cell death, which was completely abolished by caspase inhibition with the treatment of pan-caspase inhibitor z-VAD, while no cell death was induced in cells expressing RIPK3 D143N (Fig. 1B). Similar induction of apoptosis was observed in human HeLa and 293 T cells expressing the corresponding RIPK3 variants D160N, but not D142N (Fig. 1C; Fig. S1A-B). D160N RIPK3, but not WT or D142N variant, triggered the cleavage of caspase-3 and PARP, leading to apoptosis in HeLa cells (Fig. 1D). These results suggest that RIPK3 D143N does not induce spontaneous apoptosis. To investigate the physiological role of RIPK3 kinase inactivation, Ripk3D143N/D143N knock-in mice were generated (Fig. S2A). In contrast to the embryonic lethality of Ripk3D161N/D161N mice, Ripk3D143N/D143N mice were viable and fertile (Fig. 1E-F). Notably, the protein level of RIPK3, RIPK1 and MLKL in Ripk3D143N/D143N mice is consistent with littermate Ripk3+/+ mice (Fig. S2B). Histopathological examination of mouse tissues, including liver, spleen, lung, kidney, and small intestine, showed no abnormalities in Ripk3D143N/D143N mice (Fig. 1G).
A The predicted structure of mouse RIPK3 kinase domain using ChimeraX, and the inlay panel shows a magnified view of adenosine triphosphate pocket area. B Ripk3-/- MEFs were transfected with Vector, RIPK3 WT, RIPK3 D143N or RIPK3 D161N plasmid for 24 h, then cell viability and the protein level of RIPK3 were determined. C HeLa cells were transfected with Vector, RIPK3 WT, RIPK3 D142N or RIPK3 D160N plasmid for 24 h, then cell viability and the protein level of RIPK3 were determined. D Western blot analysis of RIPK3, full-caspase3, cl-caspase3, PARP and cl-PARP in HeLa cells transfected with Vector, RIPK3 WT, RIPK3 D142N or RIPK3 D160N plasmids for 24 h. E Representative image of 8-week-old Ripk3D143N/D143N mice with littermate control Ripk3+/+ mice. F Expected and observed frequency of indicated offspring genotypes from intercrosses of Ripk3D143N/+ mice. G Representative images of H&E staining in indicated tissues of Ripk3+/+ and Ripk3D143N/D143N mice. Scale bar, 100 μm. H Representative images of PI staining in MEFs of indicated mice treated with GSK’872(10 μM) for 8 h. Scale bar, 50 μm. I Cell viability of Ripk3+/+, Ripk3-/- and Ripk3D143N/D143N MEFs treated with GSK’872 or DMSO for 8 h. J Western blot analysis of full-caspase3, cl-caspase3, and RIPK3 in MEFs of indicated mice treated with GSK’872 or DMSO for 8 h. Data were shown as the mean ± SD. ****p < 0.0001; ns, not significant. Multiple comparisons were evaluated by two-way ANOVA (B-C; I) followed by Tukey HSD post-hoc test.
Evidence revealed that kinase inhibitors of RIPK3 induce apoptosis at high concentrations [37]. The RIPK3 kinase inhibitor GSK’872 has a high affinity for binding to the RIPK3 kinase domain and triggers caspase activation, leading to apoptotic cell death [31, 37]. Our findings showed that treatment with GSK’872 induced robust cell death and cleavage of caspase-3 in Ripk3+/+ MEFs, but not in Ripk3-/- or Ripk3D143N/D143N MEFs (Fig. 1H-J), which consistent with previous studies [48], indicating that the RIPK3 D143N mutation protects against GSK'872-induced apoptosis. Collectively, Ripk3D143N/D143N mice exhibited normal development without spontaneous apoptosis.
Ripk3 D143N/D143N mice-derived cells exert defective necroptosis
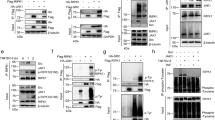
The kinase activity of RIPK3 is indispensable for necroptosis through MLKL phosphorylation [20,21,22]. To characterize the functional impact of RIPK3 D143N mutation on necroptotic signaling, primary MEFs from Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- mice were treated with necroptotic stimuli. Both Ripk3D143N/D143N and Ripk3-/- MEFs displayed complete resistance to TNF -induced necroptosis (Fig. 2A), which was consistently observed in primary BMDMs (Fig. 2B). Similar necroptosis resistance was evident in MEFs challenged with Herpes Simplex Virus Type 1 (HSV-1) infection (Fig. 2C) and BMDMs subjected to TLR3/4 activation (Fig. 2D). Notably, the phosphorylation of RIPK3 and MLKL were completely abolished by RIPK3 D143N mutation in both MEFs and BMDMs (Fig. 2E-F). The kinase activity of RIPK3 is required for its interaction with MLKL, which is critical for necroptosis execution [27]. Similar to RIPK3 D161N mutation, the RIPK3 D143N mutation disrupted the formation of RIPK3-MLKL complex (Fig. 2G), and still maintained interaction with RIPK1, though the binding appears attenuated compared to wild-type RIPK3 (Fig. S3). These results demonstrate that D143N mutation of RIPK3 prevents its activation and interaction with MLKL, thereby blocking necroptosis.
