Abstract
Neoadjuvant immunochemotherapy (nICT) has dramatically changed the treatment landscape of operable esophageal squamous cell carcinoma (ESCC), but factors influencing tumor response to nICT are not well understood. Here, using single-cell RNA sequencing paired with T cell receptor sequencing, we profile tissues from ESCC patients accepting nICT treatment and characterize the tumor microenvironment context. CXCL13+CD8+ Tex cells, a subset of exhausted CD8+ T cells, are revealed to highly infiltrate in pre-treatment tumors and show prominent progenitor exhaustion phenotype in post-treatment samples from responders. We validate CXCL13+CD8+ Tex cells as a predictor of improved response to nICT and reveal CXCL13 to potentiate anti-PD-1 efficacy in vivo. Post-treatment tumors from non-responders are enriched for CXCL13+CD8+ Tex cells with notably remarkable exhaustion phenotype and TNFRSF4+CD4+ Tregs with activated immunosuppressive function and a significant clone expansion. Several critical markers for therapeutic resistance are also identified, including LRRC15+ fibroblasts and SPP1+ macrophages, which may recruit Tregs to form an immunosuppressive landscape. Overall, our findings unravel immune features of distinct therapeutic response to nICT treatment, providing a rationale for optimizing individualized neoadjuvant strategy in ESCC.
Similar content being viewed by others
Introduction
Esophageal cancer (EC) is the seventh most common malignancy and the sixth leading cause of cancer deaths worldwide1. Esophageal squamous cell carcinoma (ESCC), accounting for almost 90% of all EC cases, is highly malignant in nature and especially prevalent in east Asia2. Due to the inconspicuous nature of its symptoms, most ESCC patients are diagnosed at an advanced stage, contributing to a 5-year overall survival rate below 20%3. For patients with resectable locally advanced ESCC, neoadjuvant chemoradiotherapy (nCRT) followed by esophagectomy is currently the standard-of-care treatment, demonstrating improved survival compared to surgery alone4,5. More than 40% of ESCC patients achieve pathological complete response (pCR) after nCRT in the CROSS trial and the NEOCRTEC5010 trial6,7. Nevertheless, nCRT strategy is also reported to increase postoperative mortality risk8,9, and the rate of distant recurrence remains high4,10. There exists a critical need for more effective and safer treatments to improve clinical outcomes of patients with operable locally advanced ESCC.
Neoadjuvant immunotherapy, particularly immune checkpoint blockade (ICB), shows superior clinical benefits and shifts the treatment paradigm across a variety of solid tumors11. For advanced-metastatic ESCC, immunochemotherapy has been approved by the U.S. Food and Drug Administration as the first-line treatment12. Neoadjuvant immunochemotherapy (nICT) has shown promising benefits for operable locally advanced ESCC, achieving a rate of pCR up to 50% without increasing surgical complication rates13,14,15. But still, the current pCR rate of this approach remains suboptimal. To optimize nICT efficacy in ESCC, it is crucial to obtain a comprehensive understanding of the mechanisms underpinning sensitivity or resistance to nICT treatment.
The tumor microenvironment (TME) harbors a heterogeneous milieu of tumor cells, immune cells, extracellular matrix, secreted molecules, and lymphatic vascular networks16. The cellular and molecular components of TME are closely associated with tumor biology and sensitivity of ICB drugs17,18,19. Elucidating the underlying responsive mechanisms of TME is the key for comprehending cancer immunotherapy20,21. Both single-cell RNA sequencing (scRNA-seq) and single-cell T-cell receptor sequencing (scTCR-seq) are powerful techniques in characterizing the cellular biology and dynamics during tumor immune response22,23. Previous studies using single-cell transcriptome have highlighted the immunosuppressive characteristics of TME in treatment-naïve ESCC tumor tissues24,25,26,27, but the response of various TME cells in locally advanced ESCC to nICT treatment remains insufficiently illuminated.
The current study performs scRNA-seq paired with scTCR-seq to investigate the cellular biology and dynamics in 18 ESCC patients undergoing nICT treatment. Our findings indicate that CD8+ Tex-CXCL13 is remarkably enriched in responders’ pre-treatment tumors and shows more obvious progenitor-exhaustion phenotype in corresponding post-treatment samples. We reveal pre-treatment CXCL13+CD8+ Tex cells to be a predictor of improved response to nICT and demonstrate that CXCL13 can potentiate anti-PD-1 efficacy in ESCC. Tumors from non-responders are enriched for CD8+ Tex-CXCL13 and CD4+ Treg-TNFRSF4 in post-treatment samples, characterized by a terminally exhausted and immune-suppressed phenotype. Moreover, cell-cell interaction analysis suggests that LRRC15+ fibroblasts and SPP1+ macrophages may recruit CD4+ Tregs to form an immune-suppressed TME, and thus inducing treatment resistance. Collectively, our study elucidates the immune features of distinct response to nICT treatment, which has potential implications for potentiating anti-PD-1 efficacy in ESCC.
Results
TME was characterized by complex and heterogenous cell populations in locally advanced ESCC before and after nICT treatment
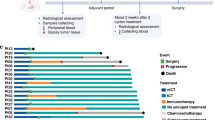
To delineate the TME features of distinct treatment response to nICT in ESCC, we analyzed a total of 7 pre-treatment biopsy and 16 post-treatment surgery samples from 18 ESCC patients receiving nICT treatment using scRNA-seq and scTCR-seq analyses (Fig. 1a). The clinicopathological characteristics of these patients are detailed in Supplementary Data 1. Pre- and post-treatment samples were obtained from endoscopic biopsy and at surgical resection, respectively, with successful matching achieved in 5 patients. ESCC patients undergoing nICT treatment were classified into responders, defined as patients with pathological complete response or major pathological response, and non-responders. Representative images reflected prominent radiological and endoscopic changes in tumors for both responders and non-responders (Fig. 1b).
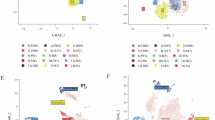
a Study design and workflow. Containing modified images from Servier Medical Art (https://smart.servier.com/), licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/). Created in BioRender. Wang, X. (2024) BioRender.com/a01w422. b Representative radiological images of responders and non-responders before and after neoadjuvant immunochemotherapy (nICT). The yellow arrows indicate the position of the tumors. pCR, pathological complete response. Non-MPR, not reach the major pathological response. c Uniform Manifold Approximation and Projection (UMAP) plots of high-quality cells (n = 128,600) from all enrolled samples (n = 23) visualizing cell-type clusters. d UMAP plot showing high-quality cells from all the samples annotated by patient origin. Samples, n = 23; patients, n = 18. e UMAP visualization of high-quality cells colored by treatment timepoints. Pre-treatment, n = 7; post-treatment, n = 16. f UMAP plot of high-quality cells from all the samples annotated by efficacy groups. Responder, n = 12; non-responder, n = 11. g Proportions of diverse cell types before and after nICT treatment. h Proportions of diverse cell types in responders and non-responders. i Expression patterns of signature genes in distinct cell clusters.
The scRNA-seq raw data of these 23 samples were then processed (Supplementary Fig. 1a–e, Supplementary Data 2). After excluding low-quality cells (Supplementary Fig. 2a, b), we acquired 128,600 high-quality cells, which were classified into 8 major cell types according to corresponding marker genes, including B cells, T cells, myeloid cells, mast cells, epithelial cells, fibroblasts, endothelial cells, smooth muscle cells and neurons (Fig. 1c). The distributions of various cell types across different patients, treatment timepoints and efficacy groups were displayed via Uniform Manifold Approximation and Projection (UMAP) visualization (Fig. 1d–f). Moreover, bar charts were applied to show the percentage of each cell type across different samples, timepoints and efficacy groups (Fig. 1g, h, Supplementary Fig. 2c, d). Each cell type was further clustered into multiple subpopulations according to the expression of canonical marker genes. As displayed, 11 clusters of T cells, 6 clusters of B cells, 12 clusters of fibroblasts, 10 clusters of myeloid cells, 6 clusters of endothelial cells and 2 clusters of epithelial cells were identified. Distinct molecular signatures of these cell subsets were summarized in the heatmap (Fig. 1i), indicating their unique cellular identities. These results revealed the complex and heterogeneous cell components in TME of both pre- and post-treatment ESCC samples.
Non-responders exhibit a highly elevated infiltration level of terminal CD8+ Tex cells along with an increased cell proportion of activated CD4+ Tregs
Further unsupervised analysis of preprocessed and normalized scRNA-seq data for T cells via UMAP revealed 11 T cell clusters, including CD8+ Tex-CXCL13, CD8+ Tex-STMN1, CD8+ Tem-GZMK, CD8+ Teff-FGFBP2, CD8+ Teff-TYROBP, CD8+ Tn-IL7R, CD8+ T-ISG15, CD4+ Treg-TNFRSF4, CD4+ Tfh/Th1-CXCL13, CD4+ Tn-CCR7 and CD4+ T-STMN1 (Fig. 2a). To identify each cluster, we constructed a dot plot of key marker genes for these various T cell clusters (Supplementary Fig. 3a). The distributions of these T cell clusters across efficacy groups and treatment timepoints were visualized through UMAP (Fig. 2b, c).
