Abstract
The spatial and temporal dynamics of forces in cells coordinate essential behaviors like division, polarization, and migration. While intracellular signaling initiates contractile ring assembly during cell division, how mechanical forces coordinate division and their energetic costs remain unclear. Here, we develop an in vitro model where myosin-induced stress drives division-like shape changes in giant unilamellar vesicles (GUVs, liposomes). Myosin activity is controlled by light patterns globally or locally at the equator. Global activation causes slow, shallow cleavage furrows due to a tug-of-war between the equatorial and polar forces. By contrast, local activation leads to faster, deeper, and symmetric division as equatorial forces dominate. Dissociating the actin cortex at the poles is crucial for inducing significant furrowing. During furrowing, actomyosin flows align actin filaments parallel to the division plane, forming a contractile ring-like structure. Mechanical power is not greatest during contraction, but is maximized just before furrowing. This study reveals the quantitative relationship between force patterning and mechanical energy during division-like shape changes, providing insights into cell division mechanics.
Similar content being viewed by others
Introduction
Cell division is fundamental to embryonic development1, cancer metastasis2, and tissue homeostasis3. Division failure leads to abnormal chromosome numbers and cell death4. In animal cells, the division plane positioning is regulated by RhoA signaling from the anaphase spindle, which locally activates F-actin polymerization and myosin II to assemble a contractile ring5,6,7,8. The contractile ring is formed through the reorganization of F-actin, driven by locally increased mechanical tension9 and compression from actomyosin cortical flows10. Division begins with cell elongation perpendicular to the division plane11 and chromosome separation towards astral microtubules12, followed by contractile ring constriction to divide the cell.
While intracellular signaling initiates F-actin assembly and myosin phosphorylation, it is mechanical forces that drive the subsequent changes in cell shape. Growing evidence suggests that not only equatorial contractile forces, but also the spatial balance across the entire cell, including contractility and elasticity at the cell poles, are crucial for successful cell division. For instance, cell elongation has been linked to the relaxation of cortical stiffness at the poles through actin clearance13,14,15,16. Conversely, globally activated contractility, including at the poles, can destabilize symmetric division17, while decreased equatorial elasticity may facilitate division18,19,20,21. Moreover, certain cell types, such as neuroblasts of C. elegans and Drosophila22,23, exhibit stable asymmetric division associated with polarized F-actin assembly24,25. However, the precise role of mechanical relaxation in successful cell division, as well as the energetic efficiency of mechanical processes during division, remains largely unexplored. While the contractile ring is formed through F-actin reorganization driven by localized tension9 and actomyosin flows10, the energetic signature of this transition remains unknown.
Understanding the mechanics and energetics of cell division is often hindered by cellular complexity, such as variable protein composition and overlapping signaling pathways. Here, we developed an in vitro model of dividing cells by encapsulating purified actin and myosin within giant unilamellar vesicles (GUVs, liposomes)26,27,28,29,30,31,32,33. This system enables precise control over both molecular inputs (e.g., protein composition) and mechanical inputs (i.e., actomyosin contractility), which is infeasible in living cells. By controlling myosin activity with light patterns, we investigate its impact on dynamics, symmetry, and mechanical energy usage of cell division-like shape changes. We correlate shape changes with liposome aspect ratio, furrow depth, and F-actin alignment under varying actin-membrane attachment strength, turnover, and F-actin architecture. Our aim is to determine how contractility regulates shape change and F-actin organization to optimize mechanical outputs during division. We identify that mechanical power is maximized during shape elongation, coinciding with contractile ring formation, while furrowing expends less mechanical power and leads to greater energy dissipation.
Results
Contractility patterning drives cell division-like shape changes
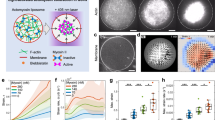
We developed an experimental system enabling cell division-like membrane deformation. Purified actin, myosin II, and actin-associated proteins were encapsulated within liposomes using the inverted emulsion method (Methods)26,34,35 (Fig. 1a). The linear F-actin nucleator formin mDia136, which is essential in dividing cells37,38, was used for F-actin polymerization to facilitate myosin force generation39. Histidine-tagged mDia1 (His-mDia1) was localized to the lipid bilayer membrane containing DOGS-NiNTA lipids (Fig. 1a) (Methods). F-actin was dispersed within the liposome volume since mDia1 may release the polymerized F-actin after elongation36 (Fig. 1b, midplane). Thus, F-actin was tethered to the membrane via His-α-actinin to form a cortical F-actin network (Fig. 1b, surface), enabling transmission of actomyosin contractility to the membrane. We expect all His-mDia1 (300 nM) and His-α-actinin (70-280 nM) to be membrane-bound since there are a sufficient number of membrane binding sites for all the histidine-tagged proteins within the range of protein concentrations we use, as estimated in our previous study40.
a Schematic of a liposome encapsulating actomyosin. His-tagged proteins were tethered to the membrane via DOGS-NiNTA lipids embedded in the lipid bilayer. Linear F-actin was nucleated by His-mDia1 and tethered to the membrane via His-α-actinin. b Confocal slices of the actin fluorescence image in the midplane and at the bottom surface of the liposome. Similar F-actin organization was consistently obtained in more than 10 independent repeats (e.g., Fig. 4b, Fig. 5a). c Schematic of the patterned light-activation of myosin. (i) Myosin is inhibited during liposome preparation. (ii) Dissociating the actin cortex at the poles by high-power laser (405 nm) to allow membrane expansion during (iii) light-activation by 488 nm laser along the equatorial plane. d Time-lapse images showing the midplane of a light-activated liposome exhibiting cell division-like deformation. e Side view (y–z plane) and furrow (x–z plane) of a three-dimensional reconstruction of the deformed liposome. f PIV of the contracting actomyosin network (left). Yellow arrows show the total displacement, u, over 5 min with vector magnitudes normalized by the maximum. The colormap represents local strain fields (right). g Mean actin flow speed within the liposome over time (n = 5 liposomes and N = 2 independent experiments in 0 nM; n = 5 and N = 2 in 35 nM; n = 5 and N = 3 in 70 nM; n = 4 and N = 2 in 140 nM; n = 5 and N = 2 in 280 nM). h Contour shape of the liposome membrane over time. Time is color-coded. i Aspect ratio \({\alpha }_{{{{\rm{shape}}}}}\) of the liposome estimated from ellipse fitting to the contour shape. j Aspect ratio over time (n = 13 liposomes and N = 3 independent experiments in 0 nM; n = 20 and N = 2 in 35 nM; n = 39 and N = 4 in 70 nM; n = 28 and N = 6 in 140 nM; n = 15 and N = 2 in 280 nM). k and l Boxplots showing the time at maximum aspect ratio (k) and maximum aspect ratio (j) (n = 13 liposomes and N = 3 independent experiments in 0 nM; n = 20 and N = 2 in 35 nM; n = 39 and N = 4 in 70 nM; n = 28 and N = 6 in 140 nM; n = 15 and N = 2 in 280 nM). Data are presented as mean ± SD (g, j) or boxplots where the interquartile range (IQR) is between Q1 (25th percentile) and Q3 (75th percentile), the center line indicates the median, whiskers are extended to Q3 + 1.5 × IQR and Q1 − 1.5 × IQR (k, l); *p < 0.05; **p < 0.01; ***p < 0.001 in a two-sided Wilcoxon rank sum test. n.s., not significant. Scale bars, 10 μm. Source data are provided as a Source Data file.
We spatio-temporally controlled actomyosin contractility using Blebbistatin, a myosin ATPase inhibitor41. Its inhibitory function is inactivated by exposure to light with a wavelength of 488 nm or shorter42,43,44. F-actin was polymerized for ~10 min at room temperature (~25 °C) after liposome preparation during which myosin was inhibited by Blebbistatin (Fig. 1c, (i)). Membrane-bound actin networks were locally dissociated at the ‘poles’ (referring to the regions at opposite ends of the periphery of the liposome) using a high-power laser with a wavelength of 405 nm to relieve contraction-induced stress (Fig. 1c, (ii)). Myosin was subsequently activated locally at the ‘equator’ using a laser with a wavelength of 488 nm applied at each image acquisition (Fig. 1c, (iii)) (Methods). This led to notable membrane deformation, resulting in the formation of a furrow at the equator enriched with actomyosin, reminiscent of a dividing cell (Fig. 1d, Movie S1). Three-dimensional reconstruction images confirmed furrowing across the equatorial plane (Fig. 1e, Supplementary Fig. 1). The liposome volume was conserved during shape changes (Supplementary Fig. 2). Particle Image Velocimetry (PIV) analysis showed that local contractility activation induced actin flow towards the equator (i.e., contractile foci induced by rupture and delocalization of the network), suggesting that locally elevated contractility may drive cell division-like furrowing (Fig. 1f). Actin flow speed was fastest at the beginning and decreased over time with comparable magnitudes at various His-α-actinin concentrations (Fig. 1g). We confirmed that myosin thick filaments could form in our experimental setup (Supplementary Fig. 3, Supplementary Fig. 4). Throughout the experiments, we used a hyper-osmotic buffer to induce membrane deflation, providing excess membrane to facilitate significant liposome shape changes (Methods). In contrast, a hypo-osmotic buffer did not cause notable membrane deformation, indicating that increased membrane tension may prevent shape changes29 (Methods) (Supplementary Fig. 5).
The liposome contour position was extracted along the membrane from membrane fluorescence images (Methods) (Fig. 1h). Membrane deformation was quantified by the aspect ratio, \({\alpha }_{{{{\rm{shape}}}}}={R}_{{{{\rm{major}}}}}/{R}_{{{{\rm{minor}}}}}\), calculated as the ratio of the major (\({R}_{{{{\rm{major}}}}}\)) to minor axis length (\({R}_{{{{\rm{minor}}}}}\)) of a fitted ellipse (Fig. 1i). We analyzed liposomes of comparable sizes with radius ~20–40 μm across different conditions to reduce variability (Methods) (Supplementary Fig. 1). The aspect ratio significantly depended on His-α-actinin concentration (Fig. 1j, Supplementary Fig. 1, Movie S2). The maximum deformation time peaked around [His-α-actinin] = 70–140 nM (~2.0 min), indicating prolonged shape change (Fig. 1k). Notably, the maximum aspect ratio peaked around [His-α-actinin] = 70 nM (\({\alpha }_{{{{\rm{shape}}}}}^{\max }=1.6\pm 0.3\), mean ± SD) (Fig. 1l). At low [His-α-actinin], weak actin-membrane attachment reduced force transmission to the membrane, while at high [His-α-actinin], dense F-actin crosslinking stiffened the membrane-bound layer, limiting deformation. It has been reported that a specific range of α-actinin concentrations can facilitate actomyosin contraction within bulk actomyosin networks crosslinked by α-actinin45. We observed similar behavior at high concentrations of crosslinkers without a His-tag, which hindered actomyosin contraction and membrane deformation ([α-actinin] = 200 nM and [Fascin] = 400 nM) (Supplementary Fig. 6, Movie S3). Together, these results suggest that fine-tuning the strength of actin-membrane attachment is essential to achieve significant cell division-like membrane deformation.
