Abstract
Cryptic fungal pathogens pose disease management challenges due to their morphological resemblance to known pathogens. Here, we investigated the genomes and phenotypes of 53 globally distributed isolates of Aspergillus section Nidulantes fungi and found 30 clinical isolates—including four isolated from COVID-19 patients—were A. latus, a cryptic pathogen that originated via allodiploid hybridization. Notably, all A. latus isolates were misidentified. A. latus hybrids likely originated via a single hybridization event during the Miocene and harbor substantial genetic diversity. Transcriptome profiling of a clinical isolate revealed that both parental subgenomes are actively expressed and respond to environmental stimuli. Characterizing infection-relevant traits—such as drug resistance and growth under oxidative stress—revealed distinct phenotypic profiles among A. latus hybrids compared to parental and closely related species. Moreover, we identified four features that could aid A. latus taxonomic identification. Together, these findings deepen our understanding of the origin of cryptic pathogens.
Similar content being viewed by others
Introduction
More than 150 million cases of severe fungal infections result in an estimated 1.7 million deaths per year worldwide1. Beyond the profound toll of lives lost, fungal pathogens pose a substantial economic burden conservatively estimated to be 11.5 billion US$ but may be as high as 48 billion US$2,3,4. The impact of fungal pathogens on human welfare will likely increase due to global climate change5. The importance of fungal diseases was recently acknowledged by the establishment of the first-ever priority pathogen list by the World Health Organization6, and it has risen in prominence as a prevalent cause of co-infection of COVID-19 patients7,8,9.
Even though a great deal is known about several aspects of fungal pathogenicity, we know surprisingly little about the repeated evolution of pathogenicity in fungi, such as the traits and genetic elements that contributed to it10. Evolutionary approaches have been highly informative both in understanding how pathogens differ from their non-pathogenic close relatives11,12,13,14 but also in discovering cryptic species, defined as species that are genetically distinct from known species but difficult to distinguish using morphology15,16. Cryptic pathogens have been identified in several fungal genera that contain pathogens, including Cryptococcus, Aspergillus, Histoplasma, and Fusarium13,15,17,18,19,20,21.
Recently, we used short-read genome sequencing data from a sample of six clinical isolates to show that Aspergillus latus is a novel cryptic fungal pathogen21. A. latus arose via allodiploid hybridization between A. spinulosporus and a close, unknown relative of A. quadrilineatus, wherein (nearly) the entire genomes of both parents were combined during hybridization. However, the small number of isolates examined, and their highly fragmented short-read genome assemblies left numerous questions unanswered—such as how the genome of the hybrids differs from the parental genomes, the timing and number of hybridization events, and genetic diversity and phenotypic variation in the species. Increased understanding of A. latus genomic and phenotypic trait variation and differences from close relatives could increase accuracy of taxonomic identification of the species and, in the longer term, aid the development of clinical diagnostic techniques.
To further understand the biology and evolutionary origins of the cryptic fungal pathogen A. latus, we generated high-quality genome assemblies using long- and/or short-read sequencing technologies on a substantially expanded set of 53 globally distributed isolates (50 clinical isolates; three type strains) from section Nidulantes of the genus Aspergillus. We found that 30 clinical isolates correspond to A. latus and the remaining 20 clinical isolates to A. spinulosporus (8), A. quadrilineatus (1), or A. nidulans (11); one previously published strain of A. latus was included in this dataset21. A. latus likely arose approximately 13 million years ago and exhibits substantial genetic diversity among gene families and biosynthetic gene clusters. Transcriptomic analysis of a clinical isolate revealed that both parental subgenomes are actively expressed and respond to environmental perturbations. Profiling of each species across infection-relevant chemical and physiological traits suggests hybridization contributes to a unique phenotypic profile. Comprehensive genomic and phenotypic characterization of A. latus identified four traits that distinguish A. latus from closely related species. Characterization of A. latus, an understudied cryptic fungal pathogen, expands our understanding of the origins of fungal pathogenicity and is a key step toward preventing and combating disease.
Results
Genome sequencing and species identification of 50 clinical isolates
The genomes of three type strains (A. latus NRRL 200 T; A. spinulosporus NRRL 2395 T; A. quadrilineatus NRRL 201 T) and 40 clinical isolates (Supplementary Data 1) were sequenced and assembled using both long- (Oxford Nanopore) and short-read (Illumina) technologies. The resulting genome assemblies are highly contiguous with mean N50 and L50 values of 3.09 ± 1.27 Mbp and 13.24 ± 21.19 scaffolds, respectively (Supplementary Fig. 1), an improvement of ~10-to-14-fold from previously available genome assemblies21. Nine additional isolates were sequenced using only short-read technology (mean N50: 262.73 ± 40.94 Kbp; mean L50: 74.00 ± 10.98 scaffolds).
Species determination via molecular phylogenetic analyzes of the taxonomically informative loci of β-tubulin and calmodulin (Supplementary Fig. 2 and 3) placed the clinical isolates into the clades of four distinct species: A. latus (29), A. spinulosporus (8), A. quadrilineatus (1) and A. nidulans (11). Notably, species determination using matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry, a standard method for microbe identification in clinical laboratories22, revealed variable accuracy for each species (p < 0.001, Fisher’s exact test; Supplementary Data 2); all examined A. latus isolates and six out of nine examined A. nidulans isolates were misidentified, whereas all A. spinulosporus isolates were accurately identified. Misidentification of A. latus is largely due to the lack of inclusion of standards for the species from MALDI-TOF databases. This finding suggests that A. latus infections are likely underreported, and that A. latus data should be included in future updates of MALDI-TOF databases. A previously sequenced and assembled clinical isolate of A. latus, MO4614921, was added to the dataset resulting in a total of 53 isolates: 50 clinical isolates and three type strains (Supplementary Fig. 4).
Clinical isolates were isolated from patients with diverse, but primarily lung-related, pathologies, such as cystic fibrosis and chronic granulomatous disease (Supplementary Data 1). Six misidentified A. latus clinical isolates came from patients co-infected with COVID-19. While some analyzes have been done on A. fumigatus COVID-19-associated pulmonary isolates9,23, identifying A. latus co-infections adds to the evidence that other Aspergillus species can also establish coinfections with SARS-CoV-224,25.
Aspergillus latus is an allodiploid hybrid that has retained the genome organization and content of both parental genomes
Genome features, phylogenomics, and macrosynteny provide unequivocal support for the hybrid origin of A. latus. The genomes of A. latus isolates are approximately twice the size, encode twice the number of genes compared to closely related species in section Nidulantes, and contain two copies of the vast majority of near-universally single-copy orthologs (or BUSCO genes; specifically, 96.68 ± 2.46% of BUSCO genes are duplicated), all features suggestive of a diploid genome (Fig. 1A; Supplementary Figs. 1 and 5). Macrosynteny analysis between A. spinulosporus NRRL 2395 T, A. quadrilineatus NRRL 201 T, and A. latus NRRL 200 T revealed large genomic segments that are syntenic with A. spinulosporus and A. quadrilineatus, further supporting an allopolyploid origin (Fig. 1B).
A The genomes of Aspergillus hybrids have larger genomes (x-axis), encode more genes (y-axis), and numerous near-universally single copy orthologs (or BUSCO genes) are present in duplicate copies (data point size). The inset table provides averages ± standard deviation for each metric for the diploid genomes of the A. latus hybrids (Alat; purple) and the haploid genomes of A. spinulosporus (Aspi; blue), A. quadrilineatus (Aqua; red), and A. nidulans (Anid; gray). B Synteny analysis between the A. latus, A. quadrilineatus, and A. spinulosporus type strains. Syntenic blocks contain a minimum of 15 genes robustly assigned to one of the parental species. Scaffolds greater than 100 kilobases are depicted. C Phylogenomic tree of the 49 clinical isolates and type strains for each species. Note that the genomes of the hybrid isolates and the A. latus type strain were split into their two subgenomes (blue: A. spinulosporus subgenome; red: A. quadrilineatus-like subgenome). Links connect subgenomes from the same isolate. D Comparative analysis of evolutionary rates using tree-based and sequence-based measures revealed parental subgenomes are evolving at indistinguishable rates [p = 1.0 for both tests; Tukey’s Honest Significant Differences Test, which was conducted after a multi-factor ANOVA test also revealed no significant differences between the two parental genomes (p = 0.054)]. E The majority of ohnolog pairs are evolving at the same rate (gray), although sometimes the A. spinulosporus subgenome evolves slightly faster than the A. quadrilineatus-like subgenome (blue) and vice versa (red). F There is little evidence of recombination between parental subgenomes; specifically, 1057 (88.30%) scaffolds had no evidence of recombination. Source data are provided as Source Data files.
Further evidence of hybridization was identified using phylogenomics; specifically, the topologies of 3746 BUSCO genes distributed across all 16 A. latus chromosomes and present in two copies were ohnologs, i.e., they showed evidence of one BUSCO gene originating from A. spinulosporus and the other from a close, but unknown, relative of A. quadrilineatus (Fig. 1C; Supplementary Fig. 6), consistent with previous findings21. Hereafter, we will refer to the gene duplicates that have originated from the allodiploidization that gave rise to A. latus as ohnolog pairs.
