Abstract
Silk fibers have been used by humans for millennia to create textiles and have recently gained the attention of scientists due to their unsurpassed mechanical properties. These properties arise from a sophisticated process by which the starting material, a liquid feedstock consisting of high-molecular-weight silk proteins, is rapidly converted within silk glands into solid fibers with a multi-scale hierarchical structure that is responsible for the material’s incredible robustness. Recently, liquid-liquid phase separation (LLPS) has emerged as a powerful framework for understanding the self-assembly behavior of silk proteins. Interestingly, LLPS-associated proteins typically exhibit disordered or dynamic conformations and have sequences rich in low-complexity multivalent repeats, reminiscent of silk protein sequences. In this review, we explore the evidence indicating that LLPS is a major aspect of silk fiber storage and assembly in both lepidopteran and spider systems. We discuss insights derived from comparative analyses of amino acid sequences, specific chemical triggers, and potential chemical interactions and contextualize the results from recent empirical investigations based on native and recombinant silk materials. We also discuss how LLPS mechanisms might be applied to the sustainable production of silk-like materials that replicate native hierarchical structures. Finally, we outline important areas for future investigations and speculate on how findings from the field of silk research may help illuminate the more general field of biomolecular condensates.
Similar content being viewed by others
Introduction
Silks are protein-based, externally acting fibers produced by arthropods as a means for their survival [1]. Silks have been extensively utilized in textile production for decades and have more recently been recognized as outstanding materials that possess remarkable mechanical properties, such as high tensile strength, extensibility, and toughness, surpassing those of many man-made materials [2,3,4,5]. The mechanical properties of silk, along with its lightweight nature, biocompatibility [6], and biodegradability [7], have made it an attractive material for applications in the biomedical [8,9,10], materials, automotive [11], and defense fields. Due to silk fibers’ superior properties, researchers have been exploring ways to produce them on a large scale and modify their performance.
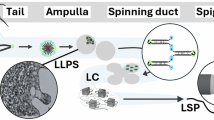
The main structural components of silk are large proteins, termed fibroins in silkworm silk and spidroins in spider silk, that are characterized by repetitive sequence motifs that drive self-assembly and ultimately confer remarkable mechanical properties. Inside the silk gland, these proteins exist as a highly concentrated liquid precursor, or silk dope, that is ready to be transformed into insoluble fiber upon processing though the spinning duct apparatus [12]. Studies have shown that various factors such as pH [13], ion concentrations [14, 15], and shear forces [16] help drive the transition of soluble silk protein into insoluble fiber. Therefore, attempts to replicate the physicochemical changes/gradients encountered by these proteins in the natural spinning process in order to produce native-like artificial silk fibers, known as biomimetic methods, have recently gained attention [17, 18].
The organization of silk proteins in the prefibrillar soluble state has been mainly viewed through three paradigms: the micelle, liquid crystalline, and liquid-liquid phase separation (LLPS) concepts. The micelle hypothesis proposes that silk proteins in solution assemble into discrete micellar structures, driven by thermodynamic interactions between their amphiphilic sequences and the aqueous environment. In this model, hydrophobic residues, primarily from repetitive domains, form a core, while hydrophilic terminal domains create a surrounding shell [19]. Observations of globular structures in silk materials may be evidence for the existence of micelles [20, 21]. On the other hand, the liquid crystalline theory proposes that during the spinning process, silk proteins adopt fluid yet compact and oriented conformations, consistent with a beads-on-a-string arrangement [22, 23]. Liquid crystallinity is thought to be crucial to the spinnability of silk dope, imparting reduced viscosity and realizing pre-arrangement of the structural elements, which enables controlled deformation and the formation of aligned fibrillar structures upon exposure to shear forces and flow. Recently, LLPS, or coacervation, as described below, has emerged as an alternative framework to understand the self-assembly behavior of silk proteins. It is important to note that despite their differences, the three hypotheses are not mutually exclusive, but are rather different prisms through which to conceptualize the highly complex and dynamic phenomenon of silk assembly. Notably, some recent reports suggest more highly orchestrated mechanisms of self-assembly. For instance, Landreh et al. suggested a hybrid assembly pathway for spider silk that incorporates elements of the LLPS and liquid-crystalline models that are mediated by different repetitive sequence features, termed liquid-liquid crystalline phase separation [24]. Additionally, Moreno-Tortolero et al. proposed that in Bombyx mori fibroin, higher-order intermediates form via stacking of N-terminal domains in response to biophysical changes in the spinning apparatus, leading to characteristic assemblies that initiate fiber formation [25].
LLPS is a spontaneous physiological process in which a homogeneous solution is separated due to a density transition and solvent release above a saturation concentration, resulting in two coexisting phases: a dilute phase and a dense phase [26, 27]. Inside cells, LLPS can compartmentalize proteins and nucleic acids into micron-scale membraneless organelles (or biomolecular condensates) that exhibit liquid characteristics; such condensates can further transform into materials with different states, ranging from viscous liquids and gels to solid aggregates [28,29,30]. Membraneless organelles possess different physiochemical properties and play a vital role in various cellular processes, including transcription, genome organization, and immune response. Furthermore, LLPS has been implicated in several pathologies of protein aggregation [31], in which liquid droplets formed from soluble proteins like TDP-43 and FUS irreversibly transition into fibrillar deposits, leading to neurodegenerative diseases.
LLPS is a product of electrochemical gradient forces within cells and is promoted by multivalent interactions [29, 32, 33]. These interactions are influenced by the spatial arrangement of molecules in droplets. As such, LLPS is highly sensitive to pH, temperature, RNA, salt concentrations, and post-translational modifications [34,35,36,37]. Many of the biomolecular condensates formed through LLPS are rich in intrinsically disordered proteins (IDPs), especially those featuring low sequence complexity and repetitive modules [38]. These arrangements generate numerous weak and transient interactions among the protein chains, resulting in de-mixing into concentrated liquid droplets while avoiding structural collapse or aggregation [36, 39, 40]. In addition to IDPs, structured proteins can also undergo LLPS, especially when they contain tandem repeats of interaction modules, such as multivalent SH3 domains and proline-rich motifs (PRMs) [33].
Similarly, silk proteins exhibit largely disordered conformations in solution and have sequences composed of low-complexity multivalent repetitive motifs; in this regard, silk proteins fit the general pattern of LLPS-associated proteins. While early observations revealed phase separation in dragline silk proteins [41], the first systematic analysis of the subject was reported by Malay et al. (2020) [42], who identified the conditions driving LLPS in recombinantly produced MaSp2 dragline spidroin and proposed the central role of LLPS in fiber self-assembly. Additional studies have added considerably to our understanding of these complex phenomena; however, many details have yet to be elucidated. This review will focus on the research progress made on the self-assembly of spider and silkworm silk fibers, particularly work related to the LLPS theory, including sequence and mechanistic considerations and future research perspectives.
Sequence analysis
The sequence-structure-function relationships of silk proteins have been explored, primarily in the context of fiber mechanics. However, less attention has been given to how these proteins navigate drastic changes in physical and chemical environments—such as extracellular secretion, storage as a concentrated liquid feedstock, and self-assembly in the spinning ducts—prior to the formation of solid fibers. It is likely that silk protein sequences have evolved to balance the competing effects of these transitional states, rather than solely to optimize the mechanical performance of the fibers. In this context, native silk sequences may be adapted for LLPS-driven self-assembly; however, the underlying molecular determinants remain to be fully elucidated.
We will consider silk protein sequences that are dominated by low-complexity, repetitive regions, including silkworm heavy-chain fibroin (Fib-H) [43, 44] and the spidroin components of major ampullate (dragline) silk (MaSp), minor ampullate silk (MiSp), and flagelliform silk (FlSp) [45]. These proteins have a common overall architecture, consisting of an extended repetitive region with low sequence complexity that is flanked by N- and C-terminal domains (NTD and CTD, respectively) with more complex sequences (Fig. 1a). Despite being independently derived in spiders and silkworms, the terminal domains play crucial roles in orchestrating fiber self-assembly and fulfill parallel biochemical functions in a remarkable case of convergent evolution (Table 1; see section below) [46]. However, in addition to being critically influenced by the terminal domains, the properties of the fiber materials are largely determined by the repetitive regions.
