Abstract
ATP synthases play a crucial role in energy production by utilizing the proton motive force (pmf) across the membrane to rotate their membrane-embedded rotor c-ring, and thus driving ATP synthesis in the hydrophilic catalytic hexamer. However, the mechanism of how pmf converts into c-ring rotation remains unclear. This study presents a 2.8 Å cryo-EM structure of the Vo domain of V/A-ATPase from Thermus thermophilus, revealing precise orientations of glutamate (Glu) residues in the c12-ring. Three Glu residues face a water channel, with one forming a salt bridge with the Arginine in the stator (a/Arg). Molecular dynamics (MD) simulations show that protonation of specific Glu residues triggers unidirectional Brownian motion of the c12-ring towards ATP synthesis. When the key Glu remains unprotonated, the salt bridge persists, blocking rotation. These findings suggest that asymmetry in the protonation of c/Glu residues biases c12-ring movement, facilitating rotation and ATP synthesis.
Similar content being viewed by others
Introduction
The molecule adenosine triphosphate (ATP) serves as the central energy currency of life1. The majority of ATP is synthesized by oxidative phosphorylation, catalyzed by ATP synthase, and dependent on generation of proton motive force (pmf) across the membranes by respiratory complexes in animal and bacterial cells1,2,3,4,5. In chloroplasts, ATP is synthesized by ATP synthase using pmf generated by light reactions5,6.
There are two types of ATP synthases: the F type (FoF1) and the V type ATP synthases7. FoF1-ATP synthases are present on the plasma membrane of bacteria, the cristae membrane of mitochondria, and the thylakoid membrane of chloroplasts1,7. FoF1-ATP synthases consist of two domains: the F1 domain, which carries out the ATP synthesis reaction using ADP and inorganic phosphate, and the membrane-embedded Fo domain, which translocates protons across the membrane8.
Certain Archea or bacteria, such as the thermophilic bacterium, Thermus thermophilus7,9 express an alternative ATPase. The subunit composition and fundamental structure of these alternative ATP synthases closely resemble those of the V-ATPase in eukaryotes, indicating a potential ancestral relationship to the eukaryotic V-ATPase10. Consequently, they are commonly referred to as V-ATPases or V/A-ATPases11.
Similar to F-ATPase, the V/A-ATPase from T. thermophilus (Tth) synthesizes ATP utilizing pmf, in contrast to eukaryotic V-ATPases, which function as proton pumps powered by ATP hydrolysis12,13. The Tth V1 domain is a motor that rotates the central rotor (DF) within the A3B3 subdomain. The A3B3 subdomain contains three catalytic sites, each comprising a structurally distinct AB dimer14. The Vo domain (E2G2d1a1c12) is composed of the stator parts, including the a-subunit, two EG peripheral stalks, and the c12 rotor ring (c12-ring), which comprises the central rotor complex together with the d1 and DF subunits of V1 (Fig. 1A). During rotation of the central rotor (DF d1c12), caused by the pmf, the DF subunit induces structural changes in the three AB dimers, leading to the cooperative synthesis of ATP at the A3B3 hexamer15.
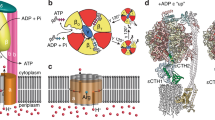
A Schematic of V/A-ATPase. The a, c12, and d subunits of the Vo domain are colored pink, blue, and green, respectively. B Schematic of the half-channel model. In T. thermophilus the a subunit of the Vo domain (a-subunit in) forms half-channels on both the periplasmic and cytoplasmic sides of the membrane. H+ can access the c12-ring through these channels, which are separated by two Arg residues. C The atomic model of the Vo domain of holo-V/A-ATPase state 1. The subunits are colored the same as in (A). The scale bar shows 20 Å. D The rotation angles, calculated by UCSF Chimera, between each state are indicated. To help visualize the rotation of the c12-ring, one of the c-subunits is colored blue. The scale bar shows 15 Å. E The interactions between the soluble arm of a-subunit and d-subunit in each state. The density maps are shown semi-transparent. The residues participating in the interactions between subunits are shown in stick representation. The scale bar is 5 Å. F The distance between c/E63 and a/R563, R622 in each state. The values indicate the distances between the Cβ of each residue. The density maps are shown as semi-transparent. The scale bar shows 5 Å.
The Vo domain rotation is driven by proton flow through the transmembrane domain of the a-subunit using a common mechanism also used by the Fo domain of FoF1. The a-subunit contains two half-channels acting as a proton entrance and a proton exit and accessible only from the extracellular and intracellular sides of the membrane16,17. Direct proton flow through the two half-channels is blocked by a positively charged Arg residue in the a subunit (Fig. 1B). Rather, the proton enters the periplasmic half-channel and moves in an anti-clockwise direction via rotation of the c-ring viewed from Vo side until it can enter the intracellular half channel for the final step of transport into the cytoplasm17. However, the precise mechanism by which the proton flow drives rotation of the c-ring remains elusive.
Overall, TthVo and eukaryotic Vo have similar structures, but they differ significantly in the structure of the c-ring. In contrast to the homomeric c12-ring of TthVo, with each composed of two transmembrane helices (2 TM, Fig. 1C), the c-ring in eukaryotic Vo is a heteromeric decamer composed of 4 TM or 5 TM c-subunits (c8, c’1, and c”1 in yeast Vo)13,18. These structural differences are likely to have important implications for how the proton flow drives c-ring rotation.
Cryo-electron microscopy (cryo-EM) has been utilized to reveal structural details of several rotary ATPases, including V- and FoF1-type ATP synthases5,8 and their membrane-embedded domains. Notably, a high-resolution structure of the Vo of yeast V-ATPase with a resolution of 2.7 Å has been determined, and molecular dynamics simulations (MD) based on this structure have advanced our understanding of the proton transport mechanism by rotation of the Vo domain18. However, the resolution of the Fo domain of ATP synthase was too low to allow identification of the water molecules involved in proton flow. In order to fully elucidate the mechanism of c-ring rotation by pmf, further high-resolution structures of ATP synthase Fo or Vo are critical.