Equatorial activation with pole relaxation maximizes deformation
We next explored how spatial patterns of actomyosin contractility influence the extent of deformation, including ‘global’ activation throughout the liposome or ‘local’ activation at the equatorial plane, with or without actin cortex dissociation at the poles (Fig. 2a). In the following, we fixed [His-α-actinin]= 70 nM to maximize deformation. Liposomes of comparable sizes were analyzed across different conditions (Supplementary Fig. 7). Without pole relaxation, liposomes exhibited no significant shape changes or polarized shapes reminiscent of cell blebbing (Fig. 2b, (i) and (ii), Movie S4–S7). The cortical actin networks appeared to block contractility-induced membrane expansion, resulting in a wrinkling-like morphology (Fig. 2b, (i) left, (ii) right, Movie S4 and S7). We occasionally observed blebbing-like behavior when pole relaxation with a 405 nm laser was not performed (Movies S5 and S7). This likely occurs because myosin-induced stress increases internal pressure, leading to F-actin breakage or crosslinker unbinding within the actin cortex. When pole relaxation is induced, this pressure increase is mitigated by the expansion of the membrane at the two poles. Indeed, division-like deformation was observed when the actin cortex was dissociated at the poles (Fig. 2b, (iii) and (iv), Movie S1, S8–S10). A bleb-like structure retracted within the furrow (Movie S8). With pole relaxation, the aspect ratio increased significantly over time (Fig. 2c), reaching a notably higher maximum aspect ratio compared to conditions without pole relaxation (Fig. 2d), highlighting the importance of pole relaxation for inducing cell division-like morphology.
a Schematic of light activation patterns. (i) Global activation within the entire liposomes. (ii) Local activation at the equatorial plane. (iii, iv) Pole relaxation by dissociating the actin cortex at the poles using a high-power 405 nm laser. b Snapshots showing the endpoint images of post-deformed liposomes exposed to different light-activation modes. c Aspect ratio over time (n = 20 liposomes and N = 4 independent experiments in global; n = 18 and N = 5 in local; n = 30 and N = 5 in global + pole; n = 39 and N = 4 in local + pole). d Boxplot showing maximum aspect ratio in (c). e Schematic showing the definition of the asymmetry parameter \({P}_{{{{\rm{act}}}}}\). The asymmetry parameter approaches 1 for asymmetric and 0 for symmetric actin distribution. The integral is taken along the membrane contour \({ds}\). f Phase diagram of maximum aspect ratio and asymmetry parameter (n = 20 liposomes and N = 4 independent experiments in global; n = 18 and N = 5 in local; n = 30 and N = 5 in global + pole; n = 39 and N = 4 in local + pole). Small points represent individual liposome. Large symbols are mean ± SD. Significance (*) is based on Fisher’s exact test between ‘local + relax.’ and ‘global + relax.’ with \({P}_{{{{\rm{act}}}}} \, > \, 0.25\) classified as asymmetric \({P}_{{{{\rm{act}}}}} \, < \, 0.25\) as symmetric. g and h Furrow radius over time normalized by its initial value, comparing different light activation patterns (g) (n = 28 liposomes and N = 9 independent experiments in global + pole; n = 40 and N = 9 in local + pole) and comparing local activation + pole relaxation with cofilin or Arp2/3 (h) (n = 40 liposomes and N = 9 independent experiments in control; n = 39 and N = 7 in cofilin; n = 20 and N = 7 in Arp2/3). The dashed line is a guide for furrow speed fitting within 0 to 1.2 min. i Boxplot showing minimum furrow radius in (g) and (h) (n = 28 liposomes and N = 9 independent experiments in global + pole; n = 40 and N = 9 in control; n = 39 and N = 7 in cofilin; n = 20 and N = 7 in Arp2/3). j Boxplot showing furrowing speed calculated from the linear fitting to the furrow radius time course within 0 to 1.2 min in (g) and (h) (n = 28 liposomes and N = 9 independent experiments in global + pole; n = 40 and N = 9 in control; n = 39 and N = 7 in cofilin; n = 20 and N = 7 in Arp2/3). Cofilin, Arp2/3, and His-VCA were used at 0.15 µM, 18 nM, and 1.5 µM, respectively. Data are presented as mean ± SD (c, f–h) or boxplots where the interquartile range (IQR) is between Q1 (25th percentile) and Q3 (75th percentile), the center line indicates the median, whiskers are extended to Q3 + 1.5 × IQR and Q1 − 1.5 × IQR (d, i, j); *p < 0.05; **p < 0.01; ***p < 0.001 in a two-sided Wilcoxon rank sum test. n.s., not significant. Scale bars, 10 μm. Source data are provided as a Source Data file.
We quantified the actin distribution using the actin asymmetry parameter \({P}_{{{{\rm{act}}}}}\) (Fig. 2e). Firstly, we calculated the center-of-mass of the liposome, \({{{{\bf{r}}}}}_{{{{\rm{c}}}}}={N}^{-1}\Sigma {{{{\bf{r}}}}}_{i}\), from membrane positions, \({{{{\bf{r}}}}}_{i}=\left({x}_{i},{y}_{i}\right),{i}=1\cdots N\). The actin fluorescence intensity was measured at each membrane position, \({I}_{{{{\rm{act}}}}}({{{{\bf{r}}}}}_{i})\). Then, the center-of-mass of the actin distribution, \({{{{\bf{r}}}}}_{{{{\rm{act}}}}},\) was calculated as the actin intensity-weighted membrane position, \({{{{\bf{r}}}}}_{{{{\rm{act}}}}}=\Sigma {{{{\bf{r}}}}}_{i}{I}_{{{{\rm{act}}}}}({{{{\bf{r}}}}}_{i})/\Sigma {I}_{{{{\rm{act}}}}}({{{{\bf{r}}}}}_{i})\). Finally, we defined the actin asymmetry, \({P}_{{{{\rm{act}}}}}\equiv \left|{{{{\bf{r}}}}}_{{{{\rm{act}}}}}-{{{{\bf{r}}}}}_{{{{\rm{c}}}}}\right|/{R}_{{{{\rm{major}}}}}\), as the deviation of the actin distribution from the center-of-mass of the liposome, normalized by the major axis length \({R}_{{{{\rm{major}}}}}\) (Fig. 2e). The asymmetry parameter approaches \({P}_{{{{\rm{act}}}}}\to 1\) for an asymmetric, polarized actin distribution, while it approaches \({P}_{{{{\rm{act}}}}}\to 0\) for a symmetric actin distribution.
We correlated liposome shape changes with the actin distribution in a phase plot of the maximum aspect ratio vs. the asymmetry parameter, providing a comprehensive view of how mechanical input (localization of actomyosin) influences morphological output (aspect ratio) (Fig. 2f). Without pole relaxation, both parameters remained low, indicating minor deformation and non-polarized actin distribution (Fig. 2f, global, local). By contrast, pole relaxation increased the aspect ratio by allowing membrane expansion at the poles while retaining a symmetric actin distribution (Fig. 2f, local + relax.). The asymmetry parameters under local and global conditions without pole relaxation are comparable to those under local conditions with pole relaxation. This is because light activation without pole relaxation does not significantly deform the membrane or change the actin distribution from its initial symmetric state. With pole relaxation, local activation induced a more symmetric actin distribution and significant deformation compared to global activation (Fig. 2f, global + relax.). These findings suggest that combining the relaxation of cortex tension at the poles with local contractility patterning tends to induce symmetric and significant cell division-like deformation of the membrane.
Actin turnover and architecture regulate furrowing dynamics
We analyzed furrow radius \({R}_{{{{\rm{f}}}}}\) across the division plane for liposomes exhibiting cell division-like deformation with pole relaxation (Methods) (Fig. 2g, Supplementary Fig. 8). Linear fitting was performed on the furrow radius vs. time plot within the first 1.2 min to estimate the approximate furrowing speed. Liposomes with comparable sizes were analyzed across different conditions (Supplementary Fig. 7). Notably, the minimum furrow radius under local activation was smaller compared to global activation (Fig. 2i), with significantly faster furrowing speed (Fig. 2j). This suggests that, under global activation, contractility at the poles may counteract equatorial contractility and decelerate furrowing46. Therefore, local activation is mechanically more suitable for inducing rapid and significant deformation than global activation.
In the presence of His-mDia1 and His-α-actinin, we added other actin-associated proteins to examine their effects on cell division-like shape changes. Cofilin ([Cofilin]/[Actin] = 0.025) accelerated furrowing (Fig. 2h, j), suggesting that softening of the actin network through enhanced actin turnover by cofilin47,48,49 facilitates cell division-like deformation. The maximum aspect ratio was increased by cofilin (Supplementary Fig. 9, Movie S11). The addition of cofilin did not significantly alter the minimum furrow radius (Fig. 2i). By contrast, phalloidin impeded polar membrane expansion and shape changes, likely due to stabilized and stiffened F-actin50 (Supplementary Fig. 9). A high cofilin concentration ([Cofilin]/[Actin]=0.625) led to actin-membrane tethering breakage and minor deformation (Supplementary Fig. 9). Conversely, branched F-actin nucleation via Arp2/3 with His-VCA ([Arp2/3] = 18 nM; [His-VCA] = 1.5 µM) decelerated furrowing and increased the minimum furrow radius (Fig. 2h–j), indicating that increased cortical F-actin branching stiffened the actin network51 or a branched F-actin architecture hindered myosin force generation39,52. The maximum aspect ratio was decreased by Arp2/3 with His-VCA (Supplementary Fig. 10, Movie S12). We confirmed that the addition of cofilin or Arp2/3 with His-VCA alters the F-actin organization inside the liposomes (Supplementary Fig. 11). These results suggest that the dynamics and the extent of furrowing can be tuned by F-actin turnover and F-actin architecture.
Furrow curvature correlates with parallel F-actin alignment
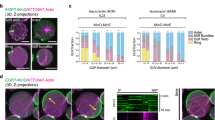
F-actin organization within the furrow plays an important role in contractile ring force generation10,53,54. Thus, we next examined F-actin organization in the furrow of post-deformed liposomes with cell division-like morphology. Notably, the maximum z-projection of confocal slices below the furrow surface exhibited distinct F-actin organization depending on furrow curvature (Fig. 3a). Mean furrow curvature, \({\kappa }_{{{{\rm{f}}}}}\), was calculated by fitting a circle to three separate points at each contour membrane position in the midplane (Methods) (Supplementary Fig. 12). In the moderately deformed liposome with low furrow curvature (\({\kappa }_{{{{\rm{f}}}}}=-0.013\) µm−1), the F-actin network was randomly oriented (Fig. 3a, left). By contrast, in the significantly deformed liposome with high furrow curvature (κf = −0.058 μm−1), the F-actin network aligned parallel to the division plane, forming a contractile ring-like organization within the furrow (Fig. 3a, right).
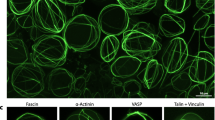
a Snapshots showing the z-projection of confocal z-stacks below the furrow surface of post-deformed liposomes with the low (left), moderate (center), and high (right) furrow curvature. Furrow curvature\(\,{\kappa }_{{{{\rm{f}}}}}\) was calculated from midplane images. b Schematic showing the definition of the orientation order parameter parallel to the division plane. The local order parameter \({S}_{{||}}(x,y)\) is calculated from the neighboring four orientation directors, which takes 1 for parallel and -1 for perpendicular to the division plane. Mean order parameter \(\langle {S}_{{||}}\rangle\) was calculated by averaging \({S}_{{||}}(x,y)\) for all points within the image. c Snapshots showing the actin fluorescence image within the furrow (top) and corresponding \({S}_{{||}}(x,y)\) (bottom). d Snapshots showing the z-projection of confocal z-stacks below the furrow surface of post-deformed liposomes with cofilin (left) or Arp2/3 with His-VCA (right). The actin organization at the furrow and \({S}_{{||}}(x,y)\) are shown below. e Boxplot showing furrow curvature (n = 43 liposomes and N = 16 independent experiments in control; n = 28 and N = 5 in cofilin; n = 16 and N = 6 in Arp2/3). f Boxplot showing the mean order parameter (n = 43 liposomes and N = 16 independent experiments in control; n = 28 and N = 5 in cofilin; n = 16 and N = 6 in Arp2/3). g Scatter plot of furrow radius vs. mean order parameter (n = 43 liposomes and N = 16 independent experiments in control; n = 28 and N = 5 in cofilin; n = 16 and N = 6 in Arp2/3). The dashed lines are linear fitting to individual data. Cofilin, Arp2/3, and His-VCA were used at 0.15 µM, 18 nM, and 1.5 µM, respectively. Data are presented as boxplots where the interquartile range (IQR) is between Q1 (25th percentile) and Q3 (75th percentile), the center line indicates the median, whiskers are extended to Q3 + 1.5 × IQR and Q1 − 1.5 × IQR (e, f); *p < 0.05; **p < 0.01; ***p < 0.001 in a two-sided Wilcoxon rank sum test (e, f) or a two-sided Pearson’s linear correlation coefficient (g). n.s., not significant. Scale bars, 10 μm. Source data are provided as a Source Data file.
