Abstract
In primates, mechanisms of both innate and adaptive immunity are well-studied in just a few species out of this very diverse taxonomic order, especially in relation to specific pathogens like SIV. Recent research has indicated there may be taxon-specific differences across primates in immune response strategies, including relative proportions of immune cell types and whether adaptive or innate responses are favored. It remains unclear which taxonomic level best explains variation in primate immune response strategies. Identifying this is important for understanding when and why these differences evolved. This review synthesizes major recent findings in primate immunology, to connect them to more generalized research on immune response strategies and present hypotheses for future research, focusing on major methodologies in the field. We demonstrate that gaining a better understanding of the evolution of primate immunity has far-reaching implications for our understanding of the evolutionary past of humans, and for present-day global health.
Similar content being viewed by others
Introduction
Cellular responses that recognize and neutralize invading pathogens are an important part of survival and maintaining homeostasis—this is the basic principle of innate immunity [1]. Gnathostomes, or jawed vertebrates, are the only known animals to also have evolved adaptive immunity, which uses immunoglobulins and T-cell receptors to launch an antigen-specific response [2]. Understanding the evolution of immunity within our branch of vertebrates, the taxonomic order Primates, is a critical context for understanding many aspects of human immunity, health, and disease. In non-human primates (hereafter primates), the mechanisms of both innate and adaptive immunity are relatively well-studied, especially in relation to specific pathogens like simian/human immunodeficiency virus (SIV/HIV) (see Glossary) [3,4,5,6,7,8]. The majority of what we know about primate immunity comes from a few primate species, mainly chimpanzees and macaques, which have been the subject of biomedical testing for decades. However, there are now almost 500 named primate species, with tremendous ecological, geographic, and phylogenetic diversity [9, 10]. In the last decade, due in large part to the development of novel methods to examine immune responses in a variety of primate species, we are beginning to generate a richer knowledge of the varied ways in which primates respond to pathogens and disease, and the factors that may influence those strategies. For example, recent research suggests that primates may differ widely in how their immune system responds to a threat, such as in the types and proportions of immune cells they produce and in how much they rely on adaptive versus innate immune responses [11,12,13,14,15,16,17]. Such findings highlight the need for a diverse approach to studying immunity across the primate taxa, because understanding this could shed light on when and why these differences evolved in the evolutionary past.
This review will synthesize major pathogen-specific findings in immunity across primates, in order to connect these findings to more generalized research on immune response strategies and present hypotheses for future research in this area. We begin with a short and accessible section on the molecular mechanisms of innate and adaptive immunity in primates. We then review important work in the study of primate immunology, focusing on major methodologies in the field. We consider these methods in three main groups: Variation in Cell-Surface Receptors, Primate Functional Genomics, and Characterizing Systemic Immune Responses Using Inflammatory Biomarkers. Each set of methods offer valuable insight into how primate immunity varies and segregates across taxonomic lines across different species. We conclude this review with a call for a more holistic approach to the study of immunology in nonhuman primates. We posit that merging our knowledge of life history and primate behavior with the growing knowledge of taxonomic differences in immune response will help guide primate evolutionary biology and immunology in new and exciting directions.
Innate and adaptive immunity in primates
We begin this review with a brief discussion of molecular mechanisms of innate and adaptive immunity in primates. Innate immunity recognizes and destroys invading pathogens by reading molecular patterns on non-host cells. It is the primary line of defense against invaders and begins almost immediately upon a pathogen entering the body [1, 18]. Neutrophils and macrophages are the most abundant innate effector cells, and natural killer lymphocytes also play an important role. Macrophages—large cells which engulf pathogens—utilize several cell surface receptors to identify foreign molecules. This includes toll-like receptors (TLRs), particularly important to understanding human evolution in the larger context of primate immunity because they have been under positive selection in our great ape lineage, which includes humans [11, 12, 19]. TLRs are cell-surface ligands that recognize pathogen associated molecular patterns (PAMPs), common markers of bacterial, viral, macroparasitic, and protozoan molecules [1]. Following this initial pattern recognition, macrophages release chemical signals to recruit other immune cells (Fig. 1).
Macrophages use TLRs to recognize pathogens and then recruit other immune cells by releasing cytokines and interferons. Created with BioRender.
Neutrophils, which represent over 50% of circulating immune cells in humans—a larger proportion than is found in other primates—are recruited to the site of infection by chemical signals from macrophages and infected endothelial cells lining the tissue [15, 20, 21]. Unlike macrophages, which live for several weeks, neutrophils are usually only functional for 1-2 days and work primarily within the blood circulation instead of within the tissue itself. Current research is aimed at understanding why humans produce higher levels of these short-lived, energetically expensive immune cells, even compared to other great apes [15, 21]. Neutrophils recognize and neutralize invaders by using cell-surface TLRs and then engulfing the foreign cell. In addition to engulfing pathogens, neutrophils can also cast extracellular nets to trap pathogens and produce reactive oxygen species, which are lethal to invaders [20].
Rather than simply reacting to a set of common foreign molecular patterns, the adaptive immune response—found only in jawed vertebrates—trains immune cells to target cell-surface molecules for each unique pathogen. This provides an advantage for the host when a pathogen may not display typical molecular patterns, or is otherwise evading the innate immune system. This training is a trade-off, because the adaptive response takes days, not minutes, to mount [18]. Nonetheless, this tailored response to unique cell-surface molecules, or antigens, has helped jawed vertebrates to inhabit a wide array of environments across the globe with high microbial variability. The main effector cells in primates are T-cells and B-cells, of which there are many subtypes. In general, T-cells are the cytotoxic effectors which kill pathogens and infected cells that display the antigen. B-cells differentiate into plasma cells, which secrete antibodies that bind to the antigen and kill the pathogen (Fig. 2). Both T- and B-cells are activated by antigen presenting cells, most commonly dendritic cells [2, 18].
This includes antigen presentation by activated dendritic cells, training cytotoxic T-cells, and release of antibodies by plasma cells. Created with BioRender.
Variation in cell-surface receptors
The study of immunology in primates is methodologically diverse and quickly changing, especially with respect to phylogenetic analyses for detecting natural selection on genes coding for a specific immune cell-surface receptor. Because this review is concerned with taxonomic differences among primates, we pay special attention to the relevance of these methods to broader phylogenetic understanding. Phylogenetic analysis provides valuable insight into when and in which common ancestor a genetic mutation occurred in the primate lineage. This becomes particularly relevant in the case of important cell-surface immune cell receptors, which are at the forefront of the host-pathogen interface. Variation in cell-surface molecules first became a focal point of primate immunology via research on SIV/HIV, because of the potential implications for human HIV treatment and prevention. We therefore discuss important examples of cell-surface receptor diversity in the context of this research, as well as expand beyond the context SIV/HIV.