Analysis of silk protein sequences. Representative full-length spidroin sequences from the orb weaver spider Araneus ventricosus (MaSp1, MaSp2, MaSp3, MiSp, and FlSp) and silkmoth heavy-chain fibroins (Fib-H) from Bombyx mori and Antheraea pernyi. A Overall architecture of the silk proteins, with the relevant domains indicated. B Amino acid composition of the core repetitive sequences. C Violin plot illustrating the distribution of Tyr residues (the putative associative units during LLPS), represented as the distance (in number of residues) between successive Tyr units along the repetitive sequences
In a simplified schema, silk fibers have a semi-crystalline organization consisting of crystalline (hard) and semi-amorphous (soft) regions, with the former providing mechanical strength through interchain crosslinking and the latter providing elasticity upon deformation [43, 47,48,49]. For MaSp and MiSp spidroins, the crystalline crosslinks are provided by poly-alanine blocks that form stacked β-sheets within the fibers, with a remarkably similar design to that found in wild silkmoth fibroins, e.g., Antheraea pernyi (Table 1). For FlSp (at least in Araneus ventricosus), crosslinking is provided by 9-residue blocks enriched in hydrophobic Val residues [50]. In contrast, for B. mori Fib-H, Gly-Ala repeats that form interlocking interchain structures constitute the crystalline component [51].
All of the above silk repetitive sequences present a pronounced compositional bias (Fig. 1b). Notably, the 4 smallest amino acids—Gly, Ala, Ser, and Pro (GASP)—account for a large majority of residues, ranging from 76–89% (Table 1); interestingly, these same 4 residues, along with Gln, are generally similarly overrepresented among LLPS-associated proteins [52, 53]. The high prevalence of these 4 residues likely enables the intrinsically disordered chains inside the silk glands to adopt dynamic, rapidly shifting conformations despite the intense molecular crowding and otherwise preventing their catastrophic aggregation (especially considering the presence of potential crosslinking sites in such confined spaces). These effects are well documented for Gly, which contributes maximum conformational flexibility, and Pro, whose unique backbone topology prevents a collapse into rigid secondary structures [54,55,56]. The high abundance of Ala is primarily associated with the \({{\rm{\beta }}}\)-sheet forming poly-Ala or Gly-Ala repeats. Interestingly, despite its relatively low \({{\rm{\beta }}}\)-sheet propensity [57] and suboptimal contribution to mechanical strength [58], Ala might be preferred for such fiber crosslinking functions due to the evolutionary need to maintain balance between the stabilization of fiber crosslinks and maintenance of conformational dynamics in the prefibrillar solution state. The polar Ser residues are likely needed to enhance the solubility of silk proteins in the silk dope through hydrogen bonding with water molecules. We speculate that GASP residues provide the underlying flexible “backbone”, or scaffold, of protein chains in solution, while the other, bulkier residues provide additional functionality relevant to self-assembly and the modulation of specific interactions.
While silk proteins share similarities in repetitive sequence composition, they also exhibit key differences [59, 60]. MaSp2 is enriched in Pro and Gln-Gln motifs, where Pro, which is also prevalent in FlSp, contributes to β-turn flexibility and dragline silk supercontraction [61,62,63]. MaSp3 has an abundance of charged residues (Arg and Asp) substituting for the Gln found in MaSp1 and MaSp2. MiSp is notably hydrophobic due to abundant Ala in poly-Ala and Gly-Ala motifs. Among silkworm fibroins, B. mori Fib-H consists primarily of repeating GAGAG(S/Y) sequences, interrupted only by 25-residue mini-domains, while A. pernyi Fib-H has an exceptionally high Ala content (45%) due to extended poly-Ala runs (13 residues on average), enhancing hydrophobicity.
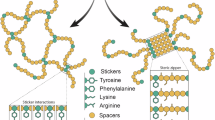
It is useful to consider the repetitive sequences of silk proteins in the context of the prevailing “stickers-and-spacers” paradigm (Fig. 2a, b) that describes how proteins achieve LLPS by balancing associative interactions (stickers) and flexible linkers (spacers) [32, 64]. Specific amino acids or motifs act as stickers that govern transient, multivalent interactions while spacers are more flexible regions that provide conformational flexibility and prevent uncontrolled aggregation, enabling the formation of liquid-like condensates [65]. While typically involving IDPs, folded domains can also act as stickers that interact with repeats of linear motifs via multivalent interactions, while the disordered linkers that connect these domains act as spacers [65].
Overview of silk protein LLPS. A Schematic representation of potential molecular interactions governing LLPS of silk proteins in the prefibrillar state. In solution, high-molecular-weight silk proteins adopt dynamic and largely disordered conformations. The ability of polymeric silk proteins to remain in solution and avoid aggregation, even at very high concentrations, depends on their having evolved optimized sequences. The stickers-and-spacers model of LLPS predicts the existence of sticker motifs, which provide transient and weakly associative interactions, interspersed among more conformationally dynamic spacer regions. In addition, silk protein repeat domains feature segments that will constitute the more strongly associative assemblies (crosslinks) in the spun fibers. B Potential sticker types in silk protein sequences, showing representative residues. C Analysis of the residue composition of well-characterized proteins involved in the formation of biomolecular condensates and representative silk proteins (repetitive regions of MaSp1 from A. ventricosus and Fib-H from B. mori), demonstrating the diversity of sequences associated with LLPS
Aromatic residues are thought to play a key role as stickers driving LLPS via π-π stacking interactions [32]. Tyr is the predominant aromatic residue in silk proteins and is present in all repetitive domains, with an abundance ranging from 3.4–6.9% (Table 1). In addition to their overall abundance, the spatial distribution of Tyr residues is crucial for modulating phase behavior, as uniform spacing helps balance LLPS and unregulated aggregation [66]. An analysis of silk protein sequences (Fig. 1c) reveals a non-clustered yet non-uniform Tyr distribution, which may reflect the need to accommodate crosslinking motifs like poly-Ala within LLPS-prone sequences. In addition to aromatics, charged, polar, and hydrophobic residues can also function as stickers [32, 67]. This may explain observed differences in the LLPS response: recombinant MaSp1, despite having a slightly lower Tyr content than an equivalent MaSp2 construct (3 vs. 3.5% of the repetitive domain), exhibited a greater LLPS propensity in response to potassium phosphate [61]. This difference may be attributed to presence of Arg (3%) in the MaSp1 construct, as Arg can enhance LLPS via π-cation interactions with Tyr [32]. Furthermore, the presence of Leu in the MaSp1 sequence (6.1%) may promote hydrophobic clustering, reducing solubility and driving phase separation (Fig. 2b).
Figure 2c compares the sequence compositions of the repetitive regions of the silk proteins MaSp1 and Fib-H with those of well-established phase-separating proteins involved in biomolecular condensate formation. Within cells, proteins such as FUS, LAF-1, and hnRNPA1 undergo liquid-liquid phase separation (LLPS) to form membraneless organelles with essential biological roles. The phase separation of these proteins is mediated by intrinsically disordered sequences, and in certain cases—such as FUS and hnRNPA1—dysregulated phase separation can lead to pathological aggregation associated with neurodegenerative diseases. An analysis demonstrates that diverse sequence compositions can support LLPS and also reveals some distinct trends. Silk proteins exhibit a relatively low sequence complexity, characterized by exceptionally high abundances of glycine (approaching 50%) and aliphatic residues, primarily alanine, as discussed above. In contrast, intracellular phase-separating proteins tend to be enriched in hydrophilic polar and charged residues. These differences highlight the varied molecular strategies that are used to achieve LLPS, emphasizing the significance of biological and functional context in determining phase behavior.