We quantified F-actin alignment within the furrow by an order parameter parallel to the division plane (Fig. 3b). First, the local order parameter was computed at each point, \({S}_{{||}}\left(x,y\right)=\cos 2\theta (x,y)\), where \(\theta (x,y)\) is the local filament orientation relative to the division plane. Thus, F-actin aligned parallel to the division plane yields \({S}_{{||}}=1\), while F-actin perpendicular to the division plane yields \({S}_{{||}}=-1\) (Fig. 3b). We then calculated the mean order parameter \(\langle {S}_{{||}}\rangle\), by averaging \({S}_{{||}}\left(x,y\right)\) within the furrow. The liposome with low furrow curvature exhibited a small mean order parameter (\(\left\langle {S}_{{||}}\right\rangle=0.29\) and \({\kappa }_{{{{\rm{f}}}}}=-0.013\) μm−1), whereas the one with high furrow curvature exhibited larger values (\(\left\langle {S}_{{||}}\right\rangle=0.75\) and κf = −0.058 μm−1) (Fig. 3a, c). It is noteworthy that F-actin orientation at the poles is perpendicular to the division plane (Supplementary Fig. 13), reminiscent of a pattern observed during mammalian cell division55,56. Previous theoretical studies suggested that this F-actin organization, parallel to the division plane at the equator and perpendicular at the poles, results from the coupling between cortical actin flow and filament orientation54.
Cofilin ([Cofilin]/[Actin]=0.025) induced a higher order parameter with higher curvature (Fig. 3d, left, and 3e, f), suggesting that softening the actin network through enhanced actin turnover47,48 may contribute to significant deformation and F-actin reorganization. Conversely, Arp2/3 with His-VCA ([Arp2/3] = 18 nM; [His-VCA] = 1.5 µM) disrupted F-actin alignment and reduced deformation (Fig. 3d, right, 3e, f), likely due to enhanced elasticity51 and hindered contraction due to F-actin branching39,52. Notably, the order parameter negatively correlated with furrow curvature, indicating that greater F-actin alignment parallel to the division plane leads to more significant deformation (Fig. 3g).
Convergent cortical flow drives parallel F-actin alignment
While F-actin alignment is correlated with furrow curvature, it remains unclear whether locally increased tension is driving the reorganization of F-actin9, or if flow-alignment coupling is inducing parallel alignment to the division plane10, as observed in living cells. To address this, we observed temporal dynamics of F-actin reorganization during furrowing (Fig. 4a). First, light activation was performed once at the liposome midplane (Fig. 4a, (i)z = R). Then, the focal plane shifted to the bottom surface (Fig. 4a, (ii)z = 0), and kept focusing on the furrow surface (i.e., on the same focal plane as the membrane) (Fig. 4a, (iii)\(z={z}_{{{{\rm{furrow}}}}}\)). This protocol ensures the initial conditions are the same as the prior experimental setup of light activation, where we continually activate and focus on the midplane of the liposome. Before light activation, the F-actin network exhibited random orientation (Fig. 4b), while post-deformed liposomes exhibited parallel alignment to the division plane (Fig. 4c). Notably, after light activation, the F-actin network transitioned from random to parallel alignment, forming a contractile ring-like organization within the furrow over time (Fig. 4d, Movie S13–S15). PIV analysis showed convergent flow towards the furrow (Fig. 4e). Consequently, the mean order parameter increased over time, accompanied by larger negative strain, indicating greater compression (Fig. 4f). The maximum increase in the order parameter (\({\left\langle {S}_{{||}}\right\rangle }_{\max }-{\left\langle {S}_{{||}}\right\rangle }_{\min }\)) strongly correlated with total strain (\(-{\epsilon }_{{{{\rm{tot}}}}}({t}_{{{{\rm{end}}}}})\)) (Fig. 4g). These results suggest that convergent cortical flow-induced compression drives F-actin alignment parallel to the division plane.
a Schematic showing the light activation protocol to track F-actin reorganization dynamics. Light activation was performed once at the midplane (i), then focused on the bottom surface (ii), and kept tracking the F-actin organization at the furrow surface (iii). b F-actin organization at the bottom surface of the liposome and corresponding order parameter before light activation. c F-actin organization at the furrow after deformation has ended. The dashed yellow rectangle is the region cropped in (d). d Time-lapse images showing the F-actin organization at the furrow (top), corresponding orientation field (middle) and order parameter (bottom) calculated within the dashed yellow rectangle. e PIV within the yellow rectangle in (d). Arrows show the total displacement, u, over 5 min, with vector magnitudes normalized by the maximum (left). The colormap represents local strain fields (right). f Mean order parameter and cumulative strain over time within the yellow rectangle in (d). g Scatter plot showing total strain vs. maximum increase of the mean order parameter \({\left\langle {S}_{{||}}\right\rangle }_{\max }-{\left\langle {S}_{{||}}\right\rangle }_{\min }\) calculated within the furrow (n = 26 liposomes and N = 16 independent experiments). The dashed line is a linear fitting. Inset shows the mean order parameter and cumulative strain over time used for the scatter plot. Data are presented as mean ± SD (g). ***p < 0.001 in a two-sided Pearson’s linear correlation coefficient (g). Scale bars, 10 μm. Source data are provided as a Source Data file.
F-actin bending leads to parallel F-actin alignment
When the F-actin network was initially aligned perpendicular to the division plane before light activation, the first light activation failed to induce furrowing (Fig. 5a). However, multiple light activations led to F-actin alignment parallel to the division plane through F-actin bending (Fig. 5b–d, Movie S16). F-actin bending occurs in the cortical flow beneath the membrane, as evidenced by multichannel confocal microscope images, where we visualized the F-actin bending and flows in the same focal plane of the membrane surface (Movies S13–S16). This indicates that, besides the conventional nematic interaction between short rod-like F-actin10,54 (Fig. 5e, (i)), the bending of long F-actin could be an additional mechanical pathway for aligning F-actin parallel to the division plane (Fig. 5e, (ii)). These results explain why multiple light activations were needed to induce significant membrane deformation: a certain level of myosin-induced stress is required to overcome the mechanical energy barrier for F-actin reorganization.
a F-actin organization at the bottom surface of the liposome (left) and corresponding order parameter (right) before the 1st light activation. b Time-lapse images showing F-actin organization at the furrow surface of the liposome in (a) after the 1st right activation (actin and membrane channel), and the end point of division (overlay). The dashed yellow rectangle is the region cropped in (c). c Time-lapse images showing the F-actin organization at the furrow (top), corresponding orientation field (middle), and order parameter (bottom) calculated within the dashed yellow rectangle in (b). d Mean order parameter over time calculated from (c). e Schematic summarizing the F-actin alignment mechanisms within the furrow. The alignment is enforced by the compressive actin flow towards the furrow, where short F-actin aligns via nematic interaction between filaments, while long F-actin aligns through bending. Scale bars, 10 μm. Source data are provided as a Source Data file.
Mechanical power is maximized during elongation
To understand how mechanical energy is utilized during division, we employed a simplified scaling model considering the balance between furrow contractility and pole tension46,57,58. For simplicity, a deforming liposome is approximated as two merged spheres (Fig. 6a). We analyzed liposomes under global activation with pole relaxation since this model better describes global contractility46. The total myosin-generated force was estimated as \({F}_{{{{\rm{myo}}}}}\sim {f}_{{{{\rm{myo}}}}}{\rho }_{{{{\rm{myo}}}}}{V}_{0}d\), where \({f}_{{{{\rm{myo}}}}}=3.4\) pN is the force per single skeletal muscle myosin II59, \({\rho }_{{{{\rm{myo}}}}}=160\) nM is the myosin concentration, \({V}_{0}=(4/3)\pi {R}_{0}^{3}\) is the initial liposome volume with a radius \({R}_{0}\), \(d=0.05\) is the unloaded duty ratio of skeletal muscle myosin II60. Forces acting on the furrow and poles were estimated based on scaling to the geometric parameters. Pole tension was expressed as \({T}_{{{{\rm{p}}}}}\sim {F}_{{{{\rm{myo}}}}}/(2{R}_{0})\). Furrow contractility depended on the furrow radius \({R}_{{{{\rm{f}}}}}\) due to increasing myosin density as the furrow deepens. This effect was approximated as \({\rho }_{{{{\rm{myo}}}}}({R}_{{{{\rm{f}}}}})\sim {\rho }_{{{{\rm{myo}}}}}{({R}_{0}/{R}_{{{{\rm{f}}}}})}^{2}\), assuming myosin density is proportional to the furrow cross-sectional area due to actomyosin contraction within the liposome. We confirmed that the alternative form of myosin density dependence (\({\rho }_{{{{\rm{myo}}}}}\left({R}_{{{{\rm{f}}}}}\right)\sim {R}_{{{{\rm{f}}}}}^{-3}\)), accounting for the contribution of an actin flow-mediated increase in myosin density, does not alter the following results (Supplementary Note 1, Supplementary Fig. 14). Furrowing force \({F}_{{{{\rm{f}}}}}\left({R}_{{{{\rm{f}}}}}\right)\) was determined by the balance between forces acting on the furrow and poles, given by \({F}_{{{{\rm{f}}}}}\left({R}_{{{{\rm{f}}}}}\right)\sim {F}_{{{{\rm{myo}}}}}\left({R}_{{{{\rm{f}}}}}\right)-{F}_{{{{\rm{myo}}}}}\), where \({F}_{{{{\rm{myo}}}}}\left({R}_{{{{\rm{f}}}}}\right)={f}_{{{{\rm{myo}}}}}{\rho }_{{{{\rm{myo}}}}}({R}_{{{{\rm{f}}}}}){V}_{0}d\).
a Schematic showing the geometric parameters of a deforming liposome. The deforming liposome is approximated as two merged spheres. b Snapshots showing the initial and final frames of the deforming liposome. c The contour shape of the liposome extracted at each frame. Time is color-coded. d Mechanical energy of division-like deformation over time (n = 14 liposomes and N = 7 independent experiments). e Mechanical power calculated from (d) over time (n = 14 liposomes and N = 7 independent experiments). f Normalized power over shifted time. The individual time course for each liposome was normalized by the maximum power, \({P}_{{{{\rm{tot}}}}}^{\max },\) and the time is shifted by the time point, \({t}_{{P}_{{{{\rm{tot}}}}}^{\max }}\), at which the power reaches its maximum (n = 14 liposomes and N = 7 independent experiments). g Schematic showing the NADH-coupled assay used for myosin ATP hydrolysis rate measurements. First, ATP is regenerated from ADP with the conversion of PEP into pyruvate (pyr.) by PK. Then, NADH is oxidized to NAD+ with the conversion of pyruvate into lactate (lac.). The decay in NADH fluorescence mirrors the amount of ATP consumed. h Time-lapse images showing a deforming liposome with NADH fluorescence decay. i NADH fluorescence over time normalized by its initial value (n = 16 liposomes and N = 5 independent experiments in NADH only, n = 20 and N = 3 in actin only; n = 15 and N = 7 in dividing). j Total mechanical energy gain (\({\varDelta E}_{{{{\rm{tot}}}}}\)) and total free energy input (\(\varDelta {G}_{{{{\rm{ATP}}}}}\)) due to myosin ATP hydrolysis over time (n = 15 and N = 7). k Total mechanical power (\({P}_{{{{\rm{tot}}}}}\)) and free energy input rate (\({{{\rm{d}}}}{G}_{{{{\rm{ATP}}}}}/{{{\rm{d}}}}t\)) calculated from (j) over time (n = 15 liposomes and N = 7 independent experiments). l The fraction of energy gain (\({\eta }_{{{{\rm{gain}}}}}\)) and energy loss (\({\eta }_{{{{\rm{loss}}}}}\)) calculated from (k) (n = 15 liposomes and N = 7 independent experiments). Data are presented as mean ± SD (d–f, i–l). Scale bars, 10 μm. Source data are provided as a Source Data file.