More than 40 species of primates have documented species-specific types of SIV, which is known as HIV in humans [8]. These viruses attack and replicate within helper T-cells. In a select few primates, SIV can cause progression to the fatal acquired immunodeficiency syndrome (AIDS). Curiously, in most other primates, progression of SIV to AIDS is not observed, meaning that these primates appear to live with SIV infection without detrimental health effects [3, 6]. This has centered non-human primates as critically important for research on treating and curing HIV, creating one of the most long-standing research traditions within primate immunology. A decade into the HIV-1 group M pandemic, the identification of the SIV lineages most closely related to HIV-1 and HIV-2 were—and arguably remain today—some of the most seminal works in primate immunology [5, 7]. These studies primarily utilized proviral DNA, the genetic footprint of the virus trying to replicate in human cells. Today, studies of SIV viruses in primates instead directly use viral RNA from fecal samples to analyze the viral genome more directly. This has led to some important new discoveries. For example, the origin of HIV-1 group O was originally attributed to a strain of SIVcpz based on a proviral DNA phylogenetic analysis [5]. However, using fecal RNA, its origin is now attributed to SIVgor in western lowland gorillas, not chimpanzees (Fig 3) [8].
Most non-human primates infected with SIV, the primate version of HIV, do not develop AIDS, whereas humans almost always do. Researchers are still exploring why this is, and it is a complex question, as an AIDS-like illness has been documented in some wild chimpanzees of Gombe National Park, Tanzania [22, 23]. This means that tolerance of SIV infection is not universal among non-human primates, either. This further highlights the need to study a wide range of primate species to understand the diversity of immune defenses. For example, some species like chimpanzees and certain African monkeys may avoid AIDS because they retain more helper T-cells and show lower inflammation during chronic infection [3, 6]. Recent research suggests that greater genetic diversity in the CD4 protein on helper T-cells may help block the virus from entering these cells, keeping their viral burden low [24]. Another cell-surface receptor, Tripartite Motif-containing protein 5 (TRIM5), is a possible mechanism for why pandemic HIV-1 has spread to over 88 million humans [25, 26]. Studies of the pandemic strain HIV-1 (known as group M) in humans and other primate cell lines suggest that this strain of HIV-1 is better able to evade the anti-viral effects of TRIM5 than other strains [25]. The TRIM5 gene varies considerably among humans, other apes, and monkeys, with the neotropical and African and Asian monkey variants of the TRIM5 receptor providing the most potent protection against HIV-1 infection in helper T-cells [27].
The Major Histocompatibility Complex (MHC) represents another important location of variation among primate taxa—in fact, it is one of the most polymorphic gene families in existence among primates [28]. The MHC codes for cell-surface proteins that bind to fragments derived from pathogens, which are displayed on a cell’s surface to be recognized by T-cell receptors [29]. Here, we discuss MHC Class I and Class II, the most polymorphic of the MHC genes and directly involved in antigen presentation. While functionally important for the cytokine-driven inflammatory response, Class III molecules are largely conserved among primates, and even mammals, making them less relevant in the context of taxonomic variability [30]. MHC Class I molecules are found on the surface of most cells within the body and have roles in innate immunity, adaptive immunity, and placenta formation in reproduction; MHC Class II molecules are mainly found on the surface of antigen-presenting cells (dendritic cells, macrophages, and B-cells), performing mainly adaptive immune functions. MHC Class I is more rapidly evolving than Class II, likely because of its diverse functions [31]—as a result of this quick evolution, only great apes have strict orthologs of all of the polymorphic human MHC class I genes (known as Human Leukocyte Antigens HLA-A, HLA-B, and HLA-C) [31, 32]. These genes have dozens, sometimes even hundreds, of different alleles present in each species, meaning there is a lot of individual variability.
This variability has functional importance, as it makes a population resilient to more pathogens, since different MHC alleles have different binding specificities for peptide fragments from pathogens. Many HLA-A, -B, and -C alleles can be organized into cross-species clades made up of different ape orthologs. For example, HLA-B Clade 1 has HLA-B and the primate orthologs like chimpanzee (Pan troglodytes) Patr-B, bonobo (Pan paniscus) Papa-B, Gorilla (Gorilla gorilla) Gogo-B, etc. In particular, -B Clade 1 contains the HLA-B*57:01 allele and the chimpanzee Patr-B*06:03 allele, both functionally associated with control of the progression of HIV-1 and SIVcpz infection, respectively [33]—yet, Papa-B Clade 1 alleles have not been identified in bonobos, despite that this clade is predicted to have been present in the common ancestor of chimpanzees and bonobos [31]. This suggests it may have been lost in the bonobo lineage, or at least driven to very low frequency. Unlike chimpanzees and humans, bonobos don’t seem to have a SIV circulating in their populations [34], a potential contributing factor to the loss of this allele.
Though cell-surface receptor diversity first gained attention in the context of SIV/HIV, immune defenses among primates are shaped by diverse genes that influence how cells recognize and respond to a variety of other pathogen threats. Natural killer cells, part of the innate immune system, use specific receptors called Killer Immunoglobulin-like Receptors (KIR) to identify and avoid attacking their own cells. The genes that control these receptors are particularly diverse in monkeys and apes, suggesting they have evolved unique immune strategies compared to other mammals [16]. Similarly, primates have distinct versions of Interferon-induced Transmembrane proteins (IFITM) genes, which help block viruses from entering cells. All apes share a specific arrangement of IFITM genes, which likely emerged after the ape lineage split from other primates [35, 36]. These examples show that primates have evolved varied immune strategies, with unique genetic adaptations in monkeys and apes that may help explain differences in susceptibility to diseases. Understanding this diversity could provide insights into why certain species are more resistant to infections and improve our understanding of immune responses across the primate order.
As the first line of defense for identifying pathogens, the extracellular domains of toll-like receptors (TLRs) constantly interface with the cell-surface receptors of pathogens. Thus, it stands to reason there should be an evolutionary race between TLRs for improving pattern recognition (and the pathogen’s evasion of this recognition) [11, 12]. While several studies examining nucleotide substitution rates through evolutionary time in primates have detected positive selection on the majority of the TLR genes, TLR4 in particular consistently displays the strongest signature of selection across several studies with different sized datasets of primate species [11, 12, 19]. One study found the extracellular domain of the cell-surface protein TLR4 was under particularly strong positive selection in the lineage of all great apes, and again in the gorilla lineage after its split from other great apes [19]. Another similar study, which looked at a larger region of the TLR genes beyond just the extracellular domains, identified the strongest positive selection to be in TLR4, within the lineage shared by African and Asian monkeys and apes [12]. TLR4 is the toll-like receptor responsible for recognizing Trypanosoma protozoans and the molecular footprint of bacterial lipopolysaccharide (a protein found on Gram-negative bacteria) [11, 12, 19]. It is reasonable that the repeated interaction of a pathogen with a host primate’s TLRs would provide this variety of selective pressures, since each TLR is responsible for recognizing a different set of molecular patterns unique to each different pathogen type (e.g., bacteria, viruses, parasites, etc.). For example, the particularly strong positive selection on TLR4 may indicate that Gram-negative bacteria were especially important selective pressures in the African and Asian monkey and ape lineages [11, 12, 19, 37]. Functional genomics research has made evaluating such phylogenetic hypotheses possible, making it critical for studying the real-world implications of taxon-level variation among primates.