Conserved nature of silk gland morphology and function
Despite the sequence diversity among different silk proteins, silk glands tend to have a consistent design and similar overall functionality (Fig. 3). This is remarkable considering that spider and moth silk spinning systems have independent evolutionary origins [46, 62, 63, 68]. In general, the gland consists of a storage compartment or sac, which contains the prefibrillar material, and a spinning duct with a narrowing geometry, where the prefibrillar material is assembled into fibrillar structures before being pulled out via a spinneret as solid fibers in the exterior of the organism [22]. Silk proteins (and other silk-associated proteins) are synthesized and secreted into the gland lumen via surrounding epithelial cells, where they are often deposited in distinct layers [69]. The morphology of the silk gland directly impacts the fiber formation process; in particular, the spinning ducts regulate finely tuned interactions between pH gradients, ion shifts, and shear forces to tightly regulate LLPS, nanofibril formation, and shear-induced chain alignment to promote the creation of hierarchically organized fibers (which can be regarded as a controlled aggregation process) while avoiding uncontrolled or premature aggregation of the densely packed protein chains in the silk dope material.
Morphology of spider silk and silkworm glands. Schematic diagram of silk glands with reported pH and ion gradients. A In the spider major ampullate gland, chaotropic ions are replaced by kosmotropic ions, which induce phase separation, while a gradient of decreasing pH induces the dimerization of NTD, which promotes fibril formation. B In the B. mori silk gland, the higher concentration of Ca2+ ions in the lumen in the PSG and MSG prevents premature fibrillation, while in the ASG, the pH decrease and localization of metal ions to the epithelial cells subsequently induce phase separation and fibril formation
pH changes constitute a major aspect of silk spinning [70] and are achieved through carbonic anhydrase activity in both spiders and silkworms. In the spider major ampullate gland, the pH gradient ranges from 7.6 in the secretory (or tail) region to 5.7 or lower near the distal end of the spinning duct [69], while in B. mori, this acidification gradient has been reported to range from pH 8.2–6.2 [71]. While relatively subtle, these changes in pH cause major changes in silk protein conformation, particularly by triggering intermolecular dimerization of spidroins via the NTD [13, 72]. In contrast, in B. mori, fibroin acidification converts a disordered NTD into dimeric β-sheet structures that undergo higher-order stacking [73]. The pH decrease also induces a structural change in the CTD of spidroin, which might trigger the propagation of \({{\rm{\beta }}}\)-sheets in the nascent fibers [74].
Ion gradients are another crucial function imparted by the silk spinning apparatus. The spider major ampullate gland (Fig. 3a) presents a distinct (yet non-quantitative) shift from a chaotropic ion to kosmotropic ion composition [14], with a progressive decrease in sodium and chlorine levels along the direction from the sac toward the spinning duct, accompanied by an increase in the levels of potassium, phosphorus, and sulfur (the latter two presumably representing phosphate and sulfate anions, respectively). The high level of NaCl in the sac is thought to prevent premature protein-protein interactions, thus helping to maintain the solubility of silk proteins prior to spinning [72, 75, 76]. Notably, exposure of recombinant MaSp1 and MaSp2 proteins to phosphate (or other kosmotropic anions) at concentrations beyond a critical threshold has been shown to efficiently trigger LLPS [42, 61, 77], as detailed below. It is worth noting that to date, only the ion composition in major ampullate glands has been elucidated [14]. While ion gradients are likely conserved across different types of spider silk glands, their specific compositions may vary to accommodate the distinct properties of each silk type.
In B. mori, silk fiber assembly is guided by metal ion gradients along the silk gland. The gland consists of three regions (Fig. 3b): the posterior silk gland (PSG), where fibroins are secreted and stabilized by high metal ion concentrations to prevent premature fibrillation [78]; the middle silk gland (MSG), where the silk feedstock is stored, sericin is produced, and Ca2+ levels remain high to further inhibit fibrillation [78,79,80,81]; and the anterior silk gland (ASG), where fibroins undergo phase separation and fibril formation before being extruded through the spinneret [82]. In the ASG, a drop in metal ion concentration within the lumen, coupled with pH shifts and shear forces, initiates the active spinning process [78].
Structural changes in silk proteins during self-assembly
Structurally ordered domains
In spidroin, the NTD folds into a globular structure consisting of 5 α-helices connected by flexible loops [13, 83,84,85]. The NTD oligomerization state is pH and salt dependent [72, 83, 86]. At neutral pH, the NTD is in monomeric form, while under mildly acidic conditions, it undergoes dimerization with antiparallel orientation, giving rise to connected chains that are essential for fibrillization. The dimerization mechanism of the NTD is preserved in different spidroin types [78, 79, 84, 87]. Spidroin NTDs thus play a crucial role in the self-assembly process, although they are not directly involved in LLPS [42].
The CTD of spidroins likewise adopts α-helical globular structures, which form parallel-oriented dimers at neutral pH [80]. The drop in pH during the spinning process has been associated with the structural unfolding of the CTD [80, 81] and has been suggested to regulate solubility [82] as well as the alignment of secondary structures formed by the repetitive domain [80] and to act as a trigger for β-sheet formation [74, 81]. Interestingly, the CTD has been shown to promote LLPS of MaSp2 in response to kosmotropic ions [42]. While part of this effect might be attributed to the effective doubling of the molecular weight of the protein as a consequence of CTD homo-dimerization, the isolated CTD also underwent LLPS in the presence of potassium phosphate at low pH, which might be related to its increased structural disorder [42].
Certain spidroin sequences also contain ordered domains within the core repetitive regions. MiSp contains a 14 kDa α-helical segment within the repetitive region, termed the “spacer” domain, that is conserved among different species [88]. Spacer domains from both A. ventricosus [89] and T. antipodiana (termed the repetitive domains in the relevant studies) [90] exhibit pH-independent conformations, while shear forces were shown to induce fibrillization [89].
Regarding silkworm fibroin, the NTD of Fib-H from B. mori exhibits a random coil structure at neutral pH; this structure is converted into dimeric β-sheet structures, which then form higher-order assemblies [73]. As in the spidroin case, the Fib-H NTD has been suggested to mediate the self-assembly of silk in response to the pH acidification gradient from the lumen of the posterior silk gland to the anterior silk gland [73].
The CTD of B. mori Fib-H is a small (5.7 kDa) domain that is connected to the fibroin light chain (Fib-L) via a disulfide bond. It remains unclear what the role of the CTD is upon acidification or changes in ion concentrations; however, one hypothesis suggests that the spinning mechanism induces micelle formation, in which the hydrophilic NTD is located on the surface while the hydrophilic CTD and Fib-L are located in the inner part of the micelles, regulating the solubility of large structures [91].
Repetitive domains
Various lines of evidence demonstrate that spidroin repetitive sequences adopt predominantly random coil conformations in the prefibrillar solution state [76, 92, 93]. A vibrational circular dichroism (VCD) study of native spidroin from T. clavipes and A. diadematus indicated that its conformation is a mixture of random coils, polyproline II (PPII) helices and some α-helices [92]. Studies using recombinant spidroins have shown that the poly-Ala region of the repetitive domain exhibits either random coil or α-helix structures, with their relative contents depending on the length of the poly-Ala blocks [93,94,95,96]. Upon spinning, spider dragline silk forms a semi-crystalline fiber, with β-sheet nanocrystals embedded within a semi-amorphous matrix. In contrast, the glycine-rich region (GGX) of MaSp silk fiber contains disordered 31 helices embedded in the amorphous region, while the polyalanine region forms a β-sheet structure in the crystalline region [97,98,99,100].