A Representative image of PI staining in Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- MEFs treated with T + S + Z (TNFα+Smac mimetic+z-VAD) for 16 h, and the quantification of PI positive cells was determined. (TNFα: 40 ng/mL, Smac mimetic: 100 nM, z-VAD: 20 μM). Scale bar, 50 μm. B Cell viability of Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- BMDMs treated with T + S + Z or DMSO as control for 24 h. C Cell viability of Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- MEFs infected with HSV-1 (MOI = 5) or Mock for 24 h. D Cell viability of Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- BMDMs treated with LPS+z-VAD (L + Z), Poly(I:C) + z-VAD (P + Z), Smac mimetic+z-VAD (S + Z) or DMSO as control for 24 h. (LPS: 20 ng/mL, Poly(I:C): 50 μg/mL). E Western blot analysis of p-RIPK1, RIPK1, p-RIPK3, RIPK3, p-MLKL, and MLKL levels in Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- MEFs treated with T + S + Z or DMSO as control for 6 h. F Western blot analysis of p-RIPK1, RIPK1, p-RIPK3, RIPK3, p-MLKL, and MLKL levels in Ripk3+/+, Ripk3D143N/D143N and Ripk3-/- BMDMs treated with LPS+z-VAD, Smac mimetic+z-VAD or DMSO as control for 6 h. G Co-immunoprecipitation analysis of MLKL interaction in lysates of 293 T reconstituted with RIPK3 WT, RIPK3 D143N, or RIPK3 D161N, and then immunoprecipitated for RIPK3. Data were shown as the mean ± SD. ****p < 0.0001, Multiple comparisons were evaluated by one-way ANOVA (A) or two-way ANOVA (B-D) followed by Tukey HSD post-hoc test.
RIPK3 D143N mutant rescues the embryonic lethality caused by caspase-8 deficiency
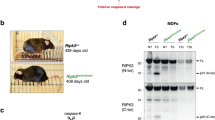
Caspase-8-deficient mice exhibit embryonic lethality due to robust necroptosis and impaired vascular, cardiac and hematopoietic development, which is entirely rescued by RIPK3 deletion [21, 49]. To examine the effect of RIPK3 D143N mutation in mouse embryonic development, Ripk3D143N/D143N mice were interbred with Caspase (Casp)8+/- mice. We found that homozygous RIPK3 D143N mutation completely rescued caspase-8 deletion-caused lethality in mice (Fig. 3A). Casp8-/-Ripk3D143N/D143N mice exhibited similar RIPK3 protein level compared to control Ripk3+/+ mice (Fig. 3B). Besides, Casp8-/-Ripk3D143N/D143N mice displayed normal fertility (Fig. 3C). Additionally, BMDMs from Ripk3D143N/D143N or Casp8-/-Ripk3D143N/D143N mice exhibit complete resistance to necroptosis induced by TLR4 or TNFR1 activation, demonstrating the key role of RIPK3 kinase activity in necroptosis (Fig. 3D-E). Together, these results establish that the RIPK3 D143N mutation effectively prevents caspase-8 deficiency-induced embryonic lethality by blocking RIPK3-dependent necroptosis.
A Expected and observed frequency of indicated offspring genotypes from intercrosses of Casp8+/-Ripk3D143N/+mice. B Western blot analysis of RIPK3 and Casp-8 in lymph node, spleen and thymus of Ripk3+/+ and Casp8-/-Ripk3D143N/D143N mice. C Expected and observed frequency of indicated offspring genotypes from intercrosses of Casp8-/-Ripk3D143N/D143N mice. D Cell viability of Ripk3+/+, Ripk3D143N/D143N, Casp8-/-Ripk3-/- and Casp8-/-Ripk3D143N/D143N MEFs treated with LPS+z-VAD, Smac mimetic+z-VAD or DMSO as control for 24 h. E The expression level of Casp-8 and RIPK3 in BMDMs of Ripk3+/+, Ripk3D143N/D143N, Casp8-/-Ripk3-/- and Casp8-/-Ripk3D143N/D143N mice. Data were presented as mean ± SD, ****p < 0.0001. Multiple comparisons were evaluated by two-way ANOVA (D) followed by Tukey HSD post-hoc test.
RIPK3 D143N mutation in mice confers less protection against TNF-induced SIRS compared to RIPK3 deletion
Emerging evidence implicates RIPK3 in the regulation of multiple pathological conditions including SIRS [40]. To delineate the kinase-dependent and -independent functions of RIPK3, Ripk3+/+, Ripk3-/-, Ripk3D143N/D143N and Ripk3D143N/+ mice were challenged with TNF, then survival rate and body temperature of mice were continuously monitored. Our findings demonstrated that RIPK3 deficiency markedly protected mice from TNF-induced lethal shock, while was mildly but significantly attenuated in Ripk3D143N/D143N mice (Fig. 4A-B). Similarly, RIPK3 D143N mutant conferred partial protection against cecum damage compared to RIPK3 deficiency (Fig. 4C). SIRS-induced IL-6 production in serum was also partially reduced in Ripk3D143N/D143N mice compared to Ripk3-/- mice (Fig. 4D). These results suggest that RIPK3 mediates TNF-induced SIRS through both kinase-dependent and -independent mechanisms.