In this study, we determined the structures of the Vo domain of the full V/A-ATPase and the isolated Vo domain using cryo-electron microscopy. The structure of the Vo domain of V/A-ATPase was almost identical to that of the isolated Vo. By analyzing the structure of the 2.8 Å resolution Vo, we identified several aligned water molecules within the two half-channels. Furthermore, MD simulations based on this structure revealed an asymmetric protonated state of the Glu residues in the two half-channels likely to be induced by the pmf. We propose a model in which the asymmetric protonation states of the Glu residues of the c-subunits on either side of the two channels provide the unidirectional bias for the Brownian motion of the c12-ring.
Results
Structure of three states of Vo domain in V/A-ATPase
In this study, we analyzed the structure of Vo domain in the holo-complex by focused refinement of Vo. We used a previous dataset of micrographs obtained from holo-V/A-ATPase incorporated into nanodiscs19. The schematic representations detailing the sequential steps of image acquisition and the subsequent reconstruction of the Vo domain structure are presented in Supplementary Fig. 1. Particles constituting each state of V/A-ATPase (state 1, state 2, and state 3) obtained under different conditions were collected and the structure of the Vo domain was reconstructed from these. Using focused refinement on the Vo domain, we obtained structures of the three rotational states of Vo domain; state 1 at 3.0 Å, state 2 at 3.6 Å, and state 3 at 3.5 Å resolution, with the d subunit position differing by 120° in each case (Fig. 1D). The structures of the c12-ring and membrane-embedded domain of a-subunit were basically identical in all three states. The distance between the soluble arm domain of the a-subunit and the d-subunit was almost the same in all three states. However, in state 1, side chain interactions were observed between a/E57,a/R54, and d/R38 (Fig. 1E). The interactions between the d-subunit and a-subunit contribute to the stabilization of state 1, potentially resulting in a higher prevalence of state 1 (53%) relative to state 2 (30%) and state 3 (17%). This result is consistent with our previous findings, which indicated that state 1 is the predominant class with state 3 represented by the lowest number of particles11. The distances between the residues forming the salt bridge between the a-subunit and the c12-ring (a/R563-c/E63) were also nearly identical among the three states (Fig. 1F). In this case, the density at the tip of the Glu residues is unclear, so the distance between the β carbons of the a/R563 and c/E63 residue was measured as indicated in Fig. 1E and F. The state 1 Vo structure was obtained from an active holo-complex undergoing ATP hydrolysis-dependent rotation, which is why it was possible to capture all three states. In order to compare the structure of active Vo with that of autoinhibited Vo, we determined the high-resolution structure of isolated Vo.
Structure of isolated Vo
The purified Vo domain incorporated into nanodiscs was subjected to single particle analysis using a cryo-EM (Titan Krios G3, Thermo Fisher) equipped with a K3 electron direct detector in electron counting mode. The strategy of single particle analysis for TthVo is summarized in Supplementary Fig. 2.
Our 3D reconstruction map of the isolated Vo complex was obtained with an overall resolution of 2.8 Å (Fig. 2A). In particular, the local resolution of the c12-ring and the interface between the a-subunit and the c12-ring approaches to at ~2.5 Å (Fig. 2A, right). This allows both discussion of the orientation of amino acid residues and identification of the water molecules responsible for proton relay (Fig. 3 and Supplementary Fig. 3).
A The density map of the isolated Vo domain. The map is colored according to the local resolution (see key). Left: overall structure. Right: the cross-section image of the region is indicated by the dotted lines in the left panel. B The salt bridge formed between a/R563, a/R622, and c/E63. The residues are shown as sticks. The density maps are shown as semi-transparent. The scale bar shows 3 Å. The left panel represents a comparison of the salt bridge structure of isolated Vo and Vo in the holo-enzyme. The superimposed image of the salt bridge of the isolated Vo (colored) and the Vo domain of holo-enzyme (gray). C The angles adopted by the c/E63 residues. The dashed semi-circle indicates the circumference of the c12-ring. The red lines indicate the angle of the E63 side chain of each c-subunit. The scale bar shows 10 Å. The insets are the magnified views of the regions indicated by the dotted rectangles. The scale bar shows 5 Å. D Plot of the angles of c/E63.
A The surface properties of the transmembrane region of a-subunit. The surface of the a-subunit is colored cyan (hydrophilic) to yellow (hydrophobic). B The coordination of water molecules in the half-channels. The cryo-EM water molecules are shown in sphere representation and colored by density from the MD simulation. The color scale illustrates the relative density of water, with the bulk density set to 1. The iso-surface of the relative water density of 0.5 obtained from MD simulation is shown with the cryo-EM water molecules. The regions indicated by the dashed rectangles are enlarged in (C, D). Enlarged views of the periplasmic (C) and cytoplasmic (D) channels. The residues forming the channels are shown in stick representation. The dashed line in (C) indicates the possible proton relay path. E The possible proton relay path. The cryo-EM water molecules are represented as spheres with the same color coding as (B–D).
As previously demonstrated, in the isolated Vo, the hydrophilic arm region of the a-subunit undergoes a conformational change, moving closer to the d-subunit and forming an autoinhibited structure that restricts the rotation of the complex of d-subunit and c12-ring relative to the stator a-subunit20. In contrast, the hydrophobic region of the a-subunit and other regions, such as the c12-ring of the isolated Vo, exhibit a remarkable similarity to the Vo domain within the holo-complex. This similarity is further illustrated by the main chain structure and the arrangement of side chains. For instance, the orientation of the a/R563 crucial for its interaction with c/E63 exhibited a remarkably close resemblance between the isolated Vo and the holo-complex Vo (Fig. 2B). Additionally, the surrounding side chains exhibit nearly identical configurations. The similarity strongly suggests that the structure of the isolated Vo is functionally active with respect to proton translocation.