Reciprocal best BLAST hit analysis between A. spinulosporus NRRL 2395 T and A. quadrilineatus NRRL 201 T identified 9443 orthologs, representing the upper limit of ohnolog pairs in the A. latus hybrids. Reciprocal best BLAST hit analysis between individual A. latus hybrid protein-coding sequences and the combined coding sequences of A. spinulosporus NRRL 2395 T and A. quadrilineatus NRRL 201 T revealed an average of 91.63 ± 2.41% (8652.26 ± 227.96) of ohnologs have been retained (Table 1). Among ohnologs where one copy has been retained, 6.26 ± 1.58% (591.58 ± 149.09) originate from the A. spinulosporus subgenome and 1.79 ± 1.16% (169.26 ± 109.79) originate from the A. quadrilineatus-like subgenome. Lastly, 0.32 ± 0.12% (29.90 ± 11.29) of ohnologs pairs were inferred to have been lost. Together, these results indicate the vast majority of ohnologs have been retained, suggestive of genome stability among the A. latus hybrids.
Subgenomes have similar evolutionary rates and recombine rarely
To further understand the genome organization and diversification of the two subgenomes, we next compared the evolutionary rates between parental subgenomes using tree- and sequence-based measures among 1712 ohnolog pairs. This analysis revealed that the subgenomes are evolving at statistically indistinguishable rates (Fig. 1D; F = 3.72, p = 0.08, Multi-factor ANOVA; p = 1.0 for pairwise comparisons for tree- and sequence-based measures, Tukey’s Honest Significant Differences Test). While one parental genome marginally evolved faster than the other at times (Fig. 1E), most ohnolog pairs (N = 1073; 62.68%) evolved at the same rate. This finding suggests that the parental subgenomes are evolving similarly in the A. latus hybrids.
The chromosome-level genome assemblies and the high nucleotide sequence divergence (7%) of the subgenomes enabled us to examine whether recombination occurs between the subgenomes of A. latus. Using the A. latus NRRL200T genome as a reference, we found that its two subgenomes display little evidence of recombination, as measured by scaffolds containing syntenic blocks of 15 or more consecutive genes assigned to one or the other parent (Fig. 1B). Across a broader sampling of isolates with highly contiguous genome assemblies (N = 22), we found that 88.30% (1057/1197) of scaffolds did not show signatures of recombination between the parental subgenomes (Fig. 1F; Supplementary Data 3; Supplementary Fig. 8; Supplementary Data 4).
Examination of the relationship between the frequency of recombination and the genome assembly quality of the A. latus isolates revealed that the number of scaffolds with evidence of recombination is significantly correlated to the number of scaffolds in a genome assembly (ρ = 0.57, p = 0.007, Spearman rank correlation) and the genome assembly N50 (ρ = −0.48, p = 0.027, Spearman rank correlation) (Supplementary Fig. 9). Therefore, we cannot exclude the possibility that at least some of the observed recombination is an artifact stemming from genome assembly errors. We conclude that A. latus hybrid genomes display little, if any, evidence of recombination between the parental genomes.
The origin of A. latus in time
The denser sampling of isolates allowed us to investigate the number and timing of the hybridization event(s) that gave rise to A. latus. The maximum likelihood phylogeny of our phylogenomic dataset, which includes all the parental subgenomes in the A. latus isolates as well as all A. spinulosporus, A. quadrilineatus, and A. nidulans isolates, suggests that A. latus originated via two distinct hybridization events – one for the environmental isolate A. latus NRRL 200 T and one for all other A. latus clinical isolates (Fig. 2). However, levels of sequence divergence are very low among most A. latus isolates (evidenced by their very short branch lengths in the phylogeny; Fig. 1C) and there is low support for the divergence of A. latus NRRL 200 T from other isolates (SH-aLRT = 71.7; UFBoot = 97; Fig. 1C; Supplementary Fig. 7). Thus, we conducted phylogenetic topology constraint tests of the alternative hypothesis that there was a single hybridization event. We used an approximately unbiased test to determine whether a constrained tree topology where all the A. spinulosporus subgenomes of A. latus isolates were monophyletic had a significantly lower likelihood score than the unconstrained maximum likelihood tree. The likelihood scores of the two trees were not significantly different in both the 3746-gene data matrix ((∆L = 139.84, p = 0.72) and a 2918-gene data matrix comprised of genes that had at least 90% taxon occupancy (∆L = 29.564, p = 0.44). Different methods of topology testing yielded the same result (Supplementary Table 1). Therefore, the most parsimonious explanation is that A. latus originated via a single hybridization event. Relaxed molecular clock analyzes suggest A. latus arose approximately 13.9–13.1 (lower bound: 16.6; upper bound: 7.4) million years ago during the Miocene (Fig. 2).
Relaxed molecular clock analyzes of the origin of each parental subgenome suggest the A. latus hybrids first arose approximately 13.7–13.1 million years ago. Divergence times are depicted next to each internode (depicted as: divergence time [lower bound, upper bound]). Species and colors are indicated as follows: A. spinulosporus (Aspi; blue), A. quadrilineatus (Aqua; red), and A. nidulans (Anid; gray). Source data are provided as a Source Data file.
Variation in A. latus gene content is pronounced among biosynthetic gene clusters
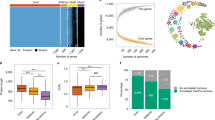
To begin examining the genetic diversity of A. latus, we assigned orthologous groups of genes (a proxy for gene families) to four different bins of occupancy: a bin of core genes present in all A. latus isolates and three different bins of accessory genes present in <100% of A. latus Isolates. This analysis primarily leveraged genome assemblies generated using long- and short-read technologies (N = 22) since fragmented genomes contribute to error in assessing gene content variation26. Accessory genes were assigned as softcore (present in nearly all isolates; N = 21), shell (present in 5-95% of isolates; 2 ≤ N < 21), or cloud (present in less than 5% of isolates; N = 1). Across 10,078 gene families, 7485 (74.27%), 1448 (14.37%), 793 (7.87%), and 352 (3.49%) were determined to be core, softcore, shell, and cloud, respectively (Fig. 3A). We then used the same approach to examine gene content variation of biosynthetic gene clusters (BGCs) involved in secondary metabolism (hereafter referred to as the BGCome). Assigning BGCs to BGC families revealed that 46 (48.42%), 9 (9.47%), 25 (26.32%), and 15 (15.79%) BGC families were assigned to the core, softcore, shell, and cloud, respectively (Fig. 3B).
A Among a total of 10,078 orthologous groups of genes, 7485 are categorized as core (present in 100% of isolates; N = 22) and 2593 as accessory (present in <100% of isolates; N < 22); among these accessory genes, 1448 are softcore (present in ≥95% and <100%; N = 21), 793 are shell (5-95% of isolates; 21 > N ≥ 2), and 352 are cloud (present in less than 5% of isolates; N = 1). B Among 95 biosynthetic gene cluster families (BGCFs), 46 are categorized as core and 49 as accessory (9 are software, 25 are shell, and 15 are cloud). C The number of accessory gene families increases as the number of strains increases, suggesting additional sequencing is needed to fully capture A. latus gene content variation. N values varied between 1 and 26,393 depending on the number of strains being analyzed. D The number of accessory BGCFs substantially increases with additional isolates suggesting the gene content variation of BGCs has also yet to be captured. Notably the core genome is larger among gene families, whereas the accessory portion is larger among BGCFs. Errors bars indicate standard deviation. E Protein sequence lengths differed among gene categories wherein core and softcore genes are longer than shell and cloud genes (N = 10,079). F As genes were less frequently observed among isolates, they were also functionally annotated less frequently. G The number of genes in BGCs differed per category wherein softcore BGCs tend to be smaller than BGCs categorized as core, shell, and cloud (N = 2511). H Few BGCs are predicted to make known secondary metabolites. Source data are provided as Source Data files. For panels E and G, statistical comparisons were made using a Kruskal–Wallis rank sum test (p < 0.01 for both tests); pairwise comparisons were made using the Dunn’s test. One, two, and three asterisks represents a significance threshold of 0.05, 0.01, and 0.001, respectively. In panels C–E, G, average values are depicted and error bars indicate the standard deviation from the mean.
Examination of the relationship between the number of gene families or BGCs in the core and accessory (softcore, shell, and cloud) gene families underscored their differing evolutionary dynamics. Core genes are substantially more common than accessory genes among A. latus isolates (Fig. 3C). In contrast, the accessory BGCome is larger than the core (Fig. 3D). Further sampling of chromosome-level assemblies will enable more precise estimates of the size of the core and accessory parts of the A. latus genome.
Proteins encoded by the core and softcore gene families significantly differ in protein sequence lengths (χ2 = 165.57, p < 0.001, df = 3; Kruskal–Wallis Rank Sum test) (Fig. 3E). Specifically, the average protein sequence length among core and softcore genes is 515.25 ± 369.06 and 517.45 ± 389.26 amino acids, respectively, and are significantly longer than shell and cloud genes that have an average length of 441.94 ± 412.28 and 401.68 ± 369.06 amino acids, respectively (p < 0.001 for both comparisons; Dunn’s test, Benjamini–Hochberg (BH) multi-test correction). Shell genes are also longer than cloud genes (p = 0.004; Dunn’s test, BH multi-test correction). Core and accessory genes also differed in the proportion of genes that could be functionally annotated, wherein a greater proportion of genes were annotated in the core and softcore gene families and fewer in the shell and cloud (Fig. 3F).
Core and accessory BGCs significantly differed in the number of genes encoded in each (χ2 = 166.19, p < 0.001, df = 3; Kruskal–Wallis Rank Sum test) (Fig. 3G). Specifically, softcore BGCs, which have an average of 10.69 ± 3.99 genes, encode significantly fewer genes than core, shell, and cloud BGCs, which have an average of 14.45 ± 4.46, 15.04 ± 6.71, and 15.10 ± 7.04 genes, respectively (p < 0.001 for all comparisons; Dunn’s test with BH multiple test correction). The products, if any, of most BGCs are not known, suggesting A. latus isolates may be capable of producing novel secondary metabolites; alternatively, numerous BGCs encoded in the A. latus genomes may be nonfunctional (Fig. 3H).