The total mechanical energy (\({E}_{{{{\rm{tot}}}}}\)) required for division comprises equatorial (\({E}_{{{{\rm{eq}}}}}\)) and polar (\({E}_{{{{\rm{pole}}}}}\)) contributions. At the equator, the furrowing force \({F}_{{{{\rm{f}}}}}\left({R}_{{{{\rm{f}}}}}\right)\) reduces the circumference of the furrow \(2\pi {R}_{{{{\rm{f}}}}}\), while polar tension \({T}_{{{{\rm{p}}}}}\) tends to reduce the surface area at the poles, \({A}_{{{{\rm{p}}}}}=4\pi {R}_{{{{\rm{p}}}}}^{2}\left(1+\cos \theta \right)\), where \({R}_{{{{\rm{p}}}}}\) is the pole radius46 (Fig. 6a). Thus, \({E}_{{{{\rm{tot}}}}}\) is written as \({E}_{{{{\rm{tot}}}}}={E}_{{{{\rm{eq}}}}}+{E}_{{{{\rm{pole}}}}}\), where \({E}_{{{{\rm{eq}}}}}\sim 2\pi {R}_{{{{\rm{f}}}}}{F}_{{{{\rm{f}}}}}({R}_{{{{\rm{f}}}}})\) and \({E}_{{{{\rm{pole}}}}}\sim {A}_{{{{\rm{p}}}}}{T}_{{{{\rm{p}}}}}\). We estimated the mechanical energy using geometric parameters extracted from membrane fluorescence images of deforming liposomes (Methods) (Fig. 6b,c). The total mechanical energy change, \(\varDelta {E}_{{{{\rm{tot}}}}}\left(t\right)={E}_{{{{\rm{tot}}}}}\left(t\right)-{E}_{{{{\rm{tot}}}}}\left(0\right)\), increased over time as division proceeded until it reached a maximum, marking the end of division (Time ~ 150 s, \(\varDelta {E}_{{{{\rm{tot}}}}}^{\max }\sim 1.0\times {10}^{-9}\) J) (Fig. 6d). The mechanical energy at the poles reached a plateau at ~60 s (\(\varDelta {E}_{{{{\rm{pole}}}}}^{\max }\sim 2.0\times {10}^{-10}\) J), reflecting the endpoint of pole elongation, while the equatorial mechanical energy kept increasing due to furrowing (\(\varDelta {E}_{{{{\rm{eq}}}}}^{\max }\sim 8.0\times {10}^{-10}\) J). Notably, the total mechanical power, \({P}_{{{{\rm{tot}}}}}={{{\rm{d}}}}{E}_{{{{\rm{tot}}}}}/{{{\rm{d}}}}t\), reached a maximum at ~60 s (\({P}_{{{{\rm{tot}}}}}^{\max }\sim 9.0\times {10}^{-12}\) J s-1) (Fig. 6e). Simultaneously, the pole mechanical power (\({P}_{{{{\rm{pole}}}}}\)) started decreasing at ~60 s, corresponding to the time point at which elongation finishes and furrowing begins. The mechanical energy landscape at equilibrium verified that the balance of contractile forces at the furrow and poles, varied under different light-activation modes, determines the final stable shape, as described in the previous study46 (Supplementary Fig. 15). It should be noted that while the actomyosin network contracts toward the center, it is not entirely depleted, as seen in the image with higher fluorescence intensity contrast; the actin network remains beneath the membrane in the polar regions (Supplementary Fig. 13).
To clarify the energy allocation between the equator and poles, we shifted the time axis by the time \({t}_{{P}_{{{{\rm{tot}}}}}^{\max }}\) at which mechanical power reaches its maximum, \({P}_{{{{\rm{tot}}}}}^{\max }\), which was normalized as \(P/{P}_{{{{\rm{tot}}}}}^{\max }\) (Fig. 6f). This rescaling reduces variability among liposomes due to differences in time at reaching \({P}_{{{{\rm{tot}}}}}^{\max }\) and its magnitude. Notably, the total mechanical power increased until \({P}_{{{{\rm{tot}}}}}^{\max }\), marking the end of the elongation at which pole mechanical power began decreasing (Fig. 6f). This suggests that elongation is energetically the most costly process during division, driven by both equatorial contraction and pole expansion, followed by a decrease in total energy cost during furrowing. Therefore, the mechanical energy expended in cell division-like deformation is maximized during contractile ring formation and shape elongation before furrowing.
Total free energy input is efficiently utilized for elongation
We have shown that mechanical power peaks during liposome elongation. However, it remains unknown how much of the free energy input derived from adenosine-triphosphate (ATP) hydrolysis by myosin is converted into mechanical power or dissipated, determining the mechanical efficiency of division. To address this, we employed the nicotinamide adenine dinucleotide (NADH)-coupled assay to measure myosin ATP hydrolysis rate during division61,62,63 (Methods) (Fig. 6g). In this assay, ATP hydrolysis produces adenosine-diphosphate (ADP), which is regenerated to ATP, yielding pyruvate. Pyruvate is oxidized with NADH to form lactate and NAD+. Since NADH fluorescence is excited at 340 nm while NAD+ is not, we can quantify myosin ATP hydrolysis rate by measuring the rate of NADH fluorescence decay61,62,63 (Methods) (Fig. 6h, i, Supplementary Fig. 16). We confirmed that NADH fluorescence decay was not due to leaking or bleaching (Supplementary Fig. 17). The measured ATP hydrolysis rate mainly originated from myosin since that of actin was significantly smaller in the present condition (Fig. 6i and Supplementary Fig. 18). Thus, the NADH fluorescence decay rate reflects the myosin ATP hydrolysis rate, which indicates motor activity, as ATP hydrolysis drives the movement of the myosin motor head. The ATP hydrolysis rate of deforming liposomes was comparable to that without deformation ([His-actinin]=0 nM), while lower than previous estimates in water-in-oil droplets63, likely due to residual influence from inactivated Blebbistatin (Supplementary Fig. 18, Movie S17-S23).
Upon global light-activation with pole relaxation, liposomes initiated division, accompanied by declining NADH fluorescence over time (Fig. 6h, i, Movie S17-S19). Note that the ATP hydrolysis rate measurement is restricted for global activation with pole relaxation since 340 nm light inactivates Blebbistatin44. The total hydrolyzed ATP (in µM) was converted to free energy (\(\varDelta {G}_{{{{\rm{ATP}}}}}\) in J) using the initial liposome volume \({V}_{0}\) and the free energy released from ATP hydrolysis under physiological conditions, ~50–70 kJ mol−1 64,65. Both total free energy input (\(\varDelta {G}_{{{{\rm{ATP}}}}}\)) and total mechanical energy (\(\varDelta {E}_{{{{\rm{tot}}}}}\)) increased over time (Fig. 6j). The free energy input per unit time remained relatively constant throughout division (\({{{\rm{d}}}}{G}_{{{{\rm{ATP}}}}}/{{{\rm{d}}}}t\sim 2.2\times {10}^{-11}\) J s−1) (Fig. 6k). Consequently, the efficiency of mechanical energy gain, \({\eta }_{{{{\rm{gain}}}}}\), defined as the ratio of mechanical power to free energy input rate, \({\eta }_{{{{\rm{gain}}}}}={P}_{{{{\rm{tot}}}}}/({{{\rm{d}}}}{G}_{{{{\rm{ATP}}}}}/{{{\rm{d}}}}t)\), mirrored the time course of mechanical power (Fig. 6l). Notably, the efficiency of mechanical energy gain (\({\eta }_{{{{\rm{gain}}}}}\)) and loss, \({\eta }_{{{{\rm{loss}}}}}=1-{\eta }_{{{{\rm{gain}}}}}\), exhibited biphasic time evolution. During the elongation, more energy input was utilized as mechanical power, increasing energy gain from \({\eta }_{{{{\rm{gain}}}}}\sim 0.45\) to\(\,{\eta }_{{{{\rm{gain}}}}}\sim 0.7\), followed by a decrease in energy gain during the furrowing. These results suggest that cell division-like deformation is a combination of energy accumulation and relaxation processes: the early elongation, forming the contractile ring, is energetically costly but dissipates less energy, while the subsequent furrowing requires less mechanical power, leading to greater dissipation.
Discussion
While cell division has been extensively studied regarding intracellular signaling5,11, less is known about its mechanical aspects5,11,66, such as how contractility patterning leads to successful division through actomyosin flows and F-actin reorganization. We isolated the mechanical contribution of actomyosin using an in vitro cell model to explore how contractility patterning regulates membrane deformation during division. Some previous in vitro studies have observed contractile ring-like structures in water-in-oil droplets and GUVs32,33,67. In these cases, the actin ring is pre-assembled or polymerized without the presence of actomyosin flows. Such an actin ring is relevant to fission yeast division, where a ring forms ‘de novo’ via F-actin polymerization at the equator68,69. By contrast, animal cells do not form an independent equatorial ring during division. Instead, a continuous actomyosin network accumulates within the furrow, often referred to as a ‘ring’, and maintains cell shape stability through force balance with the actomyosin network at the poles9,10,54. In this context, our study examines the dynamic formation of a contractile ring through imbalances in forces and flows emerging from an initially uniform actomyosin network, which is relevant to animal cell division.
By combining a light-activatable actomyosin contraction system in GUVs, developed in our recent study40, with spatio-temporal light patterning, we enabled a quantitative analysis of division-like membrane deformation dynamics coupled with actomyosin flows. This system establishes a clear causal link between myosin-induced stresses and membrane deformations, marking a significant advancement over previous studies on contraction-induced shape changes in GUVs31,32,33. Earlier works lacked the precise spatio-temporal control of myosin activity, making it difficult to quantitatively correlate myosin-induced stresses with shape changes. Additionally, it was challenging to separate the effects of myosin-induced stresses from other mechanical factors, such as actin depolymerization-induced contraction70 and actin polymerization-induced stress71, thereby limiting the clarity of the results interpretation.
Our results showed that pole relaxation is essential for significant membrane deformation, enhanced by local, rather than global, contractility activation. Compressive actin flow aligned F-actin parallel to the division plane via F-actin bending, forming a contractile ring-like structure. It is noteworthy that ring-like F-actin organization coexists with a disk-like structure in our system. Nevertheless, the F-actin network at the furrow surface aligns parallel to the division plane, supporting the existence of a ring-like F-actin structure surrounding the periphery of the equatorial furrow. The mechanical power of division was maximized during liposome elongation before furrowing. These findings highlight the importance of contractility patterning and mechanical energy allocation between the equator and poles, providing insights into cell division mechanics.
Contractility patterning in cell division involves RhoA signaling pathways6,7,8. However, whether local RhoA activation is necessary for division5,11,72 or if global activation is sufficient may depend on cell types18,20. For instance, HeLa cells require RhoA, while RhoA activity is suppressed during Rat1A cell division73, relying on equatorial collapse through Arp2/3 inactivation21. Thus, the minimal mechanical conditions that enable cell division-like shape change remain unknown. Our results demonstrated that pole relaxation drives division-like deformation regardless of contractility pattern. In cells, reducing cortical stiffness at poles via local mDia1 inhibition is necessary for division15 or optogenetic myosin inactivation at poles accelerates division, consistent with our findings16. Thus, our study provides mechanical insights that the relaxation of cortex tension at the poles sufficiently induces significant membrane deformation akin to cell division.
