Abstract
In a perfect world, scientists would develop analyses that are guaranteed to reveal the ground truth of a research question. In reality, there are countless viable workflows that produce distinct, often conflicting, results. Although reproducibility places a necessary bound on the validity of results, it is not sufficient for claiming underlying validity, eventual utility, or generalizability. In this work we focus on how embracing variability in data analysis can improve the generalizability of results. We contextualize how design decisions in brain imaging can be made to capture variation, highlight examples, and discuss how variability capture may improve the quality of results.
Similar content being viewed by others
Introduction
Central to the promise of neuroscience is the ongoing development and discovery of models linking brain structure and function to cognition, development, or clinical status. Common approaches involve generating maps of brain activation or organization through the complex analysis of imaging modalities, such as magnetic resonance imaging (MRI), electroencephalography (EEG), or magnetoencephalography (MEG). Efforts to identify associative and predictive relationships from brain imaging data have become increasingly prevalent, yielding success stories such as the identification of sex1 and age2 associations, alongside measures of health3,4 and psychiatric diagnoses5,6.
The lack of accessible ground truths in neuroscience leads to an abundance of approaches to generate derived measures of brain structure and function. Each contains undefinable amounts of uncertainty, contributing to a reproducibility crisis7. An abundance of analytic options in the field provide researchers with flexibility for how to get the most out of their data, which has the added unintended consequence of analyses being more susceptible to p-hacking or the propagation of significant results over robust ones—and thus compound the likelihood of inaccurate or irreproducible findings. While the growing accessibility of public tools (e.g., fMRIPrep8, HCP Pipelines9) and data (e.g., ABCD10, OpenNeuro11) may greatly simplify the reproduction (identical conditions) or replication (differing methods and/or data) of analyses, they do not automatically provide insight regarding whether findings can be robustly extrapolated across tools, configurations, or samples. Ensuring the reproducibility of results should remain a key priority for the scientific community, but the ability to reproduce findings is not enough for generalizability or progress.
With this in mind, we believe that the field is in need of a perspective shift, beyond merely striving for reproducibility, and towards striving for replicability, generalizability, and robustness. In this manuscript, we present a perspective that embracing reproducible variation across sets of results is necessary for generalizable findings and scientific advancement. We establish the context through which many processing and analysis related elements of experimental design can be perturbed, with an emphasis on capturing variation reproducibly across a range of analytic contexts. We discuss various studies that employ a range of techniques for capturing analytical variability, and highlight their implications on human neuroscience research. Finally, we discuss how variability capture can be accessibly applied in experiments, and discuss contexts when leveraging analytical variability can ultimately improve the quality of results. While the majority of applications covered in this manuscript are focused on brain imaging, the concepts of perturbation and ensembling are domain agnostic and widely applicable across the psychological or computational sciences.
Meaningfully incorporating variation into analytical designs
We define variability or variation in results as the change of a result based on perturbations performed to the experimental design, implementation, or, in some cases, numerical state. Throughout this manuscript we adhere to the definitions of reproducibility and replicability provided by Rougier et al. 12. which can be found succinctly summarized here13. In short, we refer to reproducibility as the ability for a finding to be regenerated in similar conditions (e.g., data, tools, analysts), while replicability refers to the ability of this finding to be regenerated in distinct conditions. For the context of this paper, which highlights human brain imaging, we consider that all results are imperfect — there does not exist a single silver-bullet analysis technique that dominates all others. It follows that each lone outcome or result of an experiment is by definition conditional upon the specific techniques used to generate them.
Results can be conditional on deliberate decisions, such as, the selection of a neuroanatomical parcellation, incidentally due to the infrastructure used, such as the algebraic solver performing optimization, or a combination of both. Each of these cases may lead to sampling bias in generating the scientific findings. Experiments that are highly conditional (e.g., do not incorporate variation across a range of alternatives at any stage) will tend to produce less generalizable results14. When paired with the publication file drawer effect15 and scientific momentum, non-generalizable findings may have long lasting effects on scientific disciplines. To increase the generalizability of results, a researcher may instead reduce the degree to which their finding is conditional by sampling variation in their approach, and attempt to replicate the study.
To avoid ambiguity, it is important to state that not all attempts to sample variation are, in practice, equal. Throughout this manuscript we will only consider attempts to sample variations that do not involve making analytical decisions that are not knowably less correct than others, but rather only those that serve as plausible alternatives. Put another way, experimental designs that are known to be inappropriate or incorrect (e.g., incompatible model specification and dataset) should always be removed from consideration, and are not considered meaningful variants in this context.
Box 1 illustrates potential levels at which variation can be introduced into experiments — including data selection, expert opinion, analytic decisions, tool selection, computational infrastructure, and numerical state. Along with each, we suggest possible (non-exhaustive) questions that prompt how variation at each stage can be explored. An important first step towards increasing generalizability is to identify which experimental components contain degrees of freedom that are likely to have a noticeable impact (bias) on the results. While generating such hypotheses is not always possible in the absence of prior exploration, consulting existing literature on methods development, for example, can inform this process. Similarly, not all sources of bias enumerated in Box 1 can be similarly perturbed. Data selection, for example, is a predominantly pre-analytic source of bias, while the others are largely introduced in the analytic or post-analytic stages. Data-related biases, such as in participant sampling, measurement technique, or incomplete data capture, cannot necessarily be perturbed to achieve external validation without great additional cost. While data sharing efforts have mitigated this cost for a variety of common designs and populations of interest, it remains that this is far from a universal luxury, often leaving the axis of data variability unexplored. Further, it is worth noting that the statistical and scientific utility of shared data decays overtime due to repeated use16. Given the breadth of literature on the issue of dataset related biases across a variety of scientific domains17,18,19,20, and the noted conceptual inconsistency with the analytical components of experimental design, we will largely direct attention on data-related biases to the above-referenced works on this issue, and focus the discussions within this perspective on specific cases that can be handled analytically.