Phenotypic profiles of hybrid isolates are distinct
Hybridization can change the phenotypic profile of organisms27. Extensive phenotyping of infection-relevant traits (i.e., growth under cell wall perturbation and integrity stresses, growth under oxidative stress, growth at 44 °C, 37 °C, and 30 °C, susceptibility to voriconazole, amphotericin B, and caspofungin, growth in minimal media, asexual spore size, and production of cleistothecia) across all isolates including the synthetic diploid strain R153XR21 of A. nidulans, revealed that hybrids were phenotypically distinct from the other three species, but were most similar to A. spinulosporus (Fig. 4A). Drug resistance and a cell wall stressor were the largest contributors to the variance along the first dimension in principal component space (Fig. 4B; Supplementary Fig. 11), whereas oxidative stress and asexual spore size, which his correlated with genome size21, contributed most to the variance along the second dimension (Fig. 4C). Notably, 14 A. latus and five A. spinulosporus isolates had high drug resistance to caspofungin compared to reference isolates of A. nidulans (varying from 2.0 to > 4.0 ug/mL and 0.03 ug/mL, respectively). Variation among the species was also observed in an invertebrate moth model of disease (p = 0.04; Log-Rank test) (Supplementary Fig. 12; Supplementary Data 6). Variation among clinically relevant traits suggests that each species may warrant unique strategies to prevent and combat disease.
A Principal component analysis of 14 infection-relevant traits reveals each species is phenotypically distinct. Note, some A. latus isolates are like some A. spinulosporus isolates. B Distributions of phenotypic profiles for the three traits that contribute most to the variance along the first and C second dimensions. The y-axis for radial growth in the presence of stressors (calcofluor white and menadione). The red dot for conidia size is a measurement from a synthetic diploid isolate. For panels B and C, N = 46. D Secondary metabolite profiling of six representative strains and E correlation analysis reveals variation between species and within A. latus. F Quantity of sterigmatocystin production in six strains (N = 24). Errors bars represent one standard deviation. G, H Near-equal RNA abundance between parental subgenomes in A. latus CI 1908 (N = 20,025). I The numbers of significantly differentially expressed genes between 37 °C and 30 °C are very similar; specifically, among upregulated genes, 218 are from the A. spinulosporus subgenome and 213 are from the A. quadrilineatus-like subgenome (χ2 = 0.058, p = 0.81, df = 1; Chi-squared test) and, among downregulated genes, 242 are from the A. spinulosporus subgenome and 232 are from the A. quadrilineatus-like subgenome (χ2 = 0.211, p = 0.65, df = 1; Chi-squared test). J Among significantly upregulated ohnologs genes, 186 belong to ohnolog pairs (93 total ohnolog pairs) were both upregulated. In contrast, 90 A. spinulosporus and 92 A. quadrilineatus-like genes were significantly upregulated but the corresponding ohnolog was not differentially expressed. Among significantly downregulated genes, 160 belong to ohnolog pairs (80 total ohnolog pairs), whereas 114 A. spinulosporus and 122 A. quadrilineatus-like genes had subgenome-specific responses. This finding suggests that the A. latus expression profile in response to environmental perturbation contains both conserved elements (both members of ohnolog pairs in the two subgenomes showing the same change of gene expression) and unique ones. Source data are provided as Source Data files. For panels B, C, G, H, box plots depict the 25th, median, and 75th percentiles; whiskers represent 1.5 times the interquartile range from the hinge; and violin plots depict the underlying distribution.
Examination of secondary metabolite profiles among six isolates—three A. latus isolates and single representatives from A. spinulosporus, A. quadrilineatus, and A. nidulans—revealed that A. latus hybrids were qualitatively more like A. quadrilineatus (Fig. 4D). Correlation analysis revealed that A. latus CI 1908 was most similar to A. quadrilineatus NRRL 201 T (Fig. 4E). Examination of the absolute concentration of a mycotoxin, sterigmatocystin, in the six isolates revealed that A. latus hybrids produced more toxin than either parental species (A. spinulosporus and A. quadrilineatus) but less than A. nidulans (Fig. 4F). Together with other phenotyping data, these results suggest that A. latus hybrids are phenotypically distinct from other species but are more like one parent for some traits, more like the other parent for others, and differ from both parents in yet others.
We next determined if A. latus gene expression patterns were biased toward one or the other parent. RNA-sequencing of A. latus CI 1908 at 30 °C and 37 °C revealed nearly equal transcript abundances in both parental subgenomes (Fig. 4G,H). For example, at 37 °C the A. spinulosporus and A. quadrilineatus-like subgenomes had average transcript per million (TPM) values of 47.72 ± 355.86 and 49.44 ± 369.27, respectively; a similar observation was made for transcript abundances at 30 °C. Examining the estimated codon usage bias of each parental genome (Supplementary Fig. 13) corroborates this finding, revealing that both subgenomes are nearly equally optimized.
Differential expression analysis between the two conditions revealed similar numbers of significantly differentially expressed genes in each parental genome (Fig. 4I; 218 and 213 upregulated genes and 242 and 232 downregulated genes in the A. spinulosporus and A. quadrilineatus subgenomes, respectively; p < 0.01 and |log2(fold change)| ≥2 for differential expression analysis). Furthermore, the numbers of differentially expressed parental genes are not significantly different in the two parental subgenomes (Upregulated genes: χ2 = 0.058, p = 0.81, df = 1; Downregulated genes: χ2 = 0.211, p = 0.65, df = 1; Chi-squared test). Among ohnologs, numerous differentially expressed genes belonged to ohnolog pairs (186 among upregulated genes; 160 among downregulated genes), revealing conserved responses to environmental stimuli (Fig. 4J). Differentially upregulated ohnologs were enriched in functional categories such as glutathione metabolic process, GO:0006749 (Supplementary Data 7); glutathione is involved in many bioprocesses including helping confer resistance against heat shock28. Ohnologs also had divergent, subgenome-specific responses. Specifically, among ohnologs, 90 and 114 genes from the A. spinulosporus subgenome and 92 and 122 genes from the A. quadrilineatus subgenomes were differentially upregulated and downregulated, respectively. Subgenome-specific genes were not enriched in any functional categories. These findings suggest that both parental subgenomes contribute to organismal function and respond to environmental perturbations.
Genomic and phenotypic traits that distinguish A. latus from close relatives
Since all A. latus isolates in the present study were misidentified by traditional methods (Supplementary Data 2), we used the newly generated genomic and phenotypic data to identify traits that taxonomically differentiate A. latus from its closest relatives (Fig. 5). At the genomic level, A. latus isolates have larger genome sizes and gene repertoires than other Aspergillus species (Fig. 1A); the larger genome size of A. latus means that the species can be distinguished from its close relatives through Fluorescence-Activated Cell Sorting (or FACS) analysis of DNA content (Supplementary Data 8). Furthermore, amplification and sequencing of single-locus molecular markers, including taxonomically informative loci, is expected to show evidence of two distinct loci that are phylogenetically distinct in a single-locus phylogeny (Fig. 1C). At the phenotypic level, A. latus spores are larger (Fig. 4C) due to their larger genome sizes21. These genomic and phenotypic differences can help taxonomically distinguish A. latus from other pathogens in section Nidulantes and may help elucidate the epidemiology and clinical burden of this cryptic pathogen.
A. latus (purple), A. spinulosporus (blue), and A. quadrilineatus (red) are indistinguishable in culture. At the genomic level, A. latus isolates have larger genome sizes and gene repertoires than other Aspergillus species and can be distinguished from its close relatives through Fluorescence-Activated Cell Sorting (or FACS) analysis of DNA content. Furthermore, amplification and sequencing of single-locus molecular markers, including taxonomically informative loci, is expected to show evidence of two distinct loci that are phylogenetically distinct in a single-locus phylogeny. At the phenotypic level, A. latus spores are larger than those of other species due to their larger genome size.
Discussion
Our study generated and analyzed extensive amounts of genomic, transcriptomic, chemical, and phenotypic data to shed light on a cryptic fungal pathogen’s evolutionary history, genetic diversity, and the signature of hybridization across infection-relevant traits. For example, evolutionary analyzes indicate that the hybrid A. latus first arose approximately 13 million years ago (Figs. 1 and 2). While our long-read genome assemblies substantially improve previous short-read assemblies, chromosome-level assemblies will help shed light on more aspects of chromosome biology among A. latus hybrids. Moreover, although we are confident one parent is A. spinulosporus, the other parent remains unknown; discovery, sequencing, and phenotyping of the unknown parent will clarify the genomic and phenotypic contribution of the other parental species. Notwithstanding these caveats, our analyzes uncovered several aspects of A. latus genome biology. For example, when one copy of a gene is lost from an ohnolog pair, A. spinulosporus gene copies were more frequently retained (Table 1). Lastly, several isolates are from patients with COVID-19-associated pulmonary aspergillosis. We anticipate that the isolates obtained from patients also infected with SARS-CoV-2 (the virus that causes COVID-19) and associated data may serve as useful resources to study COVID-19-related superinfections.