a Clustering of 27,594 T cells based on all enrolled samples (n = 23), colored and labeled by T cell clusters. b UMAP plot showing T cells colored by efficacy groups. Responder, n = 12; non-responder, n = 11. c UMAP visualization of T cells colored by treatment timepoints. Pre-treatment, n = 7; post-treatment, n = 16. d Fraction of CD8+ T cells in all CD45+ cells and fraction of CD8+ Tex-CXCL13, CD8+ Tex-STMN1, and CD8+ Tn-IL7R cells in all CD8+ T cells. Post-treatment samples from responder (Post-R), n = 6; Post-treatment samples from non-responder (Post-NR), n = 10. *p < 0.05 of Wilcoxon test. Boxes indicate median ± interquartile range; whiskers show minima and maxima. Detailed information of each sample is provided in the Source Data. e The difference in progenitor and terminal exhaustion score of CD8+ Tex-CXCL13 (left) and CD8+ Tex-STMN1 (right) between responders and non-responders. ***p < 0.0001 of Mann Whitney U test. Boxes indicate median ± interquartile range; whiskers represent 1.5-fold of the interquartile range. f Fraction of CD4+ T cells in all CD45+ cells and fraction of CD4+ Treg-TNFRSF4, CD4+ Tfh/Th1-CXCL13, and CD4+ Tn-CCR7 cells in all CD4+ T cells. **p < 0.01, ***p < 0.001 of Mann Whitney U test. Post-R, n = 6; Post-NR, n = 10. For each boxplot, the center line represents the median, the box indicates the upper and lower quartiles, and the whiskers represent minima and maxima. Source data is provided as a Source Data file. g The difference in infiltration score of CD8+ Tex-CXCL13 (left) and CD4+ Treg-TNFRSF4 (right) among pre-treatment samples of responders (Pre-R, n = 32) and non-responders (Pre-NR, n = 22), Post-R (matched with Pre-R, n = 32) and Post-NR (matched with Pre-NR, n = 22) from an independent nICT-ESCC cohort. *p < 0.05, **p < 0.01, ***p < 0.001 of Mann Whitney U test. Boxes indicate median ± interquartile range; whiskers represent 1.5-fold of the interquartile range. Detailed infiltration score of each sample is provided in the Source Data. h Representative images of multiplex IHC staining for CXCL13+CD8+ T cells from pre-treatment tumors of responders and non-responders in the independent validation cohort. Responders, n = 56; Non-responders, n = 40. Scale bar: 20 μm. i Quantification of CXCL13+CD8+ T cells by calculating cell density based on six visual fields per patient. **p < 0.01 of Mann Whitney U test. Responders, n = 56; Non-responders, n = 40. j Area under receiver operating characteristic (ROC) curves (AUC) for pre-treatment CD8+ Tex-CXCL13 signature or CD274 expression on prediction of MPR in validation cohort. Pre-R, n = 32; Pre-NR, n = 22. k Representative images of TLSs in responders (left) and non-responders (right). Scale bar: 1000 μm for general views, 200 μm for zoom views. l Difference of post-treatment tertiary lymphoid structure (TLS) abundance between responders (n = 56) and non-responders (n = 40) in external validation cohort. **p < 0.01 of Mann Whitney U test. m Scatterplots showing significant correlations between pre-treatment CXCL13+CD8+ T cell infiltration with post-treatment TLS abundance. Spearman-correlation test. Pre, n = 96; Post, n = 96.
We next explored the differences in cellular proportions in all T cell types between responders and non-responders (Supplementary Data 3). Among CD8+ T cells in post-treatment tissues, CD8+ Tex-CXCL13 and CD8+ Tex-STMN1 cells were revealed to highly infiltrate in non-responders compared with responders (Fig. 2d). Responders, on the other hand, were marked by an increased proportion of CD8+ Tn-IL7R cells in post-treatment tissues. Additionally, we also observed a notably elevated infiltration level of CD8+ Tn-IL7R cells in responders during nICT treatment (Supplementary Fig. 3b). Previous studies indicated that CD8+ Tex cells could be further classified into two phenotypes, progenitor Tex and terminal Tex, which were both closely associated with efficacy of immunotherapy28,29. Next, we calculated the progenitor and terminally exhausted signature of CD8+ Tex cells. Our results showed that CD8+ Tex-CXCL13 and CD8+ Tex-STMN1 cells in responders had prominently higher progenitor exhaustion score but lower terminal exhaustion score versus non-responders (Fig. 2e). For CD4+ T cells in post-treatment non-responders, we identified a highly increased cell proportion of CD4+ Treg-TNFRSF4 cells and an obvious decrease in the infiltration level of CD4+ Tn-CCR7 cells (Fig. 2f). In addition, CD4+ Treg-TNFRSF4 cells in post-treatment non-responders were proved to highly express the marker genes for functional activation of Tregs compared to responders (Supplementary Fig. 3d), indicating a more prominently immune-suppressive phenotype. Based on the RNA-seq data of 54 cases from an independent retrospective nICT-ESCC cohort, using ssGSEA algorithm, we validated an elevated infiltration level of CD8+ Tex-CXCL13, CD4+ Treg-TNFRSF4 and CD8+ Tex-STMN1 in post-treatment non-responders (Fig. 2g, Supplementary Fig. 3e). The cell abundance of CD4+ Tfh/Th1-CXCL13 cells increased prominently in both responders and non-responders during treatment (Supplementary Fig. 3c, f), suggesting the enhanced infiltration of CD4+ Tfh/Th1-CXCL13 cells as TME response to immunochemotherapy drugs regardless of treatment efficacy.
CD8+ Tex-CXCL13 cells are closely associated with clinical efficacy of immunotherapy in ESCC
Interestingly, in pre-treatment samples, a significant positive correlation between CXCL13 and CD274 in RNA expression levels was observed (Supplementary Fig. 3g), suggesting a potential predictive value of CXCL13 in immunotherapy. However, no significant difference in the cell proportion of CD4+ Tfh/Th1-CXCL13 cells was observed between responders and non-responders. In ESCC, CXCL13 was mainly expressed by CD4+ Tfh/Th1-CXCL13 and CD8+ Tex-CXCL13 cells (Supplementary Fig. 3h, i). Notably, the comparative analysis of CD8+ Tex-CXCL13 cell proportion cells in pre-treatment samples contradicted the findings in post-treatment samples. Compared with non-responders, responders exhibited a prominently higher infiltration score of CD8+ Tex-CXCL13 cells in pre-treatment tumors (Fig. 2g). Multiplex immunofluorescence of CXCL13 and CD8 was conducted in the pre-treatment samples from 96 ESCC patients who accepted nICT treatment (Fig. 2h). We observed significantly higher CXCL13/CD8 dual-positive cell abundance in responders versus non-responders (Fig. 2i). The subsequent ROC analysis based on the transcriptome profile of 54 pre-treatment ESCC patients demonstrated a similar C-index for predicting treatment response between CD274 expression and CD8+ Tex-CXCL13 infiltration (Fig. 2j), confirming the value of CD8+ Tex-CXCL13 cells for ESCC immunotherapy.
CXCL13 has been reported to be not only a marker for the exhausted state of CD8+ T cells30,31, but also an important chemokine controlling the recruitment and organization of B cells32,33. In this study, a total of six B cell clusters were identified according to reported marker genes, including Bmem_TNFSF13B, GCB_EGR3, Bfoc_TCL1A, Bn_IGHD, Bpro_STMN1, and PlasmaB_MZB1 (Supplementary Fig. 4a–c). Compared with non-responders, responders were shown with increased cell proportion of Bmem_TNFSF13B cells in post-treatment specimens (Supplementary Fig. 4d–f). The following CIBERSORT analysis based on RNA-seq data confirmed the highly elevated cell proportion of memory B cells in responders’ post-treatment samples (Supplementary Fig. 4g). The results of CellChat indicated a complex interaction network between T cell clusters and B cell clusters in both responders and non-responders (Supplementary Fig. 4h, i). Moreover, we also observed a high likelihood of interaction between CD8+ Tex-CXCL13 cells expressing CXCL13 and B cells expressing CXCR5 (Supplementary Fig. 4j, k).
Considering the crucial role of tertiary lymphoid structures (TLSs) in anti-tumor immunity, which were formed by tumor-infiltrating B cells organizing in aggregates surrounded by a mantle of T cells34, we next examined the abundance of TLSs in 96 post-treatment FFPE ESCC samples (Fig. 2k). As a result, TLSs were found to be more frequent in the post-treatment tumor tissues of responders (Fig. 2l), which validated the favorable role of TLSs during the process of nICT in ESCC. Notably, a statistically positive association between the abundance of post-treatment TLSs and the cell proportion of pre-treatment CD8+ Tex-CXCL13 cells was identified (Fig. 2m). These results implied that CD8+ Tex-CXCL13 cells might enhance efficacy of nICT in ESCC through recruiting TLSs.
CXCL13 enhances the efficacy of anti-PD-1 therapy in vivo
As CXCL13 was correlated with enhanced efficacy in ESCC immunotherapy, we proceeded to explore the effects of CXCL13 on the response to checkpoint inhibitors in vivo. The animal experiment was initiated when the tumor volume reached ~100 mm3, and continued for up to 30 days (Fig. 3a). Treatment of subcutaneous ESCC mice with a combination of CXCL13 and anti-PD-1 significantly inhibited tumor growth. After eight injections, the tumor volume and weight of the combinational treatment group were significantly lower than the monotherapy groups (Fig. 3b-d). Moreover, we performed flow cytometric analysis to assess the proportion of B cells and CD8+ T cells in tumors from different groups (Supplementary Fig. 5a). The percentages of CD20+ cells, CD8+ cells, and PD-1+ cells were significantly higher in the TME of the combinational treatment group (Fig. 3e–g). H & E staining of tumor tissues revealed a higher rate of pathological response in the combinational treatment group, confirming the superior anti-tumor effects of the combinational strategy (Fig. 3h). Besides, multiplex immunofluorescence of CD4 and CD8 on tumor tissues validated the increased presence of infiltrating immune cells in the TME of the combinational treatment group (Fig. 3i). No significant changes body weight or organ toxicity was observed during the entire process (Supplementary Fig. 5b, c). Taken together, these results suggested that CXCL13 might enhance the efficacy of the anti-PD-1 regimen by facilitating a more active TME.
a The design and workflow of in vivo experiments. Containing modified images from Servier Medical Art (https://smart.servier.com/), licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/). Created in BioRender. Wang, X. (2024) BioRender.com/h07a127. b Representative images of the subcutaneous tumor models treated with saline or CXCL13 or anti-PD-1 or a combination of CXCL13 and anti-PD-1. c Growth curves of the subcutaneous tumors among the indicated four groups. Short horizontal lines indicate mean ± SD (n = 5 per group). **p < 0.01, ***p < 0.001 of two-sided t-test. d Histograms showing the tumor weights of the subcutaneous tumors among the indicated four groups (n = 5 per group). ***p < 0.001 of two-sided t-test. Percentages of CD20+ cells (e), CD3+CD8+ cells (f), and PD-1+ cells (g) in the subcutaneous tumors among the indicated four groups (n = 5 per group). The corresponding quantification histograms were displayed. ***p < 0.001 of two-sided t-test. h Representative HE staining images of the subcutaneous tumors among the indicated four groups (n = 5 per group). Scale bar: 1000 or 2000 μm. i Representative images of multiplex fluorescence showing CD4+ and CD8+ cell infiltration among the indicated four groups (n = 5 per group). Scale bar: 50 μm. CD4, red. CD8, green. DAPI, blue.