Once variation has been captured, researchers must then decide how to incorporate it into their experiment. Perhaps the most common type of analyses that seek out and incorporate design variability in the psychological sciences are meta-analyses. Meta-analyses are a widely accepted technique for deriving consensus findings across multiple evaluations of similar experimental questions, and their results are often considered the strongest form of scientific evidence21. Post hoc meta-analyses, however, are limited by publication bias, and ultimately are susceptible to perpetuate widely held biases in the literature22,23. This crucially means that capturing and analyzing variability as a core part of primary experimental designs themselves is essential—when exploring the space of analytic and processing methods, this is typically referred to as multiverse analysis24.
In the following sections, we discuss several approaches for capturing and quantifying the impact of variation that have been used across all analytic stages of brain imaging experiments, and highlight the impact they have had on the interpretation of findings and the field. We will then explore various approaches that either help with the capture of variation, or demonstrate how variation can be used in analyses.
The importance of capturing analytic variation in brain imaging experiments
While there exist limitless possible representations of experiments and their components25, Fig. 1 coarsely depicts one such representation in which experiments are composed of three (more or less) sequential stages: (i) sampling and data collection, (ii) processing & curation, and (iii) analysis. Orthogonally, there exist opportunities for introducing analytical variation into each of these stages through one or more sources, as noted in Box 1: data selection, expert opinion, analytic decisions, tool selection, computational infrastructure, and numerical state. Even at this coarse resolution, combinatorially there exist thousands of possible configurations for how variation may be introduced for a given experiment.
In this illustration, we break down experiments into three coarse stages: sampling and data collection, processing and curation, and analysis. By looking at the elements of experiments one can perturb at each of the analytic stages, you can see how various forms of common experimental designs are performed. Importantly, any of these elements can be perturbed either jointly or independently, and doing such explorations is likely to provide insight into the generalizability or robustness of results.
Variability can be something that is deliberately introduced (e.g., comparing alternative processing tools), appear incidentally (e.g., switching computational systems), or a combination of both (e.g., a replication attempt of prior work with incomplete reporting). When designing such experiments, unavoidable questions arise with respect to responsibility and negligence. How many alternatives should be considered? Which elements of design should be perturbed? Where is the line between deliberate and incidental variation?
Let’s take the example of operating system variation, which has been shown to have an impact on brain imaging results26. Lacking any principled expectation for why this analysis detail may lead to significant variation in results, this was initially uncovered as an incidental source of variation. However, with the knowledge of this result, operating system selection now may become a deliberate decision for researchers. Before claiming that ignoring this source of variation would err on the side of negligence, it is essential to first ask the question of how it impacts the study being performed.
While differences due to operating systems may be statistically significant, are they impactful for the downstream derivatives being studied, the group-differences being explored, or otherwise? If the answer appears to be no, then it would be reasonable to leave it there and move on to the next source of variation. If yes, it then becomes a matter of scoping how this variation can be incorporated into an experiment. It would then be unreasonable and wasteful to expect that every operating system and version be tested moving forward, for example. In this particular case, it may be considered responsible to instead consider 2–3 alternatives that are generally considered as common alternatives, such as switching between CentOS and Ubuntu. In the general case, it also holds that exhaustively permuting every possible source of variation will inevitably be intractable. As is the case in all elements of experimental design, it is the researcher’s (and reviewers’) responsibility to justify (and evaluate) the selections based on the current understanding of which factors may limit generalizability in the domain.
The following section begins to grapple with these questions, and showcases real-world applications of the approaches described in Fig. 1 alongside what they have taught us about the state of generalizability in brain imaging research.
Impact of variability in data selection
It has long been accepted in the life sciences that the selection of participants plays a substantial role in the outcomes of scientific studies. As a result, the generalizability of results across samples places a bound on their importance. Data sharing efforts such as the Consortium of Reliability and Reproducibility27, Adolescent Brain Cognitive Development10, and Human Connectome Project28 studies have dramatically increased the availability of healthy control samples with test-retest data, and can be used to pool data into large collections or evaluate the generalizability of results in some cases. While similar resources may exist for more specific populations (e.g., to study disease, aging, or particular regions), such as the Alzheimer’s Disease Neuroimaging Initiative29 study, these resources are inconsistently available, and are thus likely to suffer from dataset decay more rapidly16. When observing phenomena with exceedingly small effect sizes, or otherwise performing a low-powered study, these open samples may provide little added value due to the broad heterogeneity, and (true) results may still fail to replicate30.
In part, these issues may be exacerbated by limitations in the reliability of either imaging or phenotypic data31,32,33. Further, it remains a challenge to produce generalizable findings while lacking epidemiologically representative samples. In these cases, naturally occurring heterogeneity is dramatically underrepresented in the data, which artificially inflates the importance of the variability which is represented. Adopting a “population neuroscience” approach17,34 is necessary as the field continues to advance. In cases when large epidemiologic samples are not available—and it is not possible to collect them—the best-case for evaluating generalizability may be through the use of external datasets containing samples with significantly different social determinants of health. Otherwise, the only available alternatives for evaluating generalizability remain limited internal validation approaches such as cross-validation35.