The high-resolution structure of the isolated Vo revealed the orientation of the side chain of each membrane-embedded c/E63 within the c12-ring. Of the c/E63 residues in the c12-ring, 9 display a distinctive conformation, bending inward toward the central axis of the ring (Fig. 2C, D and Supplementary Fig. 4). Within this hydrophobic microenvironment, the carboxyl groups of these c/E63 residues are effectively shielded from water, hindering their ionization and maintaining them in a protonated state. These protonated carboxyl groups are stabilized through the formation of hydrogen bonds with adjacent threonine residues (c/T64) in the neighboring c-subunit (Fig. 2C, insets for c(X)E63 and c(P)E63, Supplementary Fig. 4). This closed structure of c/E63, referred to as the “proton-locked state,” is evident not only in the crystal structure of the isolated Fo c-ring but also in the yeast Vo c-ring18,21,22. In contrast, the three c/E63 residues facing the hydrophilic channel do not adopt the proton-locked state. The central c/E63(Z) lies in close proximity to a/R563, indicating the formation of a salt bridge (Fig. 2B). The c(Y)/E63 and c(O)/E63 residues, situated adjacent to the c(Z)/E63 residue, do not interact with the neighboring c/T64 residues, likely due to their deprotonated state. As a result, these c/E63 residues adopt an extended conformation. The extended conformation of the Glu residues (c(Y)/E63 and c(Z)/E63) is also observed in yeast Vo, where the luminal Glu residue adjacent to the salt bridge also maintains an extended configuration. It is important to highlight that in yeast Vo, the Glu residue located at the cytosolic side (c’), equivalent to the cytosolic side in bacteria (c(O)/E63), is in a proton-locked state due to the heterogeneous c-ring structure (refer to Supplementary Fig. 5, c8c’1c”1)18. This matter is further addressed in the discussion section.
The a/R622 is positioned in close proximity to the salt bridge formed by c(Z)/E63 and a/R563 (Fig. 2B). The juxtaposition of these two Arg residues constitutes a structural motif observed in yeast and human Vo but is absent in Fo structures18,23. The two Arg residues effectively partition the hydrophilic tunnel between the a-subunit and the c12-ring into two distinct half-channels: a cytoplasmic channel and a periplasmic channel (Fig. 3A, B). The high-resolution Vo structure allowed us to identify some of the water molecules present in each channel (shown as colored spheres). In the periplasmic channel, 14 water molecules and two hydrophilic residues are likely to form a Grotthuss network (Fig. 3C). We also found a water chain consisting of water molecules and ten hydrophilic residues in the cytoplasmic side channel (Fig. 3D).
MD simulation of water molecules in Vo
We employed the atomic model of the hydrophobic domain of the a-subunit and the c12-ring for molecular dynamics (MD) simulations. Initially, we present the results of a 500 ns MD simulation illustrating the penetration of water molecules into Vo, as depicted in Fig. 3. Subsequently, following the superposition of the membrane-embedded portion of the a-subunit, position-dependent water density was determined through MDAnalysis24,25, utilizing a grid spacing of 1 Å.
We evaluated the correspondence between the water molecules identified in the Vo cryo-EM structure and the simulated water molecules from the MD simulation. Each cryo-EM water was assigned the closest grid water density (Fig. 3B and Supplementary Fig. 6). The assigned density for the cryo-EM waters displays some variability. The water molecules, as represented by red spheres, exhibit a high likelihood of existing as shown and are presumed to be bound water molecules. Cryo-EM waters situated within the two half-channels exhibit relatively high density (depicted as green spheres), suggesting the formation of the water channel that forms the Grotthuss network responsible for the proton relay26. Conversely, water located beyond the half-channel is less dense (blue spheres), suggesting that it does not contribute significantly to the proton relay.
As illustrated in Fig. 3B, simulated water fills both half-channels with a void between the cytoplasmic channel and the salt bridge (Fig. 3D). Although some water molecules go back and forth between the two half-channels, the positive charges of a/R563 and a/R622, situated at the center of the complete water channel, likely serve as a Coulombic barrier, preventing the unrestricted flow of protons along the entire channel. This structural configuration suggests that protons relayed from the periplasmic side through the water channel are transferred to c(Z)/E63, which forms a salt bridge with a/R563 at the center of the full water channel. Based on a chain of high-density water molecules toward c(Z)/E63 (dotted line in Fig. 3C, E), we suggest that a/H616 acts as a proton donor in the periplasmic channel.
Estimation of pK a of c/E63 in the water channel
The cryo-EM structure of Vo indicates that, of the 12 c/E63 residues, the c(Z)/E63 forming the salt bridge and its two neighboring c(Y)/E63 and c(O)/E63 are not in a proton-locked state. Given that the charge state differences of these Glu residues significantly impact Vo behavior, the pKa of each c/E63 was estimated using the program H++27. The equilibrated Vo structure, with lipid molecules within a 5 Å distance from the protein, from the MD simulation, served as the input structure. Lipid molecules were included to account for the membrane environment, with an inner dielectric constant set to 10.
The pKa values of cytoplasmic c(O)/E63 and periplasmic c(Y)/E63 were estimated as 6.5 and 8.1, respectively. A negative pKa value was determined for the central c(Z)/E63, which forms a salt bridge with a/R563. These results suggest that in the absence of pmf, at least c(Z)/E63 and cytoplasmic c(O)/E63 are present in a deprotonated state. The slightly higher pKa of c(Y)/E63 on the periplasmic side suggests that this residue is more prone to protonation than the c(O)/E63. In fact, there is density around that corresponding to c(Y)/E63 that appears to be water, which is also connected to residues of the a-subunit, implying hydrogen bonding between the protonated c(Y)/E63 and a water molecule (Supplementary Fig. 7). Analysis of the remaining nine c/E63 in the hydrophobic environment reveals that all of their pKa values exceed 12, confirming their protonated state.
MD simulation of c 12-ring rotation
We investigated how the protonation state of the c/E63 residues of the c12-ring effects the rotation of the c12-ring. In this simulation, we removed the hydrophilic domains of the d- and a-subunits to weaken the rotor-stator interaction of the Vo domain, thereby allowing spontaneous rotation of the c12-ring relative to the stator. We focused on three c/E63 residues: periplasmic c(Y)/E63, salt-bridge forming c(Z)/E63, and cytoplasmic c(O)/E63. In this simulation, the cytoplasmic and periplasmic c/E63 were deprotonated (c(O)/E63(COO-)) and protonated(c(Y)/E63(COOH)), respectively. For the salt-bridge forming c(Z)/E63, we examined both protonated and deprotonated states. Hereafter, we denote the protonation state of the salt-bridge forming c(Z)/E63 as c(Z)/E63(COOH) for the protonated state and c(Z)/E63(COO-) for the deprotonated state.