Non-responders are marked by significantly higher clonal expansion of CD4+ Tregs and lower clonal expansion of CD8+ Teff cells
To delineate the relationship between the characteristics of T cells and the response to nICT treatment, we next investigated changes in clonality across different time points during immunotherapy and between groups with distinct efficacy through scTCR-seq analysis. Considering the broader expression of multiple dysfunctional state marker genes of T cells in CD8+ Tex-CXCL13, including LAG3, PDCD1, CTLA4, and HAVCR2 (Supplementary Fig. 3a), we defined CD8+ Tex-CXCL13 cells as terminal CD8+ Tex. We recovered the α chains and β chains of TCR from ESCC samples at single-cell resolution (Supplementary Data 6). T cells with TCR data across patients, treatment time point, clonotypes and clone size were shown in the UMAP plots (Fig. 4a–d). We examined the clonal expansion patterns of different T cell subtypes in ESCC during immunochemotherapy, in which we observed higher percentages of high-expansion clones (n ≥ 3 cells) were mainly in CD8+ Tex and Teff cells (Fig. 4e–g, Supplementary Fig. 6a). This suggests the critical role of CD8+ Tex and Teff cells as tumor antigen-specific CD8+ T cells. Through comparing scTCR-seq data of matched pre-treatment and post-treatment samples, we found that pre-existing TCR clone types of CD8+ T cells sustained and accounted for the main part during treatment, and novel TCR clone types appeared after nICT therapy (Supplementary Fig. 6b). A similar pattern was observed in CD8+ Tex cells, suggesting their involvement in mediating therapeutic response.
a–d UMAP plots of 16,636 T cells with detected T cell receptor beta (TRB) clones from 23 samples, annotated by patient origin, treatment timepoints, clonotypes and clone size. e–g Fractions of TRB clones with different sizes (n < 3 and n ≥ 3) for different T cell clusters in pre-treatment samples (Pre, left), post-treatment samples of responders (Post-R, middle), and non-responders (Post-NR, right). h Boxplots showing Gini indices for CD4+ Tregs in different groups. Box middle lines, median; box limits, upper and lower quartiles; box whiskers, minima and maxima. *p < 0.05, **p < 0.01 of Mann Whitney U test. Pre, n = 7; Post-R, n = 6; Post-NR, n = 10. i Heatmap showing the fraction of CD8+ T cell clonotypes shared with a secondary phenotype (column) in each primary phenotype (row). Pre (left); Post-R (middle); Post-NR (right). j UMAP plot of CD8+ Tex-relevant and Tex-irrelevant cells in all samples (n = 23). k UMAP plot showing CD8+ terminal Tex and progenitor Tex (Texp) cells in all samples (n = 23). l RNA velocity analysis of CD8+ Tex clusters based on Dynamo. Arrows in the plot illustrating potential differentiation paths. m Fraction of CD8+ terminal Tex and Texp cells in CD8+ Tex-relevant cells for pre- and post-treatment responders and non-responders.
Importantly, non-responders revealed higher clonal expansion of CD4+ Tregs and lower clonal expansion of CD8+ Teff cells versus responders (Fig. 4e–g), indicating that increased tolerance by CD4+ Tregs and insufficient recognition of tumor antigen in CD8+ Teff cells may contribute to resistance to nICT treatment. The Gini index of clonal expansion of CD4+ Tregs in post-treatment non-responders was also identified to be significantly higher than responders (Fig. 4h), which was consistent with the finding that CD4+ Tregs of non-responders were highly activated. For responders, the clonal expansion of CD4+ Tregs obviously decreased in post-treatment samples compared to pre-treatment tissues. Further, to trace the lineage transitions between T cell phenotypes, we measured the fraction of clonotypes shared with a secondary phenotype in each primary phenotype. We observed significant clonotype overlaps between CD8+ T cell clusters, and the phenotype transitions were more active in post-treatment responders (Fig. 4i), especially the transition between CD8+ Tem-GZMK or CD8+ Tex-CXCL13 and CD8+ Teff-TYROBP. Moreover, CD8+ Tex shared clones with several CD8+ T cell subtypes, especially CD8+ Tem (Supplementary Fig. 6c, d), suggesting the existence of lineage transition between CD8+ Tem and CD8+ Tex35.
We selected all CD8+ T cells that shared TCRs with CD8+ Tex-CXCL13 (Fig. 4j) and referred to these T cells as Tex-relevant cells. Among the Tex-relevant cells, those not included in CD8+ Tex-CXCL13 cells (terminal Tex) were named as precursor exhausted T cells (Texp) (Fig. 4k). Texp exhibits a prominently higher expression level of marker genes enhancing anti-tumor activity of CD8+ T cells (Supplementary Fig. 6e), like CCL4 (chemoattractant for CD8+ T cells)36 and GZMK (marker of cytolytic T-cell activity). In contrast, terminal Tex was mainly marked by increased expression of checkpoint molecules, including LAG3, HAVCR2, CTLA4, and CXCL13. Importantly, the reported tumor antigen-specific T cell marker ENTPD1 (CD39) was also found to be highly expressed in terminal Tex37,38, suggesting the anti-tumor function of this group to be highly activated and exhausted. The result of GO analysis based on the upregulated DEGs of terminal Tex compared to Texp further validated the functional state of terminal Tex (Supplementary Fig. 6f). Furthermore, RNA velocity analysis on Tex-relevant cells revealed a potential transition route from Texp to terminal Tex (Fig. 4l). Our results also showed that the proportion of Texp cells in pre-treatment non-responder samples was lower than responders (Fig. 4m). Intriguingly, this proportion decreased along with an increase of terminal Tex in responders during nICT treatment. These findings indicated that the sustained high infiltration level of Texp might be critical for improved efficacy of immunotherapy for ESCC patients.
Post-treatment non-responders have a remarkable enrichment of tCAF-MMP11
Accumulating evidences have reported that cancer-associated fibroblasts (CAFs) play a key part in TME remodeling and response to cancer immunotherapy39,40,41. However, the dynamics of CAFs in ESCC during immunotherapy still lack of comprehensive exploration. Herein, we investigated the distribution of CAFs underlying distinct treatment responses on the basis of scRNA-seq data. Using previous reported marker genes, CAFs were annotated to 12 cell clusters (Fig. 5a, b), among which iCAF-CFD, iCAF-CXCL12, and iCAF-PLA2G2A were found to be highly enriched in post-treatment samples from responders, and tCAF-MMP11 was proved to remarkably increase in non-responders (Fig. 5c). Additionally, our CellChat analysis revealed an extensive and complex cell-cell communication network among various CAFs and diverse T cell clusters in post-treatment samples of both responders and non-responders (Supplementary Fig. 7a, b), suggesting a potential multi-aspect role of CAFs during nICT treatment. CAFs have been proved to be closely associated with the immunosuppressive functional state of Tregs42. The compositional single-cell data analysis using scCODA further confirmed the similar differential distribution of tCAFs and Tregs between post-treatment responders and non-responders mentioned above (Supplementary Fig. 8f). Our next focal point was the relationship between these two clusters. Notably, a statistically positive correlation was found between the cell proportion of tCAFs and that of Tregs (Fig. 5d). The results of CellChat analysis indicate a comparable number of ligand-receptor pairs between tCAF-MMP11 and CD4+ Treg-TNFRSF4 in different efficacy groups (Supplementary Fig. 7c, d). Potential cell-cell interactions were commonly observed in both responders and non-responders. Focusing on ligand-receptor pairs, in non-responders, tCAF-MMP11 produced LGALS9 as specific ligands with rich regulatory potential to induce the immunosuppressive function of CD4+ Treg-TNFRSF4 through CD44 (Supplementary Fig. 7e, f), as the function of LGALS9-CD44 signaling pathway was previous reported to enhance stability and function of adaptive CD4+ Tregs43. In our study, tCAF-MMP11 was shown to highly express LGALS9 versus other CAF clusters (Fig. 5e), and the expression level of LGALS9 was prominently higher in post-treatment non-responders compared to responders (Fig. 5f), which further validated our hypothesis.
a UMAP plot showing 13,612 CAFs colored and labeled by distinct cell clusters based on all enrolled samples (n = 23). b Bubble plot showing differential expression of marker genes for various CAF clusters. c Proportion of iCAF-CFD, iCAF-CXCL12, iCAF-PLA2G2A and tCAF-MMP11 in all CAFs. **p < 0.01 of Wilcoxon test. Post-treatment samples from responder (Post-R), n = 6; Post-treatment samples from non-responder (Post-NR), n = 10. For each boxplot, the center line represents the median, the box indicates the upper and lower quartiles, and the whiskers show minima and maxima. Detailed information of each sample in provided in the Source Data. d Scatterplot showing a significant correlation between post-treatment cell proportion of CD4+ Treg-TNFRSF4 in CD4+ T cells with that of tCAF-MMP11 in CAFs. Spearman-correlation test. Post-treatment samples, n = 16. e UMAP plot showing the distribution of FAP, MMP11, LRRC15 and LGALS9 in CAFs. f Differential expression of selected genes in tCAF-MMP11. ***p < 0.001 of two-sided Wilcoxon test. g The differential mRNA expression level of LRRC15 among distinct timepoints and efficacy groups based on the external validation cohort. ***p < 0.001 of Wilcoxon test. Pre-treatment samples of responders (Pre-R, n = 32) and non-responders (Pre-NR, n = 22); Post-R (matched with Pre-R, n = 32) and Post-NR (matched with Pre-NR, n = 22). The box indicates median ± interquartile range; whiskers represent minima and maxima. Source data are provided as a Source Data file. h The difference in infiltration score of tCAF-MMP11 among distinct timepoints and efficacy groups of the validation cohort. ***p < 0.001 of Wilcoxon test. Pre-R, n = 32; Pre-NR, n = 22; Post-R, n = 32; Post-NR, n = 22. The box indicates median ± interquartile range; whiskers represent minima and maxima. Detailed infiltration score of each sample in provided in the Source Data. i Scatterplot of correlation between the infiltration level of CD4+ Treg-TNFRSF4 with that of tCAF-MMP11 in pre-treatment samples. Spearman-correlation test. Pre-treatment samples, n = 54. j Scatterplot showing a significant correlation between the infiltration score of CD4+ Treg-TNFRSF4 with that of tCAF-MMP11 in post-treatment tissues. Spearman-correlation test. Post-treatment samples, n = 54. k Representative images of multiplex IHC staining for α-SMA+LRRC15+ CAFs and CD4+FOXP3+ T cells from post-treatment tumors of responders and non-responders. Post-R, n = 12; Post-NR, n = 15. Scale bar: 10 μm. l Cell density of LRRC15+ CAFs, CD4+FOXP3+ cells, CD4+ cells, and CD4+FOXP3- cells in post-treatment samples from responders (n = 12) and non-responders (n = 15). *p < 0.05, ***p < 0.001 of two-sided t-test. m Cell number of CD4+FOXP3+ cells at different distances from LRRC15+ CAFs. n Comparison of average distance from CD4+FOXP3+ cells to LRRC15+ CAFs between post-treatment responders and non-responders. **p < 0.01 of two-sided t-test. Post-R, n = 12; Post-NR, n = 15.