Examination of the gene content of A. latus revealed that it harbors considerable diversity, with 74.27% (7485/10,078) of gene families being core (Fig. 3). Although chromosome-level assemblies of a much larger number of isolates will be required to characterize the A. latus pangenome, comparison of A. latus to other species with well-defined pangenomes provides useful insights. For example, the Candida albicans pangenome is estimated to contain approximately 7325 gene families, 5432 (74.16%) of which are core (present in all isolates)29, whereas the A. fumigatus pangenome is more open and only 69.34% (7563/10,907) of pan-genes are core30,31. In contrast to gene families, the accessory BGCs outnumber core BGCs — 51.58% of BGCs are accessory—corroborating previous reports that biosynthetic gene clusters evolve rapidly14,32. The A. latus BGCome is less diverse than the A. fumigatus BGCome; specifically, 46 (48.42%) of A. latus BGCs are core, whereas 30.55% (11/36) of A. fumigatus BGCs are core33. A substantial portion of gene families, especially in the BGCome, remain uncharacterized and may represent an untapped source of novel biology and chemistry, especially in the context of an allodiploid organism. Additional sequencing of isolates will help shed light on the pangenome of A. latus.
A. latus has a distinct phenotypic profile compared to parental species and A. nidulans, which is one of the species that A. latus has been previously mistyped as21. We hypothesize that some phenotypes may be additive—for example, A. latus produces the mycotoxin sterigmatocystin in an amount roughly equal to the sum of the amounts produced by its parental species (Fig. 4F). Similarly, both subgenomes respond to temperature stress with similar numbers of genes, including numerous ohnolog pairs (Fig. 4G). In other cases, the hybrids are more like one parent over the other. For example, across all infection-relevant traits examined, A. latus is closer to A. spinulosporus than to A. quadrilineatus; however, this observation may partly be explained by the fact that A. spinulosporus is one of the parental species whereas A. quadrilineatus is a close relative of the other parental species (and not the actual parental species). In contrast, A. latus CI 1908 has a secondary metabolite profile more like A. quadrilineatus than A. spinulosporus. Similarity to one parent over the other may be partly due to dominant genetic effects. These observations corroborate previous findings that complex additive and non-additive effects contribute to hybrid phenotypic profiles in fungi and plants34,35.
The expression patterns of A. latus subgenomes differ from those previously observed in other allodiploid hybrids. Whereas both A. latus subgenomes are actively transcribed and nearly equally respond to environmental perturbations (Fig. 4G), the plant pathogen Verticillium longisporum displays subgenome-specific gene expression patterns36. Moreover, A. latus genomes tend to maintain allodiploidy whereas V. longisporum undergoes haploidization, a common genomic event in the aftermath of hybridization. Maintaining genetic loci after hybridization may result from selection, as previously proposed for Coccidioides fungi37. Whether selection is acting to maintain considerable portions of both parental genomes in A. latus hybrids remains an open question. Mutation rate analysis revealed that both parental subgenomes evolve at indistinguishable rates (F = 3.72, p = 0.08; Multi-factor ANOVA; Fig. 1D,E), suggesting similar selective pressures on both subgenomes. Interestingly, the coffee plant hybrid Coffea arabica, also stems from an allopolyploid event, retains (near) complete parental genomes, and exhibits no discernable expression biases between subgenomes38. However, the coffee plant originated roughly 0.5 million years ago, whereas A. latus is substantially older, suggesting that allopolyploidy may be, in some cases, a stable evolutionary and genomic state.
Accurate taxonomic identification of pathogenic microbes is the first step to their management. Our extensive population-level characterization of genomic and phenotypic diversity in A. latus facilitated the identification of several genomic and phenotypic traits that can reliably taxonomically distinguish A. latus from other species in section Nidulantes, setting the stage for the eventual development of clinical diagnostic tests that enable the medical mycology community to study the epidemiology and clinical burden of this cryptic pathogen.
Methods
Ethics statement
The principles that guide our studies are based on the Declaration of Animal Rights ratified by UNESCO on January 27, 1978, in its 8th and 14th articles. All protocols adopted in this study were approved by the local ethics committee for animal experiments from the University of São Paulo, Campus of Ribeirão Preto (Permit Number: 23.1.547.60.8; Characterization of virulence and immunopathogenicity of Aspergillus spp in the murine model). Groups of five animals were housed in individually ventilated cages and were cared for in strict accordance with the principles outlined by the Brazilian College of Animal Experimentation (COBEA) and Guiding Principles for Research Involving Animals and Human Beings, American Physiological Society. All efforts were made to minimize suffering. Animals were clinically monitored at least twice daily and humanely sacrificed if moribund (defined by lethargy, dyspnea, hypothermia and weight loss). All stressed animals were sacrificed by cervical dislocation.
Genomic DNA extraction
Isolates from Aspergillus section Nidulantes were acquired from several facilities and laboratories (Supplementary Data 1). Where possible, a preliminary species identification was performed by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS; Microflex LT Biotyper, Bruker Daltonics, Bremen, Germany) with the MSI-2 database (https://msi.happy-dev.fr/; 2019 release). The high molecular weight DNA was extracted as follows. Fungal conidia were inoculated in liquid media and let to grow at 37 °C for 16 h under agitation. The mycelia were then filtered through Miracloth, freeze-dried, and disrupted by grinding in liquid nitrogen. The ground material was transferred to a 50 mL Falcon tube, lysed in 10 mL of lysis buffer (3.75 mL Buffer A [3.19 g Sorbitol, 5 mL 1 M Tris-HCl (pH 9), 0.5 mL 0.5 M EDTA (pH 8) and ddH2O up to 50 mL], 3.75 mL Buffer B [10 mL 1 M Tris-HCl (pH 9), 5 mL 0.5 M EDTA, 5.84 g NaCl, 1 g CTAB and ddH2O up to 50 mL], 1.5 mL 5 % Sarkosyl; 1 mL 1 % PVP; 100 μL Proteinase K) and incubated 30 min at 65 °C. Subsequently, 3.35 mL of 5 M Potassium acetate (pH 7.5) were added, and the samples were incubated on ice for 30 min. After 30 min centrifugation at 5,000 g and 4°C, the supernatants were transferred to a new Falcon tube. The samples were then treated with Phenol:Chloroform:Isoamylalcohol (25:24:1), tubes were centrifuged, the aqueous phase (~8 mL) was transferred to a new tube and treated with 100 μL RNase A (10 mg/ml) for 60 min at 37°C. The genomic DNA was precipitated by adding 1/10 volume of 3 M sodium acetate and 1 volume ice-cold 96% ethanol and centrifuged 30 min at 10,000 G-force and a temperature of 4 °C. The resulting pellet was washed with 70% ethanol, dried, resuspended in 500 μL TE, and stored at -20°C until further use. The DNA concentration was estimated using a Qubit fluorometer. All other Aspergillus genomic DNA extraction were performed according to Goldman and colleagues39. The resulting DNA was sequenced using Oxford Nanopore and Illumina technologies at Nextomics Biosciences in Wuhan, China. This resulted in 10,565,680 Nanopore reads (152,924,029,920 total bases) with an average length of 15,492.31 ± 4991.34 base pairs and 1,145,484,780 Illumina reads with a length of 150 base pairs (171,822,717,000 bases). Nine isolates—which have “8954-AG” at the beginning of their identifier (Supplementary Data 1)—were sequenced using 150 bp paired-end short-read technology (NovaSeq6000) at the Vanderbilt Technologies for Advanced Genomics (VANTAGE), generating 250,464,221 reads (37,569,633,150 bases). Library preparation was conducted using the Twist Whole Genome / Metagenomics_LowpassWGS-Twist Miniprep (Twist Bioscience).
Genome assembly and annotation
Each isolate was assembled using two different approaches. First, a long sequencing read-based assembly was generated by NextDenovo, v2.3.040, using Oxford Nanopore sequencing data alone. The assembly was then polished by Nextpolish, v1.3.041, using both long- and short-sequencing read data. Second, a hybrid assembly was generated by MaSuRCA, v3.4.142, using both long- and short-sequencing data. Before assembly, reads were quality-trimmed using fastp, v0.12.643, with default settings. The two assemblies were compared for their assembly size, scaffold number, and scaffold N50 value. For the A. latus isolates, the hybrid assemblies were selected as they were about twice in size as the corresponding Nanopore-based assemblies. For other isolates, the Nanopore-based assemblies were selected as they were better in continuity. Among the nine genomes that were sequenced using only short-read technology, reads were first quality-trimmed using Trimmomatic, v0.3944 using the following parameters: LEADING:20 TRAILING:20 MINLEN:60. The resulting reads were used as input to SPAdes, v.3.14.045, for genome assembly. For genome annotation, all genome assemblies were first softmasked for repetitive sequences using RepeatMasker, v4.1.0 (http://www.repeatmasker.org; “-species” option set to “Aspergillus”). Gene models were generated using BRAKER, v2.1.446, which combines ab-initio gene predictors (AUGUSTUS, v3.3.347, and GeneMark-ES, v4.5948) and homology evidence (all Eurotiomycetes protein sequences in the OrthoDB, v1049, database).
To examine the quality of all genome assemblies examined, diverse metrics that describe genome assembly and gene content completeness were determined. Genome assembly metrics—such as assembly size, the number of scaffolds, N50, L50, and others—were calculated using BioKIT, v0.0.9 or v1.1.050. Gene content completeness was evaluated using BUSCO, v4.0.451 with the Eurotiales dataset (creation date: 2020-08-05) of 4,191 near-universally single copy orthologs (commonly referred to as BUSCO genes) from OrthoDB, v1049.