In the presence of terminal domains, acidification causes structural changes in the repeat domain of native spidroin, from random coils/α-helices to β-sheets [96]. On the other hand, the conformation of the isolated recombinant repetitive domain of spidroin does not change at low pH [93]. Interestingly, previous studies indicated that changes in ion composition are strongly related to solubility and interactive effects on the recombinant repetitive domain of spider dragline silk [76]. A higher concentration of phosphate anions promotes oligomerization of the recombinant repetitive domain, which is associated with LLPS [42, 61].
Like spidroin, native silk fibroin undergoes structural changes from silk I to silk II structures when exposed low pH and shear forces. Soluble silk I fibroin is thought to adopt either β-turn structures or helical or partially helical conformations [101]. X-ray diffraction analysis of silk fiber revealed that after spinning, silk II fiber contains antiparallel β-sheet structures[102]. Calcium ions, which are the most abundant metal ions in the silkworm gland [103], have been suggested to induce LLPS of RSF as part of its self-assembly mechanism [104].
LLPS behavior of native or recombinant silk proteins
Growing evidence suggest that silk proteins undergo phase separation, which plays a key role in their hierarchical self-assembly into fibers. Here, we review current findings on the LLPS behavior of native and recombinant silk proteins.
MaSp1 and MaSp2
In recombinant MaSp2, LLPS at neutral pH was found to be triggered by multivalent kosmotropic anions, such as phosphate, sulfate, and citrate [42], resulting in concentrated protein droplets that displayed fusion and surface wetting properties. A comparison of constructs with varying domain combinations revealed that the CTD and repetitive regions were responsible for the LLPS response. The propensity for LLPS was furthermore correlated with the number of repetitive units, with more repeats corresponding to lower ion concentration thresholds for phase separation. MaSp2 subjected to LLPS under more acidic conditions produced condensates with altered morphology, with gel-like structures at pH 6 and solidified fibrillar networks with submicron features at pH 5.0–5.5. Fibril network assembly occurred rapidly and was directly dependent on the dimerization of the NTD domains; interestingly, it also required the full complement of domains on the same polypeptide. In another report, recombinant MaSp1 was likewise found to undergo LLPS upon exposure to kosmotropic anions, albeit with a higher propensity compared with MaSp2 [61]. (Fig. 4a). As discussed above, this higher sensitivity is hypothesized to arise from differences in sequence, where Arg in the MaSp1 repetitive region enables cation-π interactions and hydrophobic Leu residues lower the overall solubility, driving phase separation. MaSp1 further showed a bimodal response to NaCl, where spontaneous LLPS occurred outside a particular concentration window (200–1000 mM background NaCl concentration in a 3 mg/ml solution at room temperature).
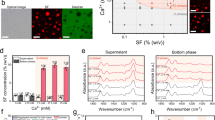
LLPS phenomena in silk proteins. A LLPS propensity of MaSp1 N-R6-C (blue shading) and MaSp2 N-R6-C (red dotted line) in response to potassium phosphate, plotted as a function of the protein concentration (x-axis) and phosphate concentration (y-axis). Under similar conditions, MaSp1 exhibited a markedly higher tendency for phase separation than MaSp2. B LLPS of miniMiSp (40 mg/mL) upon addition of 10 mM Tris buffer (pH 7.0) and 100 mM NaCl. C Phase separation of B. mori silk fibroin (SF) as a function of protein and Ca2+ concentrations. Squares correspond to no LLPS observed, triangles correspond to LLPS observed after 15 min, and circles correspond to LLPS observed immediately after mixing. The data point inside the dashed box corresponds to the inset fluorescence images on the side of the graph (scale bar 50 µm). D Kyte-Doolittle hydropathy plots for the repetitive regions of A. ventricosus MaSp2, MaSp1, B. mori Fib-H and A. ventricosus MiSp. A higher hydrophobicity score might be related to a higher propensity for LLPS and chaotropic-induced LLPS. Panel (A) reprinted with permission from Reference [61]; copyright by John Wiley and Sons Ltd. Panel (B) reprinted with permission from Reference [106]; copyright by Elsevier. Panel (C) reprinted from [104] under the Creative Commons CC BY license
Another study revealed that the propensity for LLPS could be significantly altered by changing the repetitive sequences [105]. In a recombinant mini-spidroin NT2RepCT construct based on MaSp1, the introduction of point mutations, namely Tyr to Phe and Arg to Leu, produced variants that retained the ability to undergo LLPS in response to potassium phosphate, revealing an expanded range of potential sticker residues. Interestingly, such variants exhibited variable droplet size and fluidity compared to the wild type, suggesting a highly sensitive system whose material properties can be tuned through slight changes in amino acid sequence.
Native dragline proteins also undergo LLPS under similar conditions, as observed in native silk dope samples from major ampullate gland of Trichonephila clavata [42]. In response to 0.5 M potassium phosphate (pH 8), the gland extracts readily formed spherical droplets enriched in MaSp proteins, while in 0.5 M potassium phosphate (pH 4.5), fine fibrillar networks were observed. Despite such similar results, it is important to exercise caution when comparing findings between recombinant and native silk systems. Important differences exist between these systems, notably the higher molecular weight and multicomponent spidroin composition in the native system, post-translational modifications, the potential formation of prefibrillar complexes, the presence of accessory proteins and other chemical components, all of which may influence the phase separation behavior of silk proteins.
MiSp
In a recombinant mini-MiSp system, LLPS-mediated fibril formation was induced by NaCl, while no phase separation occurred in the absence of NaCl (Fig. 4b). This contrasts with MaSp1, where NaCl maintains solubility and prevents unwanted interactions [76], suggesting possible differences in ion distribution between the major and minor ampullate glands. A recent study reported that the unstructured repetitive region of recombinant MiSp plays a vital role in LLPS-mediated fiber formation [106] and that Tyr functions as a sticker. Additionally, two studies have considered the conserved globular spacer domain to be important for MiSp [89, 90].
Silkworm silk
Studies on silk fibroin suggest a key role of LLPS in silk fiber assembly, particularly highlighting the influence of calcium ions (Ca2+) and the structural integrity of the fibroin complex. Yang et al. [104] demonstrated that Ca2+, which is abundant within silkworm silk glands, can induce LLPS in regenerated silk fibroin (RSF) under macromolecular crowding conditions. Other divalent cations, such as Zn2+, Cu2+, Mn2+, and Mg2+, along with kosmotropic anions like phosphate and citrate, also promoted LLPS of silk fibroin. Two potential mechanisms were proposed for Ca2+-induced LLPS: (1) hydrophobic interactions facilitated by charge screening and (2) salt bridge formation via electrostatic interactions between Ca2+ and carboxylic residues on the fibroin. Additionally, the binding of heavy chain fibroin (Fib-H) to light chain fibroin (Fib-L) was suggested to positively influence LLPS formation. Notably, Ca2+-induced LLPS of silk fibroin could lead to silk fiber formation upon acidification.
A previous study demonstrated that native silk fibroin (NSF) extracted directly from the silkworm gland has the inherent capability to form nanofibrils. In contrast, even at the same concentration, RSF lacked the ability to self-assemble into nanofibrils. This discrepancy suggests that the native structural integrity of silk fibroin is crucial for its self-assembly properties [107]. Recently, Zaki et al. [108] investigated the LLPS behavior of regenerated undegummed silk (RUS), which maintains native structural components, including Fib-H, Fib-L, and P25. The RUS exhibited LLPS at concentrations above 10% (wt/wt), whereas RSF did not form LLPS under similar conditions. These findings imply that preservation of the native fibroin complex is essential for LLPS and subsequent self-assembly processes. The ability of NSF and RUS to undergo LLPS and form nanofibrils underscores the importance of the native protein architecture in silk fibroin self-assembly and suggests that NSF and RUS have similar properties to those of native protein extracted from the silk glands [109].