Importantly, we also found the molecule LRRC15 to be a specific marker for tCAF-MMP11, which was highly expressed in tCAF-MMP11, especially those from non-responders (Fig. 5e, f). The activation state of tCAF-MMP11 in post-treatment non-responders was more obvious with elevated expression level of FAP, MMP11 and PDPN (Fig. 5f). Based on the RNA-seq data of our independent ESCC cohort, we further confirmed the findings from scRNA-seq analysis, and revealed the similar dynamics of LRRC15 expression and tCAF-MMP11 infiltration during nICT treatment. LRRC15 expression decreased in responders’ post-treatment specimens versus pre-treatment tissues, but no significant changes in non-responders (Fig. 5g, h). In consistent with previous studies, LRRC15+ CAFs were found to limit responsiveness to immune checkpoint blockade44. As for mRNA expression in bulk RNA-seq data, LRRC15 was revealed to positively correlate with marker genes of immunosuppressive CD4+ Tregs in post-treatment samples, but no statistically significant correlation was observed in pre-treatment samples (Supplementary Fig. 7g, h). Further ssGSEA analysis of tCAF-MMP11 and CD4+ Treg-TNFRSF4 presented comparable results (Fig. 5i, j), indicating the specific interaction between these two clusters only happened during immunochemotherapy. Through performing multiple immunofluorescence assay on markers of tCAFs (α-SMA+LRRC15+ cells) and CD4+ Tregs (CD4+FOXP3+ cells), we confirmed this association in spatial distribution (Fig. 5k). Consistently, compared with responders, we observed a simultaneous enrichment of LRRC15+ CAFs and CD4+FOXP3+ cells in post-treatment tumors of non-responders (Fig. 5l). Nevertheless, no significant difference was observed in the infiltration level of CD4+ and CD4+FOXP3+ cells between two efficacy groups, indicating that LRRC15+ CAFs selectively recruited CD4+FOXP3+ cells and formed an immunosuppressive environment in non-responders. We then investigated the distance of CD4+FOXP3+ cells to LRRC15+ CAFs and calculated the average distance in distinct efficacy groups. In post-treatment non-responders, CD4+FOXP3+ cells mainly located at a distance of 5–15 μm from surrounding LRRC15+ CAFs (Fig. 5m), much closer than that in samples sensitive to nICT treatment (Fig. 5n). Collectively, our results indicated that high infiltration of tCAF-MMP11 strongly correlated with the activation and enrichment of CD4+ Treg-TNFRSF4 cells, thus limiting the efficacy of nICT treatment in ESCC patients. The interaction network between tCAF-MMP11 and CD4+ Treg-TNFRSF4 cells driven by LGALS9-CD44 might be potential therapeutic target to enhance immunotherapy efficacy in ESCC.
SPP1+ macrophages and tumor cells highly expressing SPP1 are markers for immunotherapy resistance in post-treatment ESCC
Macrophages, representing a key component of TME, have been reported to promote tumor immune evasion via targeting receptors, transcription factors, or secreted proteins involved in downstream/upstream signaling pathways45. The phenotypic diversity of macrophages in solid tumors necessitates a comprehensive exploration of distinct subpopulations. Eight subclusters of macrophages were identified through scRNA-seq data (Fig. 6a), namely Macro-EREG, Macro-SPP1, Macro-S100A8, Macro-SLC40A1, Macro-CXCL10, Macro-VCAN, Macro-APOE, Macro-STMN1. The distribution of various myeloid clusters across different efficacy groups and treatment timepoints were studied and displayed (Fig. 6b, c). Of note, Macro-SPP1, Macro-EREG and Macro-STMN1 were observed with prominent enrichment in post-treatment non-responders compared to responders in ESCC accepting nICT treatment (Fig. 6d, Supplementary Fig. 8a). Additionally, scCODA analysis also showed a relatively higher infiltrating level of Macro-SPP1, Macro-EREG and Macro-STMN1 (Supplementary Fig. 8f). While the subpopulation Macro-SLC40A1 showed an opposite distribution. Accumulating evidences have proved that SPP1+ macrophages could promote tumor progression, mediate TME remodeling and limit response to PD-L1 blockade46,47. Consistently, upregulated DEGs of Macro-SPP1 showed a significant enrichment in signaling pathways associated with TME remodeling including cytokine/chemokine mediated signaling pathways and leukocyte migration (Fig. 6e). Cell-cell interaction analysis revealed a diverse communication network between Macro-SPP1 and various T cell clusters in both post-treatment responders and non-responders (Fig. 6f).
a. Clustering of 9703 myeloid cells in all enrolled samples (n = 23), colored and labeled by cell clusters. b UMAP plot showing myeloid cells colored by efficacy groups. Responder, n = 12; non-responder, n = 11. c UMAP visualization of myeloid cells colored by treatment timepoints. Pre-treatment, n = 6; post-treatment, n = 17. d Proportion of Macro-EREG, Macro-SPP1, Macro-SLC40A1 and Macro-STMN1 in all macrophages. *p < 0.05, ***p < 0.001 of Wilcoxon test. Post-treatment samples from responder (Post-R), n = 6; Post-treatment samples from non-responder (Post-NR), n = 10. For each boxplot, the center line represents the median, the box indicates the upper and lower quartiles, and the whiskers show minima and maxima. Source data are provided as a Source Data file. e Significant enrichment of differentially expressed marker genes of Macro-SPP1 in Gene Ontology terms. f Number of significant ligand-receptor pairs calculated via CellChat analysis from Macro-SPP1 cells to T cell subsets in post-treatment responders (left) and non-responsive tumors (right). g Heatmap displaying the activity and regulatory potential of the prioritized ligands in Macro-SPP1 driving corresponding receptors of T cells in post-treatment samples. h Scatterplot of correlation between post-treatment cell proportion of CD4+ Treg-TNFRSF4 in CD4+ T cells with that of Macro-SPP1 in macrophages. Spearman-correlation test. Post-treatment samples, n = 16. i The difference in infiltration score of Macro-SPP1 among distinct timepoints and efficacy groups of the validation cohort. ***p < 0.0001 of Mann Whitney U test. Pre-treatment samples of responders (Pre-R, n = 32) and non-responders (Pre-NR, n = 22); Post-R (matched with Pre-R, n = 32) and Post-NR (matched with Pre-NR, n = 22). Boxes indicate median ± interquartile range; whiskers represent 1.5-fold of the interquartile range. Source data are provided as a Source Data file. j Scatterplots showing correlation between the infiltration score of CD4+ Treg-TNFRSF4 with that of Macro-SPP1 in pre-treatment tumors (left) and post-treatment tissues (right), respectively. Pre-treatment samples, n = 54; Post-treatment samples, n = 54. k Number of significant ligand-receptor pairs from malignant epithelial cells to T cell subsets in post-treatment responders (left) and non-responsive tumors (right). l Heatmap displaying the activity and regulatory potential of the prioritized ligands in malignant epithelial cells driving corresponding receptors of T cells from post-treatment samples. m Area under ROC curves for post-treatment SPP1 expression on prediction of therapeutic response in FFPE validation cohort. Post-treatment samples, n = 54.
In non-responders, CCL20 was shown to be a potential specific ligand mediating the cell-cell interaction of Macro-SPP1 with CD4+ T cells, especially CD4+ Treg-TNFRSF4, with CCR6 as the targeted receptor (Fig. 6g, Supplementary Fig. 8d, e). In addition, the ligand-receptor pair SPP1-CD44, which was associated with impairment of cytotoxic T lymphocyte (CTL)48, was observed as a specific communication pathway from Macro-SPP1 to various T cell clusters for non-responders. What’s more, the cell proportion of Macro-SPP1 statistically negatively correlated with that of CD4+ Tn-CCR7 and CD8+ Tn-IL7R (Supplementary Fig. 8b, c), which were thought to play critical roles in anti-tumor immunity. We found that the cell proportion of Macro-SPP1 seemed to be positively associated with CD4+ Treg-TNFRSF4 in post-treatment samples (Fig. 6h). Through comparing the ssGSEA score of Macro-SPP1 between two efficacy groups, a similar difference was observed, with the infiltration level of Macro-SPP1 in responders decreasing prominently during nICT treatment and remaining significantly lower compared to non-responders after treatment (Fig. 6i). Notably, a significantly positive association between Macro-SPP1 and CD4+ Treg-TNFRSF4 was found in post-treatment ESCC, yet not in pre-treatment samples, which indicated that the interaction between these two clusters might be induced by immunochemotherapy (Fig. 6j). These findings revealed Macro-SPP1 to be a critical marker for therapeutic resistance to immunotherapy in ESCC, which might activate Tregs and suppress CTLs through cell-cell interaction.
We next performed clustering of epithelial cells using UMAP (Supplementary Fig. 9a), and inferred the CNV for each patient using CopyKAT to identify malignant cells (Supplementary Fig. 9b). To further portray the features of malignant cells during ESCC immunotherapy, DEGs of malignant cells between responders and non-responders were identified (Supplementary Data 7). GO analysis for the upregulated DEGs of malignant cells in post-treatment non-responders showed a prominent enrichment in regulation of apoptotic signaling pathways and response to oxidative/chemical stress (Supplementary Fig. 9c), indicating resistance to apoptosis induced by cytotoxic T cells and platinum-based chemotherapy drugs. Interestingly, SPP1 was also found to be highly expressed in residual tumor cells of post-treatment non-responders compared to responders (Supplementary Data 7). Residual ESCC tumor cells of non-responders were predicted to exhibit enhanced interaction potential with CD8+ Tn, CD8+ Tex and CD8+ Tem cells (Fig. 6k). In addition, SPP1 derived from residual ESCC tumor cells of non-responders was predicted to strongly regulate and suppress CD8+ T cell activation and confer host tumor immune tolerance through targeting CD44 (Fig. 6l). In the TCGA-ESCA cohort, high expression level of SPP1 emerged as a significant prognostic risk factor (Supplementary Fig. 9d). In our ESCC-nICT cohort, SPP1 expression of post-treatment samples was also an important predictor for response to immunochemotherapy (Fig. 6m). Collectively, both SPP1+ macrophages and tumor cells highly expressing SPP1 were markers for resistance to immunotherapy in post-treatment ESCC. SPP1, apart from being a crucial predictor of immunotherapy response, also interacts with cytotoxic CD8+ T cells within TME, ultimately limiting antitumor immunity. These findings highlighted the potential value of SPP1 as a therapeutic target to overcome immune suppression and increase the response of ESCC tumors to immunotherapies.
Discussion
Neoadjuvant immunochemotherapy (nICT) holds promise for treating locally advanced ESCC. However, not all ESCC patients could benefit from nICT treatment, due to primary sensitiveness or resistance to immunochemotherapy, the reason for which has not been fully understood. Our study used scRNA-seq paired with scTCR-seq to explore the cellular dynamics and biological features of ESCC during nICT treatment, which might shed light on therapeutic strategies against resistance to nICT treatment in ESCC patients.