Assigning genes to parent-of-origin
Determining gene parent-of-origin was done using two approaches. For BUSCO genes, a quartet decomposition approach was implemented wherein single-gene phylogenies were pruned to quartets (phylogenies with four leaves) with the type strains of A. spinulosporus NRRL2395T, A. quadrilineatus NRRL201T, A. nidulans A4, and a single-gene from a newly sequenced isolate (Supplementary Fig. 10). The parent-of-origin was determined to be the species whose sequence exhibited the shortest nodal distance to the sequence of the newly sequenced isolate in the quartet. As expected, genes from haploid isolates determined to be A. spinulosporus, A. quadrilineatus, and A. nidulans using molecular phylogenetics (Supplementary Fig. 2 and 3) were correctly assigned to their respective species (Supplementary Fig. 6) demonstrating the efficacy of this approach. Single-gene phylogenies were constructed by aligning protein sequences of BUSCO genes using MAFFT, v7.40252, with the “auto” parameter. Nucleotide sequences were threaded onto the protein alignments using the “thread_dna” function in PhyKIT, v1.10.0 or 1.19.053, to create codon-based alignments. The resulting alignments were trimmed with ClipKIT, v1.3.0 or 2.2.554, using the “smart-gap” mode. Single-gene phylogenies were inferred using IQ-TREE 2, v2.0.655. The best fitting substitution model was automatically determined according to Bayesian Information Criterion using ModelFinder56. Nodal distances were calculated using the “tip_to_tip_node_distance” function in PhyKIT, v1.10.0 or 1.19.053. This analysis was done for 3746 BUSCO genes. For all other genes, a reciprocal best BLAST hit approach was used between the coding sequences of the query hybrid genome and the concatenated coding sequences of A. spinulosporus NRRL2395T and A. quadrilineatus NRRL201T. Reciprocal best BLAST hit analysis was done using the blastn function from the BLAST+ suite, v2.3.0 + 57.
Reconstructing species- and strain-level evolutionary histories
To determine the number of hybridization events and to place the evolutionary history of the hybrid subgenomes in geologic time, a phylogenomic approach was employed. First, 3746 BUSCO genes with a median taxon occupancy of 88 out of 90 haploid and diploid subgenomes were identified (average occupancy = 79.72 ± 18.38 taxa). The trimmed codon-based alignments generated during parent-of-origin determination were concatenated together. For hybrid genomes, only sequences from the same subgenome were concatenated together. For example, A. latus NRRL200T was represented by two concatenated sequences: one representing the A. spinulosporus subgenome and the other representing the A. quadrilineatus-like subgenome. The resulting alignment had 7,033,120 sites (2,594,839 parsimony informative sites; 3,829,507 constant sites). The concatenated matrix was used to reconstruct the evolutionary history of all species and isolates using IQ-TREE 2, v2.0.655. During tree search, the number of trees in the candidate set was increased from five to ten. The best-fitting model was determined using Bayesian Information Criterion values as described above. Bipartition support was measured using two metrics: 5000 ultrafast bootstrap approximations58 and Shimodaira–Hasegawa-like approximate likelihood ratio test using 1000 replicates59.
Determining the number of hybridization events
The A. spinulosporus parental subgenomes of hybrid isolates were placed into two distinct clades in our phylogenomic analysis (Fig. 1C), suggesting two hybridization events. To evaluate if a single hybridization event also accurately describes the evolutionary history of A. latus hybrids, a topology test was conducted. The log-likelihoods of the inferred phylogeny in Fig. 1C and a maximum likelihood phylogeny where the A. spinulosporus subgenomes were constrained to be monophyletic were compared using an approximately unbiased test60 with 5000 replicates. The test was done for both the complete data matrix (Ngenes = 3746) and another data matrix comprised of genes that had at least 90% (81/90) taxon occupancy (Ngenes = 2918). To ensure our results were not sensitive to the testing approach, six other topology tests were also conducted: bootstrap proportions using resampling estimated log-likelihoods61; a one-sided Kishino-Hasegawa and weighted Kishino-Hasegawa test62; a Shimodaira–Hasegawa and weighted Shimodaira–Hasegawa test63; and an expected likelihood weight test64.
Evolutionary rate analysis of subgenomes
To determine if the parental subgenomes in the A. latus hybrids are evolving at similar or different rates, we used tree- and sequence-based measures to quantify evolutionary rate. To do so, each protein-coding gene per A. latus hybrid was concatenated into a putative group of orthologous genes based on the reciprocal best BLAST hit information. Ohnolog pairs were determined via reciprocal best BLAST analysis between A. spinulosporus NRRL 2395 T and A. quadrilineatus NRRL 201 T. Ohnolog pairs with at least 10 sequences in each ohnolog were used to evolutionary rate analysis (1712 ohnolog pairs; 3424 groups of orthologous genes). Individual groups of orthologous genes were aligned, trimmed, and subsequently used for phylogenetic tree inference using the abovementioned strategy. The tree-based evolutionary rate was measured using the “evo_rate” function in PhyKIT, v1.19.053 which is the total tree length divided by the number of taxa. The sequence-based measured as one minus the mean pairwise identity among sequences; mean pairwise identity was calculated using the “pairwise_identity” function in PhyKIT. Only ohnolog pairs that had consistent results between tree- and sequence-based measures of evolutionary rate were retained; for example, both measures had to indicate that the same parental subgenome was evolving faster or that evolutionary rates were equal in both subgenomes.
Timetree analysis
To place the evolutionary origins of A. latus in geologic time, timetree analysis was conducted using the Bayesian framework implemented in MCMCTree from PAML, v4.9d65. A 1054-gene data matrix with full taxon occupancy was used because limited taxon sampling is a known source of error in phylogenomics66. Sequences in the 1054-gene data matrix were first subsampled to those belonging to Nidulantes (N = 86) and the sister section, Versicolores (N = 2). Next, the substitution rate was estimated using a strict clock model (clock = 1) and a general time reversible model with unequal rates and unequal base frequencies67; rate heterogeneity across sites was accounted for using a discrete Gamma model with four categories68 (model = 7); the root was point calibrated to 27.55 million years ago—the split between sections Nidulantes and Versicolores based on previous whole-genome estimates69. The substitution rate was estimated to be 0.07 nucleotide substitutions per site per 10 million years.
Next, the likelihood of the alignment was approximated using a gradient and Hessian matrix—a matrix that describes the curvature of the log-likelihood surface. Estimating a gradient and Hessian matrix requires time constraints, which were set as follows: the origin of Nidulantes was constrained to an upper bound of 5.44 million years ago and a lower bound of 16.57 million years ago; the split between Nidulantes and Versicolores was set to an upper bound of 19.79 million years ago and a lower bound 35.40 million years ago). Time constraints were based on the same study used to point calibrate the root during substitution rate estimation69. Lastly, the resulting gradient and Hessian matrix divergence times were estimated using a relaxed molecular clock (clock = 2). The gamma distribution shape [a = (s/s)2] and scale [b = s/s 2] are defined by the substitution rate (s) and were set to 1 and 14.29, respectively (Supplementary Data 5). Rate variation across branches was accounted for by setting the σ2 parameter to “1 4.5.” During Markov Chain Monte Carlo analysis, a total of 5.1 million iterations were run—the first 100,000 observations were discarded and 10,000 samples were obtained by sampling every 500 iterations. The total number of iterations run is 510 times greater than the recommended minimum59. This analysis was conducted for tree topologies representing one- and two-hybridization events.
Analysis of gene content variation
Protein sequences in the dataset were clustered into orthologous groups of genes (a proxy for gene families) using OrthoFinder, v2.3.870 using an inflation value for Markov clustering of 1.5. Sequence similarity searches were done using DIAMOND, v2.0.13.15171. The resulting gene families were then binned into categories reflecting gene family occupancy following a previously established protocol30 where: core gene families are present in 100% of isolates (N = 22); softcore gene families are present in less than 100% but more than or equal to 95% of isolates (N = 21); shell gene families are present in 5-95% of isolates (21 > N ≥ 2); and cloud gene families are present in less than 5% of isolates (N = 1). The putative function of each gene family was determined by annotating the genes in each family using InterProScan, v5.53-87.072 and the Pfam73, PANTHER74, and TIGRFAM75 databases.
The patterns of core and accessory (softcore, shell, and cloud) biosynthetic gene cluster families were also examined. Biosynthetic gene cluster boundaries were predicted using antiSMASH, v4.1.076. Two biosynthetic gene clusters were determined to be putatively orthologous if at least 50% of the genes they encode are reciprocally orthologous following a previously established protocol77. The putative function of each biosynthetic gene cluster was determined by cross-referencing predicted clusters with known clusters in the MIBiG database78.
Phenotyping among infection-relevant traits
To determine growth on solid media, a drop containing 1 × 104 spores was inoculated at the center of a petri dish containing solid minimal medium [1% (w/v) glucose, original high nitrate salts, trace elements, pH 6.5]. The plates were incubated at 37 °C for 5 days. Trace elements, vitamins, and nitrate salts compositions were as described previously79.
Radial growth was used to compare how the different strains respond to cell wall stressors (congo red and calco-fluor white-CFW), temperature (30, 37, and 44 °C) and oxidative stress (H2O2 and menadione). Strains were grown in solid GMM, inoculated with 1 × 105, and incubated for 5 days at 37 °C before measuring colony diameter. To induce cell wall stress, 40 μg/mL of Congo red (Sigma–Aldrich) or CFW (Sigma–Aldrich) was added to the medium. For oxidative stress assay, plates supplemented with 0.1 mM menadione (Sigma–Aldrich) and 10 mM H2O2 (Merck) were used. Radial growth for the stresses was measured as ratios, dividing colony radial diameter (cm) of growth in the stress condition by colony radial diameter in the control (no stress) condition. Finally, the radial growth of the strains was quantified at 30 and 44 °C.
Antifungal susceptibility testing for voriconazole (Sigma–Aldrich), amphotericin B (Sigma–Aldrich) and caspofungin (Sigma–Aldrich) was performed by determining the minimal inhibitory concentration (MIC) or minimal effective concentration (MEC) (only caspofungin) according to the protocol established by the Clinical and Laboratory Standards Institute70.