Impact of variability in expert opinion
As science has evolved as an increasingly social enterprise, the many-analyst approach36 has grown in prevalence. Organizers of many-analyst studies typically provide curated datasets and specific objectives detailing the derived data to be produced and the inferences to be performed. Given the deliberate lack of instruction regarding the methods to be used, the consistency of results across problem formulations was then evaluated. This has been attempted numerous times in brain imaging, including in the analysis of task-based functional analysis by the Neuroimaging Analysis Replication and Prediction Study37 (NARPS), the generation and segmentation of diffusion-MRI based tractograms38, and with other studies ongoing at the time of writing, including the EEGManyPipelines project. In the case of NARPS37, the 70 participating teams each used unique processing and statistical testing workflows, which ultimately led to considerable disagreement in the outcomes of tested hypotheses—despite often high similarity in the functional maps used. This phenomenon was mirrored in the case of tractometry analysis, in which 42 teams presented 57 unique segmentation protocols when tasked with identifying 14 white-matter pathways38 from identically pre-processed data. Importantly, this exploration found that the variability observed across results generated from each of these largely expert-driven segmentation protocols was larger than the variation introduced by upstream processing differences. Similar results were found in the context of connectomics, in which the selection of brain parcellations—often considered a subjective design decision—were found to significantly change the distribution of derived network statistics39. These explorations demonstrate that expert preferences in decision making are a dominant factor in the generation and interpretation of results.
Impact of variability in analytic decisions
While several dominant pipeline packages exist in each modality of brain imaging, their component pieces or parameterizations vary considerably across tools. When processing identical data using 5 commonly-used fMRI preprocessing pipelines, the median correlation of derived connectomes for conceptually identical pipelines was found to range between a correlation of 0.50–0.82 across tools40. When the pipeline components were perturbed to make the tools more similar to one another, it was notable that no single stage of processing (e.g., structural normalization, denoising) was solely responsible for the observed differences. In fact, making the pipelines more similar conceptually reduced the numerical similarity of the results in several cases, demonstrating the complexity of interactions among their components and an inability to predict the similarity of results a priori. Beyond suffering from a lack of comparability across results, the reliability of derived data may be inconsistent across pipelines. Leveraging the flexibility of the Configurable Pipeline for the Analysis of Connectomes (C-PAC), reliability was evaluated for each of 192 unique pipelines41. Averaged across 28 test-retest datasets, pipeline reliability ranged from 0.85–0.95. While individual dataset-pipeline pairs peaked near 1.0, or dropped below 0.75 in some cases, there was importantly no uniformly dominant pipeline. Importantly, this lack of dominance was also observed on a more local scale, when considering datasets with highly similar acquisition parameters. Together these approaches highlight both the potentially significant bias that results generated with a single pipeline may contain, and demonstrate that the selection of a single dominant pipeline remains infeasible.
Impact of variability in tool selection
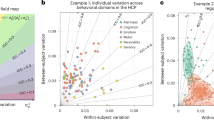
When matching the conceptual approach taken to analyze data, the implementations of algorithms can play a role in the determination of results. Recently, three task-based fMRI studies were carried out using each of the three dominant fMRI analysis libraries42. After matching the techniques and strategies applied, despite achieving qualitative similarity across the generated activation maps, thresholded statistical maps across the tools were found to range in overlap (Dice) coefficients from 0, indicating no overlap, to 0.684 (ideal: 1). When interrogating these differences further, it was found that while some steps consistently produced little difference across implementations (e.g., modeling of low-frequency signal drift), there was no single step that consistently drove significant differences43. When performing a similar exploration in the context of cortical surface estimation, the correlation between cortical thickness measures across software libraries did not exceed 0.52 or 0.66 on typical- and high-quality datasets, respectively44. Joint analysis of cortical features showed that cross-tool differences significantly overshadowed individual (i.e., participant-level) differences. Specifically, the distributions of results with respect to subject and disease varied significantly across tools, as well as the relative importance of derived features of these distributions. These findings demonstrate a severe limitation in the generalizability of learned relationships across tools individually, underscoring that variability in tool selection may be necessary to understand the trustworthiness of the presence or absence of significant results, as well as the structure of individual- and group-level relationships that may be discovered.
Impact of variability in computational infrastructure
Many of the differences described up to this point may be relatively straightforward to accept from a scientific point of view. The infrastructure underlying these elements, however, are often not considered as similarly relevant. In addition to the libraries being used, software versions may play an appreciable role in results26. In the case of cortical surface estimation, correlations across three commonly-used versions of FreeSurfer have been found to range from 0.83–0.8944 for identical data; even assuming monotonic progress of the tools, this poses significant limitations on backwards-reproducibility. When using a fixed tool version, modifying the versions of data dependencies used, such as atlas templates, was also found to lead to significant variation in results. In the case of fMRI analysis, varying the near-ubiquitous MNI152 template45,46—the version of which is often unreported—within fMRIPrep showed that identical individual data may have correlations below 0.840. More easily overlooked, the version and flavor of the operating system used during analysis can significantly influence results26. These differences are the result of changes to foundational numerical libraries, and importantly do not suggest that results generated using one configuration are superior to any other. While an individual study may not be likely to vary software, template, or operating system versions internally, the significance of these effects suggests that results are far more dependent on these hidden factors than once believed, and even well-calibrated replication attempts may fail if variability across infrastructures is not explored.
Impact of variability in numerical state
At a lower level than differences in infrastructures are numerical states, which remain far more obfuscated. While the field of uncertainty quantification exists in part to define and understand variation across numerical states, for the purposes of this review we can divide these into two categories: random effects and error. In the case of random effects, variability may be introduced into results due to the initialization of models. In one recent study, both the random state (e.g., the starting-point for optimizations) and model parameterization were found to lead to significant differences in performance of machine learning models of brain imaging data47. The random state of commonly used algorithms, such as nonlinear image normalization, has been found to similarly influence the results of these processes. From the perspective of error, often overlooked are the limitations of how modern computers handle decimal floating point data. Given a finite space for storing data, mathematical operations performed on these data are imperfect by nature48. In practice, floating point errors commonly manifest themselves in analyses through either a gradual loss of precision over the course pipelines, effectively resulting in a drop in signal-to-noise ratio (SNR) throughout, or a steep drop in precision, such as by solving poorly conditioned problems49. In the context of structural connectome estimation, the in-place perturbation of numerical errors were shown to lead to variation on the order of individual differences50. Interestingly, while this was true for individual networks and both the univariate and multivariate statistics derived from them, the group-level sample characteristics were robust to such perturbations51. This shows that just as increasing the sample size of a study may be necessary to reduce the likelihood of false positives, increasing the number of numerical states sampled in processing may be similarly necessary.