A Survival rate of Ripk3+/+ (n = 15), Ripk3D143N/+ (n = 9), Ripk3D143N/D143N (n = 11) and Ripk3-/- (n = 13) mice injected with recombinant mouse TNF-α (0.25 μg/g). B Rectal temperature of mice at indicated time points post TNF-α injection. C Representative H&E images of cecal tissues of indicated mice 4 h post TNF-α injection, and the quantification of pathological score was determined. Scale bar, 50 μm. Arrows indicate focal inflammatory infiltration and tissue damage (epithelial disruption, crypt destruction). D Serum IL-6 protein level of indicated mice 4 h post TNF-α or PBS injection. E RNA-seq was performed on cecal tissues from Ripk3+/+, Ripk3-/- and Ripk3 D143N/D143N mice 4 h post TNF-α or PBS treatment. Differentially expressed genes (DEGs) were analyzed, and heatmap of cytokine- and chemokine-related genes was shown. F Functional enrichment of differentially expressed genes (DEGs) in cecal tissues of Ripk3D143N/D143N mice compared to Ripk3-/- mice. G The expression of p-JAK1, JAK1, p-STAT1, STAT1, p-ERK and ERK in cecal tissues of mice injected with TNF-α or PBS as control. Each lane represents the sample from an individual mouse. Data were presented as mean ± SEM, *p < 0.05, ***p < 0.001, ****p < 0.0001, ns, no significance. Survival comparisons were evaluated by log-rank test (A). Multiple comparisons were evaluated by two-way ANOVA (B) and one-way ANOVA (C-D) followed by Tukey HSD post-hoc test.
To elucidate the kinase-independent function of RIPK3 in SIRS, we performed transcriptome sequencing (RNA-seq) on cecal tissues from Ripk3+/+, Ripk3-/- and Ripk3 D143N/D143N mice subjected with TNF-α or the control PBS administration. Differentially expressed genes (DEGs) analysis suggested that the expression of inflammation cytokines and chemokines were markedly repressed in Ripk3-/- mice compared to WT mice, this suppression was less evident in Ripk3 D143N/D143N mutant mice (Fig. 4E). Pathway enrichment of DEGs revealed significant activation of innate immune response and interferon (IFN) signaling pathway in Ripk3D143N/D143N mice compared to Ripk3-/- mice (Fig. 4F), indicating a kinase-independent role of RIPK3 in regulating IFN signaling pathway. Western blot analysis showed that TNF challenge induced activation of JAK-STAT1 and ERK signaling pathways in cecum from both WT and Ripk3D143N/D143N mice, whereas these responses were abolished in Ripk3-/- mice (Fig. 4G). These results demonstrated that RIPK3 deletion, but not D143N mutation, effectively suppressed JAK-STAT1 and ERK activation in TNF-induced SIRS.
Inhibition of JAK-STAT1 signaling mitigates TNF-induced SIRS in Ripk3 D143N/D143N mice
To assess the functional contribution of JAK-STAT1 and ERK signaling to TNF-induced pathogenesis in Ripk3D143N/D143N mice, TNF-challenged Ripk3D143N/D143N mice were administered with JAK1/2 inhibitor ruxolitinib, ERK inhibitor SCH772984 or vehicle control. Notably, pharmacological inhibition of JAK1/2, but not ERK, significantly ameliorated hypothermia in Ripk3D143N/D143N mice (Fig. 5A), while mitigated pathological damage in cecum (Fig. 5B), and reduced systemic IL-6 level (Fig. 5C). The RIPK1 kinase inhibitor Zharp1-211 [12] similar to ruxolitinib also prevented SIRS in Ripk3D143N/D143N mice (Fig. 5A-C), demonstrating RIPK1 kinase-dependent regulation of pathogenesis in Ripk3D143N/D143N mice. Meanwhile, RIPK1 inhibition conferred stronger protection against SIRS than JAK1/2 inhibition in TNF challenged Ripk3+/+ mice, which only partially attenuated the response (Fig. S4), those results support that both RIPK3-mediated necroptosis and RIPK3 scaffold-driven inflammatory signaling act synergistically in protection against TNF-induced SIRS. TUNEL staining analysis showed that inhibition of RIPK1 kinase or JAK1/2 but not ERK significantly reduced cell death in the cecum tissue (Fig. 5D). Notably, treatment with Zharp1-211 or ruxolitinib significantly suppressed STAT1 and ERK phosphorylation in Ripk3D143N/D143N mouse cecum (Fig. 5E). Collectively, these findings demonstrate the scaffold function of RIPK3 plays an important role in driving inflammation and pathology via JAK-STAT1 pathway in TNF-induced SIRS.
A-C Ripk3D143N/D143N mice were intraperitoneally injected with vehicle (n = 13), Zharp1-211 (5 mg/kg) (n = 6), Ruxolitinib (30 mg/kg) (n = 7) or SCH772984 (30 mg/kg) (n = 5) for 45 min, followed by the tail intravenous injection of mouse TNF-α (0.25 μg/g). Ripk3+/+ mice injected with vehicle (n = 5) as control. Body temperature loss (A) was monitored. The mice were sacrificed 4 h after TNF-α administration, and histology of the cecal tissue was analyzed by H&E staining and the representative images (B) were shown. Scale bar, 50 μm. The serum concentration of IL-6 (C) was measured. D Representative TUNEL images of cecal tissues of Ripk3D143N/D143N mice treated with vehicle, Zharp1-211, Ruxolitinib or SCH772984, and the quantification of TUNEL+ cells were shown. Scale bar, 20 μm. E The expression of p-STAT1, STAT1, p-ERK and ERK in cecal tissues of Ripk3D143N/D143N mice treated as indicated. Data were presented as mean ± SEM, *p < 0.05, **p < 0.01, ***p < 0.001, ***p < 0.0001, ns, no significance. Multiple comparisons were evaluated by two-way ANOVA (A) and one-way ANOVA (B-D) followed by Tukey HSD post-hoc test.
