Abstract
Marine and terrestrial organisms often utilise EGF/EGF-like domains in wet adhesives, yet their roles in adhesion remain unclear. Here, we investigate the Barbatia virescense byssal system and uncover an oxidation-independent, reversible, and robust adhesion mechanism where EGF/EGF-like domain tandem repetitions in adhesive proteins bind robustly to GlcNAc-based biopolymer. EGF/EGF-like-domain-containing proteins demonstrate over three-fold superior underwater adhesion to chitosan compared to the well-known strongest wet-adhesive proteins, mefp-5, and suckerin, when adhering to mica in an surface forces apparatus-based measurement. Additionally, as the degree of acetylation of chitosan decreases from 20.0 to 5.34%, the underwater adhesion energy between mefp-2 and chitosan decreases from |Wad | ≈ 41.80 to 12.92 ± 0.40 mJm−2. This finding highlights the importance of GlcNAc over GlcN in binding with EGF to formulate effective underwater adhesives, expanding our understanding of underwater adhesion and supporting EGF’s functional role in biomedical wet adhesive interfaces, hydrogels, and chitosan applications.
Similar content being viewed by others
Introduction
Adherence to wet surfaces is a challenging adhesive science and technology problem. Numerous researchers have investigated the bioadhesives produced by fouling organisms with the aim of understanding how they successfully adhere to wet surfaces in aqueous habitats. These bioadhesives have significant potential for use in bionic-interface, medical-adhesive, and underwater-adhesive applications, as well as antifouling technologies designed to mitigate energy loss by preventing fouling1,2.
In the bioadhesives field, 3,4-dihydroxyphenylalanine (Dopa), a post-translationally modified amino acid, has been a research focal point for nearly half a century3,4. Mussel adhesive proteins were initially found to contain Dopa, which inspired research into Dopa-incorporated proteins, carbohydrates, synthetic polymers, and inorganic nanoparticles with the aim of replicating the remarkable ability of organisms to adhere to wet surfaces in nature5. However, the intrinsic reactivity of Dopa coupled with translational engineering challenges and its susceptibility to oxidation present significant hurdles when considering the practical use of Dopa-containing proteins in underwater adhesion applications. Underwater adhesion in nature undoubtedly relies on a complex interplay of mechanisms that transcend Dopa; therefore, developing a deeper understanding of these mechanisms is pivotal for progressing underwater adhesives and antifouling agents.
The presence of multiple consecutive epidermal growth factor (EGF)/EGF-like domains has been identified to be a common feature of marine adhesives; such domains were first observed in mussel-derived proteins 40 years ago and subsequently observed in the biological adhesives of many marine fouling organisms, including limpets6, sea urchins7 and seastars8 and sea anemones9. Mussel foot protein-2 (mefp-2), which mainly comprises tandem repetitions of EGF/EGF-like domains, is the most abundant protein (25–40 wt%) in the mussel adhesive plaques of blue mussels (Mytilus species), the representative model organism for underwater adhesion10; however, the contributions that EGF/EGF-like domains make to underwater adhesion have not yet been elucidated.
Here, we introduce an oxidation-free, reversible, and robust underwater adhesion mechanism that relies on binding between EGF/EGF-like domains and N-acetyl-d-glucosamine (GlcNAc) residues of polysaccharides. The Barbatia virescens ark clam used in this study is an understudied species that interfacially adheres to external surfaces and surrounding tissue through a singular giant plaque (Fig. 1) composed of a GlcNAc-based biopolymer, but it is not chitin/chitosan. Furthermore, interfacial proteins with EGF/EGF-like domain repetitions (bcbp-1 and bcbp-2) were characterised at the adhesive interface with GlcNAc, which exists in the form of chitin in the marine environment, constitutes the exoskeletons of crustaceans and marine invertebrates, and is one of the main carbohydrates involved in marine biofilms and adhesives; consequently, it may be the main fouling target of marine organisms. Based on the hypothesis that EGF/EGF-like domains bind to GlcNAc residues of polysaccharides and facilitate underwater adhesion, we examined the adhesion between GlcNAc-based polymers and EGF domains using chitosan and foot protein-2 (mefp-2) from Mytilus edulis the most abundant mussel-adhesives protein. While previous studies involving mefp-2 failed to deliver wet adhesion, we show that mefp-2 exhibits three-fold stronger underwater adhesion to chitosan (with its GlcNAc residues) than mefp-511 or suckerin12, placing it among the most adhesive proteins to be measured using a surface forces apparatus (SFA).
a The exterior of the ark clam and (b) the giant plaque. c Opened ark clam showing its full body. d Transverse cross-section of the Barbatia foot tissue stained with Masson’s trichrome (right) is an a coloured illustration (left). Stainings performed 3 times on different tissues blocks. e Byssus-soft-tissue interface of the byssal system. gp, giant plaque pad; fg, foot groove; mf, muscle fibres; tg, thread gland; mg, matrix gland; bf, byssal filament; ct, collagenous tissue. (Scalebars: 1 cm for A and C, 0.5 cm for B, 1 mm for D, and 50 mm for E).
Results
B. virescens histology and its distinctive giant byssus structure
Glycine is the dominant amino acid (AA) in both the byssal filament (22.87 ± 1.01 mol%) and plaque (25.33 ± 0.22 mol%) of B. virescens based on AA-composition analysis (Supplementary Fig. 1).