The inconsistency in results across each of the domains of variation explored above necessitates a shift towards multiverse analysis in order to achieve reproducibility in the field of brain imaging. While each of the discussed approaches for capturing and evaluating the impact of methodological variations on results have historically been performed independently, they complement one another and, when used together, can paint a clearer picture of discoveries.
Incorporating and documenting the exploration of analytical variability into experiments
The above section demonstrates the often considerable significance of variability in analyses, and how it can affect published results. However, challenges remain in how to conduct such explorations broadly; in each of the described works, the evaluation of variability was the primary focus of the studies, often in lieu of testing novel downstream hypotheses or models. This is likely driven in part by the reality that capturing variability across experimental designs typically requires a several-fold increase in the computational effort required, data produced, and results analyzed. In parallel, the clear communication of such results is often hampered in publication by strict word and figure limits, particularly in high-impact journals—often necessary to ensure clear and concise reporting,—leaving little room for these types of validation alongside the communication of novel results. In this section, we will raise awareness of several initiatives, tools, and communities that can aid in both the capture and evaluation of analytical variability.
Capturing variability across data selection
As mentioned previously, the external validation of results across datasets is inherently limited by the availability of comparable data resources. Many platforms and initiatives have been developed to provide vast collections and entry points for such validations. Data resources, such as ABIDE52 for Autism Spectrum Disorders, provide accessible entry points for replication in specific disease populations. Alternatively, Neurovault53 and downstream analysis tools (e.g., NeuroQuery54, NeuroSynth55) support meta-analysis based evaluation of findings, typically for non-disease-centric cognitive neuroscience studies.
In cases where such external resources do not capture the population, modalities, or experimental paradigms of interest, techniques such as cross-validation serve as essential guides for internal validation. Importantly, cross-validation is not a silver-bullet solution, and when mis-used can lead to false confidence in results35,56,57. Even when properly applied, cross-validation still does not ensure robust estimates of generalization error given, for example, the significant presence of batch effects present in medical imaging datasets, e.g., ref. 58. Ultimately, while analytic sources of bias can be identified and overcome iteratively throughout experimental development and execution, appropriately capturing variability in data requires thoughtful proactive evaluation of the specific population(s) that are being targeted by the study, followed by a population neuroscience approach to sampling.
Capturing variability across expert opinions
Recently, cross-team explorations have taken place in the brain imaging community through self-organization on either Twitter/X or the Brainhack59 public forum. Through these communities, the 70-team NARPS project37, the 42 teams tractometry project38, and the ongoing EEGManyPipelines projects have produced incredible benchmarks for the community. Similarly, from the perspective of downstream modeling, challenge platforms such as Kaggle have been used to great effect to both estimate the variance in approaches used in the field, and evaluate out of sample performance on prediction or association tasks57. In each case, the formula for this approach remains consistent. The organizers first define a well-scoped challenge, wherein both the form of the data to be used for analysis and the scientific question are concretely defined. With these instructions, the teams then apply their chosen approach to independently solve the challenge, and report their conclusions in a standardized format, alongside the methodological details. The organizers then evaluate the variation in each of the approaches using a number of qualitative or quantitative strategies, as appropriate for the domain. The NARPS and tractometry studies referenced earlier in this section are both excellent examples that can be followed when applying this approach to other domains. In a broader sense, this approach is similar to citizen science, which has been applied to great effect in a variety of domains, and famously through platforms or communities such as Mechanical Turk and Kaggle.
Capturing variability across analytic decisions, tool selection, and computational infrastructure
While social engineering and organization can be considered as universally relevant solutions, capturing variation across pipelines and libraries unavoidably remains a domain-specific question. In the case of structural and functional MRI analysis, the Configurable Pipeline for the Analysis of Connectones60 (C-PAC) has been developed specifically to address these questions. C-PAC integrates processing tools from a variety of commonly used libraries, and provides replicated versions of several of the major fMRI preprocessing pipelines (e.g., fMRIPrep8, HCP Pipelines9). While this platform provides access to end-to-end pipelines and the perturbation of their components across libraries, it, or equivalent tools, only apply to a single scientific domain. Context-independent tools such as Boutiques61 or Common Workflow Language62 (CWL) can be used to facilitate comparisons across arbitrary pipelines and infrastructures, as it wraps tools with a consistent interface, allowing for the simplified deployment across large computing resources. Once results are generated using multiple configurations, provenance-rich versioning software such as DataLad63 can facilitate both the unambiguous storage of results, and makes it easy to either move between versions of results for re-analysis, or consider multiple versions of derived data simultaneously. While Datalad and Boutiques have largely been applied in the brain imaging community, they are flexibly domain agnostic.
Capturing variability across numerical state
Many tools that depend upon random processes, such as optimization, support the specification of random states. In addition to the repeated evaluation of tools across these random configurations, techniques such as Monte Carlo Arithmetic48 allow for the perturbation of numerical errors that are inherent to the executed software. Fuzzy64 provides transparent environments for scientists to run their software and evaluate the significance of numerical error in their results. By processing data using Fuzzy, distributions of equally-valid and likely results can be obtained across various manifestations of numerical error. It is common to evaluate results across the repeated “fuzzy” executions by computing the number of significant digits, or estimating the likelihood of pipeline failures. This has recently been used in the evaluation of the fMRIPrep Long-Term Support release8 to identify drift across pipeline package development, and successfully highlighted a release when significant algorithmic change was introduced without supervision65.
