Abstract
Hybrids between species exhibit plastic genomic architectures that could foster or slow down their adaptation. When challenged to evolve in an environment containing a UV mimetic drug, yeast hybrids have reduced adaptation rates compared to parents. We find that hybrids and their parents converge onto similar molecular mechanisms of adaptation by mutations in pleiotropic transcription factors, but at a different pace. After 100 generations, mutations in these genes tend to be homozygous in the parents but heterozygous in the hybrids. We hypothesize that a lower rate of loss of heterozygosity (LOH) in hybrids could limit fitness gain. Using genome editing, we first demonstrate that mutations display incomplete dominance, requiring homozygosity to show full impact and to entirely circumvent Haldane’s sieve, which favors the fixation of dominant mutations. Second, tracking mutations in earlier generations confirmed a different rate of LOH in hybrids. Together, these findings show that Haldane’s sieve slows down adaptation in hybrids, revealing an intrinsic constraint of hybrid genomic architecture that can limit the role of hybridization in adaptive evolution.
Similar content being viewed by others
Introduction
Hybridization rapidly generates novel genotypes that can also lead to new and sometimes extreme phenotypes1,2,3,4,5. As a result, hybrids may thrive and often outcompete their parents6,7,8. Empirical and theoretical work has shown that hybridization can promote rapid evolution2,3,9,10,11,12,13,14,15, including during evolutionary rescue, species diversification, and adaptive radiations2,11,12,16,17,18. The generation of adaptive diversity through hybridization has long been successfully employed in biotechnology for fermentation19,20,21 and in agriculture for crop improvement22,23,24.
The combination of divergent genomes in the same organism can also lead to genomic instability25,26. From microorganisms27,28 to multicellular eukaryotes such as plants29 and vertebrates30, genomic instability has been frequently observed in hybrids. While the causes of instability are not always clear, evidence points to the alteration of the molecular pathways and components responsible for genome stability themselves. For instance, cell cycle checkpoints and DNA repair pathways observed to be altered in hybrids31,32 can lead to inaccurate chromosome segregation33,34, resulting in changes in ploidy35,36,37,38, aneuploidies39,40,41, and elevated mutation rates42,43,44,45.
In spite of these potential negative consequences, genomic instability can also paradoxically enhance F1 hybrid traits. One example is the restoration of meiotic recombination through genome homogenization, which contributes to rescuing hybrid fertility46. In this scenario, recurrent loss of heterozygosity (LOH) facilitates crossing over by providing segments of sequence similarity, thereby promoting proper chromosome segregation. Similarly, whole-genome duplication can also restore fertility in interspecific hybrids47. However, the enhancement of these hybrid traits is typically observed under stable laboratory conditions. Some environmental conditions and stresses could further enhance genome instability, so what appears as a feature that enhances adaptability in benign conditions could become a liability in extreme ones. Motivated by this question, in a previous study48, we produced F1 hybrids between the budding yeasts Saccharomyces cerevisiae and Saccharomyces paradoxus to measure their adaptive potential in an environment containing a drug that increases genomic instability by mimicking UV radiation49,50. These two species diverged 5–10 million years ago51,52,53,54,55, live in similar ecological niches in nature56, and carry signs of introgression in their mitochondrial and nuclear genomes with adaptive potential in some cases46,57,58,59,60. Replicated experimental evolution across 100 generations revealed that hybrids showed smaller fitness gains than their parents in conditions mimicking UV radiation48.
Here, we investigate the genomic and genetic bases of differential adaptation rates in hybrids by testing two non-mutually exclusive hypotheses: (1) Hybrid adaptation to UV radiation is hampered by genomic instability, and (2) Hybrids do not have access to the same adaptive mutations as the parental species. We use Whole Genome Sequencing (WGS) of 270 ancestral and evolved clones of hybrid and parental genotypic backgrounds to: (1) Investigate major genomic changes in copy number, such as ploidy changes, aneuploidies, or LOH, to determine if they are more prevalent in hybrid genomes, and (2) Identify potentially differential de novo mutations in hybrids compared to parents. We functionally validate seven putative adaptive mutations with site-directed mutagenesis in both S. cerevisiae and S. paradoxus haploid backgrounds and by CRISPR-Cas9 genome editing. We examine if there are differences in the population dynamics of adaptive mutations between hybrids and parents by tracking genotype frequency over time through the historical resurrection of frozen stocks. We find that neither of our two starting hypotheses is supported. Rather, adaptation to UV mimetic conditions often proceeds through two types of mutations, nucleotide substitution and LOH, the second occurring more slowly in the hybrids, and thus slowing down their rate of adaptation.
Results
Adaptive dynamics of hybrids and parents in UV mimetic conditions
We previously evolved 90 populations (30 F1 hybrid replicates and 30 replicates of each of the parental S. cerevisiae and S. paradoxus genotypic backgrounds) for 100 generations in the presence of a DNA damaging agent, the UV mimetic drug 4-nitroquinoline 1-oxide (4-NQO), and in a control condition (rich medium) (Fig. 1a)48. We found that hybrids achieved a lower rate of adaptation compared to the parental genotypic backgrounds, with a lower average gain of fitness over the course of evolution (Fig. 1b).
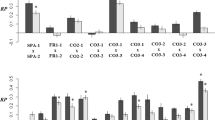
a Experimental evolution in UV mimetic and control conditions was performed using hybrid and parents for about 100 generations48 (n = 30 replicated lines for each genotypic background: S. cerevisiae, S. paradoxus or hybrid). Lines from isolated clones derived from each population were analyzed by WGS and flow cytometry (n = 269, refer to the Methods section for detailed information on the sample size). b Statistical asymptotic modeling of the growth rate of the different genotypic backgrounds as a function of the number of generations under UV mimetic conditions (n = 30 replicated lines for each genotypic background: S. cerevisiae, S. paradoxus or hybrid). In Bautista et al.48, we fitted three regression models: one linear and two non-linear (asymptotic and logarithmic) to assess the change in growth rate over time. The Akaike Information Criterion (AIC) indicated that the asymptotic model provided a better fit, followed by the logarithmic and linear models, respectively. The asymptotic model reveals that the adaptation rate was significantly lower for the hybrid than for both parents (p = 7.558e-15 and p = 4.307e-05, respectively, for S. cerevisiae and S. paradoxus in Bautista et al.48). Two-sided t-tests for pairwise comparisons were performed to compare the slope parameter of the model between genotypic backgrounds. c Growth rates of isolated clones derived from each population evolved in UV mimetic conditions correlate with growth rates of their populations of origin. Two-sided Spearman’s rank correlation coefficients (rs) and associated p-values are shown (n = 30 for S. cerevisiae, n = 30 for S. paradoxus and n = 28 for hybrid. Refer to the Methods section for detailed information on the sample size). Exact p-values are: p < 2.2e-16 for all genotypic backgrounds, p = 0.044 for S. cerevisiae, p = 0.018 for S. paradoxus and p = 0.00029 for the hybrid. The shaded band represents the 95% confidence interval.
To compare the number and type of genetic changes between evolved hybrid and parental populations, we measured DNA content by flow cytometry and sequenced the genomes of 269 isolated clones (refer to the Methods section for detailed information on the sample size) derived from the evolved populations and their ancestors (Fig. 1a and Supplementary Data 1) (genome-wide mean read depth of 100X, Supplementary Fig. 1). In parallel, we measured the growth of these isolated clones in control and UV mimetic conditions (Supplementary Fig. 2). We confirmed our previous finding that evolved hybrids showed significantly lower growth rate than parental replicated populations when evolved under UV mimetic conditions (Supplementary Fig. 2, all p < 0.001). The growth rates of isolated clones from populations and that of their populations of origin were strongly correlated (Fig. 1c: rs = 0.75, p < 0.0001), supporting that the growth of isolated clones is representative of the general fitness dynamics observed in the previous evolution experiment48. From now on, we will refer to lines when discussing observations on these isolated clones derived from their populations of origin, and we use growth rate as a proxy for fitness. While we acknowledge that growth rate measurements do not directly capture all fitness components, our findings demonstrate an improvement in growth rate, which provides evidence of selection acting on this trait.
Major genomic changes in copy number reveal increased genomic instability in UV mimetic conditions
The majority of lines remained diploid during evolution. Ploidy changes were generally not more frequent under UV mimetic conditions (Supplementary Figs. 3 and 4). Likewise, we did not observe a larger number of whole-genome level ploidy changes in hybrids in UV mimetic compared to control conditions, suggesting that ploidy changes do not account for the observed reduction in hybrid adaptation rate (Supplementary Fig. 3). Conversely, evolution under UV mimetic conditions did result in a larger number of lines with aneuploidies in hybrid and both parental genotypic backgrounds when compared to control conditions (Fig. 2a). While only the increase in aneuploidy rate observed in S. paradoxus was statistically significant (Supplementary Fig. 5) compared to control conditions, the presence of aneuploidies predominantly in UV mimetic-treated populations suggests that UV mimetic exposure may compromise genome stability across all genotypes. However, the number of aneuploidies did not correlate with the increases in growth rate observed in any group (Supplementary Fig. 6, rs = − 0.18, p > 0.05; rs = − 0.22, p > 0.05 and, rs = − 0.27, p > 0.05; for S. cerevisiae, S. paradoxus, and hybrid respectively). Hybrids also did not show a larger number of aneuploidies compared to the parents, making it unlikely to explain their reduced adaptability (Fig. 2a).
a Relative read depth per chromosome and evolved line in UV mimetic and control conditions. Colored squares show the relative read depth between evolved and ancestor lines (log2 fold change). Values with an increase in 30%-fold change represent gains in DNA content (gradient towards orange), and values with a decrease in 30%-fold change represent losses in DNA content (gradient towards purple). Rows represent individual genomes; columns show chromosomes. Hybrid lines display two chromosome sets due to the concatenated hybrid genome mapping. Homologous chromosomes are adjacent: S. cerevisiae (Scer) left, S. paradoxus (Spar) right (refer to the Methods section for detailed information). Top panels: lines evolved under control conditions. Bottom panels: lines evolved under UV mimetic conditions. Lines are ranked according to their increase in growth rate (%) under UV mimetic conditions (with top lines indicating greater increase). This ranking order is preserved under control conditions, enabling direct comparison between conditions (for control: S. cerevisiae n = 30, S. paradoxus n = 30, hybrid n = 30, and for UV mimetic conditions: S. cerevisiae n = 29, S. paradoxus n = 30 and hybrid n = 26, refer to the Methods section for detailed information on the sample size). b Example of variation in relative read depth across chromosomes for a single hybrid line (13) evolved in UV mimetic, highlighting the detection of t-LOH through simultaneous increase and decrease in relative read depth (deviations of 30% from the genome-wide median relative read depth). c Illustration of t-LOHs in hybrid genomes. d Percentage of hybrid lines carrying t-LOHs on each chromosome region. We excluded triploids from this analysis (hybrid n = 23, refer to the Methods section for detailed information on the sample size). e Number of t-LOHs per hybrid line after evolution in control and UV mimetic conditions. A two-sided Wilcoxon test p-value is shown (The exact p-value is p = 1.205e-09). We excluded triploids from this analysis (for control: hybrid n = 25 and for UV mimetic conditions: hybrid n = 23, refer to the Methods section for detailed information on the sample size).