Discussion
RIPK3 critically regulates necroptosis, inflammation and promotes apoptosis under certain conditions. Dissecting the distinct contribution of its kinase activity and non-kinase scaffold function to disease pathogenesis is crucial for developing targeted therapeutic strategies. To address this, we generated RIPK3 D143N knock-in mice, harboring a kinase-inactivate mutation. RIPK3 D143N knock-in mice were completely rescued from caspase-8 deficiency-induced embryonic lethality. Importantly, genetically employing this model, we demonstrate that RIPK3 promotes JAK-STAT1 signaling to drive inflammation and tissue damage in TNF-induced SIRS via a kinase-independent (scaffold) mechanism. Our findings establish the Ripk3D143N/D143N mouse as a valuable genetic tool for dissecting kinase-dependent and kinase-independent RIPK3 functions and uncover the critical role of RIPK3 scaffold in the pathogenesis of inflammatory disease.
Current research on RIPK3’s non-kinase functions largely depend on kinase-dead mutants and kinase inhibitors. Studies have shown that the RIPK3 D161N kinase-dead mutation triggers spontaneous caspase-8-dependent apoptosis, leading to embryonic lethality in mice [32]. RIPK3 K51A mutation does not impair mouse viability and rescues caspase-8 deficiency-induced embryonic lethality, however, this mutation markedly reduces RIPK3 protein expression in mouse tissues [37]. Besides, although the Ripk3Δ/Δ kinase mutation rescued embryonic lethality in Fadd-/- embryos, Fadd-/-Ripk3Δ/Δ mice died within 1 day after birth due to massive inflammation [50]. Additionally, RIPK3 kinase inhibitors themselves can induce apoptosis [37], limiting the study of RIPK3’s kinase-independent functions in disease contexts. Caspase-8 serves as a critical regulator of apoptosis, with its deficiency leading to embryonic lethality in mice, which can be rescued by RIPK3 deletion through prevention of caspase-8 deficiency-triggered necroptosis [49]. These challenges highlight the need for more refined approaches to elucidate the non-kinase roles of RIPK3 in disease pathogenesis. Notably, our study revealed that homozygous Ripk3D143N/D143N mice prevented caspase-8 knockout-induced embryonic lethality, with the offspring exhibiting normal viability, fertility and RIPK3 expression in tissues, which provided a powerful genetic tool for dissecting RIPK3’s kinase-dependent and kinase-independent functions. Our study shown that MEFs derived Ripk3D143N/D143N cells are protected from necroptosis without triggering spontaneous apoptosis or RIPK3 inhibitor-induced apoptosis, consistent with observations from a recent preprint [48]. However, the D143N mutation did not block GSK’872-induced apoptosis in 3T3-SA cells [37], indicating that the effects of this mutation are cell type-dependent.
While RIPK3 has been implicated in inflammatory diseases, including TNF-induced SIRS [13, 40] and graft-versus-host disease (GVHD) [12], the specific contribution of its kinase-dependent and kinase-independent functions remain unclear. In this study, we demonstrate that both RIPK3 knockout and RIPK3 D143N mutation prolonged mouse survival, mitigated hypothermia, reduced cecal injury, and decreased systemic IL-6 level in TNF-induced SIRS, with RIPK3 knockout exhibiting superior protection compared to RIPK3 D143N mutation. Our findings suggest that both kinase-dependent necroptosis and kinase-independent function of RIPK3 contribute to TNF-induced SIRS. Transcriptomic analysis of cecal tissues revealed that both Ripk3-/- and Ripk3D143N/D143N mice exhibited reduced inflammatory and chemokine gene expression compared to WT controls. However, Ripk3D143N/D143N mice displayed stronger activation of IFN signaling pathway than Ripk3-/- mice, suggesting the non-kinase activity of RIPK3 in potentiating IFN signaling. Furthermore, RIPK3 deficiency, but not RIPK3 D143N mutation, inhibited the activation of STAT1 and ERK, and inhibition of JAK-STAT1 signaling markedly alleviated TNF-induced SIRS in Ripk3D143N/D143N mice. Inflammatory cytokines act synergistically to drive cell death and amplify inflammatory responses [51]. TNF challenge in both Ripk3+/+ and Ripk3D143N/D143N mice led to a marked upregulation of inflammatory cytokines promoting the activation of inflammatory signaling pathways such as the JAK-STAT1 pathway. These results indicate that while RIPK3 kinase-dependent inflammatory necroptosis contributes to TNF-induced SIRS, the kinase-independent function of RIPK3 in promoting JAK-STAT1 activation also plays an important role in amplifying inflammation and tissue damage.