Each of the described techniques and tools takes a unique approach to capture, summarize, or describe, variability across experiments. Given the complementary nature of these techniques, it is possible that any combination of such approaches may be used concurrently. With a broad foundation of results motivating the inclusion of such variability throughout brain imaging research, what remains to be solved is a convenient and concise vehicle for incorporating and communicating these results into novel studies. In the interest of providing an actionable resource for the perspective presented here, we propose such a checklist to aid in designing and reporting multiverse experiments, that can be found online at https://www.nmind.org/variability-capture-checklist, or more strictly version-controlled on GitHub at https://github.com/nmind/. This resource can be used by authors to either inspire or directly summarize variability explorations in studies, alongside their impact on presented conclusions. The checklist is structured into three main blocks that help authors describe how variability was explored in their study, how the results of this exploration were used in analysis, and what they believe the impact of this exploration to be specifically on their findings, or how it may be extended further.
This is by no means the first checklist interested in enhancing the reproducibility and quality of research. Similar efforts, such as the COBIDAS report66, have been designed to describe an experiment from data collection through to analysis. While COBIDAS contains a section on reproducibility specifically, it focuses on software and data availability rather than the steps taken to measure or achieve generalizability. Given the primary focus of the checklist presented here is to consider the degree of bias or variation considered in an analysis, and is complementary to the COBIDAS mission.
How analytical variability can improve transparency and generalizability of results
Capturing variability affords researchers several opportunities to both evaluate the limitations or sensitivity of their work with respect to experimental configuration, and to mitigate these effects. When looking across arrays of perturbed results, techniques can be applied to either quantify confidence, identify sources of deviations, or construct more representative and generalizable results. We will discuss cases in which each of these approaches has been applied in the context of brain imaging and how they can be adapted or applied for future studies.
Quantifying confidence in results
Measures such as number of significant digits/bits, coefficient of variation, or confidence intervals, estimate the bound of quality on obtained numerical results of experiments—and subsequently temper the strength of downstream claims. Recently, significant digits— defined as the number of unchanging digits across all sampled variants of a result—were used to evaluate the stability of two structural connectome estimation pipelines51. By measuring the significant digits of every edge in the connectome, a confidence-connectome was constructed which highlighted the connections that were more or less robust50. This approach could be used to identify subsets of data most likely to contain reliable signal, even in the absence of test-retest data. Further, this study showed that numerical errors alone may contribute to variation on the level of individual differences, significantly distorting the ability to make individual predictions from data—while group-level statistics were, crucially, stable.
Pairing numerical perturbation with measurement of significant digits can be seamlessly applied across data regardless of the specific domain or properties of the analytic perturbations. Crucially, approaches such as these can be used to establish and communicate the level of trust a reader may have in the results. Using Fuzzy-tools in this context may dramatically lower the barrier to performing repeated-measures analyses67, such as a repeated-measures ANOVA68, allowing researchers to identify the impact of variability in their analysis strategy. Beyond structured analysis approaches, evaluating and reporting results across experimental perturbations also reduces the likelihood of either accidental or deliberate p-hacking in results. In these cases, pre-registration69 becomes an essential tool in ensuring accountability.
Identifying sources of analytical variation
Beyond characterizing the stability of conclusions, sources of variation can be identified by applying similar approaches at various stages of long processing workflows. While in some cases this tracing requires manual intervention due to varied infrastructures43, tools like PyTracer can be used to instrument software and construct both processing graphs and consolidated evaluations of result consistency across executions. In contexts when it is possible to perturb many conceptually-orthogonal components of a design, a metastudy approach70 can be used. Applied in the context of evaluating design elements in cognitive testing, this technique was able to identify the dependence of results on specific components of the design (e.g., subliminality of cue presentation), while identifying others as unimpactful towards driving variations (e.g., cue color).
While the specific metastudy approach cited above uses a Bayesian modeling framework, structural equation modeling techniques such as Intra-Class Effect Decomposition71 (ICED) can also be used to conveniently characterize the impact of variations across studies. In a less structured approach, brain imaging pipelines were perturbed by swapping processing steps across four tools40, and then a to-from table was constructed to demonstrate the impact of each perturbation. What resulted was a clear demonstration that not only was the impact of variation relatively evenly spread across components, but commonly undocumented pipeline components—namely, the data write-out resolution and the version of the MNI152 template used,—play a significant role. A broad exploration into the impact of variability across analytic stages can be used to inform the sampling of variation in future experiments, ensuring an efficient and broad characterization of the experimental space.
Generating representative results
Perhaps the most exciting and least often exploited avenue for incorporating variability into experimental designs in brain imaging is in the generation of robust or representative results. In these cases, the core underlying assumption is that all of the distinct results contain some portion of non-overlapping signal from one another, and thus can be complementary used. Following the Fuzzy evaluation of structural connectome estimation pipelines, we previously demonstrated that incorporating the variability observed across numerical error in the construction of predictive brain-phenotype models led to an increase of classification performance and generalizability to unseen data72. Importantly, we demonstrated that this simple dataset augmentation ensemble approach can be applied agnostically across measurements or techniques, and does not require more complex approaches than averaging of results across perturbations in order to improve their quality significantly. When exploring sampling variability, the bootstrapped analysis of stable clusters73,74 (BASC) approach has previously been developed to identify the stability of resting state networks across the dataset. Similarly, the multi-atlas MUSE approach to image segmentation75 has been developed to optimize image data normalization based on the premise that an array of distinct parcellations provide complementary information that may vary across the brain region and individual differences. In each of these approaches, typically-fixed elements of data analysis (e.g., numerical error, sample, template) were varied and the resulting array of outcomes were ensembled to refine the final results. For more details on a comparison of various ensembling approaches in the context of connectomics, please refer to ref. 72. While each of these examples demonstrate a machine learning approach to ensemble the results, meta-analytic frameworks are currently a topic of active development, and present additional challenges due to the unclear dependence of results across groups.