Figure 4A illustrates the positions of the side chains c(Z)/E63, a/R563, and a/Q619 at the start and end of the 500 ns simulation, and in both protonation states. In the Cryo-EM structure (Fig. 4A, left panel), the side chain of c(Z)/E63, which initially formed a salt bridge with a/R563, is observed to move into closer proximity with a/Q619 following the 500 ns simulation run (Fig. 4A, right-upper panel). Figure 4B illustrates the time course of the c12-ring rotation angle observed in two independent 500 ns MD runs performed in the different protonation states. The counterclockwise (view from the V1 side) ~10° rotation, which corresponds to the rotational direction for the ATP synthesis, was observed in two independent 500 ns trajectories in the protonation state c(Y)/E63(COOH), c(Z)/E63(COOH) and c(O)/E63(COO-) shown as orange lines. After this motion, the side chain of c(Z)/E63 reaches a hydrogen bonding distance (~2.5 Å) with a/Q619 (Fig. 4C, orange line). This simulation suggests that the hydrogen bonding of protonated c(Z)/E63 with a/Q619 contributes to the bias in Brownian motion. The time courses of the rotational angle show that they equilibrated in the simulations.
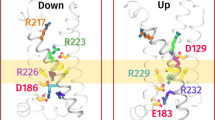
A Initial (left) and final structures with protonation (right, upper) and deprotonation (right, lower) states of c/E63 after the 500 ns simulations. The amino acid residues (c/E63, a/R563, and a/Q619) are shown in colored stick representation. The c(Y), c(Z), and c(O) subunits are shown in green, cyan, and blue, respectively. Hydrogen atoms are represented as white sticks. B Rotation angle of the c-ring observed during the simulations. Rotation trajectories in the c(Y)/E63(COOH), c(Z)/E63(COOH), c(O)/E63(COO-) simulations are shown by orange lines, while those in the c(Y)/E63(COOH), c(Z)/E63(COO-), c(O)/E63(COO-) simulations are indicated by blue lines. C The distance between E63 and Q619 during the simulation. The distances obtained in the c(Y)/E63(COOH), c(Z)/E63(COOH), c(O)/E63(COO-) simulations are shown by the orange lines, while those obtained in the c(Y)/E63(COOH), c(Z)/E63(COO-), c(O)/E63(COO-) are indicated by the blue lines. In the MD simulations independently performed three times for 100 ns under the same conditions, a rotation of about 5° in the direction of ATP synthesis was observed when c(Z)/E63 was protonated. Additionally, when protonated, the distance between c(Y)/E63(COOH) and a/Q619 approached that required for hydrogen bond formation, approximately 2 Å. These results were not obtained when c(Z)/E63 was deprotonated (Supplementary Figs. 8, 9).
In contrast, under the c(Y)/E63(COOH), c(Z)/E63(COO-), and c(O)/E63(COO-) conditions, no clear unidirectional rotation was observed during the 500 ns simulation runs (Fig. 4B, blue lines). The distance between c(Z)/E63 and a/Q619 remains approximately 6 Å under these conditions (Fig. 4C, blue line), indicating the absence of hydrogen bonding between c(Z)/E63 and a/Q619. In contrast, c(Z)/E63 and a/R563 maintained a salt bridge configuration (Fig. 4A, right-lower panel). These features were confirmed by an additional three trajectories (Supplementary Fig. 8).
These results suggest that biased Brownian motion of the c12-ring toward the periplasmic direction requires cleavage of the salt bridge between c(Z)/E63 and a/R563 as a result of protonation of c(Z)/E63. Subsequently, protonated c(Z)//E63 forms a hydrogen bond with a/Q619, as suggested by the 500 ns MD simulations (Fig. 4C).
MD simulation of forced rotation of c12-ring
We also conducted a forced-rotation simulation, applying torque to the c12-ring as previously described28. The c12-ring was rotated at a constant speed of 1 degree per nanosecond up to 30°. In conditions of c(Y)/E63(COOH), c(Z)/E63(COOH), and c(O)/E63(COO-), we observed a complete switch in the c(Z)/E63 salt-bridge interaction with a/R563. As the c12-ring rotated, the distance between c(O)/E63 and a/R563 gradually decreased, and by the end of the 30° rotation, the distance was less than 3 Å, suggesting the formation of a salt bridge between c(O)/E63 and a/R563 (Fig. 5A, C, blue line). Indeed, in the snapshot structure of the 30° rotation, c(O)/E63 had apparently formed a salt bridge with a/R563 (Fig. 5A, 30 deg panel, dashed square). At a 10° rotation (Fig. 5C, light blue line), the distance between c(Z)/E63(COOH) and a/Q619 was approximately 3 Å, suggesting hydrogen bonding between these two residues (Fig. 5A, 10 deg panel, dashed ellipse).
Snapshot structures obtained during forced-rotation simulation in c(Y)/E63(COOH), c(Z)/E63(COOH), c(O)/E63(COO-) (A), and c(Y)/E63(COOH), c(Z)/E63(COO-), c(O)/E63(COO-) (B). The c(Y), c(Z), and c(O) subunits are shown in green, cyan, and blue, respectively. Three E63 and one T64 from c-subunits, R563 and Q619 from a-subunit are shown in stick representation. The images represent the arrangement of the side chains at 0, 10, and 30 degree rotations. Distances between the salt-bridge c(Z)/E63 –a/R563 and c(O)/E63 - a/R563 are illustrated in cyan and blue lines, respectively, obtained from the c(Y)/E63(COOH), c(Z)/E63(COOH), c(O)/E63(COO-) simulation (C) and the c(Y)/E63(COOH), c(Z)/E63(COO-), c(O)/E63(COO-) simulation (D). Distances of c(Z)/E63 – a/Q619 are also shown in light blue in both figures. E Distances between c(Z)/E63 and c(Y)/T64 (blue line) and between c(Y)/E63 and c(X)/T64 (green line) during the 30° forced c12-ring rotation.
The distance between the protonated c(Y)/E63 and the adjacent c(X)/T64 is approximately 3 Å at a 10° rotation of c12-ring (Fig. 5E, green line) indicating that c(Y)/E63 is in a proton-locked state (Fig. 5A, 10 deg panel, dashed circle). When the c12-ring reaches a 30° rotation, the c(Y)/E63 becomes embedded in the membrane (Fig. 2C). Hence, it is advantageous for c(Y)/E63 to be in a proton-locked state, rendering c(Y)/E63 hydrophobic.