Global contractility activation via constitutively active RhoA destabilizes cell shape, causing cytokinesis failure74, while global RhoA activation using optogenetics leads to shallower furrow compared to local equatorial activation75. Our results showed that global contractility tends to cause asymmetric division or polarization, whereas local activation results in symmetric, faster, and deeper furrowing, consistent with cellular observations that polar contractility destabilizes division symmetry17. Thus, local contractility activation is more effective than global activation in inducing symmetric cell division-like deformation. Interestingly, asymmetric division is preferred in certain cell types for polarization, such as neuroblasts of C. elegans and Drosophila22,23, where global activation would be beneficial. Thus, we suggest that contractility patterning may primarily determine cell shape stability and resulting functions, such as polarized morphology during development and cell migration, or symmetric shape changes during cell division.
Actin-associated proteins crucial for cell division include those affecting furrowing speed, such as actin crosslinking and turnover11,76. Increased α-actinin concentration slows cytokinesis of NRK cells77, aligning with our findings, likely due to stiffened actomyosin networks with higher viscoelastic resistance46. Cofilin depletion causes cytokinetic failure78 and a thicker cortex in mitotic HeLa cells79, while its mechanical roles remained controversial76. Our results showed that cofilin can accelerate furrowing, suggesting that softened networks with enhanced turnover facilitates contraction47,48. These findings provide mechanical insights into furrowing speed regulation in cells.
During division, Rac1, which activates Arp2/3, is suppressed at the equator80. Expressing a dominant-negative MgcRacGAP mutant, which prevents equatorial Rac1 suppression, decelerates furrowing in HeLa and Rat1A cells73. Similarly, cells with constitutively active WASp mutants (an Arp2/3 activator) exhibit slower furrowing81. We observed that Arp2/3 with His-VCA decelerate furrowing, indicating that the Arp2/3-nucleated branched network may inhibit myosin force generation39,52,63. Our results suggest that F-actin architecture may be optimized in cells for prompt division.
The role of F-actin organization in contractile ring force generation has been extensively studied. Actomyosin flow towards the equator aligns F-actin parallel to the division plane54, as observed in C. elegans zygote10, which may originate from nematic interactions between short F-actin filaments. We suggest that F-actin bending induced by compressive cortical actin flow also contributes to F-actin alignment. Given that mDia1-mediated F-actin polymerization dominates during cell division37,38, such bending-induced alignment could potentially occur in cells. Observing this dynamic process in living cells is challenging due to current imaging limitations, as super-resolution imaging like structured illumination microscopy (SIM) are performed on fixed cells82,83, and polarization microscopy is limited for indirect measurement of F-actin orientation9. Thus, the dynamic F-actin reorganization process during contractile ring formation has been overlooked. Since our in vitro model of dividing cells allows visualization of F-actin reorganization dynamics, it will offer insights into the process of contractile ring formation.
The balance between equatorial and polar contractility is crucial for cell division17,58,46,57, but the underlying mechanics are not well understood. We found that mechanical power is maximized during elongation, while the furrowing expended less power, exhibiting a biphasic behavior. Furthermore, cellular energetics, particularly utilizing ATP84, is vital for understanding cell state transitions including division. Although the total amount of ATP or ATP synthesis rate decreases during cell division85,86, overlapping cellular processes, such as protein synthesis, obscure the origin of energy changes. Combining in vitro reconstitution and myosin ATP hydrolysis rate measurement, we demonstrated that cell division-like deformation is a biphasic process: efficient energy gain during contractile ring formation and shape elongation, followed by energetically inefficient furrowing with greater dissipation. These findings provide important mechanical and energetic insights into cell division.
It is important to clarify several assumptions and limitations in our mechanical power estimates. First, while myosin is known for its load-dependent behavior, we used an unloaded myosin duty ratio and force, which could potentially increase the net myosin-induced stress during membrane shape changes87. However, the use of the unloaded myosin duty ratio is validated by our observations (Supplementary Fig. 5): in the hypo-osmotic buffer, where no deformation occurs due to increased membrane tension, myosin is stalled and highly loaded. Conversely, in the hyper-osmotic buffer, deformation occurs with a deflated membrane, indicating that myosin is not stalled and remains unloaded during this process. Additionally, we noted that Blebbistatin slows down ATP consumption compared to our previous work63, suggesting that not all myosin is activated due to incomplete Blebbistatin inactivation. As a result, there may be a lower effective myosin concentration due to this incomplete inactivation. Therefore, our mechanical power estimates represent an upper bound.
It is worth mentioning about the role of osmotic pressure during division-like shape changes. Our findings indicate that membrane deformation is limited under hypo-osmotic condition due to increased membrane tension, suggesting that osmotic pressure constrains liposome deformation toward the end of division (Supplementary Fig. 5). In living cells, osmotic pressure is a major counteracting force to actomyosin contraction during cell rounding88, playing an important role in generating more blebs to maintain symmetric division89. Notably, the role of pressure-mediated stabilization in cell division symmetry, particularly through polar blebs acting as pressure bulbs to release cortical tension, has been demonstrated in dividing cells17. Thus, studying the role of osmotic pressure and membrane channels in actomyosin-driven liposome shape changes presents an interesting avenue for future research.
We note that the geometry of dividing liposomes in our study differs from that in the previous theoretical study of cell division due to the presence of a volumetric network46, but we believe that the direction of contractility at the furrow, the mechanism of force balance between the equator and poles, and the scaling form of the mechanical energies are approximately applicable. It is important to note that our goal is to provide an approximate, scaling-based estimate of mechanical work, rather than a detailed description of the combined deformation mechanics of the membrane and actomyosin cortex, including the influence of volume and surface area. More detailed descriptions will be pursued through quantitative theoretical models and numerical simulations, incorporating complex curved 3D geometries with nematic alignment90 and volumetric contributions such as bulk elasticity and osmotic pressure91. We hope our study serves as a starting point for a more detailed exploration of cell division mechanics through theory and in vitro reconstitutions.
Several factors may limit furrow depth in our system. For example, F-actin breakage or detachment from the membrane can reduce force transmission. Given a critical F-actin breakage force of ~200–400 pN92 and stress generated by myosin thick filaments (~1 μm with ~235 motors93) of ~40-230 pN considering load-dependence59,60,94, stress relaxation due to F-actin breakage could reduce net force applied to the membrane and limit furrow depth95. Some contractile networks appear to not span the entire furrow, which may be due to actin network breakage or crosslinker unbinding during contraction. This breakage or crosslinker unbinding could potentially lead to the relaxation of accumulated stress52,96, resulting in less significant furrowing. Completion of furrowing might be achieved by fully localizing the actomyosin network beneath the membrane in future studies26,35,97, using membrane-associated proteins septin98 and anillin99 to mimic cellular actin cortex and cytokinetic rings11, enhancing contractility transmission. Additionally, non-muscle myosin IIA (NMIIA)100 could produce greater forces and exhibit more significant load-dependent behavior compared to skeletal muscle myosin101,87, which may be important during cell division where cortical stiffness increases 10-fold102. Despite these limitations, our experimental system serves as a robust platform for studying the relationship between cytoskeletal force generation and membrane deformation, contributing to our understanding of cell morphodynamics.
Methods
Lipid composition
Giant unilamellar vesicles (GUVs, liposome) are formed with a combination of L-α-phosphatidylcholine from egg yolk (EPC) at 90% (840051; Avanti), 1, 2-dioleoyl-sn-glycero-3-[n(5-amino-1-carboxypentyl) iminodiacetic acidsuccinyl nickel salt at 8% (18:1 DGS-NTA(Ni)) (790404; Avanti), 1, 2-distesaroyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] at 1% (18:0 PEG2000 PE) (880120; Avanti) to reduce non-specific adhesion of proteins to the membrane40,63, and 1, 2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-[(dipyrrometheneboron difluoride)butanoyl] (18:1 TopFluor® PE) (810282; Avanti) imaged by a 488 nm laser for global activation, and 1, 2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-(Cyanine 5) (18:1 Cy5 PE) imaged by a 640 nm laser for local activation (810335; Avanti) at 1% to avoid light activation of myosin during membrane imaging.
Buffer composition
Buffers were prepared by following the method described in previous studies35,36,39,103. The inner actin polymerization (IP) buffer contains 50 mM KCl, 2 mM CaCl2, 5 mM MgCl2, 10 mM HEPES (pH 7.6), 0.8 mM DTT, 5 mM ATP, 50 mg ml−1 dextran, 175 mM sucrose. Myosin was kept inactivated by adding 68 µM Blebbistatin (B0560; Sigma-Aldrich)42,43 in the buffers. ATP is regenerated through pyruvate kinase (PK) and phosphoenolpyruvate (PEP)-based ATP regeneration system used at 1 mM PEP (10108294001; Roche) and 10 U mL−1 PK (P9136; Sigma-Aldrich). The outer buffer (OB) contains 0.2 mM CaCl2, 0.2 mM MgCl2, 10 mM HEPES (pH 7.6), 6 mM DTT, 2 mM ATP, 380 mM glucose, 0.02 mg ml-1 caseins. The osmolarity of the OB is adjusted with glucose such that the osmotic pressure in the outer buffer is ~40 mOsm/kg larger than the inner buffer measured by an automatic osmometer (5004 Micro-Osmette; Precision Systems, Inc). The outer buffer is approximately 355 mOsm/kg and the liposome is in a slightly deflated condition to allow membrane deformation. A hypo-osmotic buffer (osmotic pressure in the outer buffer is ~40 mOsm/kg lower than in the inner buffer) did not cause notable membrane deformation (Supplementary Fig. 5). The storage buffer for actin (G-buffer) contains 2 mM Tris-HCl (pH 8.0) and 0.1 mM CaCl2, 0.2 mM ATP, and 0.5 mM DTT, and 22 µM actin. The myosin storage (MS) buffer contains 4.5 M KCl, 0.1 M HEPES (pH 7.0), and 21 µM myosin. The spin-down (SD) buffer contains 0.01 mg ml−1 phalloidin, 13 µM actin, 4.2 µM myosin (in MS), 460 mM KCl, 2 mM CaCl2, 5 mM MgCl2, 10 mM HEPES (pH 7.6), 0.8 mM DTT, 5 mM ATP. The storage buffer for NADH and PEP contains 100 mM NaHCO3, and 14 mM NADH or 48.5 mM PEP, respectively. The storage buffer for PK and LDH contains 10 mM HEPES (pH 7.4), 100 mM KCl, and 2 KU mL−1 PK or 2.6 KU mL–1 LDH, respectively. All the regents were purchased from Sigma-Aldrich. The specific concentration of ATP in IP buffer has been optimized for actomyosin contraction in our assay, as developed in previous studies40,63. It is noteworthy that we benchmarked the mechanical power of actomyosin contraction and found it was slightly decreased under lower concentrations at 1 mM ATP in our previous study63. It should be noted that the ATP regeneration system maintains the ATP concentration constant, so that the absolute amount of ATP is not crucial for the regeneration system to work.