The analysis of the relative read depth revealed a particular type of alteration, for instance, in chromosomes XII and XV (Fig. 2a). Intriguingly, gains and losses occurred simultaneously in the homologous chromosomes of S. cerevisiae and S. paradoxus, i.e., when a hybrid lost a portion of the S. cerevisiae chromosome, it simultaneously gained a portion of the homologous S. paradoxus one. This prompted us to map the chromosome relative read depth to determine if there were patterns in the distribution of species-specific chromosome gains and losses. The observed changes affected large regions of chromosomes (Fig. 2b). These patterns can be caused by the reciprocal exchange of segments of homologous chromosomes in diploids during mitosis, resulting in LOH (Fig. 2c)61,62. We identified LOH as regions where there were simultaneous increases and decreases in relative read depth in homologous chromosomes along a size threshold of 20 kb (Supplementary Fig. 7). We detected two types of LOH, interstitial LOHs (i-LOHs), which often originate from gene conversion involving short exchanges, and terminal LOHs (t-LOHs), typically resulting from mitotic crossovers that encompass larger regions61,63. We identified some large i-LOHs (for example, in line 17 on chromosome XIV, Supplementary Fig. 7) but we focused on t-LOHs, which were more frequent.
The number of t-LOH events in hybrid genomes was significantly positively correlated with chromosome size (Supplementary Fig. 8a, rs = 0.81, p < 0.05). The pattern of t-LOHs showed similarities across hybrid lines, with specific regions enriched on chromosomes XII and XV (Fig. 2d). These regions have been previously identified to be susceptible to t-LOHs64. Certain t-LOHs were found to cluster around positions enriched with repetitive loci, such as the rDNA locus on chromosome XII61,65 or the gene STE4, another common t-LOH target on chromosome XV66. We found a much larger number of hybrid lines with t-LOHs when evolved under UV mimetic conditions (96% of the lines) compared to control conditions (8% of the lines) (Fig. 2e, p < 0.0001). Only one line evolved under UV mimetic conditions did not show any t-LOH (4%). This suggests that either DNA damage triggers t-LOH, thus enhancing its occurrence rate, or that t-LOH is particularly advantageous under these conditions and selected for. Advantageous LOHs have been associated with fitness increase in both the brewery and natural settings, generally under stressful environmental conditions64,67. However, LOHs in themselves may not cause variation in the rates of adaptation among hybrid lines. We indeed did not find a significant correlation between the increase in growth rate (%) and the number of t-LOH events (Supplementary Fig. 8b, rs = 0.33, p > 0.05).
In summary, although we cannot examine t-LOHs in parental genomes due to their overall homozygosity, we found that aneuploidy frequencies are similar across parental species and hybrids, suggesting that overall genome instability is not specifically increased in hybrids. There is, therefore, no support for our initial hypothesis that increased genomic instability hampers adaptation in hybrids.
Parents and hybrids have parallel access to adaptive de novo mutations in the same genes
Lower rates of adaptation to UV mimetic conditions may also be explained by hybrids not having access to the same adaptive mutations as the parental lines, as per our alternative hypothesis. For instance, some mutations could have strong background-dependent effects. We investigated whether hybrids show single nucleotide polymorphisms (SNPs) in similar genes or gene functions, focusing on missense variants, as their potential impacts are easier to interpret, and data is more robust in this category. We found a median of 20 missense variants per line in S. cerevisiae, 12 in S. paradoxus, and 42 in the hybrids (Supplementary Fig. 9), with a comparable trend in control conditions. A gene ontology (GO) analysis revealed differences in terms of the biological functions affected by mutations but also some similarities (Supplementary Fig. 10). Enrichments are not significant, but interestingly, the most enriched GO terms for S. cerevisiae included double-strand break repair via sister chromatid exchange (GO:1990414), potentially aiding in coping with 4-NQO-induced DNA breaks, and the regulation of cell differentiation (GO:0045595). For S. paradoxus, the most enriched terms included ER-associated misfolded protein catabolic processes (GO:0071712) and endocytic vesicles (GO:0030139), potentially disrupting the invagination of extracellular substances, such as the 4-NQO drug (UV mimetic conditions). In the hybrid, the most enriched terms comprised trehalose metabolic process (GO:0005991) and ABC-type transporter activity (GO:0140359), comprising efflux pumps involved in expelling xenobiotic compounds, such as 4-NQO.
We found that mutations occurred in parallel in two specific genes across all three genotypic backgrounds (Fig. 3a), suggesting that the same molecular changes provide strong selective advantages in both hybrids and parents. The most frequent parallel changes involved the pleiotropic drug response genes PDR1 and YRR1 (Fig. 3a). PDR1 and YRR1 encode zinc finger transcription factors regulating multidrug and stress responses68,69,70. Among their targets is PDR5, a well-characterized yeast efflux pump that actively transports toxic compounds out of the cell71,72. Three S. cerevisiae lines, eight S. paradoxus lines, and eight hybrid lines had non-synonymous mutations in PDR1, while each genotypic background had at least four lines with YRR1 mutations (Fig. 3a). This is a surprising level of parallelism, given that drug resistance mutations have been shown to sometimes be genotype-specific73 and that S. cerevisiae and S. paradoxus diverged 10 million years ago.
a Count of the most recurrently mutated genes in parents and hybrids. b Pdr1 structure modeled with AlphaFold 2164,165 featuring amino acid changes identified in this study (green), previously reported changes (blue), and those found both in this study and previously reported (red)74,75,76,77,78. c Mutations and chromosomal changes co-occur, affecting growth rate increases (%) across all genotypic backgrounds. A schematic of the chromosomal changes for each individual line is shown (n = 3 for S. cerevisiae, n = 8 for S. paradoxus and, n = 8 for hybrid). Matching colors indicate identical amino acid changes across genotypic backgrounds. The hybrid line 24 displays varying colors for the PDR1 mutations, as they occur at different amino acid positions. d Increase in growth rate (% change in growth rate between initial and final time points) as a function of the number of PDR1 mutated copies (sample sizes from left to right: n = 3, n = 1, n = 4, n = 3, n = 1, n = 2, n = 1 and n = 4 mutations). Boxes show the interquartile range (IQR), with the median as the center line, quartiles as box limits, and whiskers extending to 1.5 times the IQR. p-value from the Kruskal-Wallis test (above; exact p-value is p = 0.04194) and letters representing adjusted p-values comparison after false discovery rate (FDR) correction (above boxplots) are shown. e Increase in growth rate (% change in growth rate between initial and final time points) as a function of the number of PDR1 mutated copies in all genotypic backgrounds (top) or only in hybrid (bottom). Two-sided Spearman’s rank coefficient (rs) and p-value are shown (n = 3 for S. cerevisiae, n = 8 for S. paradoxus, and n = 8 for hybrid). The exact p-values are: p = 0.015 (top) and p = 0.02 (bottom). The shaded band represents the 95% confidence interval. Hybrid line 28 is denoted with (*) due to its complex pattern: whole-chromosome LOH and increase in ploidy. PDR1 mutated copies are shown in yellow in (c) and (d).
From here on, we will focus on the analysis of PDR1 since it was the most often mutated gene. Non-synonymous mutations in PDR1 were not randomly located along the gene but instead occurred in particular clusters that overlapped among all three genotypic backgrounds (Supplementary Fig. 11) when mapped on the protein tertiary structure (Fig. 3b). We found identical or similar substitutions as those from previous studies on S. cerevisiae exposed to different drugs, ethanol, and antifungals74,75,76,77,78 (Fig. 3b). Localization on the protein structure revealed the presence of a cluster also found in other species, for instance, in the pathogen Nakaseomyces glabratus (Supplementary Fig. 12a, b), for which antifungal resistance often arises from mutations in PDR179,80,81,82,83,84,85. Parallelism even occurred at the level of amino acid changes across genotypic backgrounds (Fig. 3c). For instance, a mutation at amino acid 308/307 (corresponding to the respective S. cerevisiae and S. paradoxus positions) independently occurred up to four times. This specific amino acid change has also been observed in other studies75, conferring resistance to the same UV mimetic drug we employed (4-NQO)74.
The identification of shared adaptation hotspots and shared amino acid changes suggests a common adaptive landscape across genotypic backgrounds, indicating that specific regions within PDR1 and YRR1 harbor similar potential for adaptive mutations to occur in both hybrid and parents.
Fitness increases with the copy number of PDR1 adaptive alleles
The occurrence of mutations in PDR1 across all three (hybrid and two parental) genotypic backgrounds raises an intriguing question: Why do these mutations not confer similar adaptive benefits to hybrids as they do to the parental species? We observed that some lines carrying PDR1 mutations showed particularly high increases in growth rate (%) under UV mimetic conditions within their own genotype (Fig. 3c - S. cerevisiae lines: 2, 25 and 27; S. paradoxus lines: 29, 22, 28, 27, 9 and 24; hybrid lines: 30, 27, 13, 17 and 28). A closer analysis revealed that all these lines share a common characteristic, namely the presence of multiple mutated copies of PDR1 (Fig. 3c), with the exception of two lines (line 17 of hybrid and line 9 of S. paradoxus), which are discussed in detail below. Although mutations are expected to initially be heterozygous, we observed a diversity of genotypes at the PDR1 locus. While some lines have undergone LOH that made the mutations homozygous, others show changes in ploidy, resulting in an increased number of chromosomes harboring PDR1 mutations (Fig. 3c). The regions of LOH (t-LOH) found in the hybrids on chromosome VII, which result in the homozygosity of PDR1 mutations (lines 13 and 30 in Supplementary Fig. 7), confer an adaptive advantage as homozygous lines have higher growth rates than heterozygous ones (see the comparison among lines 30 and 13 versus lines 24, 9 and 2 in Fig. 3c). An exception occurred in hybrid line 17, where heterozygosity leads to increased growth rates (%) comparable to that of homozygous lines, potentially attributed to additional mutations in genes like PDR18, which codes for a xenobiotic transporter86. Another exception is observed in line 9 of S. paradoxus (Fig. 3c). While being heterozygous for the PDR1 mutation, this line’s high improvement in growth rate is likely due to an additional mutation in the YRR1 transcription factor, resulting in simultaneous mutations of both transcription factors, potentially increasing their activity.
Although not exhaustively investigated here, our observations in YRR1 also suggest that LOHs confer a selective advantage at this locus. Mutations in the hybrid lines consistently coincide with LOH around a specific region of chromosome XV (hybrid lines 5, 6, 8, and 15, in Supplementary Fig. 7). This suggests that mutations in this gene were followed by LOH, rendering the mutation homozygous and potentially conferring a fitness advantage. We could not compare the potential benefits of homozygous vs. heterozygous YRR1 mutations as all hybrids were homozygous for this gene. But given that YRR1 encodes a transcriptional regulator, like PDR1, we suspect that the adaptive mutations displayed incomplete dominance and their full adaptive impact is only achieved following LOH. We expand on this below for PDR1.