Our study uncovered that terminal exhaustion of CD8+ Tex, along with high tolerance caused by CD4+ Tregs, are major contributors to therapeutic resistance in ESCC immunotherapy. We found post-treatment non-responders were marked by a prominently higher infiltration level of CD8+ Tex-CXCL13. CD8+ Tex has been proved to differentiate from CD8+ T cells accepting persistent tumor antigen stimulation, which is functionally exhausted with upregulated inhibitory receptors and impaired immune memory49. While immune checkpoint blockade, such as PD-1/PD-L1 inhibitors, can enhance tumor control via reinvigorating early-stage CD8+ exhausted T cells and potentiating polyfunctionality50, terminal CD8+ Tex may lack reprogrammability and responsiveness to PD-1/PD-L1 inhibitors51. Compared with responders, CD8+ Tex of post-treatment non-responders had significantly higher terminal exhaustion score but lower precursor exhaustion score, suggesting limited capacity for restoring the anti-tumor effects when exposed to immunotherapy. In TCR analysis, we noticed that the CD8+ Tex of post-treatment non-responders could barely transform into non-exhausted phenotype, especially effector CD8+ T cells. This implied that the major effector cells in non-responders after treatment might be newly recruited or expanded T cells with limited reinvigoration capacity, rather than the pre-existing CD8+ Tex. Altogether, the terminal exhaustion of CD8+ Tex may be one of the prevailing reasons behind the lower efficacy of immunochemotherapy in locally advanced ESCC.
Of note, post-treatment non-responders showed a notable increase in CD4+ Treg-TNFRSF4. Immunosuppressive CD4+ Tregs have been recognized to closely correlate with tumor progression and patients’ prognosis in ESCC52,53, and provide a main mechanism of tumor immune evasion54. Our results showed that CD4+ Treg-TNFRSF4 had higher expression of marker genes for immunosuppressive phenotype in samples with therapeutic resistance. Moreover, TCR analysis revealed that CD4+ Tregs of post-treatment non-responders had more diverse TCR clone types and higher Gini index for TCR clone type versus responders. All these findings identified a highly suppressive immune feature driven by increased CD4+ Tregs in ESCC samples resistant to immunotherapy. It is believed that removing the Treg roadblock is necessary for more effective nICT treatment in ESCC. Future researches on merging anti-Treg and immune checkpoint blockade for ESCC are needed.
Equally interestingly, CD8+ Tex-CXCL13 was found to deeply infiltrate in pre-treatment samples of responders. CXCL13 is known as a marker for dysfunctional state of CD8+ T cells30,31,55, while CXCL13+CD8+ T cells correlate with superior response to immunotherapy in multiple cancer types17,56. Consistently, our study found a positive correlation between CXCL13+CD8+ T cells’ infiltration and efficacy of nICT treatment in ESCC, suggesting CXCL13+CD8+ T cells’ potential to be predictors for effectiveness of nICT treatment. CXCL13 could be secreted into the TME by multiple cell types57, mediating immune cell recruitment58. CXCL13-proucing tumor-reactive T cells have been proved to play a central role in the maturation and distribution of tumor-associated TLSs59. CXCL13-dependent TLS formation and maturation potentiate anti-tumor immunity during immunotherapy33,60. In consistent with previous discoveries in other solid tumor61, we identified increased clustering of Bmem-TNFSF13B and GCB-EGR3 cells in post-treatment responders, which were important components of TLSs59. Previous research has linked TLSs with a favorable prognosis in treatment-naïve ESCC62, while our study filled the gap of validating higher TLS abundance in post-treatment responders and a positive association between baseline infiltrating level CXCL13+CD8+ T cells and post-treatment TLS abundance. In addition, our cell-cell interaction analysis verified the communication network between CD8+ Tex-CXCL13 and Bmem-TNFRSF13B/Bfoc-TCL1A mediated by CXCL13-CXCR5.
In conclusion, the findings in this study suggest that baseline enrichment of CD8+ Tex-CXCL13 might sensitize ESCC tumor to immunotherapy through inducing TLS formation and maturation. Furthermore, post-treatment responders exhibited a higher proportion of CD4+ Tfh/Th1-CXCL13 cells, another main cellular group of T cells expressing CXCL13. This further supports the potential of CXCL13 as a target to sensitize ESCC tumor to immunotherapy. More importantly, it is found that rmCXCL13 could significantly enhance the efficacy of anti-PD-1 therapy in subcutaneous ESCC mouse model. CXCL13 has been identified as a biomarker of germinal center activity, where the B cell antibody responses were optimized63. In line with this, we observed an increased infiltration of CD20+ cells in the TME of the treatment groups exposed to CXCL13 and increased CD8+ cells in the TME of the treatment groups exposed to anti-PD-1 therapy, indicating that these cell types could mediate the enhancement of CXCL13 on ESCC immunotherapy. Our findings shed light on the potential clinical value of combining CXCL13 with anti-PD-1 therapy in ESCC.
It is currently accepted that CAFs can form a trap to restrict infiltration and anti-tumor activity of CD8+ T cells in tumor nests, and thus regulating tumor immunity and progression40,64,65. In the present study, the elevated tCAF-MMP11 identified in post-treatment non-responders is found to be closely correlated with immunotherapy resistance, exhibiting a statistically positive association with the cell proportion of CD4+ Treg-TNFRSF4 cells. Furthermore, tCAF-MMP11 was marked by higher expression of LRRC15, which was found to be only expressed by fibroblasts. LRRC15+ CAFs was previously reported to limit anti-PD-L1 responsiveness44. Consistently, our spatial positioning analysis revealed that LRRC15+ CAFs to be highly infiltrated in ESCC tumors with primary resistance to nICT treatment, and specifically recruited CD4+ Tregs to form a highly tolerant immune feature. While Krishnamurty et al. reported LRRC15+ CAFs to limit immunotherapy efficacy through directly inhibiting CD8+ T cell effector function66, our findings showed that LRRC15+ CAFs could recruit and activate immunosuppressive CD4+ Tregs, during which LGALS9-CD44 ligand-receptor pair mediating cell-cell interaction might be a notable mechanism43. Selectively removing LRRC15+ CAFs or impairing the recruitment of CD4+ Tregs could be an attractive therapeutic option for non-responders to restore the anti-tumor effects when exposed to immunotherapy.
Of additional interest, analysis of macrophages and residual malignant epithelial cells indicated that both SPP1+ macrophages and tumor cells highly expressing SPP1 were critical markers for resistance to immunotherapy in ESCC. SPP1+ macrophages have been recognized to be linked to the progression of multiple human cancers with noticeably strong prognostic association46,67. Further CellChat analysis showed that SPP1 expressed by SPP1+ macrophages and residual ESCC tumor cells might act as an immune checkpoint to suppress CD8+ T cell activation through targeting at CD44, which was consistent with previous findings in colon cancer48. Additionally, SPP1+ macrophages in post-treatment non-responders were also revealed with a potential to recruit CD4+ Tregs through CCL20-CCR6 interaction, an important chemokine-receptor signaling during the migration of CD4+ Tregs68,69. Moreover, SPP1 secreted by tumor cells was indicated to be a strong chemokine to recruit suppressive myeloid cells and trigger the polarization of macrophages to M2-phenotype tumor-associated macrophages70,71. These findings uncover a coherent SPP1-mediated TME response that controls human cancers and affects immunotherapy efficacy. Therapeutically, the use of SPP1 inhibitors might improve treatment response in ESCC immunotherapy, though the clinical benefits require further investigation.
In summary, this study employed a combination of scRNA-seq and scTCR-seq to comprehensively characterize the immune landscape of therapeutic response to nICT treatment in locally advanced ESCC. The findings shed light on the cellular dynamics and molecular features of ESCC during immunotherapy, and highlight the potential of CXCL13 as adjuvant for better immunotherapy efficacy. We also identified the enrichment of LRRC15+ CAFs and SPP1+ macrophages as critical markers for resistance to immunotherapy. These conclusions could provide a rationale for future studies on optimal strategies to improve immunotherapy response in ESCC.
Methods
Ethics approval and consent to participate
This study was approved by the Ethical Committee of the National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences (NCC3916) and compiled with all relevant ethical regulations. All patients provided written informed consent for collecting tissue samples for research and publishing relevant clinical information. The study was performed in accordance with the Declaration of Helsinki.
Human specimens
In this study, a total of 23 fresh tumor specimens (7 pre-treatment biopsy samples & 16 post-treatment surgery samples) from 18 locally advanced ESCC patients, who received nICT followed by esophagectomy at Cancer Hospital, Chinese Academy of Medical Sciences from November, 2020 to July, 2023, were collected and enrolled in subsequent scRNA-seq combined scTCR-seq analyses. We collected the pre-treatment biopsy samples through esophagoscopy at baseline and harvested the post-treatment samples in surgery. All pre-treatment samples enrolled in analysis were collected from locally advanced ESCC tumors that were clearly visible under esophagoscopy and could be obtained through biopsy. Meanwhile, the biopsy samples for diagnosis, collected at the same site, underwent hematoxylin and eosin (H & E) staining, and were confirmed as tumors by two experienced pathologists (L.L. and Z.Y.). During the sampling for post-treatment samples from responders, Lugol’s iodine was applied for classification of regressed primary tumor bed and deeply stained para-tumor epithelial mucosa. Additionally, we retrospectively obtained 96 pairs of matched pre- and post-treatment formalin-fixed paraffin-embedded (FFPE) tumor tissues from 96 ESCC patients who underwent nICT followed by surgery in our center. Corresponding clinical data was retrieved from hospital records. On the basis of H & E staining slides from post-treatment surgical resections, the pathological response rate of each ESCC patient were evaluated by experienced pathologists (L.L. and Z.Y.) blinded to the study design. Analyses performed in this study were based on therapeutic efficacy and treatment timepoints. Sex- or gender-based analyses were not performed. Race, ethnicity, or other socially relevant groupings were not involved.
Sample processing
Pre- and post-treatment samples were obtained from the same primary tumor before and after nICT treatment. Following collection in the clinical setting, we conducted immediate blood removal for tumor samples with phosphate-buffered saline (PBS, Gibco), stored them in MACS Tissue Storage Solution (Miltenyi, 130-100-008), and transferred them on ice to the lab. Concisely, tissue samples were cut into ~1–2 mm3 pieces in the RPMI-1640 medium (Gibco) and then were enzymatically digested using the tumor Dissociation Kit, human (Miltenyi, 130-095-929) with the program 37C_m_TDK_1 on a rotor at 37 °C, following the manufacturer’s instruction. Subsequently, the dissociated cells were passed through a 100-μm SmartStrainer and centrifuged at 400 g for 5 min. After removing the supernatant, pelleted cells were suspended in the red blood cell lysis buffer (Miltenyi) and incubated on ice for 1–2 min to lyse the red blood cells. The cell pellets were then washed twice using PBS (Gibco) and re-suspended in sorting buffer, consisting of PBS supplemented with 2% fetal bovine serum (FBS, Gibco).