GUVs preparation
Actomyosin-encapsulated GUVs were prepared using the method described in previous studies26,34,35,104. All lipids were stored in chloroform at –20 °C. For preparation, the desired lipids were combined in a glass vial and dried under a stream of Argon gas to remove the solvent. The dried lipids were then dissolved in mineral oil (M5904; Sigma-Aldrich) at 2 mg ml−1. This lipid-oil mixture was sonicated in a bath sonicator for 2 h at room temperature. The mixture was then cooled to room temperature and stored at 4 °C up to a week. Next, the protein mixture was prepared as the inner phase on ice before starting GUV production. First, a solution of 22 μM monomeric actin containing 10% of labeled rhodamine-actin conjugate is obtained by incubating the actin solution in G-Buffer overnight at 4 °C. Myosin was centrifuged in the presence of polymerized actin in SD buffer to isolate the catalytically active myosin dimers before use39. Briefly, actin is polymerized for 1 h at 4 °C in SD buffer. The actin network was spiked with ATP to 1 mM, and myosin in MS buffer was added. The actin-myosin mix was then centrifuged at 4 °C for 30 min at 128360 g and the supernatant was collected. The proteins were combined in the IP buffer at the concentrations of 6 µM actin, 160 nM myosin, 300 nM His-mDia1, 70 nM His-actinin, 3 µM profilin. Proteins were freshly thawed from –80 °C fridge before use. For the ATP hydrolysis rate measurement, the NADH-coupled assay was combined with protein mix at the concentrations of 0.67 mM NADH, 1.4 mM PEP, 20 U mL−1 PK, and 32 U mL−1 LDH. To form the GUVs, in a 0.65 mL tubes (07200185; Corning Costar), 70 µL of lipid-oil mixture was added for emulsification. Then 7 µL of the protein mix was added to the lipid-oil mixture. This mixture was then syringed in a glass syringe (Hamilton) 2 to 3 times to produce emulsion. Separately, in a low absorption 0.65 mL tube, 30 µL of mineral oil was added to the top of 30 µL of outer buffer (OB). Then, the 60 µL of emulsion was added to the top of the mineral oil layer in the tube. This mixture was then centrifuged at 100 g for 15 min at 4 °C to produce GUVs. In the meantime, the 150 µL of OB was injected to coverslips. After the centrifugation, all the upper oil layer was gently aspirated, and approximately 30 µL of the bottom liposomes were then aspirated. The liposomes were gently injected into OB. F-actin polymerization was allowed for ~10 min at R.T. (~25 °C) after preparing liposomes. Thereafter, myosin was light-activated, and the dynamics of membrane deformation were recorded by timelapse images with 6 s intervals using Andor iQ software. Liposomes were imaged by Leica DMi8 inverted microscope equipped with a spinning disk confocal and a 63 × 1.4-NA oil immersion lens (Leica Microsystems).
NADH-coupled assay
Nicotinamide adenine dinucleotide (NADH)-coupled assay consists of pyruvate kinase (PK) and phosphoenolpyruvate (PEP)-based ATP regeneration system, together with lactic dehydrogenase (LDH) that oxidize NADH61,62,63. The final concentration of NADH (10128023001; Roche) is 0.67 mM, PEP (10108294001; Roche) is 1.4 mM, PK (P9136; Sigma-Aldrich) is 20 U mL−1, and LDH (L1254; Sigma-Aldrich) is 32 U mL−1. To minimize the impact of NADH assay-related enzymatic systems on the conventional actin polymerization buffer, we selected an optimal NADH concentration as low as possible but having detectable NADH signal for our measurements. Fluorescence intensity of NADH within liposomes was analyzed using Fiji/ImageJ (NIH), Microsoft Excel, and custom code written in MATLAB. The mean intensity within a circle with a half-radius of a liposome was determined and subtracted by the mean intensity of the background region. The photobleaching was corrected using exponential fitting method105 for the liposomes encapsulating NADH without any proteins (Supplementary Fig. 17). To relate the fluorescence intensity to the NADH concentration, calibration measurements were performed by adding known amounts of NADH in water-in-oil emulsion droplets63 (Supplementary Fig. 16). Because of the 1:1 stoichiometric ratio between ADP and NADH63, the slope of the NADH fluorescence decay gives ATP hydrolysis rate.
Protein concentrations
The final concentration of non-fluorescent actin (AKL99-D; Cytoskeleton Inc.) rhodamine-actin (AR05-C; Cytoskeleton Inc.) and skeletal muscle myosin II (MY02; Cytoskeleton Inc.) were 5.3 µM, 0.7 µM, and 160 nM, respectively. His-mDia1 (Methods), His-α-actinin (ab207132; abcam), profilin (PR02; Cytoskeleton Inc.) were used at 300 nM, 35-280 nM, and 3 µM, respectively. Arp2/3 (RP01P-B; Cytoskeleton Inc.) and His-VCA (Methods) were used at 18–90 nM and 1.5 µM, respectively. Cofilin (CF01; Cytoskeleton Inc.), Fascin (CS-FSC01; Cytoskeleton Inc.), and gelsolin (HPG6; Cytoskeleton Inc.) were used at 0.15-3.75 µM, 400 nM, 50 nM, respectively.
Protein purification
His-tagged purified proteins were prepared by following the method described in a previous study. Briefly, SNAP tagged mDia1(FH1C) containing a carboxy-terminal His(6x)-tag was expressed in E.coli (BL21-Codon Plus (DE3)-RP strain). The cells were induced with 500 µM IPTG at 16 °C and purified using Talon® metal affinity resin by following the manufacturer’s instructions. The mammalian SNAP-mDia1 (FH1C)-HIS plasmid was a gift from David Kovar lab. N-WASP fragment WWA (aa400–501, also called VCA) containing a carboxy-terminal His(6x)-tag was expressed in E.coli (BL21-Codon Plus (DE3)-RP strain). The cells were induced with 1 mM IPTG for 2 h at 37 °C and purified using Ni-NTA resin by following the manufacturer’s instructions, followed by another purification using the size exclusion chromatography. The pET17b KCK-ratWWA-6xHis plasmid was a gift from Cecile Sykes lab.
Fluorescent labeling of skeletal muscle myosin
Skeletal muscle myosin (rabbit, MY02; Cytoskeleton Inc.) is labeled with Alexa Fluor 647 nm C2 Maleimide under reducing conditions, following a previously described method39. Briefly, myosin is reduced in labeling buffer (50 mM HEPES, 0.5 M KCl, 1 mM EDTA, 10 mM DTT, pH 7.6) and then dialyzed overnight in a DTT-free labeling buffer. The dialyzed myosin solution is centrifuged to remove insoluble matter, and the supernatant is labeled with Alexa Fluor 647 nm C2 Maleimide, added in a 5-fold molar excess relative to myosin. The labeling reaction is performed for 1 h at 4 °C and quenched by adding 1 mM DTT. The labeled myosin is collected using a desalting column (Pierce, 5 K MWCO, 5 mL). Optical absorbance is measured at 280 nm and 647 nm to determine the concentration and degree of labeling.
Microscopy and data acquisition
Images were acquired by Leica DMi8 inverted microscope equipped with a 63 × 1.4-NA oil immersion lens (Leica Microsystems), a spinning-disk confocal (CSU22; Yokagawa), and sCMOS camera (Zyla; Andor Technology) controlled by Andor iQ3 (Andor Technology). One pixel measured 0.0938 μm pix-1. Images were recorded by timelapse with 6 s intervals up to 3–10 min after focusing on the midplane of the liposomes at room temperature (~25 °C). The NADH fluorescence was imaged using a filter set consisting of 350 ± 50 nm excitation filter, 415 nm long-pass dichroic, and 460 ± 50 nm emission filter (Leica). Illumination from a mercury halide light source (11504120; Leica EL6000 Metal Halide) was used to excite the NADH. The excitation source was shuttered after every acquisition to minimize the photobleaching.
Light activation experiments
The spinning disk confocal system equipped with a FRAPPA module (Andor Technology) was used to perform local light activation and rapidly switch the light back to collect images using the spinning disk43. For global activation, exposure times were 300 ms for 488 nm laser (10% power, 10 mW) for visualization of membrane and inactivation of Blebbistatin, 300 ms for 561 nm laser (10% power, 7.5 mW) for visualization of rhodamine-actin, and pole relaxation using high-power FRAP 405 nm laser (10% power, 10 mW) with 20 µs dwell time and 3 repeats within a square region with one side approximately 0.25 × 2R μm (R, liposome radius). For local activation, exposure times were 300 ms for 640 nm laser (10% power, 7.5 mW) for visualization of membrane, 300 ms for 561 nm laser (10% power, 7.5 mW) for visualization of rhodamine-actin, and local activation using 488 nm laser (1% power, 0.5 mW) with 20 µs dwell time and 5 repeats within a rectangular region with shorter side approximately 0.20 × 2R μm, and pole relaxation using high-power FRAP 405 nm laser (10% power, 10 mW) with 20 µs dwell time and 3 repeats within a square region with one side approximately 0.25 × 2R μm. The laser intensities were set to acquire a sufficient number of images and enough contrasts while keeping photobleaching minimal.
GUV sizes used for analysis
The liposomes with radius R approximately ~20–40 µm were analyzed in this study. It should be noted that we displayed the size distribution for liposomes used in each experiment and ensured that they were not significantly different across various conditions used for comparison (Supplementary Fig. 1, Supplementary Fig. 6, Supplementary Fig. 7, Supplementary Fig. 9, Supplementary Fig. 10). Technically, the upper limit of R~40 µm is due to the limits of region of interest in our confocal microscope with a 63× objective. On the other hand, due to the low resolution of small liposomes in our confocal microscopy magnification, it was difficult to locally break the cortex at the poles without affecting internal actomyosin contractility, as the local light exposure may influence the surrounding network. Therefore, we used larger liposomes to avoid impacts outside the poles. It is worth noting that for much smaller liposomes (e.g., R < 10 µm), significant shape changes were not observed. This may be because the total stress per unit area potentially changes with liposome size. Assuming the concentration of myosin within liposomes of different radii is constant, smaller liposomes experience less myosin-induced stress per unit surface area due to their larger surface-to-volume ratio (\(S/V\sim 1/R\)). Consequently, smaller liposomes could experience significantly smaller contractile force per unit area of the membrane compared to larger ones. Thus, we restricted our analysis to liposomes larger than R∼20 µm, where we were able to observe significant division-like membrane deformation. We ensured that the liposome size distribution used for analysis was comparable across different conditions used for comparison. Nevertheless, our results suggest that the impacts of actin-associated proteins and different light activation patterns on the extent of membrane deformation outweigh the variability due to differences in liposome size across different conditions.
Image analysis and data quantification
Particle Image Velocimetry (PIV) analysis
Quantitative image analysis on contracting actomyosin network was performed by using a custom code written in MATLAB. We measure the displacement vectors between individual frames at every spatial location \({{{\bf{u}}}}\left({{{\bf{r}}}},t\right)\) using a public domain Particle Image Velocimetry (PIV) program implemented as a Fiji/ImageJ plugin. The first, second, and third PIV window size and search window size were set to 160 and 329, 100 and 200, 60 and 120 pixels, respectively. Correlation, noise, and threshold parameters were set to 0.6, 0.20, and 5, respectively. PIV was performed between each frame with 6 s intervals. Mean compressive strain ε of the actin network is calculated from the divergence of the cumulative displacement field, \(\varepsilon (t)={\int }_{0}^{t}d{t}^{{\prime} }\int {{{\rm{d}}}}{{{\bf{r}}}}[\nabla \cdot {{{\bf{u}}}}\left({{{\bf{r}}}},{t}^{{\prime} }\right)]/\int {{{\rm{d}}}}{{{\bf{r}}}}\). The spatial integral is taken over the negative divergence within a liposome to calculate the compressive strain.
Nematic order parameter calculation
We measure the orientation field of F-actin using a public domain program OrientationJ implemented as a Fiji/ImageJ plugin. The image of the bottom surface of the liposomes were used to calculate the orientation field. The local nematic order parameter was computed from the director field around the reference point, \(S=\langle \cos 2\theta \rangle\), where 〈⋅〉 is the local average around the reference point. The four surrounding points were averaged to compute the local orientation order parameter. The mean nematic order parameter, 〈S〉, is calculated by averaging all the local nematic order parameter.