Notably, MaSp1/2, MiSp and B. mori Fib-H present very different hydropathy profiles despite having some similarities in the amino acid sequence of the repetitive regions. The repetitive regions of MaSp1/2 exhibit amphiphilic features, while silkworm silk is more hydrophobic, and MiSp is the most hydrophobic of the three (Fig. 4d). The hydrophobicity of the repetitive domain of spidroin could affect which ion species are able to promote LLPS. In the case of amphiphilic proteins like MaSp1/2, kosmotropic ions are required to induce phase separation [42, 61] while chaotropic ions promote solubility [76]. However, for moderately hydrophobic proteins such as silkworm silk, divalent cations are more effective in inducing LLPS [104], and for highly hydrophobic proteins, such as MiSp, monovalent chaotropic ions such as NaCl are sufficient to induce LLPS [106] (Fig. 4d).
Applications of silk protein LLPS
Perhaps the most significant impact of silk protein LLPS to date lies in its contributions to biomimetic methods of spinning artificial silk fibers. Studies using either recombinant MaSp1 [61] or MaSp2 [42] demonstrated that kosmotropic anion-based LLPS, when initiated in combination with acidification (at pH 5.0–5.5), triggered the rapid assembly of fine fibrillar protein networks, a result that is dependent on having the full set of functional domains [42] (Fig. 5a). Under unidirectional deformation, such network structures could readily be converted into insoluble macroscopic fibers with an internal hierarchical architecture that consists of closely packed and aligned submicron fibrils. Notably, the application of mechanical deformation during fiber formation induced the emergence of pronounced \({{\rm{\beta }}}\)-sheet conformations (i.e., the structures responsible for the mechanical strength of native silk fibers). Thus, fully biomimetic and efficient LLPS-based methods for spinning silk fibers feature a genuine hierarchical architecture.
Biomimetic spinning based on LLPS in silk protein. A Condensate structures formed upon mixing labeled MaSp2 N-R12-C with 0.5 M potassium phosphate at different pH values. Liquid droplets that exhibited dynamic fusion behavior were observed at pH 7 and 8. At lower pH, the resulting structures presented a more gel-like appearance (pH 6) or rapidly underwent self-assembly into extended fibril networks (pH 5). Scale bars, 10 µm. B Overview of the microfluidic device used for spinning MaSp2 N-R12-C fibers under biomimetic conditions. The device design includes three inlets (1, 2, and 3), one outlet, and three monitoring sections (A, B, and C). The confluence of the protein solution (1) and citrate-phosphate buffer (pH 7.0) streams (2) induced the formation of LLPS droplets (region A), while subsequent exposure to citrate-phosphate buffer at pH 5.0 (section B) produced a continuous fiber along section C with a hierarchical organization of oriented nanofibrillar bundles. Scale bar, 30 µm. Panel (A) reprinted with permission from Reference [42]; copyright by The American Association for the Advancement of Science. Panel (B) reprinted from [17] under the Creative Commons CC BY license
Furthermore, LLPS-based methods have been successfully integrated into microfluidics-based fiber spinning systems [17, 77] to create platforms that mimic the physiological changes that are induced in the spinning ducts of native silk glands. Recently, Chen et al. successfully replicated the self-assembly of spider silk by using a three-inlet microfluidic device (Fig. 5b) [17], which provided physiological and chemical triggers to structural proteins via a customized channel arrangement and temporal sequence. In their microfluidic system, MaSp2 spidroin solution readily underwent sequential LLPS and fibril formation upon exposure to kosmotropic anion solutions and acidification at points A and B, respectively. Subsequently, in region C, the condensed MaSp2 material was instantaneously transformed into a single continuous fiber under the application of shear and mechanical deformation. To initiate sample flow, negative pressure (vacuum) was applied from the outlet to mimic the natural pultrusion process [22]. Crucially, the authors found that above a critical threshold shear value, the poly-Ala blocks of MaSp2 could be converted into beta-sheet structures, resulting in water-insoluble fibers consisting of closely packed nanofibrillar bundles.
LLPS-based fiber spinning has also been explored in silkworm silk systems, particularly using regenerated undegummed silk (RUS), which retains the native structural integrity of fibroin proteins [108]. Notably, fibers spun from RUS via wet spinning displayed tightly packed nanofibrillar structures characteristic of native silk fiber, whereas those spun from degummed RSF lacked this organization, underscoring the significance of LLPS in producing biomimetic silk fibers. Moreover, the study demonstrated that RUS-derived fibers possess significantly superior mechanical properties to those of fibers derived from RSF.
LLPS has also been shown to induce phase separation in silk protein constructs with different terminal domains. Mohammadi et al. used a three-block protein consisting of spidroin repeat region flanked by cellulose-binding motifs as a precursor for functional adhesives [110]. They previously explored the essential role of LLPS in the formation of liquid-like coacervates (LLCs) and the effects of kosmotropic ions on the formation of solid-like coacervates (SLCs) [111]. Kosmotropic ions are vital in transforming \(\alpha\)-helices or random coils into \(\beta\)-sheets, which are responsible for the strength of fiber. However, it is observed that a high concentration of strong kosmotropic ions (e.g., \({{{\rm{PO}}}}_{4}^{3-}\)) promotes the formation of SLCs. Their observations suggest that the low surface energy of LLCs is particularly promising for adhesive applications as it could allow efficient infiltration into porous structures and wet internal surfaces [110]. When added to cellulose fibrils, LLC droplets easily stuck to cellulose fibers and infiltrated the fiber network, even after drying.
Other studies point to potential applications based on microscopic phase-separated silk droplets themselves. Hendra et al. used fluorescently labeled microspheres of recombinant MaSp2, designated MS-2, that were produced via LLPS in the presence of potassium phosphate and heat solidified at 60 °C, in order to explore the light guiding properties of spider dragline silk [112]. The MS-2 microspheres demonstrated robust fluorescence emission, which proved critical for their role in energy transfer experiments.
In an interesting study, Wei et al. reported the formation of artificial membraneless organelles in Escherichia coli cells upon overexpression of recombinant spidroins consisting of repetitive regions of MaSp1 and MaSp2 (termed I16 and II16, respectively) [113]. The authors observed hallmarks of LLPS associated with a dynamic liquid state, as opposed to inclusion body formation. Notably, functionalized compartments could be created via targeted colocalization of cargo proteins, such as metallothionein, forming spatially sequestered nanoreactors that could produce selenium nanoparticles.
Future perspectives
Advancing our understanding of LLPS in silk spinning systems opens several avenues for future research. One critical area is elucidating the molecular grammar governing LLPS in different types of silk proteins and their interactions during phase separation. Investigating how silk proteins, along with associated factors such as SpiCE proteins [114], contribute to the material complexity of silk can provide deeper insights into silk assembly mechanisms. Exploring the phase separation mechanisms in more complex types of silk, such as pyriform (PySp), aciniform (AcSp), aggregate (AgSp), and tubuliform (TuSp) silks, is another promising direction. Given their complex sequences and structures, these silks may undergo distinct phase separation processes. Understanding these mechanisms could shed light on the evolutionary significance of conserved motifs in silk assembly and function. The effects of post-translational modifications (PTMs) [115], such as phosphorylation, hydroxylation, and dityrosine crosslinking, on LLPS also warrant further investigation. Furthermore, investigating the molecular mechanisms of fibrillation and the existence of pre-fibrillar complexes or oligomers in various silk types is crucial. Modern computational approaches, including machine learning and AI-based modeling, can offer powerful tools for identifying key sequence motifs, simulating phase behavior, and integrating experimental data to predict and model silk LLPS and assembly of higher-order structures. Understanding these processes can inform the development of synthetic silk materials that mimic the exceptional properties of natural silk. By addressing these research areas, we will move closer to realizing the potential of silk as a supermaterial with diverse applications.
References
Craig CL. Evolution of arthropod silks. Annu Rev Entomol. 1997;42:231–67.