Primate functional genomics
Functional genomics links genes with their expression and downstream phenotypes. Fundamental techniques include RNA-seq and expression microarrays for characterizing genome-wide expression levels in in vitro tissues, at different time points and/or experimental conditions [38]. Additionally, the field is being advanced by new technologies like induced pluripotent stem cells and massively parallel reporter assays. Used in tandem with phylogenetic studies to map where functional changes occurred in evolutionary time can provide evidence regarding the effects of an evolved trait. In the context of primate immunity, this can help answer questions like whether there is more reliance on innate or adaptive immunity in each species, why the relative proportion of immune cells produced differs between species, or what contributes to different levels of systemic inflammation upon invasion by the same pathogen. Understanding what level of taxonomic organization accounts for the most variation could give clues to why these differences evolved in the evolutionary history of primates.
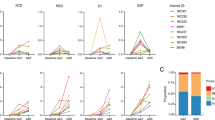
Recent work combining phylogenetic analyses and functional genomics has shed light onto the selective pressure posed by Gram-negative bacteria in the great ape lineage. Further, of relevance to human health specifically, this work includes a potential explanation for why humans in particular have an exceptionally strong immune response to stimulation with the Gram-negative bacterial protein lipopolysaccharide [39, 40]. Even small doses of lipopolysaccharide can induce powerful immune responses in humans and initiate sepsis, an extreme immune host response which leads to tissue damage and organ failure. However, African and Asian monkeys and other apes require doses of lipopolysaccharide that are an order of magnitude higher than humans to induce sepsis to a similar degree [39]. A recent study correlating primate body size to the strength of lipopolysaccharide response across nine primate species (spanning more than three orders of magnitude of body mass) suggests that body size may account for some of this variation. Larger-bodied organisms tended to have an exaggerated innate immune response to lipopolysaccharide stimulation. Specifically, this study measured transcriptional expression of innate immune genes using RNA-seq in response to this stimulation, and found an outsized increase in gene expression levels in the larger organisms [40]. RNA-seq profiles genome-wide expression levels using next-generation sequencing of transcribed RNA. However, with human and ape body mass represented with just one human and one Sumatran orangutan (Pongo abelii) [40], this study likely did not have sufficient power to detect gene expression differences that could account for why apes require doses of lipopolysaccharide an order of magnitude larger than humans to induce sepsis [39]. Thus, more work is needed to determine the relative contributions of body size, taxonomy (within similar body size categories, like humans versus other apes), and other factors affecting interspecies variability.
Tissue culture, a method which allows for the in vitro characterization of the functional impacts of genetic variation in actual primate cells, is a common technique for studying cellular responses in a safer manner than using live organisms. Using a multi-species gene expression microarray, which tracks expression levels of transcribed RNA across the genome, Barreiro and colleagues identified 793 genes that were highly expressed in tissues from human, chimpanzees, and rhesus macaques after stimulation in vitro with bacterial lipopolysaccharide molecules. Many of these genes were involved in the TLR4 pathogen recognition response [41]. Similarly, a study of the CEACAM3 receptor, which helps neutrophils recognize dangerous bacteria like Haemophilus aegyptius and Helicobacter pylori, identified different versions of the CEACAM3 gene in the lineage shared by apes and African and Asian monkeys, but found it to be absent entirely in neotropical monkeys [42]. As with TLR4 [19], they found the highest level of variability in the extracellular part of the CEACAM3 protein that directly interacts with pathogens to help immune cells recognize them [42]. This extracellular domain also shows statistical evidence of positive natural selection in the ancestor of apes and African and Asian monkeys. Crucially, Adrian and colleagues then studied the binding capabilities (to recognize H. pylori bacteria) of these different versions of the CEACAM3 protein, confirming these genetic variants do alter the ability of the different primate taxa to recognize bacterial pathogens. Such studies highlight critical variation in the genes involved with primate immune responses, and also represent a much-needed synthesis between genomic sequence phylogenies and the functional biological consequences of these sequence variations.
Hawash and colleagues characterized immune gene expression in response to both bacteria and viruses, using an in vitro study on innate immune cells from humans, chimpanzees (Pan troglodytes), olive baboons (Papio anubis), and rhesus macaques (Macaca mulatta) [15]. Their analysis centered around splitting these species in two phylogenetic categories: apes vs. African and Asian monkeys. In response to stimulation by both bacterial lipopolysaccharide and gardiquimod (mimics single-stranded RNA viruses), the transcriptional profiles from RNA-seq indicated three times more ligand-specific targeting of either the lipopolysaccharide or gardiquimod conditions by the African and Asian monkeys than by the apes. They found that apes essentially produce a stronger, but less pathogen-specific, immune response than the African and Asian monkeys [15]. Their methodology and phylogenetic framing of results is a promising avenue of future research. A broader sample of taxa would unite functional and phylogenetic approaches and shed light onto evolutionary reasons behind these differences in immune response strategies. Given that neotropical monkeys do not display comparable signatures of selection on TLR4 to apes and African and Asian Monkeys [11, 12, 19], it would be useful to compare their transcriptional profiles under the lipopolysaccharide condition, which could be informative of the selective pressure posed by Gram-negative bacteria in each lineage.
A significant barrier to expanding species representation is the difficulty of obtaining cell lines for use in functional studies. Induced pluripotent stem cells (iPSCs) offer a solution, as they can be differentiated into tissues that are typically difficult to collect from living primates, helping bridge this gap. iPSCs are also renewable, preserving access to these tissues long-term. This is especially valuable for functional studies involving apes, as tissue collection from chimpanzees is no longer permitted in the United States [38]. Other new methods, such as massively parallel reporter assays (MPRA), can simultaneously test how thousands of genome segments affect gene regulatory activity [43]. MPRAs can screen for enhancer function in primate cell lines (enhancers are DNA regions that increase the transcriptional output of target genes), providing a genome-wide picture of transcriptional regulation. This is important in an evolutionary context, as many functional studies primarily involve protein-coding genes. However, non-coding regulatory regions, which make up the majority of primate genomes, are also undoubtedly subject to selective pressures from pathogens and other evolutionary constraints [44].