Contour analysis of the membrane deformation
The position of the membrane was extracted from the image of the membrane channel using a public domain program JFilament implemented as a Fiji/ImageJ plugin. The extracted contour position of the membrane was imported to MATLAB. The center-of-mass of the liposome, \({{{{\bf{r}}}}}_{{{{\rm{c}}}}}={N}^{-1}\Sigma {{{{\bf{r}}}}}_{i}\), was calculated from the position of the membrane, \({{{{\bf{r}}}}}_{i}=\left({x}_{i},{y}_{i}\right),{i}=1\cdots N\). The origin of the polar coordinate was then centered at the center-of-mass of the liposome by subtracting the center-of-mass of the contour position from each position of the membrane as \({{{{\bf{r}}}}}_{i}-{{{{\bf{r}}}}}_{{{{\rm{c}}}}}\). To calculate the aspect ratio of a liposome \({\alpha }_{{{{\rm{alpha}}}}}={R}_{{{{\rm{major}}}}}/{R}_{{{{\rm{minor}}}}}\), the major axis length \({R}_{{{{\rm{major}}}}}\) and minor axis length \({R}_{{{{\rm{minor}}}}}\) of a liposome was determined by fitting an ellipse to the contour position (Fig. 1i). The local curvature of the membrane was determined by fitting a circle to the three separated points around each membrane position by Pratt-Newton method (Supplementary Fig. 12). The geometric parameters were extracted from the contour position of the membrane for the calculation of mechanical energy (Fig. 6). Briefly, the furrow radius \({R}_{{{{\rm{f}}}}}\) and \({R}_{{{{\rm{major}}}}}\) was directly extracted from the image through kymograph analysis (Supplementary Fig. 8) and fitting an ellipse (Fig. 1i), respectively. Thereafter, the pole radius \({R}_{{{{\rm{p}}}}}\) and \(\cos \theta\) was determined by the relationship\(\,{R}_{{{{\rm{p}}}}}=({R}_{{{{\rm{major}}}}}^{2}+{R}_{{{{\rm{f}}}}}^{2})/(2{R}_{{{{\rm{major}}}}})\) and \(\cos \theta={({R}_{{{{\rm{p}}}}}^{2}-{R}_{{{{\rm{f}}}}}^{2})}^{1/2}/{R}_{{{{\rm{p}}}}}\) using \({R}_{{{{\rm{f}}}}}\) and \({R}_{{{{\rm{major}}}}}\).
Particle Tracking Velocimetry (PTV) analysis
Particle-tracking analysis was performed by using the Fiji plugin ParticleTracker106 to extract the position of the myosin spots (Supplementary Fig. 3). The particle detection radius, cutoff, per/abs, link range, and displacement were set to 15, 0.001, 0.5, 2, and 10, respectively. Dynamics was set to Straight lines mode. The extracted particle positions were imported into MATLAB for instantaneous velocity analysis.
Statistics & reproducibility
Statistical tests comparing distributions were done with the Wilcoxon rank sum test. A two-sided Pearson’s correlation coefficient was used to assess the relationship between two variables. All data displayed as a single value with an error bar is quoting the mean +/- standard deviation. Data are presented as boxplots where the interquartile range (IQR) is between Q1 (25th percentile) and Q3 (75th percentile), the center line indicates the median, whiskers are extended to Q3 + 1.5 × IQR and Q1 − 1.5 × IQR. The symbols *, **, and *** represent p < 0.05, 0.01, and 0.001 respectively. The sample size was determined by experimental feasibility and adjusted when it was deemed sufficient, ensuring that sample statistics were not affected by changes in sample size. Liposomes lacking actin fluorescence were excluded from the analysis as their behavior should not be affected by actomyosin contractility. Sample sizes (n) are indicated at the relevant locations in the manuscript. Experiments were independently replicated at least two times, and all replications were successful. Independent experiments (N) are indicated at the relevant locations in the manuscript.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data supporting the finding of this manuscript are available from the corresponding authors upon request because of the large size of the time lapse image data. The data generated in this study are provided in the Supplementary Information and Source Data file. Source data are provided with this paper.
Code availability
Code supporting the findings of this manuscript are available from the corresponding authors upon request. Fiji [https://imagej.net/software/fiji/downloads] was used for basic image processing. Particle image velocimetry (PIV) was performed in Fiji using a public domain plugin [https://sites.google.com/site/qingzongtseng/piv#h.39ycjstb0wmn]107. Orientation analysis was performed in Fiji using a public domain plugin [https://bigwww.epfl.ch/demo/orientation/]108. Particle tracking velocimetry (PTV) was performed using a public domain plugin [https://sbalzarini-lab.org/?q=downloads/]106. MATLAB (R) version 2021b was used for data analysis and graph production. A reporting summary for this Article is available as a Supplementary Information file.
Change history
03 December 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41467-024-54985-6
References
Zernicka-Goetz, M. Cleavage pattern and emerging asymmetry of the mouse embryo. Nat. Rev. Mol. Cell Biol. 6, 919–928 (2005).
Fares, J., Fares, M. Y., Khachfe, H. H., Salhab, H. A. & Fares, Y. Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct. Target. Ther. 5, 28 (2020).
Mao, Y. & Wickström, S. A. Mechanical state transitions in the regulation of tissue form and function. Nat. Rev. Mol. Cell Biol. 25, 654–670 (2024).
Hosea, R., Hillary, S., Naqvi, S., Wu, S. & Kasim, V. The two sides of chromosomal instability: drivers and brakes in cancer. Signal Transduct. Target. Ther. 9, 75 (2024).
Eggert, U. S., Mitchison, T. J. & Field, C. M. Animal cytokinesis: from parts list to mechanisms. Annu. Rev. Biochem. 75, 543–566 (2006).
Fededa, J. P. & Gerlich, D. W. Molecular control of animal cell cytokinesis. Nat. Cell Biol. 14, 440–447 (2012).
Chircop, M. Rho GTPases as regulators of mitosis and cytokinesis in mammalian cells. Small GTPases 5, e29770 (2014).
Pollard, T. D. & O’Shaughnessy, B. Molecular mechanism of cytokinesis. Annu. Rev. Biochem. 88, 661–689 (2019).
Spira, F. et al. Cytokinesis in vertebrate cells initiates by contraction of an equatorial actomyosin network composed of randomly oriented filaments. Elife 6, e30867 (2017).
Reymann, A. C., Staniscia, F., Erzberger, A., Salbreux, G. & Grill, S. W. Cortical flow aligns actin filaments to form a furrow. Elife 5, e17807 (2016).
Green, R. A., Paluch, E. & Oegema, K. Cytokinesis in animal cells. Annu. Rev. Cell Dev. Biol. 28, 29–58 (2012).
Stricker, J., Falzone, T. & Gardel, M. L. Mechanics of the F-actin cytoskeleton. J. Biomech. 43, 9–14 (2010).
Kiyomitsu, T. & Cheeseman, I. M. Cortical dynein and asymmetric membrane elongation coordinately position the spindle in anaphase. Cell 154, 391–402 (2013).
Rodrigues, N. T. et al. Kinetochore-localized PP1-Sds22 couples chromosome segregation to polar relaxation. Nature 524, 489–492 (2015).
Chen, A. et al. Inhibition of polar actin assembly by astral microtubules is required for cytokinesis. Nat. Commun. 12, 2409 (2021).
Yamamoto, K. et al. Optogenetic relaxation of actomyosin contractility uncovers mechanistic roles of cortical tension during cytokinesis. Nat. Commun. 12, 7145 (2021).
Sedzinski, J. et al. Polar actomyosin contractility destabilizes the position of the cytokinetic furrow. Nature 476, 462–466 (2011).
O’Connell, C. B., Warner, A. K. & Wang, Y. Distinct roles of the equatorial and polar cortices in the cleavage of adherent cells. Curr. Biol. 11, 702–707 (2001).
Wang, Y. L. The mechanism of cytokinesis: reconsideration and reconciliation. Cell Struct. Funct. 26, 633–638 (2001).
Wang, Y. L. The mechanism of cortical ingression during early cytokinesis: thinking beyond the contractile ring hypothesis. Trends Cell Biol 15, 581–588 (2005).
D’Avino, P. P., Savoian, M. S. & Glover, D. M. Cleavage furrow formation and ingression during animal cytokinesis: a microtubule legacy. J. Cell Sci. 118, 1549–1558 (2005).
Ou, G., Stuurman, N., D’Ambrosio, M. & Vale, R. D. Polarized myosin produces unequal-size daughters during asymmetric cell division. Science 330, 677–680 (2010).
Cabernard, C., Prehoda, K. E. & Doe, C. Q. A spindle-independent cleavage furrow positioning pathway. Nature 467, 91–94 (2010).
Daeden, A. et al. Polarized branched Actin modulates cortical mechanics to produce unequal-size daughters during asymmetric division. Nat. Cell Biol. 25, 235–245 (2023).
Cazzagon, G., Roubinet, C. & Baum, B. Polarized SCAR and the Arp2/3 complex regulate apical cortical remodeling in asymmetrically dividing neuroblasts. iScience 26, 107129 (2023).
Pontani, L. L. et al. Reconstitution of an actin cortex inside a liposome. Biophys. J. 96, 192–198 (2009).
Carvalho, K. et al. Cell-sized liposomes reveal how actomyosin cortical tension drives shape change. Proc. Natl. Acad. Sci. USA 110, 16456–16461 (2013).
Miyata, H. & Hotani, H. Morphological changes in liposomes caused by polymerization of encapsulated actin and spontaneous formation of actin bundles. Proc. Natl. Acad. Sci. USA. 89, 11547–11551 (1992).
Simon, C. et al. Actin dynamics drive cell-like membrane deformation. Nat. Phys. 15, 602–609 (2019).
Tsai, F.-C. & Koenderink, G. H. Shape control of lipid bilayer membranes by confined actin bundles. Soft Matter 11, 8834–8847 (2015).
Loiseau, E. et al. Shape remodeling and blebbing of active cytoskeletal vesicles. Sci. Adv. 2, e1500465 (2016).
Litschel, T. et al. Reconstitution of contractile actomyosin rings in vesicles. Nat. Commun. 12, 2254 (2021).
Bashirzadeh, Y., Moghimianavval, H. & Liu, A. P. Encapsulated actomyosin patterns drive cell-like membrane shape changes. iScience 25, 104236 (2022).
Murrell, M. et al. Spreading dynamics of biomimetic actin cortices. Biophys. J. 100, 1400–1409 (2011).
Sakamoto, R. et al. Membrane tension induces F-actin reorganization and flow in a biomimetic model cortex. Commun. Biol. 6, 325 (2023).
Yadav, V. et al. Filament nucleation tunes mechanical memory in active polymer networks. Adv. Funct. Mater. 29, 1905243 (2019).
Bovellan, M. et al. Cellular control of cortical actin nucleation. Curr. Biol. 24, 1628–1635 (2014).
Rosa, A., Vlassaks, E., Pichaud, F. & Baum, B. Ect2/Pbl acts via Rho and polarity proteins to direct the assembly of an isotropic actomyosin cortex upon mitotic entry. Dev. Cell 32, 604–616 (2015).
Muresan, C. G. et al. F-actin architecture determines constraints on myosin thick filament motion. Nat. Commun. 13, 7008 (2022).
Sakamoto, R. & Murrell, M. P. Composite branched and linear F-actin maximize myosin-induced membrane shape changes in a biomimetic cell model. Commun. Biol. 7, 840 (2024).
Straight, A. F. et al. Dissecting temporal and spatial control of cytokinesis with a myosin II inhibitor. Science 299, 1743–1747 (2003).
Schuppler, M., Keber, F. C., Kroger, M. & Bausch, A. R. Boundaries steer the contraction of active gels. Nat. Commun. 7, 13120 (2016).
Linsmeier, I. et al. Disordered actomyosin networks are sufficient to produce cooperative and telescopic contractility. Nat. Commun. 7, 12615 (2016).
Sakamoto, T., Limouze, J., Combs, C. A., Straight, A. F. & Sellers, J. R. Blebbistatin, a myosin II inhibitor, is photoinactivated by blue light. Biochemistry 44, 584–588 (2005).
Bendix, P. M. et al. A quantitative analysis of contractility in active cytoskeletal protein networks. Biophys. J. 94, 3126–3136 (2008).
Turlier, H., Audoly, B., Prost, J. & Joanny, J. F. Furrow constriction in animal cell cytokinesis. Biophys. J. 106, 114–123 (2014).
McCall, P. M., MacKintosh, F. C., Kovar, D. R. & Gardel, M. L. Cofilin drives rapid turnover and fluidization of entangled F-actin. Proc. Natl. Acad. Sci. USA. 116, 12629–12637 (2019).
Sun, Z. G. et al. Cofilin-mediated actin filament network flexibility facilitates 2D to 3D actomyosin shape change. Eur. J. Cell Biol. 103, 151379 (2024).
Zhang, X. F. et al. Regulation of axon growth by myosin II-dependent mechanocatalysis of cofilin activity. J. Cell Biol. 218, 2329–2349 (2019).