Vierra C, Hsia Y, Gnesa E, Tang S, Jeffery F. Spider silk composites and applications. In: Cuppoletti J, editor. Metal, Ceramic and Polymeric Composites for Various Uses. Rijeka: IntechOpen; 2011.
Numata K. Biopolymer material and composite. In: Numata K, editor. Biopolymer Science for Proteins and Peptides. Elsevier; 2021. p. 205–46.
Malay AD, Craig HC, Chen J, Oktaviani NA, Numata K. Complexity of spider dragline silk. Biomacromolecules. 2022;23:1827–40.
Numata K, Kaplan DL. Silk proteins: designs from nature with multipurpose utility and infinite future possibilities. Adv Mater. 2024;37:2411256.
Allmeling C, Jokuszies A, Reimers K, Kall S, Vogt PM. Use of spider silk fibres as an innovative material in a biocompatible artificial nerve conduit. J Cell Mol Med. 2006;10:770–7.
Gosline JM, Denny MW, DeMont ME. Spider silk as rubber. Nature. 1984;309:551–2.
Wendt H, Hillmer A, Reimers K, Kuhbier JW, Schäfer-Nolte F, Allmeling C, et al. Artificial skin – culturing of different skin cell lines for generating an artificial skin substitute on cross-weaved spider silk fibres. PLoS One. 2011;6:e21833.
Yu H, Chen G, Li L, Wei G, Li Y, Xiong S, et al. Spider minor ampullate silk protein nanoparticles: an effective protein delivery system capable of enhancing systemic immune responses. MedComm. 2024;5:e573.
Hennecke K, Redeker J, Kuhbier JW, Strauss S, Allmeling C, Kasper C, et al. Bundles of spider silk, braided into sutures, resist basic cyclic tests: potential use for flexor tendon repair. PLoS One. 2013;8:e61100.
Numata K. How to define and study structural proteins as biopolymer materials. Polym J. 2020;52:1043–56.
Hijirida DH, Do KG, Michal C, Wong S, Zax D, Jelinski LW. 13C NMR of Nephila clavipes major ampullate silk gland. Biophys J. 1996;71:3442–7.
Askarieh G, Hedhammar M, Nordling K, Saenz A, Casals C, Rising A, et al. Self-assembly of spider silk proteins is controlled by a pH-sensitive relay. Nature. 2010;465:236–8.
Knight DP, Vollrath F. Changes in element composition along the spinning duct in a Nephila spider. Naturwissenschaften. 2001;88:179–82.
Brookstein O, Shimoni E, Eliaz D, Kaplan-Ashiri I, Carmel I, Shimanovich U. Metal ions guide the production of silkworm silk fibers. Nat Commun. 2024;15:6671.
Knight DP, Vollrath F. Liquid crystals and flow elongation in a spider’s silk production line. Proc R Soc Lond B Biol Sci. 1999;266:519–23.
Chen J, Tsuchida A, Malay AD, Tsuchiya K, Masunaga H, Tsuji Y, et al. Replicating shear-mediated self-assembly of spider silk through microfluidics. Nat Commun. 2024;15:527.
Rising A, Harrington MJ. Biological materials processing: time-tested tricks for sustainable fiber fabrication. Chem Rev. 2022;123:2155–99.
Jin H-J, Kaplan DL. Mechanism of silk processing in insects and spiders. Nature. 2003;424:1057–61.
Lin TY, Masunaga H, Sato R, Malay AD, Toyooka K, Hikima T, et al. Liquid crystalline granules align in a hierarchical structure to produce spider dragline microfibrils. Biomacromolecules. 2017;18:1350–5.
Parent LR, Onofrei D, Xu D, Stengel D, Roehling JD, Addison JB, et al. Hierarchical spidroin micellar nanoparticles as the fundamental precursors of spider silks. Proceedings Natl Acad Sci. 2018;115:11507–12.
Vollrath F, Knight DP. Liquid crystalline spinning of spider silk. Nature. 2001;410:541–8.
Vollrath F, Porter D. Silks as ancient models for modern polymers. Polymer. 2009;50:5623–32.
Landreh M, Osterholz H, Chen G, Knight SD, Rising A, Leppert A. Liquid-liquid crystalline phase separation of spider silk proteins. Commun Chem. 2024;7:260.
Moreno-Tortolero RO, Luo Y, Parmeggiani F, Skaer N, Walker R, Serpell LC, et al. Molecular organization of fibroin heavy chain and mechanism of fibre formation in Bombyx mori. Commun Biol. 2024;7:786.
Peran I, Mittag T. Molecular structure in biomolecular condensates. Curr Opin Struct Biol. 2020;60:17–26.
Harmon TS, Holehouse AS, Rosen MK, Pappu RV. Intrinsically disordered linkers determine the interplay between phase separation and gelation in multivalent proteins. Elife. 2017;6:e30294.
Patel A, Lee HO, Jawerth L, Maharana S, Jahnel M, Hein MY, et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell. 2015;162:1066–77.
Zhang H, Ji X, Li P, Liu C, Lou J, Wang Z, et al. Liquid-liquid phase separation in biology: mechanisms, physiological functions and human diseases. Sci China Life Sci. 2020;63:953–85.
Wegmann S, Eftekharzadeh B, Tepper K, Zoltowska KM, Bennett RE, Dujardin S, et al. Tau protein liquid–liquid phase separation can initiate tau aggregation. EMBO J. 2018;37:e98049.
Zbinden A, Pérez-Berlanga M, De Rossi P, Polymenidou M. Phase separation and neurodegenerative diseases: a disturbance in the force. Dev Cell. 2020;55:45–68.
Wang J, Choi JM, Holehouse AS, Lee HO, Zhang X, Jahnel M, et al. A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell. 2018;174:688–99.
Li P, Banjade S, Cheng HC, Kim S, Chen B, Guo L, et al. Phase transitions in the assembly of multivalent signalling proteins. Nature. 2012;483:336–40.
Iserman C, Roden CA, Boerneke MA, Sealfon RSG, McLaughlin GA, Jungreis I, et al. Genomic RNA elements drive phase separation of the SARS-CoV-2 nucleocapsid. Mol Cell. 2020;80:1078–91.
Chen H, Cui Y, Han X, Hu W, Sun M, Zhang Y, et al. Liquid–liquid phase separation by SARS-CoV-2 nucleocapsid protein and RNA. Cell Res. 2020;30:1143–5.
Alberti S, Gladfelter A, Mittag T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell. 2019;176:419–34.
Henninger JE, Oksuz O, Shrinivas K, Sagi I, LeRoy G, Zheng MM, et al. Cissé II, Chakraborty AK, Young RA. RNA-mediated feedback control of transcriptional condensates. Cell. 2021;184:207–25.
Banani SF, Lee HO, Hyman AA, Rosen MK. Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol. 2017;18:285–98.
Shin Y, Brangwynne CP. Liquid phase condensation in cell physiology and disease. Science. 1979;2017:357 eaaf4382.
Dignon GL, Best RB, Mittal J. Biomolecular phase separation: from molecular driving forces to macroscopic properties. Annu Rev Phys Chem. 2020;71:53–75.
Exler JH, Hümmerich D, Scheibel T. The amphiphilic properties of spider silks are important for spinning. Angewandte Chem Int Ed. 2007;46:3559–62.
Malay AD, Suzuki T, Katashima T, Kono N, Arakawa K, Numata K. Spider silk self-assembly via modular liquid-liquid phase separation and nanofibrillation. Sci Adv. 2020;6:eabb6030.
Zhou CZ, Confalonieri F, Medina N, Zivanovic Y, Esnault C, Yang T, et al. Fine organization of Bombyx mori fibroin heavy chain gene. Nucleic Acids Res. 2000;28:2413–9.
Sezutsu H, Yukuhiro K. Dynamic rearrangement within the Antheraea pernyi silk fibroin gene is associated with four types of repetitive units. J Mol Evol. 2000;51:329–38.