Fitness increased as a function of the number of PDR1 mutated copies in all three genotypic backgrounds (Fig. 3d, e top panel rs = 0.55, p < 0.05). This effect was particularly strong in hybrids (Fig. 3e bottom panel, rs = 0.79, p < 0.05). Among S. paradoxus and hybrid lines, the two with the largest observed increase in growth rate within their respective genotypic contexts (134% in S. paradoxus line 24 and 123% in hybrid line 28) each carried three copies of the PDR1 mutant alleles (Fig. 3c, d). Hybrid line 28 showed a complex pattern in which a whole-chromosome LOH was combined with an increase in ploidy (Fig. 3c*, Supplementary Figs. 13 and 7).
Taken together, our findings demonstrate an association between genomic changes leading to the amplification of mutated PDR1 copies and increases in growth rate (%) in UV mimetic conditions. Hybrids with PDR1 mutations do show improvements in growth rate (%) but to a smaller extent than both parental lines. This advantage could derive from the fact that parental genomes achieve higher levels of homozygosity for these mutations (Fig. 3c): The LOH frequency in hybrids containing PDR1 mutations was 37.5% (3/8 hybrid lines had a homozygous PDR1 mutation), whereas it was 100% across the parental S. cerevisiae lines (3/3) and 62.5% across S. paradoxus lines (5/8), making PDR1 mutations more visible to selection in parental lines. Thus, the limiting factor may not be the rate and type of mutation but their limited allelic amplification in hybrids (Fig. 4a).
a Schematic of the hypotheses tested. A single-parent species is depicted for simplicity. b Growth rate in UV mimetic conditions (4 µM of 4-NQO) of the four PDR1 zygosity types constructed by CRISPR-Cas9 (sample sizes from left to right: n = 24, n = 24, n = 24, n = 20 and n = 28 mutants). p-value from the two-sided ANOVA test (above; exact p-value is p < 2e-16) is shown; letters represent Tukey post hoc pairwise groups (above boxplots). Hemizygous is represented in light gray. c Temporal dynamics of populations 13 (hybrid) and 29 (S. paradoxus) evolved in UV conditions. Relative frequencies of the three zygosities over time are shown. The relative frequency is displayed from one time point before the first detection of the PDR1 mutation to up to two-time points after. The panel shows a representative example; see Supplementary Fig. 17 for a comprehensive analysis (n = 5–10 isolated clones per time point). d Temporal dynamics of the evolutionary history across the three zygosities (n = 2 S. cerevisiae populations, n = 4 S. paradoxus populations, and n = 3 hybrid populations, each population has n = 5–10 isolated clones per time point). The dark-colored interval represents the number of generations in which the first homozygous mutation was detected. e Growth rate in UV mimetic conditions (4 µM of 4-NQO) of hybrid zygosities (heterozygous PDR1, homozygous PDR1 or no mutation in PDR1) per population over time (from isolated clones of the experimental evolution frozen population stocks). For populations 13, 28, and 30, the numbers of isolated clones (n) were respectively: heterozygous (12, 40, 19), homozygous (6, 6, 11), and non-mutated (29, 48, 93). p-value from a two-sided ANOVA test is shown (exact p-values: p = 2.94e-08 for population 13, p < 2e-16 for population 28, and p = 0.000316 for population 30). Boxes in (b) and (e) show the interquartile range (IQR), with the median as the center line, quartiles as box limits, and whiskers extending to 1.5 times the IQR.
The challenge of attaining homozygosity accounts for the reduced adaptive potential of hybrids
The likelihood and rate of a mutation becoming fixed in a population is shaped by the architecture of the trait under selection, including allelic dominance, and the strength of selection. Therefore, not only does an adaptive mutation need to occur, but it also needs to be in a proper genotype for the individuals to fully benefit from it87. Dominant mutations are more likely to become fixed than recessive ones, a principle referred to as Haldane’s sieve88,89,90. This poses a problem to non-obligate sexual species and in systems that only rarely reproduce sexually, like many unicellular microbes such as fungi. In asexual populations, recessive or incompletely dominant beneficial mutations can bypass Haldane’s sieve by achieving homozygosity through LOH91, although completely recessive alleles would be invisible to selection and could thus be lost before an LOH occurs. The rate of LOH is not uniform among genotypes, and it has been shown to be lower in heterozygous ones92. In hybrids, which are highly heterozygous, LOH rates are even more limited (Fig. 4a)41,44. Once a beneficial mutation occurs in the parental genomes, high rates of mitotic recombination can rapidly lead to LOH at the site where the mutation occurred and make a mutation homozygous. This would occur at a lower rate in hybrids, slowing down the rate of adaptation.
In order to test if Haldane’s sieve slows down adaptation in hybrids, we need to assess these conditions: (1) The initial PDR1 mutation is recessive or incompletely dominant, requiring homozygosity to unmask its full fitness benefits. We thus predict that homozygous PDR1 mutants will display higher fitness than heterozygous PDR1 mutants; (2) LOH occurs at a slower pace in hybrid genomes41,44,92.
We first validated the adaptiveness of specific PDR1 mutations. Site-directed mutagenesis was used to introduce seven candidate mutations (G280R, G280S, M308I, and G1042W for S. cerevisiae; G279R, G279S, and G281V for S. paradoxus) on a plasmid carrying either S. cerevisiae or S. paradoxus PDR1 gene. After introducing these plasmids into an S. cerevisiae strain (BY4741) deleted for PDR1 (pdr1Δ), we found that the mutations conferred significantly higher growth rates in the presence of the UV mimetic drug compared to the wild-type (WT) (Supplementary Fig. 14a). These mutations also conferred benefits in the parental backgrounds with slight variations in the extent of effects (Supplementary Fig. 15). To further confirm the effects derived from the transcription factor activity of PDR1, we measured expression of the downstream drug efflux pump Pdr571,72. We fused Pdr5 to a Green Fluorescent Protein (mEGFP) and measured its expression in the same strain (BY4741) containing PDR1 mutations on a plasmid. The Pdr5-mEGFP strain exhibited higher fluorescence levels when carrying a plasmid containing specific PDR1 mutations, compared to wild-type (WT) (Supplementary Fig. 14b). Consistent with this observation, we found that the same PDR1 mutations also lead to resistance to several drugs other than 4-NQO (Supplementary Fig. 12c).
To assess the dominance of PDR1 mutations in a reference genomic background, we generated homozygous and heterozygous diploid mutants using CRISPR-Cas9 genome editing and by mating S. cerevisiae lab strains BY4741 and BY4742. We focused on the M308I mutation because it showed the most significant increase in growth rate (%) and activation of the Pdr5 efflux pump (Supplementary Fig. 14). We found that in the homozygous state, this mutation conferred a higher fitness advantage than in the heterozygous state (Fig. 4b), confirming its incomplete dominance. The hemizygote also showed a higher growth rate than the heterozygote, suggesting that the mutated allele confers higher benefits in the absence of the WT allele. We tested this assumption in S. paradoxus, specifically in line 27, which contained the same mutation in PDR1 (M307I) (Fig. 3c). We observed a similar pattern of incomplete dominance when comparing the various genotypes (see Supplementary Fig. 16). Since we lacked an evolved S. cerevisiae line with this specific mutation (Fig. 3c), we crossed haploid strains of the constructed PDR1 mutant (M308I) in the BY4741/2 genetic backgrounds and of the line 27 of S. paradoxus to generate both homozygous and heterozygous hybrids for the PDR1 locus. The pattern remained consistent among the hybrids, where homozygous mutants exhibited enhanced growth rates compared to their heterozygous counterparts (Supplementary Fig. 16). Taken together, our findings confirm the first key assumption that (1) the initial PDR1 mutation exhibits incomplete dominance and requires homozygosity to fully contribute to adaptation.
We archived populations regularly over the course of experimental evolution48, so we could revive the frozen stocks and isolate some clones to determine the timing of appearance of the various PDR1 mutations. We revived ~ 350 clones from nine hybrid and parental populations that harbor homozygous PDR1 mutations at the end of the experimental evolution (based on WGS data) and sequenced the PDR1 locus using amplicon Sanger sequencing. We identified three types of zygosity at the PDR1 locus at intermediate time points (Fig. 4c): heterozygous, homozygous WT, and homozygous mutant. As expected for de novo mutations in diploids, mutations were consistently first detected as heterozygous (Fig. 4d), but in some cases, in the parental populations, the LOH was so rapid that we also detected homozygous mutants (Fig. 4c left).
This trend aligns with the increase in growth rate (%) recorded during experimental evolution in their populations of origin but also when analyzing specific isolated clones48 (Supplementary Figs. 17 and 18, respectively). Mutations also became homozygous later (Fig. 4d), but with the important difference that the parents become homozygous at a higher frequency than the hybrids. This pattern persisted across all experimental populations: Homozygous genotypes appeared quickly after the initial PDR1 mutations occurred, and spread rapidly in the parental populations, whereas in the hybrids, even by generation 95, a high proportion of heterozygotes were still observed (Fig. 4d, see Supplementary Fig. 17 for detailed analysis). Supporting this trend, we saw in some of the parents (population 2 and 25 of S. cerevisiae, population 27 and 29 of S. paradoxus) that the emergence of homozygous and heterozygous mutations coincided, indicating that the mechanism of adaptation through LOH can operate quickly (Fig. 4c, d). The relative frequency of each mutation in each population through time further shows that the proportion of homozygous mutants was much lower in hybrids compared to parental species (Fig. 4d). To verify that the low homozygote frequency in hybrids was not due to reduced homozygote fitness in the specific hybrid genotypic background, we compared hybrid growth rates across generations and populations and confirmed that homozygotes were fitter than heterozygotes and homozygotes WT (Fig. 4e and Supplementary Fig. 18 for growth rate measurements across generations). These findings validate our final assumption that (2) LOH occurs at a slower pace in hybrids.
Discussion
Hybridization is a recurring phenomenon in nature that has captured the interest of scientists for decades1,9,14,93,94,95,96 in fundamental fields but also in applied research such as in agriculture and medical microbiology23,24,27,97,98. The adaptive and non-adaptive roles of hybridization have been studied extensively8,46,99,100,101,102,103,104,105,106,107. However, investigations on the negative consequences have mostly focused on reproductive isolation108,109 and less on the potential reduction in adaptive rates of hybrids. We previously evolved hybrids of S. cerevisiae and S. paradoxus species during 100 generations under stress that mimics UV radiation and observed a reduced adaptive potential of hybrids48. We used this system to investigate what could reduce the hybrid rate of adaptation.
We first observed that hybrid genomes exhibit similar rates of aneuploidy compared to parental species. Indeed, our findings reveal markedly lower levels of aneuploidy compared to prior investigations conducted in contexts such as experimental evolution under selective pressures110, industrial or natural environments26. This notable variation in aneuploidy prevalence among hybrids hints at the potential influence of environment-dependent genetic effects, as demonstrated in a previous study using the same set of strains41.