In conditions of c(Y)/E63(COOH), c(Z)/E63(COO-), and c(O)/E63(COO-), the salt bridge interaction between c(Z)/E63 and a/R563 persisted at the end of the 10° rotation of the c12-ring (Fig. 5B, 10 deg panel). In fact, the distance between c(Z)/E63 and a/R563 remained within 4 Å during the whole 30° rotation (Fig. 5D, cyan line), hindering the formation of a new salt bridge between c(O)/E63 and a/R563 (Fig. 5B, 30 deg panel). These results indicate that protonation of c(Z)/E63 is necessary for both dissociation of the salt bridge with a/R563 and for smooth rotation of the c12-ring, ruling out a scenario that the 30° rotation precedes the protonation of c(Z)/E63.
Discussion
We determined the structure of the Vo domain in both the holo-enzyme and from isolated Vo of V/A-ATPase. Consistent with previous studies, there were marked differences in the distribution of molecules corresponding to different rotational states observed for the V/A-ATPase, with a significantly higher distribution ratio for state 1. In this study, we constructed an atomic model of the Vo domain based on the high-resolution structure of state 1, confirming the interaction between a/E57 (stator) and d/R38 (rotor) as depicted in Fig. 1B. This suggests that the interaction between a/E57 and d/R38 stabilizes state 1, leading to a molecular distribution biased towards state 1.
We elucidated the high-resolution structure of the isolated Vo domain, enabling us to determine the side chain orientation of c/E63 in each of the 12 protomers of the c12-ring, a residue which plays a crucial role in proton translocation. Our high-resolution structural analysis of the Vo domain revealed that 9 of the 12 c/E63 residues embedded in the membrane assume a proton-locked state18. In this state, the protonated carboxyl group forms a hydrogen bond with the c/T64 residue in the adjacent protomer (Fig. 2C and Supplementary Fig. 4), thereby stabilizing it within the hydrophobic environment.
In contrast, of the remaining three c/E63 residues situated in the hydrophilic water channel, the central c(Z)/E63 is positioned close to a/R563, and calculated pKa values indicate that these residues form a salt bridge. Furthermore, the c/E63 residues on either side of c(Z)/E63 are not in the proton-locked state and adopt an extended structure. These results are consistent with the recent solid-state NMR study using FoF1 from thermophilic Bacillus sp. PS3, where at least three Glu residues adopt the deprotonated state in the Fo c10-ring29.
Recent cryo-EM structures of the Fo and Vo domains have implied that the conserved Glu residue in the c-ring and the Arg residue in the stator a-subunit are close to each other, suggesting that they may form a salt bridge18,23. Our high-resolution structure of the TthVo domain confirms that the salt bridge forms between c(Z)/E63 and a/R563. This crucial finding strongly suggests that the c12-ring cannot undergo rotation unless this salt bridge is cleaved. A similarly stable a-c salt bridge has been reported in the Vo structure of yeast, with MD simulations proposing an energy requirement of ~8 kcal/mol for cleaving this salt bridge18. Therefore, it is crucial to develop a model that accounts for the breakage of these salt bridges in order to gain a comprehensive understanding of how the proton motive force generates Brownian motion of the c-ring that is biased in the direction of ATP synthesis.
The high-resolution structure of Vo presented here combined with MD simulations based on this structure have allowed identification of water molecules that fill both the periplasmic and cytoplasmic half-channels, thus forming a complete water channel (Fig. 4). The positive charges of a/R563 and a/R622 block direct proton flow through the two half-channels. Instead, the water channel structure enables the protonation of c(Z)/E63 in the salt bridge by relaying protons to the location of this residue under the influence of the pmf. Because the estimated low pKa value is unsuitable for accepting a proton, this would require sufficient water molecules around the salt bridge to increase the pKa of c(Z)/E63 from the estimated one. A comparable water channel structure was observed in MD simulations conducted on the Fo structure30. We note that the all-atom MD simulation used in this work, as well as other Fo studies30,31 does not allow protonation or deprotonation events to occur during the simulation. For a direct simulation of proton transfer, hybrid quantum mechanics/molecular mechanics (QM/MM) simulation is needed32.
The MD simulations conducted in this study unequivocally demonstrated that rotation of the c12-ring requires cleavage of the salt bridge between the c12-ring and the a-subunit. Protonation of the salt bridge forming c(Z)/E63 resulted in an observed unidirectional Brownian motion of approximately 10° in the direction of ATP synthesis (Fig. 4). Subsequently, the protonated c(Z)/E63 reached hydrogen bond distance with a/Q619. This outcome suggests that hydrogen bonding arising from the protonation of c(Z)/E63 may contribute to the biased Brownian motion of the c12-ring. The importance of the charged states of the three Glu residues in the c12-ring for generating rotational force has been previously discussed33. However, the results of our current MD simulation, based on the high-resolution structure of Vo, are to the best of our knowledge, novel in that they identify an intermediate structure where a/Q619 interacts with the Glu residues.
All-atom MD simulations with forced rotation of the c12-ring yielded consistent results. In the setup with a deprotonated c(Z)/E63, the distance between c(Z)/E63 and a/R563 remained nearly constant after the forced rotation of the c12-ring by 30° relative to the a-subunit, strongly suggesting maintenance of the salt bridge (Fig. 5D). Consequently, neither hydrogen bonding between c(Z)/E63 and a/Q619 nor interaction between c(O)/E63 and a/R563 occurred (Fig. 5B). In MD simulations under conditions where c(Z)/E63 is protonated, the distance between c(Z)/E63 and a/R563 gradually increased as the c12-ring rotated (Fig. 5C). Simultaneously, the distance between c(O)/E63 and a/R563 decreased, indicating the formation of a new salt bridge. These results establish that protonation of both the periplasmic c(Y)/E63 and the salt bridge forming c(Z)/E63 is essential for the c12-ring to be rotated by the pmf.
The suggested mechanism of proton flow-driven rotation in the direction of ATP synthesis is illustrated in Fig. 6. A proton relayed through the water channel leads to sequential protonation of c(Y)/E63 and then c(Z)/E63. Protonation of c(Z)/E63 eliminates the Coulomb attraction between a/R563 and c(Z)/E63, thereby exposing the positive charge of a/R563. The protonated c(Z)/E63 then forms a hydrogen bond with a/Q619, with a 10° rotation in the direction of ATP synthesis. The extended c(Y)/E63 adopts a proton-locked state in the water channel after 10° rotation of the c12-ring, resulting in a hydrophobic interaction between the proton-locked c(Y)/E63 and the membrane. The subsequent Coulomb attraction between a/R563 and the deprotonated c(O)/E63 in the c12-ring biases Brownian motion in the direction of ATP synthesis.