The diameters of 100 spores for each isolate were measured under a Carl Zeiss (Jena, Germany) AxioObserver.Z1 fluorescent microscope equipped with a 100-W HBO mercury lamp using the 100x magnification oil immersion objective and the software AxioVision, v.3.1.
For the crossing experiments, the lineages were inoculated with minimal medium and incubated for two days at 37 °C. For sexual cycle induction, the plates were sealed with tape to reduce oxygen tension. After incubation for 14 days at 30 °C and 37 °C, the cleistothecia were isolated using a magnifier and cleaned in a petri dish containing 4% (p/v) of agar. Cleistothecia were broken into microtubes containing water to release the ascospores, which were inoculated in minimal medium and incubated at 37 °C to check their viability.
To visualize phenotypic variation between strains of A. latus, A. spinulosporus, A. quadrilineatus, and A. nidulans, a principal component analysis (PCA) using the factoextra, v2.8 (https://cran.r-project.org/web/packages/factoextra), and FactoMineR, v1.0.7 (https://cran.r-project.org/web/packages/FactoMineR), packages in R, v4.3.0, were utilized. More specifically, the dataset included measures of fungal growth in response to cell wall (Calcofluor White and Congo red at 40 μg/mL), oxidative (menadione, at 0.07 mM and 0.1 mM concentrations, and H2O2 for three days), and temperature (44 °C, 37 °C, and 30 °C) stressors, in addition to various antifungal medications (voriconazole, amphotericin B, and caspofungin). Other physical characteristics in the growth of each isolate, such as asexual spore (conidial) size and the presence or absence of a viable cleistothecium, were also included for a total of 14 variables used in this PCA. Data were scaled to unit variance before the analysis (scale.unit = TRUE), and 5 dimensions were kept in the final results (ncp = 5).
Survival curves
Galleria mellonella larvae were used to investigate the virulence of the different strains. The larvae were cultivated and prepared as described previously21. The larvae used for the infection were in the last stage of development (sixth week). All selected larvae weighed ∼300 mg and were restricted to food for 24 h before the experiment. The fresh conidia of each specie were counted using a hemocytometer and adjusted to the final concentration of 2 × 108 conidia/mL. Five microliter of each inoculum was injected with a Hamilton syringe (7000.5 KH) through the last left ear (n = 10/group), resulting in 1 × 106 conidia. The control group was inoculated with phosphate buffered saline (PBS). After infection, the larvae were kept with food restrictions, at 37 °C in Petri dishes in the dark and scored daily. The larvae were considered dead due to lack of movement in response to touch. The viability of the inoculum administered was determined by serial dilution of the conidia in YAG medium and incubating the plates at 37 °C for 72 h. The experiment was repeated twice. We separated and assembled the groups with the larvae (n = 10) in Petri dishes. The groups are composed of larvae that are approximately 300 mg in weight and 2 cm long. Moth sex was not accounted for due to the impossibility of visually determining sex at the sixth week of larval development.
Fermentation, extraction, and isolation for chemotyping
To identify the secondary metabolites biosynthesized by A. nidulans, A. latus, A. quadrilineatus, and A. spinulosporus, these species were grown as large-scale fermentations to isolate and characterize the secondary metabolites, similar to methods previously described80. To inoculate oatmeal cereal media (Old Fashioned Breakfast Quaker oats), agar plugs from fungal stains grown on potato dextrose agar; Difco were excised from the edge of the Petri dish culture and transferred to separate liquid seed medium that contained 10 ml YESD broth (2% soy peptone, 2% dextrose, and 1% yeast extract; 5 g of yeast extract, 10 g of soy peptone, and 10 g of D-glucose in 500 mL of deionized H2O) and allowed to grow at 23 °C with agitation at 100 rpm for 3 days. The YESD seed cultures of the fungi were subsequently used to inoculate solid-state oatmeal fermentation cultures, which were either grown at room temperature (~23 °C under 12 h light/dark cycles for 16 days), 30 °C for 12 days, or 37 °C; all growth at the latter two temperatures was carried out in an incubator (VWR International). The oatmeal cultures were prepared in 250 ml Erlenmeyer flasks that contained 10 g of autoclaved oatmeal (10 g of oatmeal with 17 mL of deionized H2O and sterilized for 15–20 min at 121 °C). For all fungal strains, six flasks of oatmeal cultures were grown at 37 °C and 30 °C, except for A. quadrilineatus, which wasn’t grown at 37 °C. Only one flask of each strain was grown for the growth at room temperature.
The cultures were extracted by adding 60 mL of 1:1 mixture of MeOH-CHCl3, chopping thoroughly with a spatula, and shaking overnight (~16 h) at ~100 rpm at room temperature. The culture was filtered in vacuo, and 90 mL CHCl3 and 150 mL H2O were added to the filtrate. The sample was transferred to a separatory funnel and the organic layer was drawn off and evaporated to dryness in vacuo. The dried organic layer was reconstituted in 100 mL of (1:1) MeOH–CH3CN and 100 mL of hexanes, transferred to a separatory funnel and shaken vigorously. The defatted organic layer (MeOH–CH3CN) was evaporated to dryness in vacuo.
Sterigmatocystin calibration curve
High-resolution electrospray ionization mass spectrometry (HRESIMS) experiments utilized a Thermo LTQ Orbitrap mass spectrometer (Thermo Fisher Scientific) that was equipped with an electrospray ionization source. This was coupled to an Acquity ultra-high-performance liquid chromatography (UHPLC) system (Waters Corp.), using a flow rate of 0.3 mL/min and a BEH C18 column (2.1 × 50 mm, 1.7 μm) that was operated at 40°C. The mobile phase consisted of CH3CN–H2O (Fischer Optima LC-MS grade; both were acidified with 0.1% formic acid). The gradient started at 15% CH3CN and increased linearly to 100% CH3CN over 8 min, where it was held for 1.5 min before returning to starting conditions to re-equilibrate.
Extracts were analyzed with four biological replicates, and each of these in technical triplicates, all in the positive ion mode with a resolving power of 35,000. The samples were each prepared at a concentration of 0.2 mg/mL and were dissolved in MeOH and injected with a volume of 3 μL. To eliminate the possibility of sample carryover, a blank (MeOH) was injected between every sample. The sterigmatocystin calibration curve was analyzed in triplicate and ran at concentrations of 102.4, 51.2, 25.6, 6.4, 3.2, 1.6, 0.8 μg/mL. To ascertain the absolute concentration of sterigmatocystin in these samples, batch process methods were run using Thermo Xcalibur (Thermo Fisher Scientific). We used a mass range of 325.0687–325.0719 Da for sterigmatocystin at a retention time of 5.00 min with a 7.00-second window. The calibration curve was analyzed quadratically with a weight index of 1/X since these compounds did not ionize linearly at this large concentration range. The equation used to calculate the absolute concertation of sterigmatocystin in these samples was, Y = 1.99463e + 007 + 5.0315e + 007*X-212053*X2 with a R2 of 0.9987. The absolute concentration of sterigmatocystin per flask of fungal growth was calculated by converting the concentration in the sample injection to the amount in the total extract, followed by averaging the biological replicates.
Chemometric analysis
Untargeted UPLC-MS datasets for each sample were individually aligned, filtered, and analyzed using MZmine 2.20 software (https://sourceforge.net/projects/mzmine/)81 Peak detection was achieved using the following parameters: noise level (absolute value), 5 × 105; minimum peak duration, 0.05 min; m/z variation tolerance, 0.05; and m/z intensity variation, 20%. Peak list filtering and retention time alignment algorithms were used to refine peak detection. The join algorithm integrated all sample profiles into a data matrix using the following parameters: m/z and retention time balance set at 10.0 each, m/z tolerance set at 0.001, and RT tolerance set at 0.5 min. The resulting data matrix was exported to Excel (Microsoft) for analysis as a set of m/z – retention time pairs with individual peak areas detected in triplicate analyses. Samples that did not possess detectable quantities of a given marker ion were assigned a peak area of zero to maintain the same number of variables for all sample sets. Ions that did not elute between 2 and 8 min and/or had an m/z ratio <200 or >800 Da were removed from analysis. Relative standard deviation was used to understand the quantity of variance between the technical replicate injections, which may differ slightly based on instrument variance. A cutoff of 40% was used at any given m/z—retention time pair across the technical replicate injections of one biological replicate; if the variance was greater than the cutoff, it was assigned a peak area of zero. The data underwent standard scaling to normalize the data. Final chemometric analysis was performed with Python. The network graphs were generated using the standard scaled data with Pandas.
Flow cytometry analysis for determination of spores DNA content
Asexual spores were collected, centrifuged (13,000 rpm for 3 min), and washed with sterile PBS. Spores were stained following the protocol previously described82. Briefly, after harvesting, spores were fixed overnight with 70% ethanol (v/v) at 4 °C, washed and resuspended in 850 μL of sodium citrate buffer (50 mM; pH = 7.5) and dissociated by sonication using four ultrasound pulses at 40 W for 2 s with an interval of 1 to 2 s between pulses. Spores were treated with RNase A (0.50 mg/mL; Invitrogen, USA; 1 h; 50 °C) and then with proteinase K (1 mg/mL; Sigma–Aldrich, St. Louis, Missouri, USA; 2 h; 50 °C). Spore DNA was stained with SYBR Green 10,000× (Invitrogen, USA; 0.2% (v/v)), overnight at 4 °C. Before analysis, spores were treated with Triton® X-100 (Sigma–Aldrich; 0.25% (v/v)). Samples were acquired in an LSRII flow cytometer (Becton Dickinson, NJ, USA) with a 488-nanometer excitation laser with Diva as the acquisition software. A minimum of 30,000 spores per sample were analyzed with an acquisition protocol defined to measure forward scatter and side scatter on a logarithmic scale and green fluorescence on a linear scale. Results were analyzed with the FlowJo software (BD Biosciences, USA).