Kono N, Nakamura H, Ohtoshi R, Moran DAP, Shinohara A, Yoshida Y, et al. Orb-weaving spider Araneus ventricosus genome elucidates the spidroin gene catalogue. Sci Rep. 2019;9:8380.
Andersson M, Johansson J, Rising A. Silk spinning in silkworms and spiders. Int J Mol Sci. 2016;17:1290.
Termonia Y. Molecular modeling of spider silk elasticity. Macromolecules. 1994;27:7378–81.
Thiel BL, Guess KB, Viney C. Non-periodic lattice crystals in the hierarchical microstructure of spider (major ampullate) silk. Biopolymers. 1997;41:703–19.
Fazio V, Malay AD, Numata K, Pugno NM, Puglisi G. A physically-based machine learning approach inspires an analytical model for spider silk supercontraction. Adv Funct Mater. 2025;35:2420095.
Lefèvre T, Pézolet M. Unexpected β-sheets and molecular orientation in flagelliform spider silk as revealed by Raman spectromicroscopy. Soft Matter. 2012;8:6350–7.
Asakura T, Ohata T, Kametani S, Okushita K, Yazawa K, Nishiyama Y, et al. Intermolecular packing in B. mori silk fibroin: multinuclear NMR study of the model peptide (Ala-Gly)15 defines a heterogeneous antiparallel antipolar mode of assembly in the silk II form. Macromolecules. 2015;48:28–36.
Ozawa Y, Anbo H, Ota M, Fukuchi S. Classification of proteins inducing liquid-liquid phase separation: sequential, structural and functional characterization. J Biochem. 2023;173:255–64.
Hardenberg M, Horvath A, Ambrus V, Fuxreiter M, Vendruscolo M. Widespread occurrence of the droplet state of proteins in the human proteome. Proceedings Natl Acad Sci. 2020;117:33254–62.
Rauscher S, Baud S, Miao M, Keeley FW, Pomès R. Proline and glycine control protein self-organization into elastomeric or amyloid fibrils. Structure. 2006;14:1667–76.
Theillet FX, Kalmar L, Tompa P, Han KH, Selenko P, Dunker AK, et al. The alphabet of intrinsic disorder. Intrinsically Disord Proteins. 2013;1:e24360.
Muiznieks LD, Weiss AS, Keeley FW. Structural disorder and dynamics of elastin. Biochemistry Cell Biol. 2010;88:239–50.
Chou PY, Fasman GD. Conformational parameters for amino acids in helical, β-sheet, and random coil regions calculated from proteins. Biochemistry. 1974;13:211–22.
Johansson J, Rising A. Doing what spiders cannot—a road map to supreme artificial silk fibers. ACS Nano. 2021;15:1952–9.
Hayashi CY, Shipley NH, Lewis RV. Hypotheses that correlate the sequence, structure, and mechanical properties of spider silk proteins. Int J Biol Macromol. 1999;24:271–5.
Malay AD, Arakawa K, Numata K. Analysis of repetitive amino acid motifs reveals the essential features of spider dragline silk proteins. PLoS One. 2017;12:e0183397.
Malay AD, Oktaviani NA, Chen J, Numata K. Spider silk: Rapid, bottom-up self-assembly of MaSp1 into hierarchically structured fibers through biomimetic processing. Adv Funct Mater. 2024;35:2408175.
Sehnal F, Akai H. Insect silk glands: their types, development and function, and effects of environmental factors and morphogenetic hormones on them. Int J Insect Morphol Embryol. 1990;19:79–132.
Sonavane S, Westermark P, Rising A, Holm L. Regionalization of cell types in silk glands of Larinioides sclopetarius suggest that spider silk fibers are complex layered structures. Sci Rep. 2023;13:22273.
Rubinstein M, Semenov AN. Thermoreversible gelation in solutions of associating polymers. 2. linear dynamics. Macromolecules. 1998;31:1386–97.
Choi JM, Holehouse AS, Pappu RV. Physical principles underlying the complex biology of intracellular phase transitions. Annu Rev Biophys. 2020;49:107–33.
Martin EW, Holehouse AS, Peran I, Farag M, Incicco JJ, Bremer A, et al. Valence and patterning of aromatic residues determine the phase behavior of prion-like domains. Science. 2020;367:694–9.
Murthy AC, Dignon GL, Kan Y, Zerze GH, Parekh SH, Mittal J, et al. Molecular interactions underlying liquid−liquid phase separation of the FUS low-complexity domain. Nat Struct Mol Biol. 2019;26:637–48.
Kovoor J. Comparative structure and histochemistry of silk-producing organs in arachnids. In: Nentwig W, ed. Ecophysiology of Spiders. Berlin, Heidelberg: Springer Berlin Heidelberg; 1987. p. 160–86.
Sonavane S, Hassan S, Chatterjee U, Soler L, Holm L, Mollbrink A, et al. Origin, structure, and composition of the spider major ampullate silk fiber revealed by genomics, proteomics, and single-cell and spatial transcriptomics. Sci Adv. 2025;10:eadn0597.
Vollrath F, Knight DP, Hu XW. Silk production in a spider involves acid bath treatment. Proc R Soc Lond B Biol Sci. 1998;265:817–20.
Domigan LJ, Andersson M, Alberti KA, Chesler M, Xu Q, Johansson J, et al. Carbonic anhydrase generates a pH gradient in Bombyx mori silk glands. Insect Biochem Mol Biol. 2015;65:100–6.
Hagn F, Thamm C, Scheibel T, Kessler H. pH-dependent dimerization and salt-dependent stabilization of the N-terminal domain of spider dragline silk—implications for fiber formation. Angewandte Chem Int Ed. 2011;50:310–3.
He YX, Zhang NN, Li WF, Jia N, Chen BY, Zhou K, et al. N-Terminal domain of Bombyx mori fibroin mediates the assembly of silk in response to pH decrease. J Mol Biol. 2012;418:197–207.
De Oliveira DH, Gowda V, Sparrman T, Gustafsson L, Pires RS, Riekel C, et al. Structural conversion of the spidroin C-terminal domain during assembly of spider silk fibers. Nat Commun. 2024;15:4670.
Leclerc J, Lefèvre T, Gauthier M, Gagné SM, Auger M. Hydrodynamical properties of recombinant spider silk proteins: Effects of pH, salts and shear, and implications for the spinning process. Biopolymers. 2013;99:582–93.
Oktaviani NA, Matsugami A, Hayashi F, Numata K. Ion effects on the conformation and dynamics of repetitive domains of a spider silk protein: implications for solubility and β-sheet formation. Chem Commun. 2019;55:9761–4.
Rammensee S, Slotta U, Scheibel T, Bausch AR. Assembly mechanism of recombinant spider silk proteins. Proc Natl Acad Sci USA. 2008;105:6590–5.
Heiby JC, Rajab S, Rat C, Johnson CM, Neuweiler H. Conservation of folding and association within a family of spidroin N-terminal domains. Sci Rep. 2017;7:16789.
Otikovs M, Chen G, Nordling K, Landreh M, Meng Q, Jörnvall H, et al. Diversified structural basis of a conserved molecular mechanism for pH-dependent dimerization in spider silk N-terminal domains. Chembiochem. 2015;16:1720–4.
Hagn F, Eisoldt L, Hardy JG, Vendrely C, Coles M, Scheibel T, et al. A conserved spider silk domain acts as a molecular switch that controls fibre assembly. Nature. 2010;465:239–42.
Andersson M, Chen G, Otikovs M, Landreh M, Nordling K, Kronqvist N, et al. Carbonic anhydrase generates CO2 and H+ that drive spider silk formation via opposite effects on the terminal domains. PLoS Biol. 2014;12:e1001921.
Ittah S, Cohen S, Garty S, Cohn D, Gat U. An essential role for the C-terminal domain of a dragline spider silk protein in directing fiber formation. Biomacromolecules. 2006;7:1790–5.