A A schematic of the interface between the a-subunit and the c12-ring. The ring is surrounded by lipids, as shown here. B is a magnified view of the interface. C The rotation scheme of c12-ring. top: The protonation of c(Z)/E63 causes cleavage of the salt bridge with the Arg residues of the a-subunit. middle: The deprotonated c(O)/E63 is attracted to the Arg residues of the a-subunit by Coulombic interaction (red arrow). The Brownian motion of the protonated (hydrophobic) c(Y)/E63 and c(Z)/E63 is biased toward the membrane (black arrows). bottom: After a 30° rotation, c(O)/E63 forms a new salt bridge with the Arg residues. c(P)/E63 is deprotonated.
Together, the protonation of periplasmic c(Y)/E63, hydrogen bonding of protonated c(Z)/E63 with a/Q619, and the Coulomb attraction between a/R563 and the deprotonated cytoplasmic c(O)/E63 collectively contribute to a unidirectional Brownian motion of the c12-ring, driving it in the ATP synthesis direction. Recently published MD simulations of Fo also show that the c-ring rotates through 10° intermediates, suggesting that both Vo and Fo use the same system to convert the pmf into c-ring rotation31.
The structure of the Vo domain of V/A-ATPase, which functions as an ATP synthase, closely resembles that of the Vo domain in yeast V-ATPase, which functions as a proton pump. However, in the yeast Vo domain, the cytoplasmic Glu residue situated in the c’-subunit (equivalent to c(O)/E63 in TthVo) exhibits a large arc of 54° between c’ and c” subunits, resulting in the proton-locked state of Glu in the c’-subunit (Supplementary Fig. 5). Hence, even if the Glu in the c”-subunit involved in forming salt bridges is protonated by the pmf from the luminal side, Coulomb attraction between the Glu residue in the c’-subunit and the a-subunit Arg was not generated. As a result, the heterogeneous structure of the c-ring in yeast Vo hampers rotation of the c-ring in the ATP synthetic direction induced by the pmf. A similar heterogenous c-ring structure is also present in the mammalian V-ATPase, where the distance between the Glu residues on the c-ring is long. As a result, the distance between a/Arg and c/Glu is greater in the eukaryotic Vo, which is disadvantageous for the generation of rotational force in the direction of ATP synthesis by pmf, where Coulombic forces play a significant role.
Methods
Protein preparation
T. thermophilus V/A-ATPase was expressed with a C-terminal His-3 tag on the c-subunit using a modified operon generated by the integration vector system34. The purification of His-tagged Vo was conducted following previously described methods20. In brief, T. thermophilus membranes were suspended in a buffer containing 10% Triton X-100 and sonicated to solubilize the membrane proteins. After ultracentrifugation, the supernatant containing V/A-ATPase was applied to a Ni2+-NTA column. The fractions containing V/A-ATPase were dialyzed against 20 mM Tris-HCl (pH 8.0), 1 mM EDTA for 2 days at 4 °C. The combined fraction was then applied to a Resource Q column (BioRad). Eluted fractions were analyzed by SDS-PAGE, and the fractions containing Vo and V/A-ATPase were concentrated separately using Amicon 100 K molecular weight cut-off filters (Millipore). For nanodisc incorporation, 25 mM DMPC (Avanti) solubilized in 5% DDM was used20. The concentrated Vo fraction, the scaffold protein MSP1E3D1, and DMPC were mixed in a 1:12:600 molar ratio and incubated for 30 min at room temperature. Then, 200 μL of Bio Beads SM-2 equilibrated with wash buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl) were added into 500 μL of the protein-lipid mixture. After 2 h of incubation at 4 °C with gentle stirring, an additional 300 μL of Bio Beads was added to the mixture prior to overnight incubation at 4 °C to yield the nanodiscs. The supernatant of the mixture containing the nanodisc-Vo was applied to a Superdex200 Increase 10/300 column equilibrated in wash buffer. Individual fractions were analyzed by SDS-PAGE and concentrated to approximately 4 mg/mL. The prepared nanodisc-Vo was stored at 4 °C and used for cryo-grid preparation within several days.
Cryo-EM data acquisition
The cryo-EM dataset for holo-VoV1 used in this study were collected in a previous study we reported11. The grid preparation and the condition of the cryo-EM acquisition were as previously described11.
To prepare the cryo-EM grid for the isolated Vo, 2.7–3.0 μL of the sample solution was loaded onto a glow-discharged Quantifoil Cu R1.2/1.3 and blotted by a Vitrobot IV followed by vitrification with liquid ethane. Cryo-EM movies were automatically collected using a Titan Krios (Thermo Fisher, USA) equipped with a K3 BioQuantum camera (Gatan, USA) using serialEM-software35. The conditions of cryo-EM acquisition used were a nominal magnification: 81,000, the pixel size: 0.88 Å/pix, the total electron dose: 60 electrons/Å2, the corrected spherical abbreviation: 0.079 mm.
Image processing
For the Vo domain from holo-VoV1, the image processing was performed using Relion36 and CryoSPARC v4.037. The collected movies were subjected to motion correction using MotionCor238 and contrast transfer function (CTF) estimation using Ctffind439. As a result of the manual inspection and curation of the micrographs, 15,466 micrographs were used. The particles were picked using Topaz, a machine-learning-based particle picker40. The picked 1,671,397 particles were imported into cryoSPARC, and subjected to several rounds of 2D classification in order to select the good particles. These good particles were subjected to heterogeneous refinement. The refined particles were classified into three states of VoV1. States 1, 2, and 3 contained 84,150, 48,316, and 26,525 particles, respectively. These particles were re-imported to Relion and subjected to Refine3D, CtfRefine, and Postprocess jobs. Then, the particles were subjected to Bayesian Polishing. The polished particles were used for the final 3D reconstruction. The results of Refine3D for each state of VoV1, yielded density maps for states 1, 2, and 3 at resolutions of 2.9, 3.2, and 3.4 Å, respectively. Further focused refinement using the mask focusing on the Vo domain provided density maps of the Vo domain only at 3.0, 3.6, and 3.6 Å for states 1, 2, and 3, respectively.