Our results also reveal that hybrid and parental species have access to the same adaptive changes in key genes. In principle, adaptation can, therefore, occur through the same mechanisms and at the same rate. We examined the cases of mutations that impacted a transcriptional factor involved in drug resistance because they displayed strong parallelism. The mutations displayed incomplete dominance, thus partially avoiding Haldane’s sieve - the bias against the establishment of recessive beneficial mutations. However, achieving homozygosity through LOH allowed them to fully circumvent this filter88,89,90. Experiments involving the evolution of heterozygous yeast populations have also shown that LOH frequently unmasks beneficial recessive alleles, which can confer significant fitness advantages91,111,112,113. Because LOH depends on recombination and recombination depends on sequence identity41,44,92, LOH occurs at a slower rate in hybrids, ultimately contributing to slowing down hybrid adaptation. This contrasts with de novo mutations, which accumulate in yeast hybrid genotypes at rates that are similar to those observed in the parental species114. Similarly, the absence of a higher frequency of aneuploidy in hybrids cannot compensate for the delayed onset of LOH in increasing mutated gene copy numbers. While this reduced rate of LOH represents one potential mechanism influencing hybrid adaptation, it is likely not the sole driver. Additional exploration may uncover alternative genetic interactions contributing to the lower adaptation of hybrid lines.
T-LOH events frequently originate in conserved regions because they contain similar sequences between chromosome pairs, facilitating the initiation of recombination. In the case of PDR1, since it is located near the centromere, recombination might be initiated in centromeric sequences that are highly conserved among related budding yeast species115,116. Other elements, such as highly repetitive retrotransposons, can also trigger the initiation of mitotic recombination117. The observed high frequency of YRR1 mutations associated with t-LOHs in all examined hybrid lines (4/4) suggests a potentially elevated rate of LOH for this gene. This could be attributed to increased sequence homology between recombination-triggering regions. For instance, the t-LOHs we observed involving YRR1 may have originated around a Ty1 long-terminal repeat (LTR) (YORCdelta11). It is thus possible that not all loci, where adaptive mutations could occur, are limited to contributing to adaptation because of the low LOH rate in hybrids. Although not explored here, t-LOH in hybrids could also limit adaptation in other ways than simply occurring at a lower rate around the adaptive mutations. Since mitotic recombination often extends along the entire length of a chromosome arm61,63 in heterozygous genomes62, an LOH that renders a beneficial mutation homozygous could also bring along other molecular changes or combination of changes that would negatively impact fitness66. For example, Feri et al.118 demonstrated in Candida albicans the presence of several recessive lethal alleles that restrict LOH events. We also cannot disregard the importance of the hybrid background itself and the impact of interactions between divergent genomes, such as epistasis and hybrid incompatibilities119,120,121,122 which could modulate the fitness consequences of LOH.
Our findings contribute to the understanding of the genomic factors shaping asexual microbes. Such hybrids often evolve during domestication, for instance, many beer yeasts are among-species hybrids123,124 and these hybrids are known to be largely sterile, i.e., to not have access to sexual reproduction125. Even fungal pathogens evolve through recurrent hybridization events27,98,126,127,128,129,130, acquire antifungal resistance and adapt to new hosts with de novo mutations and LOH131,132,133,134,135. It has been shown for instance that to confer full resistance to antifungals, a mutation in a transcriptional regulator needed to be followed by an LOH in the fungal pathogen Candida albicans136. Antimicrobial resistance has also been shown to depend on an LOH event in S. cerevisiae that is required again to render an LOF mutation homozygous137,138. Understanding which conditions could slow down the rate of LOH, such as heterozygosity along the chromosome as we exemplify here, or the physical linkage to potentially deleterious mutations118, is therefore key to understanding evolution in an applied context such as antimicrobial resistance. Similarly, cancer cells reproduce somatically and usually evolve by LOH, as some of the mutations associated with tumor progression involve the loss of functional dominant alleles of tumor suppressors139,140. The phenomenon we uncovered here, whereby some genotypes experience lower rates of LOH, thus has also consequences that extend beyond the study of hybrids.
Methods
Experimental crosses and previous experimental evolution
The S. cerevisiae and S. paradoxus strains used were described in Bautista et al.48 and were derived from the natural strains LL13_054 and MSH-604 isolated in North American forests13,47 (Supplementary Data 2). To prevent mating type switching in haploids, the HO locus was replaced with resistance cassettes (HPHNT1 for Hygromycin B resistance and NATMX4 for Nourseothricin resistance)141. A total of 90 experimental strains were constructed by crossing haploid strains (30 S. cerevisiae, 30 S. paradoxus, and 30 hybrids), so that each starting parental and hybrid diploid population is the result of an independent mating event as described in Bautista et al.48. The 90 populations were evolved for 100 generations (Fig. 1a) as described in Bautista et al.48. Briefly, we used as environments a non-DNA damaging growth condition we call control (YPD medium consists of 10 g/L yeast extract, 20 g/L tryptone, and 20 g/L glucose) and a DNA damaging growth condition, which was identical except for the addition of a UV mimetic molecule49 4-Nitroquinoline 1-oxide (4-NQO) (Sigma–Aldrich, cat. no. N8141, batch #WXBC3635V, Canada).
Growth assays on individual clones
Growth assays were conducted on individual clones, which were used for genome sequencing and isolated from the glycerol stocks from experimental evolution48 (Supplementary Data 1). Ancestor strains (n = 90, once per clone) as well as the clones evolved in YPD (n = 90, once per clone) and in YPD + 4-NQO (n = 90, once per clone) were pre-cultured in 1 mL of YPD in 96 deep-well plates and incubated for 24 h at 25 °C. Subsequently, 20 µL of these pre-cultures were grown in 96-well flat-bottomed culture plates in 180 µL of medium (YPD or YPD + 4 µM of 4-NQO), resulting in an initial OD595 of approximately 0.1. A transfer cycle was performed at 24 h after approximately ~ 5 generations in rich conditions. Each culture was diluted approximately 30-fold by transferring 6 µL of grown culture into 194 µL of fresh medium to initiate a new round of growth at an OD595 starting at about 0.03 (this second cycle is used to calculate growth parameters, such as the growth rate). Incubation at 25 °C was performed directly in three temperature-controlled spectrophotometers (Infinite® 200 PRO, Tecan, Reading, UK) that read the OD595 at intervals of 15 min throughout the two cycles performed. All samples were randomized across plates, temperature-controlled spectrophotometers, and days.
DNA extraction, library construction, and Whole Genome Sequencing (WGS)
We obtained whole-genome sequences of 270 individual clones derived from the 270 experimental populations48 (Supplementary Data 1). We extracted genomic DNA from overnight YPD cultures derived from each clone according to the manufacturer’s instructions (MasterPure™ Yeast DNA Purification Kit, Biosearch Technologies - Lucigen, Wisconsin, USA) and purified on Axygen™ AxyPrep Magnetic PCR Clean-up SPRI beads (Axygen Inc, New York, USA). Five DNA pooled libraries were prepared using RIPTIDE™ High Throughput rapid DNA library prep in 96-well plate format (iGenomX, South San Francisco, USA)142. The quality of the libraries was verified using an Agilent BioAnalyzer 2100 electrophoresis system (Genomic Analysis Platform of the Institute of Integrative Biology and Systems of Université Laval, Québec, Canada). Pooled libraries were sequenced using paired-end 150 bp reads on different lanes of an Illumina NovaSeq 6000 (Illumina, San Diego, USA) at the Genome Quebec Innovation Center (Montréal, Canada). We excluded sequences of the S. cerevisiae line 16 evolved in UV mimetic due to poor performance in the construction of the library (resulting in a final sample size of n = 269).
Flow cytometry analysis of ploidy
DNA content was measured by flow cytometry using the SYTOX™ green staining assay (Thermo Fisher, Waltham, USA)41,47. Diploid strains of the S. cerevisiae isolate LL13_054 were used as the diploid control. Given the different genetic backgrounds used in our study, the inference of polyploidy was done by comparing strains and looking for outliers, as most strains aligned with each other and with the diploid control. Whole-genome ploidy changes were expected to result in a shift toward higher DNA content values, as shown in Supplementary Fig. 19. For the hybrid strains, WGS analysis enabled us to determine the specific ploidy state as triploid, due to the gain of an entire genome copy from one of the parental genotypic backgrounds (Supplementary Figs. 7 and 19). The 270 individual clones derived from the 270 experimental populations (Supplementary Data 1) from Bautista et al.48 and used for WGS (n = 269) were thawed from glycerol stocks and grown on solid YPD omnitray plates (25 °C, 72 h). They were inoculated into 1 mL of YPD in 96 deepwell plates and incubated for 24 h at 25 °C. Cells were subsequently prepared for flow cytometry as in Gerstein et al.143. They were fixed in 70% ethanol and kept frozen at −20 °C for further analysis. RNA was degraded using 0.25 mg mL−1 RNase A during an overnight incubation at 37 °C. Cells were washed twice with sodium citrate (50 mM, pH 7) and stained with a SYTOX™ green concentration of 0.6 μM for 1 h at 25 °C in the dark. Cell concentration was adjusted in sodium citrate (50 mM, pH 7) to be less than 500 cells/μL. Five thousand cells from each of the 270 samples were analyzed in 96-well plates in a CytoFLEX Platform flow cytometer (Beckman Coulter, California, USA) at the Feldan Therapeutics facility (Québec, Canada). Cells were excited with the blue laser at 488 nm, and fluorescence was measured in a green fluorescence detection channel (525/40 nm). The distributions of the green fluorescence values were processed to find the two main density peaks, which correspond to the two cell populations in the G1 and G2 phases, respectively. Although two peaks (G1 and G2) are expected when lines are in the exponential phase, due to variations in growth, some lines may have reached the stationary phase (single peak in G0, which has the same DNA content as G1). Therefore, only those samples clearly exceeding the first peak of the diploid control were considered polyploids (Supplementary Fig. 19).
Quality assessment and read mapping of next-generation sequencing data
Raw reads from barcoded samples of the five libraries were demultiplexed using DemuxFastqs from fgbio tools144 v1.5.0 (Supplementary Data 3). Reads were trimmed using Trimmomatic145 v0.36 with parameters ILLUMINACLIP:Trimm_seqs.fa:6:20:10 and using Trimm_seqs.fa (Supplementary Data 4) as a list of adapter sequences used. To assess the quality of both pre- and post-trimming sequencing reads, we used FastQC v0.11.9146 and MultiQC v1.11147.
Reads from S. cerevisiae samples were mapped to the indexed reference genome of S. cerevisiae strain YPS128148, which in our study is named LL13_054, and S. paradoxus samples were mapped to the S. paradoxus SpB (named MSH-604) genome148,149. Reads from hybrid lines were mapped to a concatenated genome comprising the two respective parental genomes end to end. The BWA-MEM algorithm150 v0.7.17 was used for mapping. Mapped reads were processed by genome-sorting algorithms using samtools v1.8151, and quality was assessed by mapping coverage with goleft v0.2.2152. The average mean read depth across samples was about 100X (Supplementary Fig. 1). We used Picard tools v2.26.11153 for adding Read Groups with AddOrReplaceReadGroups, and we removed duplicate reads with MarkDuplicates with parameter REMOVE_DUPLICATES = true.