While variability-based augmentation is an important concept and powerful technique to improve generalizability, it is not a silver bullet that will resolve all analytical biases or issues. In fact, there are two scenarios in which the method can actually worsen the situation. First, in cases where an analysis introduces two or more alternatives that are highly collinear, it is possible that the method will increase the variability without substantially improving the generalizability—with the end result being a reduction in the overall signal-to-noise ratio (SNR) of the experiment. Second, in cases where poor techniques are incorporated, the variation added may not be particularly meaningful, and as a result also lead to a reduction in SNR. In each of these extremes, the act of defining a threshold for techniques that are too similar, or poor, respectively, remains empirical work. One recently proposed solution to identifying a threshold is through variance-driven truncation72. In this approach, the researcher constructs models using the unchanged portion of the signal across perturbations, and then compares the outcome against the initial result prior to perturbation. If highly similar, then a researcher knows that variability capture across the particular domain tested would add little variation to improve the generalizability of the results; if dramatically different, then variation may contribute sufficient noise to overpower the remaining signal. In cases where there is a reduction in model performance, but the relationship remains trending towards significance, we expect the augmentation of models across perturbations to be beneficial in terms of improving results generalizability. Ultimately, the importance of evaluating the validity of perturbations performed in each particular experimental context cannot be overstated, and requires careful and ongoing consideration.
Discussion
In this piece, we have clarified that robust or generalizable findings are those that hold across many degrees of variation. A natural consequence of the exploratory and iterative nature of scientific advancement is that the outcomes of all experiments are imperfect, and we can accelerate discovery by observing these outcomes across as many degrees of experimental freedom as possible—such as processing approach, datasets, or inferential techniques. There are a growing number of approaches that facilitate variability capture and incorporation across a range of experimental stages that can be used to temper confidence, or improve overall result quality.
A key consideration when applying the techniques presented here are the associated increases in the cost of running experiments. Varying parameters of processing, whether through numerical error, processing parameters, or otherwise, may lead to a several fold multiple in time for completing experiments, cost, and of course the associated environmental impact due to energy consumption. To balance each of these trade-offs, it may often be important to perform variability capture in preliminary studies so that the effect of design variability may be understood for the proposed experiment. However, maintaining a close correspondence between the preliminary and full-scale study in such cases is essential; while one study may demonstrate a relative sorting of the impact of variability sources, a lack of theoretical understanding of these variations makes extrapolating such relationships across experiments dangerous. Broadly adopting open science practices—data and code sharing, open access publishing—will further lower the barrier to both producing and evaluating results and the impact of variations transparently, and reduce duplication of effort. Cost can further be mitigated in many cases by partnerships with accessible computing resources made available to scientists, such as the Digital Alliance in Canada, the NSF-supported ACCESS network in the United States. Each of these academic-facing networks aim to provide low-to-no-cost solutions for large-scale computational analysis for researchers.
With respect to non-financial costs associated with incorporating variability into studies, such as additional time, planning, and experimental complexity—and therefore likelihood of software bugs—the authors again refer to the adoption of open science practices, in particular as many of these costs may have already been paid by the community. Leveraging existing open-science software solutions, such as fMRIPrep and C-PAC configurable pipelines in brain imaging, can dramatically reduce the likelihood of introducing new bugs in processing and analysis, and each contain years of insight in their designs as to which analytic designs may make sense and should be implemented/considered. Active support forums such as NeuroStars provide avenues for collaboration, seeking feedback, and getting assistance or clarification.
Alongside our showcased examples of variability capture throughout brain imaging and psychology research, we have discussed techniques that can be applied to induce, capture, and leverage variability across the computational sciences more broadly. Ultimately, we hope this piece will aid in the generation of more robust and generalizable scientific contributions in the life sciences and beyond.
References
Ritchie, S. J. et al. Sex differences in the adult human brain: evidence from 5216 UK biobank participants. Cereb. Cortex 28, 2959–2975 (2018).
Aycheh, H. M. et al. Biological brain age prediction using cortical thickness data: a large scale cohort study. Front. Aging Neurosci. 10, 252 (2018).
Hidese, S. et al. Association of obesity with cognitive function and brain structure in patients with major depressive disorder. J. Affect. Disord. 225, 188–194 (2018).
Ledig, C., Schuh, A., Guerrero, R., Heckemann, R. A. & Rueckert, D. Structural brain imaging in Alzheimer’s disease and mild cognitive impairment: biomarker analysis and shared morphometry database. Sci. Rep. 8, 11258 (2018).
Tolan, E. & Isik, Z. Graph theory based classification of brain connectivity network for autism spectrum disorder. in Bioinformatics and Biomedical Engineering 520–530 (Springer International Publishing, 2018).
Zhu, X., Du, X., Kerich, M., Lohoff, F. W. & Momenan, R. Random forest based classification of alcohol dependence patients and healthy controls using resting state MRI. Neurosci. Lett. 676, 27–33 (2018).
Open Science Collaboration. PSYCHOLOGY. Estimating the reproducibility of psychological science. Science 349, aac4716 (2015).
Esteban, O. et al. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16, 111–116 (2019).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80, 105–124 (2013).
Volkow, N. D. et al. The conception of the ABCD study: from substance use to a broad NIH collaboration. Dev. Cogn. Neurosci. 32, 4–7 (2018).
Markiewicz, C. J. et al. OpenNeuro: an open resource for sharing of neuroimaging data. bioRxiv 2021.06.28.450168 https://doi.org/10.1101/2021.06.28.450168 (2021).
Rougier, N. P. et al. Sustainable computational science: the ReScience initiative. PeerJ Comput Sci. 3, e142 (2017).
Plesser, H. E. Reproducibility vs. replicability: a brief history of a confused terminology. Front. Neuroinform. 11, 76 (2017).