Characterizing systemic immune responses using inflammatory biomarkers
There has been a recent push to understand how primate immunity varies across taxonomic lines at the systemic level (i.e., involving multiple immune cell types), beyond characterizing one single type of immune cell receptor. In addition to statistical phylogenetics or functional genomics from in vitro assays, it is important to study immunity in the full context of the body system in live primates. Sampling inflammatory biomarkers (both invasively and non-invasively) are key methods commonly used in studying differences in systemic immune responses among primates and humans, which we explore in this section of the review.
Immune senescence and increased inflammation, two key processes involved in human aging, are promising areas for broader phylogenetic comparisons of immune function. Most research, other than in humans, has occurred in rhesus macaques (Macaca mulatta) as representatives of African and Asian monkeys. Two rhesus macaque studies, one focused on immune senescence in innate immune cells like neutrophils, basophils, and monocytes [21], and the other on T-cells and monocytes [13], generally report that rhesus macaques are similar to humans in exhibiting immune senescence at old age [45]. For example, Asquith and colleagues note a similar trend of decreasing naive T-cells and increasing terminally differentiated T-cells with age [13]. Additionally, serum levels of pro-inflammatory Interleukin-1 receptor antagonist (IL-1Ra) cytokine were positively correlated with age in free-ranging female rhesus macaques at Cayo Santiago, Puerto Rico but showed no relationship with female reproductive status [46]. The similarity in aging patterns between macaques and humans may directly reflect evolutionary trends in immune senescence in haplorrhines, or could instead be a by-product of longer lifespans and other differing life history strategies between strepsirrhines and haplorrhines.
Expanding aging studies beyond macaques has already provided some insight into which other primate groups mirror aging in humans, and whether these differences result from lifespan alone. For example, little is known about strepsirrhine immune senescence with respect to innate immune cells and T-cells, highlighting the importance of expanding investigation to more species in this context. However, a few strepsirrhine species have been considered in other aging-related processes like inflammation. Unlike in humans and rhesus macaques, a study of age-related inflammation in captive sifakas (Propithecus coquereli) and ring-tailed lemurs (Lemur catta) found no evidence for age as a predictor of increased inflammatory biomarkers or oxidative damage, despite the different life history strategies of these two strepsirrhines (e.g., sifakas live longer and have later reproductive maturation than lemurs) [47]. This suggests that expanding strepsirrhine studies to include immune senescence of innate immune cells and T-cells would be very revealing of phylogenetic differences in immune senescence across primates, especially between strepsirrhines versus haplorrhines. Furthermore, a framework for studying the socio-environmental influences on aging in a wider sample of haplorrhines beyond humans and macaques has been recently proposed for a species of neotropical monkey, white-faced capuchins (Cebus imitator) of Santa Rosa National Park, Costa Rica, given their access to long-term longitudinal data on behavior, biomarkers, and environmental conditions [48]. Monitoring immune function in conjunction with aging and reproductive status could evaluate how life history strategies affect immune function, possibly informing the evolution of taxon-specific phenotypes in these neotropical monkeys and how they differ from apes and African and Asian monkeys or strepsirrhines.
Unfortunately, it is currently difficult both to compare taxon-specific differences in immune function and monitor them longitudinally in a particular species, especially in free-ranging populations located in remote areas. The majority of in vivo research has been conducted on primate populations suitable for invasive biological sample collection (e.g., blood draws) that require capture and anesthetization, which are stressful procedures that can be difficult to safely implement in wild populations, especially in arboreal species [49]. Recently, more research has utilized non-invasive markers of immune function to promote the inclusion of understudied species in immunological research, such as strepsirrhines and neotropical monkeys, and enable functional comparison across primate taxon. Three non-invasive markers of immune function, urinary neopterin, urinary suPAR, and fecal immunoglobulin A (IgA), have been successfully validated in primates and offer promising avenues for non-invasive immunological research in wild primate populations.
Neopterin, a biomarker excreted primarily from innate effector cells, such as macrophages [50], is currently the most widely utilized immune biomarker in primate field studies. Higham and colleagues validated neopterin as a reliable indicator of viral infection in primates and illustrated a strong correlation between serum and urinary neopterin values [51], paving the way for neopterin to be used in non-invasive research. Urinary neopterin has since been validated in a range of taxa (e.g., apes [52], African and Asian monkeys [53], and neotropical primates [54]) and is now considered a reliable marker of immunosenescence [53,54,55] and a possible indicator of habitat quality [52, 54] and environmental conditions like temperature [56]. Moreover, increases in neopterin levels were correlated with acute outbreaks of respiratory disease in wild chimpanzees (Pan troglodytes) [57] and bonobos (Pan paniscus) [58], while lower neopterin levels were associated with increased richness of gastrointestinal helminths in wild geladas (Theropithecus gelada) [59]. Neopterin levels have also been correlated with infection outcomes in SIV-infected mandrills (Mandrillus sphinx) [60]. Taken together, these findings demonstrate the utility of neopterin across studies of generalized immune activation and disease-specific research.
Urinary suPAR is a proinflammatory biomarker and can also be measured noninvasively in urine; however, it has been utilized in fewer studies and has only been validated for use in rhesus macaques. Urinary suPAR is the soluble form of the urokinase plasminogen activator receptor which is the binding site for the urokinase protein (uPAR), an enzyme found on both innate and adaptive effector cells that plays a crucial role in cell remodeling [61, 62]. Unlike neopterin, which showed minimal response to tissue damage, urinary suPAR was shown to reflect surgically inflicted tissue damage [61], likely making it a useful biomarker for non-invasive monitoring of wound severity and healing. However, urinary suPAR exhibited a weaker response to viral infection than neopterin [61] and did not reflect age-related changes in immune function [63]. Therefore, urinary neopterin and suPAR likely indicate different immune processes and could possibly be used to evaluate taxon-specific differences in immune function to distinct immune perturbations (e.g. viral infection versus wound healing).
Distinct from neopterin and suPAR, immunoglobulin A (IgA) is an antigen crucial to mucosal-based immunity that helps protect the epithelium from opportunistic pathogens. Fecal IgA, specifically, represents the amount of secretory IgA in the gastrointestinal tract and has been shown to reflect the function of the intestinal mucosal immune system [64, 65]. In contrast to urinary neopterin and suPAR, which measure systemic immune activation, fecal IgA is a promising biomarker to quantify immune function specific to the gastrointestinal tract which is constantly exposed to toxins and enteric pathogens including intestinal parasites. For example, higher IgA has been associated with lower whipworm parasite egg counts in wild baboons (Papio cynocephalus) [66]. Fecal IgA may also be an ideal biomarker to study the relationship between the gastrointestinal microbiome and host immunity to expand our understanding of how microbial communities assist in the development and maintenance of host immune function. Fecal IgA has primarily been used as a marker of stress in captive primates [67, 68] because chronically elevated cortisol levels can cause immunosuppression, decreasing fecal IgA. In wild chimpanzees, fecal IgA was shown to reflect the same seasonal patterns as respiratory illness with higher levels of fecal IgA during the late dry season, which is the period typically associated with higher rates of respiratory infection [69]. However, this study also noted that seasonal changes in nutritional stress and parasite infection could be correlated with IgA concentrations. In addition, Behringer and colleagues caution from their study of Barbary macaques (Macaca sylvanus) that the strong responsiveness of fecal IgA to Hypothalamic-Pituitary-Adrenal-axis activity may limit the interpretability of IgA as a measure of immunity [70]. Thus, more work is necessary to determine whether fecal IgA is a suitable biomarker of immune system function in primates. Work expanding species representation through IgA validation studies, as recently done in red fronted lemurs (Eulemur rufifrons) [71], will help characterize the immunological correlates of IgA in more primate species.