Analysis of sequencing read depth (aneuploidies and loss of heterozygosity)
Mean read depths over 1 kbp windows were obtained with BamStats04 from Jvarkit tools v2021.08.10154 and make windows from bedtools tools v2.30.0155. We first eliminated some sequences of the hybrid lines evolved in UV mimetic because the sequencing depth was low (lines 1 and 21) or because the content of one of the parental genomes was the majority (lines 10 and 25) (Supplementary Figs. 20–22). We removed these two lines (10 and 25) from the rest of the non-WGS analysis due to the uncertainty about their origin. We also verified the results of the Flow cytometry analysis of ploidy section and compared DNA content by measuring average read depth across genomes (Supplementary Fig. 19). We computed the median chromosome read depth and the median whole genome read depth for each line. In order to detect the number of gained or lost chromosomes, we divided each chromosome’s median read depth by the genome-wide median read depth. We standardized this value by the value of the corresponding ancestors to obtain the relative read depth (log2 fold change). Values with a chromosome median read depth higher than the genome-wide median represent gains in DNA content (gradient towards red in Fig. 2a, b, Supplementary Figs. 7 and 13a), and values with a median read depth lower than the genome-wide median represent losses in DNA content (gradient towards purple in Fig. 2a, b, Supplementary Figs. 7 and 13a) for each individual chromosome. We also computed the number of chromosomes with aneuploidies per line by considering an aneuploidy as a deviation (increase or decrease) of 30% with respect to the genome-wide median read depth as in Bendixsen et al.26. The positional coverage mapping of hybrid genomes unveiled terminal regions with pronounced increases in read depth in one parental chromosome copy and concurrent decreases in the other copy (Supplementary Fig. 7), revealing the presence of reciprocal crossovers between chromosomes. These tracts that extend to the telomeres and usually measure between 50–100 kb correspond to terminal-loss of heterozygosity (t-LOH) regions61,62,65. To quantitatively assess the number of t-LOH events in hybrid lines, we identified regions with simultaneous increases and decreases in read depth (deviation of 30% with respect to the genome-wide median read depth) in both chromosomal copies exceeding a size threshold of 20 kb. We excluded triploid strains from the t-LOH analyses due to the impossibility of detecting increases and decreases in read-depth. Therefore, the sample size for the evolved UV mimetic hybrids is n = 28 (excluding lines 10 and 25) for non-WGS analyses (such as in Flow cytometry analysis of ploidy or Growth assays on individual clones sections); n = 26 (excluding lines 1, 10, 21, and 25) when analyzing coverage patterns; n = 23 in UV mimetic conditions or n = 25 in control conditions when analyzing general LOH patterns (since we also exclude triploids which are displayed in Supplementary Fig. 19).
Functional analysis of de novo mutations
We eliminated some sequences of the hybrid lines evolved in UV mimetic because the sequencing depth was low (lines 1 and 21) or because the content of one of the parental genomes was the majority (lines 10 and 25) (Supplementary Figs. 20– 22). SNP calling was performed with Haplotype Caller (gatk-v4.1.4.1)156,157. Before generating the GVCFs (Genomic Variant Call Format files), we added an RG (read group) tag to individual BAM files. After SNP calling, genotyping of the GVCFs was performed with GenotypeGVCFs. For variant filtration, we applied standard hard filters with options: QUAL by depth (QD) < 2.0, mapping quality (MQ) < 40.0, Fisher’s exact tests of strand bias (FS) > 60.0, symmetric odds ratio test of strand bias (SOR) > 3.0, mapping quality rank sum test (MQRankSum) < − 12.5, rank sum test for site position within reads (ReadPosRankSum) < − 8.0, Genotype Quality (GQ) < 20, and Coverage (DP) < 3. We selected variants that passed the filters described above, and we excluded INDELS, focusing exclusively on substitutions (SNP) for subsequent analysis. We excluded the pre-existing genetic variation relative to the reference genome, by removing any variant that was already present in the ancestral strain for each evolved line (Supplementary Fig. 9). For annotation purposes, we used the S. cerevisiae genome assembly R64-1-1 (Saccharomyces cerevisiae S288c assembly from Saccharomyces Genome Database, INSDC Assembly GCA_000146045.2, Sep 2011) for S. cerevisiae genomes. Subsequently, we generated maps annotations with Liftoff (v1.6.3)158 of the S. paradoxus genome from the S. cerevisiae genome assembly R64-1-1 (Saccharomyces cerevisiae S288c assembly from Saccharomyces Genome Database, INSDC Assembly GCA_000146045.2, Sep 2011). We finally generated a combined annotated genome for hybrid analysis. The variants were ultimately annotated using Ensembl Variant Effect Predictor (VEP) v110159. We examined missense variants to perform Gene Ontology (GO) analysis using bioMart160, simplifyEnrichment161, and clusterProfiler162.
Validation of the adaptiveness of mutations
We used ChimeraX v1.5163 to visualize Pdr1 amino acid changes we found throughout the experimental evolution and other mutations found in the literature74,75,76,77,78 (Supplementary Data 5) on the AlphaFold 2164,165 generated structure for Pdr1 (AF-P12383-F1, Supplementary Data 6). Subsequently, we analyzed the Nakaseomyces glabratus Pdr1 protein by superimposing structures and amino acid changes79,82,83 across species (Supplementary Data 7; AF-B9VI40-F1, Supplementary Data 8). We next used a set of plasmids derived from the MoBy-ORF library, in which genes are controlled by their native promoter and terminator166, to express the PDR1 sequences from either S. cerevisiae (BY4741) or S. paradoxus (MSH-604). Candidate mutations (G280R, G280S, M308I, and G1042W, being G1041W in LL13_054 strain for the S. cerevisiae sequence, G279R, G279S, and G281V for the S. paradoxus sequence) were inserted by site-directed mutagenesis. As controls, we used the plasmid without the PDR1 gene (Empty) or the plasmid containing the Wild-Type (WT) PDR1 sequence. We introduced these plasmids following a modified lithium acetate transformation protocol167 in an S. cerevisiae lab strain BY4741, and in natural strains S. cerevisiae LL13_054 and S. paradoxus MSH-604 in both WT and pdr1Δ (previously constructed by replacing PDR1 locus with a NATMX4 module) backgrounds. Growth assays were performed following the same steps as described above in the Growth assays on individual clones section. We added extra conditions to the ones previously used (control and 4 µM of 4-NQO): 8 µM and 10 µM of 4-NQO (Supplementary Fig. 15). We also conducted an assay on the S. cerevisiae lab strain BY4741 pdr1Δ, containing the same mutations on the same plasmids, and exposed them to antifungal azoles. After adjusting cell density to an OD595 of 1, we made three serial dilutions 1/5 in 200 μL of water (40 μL of cells in 160 μL of water). We spotted 5 μL of each dilution on YPD + 0.2% DMSO (control), YPD + 16 μg/mL Fluconazole (FLC), YPD + 2 μg/mL Itraconazole (ITR) or YPD + 0.5 μg/mL Voriconazole (VRC) and incubated at 30 °C for 48 h. We assessed the expression of the downstream Pdr5p drug efflux pump by fusing Pdr5 to a Green Fluorescent Protein (mEGFP) (Pdr5-mEGFP) in a BY4741 pdr1Δ lab strain expressing PDR1 mutants from the above-described pMoBY plasmids. For a comprehensive list of strains, refer to Supplementary Data 2, and for a comprehensive list of oligonucleotide sequences, refer to Supplementary Data 9.
Incomplete-dominance assay
To evaluate the dominance of the PDR1 mutations, we created S. cerevisiae diploids harboring either homozygous or heterozygous M308I substitutions using CRISPR-Cas9 genome editing and mating BY4741 and BY4742 haploids (Supplementary Data 2). We replaced the PDR1 locus with NATMX4 (in BY4741) or HPHNT1 (in BY4742) modules specifically targeted by two different gRNA (guide RNA) using a modified protocol from Ryan et al.168. Yeast cells were transformed following a modified lithium acetate transformation protocol167 with a pCAS-NAT or pCAS-HPH plasmid (Addgene plasmid 6084747 modified by Després et al.169 and Lemieux et al.170 using the same approach as in Dionne et al.171) expressing both the gRNA (guide RNA) of interest (targeting NATMX4 or HPHNT1), the Streptococcus pyogenes Cas9172 and a donor DNA sequence featuring 40 bp homology arms flanking the PDR1 DNA sequence. The donor DNA sequences were: PDR1 sequence (WT), PDR1 sequence with the desired mutation (M308I), or PDR1 sequence with a stop codon instead of the first methionine (M1Stop) (Oligonucleotide sequences can be found in Supplementary Data 9). Finally, we mated BY4741 and BY4742 haploids with the desired mutations to create the following diploid strains: WT homozygous (pdr1Δ::PDR1/pdr1Δ::PDR1), M308I homozygous (pdr1Δ::PDR1(M308I)/pdr1Δ::PDR1(M308I)), heterozygous (pdr1Δ::PDR1/pdr1Δ::PDR1(M308I) or pdr1Δ::PDR1(M308I)/pdr1Δ::PDR1) or hemizygous (pdr1Δ::PDR1(M308I)/pdr1Δ::M1Stop) (refer to Supplementary Data 2 for a comprehensive list of strains). We selected diploid strains using double selection (Synthetic complete (SC) medium -lysine -methionine which is used as a standard selection method for BY background). SC medium consists of 1.74 g/L yeast nitrogen base without ammonium sulfate, 20 g/L glucose, and 5 g/L ammonium sulfate. Growth assays were performed following the same steps as described above in the Growth assays on individual clones section.
Incomplete-dominance assay on the three genotypic backgrounds
Since CRISPR-Cas9 editing was not feasible in the wild-type ancestral strains (S. paradoxus: MSH-604 or S. cerevisiae: LL13_054), we sporulated line 27 of Saccharomyces paradoxus (strain MSH-604) containing the M307I mutation (Fig. 3c) to obtain haploid cells for subsequent crosses with the BY4741 and BY4742 strains described above. For sporulation, diploid cells were grown overnight in 10 mL of selective medium, then transferred to fresh medium for a 3 h incubation with shaking at 30 °C. After centrifugation and washing with 20 mL of sterile water, the cells were resuspended in 10 mL of YEPA medium (10 g/L yeast extract, 20 g/L tryptone, and 20 g/L potassium acetate (CH3CO2K)) and incubated overnight at 30 °C with shaking. Subsequently, the cells were washed twice with 20 mL of sterile water and then resuspended in 25 mL of SPM medium (3 g/L potassium acetate (CH3CO2K) and 0.2 g/L raffinose) for 3–5 days to induce sporulation.
For yeast tetrad dissection, asci were incubated in 100 μL of zymolyase 20 T (200 μg/mL in water) for 20 min and then resuspended in 100 μL of sorbitol 1 M. Samples were spread onto Petri dishes containing YPD medium and dissected under a microscope before being incubated for 3 days at 30 °C. Finally, we mated BY4741, BY4742, S. paradoxus line 27 MATa and S. paradoxus line 27 MATα haploids with the desired mutations to create diploid strains: WT homozygous (pdr1Δ::PDR1 / pdr1Δ::PDR1), M308I/M307I homozygous (pdr1Δ::PDR1(M308I/M307I) / pdr1Δ::PDR1(M308I/M307I)) or heterozygous (pdr1Δ::PDR1 / pdr1Δ::PDR1(M308I/M307I) or pdr1Δ::PDR1(M308I/M307I) / pdr1Δ::PDR1) in the three backgrounds S. cerevisiae (diploid BY4743), S. paradoxus (MSH-604) and hybrid (refer to Supplementary Data 2 for a comprehensive list of strains). We then selected diploid strains using double selection (YPD + Nourseothricin + Hygromycin B) or by colony PCR screening173 to identify diploid colonies when double selection was not feasible (when crossing BY4741/2 strains with S. paradoxus haploid strains MSH-604). Growth assays were performed following the same steps as described above in the Growth assays on individual clones section.