Isambert, H. et al. Flexibility of actin filaments derived from thermal fluctuations. Effect of bound nucleotide, phalloidin, and muscle regulatory proteins. J. Biol. Chem. 270, 11437–11444 (1995).
Pujol, T., du Roure, O., Fermigier, M. & Heuvingh, J. Impact of branching on the elasticity of actin networks. Proc. Natl. Acad. Sci. USA. 109, 10364–10369 (2012).
Jung, W. et al. Dynamic motions of molecular motors in the actin cytoskeleton. Cytoskeleton 76, 517–531 (2019).
Reymann, A. C. et al. Actin network architecture can determine myosin motor activity. Science 336, 1310–1314 (2012).
Salbreux, G., Prost, J. & Joanny, J. F. Hydrodynamics of cellular cortical flows and the formation of contractile rings. Phys. Rev. Lett. 103, 058102 (2009).
Fishkind, D. J. & Wang, Y. L. Orientation and three-dimensional organization of actin filaments in dividing cultured cells. J. Cell Biol. 123, 837–848 (1993).
Oegema, K., Savoian, M. S., Mitchison, T. J. & Field, C. M. Functional analysis of a human homologue of the Drosophila actin binding protein anillin suggests a role in cytokinesis. J. Cell Biol. 150, 539–552 (2000).
Yoneda, M. & Dan, K. Tension at the surface of the dividing sea-urchin egg. J. Exp. Biol. 57, 575–587 (1972).
Robinson, D. N., Cavet, G., Warrick, H. M. & Spudich, J. A. Quantitation of the distribution and flux of myosin-II during cytokinesis. BMC Cell Biol. 3, 4 (2002).
Finer, J. T., Simmons, R. M. & Spudich, J. A. Single myosin molecule mechanics: piconewton forces and nanometre steps. Nature 368, 113–119 (1994).
Uyeda, T. Q. P., Kron, S. J. & Spudich, J. A. Myosin step size - estimation from slow sliding movement of actin over low-densities of heavy-meromyosin. J. Mol. Biol. 214, 699–710 (1990).
Chen, D. T. N., Heymann, M., Fraden, S., Nicastro, D. & Dogic, Z. ATP consumption of eukaryotic flagella measured at a single-cell level. Biophys. J. 109, 2562–2573 (2015).
Haithcock, J. et al. The kinetic mechanism of mouse myosin VIIA. J. Biol. Chem. 286, 8819–8828 (2011).
Sakamoto, R. & Murrell, M. P. F-actin architecture determines the conversion of chemical energy into mechanical work. Nat. Commun. 15, 3444 (2024).
Kushmerick, M. J. & Davies, R. E. The chemical energetics of muscle contraction. II. The chemistry, efficiency and power of maximally working sartorius muscles. Appendix. Free energy and enthalpy of atp hydrolysis in the sarcoplasm. Proc. R Soc. Lond. B Biol. Sci. 174, 315–353 (1969).
Milo, R., Phillips, R. Cell Biology by the Numbers (1st ed.). Garland Science (2015).
Ramkumar, N. & Baum, B. Coupling changes in cell shape to chromosome segregation. Nat. Rev. Mol. Cell Biol. 17, 511–521 (2016).
Miyazaki, M., Chiba, M., Eguchi, H., Ohki, T. & Ishiwata, S. Cell-sized spherical confinement induces the spontaneous formation of contractile actomyosin rings in vitro. Nat. Cell Biol. 17, 480–489 (2015).
Vavylonis, D., Wu, J. Q., Hao, S., O’Shaughnessy, B. & Pollard, T. D. Assembly mechanism of the contractile ring for cytokinesis by fission yeast. Science 319, 97–100 (2008).
Pollard, T. D. & Wu, J. Q. Understanding cytokinesis: lessons from fission yeast. Nat. Rev. Mol. Cell Biol. 11, 149–155 (2010).
Mendes Pinto, I., Rubinstein, B., Kucharavy, A., Unruh, J. R. & Li, R. Actin depolymerization drives actomyosin ring contraction during budding yeast cytokinesis. Dev. Cell 22, 1247–1260 (2012).
Carvalho, K. et al. Actin polymerization or myosin contraction: two ways to build up cortical tension for symmetry breaking. Phil. Trans. R. Soc. B. Biol. Sci. 368, 20130005 (2013).
Kelkar, M., Bohec, P. & Charras, G. Mechanics of the cellular actin cortex: From signalling to shape change. Curr. Opin. Cell Biol. 66, 69–78 (2020).
Yoshizaki, H. et al. Cell type-specific regulation of RhoA activity during cytokinesis. J. Biol. Chem. 279, 44756–44762 (2004).
Zanin, E. et al. A conserved RhoGAP limits M phase contractility and coordinates with microtubule asters to confine RhoA during cytokinesis. Dev. Cell 26, 496–510 (2013).
Wagner, E. & Glotzer, M. Local RhoA activation induces cytokinetic furrows independent of spindle position and cell cycle stage. J. Cell Biol. 213, 641–649 (2016).
Chugh, P. & Paluch, E. K. The actin cortex at a glance. J. Cell Sci. 131, jcs186254 (2018).
Mukhina, S., Wang, Y. L. & Murata-Hori, M. Alpha-actinin is required for tightly regulated remodeling of the actin cortical network during cytokinesis. Dev. Cell 13, 554–565 (2007).
Wiggan, O., Shaw, A. E., DeLuca, J. G. & Bamburg, J. R. ADF/cofilin regulates actomyosin assembly through competitive inhibition of myosin II binding to F-actin. Dev. Cell 22, 530–543 (2012).
Chugh, P. et al. Actin cortex architecture regulates cell surface tension. Nat. Cell Biol. 19, 689–697 (2017).
Yoshizaki, H. et al. Activity of Rho-family GTPases during cell division as visualized with FRET-based probes. J. Cell Biol. 162, 223–232 (2003).
Moulding, D. A. et al. Excess F-actin mechanically impedes mitosis leading to cytokinesis failure in X-linked neutropenia by exceeding Aurora B kinase error correction capacity. Blood 120, 3803–3811 (2012).
Henson, J. H. et al. The ultrastructural organization of actin and myosin II filaments in the contractile ring: new support for an old model of cytokinesis. Mol. Biol. Cell 28, 613–623 (2017).
Fenix, A. M. et al. Expansion and concatenation of non-muscle myosin IIA filaments drive cellular contractile system formation during interphase and mitosis. Mol. Biol. Cell 27, 1465–1478 (2016).
Fox, C. J., Hammerman, P. S. & Thompson, C. B. Fuel feeds function: energy metabolism and the T-cell response. Nat. Rev. Immunol. 5, 844–852 (2005).
Maeshima, K. et al. A Transient Rise in Free Mg(2 + ) Ions Released from ATP-Mg Hydrolysis Contributes to Mitotic Chromosome Condensation. Curr. Biol. 28, 444–451 (2018).
Kang, J. H. et al. Monitoring and modeling of lymphocytic leukemia cell bioenergetics reveals decreased ATP synthesis during cell division. Nat. Commun. 11, 4983 (2020).
Stam, S., Alberts, J., Gardel, M. L. & Munro, E. Isoforms confer characteristic force generation and mechanosensation by myosin ii filaments. Biophys. J. 108, 1997–2006 (2015).
Stewart, M. P. et al. Hydrostatic pressure and the actomyosin cortex drive mitotic cell rounding. Nature 469, 226–230 (2011).
Wang, X. et al. Effects of the Laplace pressure on the cells during cytokinesis. iScience 24, 102945 (2021).
Khoromskaia, D. & Salbreux, G. Active morphogenesis of patterned epithelial shells. eLife 12, e75878 (2023).
Tinevez, J. Y. et al. Role of cortical tension in bleb growth. Proc. Natl. Acad. Sci. USA. 106, 18581–18586 (2009).
Tsuda, Y., Yasutake, H., Ishijima, A. & Yanagida, T. Torsional rigidity of single actin filaments and actin-actin bond breaking force under torsion measured directly by in vitro micromanipulation. Proc. Natl. Acad. Sci. USA. 93, 12937–12942 (1996).
Morimoto, K. & Harrington, W. F. Substructure of the thick filament of vertebrate striated muscle. J. Mol. Biol. 83, 83–97 (1974).
Piazzesi, G. et al. Skeletal muscle performance determined by modulation of number of myosin motors rather than motor force or stroke size. Cell 131, 784–795 (2007).
Jung, W., Murrell, M. P. & Kim, T. F-actin fragmentation induces distinct mechanisms of stress relaxation in the actin cytoskeleton. Acs Macro Lett. 5, 641–645 (2016).
Jung, W., Murrell, M. P. & Kim, T. F-actin cross-linking enhances the stability of force generation in disordered actomyosin networks. Comput. Part. Mech. 2, 317–327 (2015).
Dürre, K. et al. Capping protein-controlled actin polymerization shapes lipid membranes. Nat. Commun. 9, 1630 (2018).
Mostowy, S. & Cossart, P. Septins: the fourth component of the cytoskeleton. Nat. Rev. Mol. Cell Biol. 13, 183–194 (2012).
D’Avino, P. P. How to scaffold the contractile ring for a safe cytokinesis - lessons from Anillin-related proteins. J. Cell Sci. 122, 1071–1079 (2009).
Vicente-Manzanares, M., Ma, X. F., Adelstein, R. S. & Horwitz, A. R. Non-muscle myosin II takes centre stage in cell adhesion and migration. Nat. Rev. Mol. Cell Biol. 10, 778–790 (2009).
Kovács, M., Thirumurugan, K., Knight, P. J. & Sellers, J. R. Load-dependent mechanism of nonmuscle myosin 2. Proc. Natl. Acad. Sci. USA. 104, 9994–9999 (2007).
Matzke, R., Jacobson, K. & Radmacher, M. Direct, high-resolution measurement of furrow stiffening during division of adherent cells. Nat. Cell Biol. 3, 607–610 (2001).
Murrell, M. P. & Gardel, M. L. F-actin buckling coordinates contractility and severing in a biomimetic actomyosin cortex. Proc. Natl. Acad. Sci. USA. 109, 20820–20825 (2012).
Murrell, M. P. et al. Liposome adhesion generates traction stress. Nat. Phys. 10, 163–169 (2014).
Miura, K. Bleach correction ImageJ plugin for compensating the photobleaching of time-lapse sequences. F1000Res. 9, 1494 (2020).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Tseng, Q. et al. Spatial organization of the extracellular matrix regulates cell–cell junction positioning. Proc. Natl. Acad. Sci. USA. 109, 1506–1511 (2012).
Püspöki, Z., Storath, M., Sage, D. & Unser, M. Transforms and Operators for Directional Bioimage Analysis: A Survey. Adv. Anat. Embryol. Cell Biol. 219, 69–93 (2016).
Acknowledgements
The authors acknowledge funding ARO MURI W911NF-14-1-0403 and M.P.M, NIH RO1 GM126256 to M.P.M, NIH U54 CA209992 to M.P.M, and Human Frontier Science Program (HFSP) grant # RGY0073/2018 to M.P.M. M.P.M. and R.S. acknowledge support from Yale start-up funds. R.S. acknowledges support from the Overseas Postdoctoral Fellowships of the Uehara Memorial Foundation and Japan Society for the Promotion of Science (JSPS). Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the NSF, NIH, ARO, HFSP, Uehara Memorial Foundation, or JSPS.
Author information
Authors and Affiliations
Contributions
R.S. and M.P.M. designed the experimental work. R.S. acquired the experimental data. R.S. and M.P.M. contributed new reagents/analytic tools. R.S. analyzed experimental data. R.S. drafted the paper. R.S. and M.P.M. edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sakamoto, R., Murrell, M.P. Mechanical power is maximized during contractile ring-like formation in a biomimetic dividing cell model. Nat Commun 15, 9731 (2024). https://doi.org/10.1038/s41467-024-53228-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53228-y