RNA-sequencing extraction and analysis
The RNA extraction was performed according to dos Reis et al. 83 with modifications83. Briefly, 106 spores/mL of each Aspergillus strain were inoculated in 50 mL of liquid minimal medium and grown at 37 °C for 16 h under agitation. Further, the mycelia were filtered through miracloth, freeze-dried, and disrupted by grinding in liquid nitrogen. Total RNA was extracted using TRIzol (Invitrogen), treated with RQ1 RNase-free DNase I (Promega) and purified using the RNAeasy kit (Qiagen) according to the manufacturer’s instructions. RNA samples were quantified using a NanoDrop and its RNA quality was verified using an Agilent Bioanalyzer 2100 (Agilent Technologies). RNAs selected for further analysis had a minimum RNA integrity number (RIN) value of 8.0. For RNA-sequencing, the cDNA libraries were constructed using the TruSeq Total RNA with Ribo Zero (Illumina, San Diego, CA, USA). From 0.1–1 μg of total RNA, the ribosomal RNA was depleted, and the remaining RNA was purified, fragmented, and prepared for complementary DNA (cDNA) synthesis, according to the manufacturer’s instructions. The libraries were validated following the Library Quantitative PCR (qPCR) Quantification Guide (Illumina). Following, the RNA-seq was carried out by paired-end sequencing on the Illumina NextSeq 500 Sequencing System using NextSeq High Output (2 × 150) kit, according to the manufacturer’s instructions. Using the same approach, RNA was also extracted from spores grown at 30 °C.
RNA was sequenced at Vanderbilt Technologies for Advanced Genomics (VANTAGE) on an Illumina NovaSeq machine (paired-end, 150 bp length for each read). All samples passed quality control checks and had acceptable RNA Integrity Numbers. Reads were trimmed with Trimmomatic, v0.3944, using the following options “TruSeq2-PE.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36”. Filtered reads for each condition and replicate were then mapped to the hybrid genome assembly and annotation file using HISAT2, v2.1.084, with default parameters. The resulting SAM files from HISAT2 were then sorted and compressed to BAM files with SAMtools, v1.985. Gene abundance for each condition and replicate were estimated by first filtering the alignment files for only properly paired reads using SAMtools (samtools view -b -f 2 <input.bam > <input.proper.pairs.only.bam > ). Per-sample gene abundances were then quantified in terms of transcripts per million (TPM) using TPMcalculator, v0.0.386. To identify genes that appear differentially expressed between the two treatment conditions, we extracted per-sample unique read counts from each alignment file using from HTseq87 and then passed these counts to DESeq2, v1.3488, to estimate differential expression between conditions. Genes were considered to be significantly differentially expressed if their FDR-corrected (Benjamini and Hochberg method) p-value (padj) was lower than 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The genome sequencing and transcriptomic data generated in this study have been deposited in NCBI under BioProject accession code PRJNA987017. The short-read and long-read genome sequencing data generated in this study have been deposited in the NCBI Sequence Read Archive under accession codes SRR25010807-SRR25010890. The transcriptomic data generated in this study have been deposited in the Sequence Read Archive under the accessions SRR25016725-SRR25016730. Processed data including results and key intermediate files—genome assemblies, annotations, orthology inference, and biosynthetic gene cluster boundaries, for example—are available at figshare (https://doi.org/10.6084/m9.figshare.23589570). Data Source files have also been uploaded to figshare. Source data are provided with this paper.
References
Kainz, K., Bauer, M. A., Madeo, F. & Carmona-Gutierrez, D. Fungal infections in humans: the silent crisis. MIC 7, 143–145 (2020).
Benedict, K., Jackson, B. R., Chiller, T. & Beer, K. D. Estimation of direct healthcare costs of fungal diseases in the United States. Clin. Infect. Dis. 68, 1791–1797 (2019).
Benedict, K., Whitham, H. K. & Jackson, B. R. Economic burden of fungal diseases in the United States. Open Forum Infect. Dis. 9, ofac097 (2022).
Rayens, E. & Norris, K. A. Prevalence and healthcare burden of fungal infections in the United States, 2018. Open Forum Infect. Dis. 9, ofab593 (2022).
Nnadi, N. E. & Carter, D. A. Climate change and the emergence of fungal pathogens. PLoS Pathog. 17, e1009503 (2021).
Fisher, M. C. & Denning, D. W. The WHO fungal priority pathogens list as a game-changer. Nat. Rev. Microbiol. 21, 211–212 (2023).
Van De Veerdonk, F. L. et al. COVID-19-associated Aspergillus tracheobronchitis: the interplay between viral tropism, host defence, and fungal invasion. Lancet Respiratory Med. 9, 795–802 (2021).
Jabeen, K., Farooqi, J., Irfan, M., Ali, S. A. & Denning, D. W. Diagnostic dilemma in COVID-19-associated pulmonary aspergillosis. Lancet Infect. Dis. 21, 767 (2021).
Steenwyk, J. L. et al. Genomic and phenotypic analysis of COVID-19-associated pulmonary aspergillosis isolates of Aspergillus fumigatus. Microbiol Spectr. 9, e00010-21 (2021).
Rokas, A. Evolution of the human pathogenic lifestyle in fungi. Nat. Microbiol. 7, 607–619 (2022).
Fedorova, N. D. et al. Genomic islands in the pathogenic filamentous fungus Aspergillus fumigatus. PLoS Genet 4, e1000046 (2008).
Muñoz, J. F. et al. Genomic insights into multidrug-resistance, mating and virulence in Candida auris and related emerging species. Nat. Commun. 9, 5346 (2018).
Passer, A. R. et al. Genetic and genomic analyses reveal boundaries between species closely related to Cryptococcus Pathogens. mBio 10, e00764–19 (2019).
Steenwyk, J. L. et al. Variation among biosynthetic gene clusters, secondary metabolite profiles, and cards of virulence across Aspergillus Species. Genetics 216, 481–497 (2020).
Steenwyk, J. L., Rokas, A. & Goldman, G. H. Know the enemy and know yourself: Addressing cryptic fungal pathogens of humans and beyond. PLoS Pathog. 19, e1011704 (2023).
Balajee, S. A., Gribskov, J. L., Hanley, E., Nickle, D. & Marr, K. A. Aspergillus lentulus sp. nov., a New Sibling Species of A. fumigatus. Eukaryot. Cell 4, 625–632 (2005).
Franzot, S. P., Salkin, I. F. & Casadevall, A. Cryptococcus neoformans var. grubii: Separate Varietal Status for Cryptococcus neoformans Serotype A Isolates. J. Clin. Microbiol. 37, 838–840 (1999).
O’Donnell, K., Kistler, H. C., Tacke, B. K. & Casper, H. H. Gene genealogies reveal global phylogeographic structure and reproductive isolation among lineages of Fusarium graminearum, the fungus causing wheat scab. Proc. Natl Acad. Sci. Usa. 97, 7905–7910 (2000).
Hagen, F. et al. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. 78, 16–48 (2015).
Sepúlveda, V. E., Márquez, R., Turissini, D. A., Goldman, W. E. & Matute, D. R. Genome sequences reveal cryptic speciation in the human pathogen Histoplasma capsulatum. mBio. 8, e01339–17 (2017).
Steenwyk, J. L. et al. Pathogenic allodiploid hybrids of Aspergillus fungi. Curr. Biol. 30, 2495–2507.e7 (2020).
Bader, O. Fungal species identification by MALDI-ToF mass spectrometry. Methods Mol. Biol. 1508, 323–337 (2017).
Mead, M. E. et al. COVID-19-associated pulmonary aspergillosis isolates are genomically diverse but similar to each other in their responses to infection-relevant stresses. Microbiol Spectr. 11, e05128–22 (2023).
Salmanton-García, J. et al. COVID-19-associated pulmonary Aspergillosis, March-August 2020. Emerg. Infect. Dis. 27, 1077–1086 (2021).
Kariyawasam, R. M. et al. Defining COVID-19–associated pulmonary aspergillosis: systematic review and meta-analysis. Clin. Microbiol. Infect. 28, 920–927 (2022).
Marcet-Houben, M. et al. Chromosome-level assemblies from diverse clades reveal limited structural and gene content variation in the genome of Candida glabrata. BMC Biol. 20, 226 (2022).
Abbott, R. et al. Hybridization and speciation. J. Evol. Biol. 26, 229–246 (2013).
Pócsi, I., Prade, R. A. & Penninckx, M. J. Glutathione, Altruistic Metabolite in Fungi. in Advances in Microbial Physiology vol. 49 1–76 (Elsevier, 2004).
McCarthy, C. G. P. & Fitzpatrick, D. A. Pan-genome analyses of model fungal species. Microb. Genomics 5, e000243 (2019).
Barber, A. E. et al. Aspergillus fumigatus pan-genome analysis identifies genetic variants associated with human infection. Nat. Microbiol 6, 1526–1536 (2021).
Horta, M. A. C. et al. Examination of genome-wide ortholog variation in clinical and environmental isolates of the fungal pathogen Aspergillus fumigatus. mBio. 13, e01519–e01522 (2022).
Rokas, A., Mead, M. E., Steenwyk, J. L., Raja, H. A. & Oberlies, N. H. Biosynthetic gene clusters and the evolution of fungal chemodiversity. Nat. Prod. Rep. 37, 868–878 (2020).
Lind, A. L. et al. Drivers of genetic diversity in secondary metabolic gene clusters within a fungal species. PLoS Biol. 15, e2003583 (2017).