Our understanding of systemic immune function in primates will benefit greatly from integrative work on senescence, gene expression data, and measures of inflammatory biomarkers. In particular, non-invasive methods for characterizing biomarkers like suPAR, neopterin, and IgA are rapidly improving, and additional markers are continuing to be validated and deployed (Table 1). This is critical for expanding species representation in primate immunity research, as the ecological and geographic diversity of primates presents challenges for using invasive methods, particularly in free ranging populations. In the following section, we outline a framework for future research that is based on the current available data in hopes of generating advancements in this area.
Future perspectives
Our review illustrates that while sufficient gaps remain in our understanding of the evolution of primate immunity due to lack of data from many species in the primate order, existing data provide intriguing evidence of variation in immune response that broadly occurs along taxonomic designations (Fig. 4). We propose that taxonomic differences in immune response and senescence with age may be informed by life history differences.
This phylogenetic tree shows approximate evolutionary relationships between species, and colored stars each indicate a putative taxonomic difference [3, 12, 15, 16, 19, 39]. Branch lengths are not to exact scale and placement of stars along each branch do not reflect exact timing of that event. Created with BioRender.
For example, positive selection on the TLR4 extracellular domain which recognizes lipopolysaccharide on Gram-negative bacteria [19], and evidence of increased reliance on generalized innate immune strength over specificity [15], suggest there may have been a wide variety of bacterial pathogens present in the great ape ancestral environment as a selective pressure [11, 12, 15, 19, 41]. However, great ape habitat range overlaps with that of many African monkeys, suggesting that additional factors warrant consideration. Great apes exhibit markedly prolonged periods of offspring development compared to other primates, which would increase early-life exposure to pathogens, including bacteria. A stronger innate immune response during development—especially one that has not been subject to immune senescence, which is known to be a broadly similar process in both aging macaque monkeys and humans—would thus require evolutionary adaptations for immune strength and improved TLR4 recognition [13, 21, 72]. It is well-established that human children have a much stronger innate immune response and rely on this generalized approach to fighting pathogens, rather than the specificity of adaptive immunity [73]. Perhaps the prolonged childhood that occurs in great apes, in comparison to other primates, has selected for strong innate immunity and repeatedly selected for improved TLR4 recognition.
To test this hypothesis, comparable patterns of immune aging should be confirmed in great apes to facilitate a useful comparison to rhesus macaque studies, as well as studying a greater variety of African and Asian monkeys. For example, an interesting great ape to study in this context would be the orangutan. They have the longest developmental period of all great apes, but are the least social [74], potentially reducing exposure to certain pathogens with improved transmission in high-density populations. A much larger dataset with species diversity that includes both functional genomic and life history data on immune response strategies could help to explain these specializations among great apes, especially in light of potential differences in patterns of age-related inflammation seen in two species of strepsirrhines versus in humans and macaques [47].
Recent advances in computational methods for detecting selection may offer additional insights—for example, methods which specifically detect directional selection in regulatory regions, like transcription factor binding sites, can highlight evolutionary constraints that differ from those in protein coding regions [75]. And, phylogeny-aware models like Ornstein–Uhlenbeck could be potentially useful for characterizing the evolution of immunity-related traits through evolutionary time in primates, as they have recently been applied to account for genetic variation within species which can mimic stabilizing selection towards the optimum within the model [76, 77]. Furthermore, TLR genetic loci under positive selection should also be functionally characterized in greater detail [11, 12, 19], whereby identifying the consequences of mutations in these TLRs within a phylogenetic context will provide clues to which pathogens exerted the greatest selective pressure on primate ancestral lineages. For example, this could potentially help resolve outstanding questions about positive selection on the TLR4 gene regarding which gene regions are most important for Gram-negative bacteria recognition by innate immune cells. One study found the TLR4 extracellular domain to be under strong positive selection starting in the lineage of all great apes [19], whereas another similar study, which looked at more parts of the gene beyond the coding region for the extracellular domain, identified positive selection within the lineage shared by both African and Asian monkeys and apes [12]. Resolving such questions could reveal specific mechanisms underlying strong innate immunity in great apes, and how this may relate to life history characteristics.
Conclusion
Because humans are also great apes, the evolutionary history of our immune response strategies has enormous implications for treating human diseases. This has been an important consideration for many decades regarding HIV, but of course has broad potential for many other infectious and autoimmune diseases. There are now almost 500 named primate species, so the dearth of comprehensive data suitable for cross-species comparison is one of the largest obstacles facing the field, at present [9, 10]. Currently, it is not possible to speculate what level of taxonomic organization accounts for the most variation in immune response strategies in primates. However, as this review illustrates, novel research can be combined with existing knowledge regarding life history strategies among primates to guide new lines of inquiry. Obtaining data for all species across all areas of immunity research is obviously intractable. Throughout this review, we instead advocate for a practical approach to increasing comparative data across species, recommending increased species representation from certain phylogenetic lineages in specific areas of research that are most informative of phylogenetic.
Data availability
This review did not generate original data, code, or reagents.
References
Buchmann K. Evolution of Innate Immunity: Clues from Invertebrates via Fish to Mammals. Front Immunol. 2014;5:459. https://doi.org/10.3389/fimmu.2014.00459.
Litman GW, Rast JP, Fugmann SD. The origins of vertebrate adaptive immunity. Nat Rev Immunol. 2010;10:543–53. https://doi.org/10.1038/nri2807.
Pandrea I, Apetrei C. Where the Wild Things Are: Pathogenesis of SIV Infection in African Nonhuman Primate Hosts. Curr HIV-AIDS Rep. 2010;7:28–36. https://doi.org/10.1007/s11904-009-0034-8.
Compton AA, Malik HS, Emerman M. Host gene evolution traces the evolutionary history of ancient primate lentiviruses. Philosophical Trans R Soc B Biol Sci. 2013;368:20120496. https://doi.org/10.1098/rstb.2012.0496.