In our previous study, we found that RIPK1, but not RIPK3, could directly interact with JAK1 [12]. In contrast, the present results demonstrate that the scaffold function of RIPK3 promotes the activation of the JAK/STAT1 pathway, and that this process depends on the kinase activity of RIPK1. These findings suggest that the RIPK3 D143N mutation retains the scaffold function to coordinate with RIPK1 to activate its kinase function, which in turn promotes the activation of the JAK-STAT1 pathway.
In summary, our findings establish Ripk3D143N/D143N mice as a valuable genetic tool for investigating the kinase-dependent and scaffold functions of RIPK3, providing crucial insights for therapeutic targeting of RIPK3 kinase and scaffold functions in inflammatory disorders.
Materials and methods
Reagents and antibodies
Recombinant mouse TNF-α was purchased from GenScript. Smac mimetic was kindly provided by Dr. Xiaodong Wang (National Institute of Biological Sciences, Beijing). z-VAD was purchased from Selleck. LPS was purchased from Sigma Aldrich (L2630). Poly(I:C) was purchased from InvivoGen (tlrl-pic). GSK’872 was purchased from Selleck. Zharp1-211 was synthesized as previously described [12]. The following antibodies were used: mouse p-RIPK1 (53286, CST), RIPK1 (610459, BD), mouse p-RIPK3 (91702, CST), mouse RIPK3 (2283, Prosci), human RIPK3(13526, CST), mouse p-MLKL(ab196436, abcam), mouse MLKL (AP14272b, abgent), Fl-Caspase-3 (9662, CST), Cl-caspase-3 (9661,CST), Fl-caspase-8 (4790, CST), PARP (9542, CST), p-JAK1 (74129, Abcam), JAK1 (50996, CST), p-STAT1 (9167, CST), STAT1 (14994, CST), p-ERK (4370, CST), ERK (9102,CST), Anti-FLAG®M2 Affinity Gel (A2220, Sigma), Anti-Flag HRP (A8592, Sigma), Actin (A2066, Sigma), GAPDH (R1210-1, HUABIO).
Cell culture
Human cervical carcinoma HeLa cells and embryonic kidney 293 T cells were from ATCC. Mouse embryonic fibroblast (MEF) and bone marrow-derived macrophages (BMDMs) were obtained as previously described [20, 52]. HeLa, 293 T and MEF were cultured in DMEM medium supplemented with 10% FBS and 1% penicillin-streptomycin. BMDMs were maintained in RPMI-1640 medium supplemented with 10% FBS and 1% penicillin-streptomycin. All cells were cultured at 37 °C and 5% CO2.
Mice
Mouse line C57BL/6 J Ripk3-/- was kindly provided by Dr. Xiaodong Wang (National Institute of Biological Sciences, Beijing). C57BL/6 J Ripk3D143N/D143N, Ripk3D143N/+, Ripk3+/+, Caspase8-/-Ripk3-/-, Caspase8-/-Ripk3D143N/D143N and Caspase8+/-Ripk3D143N/+ mice were constructed in our laboratory. 6-8-week-old female mice were used for the experiment. Mice were randomly assigned to the experimental groups. All mice were maintained in the specific pathogen-free (SPF) facility of the Suzhou Institute of Systems Medicine.
Cell viability assay
The cell viability was determined by measuring ATP levels using the Cell Titer-Glo Luminescent Cell Viability Assay kit (Promega, USA) according to the manufacturer’s instructions.
Western blot analysis
Cell pellets or tissue samples were suspended using lysis buffer (20 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% Triton X-100, 10% glycerol, 25 mM β-glycerol phosphate, 1 mM Na3VO4) containing PMSF and protease inhibitors. Cell lysate was incubated on ice for 20 min, followed by centrifugation at 13 000×g for 20 min at 4 °C. Then supernatants were collected and subjected to western blot analysis of the indicated proteins.
Cytometric bead array
Protein level of IL-6 in serum of mice subjected with TNF-α administration was determined using the CBA Mouse Inflammation Kit (BD, 552364) according to the manufacturer’s protocols.
PI and TUNEL staining
Cells were seeded in plates and treated as indicated, then Propidium iodide (PI) staining was conducted to evaluate cell death. TUNEL staining for detection of dead cells in cecal tissues was performed according to manufacturer’s protocols (C1086, Beyotime).
H&E staining
Cecal tissues of mice were fixed in 4% PFA and embedded in paraffin, then H&E staining was performed according to manufacturer’s protocols. The cecum histopathological scoring evaluates three key parameters: (1) Inflammatory infiltration (0=normal; 0.5=scattered infiltrates without wall thickening; 1=dense infiltrates without thickening; 2=dense infiltrates with marked wall thickening), (2) Villus morphology (0=normal; 0.5=shortened length; 1=shortened length with reduced density; 2=focal villus loss), and (3) Paneth cells/crypt glands (0=normal; 0.5=Paneth cell reduction; 1=mild reduction in both cell types; 2=severe depletion). Scores progress from 0 (normal) to 2 (severe) for each parameter, with intermediate 0.5 and 1 scores reflecting gradations of pathological changes.
TNF-induced systemic inflammatory response syndrome (SIRS) model
C57BL/6 mice were intravenously injected with mouse TNF-α (0.25 µg/g). Mouse mortality and anal temperature were monitored for the indicated time. Serum was collected 4 h after TNF-α administration for further analysis. For inhibitor experiments, RIPK1 inhibitor Zharp1-211 [12], JAK1/2 inhibitor ruxolitinib [53], and ERK inhibitor SCH772984 [54] were diluted in 7.3% Cremophor EL, and then dissolved in sterile 67.5% PBS containing 22.5% PEG400 plus 2.7% DMSO, and intraperitoneally injected 45 min before TNF-α administration.