Charter, R. A. Methodological commentary: effect of measurement error on tests of statistical significance. J. Clin. Exp. Neuropsychol. 19, 458–462 (1997).
Rosenthal, R. The file drawer problem and tolerance for null results. Psychol. Bull. 86, 638 (1979).
Thompson, W. H., Wright, J., Bissett, P. G. & Poldrack, R. A. Meta-research: dataset decay and the problem of sequential analyses on open datasets. Elife 9, e53498 (2020).
Paus, T. Population neuroscience: why and how. Hum. Brain Mapp. 31, 891–903 (2010).
Nielsen, M., Haun, D., Kärtner, J. & Legare, C. H. The persistent sampling bias in developmental psychology: a call to action. J. Exp. Child Psychol. 162, 31–38 (2017).
Charpentier, C. J. et al. How representative are neuroimaging samples? Large-scale evidence for trait anxiety differences between fMRI and behaviour-only research participants. Soc. Cogn. Affect. Neurosci. 16, 1057–1070 (2021).
Li, J. et al. Cross-Ethnicity/race Generalization Failure of RSFC-Based Behavioral Prediction and Potential Consequences. https://juser.fz-juelich.de/record/910372 (2022).
Guyatt, G. H. et al. Users’ guides to the medical literature: IX. A method for grading health care recommendations. JAMA 274, 1800–1804 (1995).
Jennings, R. G. & Van Horn, J. D. Publication bias in neuroimaging research: implications for meta-analyses. Neuroinformatics 10, 67–80 (2012).
Traut, N. et al. Cerebellar volume in autism: literature meta-analysis and analysis of the autism brain imaging data exchange cohort. Biol. Psychiatry 83, 579–588 (2018).
Steegen, S., Tuerlinckx, F., Gelman, A. & Vanpaemel, W. Increasing transparency through a multiverse analysis. Perspect. Psychol. Sci. 11, 702–712 (2016).
Guest, O. & Martin, A. E. How computational modeling can force theory building in psychological science. Perspect. Psychol. Sci. 16, 789–802 (2021).
Glatard, T. et al. Reproducibility of neuroimaging analyses across operating systems. Front. Neuroinform. 9, 12 (2015).
Zuo, X.-N. et al. An open science resource for establishing reliability and reproducibility in functional connectomics. Sci. Data 1, 140049 (2014).
Van Essen, D. C. et al. The WU-Minn human connectome project: an overview. Neuroimage 80, 62–79 (2013).
Jack, C. R. Jr et al. The Alzheimer’s Disease Neuroimaging Initiative (ADNI): MRI methods. J. Magn. Reson. Imaging 27, 685–691 (2008).
Button, K. S. et al. Power failure: why small sample size undermines the reliability of neuroscience. Nat. Rev. Neurosci. 14, 365–376 (2013).
Regier, D. A. et al. DSM-5 field trials in the United States and Canada, Part II: test-retest reliability of selected categorical diagnoses. Am. J. Psychiatry 170, 59–70 (2013).
Nikolaidis, A. et al. Suboptimal phenotypic reliability impedes reproducible human neuroscience. bioRxiv 2022.07.22.501193 https://doi.org/10.1101/2022.07.22.501193 (2022).
Gell, M. et al. The burden of reliability: how measurement noise limits brain-behaviour predictions. bioRxiv 2023.02.09.527898 https://doi.org/10.1101/2023.02.09.527898 (2023).
Falk, E. B. et al. What is a representative brain? Neuroscience meets population science. Proc. Natl. Acad. Sci. USA 110, 17615–17622 (2013).
Varoquaux, G. Cross-validation failure: small sample sizes lead to large error bars. Neuroimage 180, 68–77 (2018).
Silberzahn, R. et al. Many analysts, one data set: making transparent how variations in analytic choices affect results. Adv. Methods Pract. Psychol. Sci. 1, 337–356 (2018).
Botvinik-Nezer, R. et al. Variability in the analysis of a single neuroimaging dataset by many teams. Nature 582, 84–88 (2020).
Schilling, K. G. et al. Tractography dissection variability: what happens when 42 groups dissect 14 white matter bundles on the same dataset? Neuroimage 243, 118502 (2021).
Fornito, A., Zalesky, A. & Bullmore, E. T. Network scaling effects in graph analytic studies of human resting-state FMRI data. Front. Syst. Neurosci. 4, 22 (2010).
Li, X. et al. Moving beyond processing and analysis-related variation in neuroscience. bioRxiv 2021.12.01.470790 https://doi.org/10.1101/2021.12.01.470790 (2021).
Bridgeford, E. W. et al. Eliminating accidental deviations to minimize generalization error and maximize replicability: Applications in connectomics and genomics. PLoS Comput. Biol. 17, e1009279 (2021).
Bowring, A., Maumet, C. & Nichols, T. E. Exploring the impact of analysis software on task fMRI results. Hum. Brain Mapp. 40, 3362–3384 (2019).
Bowring, A., Nichols, T. E. & Maumet, C. Isolating the sources of pipeline-variability in group-level task-fMRI results. Hum. Brain Mapp. https://doi.org/10.1002/hbm.25713 (2021).
Bhagwat, N. et al. Understanding the impact of preprocessing pipelines on neuroimaging cortical surface analyses. Gigascience 10, giaa155 (2021).
Fonov, V. S., Evans, A. C., McKinstry, R. C., Almli, C. R. & Collins, D. L. Unbiased nonlinear average age-appropriate brain templates from birth to adulthood. Neuroimage Suppl. 1, S102 (2009).
Lancaster, J. L. et al. Bias between MNI and Talairach coordinates analyzed using the ICBM-152 brain template. Hum. Brain Mapp. 28, 1194–1205 (2007).
Bouthillier, X. et al. Accounting for variance in machine learning benchmarks. Proceedings of Machine Learning and Systems 3, (2021).