Jaudzems K, Askarieh G, Landreh M, Nordling K, Hedhammar M, Jörnvall H, et al. pH-dependent dimerization of spider silk N-terminal domain requires relocation of a wedged tryptophan side chain. J Mol Biol. 2012;422:477–87.
Oktaviani NA, Malay AD, Matsugami A, Hayashi F, Numata K. Unusual pKa values mediate the self-assembly of spider dragline silk proteins. Biomacromolecules. 2023;24:1604–16.
Oktaviani NA, Malay AD, Goto M, Nagashima T, Hayashi F, Numata K. NMR assignment and dynamics of the dimeric form of soluble C-terminal domain major ampullate spidroin 2 from Latrodectus hesperus. Biomol NMR Assign. 2023;17:249–55.
Kronqvist N, Otikovs M, Chmyrov V, Chen G, Andersson M, Nordling K, et al. Sequential pH-driven dimerization and stabilization of the N-terminal domain enables rapid spider silk formation. Nat Commun. 2014;5:3254.
Sarr M, Kitoka K, Walsh-White KA, Kaldmäe M, Metlāns R, Tārs K, et al. The dimerization mechanism of the N-terminal domain of spider silk proteins is conserved despite extensive sequence divergence. Journal Biol Chem. 2022;298:101913.
Gao Z, Lin Z, Huang W, Lai CC. Fan Js, Yang D. Structural characterization of minor ampullate spidroin domains and their distinct roles in fibroin solubility and fiber formation. PLoS One. 2013;8:1–11.
Qi X, Wang H, Wang K, Wang Y, Leppert A, Iashchishyn I, et al. Spiders use structural conversion of globular amyloidogenic domains to make strong silk fibers. Adv Funct Mater. 2024;34:2470130.
Yang Y, Gao Z, Yang D. pH-dependent self-assembly mechanism of a single repetitive domain from a spider silk protein. Int J Biol Macromol. 2023;242:124775.
Hao Z, Long D, Zhang Y, Umuhoza D, Dai J, Xu Z, et al. New insight into the mechanism of in vivo fibroin self-assembly and secretion in the silkworm, Bombyx mori. Int J Biol Macromol. 2021;169:473–9.
Lefèvre T, Leclerc J, Rioux-Dubé J-F, Buffeteau T, Paquin MC, Rousseau M-E, et al. Conformation of spider silk proteins in situ in the intact major ampullate gland and in solution. Biomacromolecules. 2007;8:2342–4.
Oktaviani NA, Matsugami A, Malay AD, Hayashi F, Kaplan DL, Numata K. Conformation and dynamics of soluble repetitive domain elucidates the initial β-sheet formation of spider silk. Nat Commun. 2018;9:2121.
Otikovs M, Andersson M, Jia Q, Nordling K, Meng Q, Andreas LB, et al. Degree of biomimicry of artificial spider silk spinning assessed by NMR spectroscopy. Angew. Chem Int Ed. 2017;129:12745–9.
Polling S, Ormsby AR, Wood RJ, Lee K, Shoubridge C, Hughes JN, et al. Polyalanine expansions drive a shift into α-helical clusters without amyloid-fibril formation. Nat Struct Mol Biol. 2015;22:1008–15.
Xu D, Guo C, Holland GP. Probing the impact of acidification on spider silk assembly kinetics. Biomacromolecules. 2015;16:2072–9.
Gosline JM, Guerette PA, Ortlepp CS, Savage KN. The mechanical design of spider silks: from fibroin sequence to mechanical function. Journal Exp Biol. 1999;202:3295–303.
Jenkins JE, Sampath S, Butler E, Kim J, Henning RW, Holland GP, et al. Characterizing the secondary protein structure of black widow dragline silk using solid-state NMR and X-ray diffraction. Biomacromolecules. 2013;14:3472–83.
Holland GP, Creager MS, Jenkins JE, Lewis RV, Yarger JL. Determining secondary structure in spider dragline silk by carbon−carbon correlation solid-state NMR spectroscopy. J Am Chem Soc. 2008;130:9871–7.
Kümmerlen J, van Beek JD, Vollrath F, Meier BH. Local structure in spider dragline silk investigated by two-dimensional spin-diffusion nuclear magnetic resonance. Macromolecules. 1996;29:2920–8.
Asakura T. Structure of silk I (Bombyx mori silk fibroin before spinning) -type II β-turn, not α-helix. Molecules. 2021;26:3706.
Marsh RE, Corey RB, Pauling L. An investigation of the structure of silk fibroin. Biochim Biophys Acta. 1955;16:1–34.
Wang X, Li Y, Liu Q, Chen Q, Xia Q, Zhao P. In vivo effects of metal ions on conformation and mechanical performance of silkworm silks. Biochimica et Biophysica Acta (BBA) - Gen Subj. 2017;1861:567–76.
Yang S, Yu Y, Jo S, Lee Y, Son S, Lee KH. Calcium ion-triggered liquid-liquid phase separation of silk fibroin and spinning through acidification and shear stress. Nat Commun. 2024;15:1–15.
Leppert A, Chen G, Lama D, Sahin C, Railaite V, Shilkova O, et al. Liquid–liquid phase separation primes spider silk proteins for fiber formation via a conditional sticker domain. Nano Lett. 2023;23:5836–41.
Li J, Yang GZ, Li X, Tan HL, Wong ZW, Jiang S, et al. Nanoassembly of spider silk protein mediated by intrinsically disordered regions. Int J Biol Macromol. 2024;271:132438.
Koebley SR, Thorpe D, Pang P, Chrisochoides P, Greving I, Vollrath F, et al. Silk reconstitution disrupts fibroin self-assembly. Biomacromolecules. 2015;16:2796–804.
Zaki M, Rajkhowa R, Holland C, Razal JM, Hegh DY, Mota-Santiago P, et al. Recreating silk’s fibrillar nanostructure by spinning solubilized, undegummed silk. Adv Mater. 2025;37:2413786.
Laity PR, Baldwin E, Holland C. Changes in silk feedstock rheology during cocoon construction: the role of calcium and potassium ions. Macromol Biosci. 2019;19:1800188.
Mohammadi P, Beaune G, Stokke BT, Timonen JVI, Linder MB. Self-coacervation of a silk-like protein and its use as an adhesive for cellulosic materials. ACS Macro Lett. 2018;7:1120–5.
Mohammadi P, Aranko AS, Lemetti L, Cenev Z, Zhou Q, Virtanen S, et al. Phase transitions as intermediate steps in the formation of molecularly engineered protein fibers. Commun Biol. 2018;1:86.
Hendra, Yamagishi H, Heah WY, Malay AD, Numata K, Yamamoto Y. Micrometer‐scale optical web made of spider dragline fibers with optical gate operations. Adv Optical Mater. 2023;11:202202563.
Wei S-P, Qian Z-G, Hu C-F, Pan F, Chen M-T, Lee SY, et al. Formation and functionalization of membraneless compartments in Escherichia coli. Nat Chem Biol. 2020;16:1143–8.
Kono N, Nakamura H, Mori M, Yoshida Y, Ohtoshi R, Malay AD, et al. Multicomponent nature underlies the extraordinary mechanical properties of spider dragline silk. Proceedings Natl Acad Sci. 2021;118:e2107065118.
Craig HC, Malay AD, Hayashi F, Mori M, Arakawa K, Numata K. Posttranslational modifications in spider silk influence conformation and dimerization dynamics. MRS Bull. 2024;49:1192–204.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lay, M.G., Oktaviani, N.A., Malay, A.D. et al. Exploring the self-assembly of silk proteins through liquid-liquid phase separation. Polym J 57, 799–814 (2025). https://doi.org/10.1038/s41428-025-01040-w
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41428-025-01040-w
This article is cited by
-
Silk-based biomaterial scaffolds for dental tissue regeneration
Communications Materials (2026)