Bulk RNA-seq analysis
C57BL/6 Ripk3+/+, Ripk3-/- and Ripk3D143N/D143N mice were intravenously injected with mouse TNF-α (0.25 µg/g) for 4 h, and then cecum tissues were harvested. Total RNA of tissue samples was extracted using TRIzol reagent (Vazyme) and sequenced on the Illumina HiSeq platform. Gene expression levels were quantified using the FPKM method with HTSeq. Differential gene expression analysis was conducted with significant differences defined as a log2fold-change ≥ 0.5 and a P value < 0.05. The raw data are available in the NCBI Gene Expression Omnibus (GEO) database under accession number GSE305184. All data are available in the main text or the supplementary materials.
Statistical analysis
Data were represented as means ± SD or means ± SEM. All experiments were repeated at least 3 times with similar results. Statistical analysis was conducted using GraphPad Prism. Statistical differences between groups were evaluated using one-way analysis of variance (ANOVA) or two-way ANOVA followed by Tukey HSD post-hoc test. Survival curves were plotted using Kaplan-Meier estimates and compared using the log-rank test. Graphical abstract was created in BioRender. Yu, X. (2026) https://BioRender.com/p99d242.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Kerr JFR, Wyllie AH, Currie AR. Apoptosis: a basic biological phenomenon with wideranging implications in tissue kinetics. Br J Cancer. 1972;26:239–57.
Vitale I, Pietrocola F, Guilbaud E, Aaronson SA, Abrams JM, Adam D, et al. Apoptotic cell death in disease-Current understanding of the NCCD 2023. Cell Death Differ. 2023;30:1097–154.
Micheau O, Tschopp J. Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell. 2003;114:181–90.
Wang L, Du F, Wang X. TNF-α induces two distinct caspase-8 activation pathways. Cell. 2008;133:693–703.
Yuan J, Ofengeim D. A guide to cell death pathways. Nat Rev Mol Cell Biol. 2024;25:379–95.
He S, Wang X. RIP kinases as modulators of inflammation and immunity. Nat Immunol. 2018;19:912–22.
Linkermann A, Green DR. Necroptosis. N Engl J Med. 2014;370:455–65.
Holler N, Zaru R, Micheau O, Thome M, Attinger A, Valitutti S, et al. Fas triggers an alternative, caspase-8–independent cell death pathway using the kinase RIP as effector molecule. Nat Immunol. 2000;1:489–95.
Lin Y, Choksi S, Shen H-M, Yang Q-F, Hur GM, Kim YS, et al. Tumor necrosis factor-induced nonapoptotic cell death requires receptor-interacting protein-mediated cellular reactive oxygen species accumulation *. J Biol Chem. 2004;279:10822–8.
Vercammen D, Beyaert R, Denecker G, Goossens V, Van Loo G, Declercq W, et al. Inhibition of caspases increases the sensitivity of L929 cells to necrosis mediated by tumor necrosis factor. J Exp Med. 1998;187:1477–85.
Chan FK-M, Shisler J, Bixby JG, Felices M, Zheng L, Appel M, et al. A role for tumor necrosis factor receptor-2 and receptor-interacting protein in programmed necrosis and antiviral responses *. J Biol Chem. 2003;278:51613–21.
Yu X, Ma H, Li B, Ji Y, Du Y, Liu S, et al. A novel RIPK1 inhibitor reduces GVHD in mice via a nonimmunosuppressive mechanism that restores intestinal homeostasis. Blood. 2023;141:1070–86.
Duprez L, Takahashi N, Van Hauwermeiren F, Vandendriessche B, Goossens V, Vanden Berghe T, et al. RIP kinase-dependent necrosis drives lethal systemic inflammatory response syndrome. Immunity. 2011;35:908–18.
Yuan J, Amin P, Ofengeim D. Necroptosis and RIPK1-mediated neuroinflammation in CNS diseases. Nat Rev Neurosci. 2019;20:19–33.
Zhou W, Yuan J. Necroptosis in health and diseases. Semin Cell Developmental Biol. 2014;35:14–23.
Conrad M, Angeli JPF, Vandenabeele P, Stockwell BR. Regulated necrosis: disease relevance and therapeutic opportunities. Nat Rev Drug Discov. 2016;15:348–66.
Kaiser WJ, Upton JW, Mocarski ES. Viral modulation of programmed necrosis. Curr Opin Virol. 2013;3:296–306.
Upton JW, Kaiser WJ, Mocarski ES. Virus Inhibition of RIP3-Dependent Necrosis. Cell Host Microbe. 2010;7:302–13.
Oberst A, Dillon CP, Weinlich R, McCormick LL, Fitzgerald P, Pop C, et al. Catalytic activity of the caspase-8–FLIPL complex inhibits RIPK3-dependent necrosis. Nature. 2011;471:363–7.
He S, Wang L, Miao L, Wang T, Du F, Zhao L, et al. Receptor Interacting Protein Kinase-3 Determines Cellular Necrotic Response to TNF-α. Cell. 2009;137:1100–11.