Preparation and sequencing of single-cell RNA-seq and TCR-seq libraries
As single cells were re-suspended in PBS containing 2% FBS (Gibco) to a final concentration of 500–1200 cells/μl as determined by the Rigel S2 cell counter (Countstar), ~8000 cells were captured in droplets to generate nanoliter-scale Gel beads in Emulsion (GEMs). GEMs were then reverse transcribed in a C1000 Touch Thermal Cycler (Bio-Rad) programmed at 53 °C for 45 min, 85 °C for 5 min, and held at 4 °C. After reverse transcription and cell barcoding, the emulsions were broken and cDNA was isolated and purified with Cleanup Mix containing DynaBeads and SPRIselect reagent (Thermo Fisher Scientific), followed by PCR amplification. Amplified cDNA was then used for both 5’ gene expression library construction and TCR V(D)J targeted enrichment. The 5’ gene expression library was prepared using the Chromium Next GEM Single Cell 5’ Library & Gel Bead Kit v1.1 (10x Genomics, PN-1000165), and the TCR library was prepared using the Chromium Single Cell V(D)J Enrichment Kit (10x Genomics, PN-1000005) according to the manufacturer’s user guide. After quality assessment, the single-cell RNA and TCR V(D)J libraries were sequenced on the Illumina NovaSeq platform with 150 bp paired-end reads.
Single-cell RNA-Seq data processing
Raw gene expression matrices were generated for each sample by the Cell Ranger (version 5.0.0) provided by 10x Genomics, and the pipeline was coupled with the human reference genome (GRCh38). The output-filtered gene expression matrices were further analyzed by R software (version 3.6.1) with the Seurat package (version 3.2.0, Supplementary Software 1). Cells with low quality were removed based on the following criteria: (1) <200 genes; (2) expressed gene counts <3; (3) >10% UMIs derived from the mitochondrial genome. After removing low-quality cells, the gene expression matrices were normalized by the NormalizeData function, and features with high cell-to-cell variation were calculated using the FindVariableFeatures function.
Dimension reduction and unsupervised clustering for scRNA-seq data
To reduce the dimensionality of the datasets, the RunPCA function was conducted with default parameters on linear-transformation scaled data generated by the ScaleData function. Next, the ElbowPlot and DimHeatmap functions were used to identify the proper dimensions of each dataset. Finally, we clustered cells using the FindNeighbors and FindClusters functions and performed non-linear dimensional reduction with the RunUMAP function with default settings. The Harmony algorithm was utilized to correct the batch effect of T cells.
Cell type annotation and cluster markers identification
After non-linear dimensional reduction and projection of all cells into two-dimensional space by UMAP, cells clustered together according to common features. In the process of cell cluster annotation, we referred to the differentially expressed genes (DEGs) of each cell cluster. DEGs were filtered using the t-test under a minimum absolute value of log (fold change) of 0.25 (|log (fold change) |≥0.25) and a p-value of 0.01. Cell clusters were then classified and annotated based on expressions of canonical markers of particular cell types.
Construction of cell-to-cell interaction network and ligand-receptor analysis
We investigated the cell-to-cell interactions among diverse cell types using the R package “CellChat” (v1.1.3, Supplementary Software 1), which could quantitatively infer and analyze intercellular communication networks from scRNA-seq data72. We mainly utilized the “netVisual_circle” function of CellChat, and the corresponding results were presented via crosstalk maps and ligand-receptor heatmaps.
Single-cell TCR data processing
The Cell Ranger toolkit (version 5.0.0, Supplementary Software 1) provided by 10x Genomics was applied to align TCR-seq reads to the human reference genome (GRCh38) and assemble TCR sequences. Preliminary TCR sequences were filtered to keep those characterized with high confidence, full-length, productive, and assigned with a valid cell barcode and an unambiguous chain type. Each cell was assigned one alpha and one beta chain with the highest UMI counts. Cells with identical beta chains were defined as clonal cells and considered to originate from the same ancestry.
Gini index calculation
The Gini coefficient was computed using the R package “DescTools” (Version 0.99.47). The detailed calculation procedure has been described previously73, and the equation used is as follows:
where \({p}_{i}\) and \({p}_{j}\) represent the frequency of the ith and jth TRB clonotype, \(N\) is the number of TRB clonotypes, and the \(\bar{p}\) is the average of clonotype frequencies.
Clonal analysis
A clonotype in a specific cell cluster, comprising no fewer than 3 cells, was considered expanded, regardless of the annotation of other cells within the same clonotype. A cutoff of 3 was selected, representing the threshold for a high-quality clonotype. For CD8+ T cells, “Tex-relevant cells” were defined as CD8+ T cells sharing the same TCR with CD8+ Tex-CXCL13, and the resting cells were designated as “Tex-irrelevant cells”. Within Tex-relevant cells, those not annotated as CD8+ Tex-CXCL13 were identified as precursor exhausted T (Texp) cells.
RNA velocity analysis
RNA velocity values for each gene of each cell were calculated by scVelo, and the computationally inferred pseudotime of all cells was further estimated based on RNA velocities. To confirm the cell state transitions, we performed another RNA velocity-based algorithm, Dynamo, following the standard workflow.
Copy number variation (CNV) estimation
We utilized CopyKAT 0.1.0 (Copynumber Karyotyping of Tumors, Version 0.1.0, Supplementary Software 1), a computational tool that can identify genome-wide aneuploidy based on integrative Bayesian approaches (https://github.com/navinlabcode/copykat), to infer CNV as described by previous study74. CopyKAT used a Gaussian Mixture Model (GMM) to estimate the variance of each single-cell cluster. Given the prevalence of aneuploidy in human cancers (90%), CopyKAT defined the cluster with minimal estimated variance as the “confident diploid cells”, and cells displaying extensive genome-wide CNV were considered tumor cells.
Identification of DEGs in malignant cells and pathway enrichment
We identified the DEGs in malignant cells between responders and non-responders using the t-test. The selection criteria for DEGs included a minimum absolute value of log (fold change) of 0.25 (|log (fold change)| ≥ 0.25) and a p-value of 0.01. Based on these DEGs, we performed the pathway enrichment using Gene Ontology (GO) method.
RNA sequencing
For validation, based on the retrospective extracted ESCC cohort, we screened out 54 patients with paired high-quality pre- and post-treatment FFPE sections (Supplementary Data 5), from which total RNA was extracted using the miRNeasy FFPE kit (Qiagen) according to the manufacturer’s protocol. The extracted total RNA was fragmented through divalent cations at high temperature. Random primers were applied to reverse transcribe the interrupted total RNA fragment to obtain the first and second strands of cDNA. The reverse transcription product was subjected to terminal repair, followed by addition of an A base at the 3’ end. Subsequently, the fragments were ligated with an Illumina universal adapter. After PCR amplified, the product was hybridized in the hybridization buffer. The sequence specific probe was used to capture the coding region of the transcriptome. After purification, the captured product was enriched by PCR and purified to obtain the final sequencing library. Library sizes were detected via an Agilent 2100 Bioanalyzer, and library concentrations were examined using Qubit 2.0. The sequencing was performed according to Illumina’s standard protocol.
In this project, read quality was calculated for all samples using FastQC software (v0.11.9) with the default parameters. PE 150 was applied to sequence the samples. Fastp software (v0.20.1) was used to remove the adapter sequence of raw data (Fastq)75, and reads longer than 75 bp after adapter removal were selected for downstream analysis, resulting in a set of reads referred to as ‘clean reads’. Clean reads of each sample were aligned to reference genome using STAR software with default parameters. RSEM (v1.3.3) was applied to accurately quantify gene or transcript expression levels from RNA-seq data76. The edgeR (v3.28.1) package in R was used to perform TMM (trimmed mean of M values) and CPM (count per million) normalization for the gene expression77. Then, the gene expression matrix was transformed to TPM (transcripts per million) format for following analysis. The RNA-seq data of paired pre- and post-treatment FFPE samples from these 54 patients was applied as an external independent validation cohort, named nICT-ESCC, to verify these findings derived from single-cell analysis.
Gene signature calculation
In scRNA-seq analysis, gene signature calculation was performed via Vision78. The progenitor exhaustion and terminal exhaustion signatures were derived from a previous study defining T cell states associated with response to checkpoint immunotherapy28. Cell type specific signatures of each cell cluster were defined as the top 30 DEGs ranked by log-transformed the ratio of normalized gene expression in this cell cluster relative to other clusters within corresponding major cell compartments (log2_avg). Cell type specific signatures of CD8+ Tex-CXCL13, CD8+ Tex-STMN1, CD4+ Treg-TNFRSF4, CD4+ Tfh/Th1-CXCL13, tCAF-MMP11, Macro-EREG and Macro-SPP1were extracted (Supplementary Data 4). To validate the distributions of distinct cell clusters revealed by single-cell analysis, we utilized the gene signatures of selected cell clusters mentioned above and calculated their expression score via ssGSEA based on the RNA-seq data of 54 pairs of FFPE samples.
Multiplex immunofluorescence staining
We collected 96 FFPE specimens of pre-treatment biopsy from the retrospective ESCC cohort with nICT treatment and performed immunofluorescence staining of CD8 and CXCL13 on these tumor biopsy sections. A mixture of primary antibodies against CXCL13 (1:200, Abcam, ab246518) and CD8 (1:500, Servicebio, GB12068) was incubated at 4 °C overnight, followed by the incubation of the corresponding secondary antibodies for 50 min. CXCL13 and CD8 were labeled red and green, respectively. The nuclei were stained with DAPI (4′,6-diamidino-2-phenylindole). The stained slides were scanned via the Panoramic MIDI (3DHISTECH). We defined the average number of CXCL13/CD8 dual-positive cells of 8 high-power fields as the CXCL13/CD8 dual-positive cell abundance. For post-treatment samples, primary antibodies for LRRC15 (1:200, Abcam, ab150376), α-SMA (1:2000, Abcam, ab124964), CD4 (1:500, Abcam, ab133616) and FOXP3 (1:50, CST, 98377 s) were incubated for 1 h at room temperature, and horseradish peroxidase-labeled secondary antibodies were incubated at room temperature for 30 min, followed by TSA fluorescent dye working solution incubation. After multi-antigen staining, nuclei were stained with DAPI. The stained slides were enclosed by Nail Polish, scanned via the SLIDEVIEW VS200 (Olympus), and analyzed using OlyVIA software.
Tertiary lymphoid structure evaluation
To explore the association between tertiary lymphoid structure (TLS) and clinical efficacy, we collected the post-treatment H & E slides of 96 ESCC patients from the retrospective ESCC cohort receiving neoadjuvant immunotherapy. As previously described79, briefly, TLSs were defined as lymphoid aggregates of B lymphocytes (a mixture of a variable proportion of T lymphocytes and plasma cells in most cases). For each slide, TLS abundance was calculated by the number of TLS in the area of the tumor bed divided by the area of the esophagus on the slide.