Gao F, Bailes E, Robertson DL, Chen Y, Rodenburg CM, Michael SF, et al. Origin of HIV-1 in the chimpanzee Pan troglodytes troglodytes. Nature. 1999;397:436–41. https://doi.org/10.1038/17130.
Liovat A-S, Jacquelin B, Ploquin M, Barre-Sinoussi F, Muller-Trutwin M. African Non Human Primates Infected by SIV - Why Dont they Get Sick? Lessons from Studies on the Early Phase of Non-Pathogenic SIV Infection. Curr Hiv Res. 2009;7:39–50. https://doi.org/10.2174/157016209787048546.
Hirsch VM, Olmsted RA, Murphey-Corb M, Purcell RH, Johnson PR. An African primate lentivirus (SIVsm) closely related to HIV-2. Nature. 1989;339:389–92. https://doi.org/10.1038/339389a0.
D’arc M, Ayouba A, Esteban A, Learn GH, Boué V, Liegeois F, et al. Origin of the HIV-1 group O epidemic in western lowland gorillas. Proc Natl Acad Sci. 2015;112:E1343–52. https://doi.org/10.1073/pnas.1502022112.
Rylands AB, Mittermeier RA. Primate taxonomy: Species and conservation. Evol Anthropol Issues N. Rev. 2014;23:8–10. https://doi.org/10.1002/evan.21387.
Fleagle JG, Seiffert ER. Chapter 20 - The Phylogeny of Primates. In Evolutionary Neuroscience (Second Edition) (ed JH Kaas). 2020, pp. 483–518. Elsevier Academic Press. https://doi.org/10.1016/b978-0-12-820584-6.00020-9
Wlasiuk G, Khan S, Switzer WM, Nachman MW. A History of Recurrent Positive Selection at the Toll-Like Receptor 5 in Primates. Mol Biol Evol. 2009;26:937–49. https://doi.org/10.1093/molbev/msp018.
Wlasiuk G, Nachman MW. Adaptation and Constraint at Toll-Like Receptors in Primates. Mol Biol Evol. 2010;27:2172–86. https://doi.org/10.1093/molbev/msq104.
Asquith M, Haberthur K, Brown M, Engelmann F, Murphy A, Al-Mahdi Z, et al. Age-dependent changes in innate immune phenotype and function in rhesus macaques (Macaca mulatta). Pathobiol Aging Age-Relat Dis. 2012;2:18052. https://doi.org/10.3402/pba.v2i0.18052.
Gombart AF, Saito T, Koeffler HP. Exaptation of an ancient Alu short interspersed element provides a highly conserved vitamin D-mediated innate immune response in humans and primates. BMC Genom. 2009;10:321. https://doi.org/10.1186/1471-2164-10-321.
Hawash MBF, Sanz-Remón J, Grenier J-C, Kohn J, Yotova V, Johnson Z, et al. Primate innate immune responses to bacterial and viral pathogens reveals an evolutionary trade-off between strength and specificity. Proc Natl Acad Sci USA. 2021;118:e2015855118. https://doi.org/10.1073/pnas.2015855118.
Parham P, Abi-Rached L, Matevosyan L, Moesta AK, Norman PJ, Aguilar AMO, et al. Primate-specific regulation of natural killer cells. J Med Primatol. 2010;39:194–212. https://doi.org/10.1111/j.1600-0684.2010.00432.x.
Lagishetty V, Liu NQ, Hewison M. Vitamin D metabolism and innate immunity. Mol Cell Endocrinol. 2011;347:97–105. https://doi.org/10.1016/j.mce.2011.04.015.
Iwasaki A, Medzhitov R. Control of adaptive immunity by the innate immune system. Nat Immunol. 2015;16:343–53. https://doi.org/10.1038/ni.3123.
Nakajima T, Ohtani H, Satta Y, Uno Y, Akari H, Ishida T, et al. Natural selection in the TLR-related genes in the course of primate evolution. Immunogenetics. 2008;60:727–35. https://doi.org/10.1007/s00251-008-0332-0.
Mayadas TN, Cullere X, Lowell CA. The Multifaceted Functions of Neutrophils. Annu Rev Pathol Mech Dis. 2014;9:181–218. https://doi.org/10.1146/annurev-pathol-020712-164023.
He Z, Allers C, Sugimoto C, Ahmed N, Fujioka H, Kim W-K, et al. Rapid Turnover and High Production Rate of Myeloid Cells in Adult Rhesus Macaques with Compensations during Aging. J Immunol. 2018;200:4059–67. https://doi.org/10.4049/jimmunol.1800207.
Terio KA, Kinsel MJ, Raphael J, Mlengeya T, Lipende I, Kirchhoff CA, et al. Pathologic Lesions in Chimpanzees (Pan trogylodytes schweinfurthii) from Gombe National Park, Tanzania, 2004-2010. J Zoo Wildl Med. 2011;42:597–607. https://doi.org/10.1638/2010-0237.1.
Keele BF, Jones JH, Terio KA, Estes JD, Rudicell RS, Wilson ML, et al. Increased mortality and AIDS-like immunopathology in wild chimpanzees infected with SIVcpz. Nature. 2009;460:515–9. https://doi.org/10.1038/nature08200.
Russell RM, Bibollet-Ruche F, Liu W, Sherrill-Mix S, Li Y, Connell J, et al. CD4 receptor diversity represents an ancient protection mechanism against primate lentiviruses. Proc Natl Acad Sci. 2021;118:e2025914118. https://doi.org/10.1073/pnas.2025914118.
Zuliani-Alvarez L, Govasli ML, Rasaiyaah J, Monit C, Perry SO, Sumner RP, et al. Evasion of cGAS and TRIM5 defines pandemic HIV. Nat Microbiol. 2022;7:1762–76. https://doi.org/10.1038/s41564-022-01247-0.
Qiu S, Liu H, Jian Z, Fan Z, Liu S, Xing J, et al. Characterization of the primate TRIM gene family reveals the recent evolution in primates. Mol Genet Genomics. 2020;295:1281–94. https://doi.org/10.1007/s00438-020-01698-2.
Song B, Javanbakht H, Perron M, Park DH, Stremlau M, Sodroski J. Ret.rovirus Restriction by TRIM5α Variants from Old World and New World Primates. J Virol. 2005;79:3930–7. https://doi.org/10.1128/jvi.79.7.3930-3937.2005.
Heijmans CMC, Groot NG, de, Bontrop RE. Comparative genetics of the major histocompatibility complex in humans and nonhuman primates. Int J Immunogenet. 2020;47:243–60. https://doi.org/10.1111/iji.12490.
Fortier AL, Pritchard JK. 2025 The Primate major histocompatibility complex: an illustrative example of gene family evolution. https://doi.org/10.7554/elife.103545.1.
Deakin JE, Papenfuss AT, Belov K, Cross JG, Coggill P, Palmer S, et al. Evolution and comparative analysis of the MHC Class III inflammatory region. BMC Genom. 2006;7:281. https://doi.org/10.1186/1471-2164-7-281.