Allele frequency dynamics on PDR1 mutants
Whole population samples were archived regularly during the evolution experiment48. From these samples, we isolated clones to estimate the timing of the appearance of the various PDR1 mutations and to track the dynamics between homozygous and heterozygous zygosities in the population. We revived approximately 350 clones from nine populations (Supplementary Data 1) and sequenced the PDR1 locus using amplicon Sanger sequencing (Oligonucleotide sequences can be found in Supplementary Data 10). We designed specific primers for each mutation (Oligonucleotide sequences can be found in Supplementary Data 9) and used standard colony PCR to amplify those regions. We performed an exhaustive zygosity analysis across time points (each time point represents five generations) to quantify the ratio of homozygous to heterozygous variants. We sequenced clones starting from the first detection of PDR1 mutations until the homozygous frequency exceeded 80%, or if this did not occur, we sequenced until time point 19 (corresponding to 95 generations). Growth assays in isolated hybrid clones were performed following the same steps as described above in the Growth assays on individual clones section.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The sequencing data generated in this study have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject PRJNA1045261. The demultiplexing process details are available at Zenodo174 (https://doi.org/10.5281/zenodo.10389557) and Supplementary Data 3. The processed data are available at Zenodo174 (https://doi.org/10.5281/zenodo.10389557). PDB-formatted files of the AF2-generated models are Supplementary Data 6 and 8, and can also be found at Zenodo174 (https://doi.org/10.5281/zenodo.10389557). Strains are available upon request.
Code availability
Data and scripts for figure generation can be found at Zenodo174 (https://doi.org/10.5281/zenodo.10389557) and on GitHub (https://github.com/cbautistaro/Bautista2024_LOH_project_NatureCommunications). Data were analyzed using bash and R version 4.2.0.
References
Lewontin, R. C. & Birch, L. C. Hybridization as a source of variation for adaptation to new environments. Evolution 20, 315–336 (1966).
Ballerini, E. S. Rapidly Evolving Genes and Genetic Systems 256–265 (2012).
Kagawa, K. & Takimoto, G. Hybridization can promote adaptive radiation by means of transgressive segregation. Ecol. Lett. 21, 264–274 (2018).
Steensels, J., Gallone, B. & Verstrepen, K. J. Interspecific hybridization as a driver of fungal evolution and adaptation. Nat. Rev. Microbiol. 19, 485–500 (2021).
Porretta, D. & Canestrelli, D. The ecological importance of hybridization. Trends Ecol. Evol. 38, 1097–1108 (2023).
François, J. M. et al. A quasi-domesticate relic hybrid population of saccharomyces cerevisiae× S. paradoxus adapted to olive brine. Front. Genet. 10, 449 (2019).
Martin-Roy, R., Nygård, E., Nouhaud, P. & Kulmuni, J. Differences in thermal tolerance between parental species could fuel thermal adaptation in hybrid wood ants. Am. Nat. 198, 278–294 (2021).
Stelkens, R. & Bendixsen, D. P. The evolutionary and ecological potential of yeast hybrids. Curr. Opin. Genet. Dev. 76, 101958 (2022).
Grant, B. R. & Grant, P. R. High survival of Darwin’s finch hybrids: Effects of beak morphology and diets. Ecology 77, 500–509 (1996).
Seehausen, O. Hybridization and adaptive radiation. Trends Ecol. Evol. 19, 198–207 (2004).
Schumer, M., Rosenthal, G. G. & Andolfatto, P. How common is homoploid hybrid speciation? Evolution 68, 1553–1560 (2014).
Stelkens, R., Brockhurst, M. A., Hurst, G. D. D. & Greig, D. Hybridization facilitates evolutionary rescue. Evol. Appl. 7, 1209–1217 (2014).
Leducq, J.-B. et al. Speciation driven by hybridization and chromosomal plasticity in a wild yeast. Nat. Microbiol. 1, 15003 (2016).
Mitchell, N., Owens, G. L., Hovick, S. M., Rieseberg, L. H. & Whitney, K. D. Hybridization speeds adaptive evolution in an eight-year field experiment. Sci. Rep. 9, 6746 (2019).
Kulmuni, J., Wiley, B. & Otto, S. P. On the fast track: hybrids adapt more rapidly than parental populations in a novel environment. Evol. Lett. 8, 128–136 (2023).
Rieseberg, L. H. et al. Major ecological transitions in wild sunflowers facilitated by hybridization. Science 301, 1211–1216 (2003).
Meier, J. I. et al. Ancient hybridization fuels rapid cichlid fish adaptive radiations. Nat. Commun. 8, 14363 (2017).
Vedder, D. et al. Hybridization may aid evolutionary rescue of an endangered East African passerine. Evol. Appl. 15, 1177–1188 (2022).
Peris, D. et al. Hybridization and adaptive evolution of diverse Saccharomyces species for cellulosic biofuel production. Biotechnol. Biofuels 10, 78 (2017).
Sipiczki, M. Interspecies hybridisation and genome chimerisation in saccharomyces: Combining of gene pools of species and its biotechnological perspectives. Front. Microbiol. 9, 3071 (2018).
Lopandic, K. Saccharomyces interspecies hybrids as model organisms for studying yeast adaptation to stressful environments. Yeast 35, 21–38 (2018).
Crow, J. F. 90 years ago: the beginning of hybrid maize. Genetics 148, 923–928 (1998).
Campbell, L. G., Snow, A. A. & Ridley, C. E. Weed evolution after crop gene introgression: greater survival and fecundity of hybrids in a new environment. Ecol. Lett. 9, 1198–1209 (2006).
Matsuoka, Y. Evolution of polyploid triticum wheats under cultivation: the role of domestication, natural hybridization and allopolyploid speciation in their diversification. Plant Cell Physiol. 52, 750–764 (2011).
Baack, E. J. & Rieseberg, L. H. A genomic view of introgression and hybrid speciation. Curr. Opin. Genet. Dev. 17, 513–518 (2007).
Bendixsen, D. P., Peris, D. & Stelkens, R. Patterns of genomic instability in interspecific yeast hybrids with diverse ancestries. Front. Fungal Biol. 2, 742894 (2021).
Pryszcz, L. P. et al. Correction: The Genomic Aftermath of Hybridization in the Opportunistic Pathogen Candida metapsilosis. PLoS Genet. 12, e1006202 (2016).
Morard, M. et al. Genomic instability in an interspecific hybrid of the genus Saccharomyces: a matter of adaptability. Microb. Genom. 6, https://doi.org/10.1099/mgen.0.000448 (2020).
Anisimova, I. N. et al. Genomic instability in sunflower interspecific hybrids. Genetika 45, 934–943 (2009).
Metcalfe, C. J. et al. Genomic instability within centromeres of interspecific marsupial hybrids. Genetics 177, 2507–2517 (2007).
Aguilera, A. & Gómez-González, B. Genome instability: a mechanistic view of its causes and consequences. Nat. Rev. Genet. 9, 204–217 (2008).
Herbst, R. H. et al. Heterosis as a consequence of regulatory incompatibility. BMC Biol. 15, 38 (2017).
Dion-Côté, A.-M. & Barbash, D. A. Beyond speciation genes: an overview of genome stability in evolution and speciation. Curr. Opin. Genet. Dev. 47, 17–23 (2017).
Maheshwari, S. & Barbash, D. A. The genetics of hybrid incompatibilities. Annu. Rev. Genet. 45, 331–355 (2011).
Alves, M. J., Coelho, M. M. & Collares-Pereira, M. J. Evolution in action through hybridisation and polyploidy in an Iberian freshwater fish: a genetic review. Genetica 111, 375–385 (2001).
Soltis, P. S. & Soltis, D. E. The role of hybridization in plant speciation. Annu. Rev. Plant Biol. 60, 561–588 (2009).
Marcet-Houben, M. & Gabaldón, T. Beyond the whole-genome duplication: Phylogenetic evidence for an ancient interspecies hybridization in the baker’s yeast lineage. PLoS Biol. 13, e1002220 (2015).
Alix, K., Gérard, P. R., Schwarzacher, T. & Heslop-Harrison, J. S. P. Polyploidy and interspecific hybridization: partners for adaptation, speciation and evolution in plants. Ann. Bot. 120, 183–194 (2017).
Chheda, H. R., De Wet, J. M. J. & Harlan, J. R. Aneuploidy inBothriochloaHybrids. Caryologia 14, 205–217 (1961).
Gilchrist, C. & Stelkens, R. Aneuploidy in yeast: Segregation error or adaptation mechanism? Yeast 36, 525–539 (2019).
Marsit, S., Hénault, M., Charron, G., Fijarczyk, A. & Landry, C. R. The neutral rate of whole-genome duplication varies among yeast species and their hybrids. Nat. Commun. 12, 3126 (2021).
Yang, S. et al. Parent–progeny sequencing indicates higher mutation rates in heterozygotes. Nature 523, 463–467 (2015).
Xie, Z. et al. Mutation rate analysis via parent–progeny sequencing of the perennial peach. I. A low rate in woody perennials and a higher mutagenicity in hybrids. Proc. R. Soc. B Biol. Sci. 283, 20161016 (2016).
Tattini, L. et al. Accurate tracking of the mutational landscape of diploid hybrid genomes. Mol. Biol. Evol. 36, 2861–2877 (2019).
Krasovec, M. The spontaneous mutation rate of Drosophila pseudoobscura. G3 11, (2021).
D’Angiolo, M. et al. A yeast living ancestor reveals the origin of genomic introgressions. Nature 587, 420–425 (2020).
Charron, G., Marsit, S., Hénault, M., Martin, H. & Landry, C. R. Spontaneous whole-genome duplication restores fertility in interspecific hybrids. Nat. Commun. 10, 4126 (2019).
Bautista, C., Marsit, S. & Landry, C. R. Interspecific hybrids show a reduced adaptive potential under DNA damaging conditions. Evol. Appl. 14, 758–769 (2021).
Felkner, I. C. & Kadlubar, F. Parallel between ultraviolet light and 4-nitroquinoline-1-oxide sensitivity in Bacillus subtilis. J. Bacteriol. 96, 1448–1449 (1968).
Wang, L.-E. et al. 4-nitroquinoline-1-oxide-induced mutagen sensitivity and risk of cutaneous melanoma: a case-control analysis. Melanoma Res. 26, 181–187 (2016).
Liti, G., Barton, D. B. H. & Louis, E. J. Sequence diversity, reproductive isolation and species concepts in Saccharomyces. Genetics 174, 839–850 (2006).
Kellis, M., Patterson, N., Endrizzi, M., Birren, B. & Lander, E. S. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423, 241–254 (2003).
Tirosh, I., Reikhav, S., Levy, A. A. & Barkai, N. A yeast hybrid provides insight into the evolution of gene expression regulation. Science 324, 659–662 (2009).