For the isolated Vo, all image processing was performed using cryoSPARC v4.0. The workflows are summarized in Supplementary Fig. 2. The collected movies were applied to motion correction and CTF estimation. As a result of manual inspection and curation of the micrographs, 8719 micrographs were used. A blob particle picker was used for the initial particle picking for the isolated Vo. The picked particles were subjected to several rounds of 2D classification. Some good 2D average images were selected and then used with a Template particle picker. The picked 4,391,283 particles were selected using several rounds of heterogeneous refinement and ab initio reconstruction. The selected 370,215 particles were subjected to non-uniform refinement, with the optimizing per-particle defocus option. This refinement resulted in a density map for the isolated Vo with a resolution of 2.8 Å.
Model building and refinement
To build the atomic model of the isolated Vo, the atomic model of the enzyme we previously reported (PDBID: 6LY9) was used as the initial model20. The model was fitted into the density map as a rigid body, then manually refined using COOT software41. Water molecules within the model were placed manually. The manually corrected model was refined using the Servalcat software42. This iterative process was performed for several rounds to correct the remaining errors until the model satisfied the geometry. The model quality was assessed by Molprobity scores43. The statistics of maps and models presented here are summarized in Table S1. All figures were prepared using UCSF chimeraX software44.
Molecular dynamics simulations
Two simulation systems were set up. The first system includes the a, c12, and d subunits, while the second system only includes the a-subunit and c12-ring to avoid the formation of the autoinhibited interactions between the a and d subunits. After building missing residue structures with Modeller (version 10.4)45, histidine protonation types were determined with the PDB2PQR server46. Based on the high pKa predicted by PROPKA347,48, a/D365 was protonated. The membrane-embedded c(P–X)/E63 were also protonated. A disulfide bond was introduced between d/C267 and d/C322. With the CHARMM-GUI Membrane Builder49, the Vo structure was embedded in a 160 × 160 Å POPC membrane with 150 mM NaCl in solution. The all-atom CHARMM 36 m and CHARMM 36 force fields used for protein and lipids, respectively, have been extensively tested for membrane proteins50,51. The total number of atoms is ~427,000.
The systems were then minimized and equilibrated with NAMD 2.1252,53. After 5000 steps of minimization, they were equilibrated in three stages: (1) 100 ps equilibration at T = 100 K and no pressure control with restraint of non-hydrogen protein and lipid atoms, (2) 100 ps equilibration at T = 200 K and 1 atm pressure with the restraint of non-hydrogen protein atoms, (3) 1 ns equilibration at T = 310 K and 1 atm pressure with the restraint of non-hydrogen protein atoms. The restraint force constant was 1 kcal/molÅ2 for each atom. With this scheme, water, ions, and lipid molecules were well equilibrated, and the protein structure was maintained. The systems were simulated at T = 310 K and 1 atm pressure without the above restraints in the production run. For the second system lacking the d-subunit, the Cα atoms of residues 45–270 of the a-subunit that normally interact with the d-subunit were restrained to avoid any unnatural structural change. The integration timestep was 2 fs. Langevin dynamics with a 1 ps damping coefficient was used for temperature control, and the Nose–Hoover Langevin piston was used for pressure control. Each trajectory has different random forces in Langevin dynamics, making the results nondeterministic.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM density maps and atomic models obtained in this study have been deposited to both the EMDB and PDB. The accession codes (EMDBID and PDBID) are EMD-39661 and 8YXZ, EMD-39662 and 8YY0, EMD-39663 and 8YY1, for states 1–3 of the Vo domain from holo-enzyme, and EMD-39644 and 8YWT, for the isolated Vo domain. MD simulation movies are provided as Supplementary Movies 1–3.
Code availability
The code used for the molecular dynamics simulations has been uploaded to https://github.com/OkazakiLab/vo_files.
References
Boyer, P. D. The binding change mechanism for ATP synthase–some probabilities and possibilities. Biochim. Biophys. Acta 1140, 215–250 (1993).
Mitchell, P. Chemiosmotic coupling in oxidative and photosynthetic phosphorylation. Biol. Rev. Camb. Philos. Soc. 41, 445–502 (1966).
Senior, A. E. ATP synthesis by oxidative phosphorylation. Physiol. Rev. 68, 177–231 (1988).
Yoshida, M., Muneyuki, E. & Hisabori, T. ATP synthase - a marvellous rotary engine of the cell. Nat. Rev. Mol. Cell Biol. 2, 669–677 (2001).
Kuhlbrandt, W. Structure and mechanisms of F-type ATP synthases. Annu. Rev. Biochem. 88, 515–549 (2019).
Hahn, A., Vonck, J., Mills, D. J., Meier, T. & Kuhlbrandt, W. Structure, mechanism, and regulation of the chloroplast ATP synthase. Science 360, eaat4318 (2018).
Yokoyama, K. & Imamura, H. Rotation, structure, and classification of prokaryotic V-ATPase. J. Bioenerg. Biomembr. 37, 405–410 (2005).
Guo, H. & Rubinstein, J. L. Cryo-EM of ATP synthases. Curr. Opin. Struct. Biol. 52, 71–79 (2018).
Yokoyama, K., Oshima, T. & Yoshida, M. Thermus thermophilus membrane-associated ATPase. Indication of a eubacterial V-type ATPase. J. Biol. Chem. 265, 21946–21950 (1990).
Gogarten, J. P. et al. Evolution of the vacuolar H+-ATPase: implications for the origin of eukaryotes. Proc. Natl Acad. Sci. USA 86, 6661–6665 (1989).
Nakanishi, A., Kishikawa, J. I., Tamakoshi, M., Mitsuoka, K. & Yokoyama, K. Cryo EM structure of intact rotary H(+)-ATPase/synthase from Thermus thermophilus. Nat. Commun. 9, 89 (2018).
Yokoyama, K., Akabane, Y., Ishii, N. & Yoshida, M. Isolation of prokaryotic V0V1-ATPase from a thermophilic eubacterium Thermus thermophilus. J. Biol. Chem. 269, 12248–12253 (1994).
Forgac, M. Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat. Rev. Mol. Cell Biol. 8, 917–929 (2007).