Wroblewski EE, Guethlein LA, Norman PJ, Li Y, Shaw CM, Han AS, et al. Bonobos Maintain Immune System Diversity with Three Functional Types of MHC-B. J Immunol. 2017;198:3480–93. https://doi.org/10.4049/jimmunol.1601955.
Adams EJ, Parham P. Species-specific evolution of MHC class I genes in the higher primates. Immunol Rev. 2001;183:41–64. https://doi.org/10.1034/j.1600-065x.2001.1830104.x.
Groot, de NG, Heijmans, Zoet CMC, Ru YM, de AH, et al. AIDS-protective HLA-B*27/B*57 and chimpanzee MHC class I molecules target analogous conserved areas of HIV-1/SIVcpz. Proc Natl Acad Sci. 2010;107:15175–80. https://doi.org/10.1073/pnas.1009136107.
Li Y, Ndjango J-B, Learn GH, Ramirez MA, Keele BF, Bibollet-Ruche F, et al. Eastern Chimpanzees, but Not Bonobos, Represent a Simian Immunodeficiency Virus Reservoir. J Virol. 2012;86:10776–91. https://doi.org/10.1128/jvi.01498-12.
Schelle L, Abrantes J, Baldauf H-M, Esteves PJ. Evolution of primate interferon-induced transmembrane proteins (IFITMs): a story of gain and loss with a differentiation into a canonical cluster and IFITM retrogenes. Front Microbiol. 2023;14:1213685. https://doi.org/10.3389/fmicb.2023.1213685.
Liao Y, Goraya MU, Yuan X, Zhang B, Chiu S-H, Chen J-L. Functional Involvement of Interferon-Inducible Transmembrane Proteins in Antiviral Immunity. Front Microbiol. 2019;10:1097. https://doi.org/10.3389/fmicb.2019.01097.
Kumar H, Kawai T, Akira S. Pathogen recognition in the innate immune response. Biochem J. 2009;420:1–16. https://doi.org/10.1042/bj20090272.
Housman G, Gilad Y. Prime time for primate functional genomics. Curr Opin Genet Dev. 2020;62:1–7. https://doi.org/10.1016/j.gde.2020.04.007.
Brinkworth JF, Valizadegan N. Sepsis and the evolution of human increased sensitivity to lipopolysaccharide. Evol Anthr Issues N Rev. 2021;30:141–57. https://doi.org/10.1002/evan.21887.
McMinds R, Jiang RHY, Adapa SR, Ruhs EC, Munds RA, Leiding JW, et al. Bacterial sepsis triggers stronger transcriptomic immune responses in larger primates. Proc B. 2024;291:20240535. https://doi.org/10.1098/rspb.2024.0535.
Barreiro LB, Marioni JC, Blekhman R, Stephens M, Gilad Y. Functional Comparison of Innate Immune Signaling Pathways in Primates. PLoS Genet. 2010;6:e1001249. https://doi.org/10.1371/journal.pgen.1001249.
Adrian J, Bonsignore P, Hammer S, Frickey T, Hauck CR. Adaptation to Host-Specific Bacterial Pathogens Drives Rapid Evolution of a Human Innate Immune Receptor. Curr Biol. 2019;29:616–630.e5. https://doi.org/10.1016/j.cub.2019.01.058.
Klein JC, Agarwal V, Inoue F, Keith A, Martin B, Kircher M, et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays. Nat Methods. 2020;17:1083–91. https://doi.org/10.1038/s41592-020-0965-y.
Romero IG, Lea AJ. Leveraging massively parallel reporter assays for evolutionary questions. Genome Biol. 2023;24:26. https://doi.org/10.1186/s13059-023-02856-6.
Weiskopf D, Weinberger B, Grubeck-Loebenstein B. The aging of the immune system. Transpl Int. 2009;22:1041–50. https://doi.org/10.1111/j.1432-2277.2009.00927.x.
Hoffman CL, Higham JP, Heistermann M, Coe CL, Prendergast BJ, Maestripieri D. Immune function and HPA axis activity in free-ranging rhesus macaques. Physiol Behav. 2011;104:507–14. https://doi.org/10.1016/j.physbeh.2011.05.021.
Guevara EE, Grebe NM, Lawler RR, Crowley A, Lo S, Paietta EN, et al. Comparison of age-related inflammation and oxidative stress in two lemur species. J Comp Physiol B 2025;1–12. https://doi.org/10.1007/s00360-025-01619-y
Campos FA, Wikberg EC, Orkin JD, Park Y, Snyder-Mackler N, Hernandez SC, et al. Wild capuchin monkeys as a model system for investigating the social and ecological determinants of ageing. Philos Trans B. 2024;379:20230482. https://doi.org/10.1098/rstb.2023.0482.
Crofoot MC, Norton TM, Lessnau RG, Viner TC, Chen TC, Mazzaro LM, et al. Field Anesthesia and Health Assessment of Free-ranging Cebus capucinus in Panama. Int J Primatol. 2009;30:125. https://doi.org/10.1007/s10764-009-9333-6.
Murr C, Widner B, Wirleitner B, Fuchs D. Neopterin as a marker for immune system activation. Curr Drug Metab. 2002;3:175–87. https://doi.org/10.2174/1389200024605082.
Higham JP, Kraus C, Stahl-Hennig C, Engelhardt A, Fuchs D, Heistermann M. Evaluating noninvasive markers of nonhuman primate immune activation and inflammation. Am J Phys Anthropol. 2015;158:673–84. https://doi.org/10.1002/ajpa.22821.
Behringer V, Stevens JMG, Leendertz FH, Hohmann G, Deschner T. Validation of a Method for the Assessment of Urinary Neopterin Levels to Monitor Health Status in Non-human-primate Species. Front Physiol. 2017;8:51. https://doi.org/10.3389/fphys.2017.00051.
Müller N, Heistermann M, Strube C, Schülke O, Ostner J. Age, but not anthelmintic treatment, is associated with urinary neopterin levels in semi-free ranging Barbary macaques. Sci Rep. 2017;7:41973. https://doi.org/10.1038/srep41973.
Lucore JM, Marshall AJ, Brosnan SF, Benítez ME. Validating Urinary Neopterin as a Biomarker of Immune Response in Captive and Wild Capuchin Monkeys. Front Vet Sci. 2022;9:918036. https://doi.org/10.3389/fvets.2022.918036.
Negrey JD, Behringer V, Langergraber KE, Deschner T. Urinary neopterin of wild chimpanzees indicates that cell-mediated immune activity varies by age, sex, and female reproductive status. Sci Rep. 2021;11:9298. https://doi.org/10.1038/s41598-021-88401-6.