Borneman, A. R. & Pretorius, I. S. Genomic insights into the Saccharomyces sensu stricto complex. Genetics 199, 281–291 (2015).
Shen, X.-X. et al. Tempo and mode of genome evolution in the budding yeast subphylum. Cell 175, 1533–1545.e20 (2018).
Sniegowski, P. D., Dombrowski, P. G. & Fingerman, E. Saccharomyces cerevisiae and Saccharomyces paradoxus coexist in a natural woodland site in North America and display different levels of reproductive isolation from European conspecifics. FEMS Yeast Res. 1, 299–306 (2002).
Peris, D. et al. Mitochondrial introgression suggests extensive ancestral hybridization events among Saccharomyces species. Mol. Phylogenet. Evol. 108, 49–60 (2017).
Barbosa, R. et al. Evidence of natural hybridization in brazilian wild lineages of saccharomyces cerevisiae. Genome Biol. Evol. 8, 317–329 (2016).
Peter, J. et al. Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556, 339–344 (2018).
Gallegos-Casillas, P. et al. Yeast diversity in open agave fermentations across Mexico. Yeast 41, 35–51 (2023).
Sui, Y. et al. Genome-wide mapping of spontaneous genetic alterations in diploid yeast cells. Proc. Natl. Acad. Sci. USA 117, 28191–28200 (2020).
Dutta, A., Dutreux, F. & Schacherer, J. Loss of heterozygosity results in rapid but variable genome homogenization across yeast genetic backgrounds. Elife 10, https://doi.org/10.7554/elife.70339 (2021).
Yim, E., O’Connell, K. E., St Charles, J. & Petes, T. D. High-resolution mapping of two types of spontaneous mitotic gene conversion events in Saccharomyces cerevisiae. Genetics 198, 181–192 (2014).
Large, C. R. L. et al. Genomic stability and adaptation of beer brewing yeasts during serial repitching in the brewery. Preprint at bioRxiv https://doi.org/10.1101/2020.06.26.166157 (2020).
Smukowski Heil, C. Loss of heterozygosity and its importance in evolution. J. Mol. Evol. 91, 369–377 (2023).
Fisher, K. J., Vignogna, R. C. & Lang, G. I. Overdominant mutations restrict adaptive loss of heterozygosity at linked loci. Genome Biol. Evol. 13, https://doi.org/10.1093/gbe/evab181 (2021).
James, T. Y. et al. Adaptation by loss of heterozygosity in saccharomyces cerevisiae clones under divergent selection. Genetics 213, 665–683 (2019).
Balzi, E., Chen, W., Ulaszewski, S., Capieaux, E. & Goffeau, A. The multidrug resistance gene PDR1 from Saccharomyces cerevisiae. J. Biol. Chem. 262, 16871–16879 (1987).
Le Crom, S. et al. New insights into the pleiotropic drug resistance network from genome-wide characterization of the YRR1 transcription factor regulation system. Mol. Cell. Biol. 22, 2642–2649 (2002).
Ottilie, S. et al. Adaptive laboratory evolution in S. cerevisiae highlights role of transcription factors in fungal xenobiotic resistance. Commun. Biol. 5, 128 (2022).
Leppert, G. et al. Cloning by gene amplification of two loci conferring multiple drug resistance in Saccharomyces. Genetics 125, 13–20 (1990).
Harris, A. et al. Structure and efflux mechanism of the yeast pleiotropic drug resistance transporter Pdr5. Nat. Commun. 12, 5254 (2021).
Gerstein, A. C. & Berman, J. Candida albicans genetic background influences mean and heterogeneity of drug responses and genome stability during evolution in fluconazole. mSphere 5, https://doi.org/10.1128/msphere.00480-20 (2020).
Carvajal, E., van den Hazel, H. B., Cybularz-Kolaczkowska, A., Balzi, E. & Goffeau, A. Molecular and phenotypic characterization of yeast PDR1 mutants that show hyperactive transcription of various ABC multidrug transporter genes. Mol. Gen. Genet. 256, 406–415 (1997).
Anderson, J. B., Sirjusingh, C. & Ricker, N. Haploidy, diploidy and evolution of antifungal drug resistance in Saccharomyces cerevisiae. Genetics 168, 1915–1923 (2004).
Mutlu, N., Garipler, G., Akdoğan, E. & Dunn, C. D. Activation of the pleiotropic drug resistance pathway can promote mitochondrial DNA retention by fusion-defective mitochondria in Saccharomyces cerevisiae. G3 4, 1247–1258 (2014).
Xia, H. et al. Evolutionary and reverse engineering in Saccharomyces cerevisiae reveals a Pdr1p mutation-dependent mechanism for 2-phenylethanol tolerance. Microb. Cell Fact. 21, 269 (2022).
Taylor, M. B. et al. yEvo: experimental evolution in high school classrooms selects for novel mutations that impact clotrimazole resistance in Saccharomyces cerevisiae. G3 12, jkac246 (2022).
Tsai, H.-F., Krol, A. A., Sarti, K. E. & Bennett, J. E. Candida glabrata PDR1, a transcriptional regulator of a pleiotropic drug resistance network, mediates azole resistance in clinical isolates and petite mutants. Antimicrob. Agents Chemother. 50, 1384–1392 (2006).
Ferrari, S. et al. Gain of function mutations in CgPDR1 of Candida glabrata not only mediate antifungal resistance but also enhance virulence. PLoS Pathog. 5, e1000268 (2009).
Vale-Silva, L., Ischer, F., Leibundgut-Landmann, S. & Sanglard, D. Gain-of-function mutations in PDR1, a regulator of antifungal drug resistance in Candida glabrata, control adherence to host cells. Infect. Immun. 81, 1709–1720 (2013).
Whaley, S. G. et al. Jjj1 Is a negative regulator of Pdr1-mediated fluconazole resistance in Candida glabrata. mSphere 3, https://doi.org/10.1128/msphere.00466-17 (2018).
Khakhina, S., Simonicova, L. & Moye-Rowley, W. S. Positive autoregulation and repression of transactivation are key regulatory features of the Candida glabrata Pdr1 transcription factor. Mol. Microbiol. 107, 747–764 (2018).
Tian, Y. et al. A gain-of-function mutation in PDR1 of Candida glabrata decreases EPA1 expression and attenuates adherence to epithelial cells through enhancing recruitment of the Mediator subunit Gal11A. Microbiol. Res. 239, 126519 (2020).
Ksiezopolska, E. et al. Narrow mutational signatures drive acquisition of multidrug resistance in the fungal pathogen Candida glabrata. Curr. Biol. 31, 5314–5326 (2021).
Cabrito, T. R., Teixeira, M. C., Singh, A., Prasad, R. & Sá-Correia, I. The yeast ABC transporter Pdr18 (ORF YNR070w) controls plasma membrane sterol composition, playing a role in multidrug resistance. Biochem. J. 440, 195–202 (2011).
Lang, G. I., Botstein, D. & Desai, M. M. Genetic variation and the fate of beneficial mutations in asexual populations. Genetics 188, 647–661 (2011).
Haldane, J. B. S. A mathematical theory of natural and artificial selection, part V: Selection and mutation. Math. Proc. Camb. Philos. Soc. 23, 838–844 (1927).
Orr, H. A. & Betancourt, A. J. Haldane’s sieve and adaptation from the standing genetic variation. Genetics 157, 875–884 (2001).
Ronfort, J. & Glemin, S. Mating system, Haldane’s sieve, and the domestication process. Evolution 67, 1518–1526 (2013).
Gerstein, A. C., Kuzmin, A. & Otto, S. P. Loss-of-heterozygosity facilitates passage through Haldane’s sieve for Saccharomyces cerevisiae undergoing adaptation. Nat. Commun. 5, 3819 (2014).
Tutaj, H., Pirog, A., Tomala, K. & Korona, R. Genome-scale patterns in the loss of heterozygosity incidence in Saccharomyces cerevisiae. Genetics 221, https://doi.org/10.1093/genetics/iyac032 (2022).
MacDougal, D. T. Hybridization of wild plants. Bot. Gaz. 43, 45–58 (1906).
Heiser, C. B. Hybridization between the sunflower species helianthus annuus and H. petiolaris. Evolution 1, 249–262 (1947).
Morales, L. & Dujon, B. Evolutionary role of interspecies hybridization and genetic exchanges in yeasts. Microbiol. Mol. Biol. Rev. 76, 721–739 (2012).
Wei, X. et al. The lingering effects of Neanderthal introgression on human complex traits. Elife 12, e80757 (2023).
Pryszcz, L. P., Németh, T., Gácser, A. & Gabaldón, T. Genome comparison of Candida orthopsilosis clinical strains reveals the existence of hybrids between two distinct subspecies. Genome Biol. Evol. 6, 1069–1078 (2014).
Mixão, V. & Gabaldón, T. Hybridization and emergence of virulence in opportunistic human yeast pathogens. Yeast 35, 5–20 (2018).
Rieseberg, L. H., Archer, M. A. & Wayne, R. K. Transgressive segregation, adaptation and speciation. Heredity 83, 363–372 (1999).
Burton, T. L. & Husband, B. C. Fitness differences among diploids, tetraploids, and their triploid progeny in Chamerion angustifolium: mechanisms of inviability and implications for polyploid evolution. Evolution 54, 1182–1191 (2000).
Nolte, A. W. & David Sheets, H. Shape based assignment tests suggest transgressive phenotypes in natural sculpin hybrids (Teleostei, Scorpaeniformes, Cottidae). Front. Zool. 2, https://doi.org/10.1186/1742-9994-2-11 (2005).
Shahid, M., Han, S., Yoell, H. & Xu, J. Fitness distribution and transgressive segregation across 40 environments in a hybrid progeny population of the human-pathogenic yeast Cryptococcus neoformans. Genome 51, 272–281 (2008).
Zanders, S. E. et al. Genome rearrangements and pervasive meiotic drive cause hybrid infertility in fission yeast. Elife 3, e02630 (2014).
Shapira, R., Levy, T., Shaked, S., Fridman, E. & David, L. Extensive heterosis in growth of yeast hybrids is explained by a combination of genetic models. Heredity 113, 316–326 (2014).
Stelkens, R., Brockhurst, M. A., Hurst, G. D. D., Miller, E. L. & Greig, D. The effect of hybrid transgression on environmental tolerance in experimental yeast crosses. J. Evol. Biol. 27, 2507–2519 (2014).
Bernardes, J. P., Stelkens, R. B. & Greig, D. Heterosis in hybrids within and between yeast species. J. Evol. Biol. 30, 538–548 (2017).
Moran, B. M. et al. The genomic consequences of hybridization. Elife 10, https://doi.org/10.7554/eLife.69016 (2021).
Fierst, J. L. & Hansen, T. F. Genetic architecture and postzygotic reproductive isolation: evolution of Bateson–Dobzhansky–Muller incompatibilities in a polygenic model. Evolution 64, 675–693 (2010).
Schumer, M., Cui, R., Rosenthal, G. G. & Andolfatto, P. Reproductive isolation of hybrid populations driven by genetic incompatibilities. PLoS Genet. 11, e1005041 (2015).
Zhang, Z. et al. Recombining your way out of trouble: The genetic architecture of hybrid fitness under environmental stress. Mol. Biol. Evol. 37, 167–182 (2020).