Parker, D. S. Monte Carlo Arithmetic: Exploiting Randomness in Floating-Point Arithmetic (University of California (Los Angeles). Computer Science Department, 1997).
Skare, S., Hedehus, M., Moseley, M. E. & Li, T. Q. Condition number as a measure of noise performance of diffusion tensor data acquisition schemes with MRI. J. Magn. Reson. 147, 340–352 (2000).
Kiar, G. et al. Comparing perturbation models for evaluating stability of neuroimaging pipelines. Int. J. High Perform. Comput. Appl. 34, 491–501 (2020).
Kiar, G. et al. Numerical uncertainty in analytical pipelines lead to impactful variability in brain networks. bioRxiv 2020.10.15.341495 https://doi.org/10.1101/2020.10.15.341495 (2021).
Di Martino, A. et al. The autism brain imaging data exchange: towards a large-scale evaluation of the intrinsic brain architecture in autism. Mol. Psychiatry 19, 659–667 (2014).
Gorgolewski, K. J. et al. NeuroVault.org: a web-based repository for collecting and sharing unthresholded statistical maps of the human brain. Front. Neuroinform. 9, 8 (2015).
Dockès, J. et al. NeuroQuery, comprehensive meta-analysis of human brain mapping. Elife 9, e53385 (2020).
Yarkoni, T., Poldrack, R. A., Nichols, T. E., Van Essen, D. C., & Wager, T. D. Large-scale automated synthesis of human functional neuroimaging data. Nat. Methods 8, 665–670 (2011).
Krstajic, D., Buturovic, L. J., Leahy, D. E. & Thomas, S. Cross-validation pitfalls when selecting and assessing regression and classification models. J. Cheminform. 6, 10 (2014).
Traut, N. et al. Insights from an autism imaging biomarker challenge: promises and threats to biomarker discovery. Neuroimage 255, 119171 (2022).
Olivetti, E., Greiner, S. & Avesani, P. ADHD diagnosis from multiple data sources with batch effects. Front. Syst. Neurosci. 6, 70 (2012).
Gau, R. et al. Brainhack: developing a culture of open, inclusive, community-driven neuroscience. Neuron 109, 1769–1775 (2021).
Craddock, C. et al. Towards automated analysis of connectomes: the configurable pipeline for the analysis of connectomes (c-pac). Front. Neuroinform. 42, 10–3389 (2013).
Glatard, T. et al. Boutiques: a flexible framework to integrate command-line applications in computing platforms. Gigascience 7, giy016 (2018).
Amstutz, P. et al. Common Workflow Language, draft 3. https://doi.org/10.6084/m9.figshare.3115156.v1 (2016).
Halchenko, Y. et al. DataLad: distributed system for joint management of code, data, and their relationship. J. Open Source Softw. 6, 3262 (2021).
Kiar, G. et al. Verificarlo/fuzzy: Fuzzy v0.5.0. https://doi.org/10.5281/zenodo.5027708 (2021).
Chatelain, Y. et al. A numerical variability approach to results stability tests and its application to neuroimaging. arXiv [physics.med-ph] (2023).
Gau, R. et al. COBIDAS checklist. Preprint at https://doi.org/10.17605/OSF.IO/ANVQY (2019).
Salkind, N. J. (Ed.) Encyclopedia of research design. SAGE Publications, Inc., https://doi.org/10.4135/9781412961288 (2010).
Gueorguieva, R. & Krystal, J. H. Move over ANOVA: progress in analyzing repeated-measures data and its reflection in papers published in the archives of general psychiatry. Arch. Gen. Psychiatry 61, 310–317 (2004).
Simmons, J., Nelson, L. & Simonsohn, U. Pre‐registration: why and how. J. Consum. Psychol. 31, 151–162 (2021).
Baribault, B. et al. Metastudies for robust tests of theory. Proc. Natl. Acad. Sci. USA 115, 2607–2612 (2018).
Brandmaier, A. M. et al. Assessing reliability in neuroimaging research through intra-class effect decomposition (ICED). Elife 7, e35718 (2018).
Kiar, G., Chatelain, Y., Salari, A., Evans, A. C. & Glatard, T. Data augmentation through Monte Carlo arithmetic leads to more generalizable classification in connectomics. Neurons Behav. Data Theory https://doi.org/10.51628/001c.28328 (2021).
Bellec, P., Rosa-Neto, P., Benali, H. & Evans, A. C. Multi-level bootstrap analysis of stable clusters (BASC) in resting-state fMRI. NeuroImage 47, S123 (2009).
Nikolaidis, A. et al. Bagging improves reproducibility of functional parcellation of the human brain. Neuroimage 214, 116678 (2020).
Doshi, J. et al. MUSE: MUlti-atlas region Segmentation utilizing Ensembles of registration algorithms and parameters, and locally optimal atlas selection. Neuroimage 127, 186–195 (2016).
Acknowledgements
Research reported in this publication was supported in part by the National Institute Of Mental Health under Award Numbers RF1MH130859 and R01MH120482 (G.K., M.P.M.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We would also like to offer our thanks to Emily Hector and Ted Satterthwaite, for their insightful reflections and feedback on this work throughout its development.
Author information
Authors and Affiliations
Contributions
G.K. was responsible for the conceptualization, funding acquisition, visualization, and primary writing of this work. J.A.M., T.X., J.T.V., and T.G. were responsible for contributing to the conceptualization and writing/editing of this work. M.P.M. was responsible for contributing to the conceptualization, funding acquisition, and writing/editing of this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dylan Nielson, Aaron Peikert and Dustin Scheinost for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kiar, G., Mumford, J.A., Xu, T. et al. Why experimental variation in neuroimaging should be embraced. Nat Commun 15, 9411 (2024). https://doi.org/10.1038/s41467-024-53743-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53743-y