Cho Y, Challa S, Moquin D, Genga R, Ray TD, Guildford M, et al. Phosphorylation-Driven Assembly of the RIP1-RIP3 Complex Regulates Programmed Necrosis and Virus-Induced Inflammation. Cell. 2009;137:1112–23.
Zhang D-W, Shao J, Lin J, Zhang N, Lu B-J, Lin S-C, et al. RIP3, an Energy Metabolism Regulator That Switches TNF-Induced Cell Death from Apoptosis to Necrosis. Science. 2009;325:332–6.
He S, Liang Y, Shao F, Wang X. Toll-like receptors activate programmed necrosis in macrophages through a receptor-interacting kinase-3–mediated pathway. Proc Natl Acad Sci. 2011;108:20054–9.
Kaiser WJ, Sridharan H, Huang C, Mandal P, Upton JW, Gough PJ, et al. Toll-like Receptor 3-mediated Necrosis via TRIF, RIP3, and MLKL *. J Biol Chem. 2013;288:31268–79.
Jiao H, Wachsmuth L, Kumari S, Schwarzer R, Lin J, Eren RO, et al. Z-nucleic-acid sensing triggers ZBP1-dependent necroptosis and inflammation. Nature. 2020;580:391–5.
Yang D, Liang Y, Zhao S, Ding Y, Zhuang Q, Shi Q, et al. ZBP1 mediates interferon-induced necroptosis. Cell Mol Immunol. 2020;17:356–68.
Sun L, Wang H, Wang Z, He S, Chen S, Liao D, et al. Mixed Lineage Kinase Domain-like Protein Mediates Necrosis Signaling Downstream of RIP3 Kinase. Cell. 2012;148:213–27.
Wang H, Sun L, Su L, Rizo J, Liu L, Wang L-F, et al. Mixed Lineage Kinase Domain-like Protein MLKL Causes Necrotic Membrane Disruption upon Phosphorylation by RIP3. Mol Cell. 2014;54:133–46.
Chen X, Li W, Ren J, Huang D, He W, Song Y, et al. Translocation of mixed lineage kinase domain-like protein to plasma membrane leads to necrotic cell death. Cell Res. 2014;24:105–21.
Kaczmarek A, Vandenabeele P, Krysko DV. Necroptosis: the release of damage-associated molecular patterns and its physiological relevance. Immunity. 2013;38:209–23.
Xia K, Zhu F, Yang C, Wu S, Lin Y, Ma H, et al. Discovery of a Potent RIPK3 Inhibitor for the Amelioration of Necroptosis-Associated Inflammatory Injury. Front Cell Dev Biol. 2020;8.
Newton K, Dugger DL, Wickliffe KE, Kapoor N, De Almagro MC, Vucic D, et al. Activity of protein kinase RIPK3 determines whether cells die by necroptosis or apoptosis. Science. 2014;343:1357–60.
Sun X, Lee J, Navas T, Baldwin DT, Stewart TA, Dixit VM. RIP3, a novel apoptosis-inducing kinase *. J Biol Chem. 1999;274:16871–5.
Yu PW, Huang BCB, Shen M, Quast J, Chan E, Xu X, et al. Identification of RIP3, a RIP-like kinase that activates apoptosis and NFκB. Curr Biol. 1999;9:539–42.
Kasof GM, Prosser JC, Liu D, Lorenzi MV, Gomes BC. The RIP-like kinase, RIP3, induces apoptosis and NF-κB nuclear translocation and localizes to mitochondria. FEBS Lett. 2000;473:285–91.
Tenev T, Bianchi K, Darding M, Broemer M, Langlais C, Wallberg F, et al. The ripoptosome, a signaling platform that assembles in response to genotoxic stress and loss of IAPs. Mol Cell. 2011;43:432–48.
Mandal P, Berger SB, Pillay S, Moriwaki K, Huang C, Guo H, et al. RIP3 induces apoptosis independent of pronecrotic kinase activity. Mol Cell. 2014;56:481–95.
Kuriakose T, Man SM, Subbarao Malireddi RK, Karki R, Kesavardhana S, Place DE, et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci Immunol. 2016;1:aag2045–aag2045.
Gautam A, Boyd DF, Nikhar S, Zhang T, Siokas I, Van de Velde L-A, et al. Necroptosis blockade prevents lung injury in severe influenza. Nature. 2024;628:835–43.
Berger SB, Kasparcova V, Hoffman S, Swift B, Dare L, Schaeffer M, et al. Cutting Edge: RIP1 Kinase Activity Is Dispensable for Normal Development but Is a Key Regulator of Inflammation in SHARPIN-Deficient Mice. J Immunol. 2014;192:5476–80.
Prado-Acosta M, Jeong S, Utrero-Rico A, Goncharov T, Webster JD, Holler E, et al. Inhibition of RIP1 improves immune reconstitution and reduces GVHD mortality while preserving graft-versus-leukemia effects. Sci Transl Med. 2023;15:eadf8366.
Ito Y, Ofengeim D, Najafov A, Das S, Saberi S, Li Y, et al. RIPK1 mediates axonal degeneration by promoting inflammation and necroptosis in ALS. Science. 2016;353:603–8.
Sharma A, Matsuo S, Yang W-L, Wang Z, Wang P. Receptor-interacting protein kinase 3 deficiency inhibits immune cell infiltration and attenuates organ injury in sepsis. Crit Care. 2014;18:R142.