Heil, C. S. S. et al. Loss of heterozygosity drives adaptation in hybrid yeast. Mol. Biol. Evol. 34, 1596–1612 (2017).
Marad, D. A., Buskirk, S. W. & Lang, G. I. Altered access to beneficial mutations slows adaptation and biases fixed mutations in diploids. Nat. Ecol. Evol. 2, 882–889 (2018).
Johnson, M. S. et al. Phenotypic and molecular evolution across 10,000 generations in laboratory budding yeast populations. Elife 10, https://doi.org/10.7554/elife.63910 (2021).
Fijarczyk, A., Hénault, M., Marsit, S., Charron, G. & Landry, C. R. Heterogeneous mutation rates and spectra in yeast hybrids. Genome Biol. Evol. 13, https://doi.org/10.1093/gbe/evab282 (2021).
Panzeri, L., Landonio, L., Stotz, A. & Philippsen, P. Role of conserved sequence elements in yeast centromere DNA. EMBO J. 4, 1867–1874 (1985).
Liebman, S. W., Symington, L. S. & Petes, T. D. Mitotic recombination within the centromere of a yeast chromosome. Science 241, 1074–1077 (1988).
Kim, J. M., Vanguri, S., Boeke, J. D., Gabriel, A. & Voytas, D. F. Transposable elements and genome organization: a comprehensive survey of retrotransposons revealed by the complete Saccharomyces cerevisiae genome sequence. Genome Res. 8, 464–478 (1998).
Feri, A. et al. Analysis of repair mechanisms following an induced double-strand break uncovers recessive deleterious alleles in the candida albicans diploid genome. MBio. 7, e01109-16 (2016).
Chang, A. S. & Noor, M. A. F. Epistasis modifies the dominance of loci causing hybrid male sterility in the Drosophila pseudoobscura species group. Evolution 64, 253–260 (2010).
Beukeboom, L. W., Koevoets, T., Morales, H. E., Ferber, S. & van de Zande, L. Hybrid incompatibilities are affected by dominance and dosage in the haplodiploid wasp Nasonia. Front. Genet. 6, 140 (2015).
Ono, J., Gerstein, A. C. & Otto, S. P. Widespread genetic incompatibilities between first-step mutations during parallel adaptation of saccharomyces cerevisiae to a common environment. PLoS Biol. 15, e1002591 (2017).
Thompson, K. A. Experimental hybridization studies suggest that pleiotropic alleles commonly underlie adaptive divergence between natural populations. Am. Nat. 196, E16–E22 (2020).
Krogerus, K., Magalhães, F., Vidgren, V. & Gibson, B. Novel brewing yeast hybrids: creation and application. Appl. Microbiol. Biotechnol. 101, 65–78 (2017).
Masneuf, I., Hansen, J., Groth, C., Piskur, J. & Dubourdieu, D. New hybrids between Saccharomyces sensu stricto yeast species found among wine and cider production strains. Appl. Environ. Microbiol. 64, 3887–3892 (1998).
Fukuda, N. Crossbreeding of yeasts domesticated for fermentation: Infertility challenges. Int. J. Mol. Sci. 21, https://doi.org/10.3390/ijms21217985 (2020).
Samarasinghe, H. & Xu, J. Hybrids and hybridization in the cryptococcus neoformans and Cryptococcus gattii species complexes. Infect. Genet. Evol. 66, 245–255 (2018).
Mixão, V. & Gabaldón, T. Genomic evidence for a hybrid origin of the yeast opportunistic pathogen Candida albicans. BMC Biol. 18, 48 (2020).
Hessenauer, P. et al. Hybridization and introgression drive genome evolution of Dutch elm disease pathogens. Nat. Ecol. Evol. 4, 626–638 (2020).
Gabaldón, T. Hybridization and the origin of new yeast lineages. FEMS Yeast Res. 20, https://doi.org/10.1093/femsyr/foaa040 (2020).
Del Olmo, V. et al. Origin of fungal hybrids with pathogenic potential from warm seawater environments. Nat. Commun. 14, 6919 (2023).
Diogo, D., Bouchier, C., d’Enfert, C. & Bougnoux, M.-E. Loss of heterozygosity in commensal isolates of the asexual diploid yeast Candida albicans. Fungal Genet. Biol. 46, 159–168 (2009).
Forche, A. et al. Stress alters rates and types of loss of heterozygosity in Candida albicans. MBio 2, https://doi.org/10.1128/mbio.00129-11 (2011).
Bennett, R. J., Forche, A. & Berman, J. Rapid mechanisms for generating genome diversity: whole ploidy shifts, aneuploidy, and loss of heterozygosity. Cold Spring Harb. Perspect. Med. 4, https://doi.org/10.1101/cshperspect.a019604 (2014).
Ford, C. B. et al. The evolution of drug resistance in clinical isolates of Candida albicans. Elife 4, e00662 (2015).
Rokas, A. Evolution of the human pathogenic lifestyle in fungi. Nat. Microbiol. 7, 607–619 (2022).
Coste, A. et al. A mutation in Tac1p, a transcription factor regulating CDR1 and CDR2, is coupled with loss of heterozygosity at chromosome 5 to mediate antifungal resistance in Candida albicans. Genetics 172, 2139–2156 (2006).
Li, J. et al. Shared molecular targets confer resistance over short and long evolutionary timescales. Mol. Biol. Evol. 36, 691–708 (2019).
Vázquez-García, I. et al. Clonal heterogeneity influences the fate of new adaptive mutations. Cell Rep. 21, 732–744 (2017).
Ryland, G. L. et al. Loss of heterozygosity: what is it good for? BMC Med. Genomics 8, 1–12 (2015).
Zhang, X. & Sjöblom, T. Targeting loss of heterozygosity: A novel paradigm for cancer therapy. Pharmaceuticals 14, https://doi.org/10.3390/ph14010057 (2021).
Güldener, U., Heck, S., Fielder, T., Beinhauer, J. & Hegemann, J. H. A new efficient gene disruption cassette for repeated use in budding yeast. Nucleic Acids Res. 24, 2519–2524 (1996).
Siddique, A. et al. RiptideTM high throughput NGS library prep for genotyping in populations. J. Biomol. Tech. 30, S35–S36 (2019).
Gerstein, A. C., Chun, H.-J. E., Grant, A. & Otto, S. P. Genomic convergence toward diploidy in Saccharomyces cerevisiae. PLoS Genet. 2, e145 (2006).
Fgbio. fgbio http://fulcrumgenomics.github.io/fgbio/.
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Andrews, S. et al. FastQC: a quality control tool for high throughput sequence data. (2010).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Yue, J.-X. et al. Contrasting evolutionary genome dynamics between domesticated and wild yeasts. Nat. Genet. 49, 913–924 (2017).
Eberlein, C. et al. Hybridization is a recurrent evolutionary stimulus in wild yeast speciation. Nat. Commun. 10, 923 (2019).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Pedersen, B. S., Collins, R. L., Talkowski, M. E. & Quinlan, A. R. Indexcov: fast coverage quality control for whole-genome sequencing. GigaScience 6, 1–6 (2017).
Lindenbaum, P. JVarkit: java-based utilities for Bioinformatics. figshare 10, m9 (2015).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2018).
Van der Auwera, G. A. & O’Connor, B. D. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra. (O’Reilly Media, Inc., 2020).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Smedley, D. et al. BioMart–biological queries made easy. BMC Genomics 10, 22 (2009).
Gu, Z. & Hübschmann, D. Simplifyenrichment: A bioconductor package for clustering and visualizing functional enrichment results. Genom. Proteom. Bioinforma. 21, 190–202 (2023).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Ho, C. H. et al. A molecular barcoded yeast ORF library enables mode-of-action analysis of bioactive compounds. Nat. Biotechnol. 27, 369–377 (2009).
Kaiser, C. Methods in yeast genetics: a Cold Spring Harbor Laboratory Course Manual 133–134 (1994).
Ryan, O. W., Poddar, S. & Cate, J. H. D. CRISPR-Cas9 Genome Engineering in Saccharomyces cerevisiae Cells. Cold Spring Harb. Protoc. 2016, https://doi.org/10.1101/pdb.prot086827 (2016).
Després, P. C. et al. Asymmetrical dose responses shape the evolutionary trade-off between antifungal resistance and nutrient use. Nat. Ecol. Evol. 6, 1501–1515 (2022).
Lemieux, P., Bradley, D., Dubé, A. K., Dionne, U. & Landry, C. R. Dissection of the role of a SH3 domain in the evolution of binding preference of paralogous proteins. Genetics, 226, https://doi.org/10.1093/genetics/iyad175 (2024).
Dionne, U. et al. Protein context shapes the specificity of SH3 domain-mediated interactions in vivo. Nat. Commun. 12, 1597 (2021).
Ryan, O. W. et al. Selection of chromosomal DNA libraries using a multiplex CRISPR system. Elife 3, https://doi.org/10.7554/elife.03703 (2014).
Huxley, C., Green, E. D. & Dunham, I. Rapid assessment of S. cerevisiae mating type by PCR. Trends Genet. 6, 236 (1990).
Bautista, C. Hybrid adaptation is hampered by Haldane’s sieve. Zenodo. https://doi.org/10.5281/zenodo.10389557 (2024).
Acknowledgements
We thank Nadia Aubin-Horth for feedback on the manuscript, Alexandre Dubé, and Cynthia Gagné-Thivierge for assistance in the laboratory work, Angel Cisneros, Lorena Ament, Alexandre Andreas Rego, Damien-Biot Pelletier, Adarsh Jay, and David Bradley for help with data analysis and members of the Landry Lab for helpful discussions. We thank the “Centre d’expertise et de services Génome Québec” and Feldan Thérapeutiques Québec, especially Vanessa Théberge and David Guay, for their services in sequencing and flow cytometry analysis, respectively. This work was funded by a Natural Sciences and Engineering Research Council of Canada (NSERC) grant to C.R.L. RGPIN-2020-04844. C.B. is supported by “la Caixa” Foundation (ID 100010434), under the agreement (LCF/BQ/AA16/11580051) and by the Fonds de recherche du Québec - Nature et technologies (FRQNT) (#274987) doctoral fellowships. M.U. was funded by the Human Frontier Science Program (HFSP) initiative “Science for Scientists” to support Ukrainian scientists. C.R.L. holds the Canada Research Chair in Cellular Systems and Synthetic Biology.
Author information
Authors and Affiliations
Contributions
C.B., I.G.A., and C.R.L. designed the research. CB performed the experiments collected and analyzed the data with the assistance of I.G.A., M.U., A.F., D.B., R.S., and C.RL. C.B. wrote the manuscript with the assistance of CRL. C.B., I.G.A., M.U., A.F., D.B., R.S., and C.R.L. edited the manuscript. C.R.L. was responsible for funding acquisition. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bautista, C., Gagnon-Arsenault, I., Utrobina, M. et al. Hybrid adaptation is hampered by Haldane’s sieve. Nat Commun 15, 10319 (2024). https://doi.org/10.1038/s41467-024-54105-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-54105-4
This article is cited by
-
Evolutionary causes and consequences of gene duplication
Nature Reviews Genetics (2026)