Fournier, T. et al. Extensive impact of low-frequency variants on the phenotypic landscape at population-scale. eLife 8, e49258 (2019).
Xie, J. et al. Large-scale genomic and transcriptomic profiles of rice hybrids reveal a core mechanism underlying heterosis. Genome Biol. 23, 264 (2022).
Depotter, J. R. L. et al. The interspecific fungal hybrid Verticillium longisporum displays subgenome-specific gene expression. mBio 12, e01496–21 (2021).
Neafsey, D. E. et al. Population genomic sequencing of Coccidioides fungi reveals recent hybridization and transposon control. Genome Res. 20, 938–946 (2010).
Salojärvi, J. et al. The genome and population genomics of allopolyploid Coffea arabica reveal the diversification history of modern coffee cultivars. Nat. Genet 56, 721–731 (2024).
Golduran, G. H. & Morris, N. R. [3] Aspergillus nidulans as a model system for cell and molecular biology studies. in Methods in Molecular Genetics vol. 6 48–65 (Elsevier, 1995).
Hu, J. et al. An Efficient Error Correction and Accurate Assembly Tool for Noisy Long Reads. http://biorxiv.org/lookup/doi/10.1101/2023.03.09.531669 (2023) https://doi.org/10.1101/2023.03.09.531669.
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Zimin, A. V. et al. The MaSuRCA genome assembler. Bioinformatics 29, 2669–2677 (2013).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Computational Biol. 19, 455–477 (2012).
Brůna, T., Hoff, K. J., Lomsadze, A., Stanke, M. & Borodovsky, M. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genomics Bioinforma. 3, lqaa108 (2021).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 18, 1979–1990 (2008).
Kriventseva, E. V. et al. OrthoDB v10: sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res. 47, D807–D811 (2019).
Steenwyk, J. L. et al. BioKIT: a versatile toolkit for processing and analyzing diverse types of sequence data. Genetics 221, iyac079 (2022).
Manni, M., Berkeley, M. R., Seppey, M. & Zdobnov, E. M. BUSCO: assessing genomic data quality and beyond. Curr. Protoc. 1, e323 (2021).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: improvements in performance and usability. Mol. Biol. Evolution 30, 772–780 (2013).
Steenwyk, J. L. et al. PhyKIT: a broadly applicable UNIX shell toolkit for processing and analyzing phylogenomic data. Bioinformatics 37, 2325–2331 (2021).
Steenwyk, J. L., Buida, T. J., Li, Y., Shen, X.-X. & Rokas, A. ClipKIT: a multiple sequence alignment trimming software for accurate phylogenomic inference. PLoS Biol. 18, e3001007 (2020).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evolution 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinforma. 10, 421 (2009).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evolution 35, 518–522 (2018).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Shimodaira, H. An approximately unbiased test of phylogenetic tree selection. Syst. Biol. 51, 492–508 (2002).
Kishino, H., Miyata, T. & Hasegawa, M. Maximum likelihood inference of protein phylogeny and the origin of chloroplasts. J. Mol. Evol. 31, 151–160 (1990).
Kishino, H. & Hasegawa, M. Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in hominoidea. J. Mol. Evol. 29, 170–179 (1989).
Shimodaira, H. & Hasegawa, M. Multiple Comparisons of Log-Likelihoods with Applications to Phylogenetic Inference. Mol. Biol. Evolution 16, 1114–1116 (1999).
Strimmer, K. & Rambaut, A. Inferring confidence sets of possibly misspecified gene trees. Proc. Biol. Sci. 269, 137–142 (2002).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Steenwyk, J. L., Li, Y., Zhou, X., Shen, X.-X. & Rokas, A. Incongruence in the phylogenomics era. Nature Reviews Genetics https://doi.org/10.1038/s41576-023-00620-x (2023).
Tavaré, S. Some probabilistic and statistical problems in the analysis of DNA sequences. Lect. Math. Life Sci. (Am. Math. Soc.) 17, 57–86 (1986).
Yang, Z. Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J. Mol. Evol. 39, 306–314 (1994).
Steenwyk, J. L., Shen, X.-X., Lind, A. L., Goldman, G. H. & Rokas, A. A robust phylogenomic time tree for biotechnologically and medically important fungi in the genera Aspergillus and Penicillium. mBio. 10, e00925–19 (2019).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Buchfink, B., Ashkenazy, H., Reuter, K., Kennedy, J. A. & Drost, H.-G. Sensitive Clustering of Protein Sequences at Tree-of-Life Scale Using DIAMOND DeepClust. http://biorxiv.org/lookup/doi/10.1101/2023.01.24.525373 (2023) https://doi.org/10.1101/2023.01.24.525373.
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Mistry, J. et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Mi, H. et al. PANTHER version 16: a revised family classification, tree-based classification tool, enhancer regions and extensive API. Nucleic Acids Res. 49, D394–D403 (2021).
Haft, D. H., Selengut, J. D. & White, O. The TIGRFAMs database of protein families. Nucleic Acids Res. 31, 371–373 (2003).
Blin, K. et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 45, W36–W41 (2017).
Lind, A. L. et al. Examining the evolution of the regulatory circuit controlling secondary metabolism and development in the fungal genus Aspergillus. PLoS Genet 11, e1005096 (2015).
Kautsar, S. A. et al. MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res. 48, D454–D458 (2020).
Käfer, E. Meiotic and Mitotic Recombination in Aspergillus and Its Chromosomal Aberrations. in Advances in Genetics vol. 19 33–131 (Elsevier, 1977).
Mead, M. E. et al. Characterizing the pathogenic, genomic, and chemical traits of Aspergillus fischeri, a close relative of the major human fungal pathogen Aspergillus fumigatus. mSphere 4, e00018–e00019 (2019).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinforma. 11, 395 (2010).
Almeida, A. J. et al. Genome size and ploidy of Paracoccidioides brasiliensis reveals a haploid DNA content: Flow cytometry and GP43 sequence analysis. Fungal Genet. Biol. 44, 25–31 (2007).
Reis, Dos et al. Screening of chemical libraries for new antifungal drugs against aspergillus fumigatus reveals sphingolipids are involved in the mechanism of action of miltefosine. mBio. 12, e01458–21 (2021).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Vera Alvarez, R., Pongor, L. S., Mariño-Ramírez, L. & Landsman, D. TPMCalculator: one-step software to quantify mRNA abundance of genomic features. Bioinformatics 35, 1960–1962 (2019).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank the Rokas lab and Dr. Judith Berman for helpful discussion and feedback. J.L.S. and A.R. were funded by the Howard Hughes Medical Institute through the James H. Gilliam Fellowships for Advanced Study program. J.L.S. is a Howard Hughes Medical Institute Awardee of the Life Sciences Research Foundation. Research in A.R.’s lab is supported by grants from the National Science Foundation (DEB-2110404), the National Institutes of Health/National Institute of Allergy and Infectious Diseases (R01 AI153356), and the Burroughs Wellcome Fund. Research in G.H.G’s lab is supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) grants numbers 2016/07870-9 and 2021/04977-5 (G.H.G.) and the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) grant numbers 301058/2019-9 and 404735/2018-5 (G.H.G.), both from Brazil. Research in F.R.’s Lab was supported by the European Union’s Horizon 2020 research and innovation program under grant agreement no. 847507 (HDM-FUN). Research in D.C.’s Lab is supported by grants PID2021-128001OB-I00 and TED2021-129601B-I00 funded by MICIU/AEI/ 10.13039/501100011033, and by “ERDF A way of making Europe” and the “European Union NextGenerationEU/PRTR”. A.Z.S.’s lab is supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) grants numbers 2022/16198-3 and 2022/12553-3.
Author information
Authors and Affiliations
Contributions
Conceptualization: JLS, GHG, and AR. Methodology: JLS, SK, RWB, MEM, DC, RSL, KL, FR, NHO, XZ, GHG, and AR. Software: NA. Validation: JLS, SK, RWB, CB, DR, MEM, CDR, HAR, YL, ACC, PAdC, TFdR, MA-M, AA, DG-H, ED, LP, CFP, AZS, DC, RSL, KL, ET, FR, NHO, XZ, GHG, and AR. Formal analysis: JLS, SK, RWB, CB, DR, CDR, HAR, PAdC, TFdR, AG, MA-M, AA, DG-H, ED, LP, CFP, AZS, and ZH. Investigation: JLS, SK, RWB, CB, DR, MEM, CDR, HAR, YL, ACC, PAdC, TFdR, AG, MA-M, AA, DG-H, ED, LP, CFP, AZS, DC, RSL, KL, ET, FR, and ZH. Resources: DC, RSL, KL, ET, FR, NHO, XZ, GHG, AR. Data curation: JLS, SK, RWB, HAR, YL, AG, ED, KL, GHG, and AR. Writing – Original Draft: JLS, SK, NHO, XZ, GHG, and AR. Writing – Review & Editing: All authors. Visualization: JLS, SK, KL, GHG, and AR. Supervision: KL, NHO, GHG, and AR. Project administration: JLS, GHG, and AR. Funding acquisition: JLS, FR, DC, AZS, GHG, and AR.
Corresponding authors
Ethics declarations
Competing interests
J.L.S. held a scientific advisory position for WittGen Biotechnologies during a portion of the project. J.L.S. is an advisor for ForensisGroup Incorporated. A.R. is a scientific consultant for LifeMine Therapeutics, Inc. G.H.G. is a scientific consultant for Innovation Pharmaceuticals Inc. The other authors declare no other competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Steenwyk, J.L., Knowles, S., Bastos, R.W. et al. Evolutionary origin and population diversity of a cryptic hybrid pathogen. Nat Commun 15, 8412 (2024). https://doi.org/10.1038/s41467-024-52639-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52639-1