Lucore JM, Beehner JC, White AF, Sinclair LF, Martins VA, Kovalaskas SA, et al. High temperatures are associated with decreased immune system performance in a wild primate. Sci Adv. 2024;10:eadq6629. https://doi.org/10.1126/sciadv.adq6629.
Wu DF, Behringer V, Wittig RM, Leendertz FH, Deschner T. Urinary neopterin levels increase and predict survival during a respiratory outbreak in wild chimpanzees (Taï National Park, Côte d’Ivoire). Sci Rep. 2018;8:13346. https://doi.org/10.1038/s41598-018-31563-7.
Kreyer M, Behringer V, Deimel C, Fruth B. Neopterin Levels in Bonobos Vary Seasonally and Reflect Symptomatic Respiratory Infections. Ecohealth. 2023;20:93–104. https://doi.org/10.1007/s10393-023-01633-y.
Schneider-Crease IA, Feder JA, Baniel A, McCann C, Haile AA, Abebe B, et al. Urinary neopterin reflects immunological variation associated with age, helminth parasitism, and the microbiome in a wild primate. Sci Rep. 2022;12:21307. https://doi.org/10.1038/s41598-022-25298-9.
Dibakou SE, Souza A, Boundenga L, Givalois L, Mercier-Delarue S, Simon F, et al. Ecological, parasitological and individual determinants of plasma neopterin levels in a natural mandrill population. Int J Parasitol Parasites Wildl. 2020;11:198–206. https://doi.org/10.1016/j.ijppaw.2020.02.009.
Higham JP, Stahl-Hennig C, Heistermann M. Urinary suPAR: a non-invasive biomarker of infection and tissue inflammation for use in studies of large free-ranging mammals. Roy Soc Open Sci. 2020;7:191825. https://doi.org/10.1098/rsos.191825.
Thunø M, Macho B, Eugen-Olsen J. suPAR: The Molecular Crystal Ball. Dis Markers. 2009;27:157–72. https://doi.org/10.1155/2009/504294.
Cooper EB, Watowich MM, Beeby N, Whalen C, Unit CBR, Montague MJ, et al. Concentrations of urinary neopterin, but not suPAR, positively correlate with age in rhesus macaques. Front Ecol Evol. 2022;10:1007052. https://doi.org/10.3389/fevo.2022.1007052.
Corthésy B. Multi-Faceted Functions of Secretory IgA at Mucosal Surfaces. Front Immunol. 2013;4:185 https://doi.org/10.3389/fimmu.2013.00185.
Mantis NJ, Rol N, Corthésy B. Secretory IgA’s complex roles in immunity and mucosal homeostasis in the gut. Mucosal Immunol. 2011;4:603–11. https://doi.org/10.1038/mi.2011.41.
Gesquiere LR, Habig B, Hansen C, Li A, Freid K, Learn NH, et al. Noninvasive measurement of mucosal immunity in a free-ranging baboon population. Am J Primatol. 2020;82:e23093. https://doi.org/10.1002/ajp.23093.
Huang S, Li L, Wu J, Li C, Bai J, Sun Y, et al. Seasonal variations in immunoreactive cortisol and fecal immunoglobulin levels in Sichuan golden monkey (Rhinopithecus roxellana). Turk J Zool. 2014;38:642–50. https://doi.org/10.3906/zoo-1309-39.
Paramastri Y, Royo F, Eberova J, Carlsson H-E, Sajuthi D, Fernstrom A-L, et al. Urinary and fecal immunoglobulin A, cortisol and 11-17 dioxoandrostanes, and serum cortisol in metabolic cage housed female cynomolgus monkeys (Macaca fascicularis). J Med Primatol. 2007;36:355–64. https://doi.org/10.1111/j.1600-0684.2007.00229.x.
Lantz EL, Lonsdorf EV, Heintz MR, Murray CM, Lipende I, Travis DA, et al. Non-invasive quantification of immunoglobulin A in chimpanzees (Pan troglodytes schweinfurthii) at Gombe National Park, Tanzania: Immunoglobulin A in Chimpanzees at Gombe. Am J Primatol. 2016;80:e22558. https://doi.org/10.1002/ajp.22558.
Behringer V, Müller-Klein N, Strube C, Schülke O, Heistermann M, Ostner J. Responsiveness of fecal immunoglobulin A to HPA-axis activation limits its use for mucosal immunity assessment. Am J Primatol. 2021;83:e23329. https://doi.org/10.1002/ajp.23329.
Pethig L, Behringer V, Kappeler PM, Fichtel C, Heistermann M. Establishment and Validation of Fecal Secretory Immunoglobulin A Measurement for Intestinal Mucosal Health Assessment in Wild Lemurs. Am J Primatol. 2024;86:e23694. https://doi.org/10.1002/ajp.23694.
Cinco IR, Napier EG, Rhoades NS, Davies MH, Allison DB, Kohama SG, et al. Immunological and microbial shifts in the aging rhesus macaque lung during nontuberculous mycobacterial infection. mBio. 2024;15:e00829–24. https://doi.org/10.1128/mbio.00829-24.
Simon AK, Hollander GA, McMichael A. Evolution of the immune system in humans from infancy to old age. Proc R Soc B Biol Sci. 2015;282:20143085. https://doi.org/10.1098/rspb.2014.3085.
Noordwijk van MA, Atmoko SSU, Knott CD, Kuze N, Morrogh-Bernard HC, Oram F, et al. The slow ape: High infant survival and long interbirth intervals in wild orangutans. J Hum Evol. 2018;125:38–49. https://doi.org/10.1016/j.jhevol.2018.09.004.
Andrews, Fan G, Pratt K, Phalke HE, Zoonomia Consortium N, Karlsson EK, et al. Mammalian evolution of human cis-regulatory elements and transcription factor binding sites. Science. 2023;380:eabn7930. https://doi.org/10.1126/science.abn7930.
Bartoszek K, Glémin S, Kaj I, Lascoux M. Using the Ornstein–Uhlenbeck process to model the evolution of interacting populations. J Theor Biol. 2017;429:35–45. https://doi.org/10.1016/j.jtbi.2017.06.011.
Rohlfs RV, Harrigan P, Nielsen R. Modeling Gene Expression Evolution with an Extended Ornstein–Uhlenbeck Process Accounting for Within-Species Variation. Mol Biol Evol. 2014;31:201–11. https://doi.org/10.1093/molbev/mst190.
Author information
Authors and Affiliations
Contributions
MEB and SKJ conceptualized the project. SKJ wrote the initial manuscript, and JML, JL, EVL, and MEB provided extensive feedback and additional contributions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Joseph, S.K., Lucore, J.M., Lindo, J. et al. Taxonomic variation in immune response strategies among primates. Genes Immun 26, 539–548 (2025). https://doi.org/10.1038/s41435-025-00363-1
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41435-025-00363-1