Imamura, H. et al. Evidence for rotation of V1-ATPase. Proc. Natl Acad. Sci. USA 100, 2312–2315 (2003).
Nakanishi, A., Kishikawa, J. I., Mitsuoka, K. & Yokoyama, K. Cryo-EM analysis of V/A-ATPase intermediates reveals the transition of the ground-state structure to steady-state structures by sequential ATP binding. J. Biol. Chem. 299, 102884 (2023).
Rastogi, V. K. & Girvin, M. E. Structural changes linked to proton translocation by subunit c of the ATP synthase. Nature 402, 263–268 (1999).
Oster, G. & Wang, H. ATP synthase: two motors, two fuels. Structure 7, R67–R72 (1999).
Roh, S. H. et al. Cryo-EM and MD infer water-mediated proton transport and autoinhibition mechanisms of V(o) complex. Sci. Adv. 6, eabb9605 (2020).
Kishikawa, J. I. et al. Structural snapshots of V/A-ATPase reveal the rotary catalytic mechanism of rotary ATPases. Nat. Commun. 13, 1213 (2023).
Kishikawa, J. I. et al. Mechanical inhibition of isolated V(o) from V/A-ATPase for proton conductance. Elife 9, e56862 (2020).
Murata, T., Yamato, I., Kakinuma, Y., Leslie, A. G. & Walker, J. E. Structure of the rotor of the V-type Na+-ATPase from Enterococcus hirae. Science 308, 654–659 (2005).
Guo, H., Bueler, S. A. & Rubinstein, J. L. Atomic model for the dimeric F(O) region of mitochondrial ATP synthase. Science 358, 936–940 (2017).
Mazhab-Jafari, M. T. et al. Atomic model for the membrane-embedded VO motor of a eukaryotic V-ATPase. Nature 539, 118–122 (2016).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Sharpee, T. O. et al. 25th Annual computational neuroscience meeting: CNS-2016. BMC Neurosci. 17, 54 (2016).
Tani, K. & Fujiyoshi, Y. Water channel structures analysed by electron crystallography. Biochim. Biophys. Acta 1840, 1605–1613 (2014).
Gordon, J. C. et al. H++: a server for estimating pKas and adding missing hydrogens to macromolecules. Nucleic Acids Res. 33, W368–W371 (2005).
Okazaki, K. & Hummer, G. Phosphate release coupled to rotary motion of F1-ATPase. Proc. Natl Acad. Sci. USA 110, 16468–16473 (2013).
Todokoro, Y. et al. Chemical conformation of the essential glutamate site of the c-ring within thermophilic Bacillus F(o)F(1)-ATP synthase determined by solid-state NMR based on its isolated c-ring structure. J. Am. Chem. Soc. 144, 14132–14139 (2022).
Parkin, D. & Takano, M. Coulombic organization in membrane-embedded rotary motor of ATP synthase. J. Phys. Chem. B 127, 1552–1562 (2023).
Blanc, F. E. C. & Hummer, G. Mechanism of proton-powered c-ring rotation in a mitochondrial ATP synthase. Proc. Natl Acad. Sci. USA 121, e2314199121 (2024).
Kaila, V. R., Wikström, M. & Hummer, G. Electrostatics, hydration, and proton transfer dynamics in the membrane domain of respiratory complex I. Proc. Natl Acad. Sci. USA 111, 6988–6993 (2014).
Balmoos, C. Alternative proton binding mode in ATP synthases. J. Bioenerg. Biomembr. 39, 441–445 (2007).
Yokoyama, K., Nakano, M., Imamura, H., Yoshida, M. & Tamakoshi, M. Rotation of the proteolipid ring in the V-ATPase. J. Biol. Chem. 278, 24255–24258 (2003).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Yamashita, K., Palmer, C. M., Burnley, T. & Murshudov, G. N. Cryo-EM single-particle structure refinement and map calculation using Servalcat. Acta Crystallogr. D. Struct. Biol. 77, 1282–1291 (2021).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF chimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. 86, 2.9.1–2.9.37 (2016).
Jurrus, E. et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 27, 112–128 (2018).
Olsson, M. H., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Richter, M., Marquetand, P., González-Vázquez, J., Sola, I. & González, L. Correction to “SHARC - ab initio molecular dynamics with surface hopping in the adiabatic representation including arbitrary couplings” [J. Chem. Theory Comput. 2011, 7, 1253–1258]. J. Chem. Theory Comput. 8, 374 (2012).
Wu, E. L. et al. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Yu, Y. et al. CHARMM36 lipid force field with explicit treatment of long-range dispersion: parametrization and validation for phosphatidylethanolamine, phosphatidylglycerol, and ether lipids. J. Chem. Theory Comput. 17, 1581–1595 (2021).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Phillips, J. C. et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 153, 044130 (2020).
Acknowledgements
We are grateful to all the members of the Yokoyama Lab for their continuous support and technical assistance. Our research was supported by Grant-in-Aid for Scientific Research (JSPS KAKENHI) Grant Numbers 23H02453 to K.Y., 20K06514 to J.K., 22H02595 and 23K23858 to K.O. and Takeda Science foundation funding to K.Y. Our research was also supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED under Grant Number JP17am0101001 (support number 1312), and Grants-in-Aid from the “Nanotechnology Platform” of the Ministry of Education, Culture, Sports, Science and Technology (MEXT). This work was also supported by JST CREST to K.M. (Grant Number. JPMJCR1865). The computation was partially performed using the Research Center for Computational Science, Okazaki, Japan (Project: 22-IMS-C189 and 23-IMS-C201).
Author information
Authors and Affiliations
Contributions
K.Y., J.K., and K.O. designed, performed, and analyzed the experiments. A.N. and Y.N. analyzed the data and contributed to the preparation of the samples. K.M. provided technical support and conceptual advice. K.Y. designed and supervised the experiments and wrote the manuscript. All authors discussed the results and commented on the manuscript. The authors declare no conflicts of interest associated with this manuscript. All data were available in the manuscript or in the supplementary materials.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Stephan Wilkens, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kishikawa, Ji., Nishida, Y., Nakano, A. et al. Rotary mechanism of the prokaryotic Vo motor driven by proton motive force. Nat Commun 15, 9883 (2024). https://doi.org/10.1038/s41467-024-53504-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53504-x