Newton K, Dugger DL, Maltzman A, Greve JM, Hedehus M, Martin-McNulty B, et al. RIPK3 deficiency or catalytically inactive RIPK1 provides greater benefit than MLKL deficiency in mouse models of inflammation and tissue injury. Cell Death Differ. 2016;23:1565–76.
Sun Y, Ji L, Liu W, Sun J, Liu P, Wang X, et al. Influenza virus infection activates TAK1 to suppress RIPK3-independent apoptosis and RIPK1-dependent necroptosis. Cell Commun Signal. 2024;22:372.
Thapa RJ, Ingram JP, Ragan KB, Nogusa S, Boyd DF, Benitez AA, et al. DAI Senses Influenza A Virus Genomic RNA and Activates RIPK3-Dependent Cell Death. Cell Host Microbe. 2016;20:674–81.
Nogusa S, Thapa RJ, Dillon CP, Liedmann S, Oguin TH, Ingram JP, et al. RIPK3 Activates Parallel Pathways of MLKL-Driven Necroptosis and FADD-Mediated Apoptosis to Protect against Influenza A Virus. Cell Host Microbe. 2016;20:13–24.
Chiou S, Patel KM, Preaudet A, Rickard JA, Horne CR, Young SN, et al. The kinase domain of RIPK3 tunes its scaffolding functions. 2025;2025.04.29.651198.
Kaiser WJ, Upton JW, Long AB, Livingston-Rosanoff D, Daley-Bauer LP, Hakem R, et al. RIP3 mediates the embryonic lethality of caspase-8-deficient mice. Nature. 2011;471:368–72.
Zhao Q, Yu X, Zhang H, Liu Y, Zhang X, Wu X, et al. RIPK3 mediates necroptosis during embryonic development and postnatal inflammation in fadd-deficient mice. Cell Rep. 2017;19:798–808.
Deng B, Wang J, Yang T, Deng Z, Yuan J, Zhang B, et al. TNF and IFNγ-induced cell death requires IRF1 and ELAVL1 to promote CASP8 expression. J Cell Biol. 2024;223:e202305026.
Hou J, Ju J, Zhang Z, Zhao C, Li Z, Zheng J, et al. Discovery of potent necroptosis inhibitors targeting RIPK1 kinase activity for the treatment of inflammatory disorder and cancer metastasis. Cell Death Dis. 2019;10:493.
Marcuzzi A, Rimondi E, Melloni E, Gonelli A, Grasso AG, Barbi E, et al. New applications of JAK/STAT inhibitors in pediatrics: current use of ruxolitinib. Pharmaceuticals. 2022;15:374.
Wang C, Fei K, Liu L, Duan J, Wang Z, Li S, et al. Abnormal activation of NF-κB and MAPK signaling pathways affect osimertinib resistance and influence the recruitment of myeloid-derived suppressor cells to shape the immunosuppressive tumor immune microenvironment. Thorac Cancer. 2023;14:1843–56.
Acknowledgements
We thank Dr. Xiaodong Wang (National Institute of Biological Sciences (NIBS), Beijing, China) for kindly providing Smac mimetic and Ripk3-/- mice. This work was supported by the National Key Research and Development Program of China (No. 2022YFC2502700), the National Natural Science Foundation of China (31830051, 32370810, 82371876, 32500645), the CAMS Innovation Fund for Medical Sciences (2023-I2M-2-005, 2024-I2M-TS-032, 2025-I2M-XHJC-051, 2022-I2M-2-004, 2021-I2M-1-041, 2021-I2M-1-047, and 2021-I2M-1-061), Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (2021-PT180-001, 2019PT310028, 2017NL31004, 2017NL31002), the Special Research Fund for Central Universities, Peking Union Medical College (3332022077, 3332025146), Basic Research Program of Jiangsu (BK20243030, BK20250445), the Suzhou Municipal Key Laboratory (SZS2022005), and the NCTIB Fund for R&D platform for Cell and Gene Therapy.
Author information
Authors and Affiliations
Contributions
S.D.H., X.Y. and X.Z. designed this study and wrote the manuscript. Y.D., J.L., C.Z. and S.Q.H. designed and performed the majority of the experiments, analyzed the data, and wrote the manuscript. Z.L. synthesized the chemical compound and analyzed the data, J.Q analyzed the bulk RNA-seq data, Q.L., X.X. and C.Z. provided technical assistance and expertise for TNF-induced SIRS experiments, L.X. and F.M. provided technical assistance for data analysis.
Corresponding authors
Ethics declarations
Competing interests
X. Z. and S.D.H. are co-founders, consultants, and shareholders of Accro Bioscience Inc, which supports research in their labs. The remaining authors declare no competing financial interests.
Ethics approval statement
All animal experiments were performed in accordance with protocols approved by the Suzhou Institutes of Systems Medicine Institutional Animal Care and Use Committee (ISM-IACUC-0037-R).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Du, Y., Li, J., Zhao, C. et al. A viable kinase-inactive RIPK3 D143N mouse model reveals its scaffold function in driving TNF-induced inflammatory disorder. Cell Death Discov. 12, 107 (2026). https://doi.org/10.1038/s41420-026-02962-x
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41420-026-02962-x