Cell culture
The mouse ESCC cell line (mEC25) was kindly provided by Dr. Li Fu from Guangdong Provincial Key Laboratory of Regional Immunity and Diseases, Department of Pharmacology and Shenzhen University International Cancer Center, Shenzhen University Health Science Center80,81. Cell line authentication was performed by Dr. Li Fu using short tandem repeat profiling. Additionally, HE and IHC staining were also conducted to confirm the tissue source from esophagus. The mEC25 cells were cultured in DMEM high-glucose medium with 10% FBS and incubated with 5% CO2 at 37 °C. All cells were tested negative for mycoplasma contamination using TransDetect® PCR Mycoplasma Detection Kit (Transgen Biotech, FM311-01).
Animal experiments
All animal experiments were carried out strictly in accordance with the protocols approved by the Institutional Animal Care and Use Committee of Cancer Hospital, Chinese Academy of Medical Sciences. As this study focused on exploring the immune features of different treatment response to nICT in ESCC and the majority was male patients, sex as a biological variable was not considered in the study design, and only male mice were used in experiments. Male C57BL/6 mice, aged 5–6 weeks, were purchased from Changchun YISI Experimental Animal Technology Company and acclimated for 7 days in our animal facility before experimentation. Mice were maintained in pathogen-free cages keeping on ventilated racks with adequate food and water supply, sufficient nesting and bedding materials, appropriate temperature and humidity, as well as a 12-h cycle of light and darkness. To establish the tumor model, 50 μl single-cell suspensions of mEC25 cells (5 × 106) mixed with 50 μl Matrigel (BD Pharmingen, catalog No. 354234) were subcutaneously injected into the right flank of mice. Tumor volume (mm3) was measured every 3 days and calculated as (length × width2)/2. Saline control, rmCXCL13 (5 μg per mouse, R&D, catalog No. 470-BC-025), and anti-murine PD-1 (200 μg per mouse, BioXCell, catalog No. BE0033-2) were injected intraperitoneally 7 days after tumor implantation and every 3 days thereafter until the end of the experiment. For the combinational therapy, mice were treated with both rmCXCL13 and anti-murine PD-1. After eight injections, the mice were euthanized by cervical dislocation. No tumor reached 2000 mm3 in volume (this animal license limit was not exceeded). And then the heart, lung, liver, spleen, and kidney were harvested for HE staining to study potential changes in organ morphology. Tumors were also harvested for subsequent flow cytometry and multiplex immunofluorescence analysis. For flow cytometry analysis, the tumors were digested into single-cell suspensions and immunostained with several fluorochrome-tagged monoclonal antibodies, including anti-PD-1 (1:100, Biolegend, 135210), anti-CD20 (1:100, Biolegend, 150422), anti-CD3 (1:100, Biolegend, 100203), and anti-CD8 (1:100, Biolegend, 126608). The stained cells were analyzed on a flow cytometer (BD, C6). For multiplex immunofluorescence analysis, the tumor tissues were formaldehyde-fixed and sectioned into slides, which were stained with anti-CD4 (1:1000, Abcam, ab288724) and anti-CD8 (1:2000, Abcam, ab217344). The color setting for immunofluorescence was as follows: red-CD4, green-CD8. Cell nuclei were counterstained with DAPI.
Statistics
Statistical analyses were performed as described in the figure legends. In brief, the t-test or Wilcox-test was used to determine the significance of continuous variables. Compositional changes of various cell types between distinct efficacy groups were also detected by scCODA (https://github.com/theislab/scCODA), a Bayesian model for compositional single-cell data analysis82. We evaluated the relevance between continuous variables using Spearman’s correlation or Pearson’s correlation. Statistical tests were conducted in R (v. 3.6.1). A two-tailed p < 0.05 was regarded as statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq, scTCR-seq and bulk RNA-seq data of this study have been deposited to GSA for Human (https://ngdc.cncb.ac.cn/gsa-human/) with an accession number of HRA003591 for single cell sequencing data and HRA007035 for FFPE RNA sequencing data, respectively. The data is available under restricted access and will be provided for scientific research upon request complying with the law due to human patient privacy concerns. Readers can obtain access to the data via initiating application requests through the OMIX system of GSA for Human. Data access requests will be responded within 7 days and once access has been granted, the data can be download within 2 weeks. Requests for access to the patient-level data from this study can be submitted via email to the corresponding author with a detailed proposal for approval. The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
Code availability
The source code employed for data processing and analysis in this study is available in Supplementary Software 1.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 71, 209–249 (2021).
Chen, W. et al. Cancer statistics in China, 2015. CA A Cancer J. Clin. 66, 115–132 (2016).
Abnet, C. C., Arnold, M. & Wei, W. Q. Epidemiology of esophageal squamous cell carcinoma. Gastroenterology 154, 360–373 (2018).
Eyck, B. M. et al. Ten-year outcome of neoadjuvant chemoradiotherapy plus surgery for esophageal cancer: the randomized controlled CROSS trial. J. Clin. Oncol. 39, 1995–2004 (2021).
Yang, H. et al. Long-term efficacy of neoadjuvant chemoradiotherapy plus surgery for the treatment of locally advanced esophageal squamous cell carcinoma: the NEOCRTEC5010 randomized clinical trial. JAMA Surg. 156, 721–729 (2021).
van Hagen, P. et al. Preoperative chemoradiotherapy for esophageal or junctional cancer. N. Engl. J. Med. 366, 2074–2084 (2012).
Yang, H. et al. Neoadjuvant chemoradiotherapy followed by surgery versus surgery alone for locally advanced squamous cell carcinoma of the esophagus (NEOCRTEC5010): a phase iii multicenter, randomized, open-label clinical trial. J. Clin. Oncol. 36, 2796–2803 (2018).
Pasquali, S. et al. Survival after neoadjuvant and adjuvant treatments compared to surgery alone for resectable esophageal carcinoma: a network meta-analysis. Ann. Surg. 265, 481–491 (2017).
Chan, K. K. W. et al. Neoadjuvant treatments for locally advanced, resectable esophageal cancer: a network meta-analysis. Int. J. Cancer 143, 430–437 (2018).
Liu, S. et al. Recurrence patterns after neoadjuvant chemoradiotherapy compared with surgery alone in oesophageal squamous cell carcinoma: results from the multicenter phase III trial NEOCRTEC5010. Eur. J. Cancer 138, 113–121 (2020).
Topalian, S. L. et al. Neoadjuvant immune checkpoint blockade: a window of opportunity to advance cancer immunotherapy. Cancer Cell 41, 1551–1566 (2023).
Patel, M. A., Kratz, J. D., Lubner, S. J., Loconte, N. K. & Uboha, N. V. Esophagogastric cancers: integrating immunotherapy therapy into current practice. J. Clin. Oncol. 40, 2751–2762 (2022).
Li, Y. et al. Comparing a PD-L1 inhibitor plus chemotherapy to chemotherapy alone in neoadjuvant therapy for locally advanced ESCC: a randomized Phase II clinical trial: a randomized clinical trial of neoadjuvant therapy for ESCC. BMC Med. 21, 86 (2023).
Yan, X. et al. Tislelizumab combined with chemotherapy as neoadjuvant therapy for surgically resectable esophageal cancer: a prospective, single-arm, phase II study (TD-NICE). Int. J. Surg. 103, 106680 (2022).
Chen, X. et al. Neoadjuvant sintilimab and chemotherapy in patients with potentially resectable esophageal squamous cell carcinoma (KEEP-G 03): an open-label, single-arm, phase 2 trial. J. Immunother. Cancer 11, https://doi.org/10.1136/jitc-2022-005830 (2023).
Bejarano, L., Jordāo, M. J. C. & Joyce, J. A. Therapeutic targeting of the tumor microenvironment. Cancer Discov. 11, 933–959 (2021).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593.e1578 (2021).
Kieffer, Y. et al. Single-cell analysis reveals fibroblast clusters linked to immunotherapy resistance in cancer. Cancer Discov. 10, 1330–1351 (2020).
Goubet, A. G. et al. Escherichia coli-specific CXCL13-producing TFH are associated with clinical efficacy of neoadjuvant PD-1 blockade against muscle-invasive bladder cancer. Cancer Discov. 12, 2280–2307 (2022).
Ren, X. et al. Insights gained from single-cell analysis of immune cells in the tumor microenvironment. Annu. Rev. Immunol. 39, 583–609 (2021).
Sadeghi Rad, H. et al. Understanding the tumor microenvironment for effective immunotherapy. Med. Res. Rev. 41, 1474–1498 (2021).
Pai, J. A. & Satpathy, A. T. High-throughput and single-cell T cell receptor sequencing technologies. Nat. Methods 18, 881–892 (2021).
Krishna, C. et al. Single-cell sequencing links multiregional immune landscapes and tissue-resident T cells in ccRCC to tumor topology and therapy efficacy. Cancer Cell 39, 662–677.e666 (2021).
Zheng, Y. et al. Immune suppressive landscape in the human esophageal squamous cell carcinoma microenvironment. Nat. Commun. 11, 6268 (2020).
Chen, Z. et al. Dissecting the single-cell transcriptome network underlying esophagus non-malignant tissues and esophageal squamous cell carcinoma. EBioMedicine 69, 103459 (2021).
Zhang, X. et al. Dissecting esophageal squamous-cell carcinoma ecosystem by single-cell transcriptomic analysis. Nat. Commun. 12, 5291 (2021).
Dinh, H. Q. et al. Integrated single-cell transcriptome analysis reveals heterogeneity of esophageal squamous cell carcinoma microenvironment. Nat. Commun. 12, 7335 (2021).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e1020 (2018).
Liu, B. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022).
Li, H. et al. Dysfunctional CD8 T cells form a proliferative, dynamically regulated compartment within human melanoma. Cell 176, 775–789.e718 (2019).
van der Leun, A. M., Thommen, D. S. & Schumacher, T. N. CD8(+) T cell states in human cancer: insights from single-cell analysis. Nat. Rev. Cancer 20, 218–232 (2020).
Chaurio, R. A. et al. TGF-β-mediated silencing of genomic organizer SATB1 promotes Tfh cell differentiation and formation of intra-tumoral tertiary lymphoid structures. Immunity 55, 115–128.e119 (2022).
Ng, K. W. et al. Antibodies against endogenous retroviruses promote lung cancer immunotherapy. Nature 616, 563–573 (2023).
Cabrita, R. et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577, 561–565 (2020).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med 25, 1251–1259 (2019).
Castellino, F. et al. Chemokines enhance immunity by guiding naive CD8+ T cells to sites of CD4+ T cell-dendritic cell interaction. Nature 440, 890–895 (2006).
Simoni, Y. et al. Bystander CD8(+) T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Oliveira, G. et al. Phenotype, specificity and avidity of antitumour CD8(+) T cells in melanoma. Nature 596, 119–125 (2021).
Obradovic, A. et al. Immunostimulatory cancer-associated fibroblast subpopulations can predict immunotherapy response in head and neck cancer. Clin. Cancer Res. 28, 2094–2109 (2022).
Li, C. et al. Spatial and single-cell transcriptomics reveal a cancer-associated fibroblast subset in HNSCC that restricts infiltration and anti-tumor activity of CD8+ T cells. Cancer Res. https://doi.org/10.1158/0008-5472.can-23-1448 (2023).
Ma, C. et al. Pan-cancer spatially resolved single-cell analysis reveals the crosstalk between cancer-associated fibroblasts and tumor microenvironment. Mol. Cancer 22, 170 (2023).
Costa, A. et al. Fibroblast heterogeneity and immunosuppressive environment in human breast cancer. Cancer Cell 33, 463–479.e410 (2018).
Wu, C. et al. Galectin-9-CD44 interaction enhances stability and function of adaptive regulatory T cells. Immunity 41, 270–282 (2014).
Dominguez, C. X. et al. Single-cell RNA sequencing reveals stromal evolution into LRRC15(+) myofibroblasts as a determinant of patient response to cancer immunotherapy. Cancer Discov. 10, 232–253 (2020).
Nasir, I. et al. Tumor macrophage functional heterogeneity can inform the development of novel cancer therapies. Trends Immunol. 44, 971–985 (2023).
Bill, R. et al. CXCL9:SPP1 macrophage polarity identifies a network of cellular programs that control human cancers. Science 381, 515–524 (2023).
Qi, J. et al. Single-cell and spatial analysis reveal interaction of FAP(+) fibroblasts and SPP1(+) macrophages in colorectal cancer. Nat. Commun. 13, 1742 (2022).
Klement, J. D. et al. An osteopontin/CD44 immune checkpoint controls CD8+ T cell activation and tumor immune evasion. J. Clin. Investig. 128, 5549–5560 (2018).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Spitzer, M. H. et al. Systemic immunity is required for effective cancer immunotherapy. Cell 168, 487–502.e415 (2017).
Miller, B. C. et al. Subsets of exhausted CD8(+) T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Liu, J. Y. et al. CTL- vs Treg lymphocyte-attracting chemokines, CCL4 and CCL20, are strong reciprocal predictive markers for survival of patients with oesophageal squamous cell carcinoma. Br. J. Cancer 113, 747–755 (2015).
Pan, C. et al. Phenotypic profiling and prognostic significance of immune infiltrates in esophageal squamous cell carcinoma. Oncoimmunology 10, 1883890 (2021).
Kang, J. H. & Zappasodi, R. Modulating Treg stability to improve cancer immunotherapy. Trends Cancer 9, 911–927 (2023).
Clarke, J. et al. Single-cell transcriptomic analysis of tissue-resident memory T cells in human lung cancer. J. Exp. Med. 216, 2128–2149 (2019).
Workel, H. H. et al. A transcriptionally distinct CXCL13(+)CD103(+)CD8(+) T-cell population is associated with b-cell recruitment and neoantigen load in human cancer. Cancer Immunol. Res 7, 784–796 (2019).
Hsieh, C. H. et al. Potential role of CXCL13/CXCR5 signaling in immune checkpoint inhibitor treatment in cancer. Cancers 14, https://doi.org/10.3390/cancers14020294 (2022).
Thommen, D. S. et al. A transcriptionally and functionally distinct PD-1(+) CD8(+) T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat. Med. 24, 994–1004 (2018).
Fridman, W. H. et al. Tertiary lymphoid structures and B cells: an intratumoral immunity cycle. Immunity 56, 2254–2269 (2023).
Kinker, G. S. et al. Mature tertiary lymphoid structures are key niches of tumour-specific immune responses in pancreatic ductal adenocarcinomas. Gut 72, 1927–1941 (2023).
Carril-Ajuria, L. et al. Baseline circulating unswitched memory B cells and B-cell related soluble factors are associated with overall survival in patients with clear cell renal cell carcinoma treated with nivolumab within the NIVOREN GETUG-AFU 26 study. J. Immunother. Cancer 10, https://doi.org/10.1136/jitc-2022-004885 (2022).
Nakamura, S. et al. Tertiary lymphoid structures correlate with enhancement of antitumor immunity in esophageal squamous cell carcinoma. Br. J. Cancer 129, 1314–1326 (2023).
Havenar-Daughton, C. et al. CXCL13 is a plasma biomarker of germinal center activity. Proc. Natl Acad. Sci. USA 113, 2702–2707 (2016).
Grout, J. A. et al. Spatial positioning and matrix programs of cancer-associated fibroblasts promote T-cell exclusion in human lung tumors. Cancer Discov. 12, 2606–2625 (2022).
Liu, Y. et al. Identification of a tumour immune barrier in the HCC microenvironment that determines the efficacy of immunotherapy. J. Hepatol. 78, 770–782 (2023).
Krishnamurty, A. T. et al. LRRC15(+) myofibroblasts dictate the stromal setpoint to suppress tumour immunity. Nature 611, 148–154 (2022).
Wu, J., Shen, Y., Zeng, G., Liang, Y. & Liao, G. SPP1(+) TAM subpopulations in tumor microenvironment promote intravasation and metastasis of head and neck squamous cell carcinoma. Cancer Gene Ther. https://doi.org/10.1038/s41417-023-00704-0 (2023).
Rutihinda, C. et al. Inhibition of the CCR6-CCL20 axis prevents regulatory T cell recruitment and sensitizes head and neck squamous cell carcinoma to radiation therapy. Cancer Immunol. Immunother. 72, 1089–1102 (2023).
Cook, K. W. et al. CCL20/CCR6-mediated migration of regulatory T cells to the Helicobacter pylori-infected human gastric mucosa. Gut 63, 1550–1559 (2014).
Liu, L. et al. Construction of TME and Identification of crosstalk between malignant cells and macrophages by SPP1 in hepatocellular carcinoma. Cancer Immunol. Immunother. 71, 121–136 (2022).
Brina, D. et al. The Akt/mTOR and MNK/eIF4E pathways rewire the prostate cancer translatome to secrete HGF, SPP1 and BGN and recruit suppressive myeloid cells. Nat. Cancer 4, 1102–1121 (2023).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Chiffelle, J. et al. T-cell repertoire analysis and metrics of diversity and clonality. Curr. Opin. Biotechnol. 65, 284–295 (2020).
Gao, R. et al. Delineating copy number and clonal substructure in human tumors from single-cell transcriptomes. Nat. Biotechnol. 39, 599–608 (2021).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
DeTomaso, D. et al. Functional interpretation of single cell similarity maps. Nat. Commun. 10, 4376 (2019).
Vanhersecke, L. et al. Mature tertiary lymphoid structures predict immune checkpoint inhibitor efficacy in solid tumors independently of PD-L1 expression. Nat. Cancer 2, 794–802 (2021).
Huang, T., Yang, J., Liu, B. & Fu, L. A new mouse esophageal cancer cell line (mEC25)-derived pre-clinical syngeneic tumor model for immunotherapy. Cancer Commun. 40, 316–320 (2020).
Huang, T. X. et al. Targeting cancer-associated fibroblast-secreted WNT2 restores dendritic cell-mediated antitumour immunity. Gut 71, 333–344 (2022).
Büttner, M., Ostner, J., Müller, C. L., Theis, F. J. & Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat. Commun. 12, 6876 (2021).
Acknowledgements
This study was supported by National Key R&D Program of China (2021YFC2500900 to S.G.G.); CAMS Innovation Fund for Medical Sciences (CIFMS) (2022-I2M-C&T-B-058 to Z.L.Y.); National Natural Sciences Foundation (82273129 to S.G.G., 82203827 and 82472808 to Z.L.Y., 82102886 to J.C.X.); Young Elite Scientists Sponsorship Program by CAST (2022QNRC001 to Z.L.Y.); Beijing Nova Program (Z211100002121055 to Z.L.Y., 20220484119 to J.C.X.); Beijing Natural Science Foundation (7222146 to Z.L.Y., 7242114 to J.C.X.); Beijing Hope Run Special Fund of Cancer Foundation of China (LC2021B22 to Z.L.Y.); Guangdong Association of Clinical Trials (GACT) /Chinese Thoracic Oncology Group (CTONG) and Guangdong Provincial Key Lab of Translational Medicine in Lung Cancer (Grant No. 2017B030314120 to J.C.X.). We thank Doctor Xinfeng Wang from Department of Thoracic Surgery, National Cancer Center/National Clinical Research Center for Cancer/Cancer Hospital for his assistance in the drawing of Figs. 1a and 3a via BioRender (https://www.biorender.com/). In addition, Figs. 1a and 3a contain modified images from Servier Medical Art (https://smart.servier.com/), which have been licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/).
Author information
Authors and Affiliations
Contributions
S.G.G., and L.Y.X. conceived the study and designed the experiments. Z.L.Y., X.W.C., and B.Z. L. performed the experiments, conducted analysis and interpretation of the data. Z.L.Y., X.W.C., G.Y.B., and S.G.G. conducted surgery and collected samples from patients. X.W.C., J.C.X., and Q.Y.C. contributed to reagents/analysis tools. X.W.C., Z.L.Y., and S.G.G. composed, reviewed, and revised the manuscript. Y.P., Q.L., S.B.W., H.T., and W.G. were responsible for administrative or material support. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sarah Derks and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, Z., Tian, H., Chen, X. et al. Single-cell sequencing reveals immune features of treatment response to neoadjuvant immunochemotherapy in esophageal squamous cell carcinoma. Nat Commun 15, 9097 (2024). https://doi.org/10.1038/s41467-024-52977-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52977-0
This article is cited by
-
Single-cell omics in tumor lymph node metastasis: mechanisms and therapeutic implications
Molecular Cancer (2026)
-
18F-NOTA-FAPI-04 PET/CT metabolic parameters combined with hematological biomarkers predict pathological complete response after neoadjuvant chemoradiotherapy in esophageal squamous cell carcinoma
European Journal of Nuclear Medicine and Molecular Imaging (2026)
-
Cancer-associated fibroblasts in the tumor microenvironment: heterogeneity, crosstalk mechanisms, and therapeutic implications
Molecular Cancer (2025)
-
Intratumoral microbiota-aided fusion radiomics model for predicting tumor response to neoadjuvant chemoimmunotherapy in triple-negative breast cancer
Journal of Translational Medicine (2025)
-
Neoadjuvant chemotherapy with or without PD-1/PD-L1 inhibitors in resectable esophageal squamous cell carcinoma: a meta-analysis based on randomized controlled trials
BMC Gastroenterology (2025)