However, unlike the Mytilus mussel byssus, which is primarily composed of collagenous proteins13, the B. virescens byssus comprises a mixture of collagenous and non-collagenous proteins. X-ray diffractometry reveals the absence of the characteristic diffraction peak (D = 0.287 nm) representing collagen, indicating that the translational axial rise per amino acid residue along the collagen triple helix is lacking in the B. virescens byssal filament (Supplementary Fig. 2). This indicates that collagenous proteins are not the dominant constituents of the byssus. Furthermore, Masson’s trichrome (MT) stains the B. virescens byssal filament purplish in colour, unlike pure collagen, which is typically stained blue (Fig. 2c). However, Sirius red staining reveals a strong positive signal, which is indicative of collagen. Additionally, upon isolating and identifying the byssal filament proteins using electrospray ionisation–liquid chromatography–tandem mass spectrometry (ESI-LC-MS/MS), the presence of collagenous proteins is confirmed (Supplementary Data 1).
Histochemical staining of the (a–c) byssal filament, (g) foot, and (j–l) thread gland of B. virescens. Lectin-labelled (d–f) byssal filament and (h, i) foot tissue of B. virescens. Stainings performed 3 times on different tissues blocks. m Amino acid analysis profiles of a hydrolysed byssal filament (green) and d-glucosamine (black). n X-ray diffraction pattern of a deproteinised/depigmented byssal filament (green) against that of chitin from shrimp shells as a control (black). SR, Sirius red; AP, Alcian-PAS; MT, Masson’s trichrome; RCA, Ricinus communis agglutinin; WGA, wheat germ agglutinin; s-WGA, succinylated WGA. Scalebars: 20 mm for (a–f), 200 mm for (g–i), and 5 mm for (j–l). The source data are provided as a Source Data file.
We used lectin histochemistry based on 13 types of biotinylated lectin to investigate the carbohydrate distribution in the byssus and byssal glands because sugars also contribute to underwater adhesion14. Of the 13 lectins tested, only wheat germ agglutinin (WGA) and succinylated WGA (s-WGA) provided positive results for byssal filament bodies and thread vesicles (Fig. 2d–f, h–i). While WGA recognises GlcNAc and terminal sialic acid residues in glycoconjugates, which belong to the family of carbohydrates covalently linked to chemical species, such as proteins, peptides, lipids, and other compounds, its binding to these residues can be distinguished through WGA succinylation; s-WGA exclusively reacts with β-N-acetylglucosamine (β-GlcNAc) residues, as it carries a negative charge. Therefore, we infer that the byssal filament body is composed of fibres rich in β-GlcNAc, which is similar to chitin/chitosan. In addition, the surface of the byssal filament was positively stained using the Alcian blue (at pH 2.5)/periodic acid–Schiff (PAS) sequence (Alcian-PAS) as well as in the WGA and Ricinus communis agglutinin (RCA) lectin-binding assays (Fig. 2b, d, e). Considering that Alcian blue stains carboxyl-group-containing sugars (e.g., sialic acids and uronic acids) at pH 2.514, these results suggest that a mucosubstance layer that includes sialic acid and other glycocomponents exists on the byssal filament surface.
d-Glucosamine (GlcN), which constitutes 1.65 ± 0.39 mol% of the material, was identified as a chitin/chitosan or GlcNAc acid-hydrolysis product during amino acid analysis of Barbatia byssal filaments (Fig. 2m, Supplementary Fig. 1); it was not observed following deproteinisation/depigmentation (Supplementary Fig. 1). In addition, deproteinised/depigmented byssal filaments were subjected to powder X-ray diffractometry (PXRD) to confirm whether the byssal filaments contains chitin or chitosan. The diffraction peaks of the α-chitin control from shrimp shells are in good agreement with the characteristic antiparallel crystalline pattern of α-chitin; specifically the (020), (110), (120), and (130) planes, as previously reported15. However, these peaks were not observed in the X-ray diffraction pattern of deproteinised/depigmented byssal filament powder (Fig. 2n). Therefore, we infer that the B. virescens byssus uses GlcNAcylated proteins in contrast to other marine organisms (e.g., squids, crabs, shrimp, and sea spiders)16 that use chitin to create sturdy structures.
An alternative pattern (d-spacing = 16.03 ± 1.43 nm, n = 20) was observed when the ultrastructure of a vertical section of the byssal filament was characterised by transmission electron microscopy (TEM) (Supplementary Fig. 4a); this pattern was also detected using small-angle X-ray scattering (SAXS) (Supplementary Fig. 4b–e). The first SAXS peak in the radial integrated plot (with q = 0.039 Å−1/d-spacing = 16.10 nm) is ascribable to the fibrils based on the d-spacing (Supplementary Fig. 4c). Furthermore, we generated 1D azimuthal intensity profiles (I(χ)) at each scanning point to extract data that describe the average fibril orientation and the extent of fibrillar alignment (Supplementary Fig. 4d). The acquired SAXD pattern clearly indicates that the byssal fibrils are aligned parallel to the thread axis (Supplementary Fig. 4e).
EGF/EGF-like domain-rich proteins are found in the byssus/tissue interfacial contact region
Based on the discovery that the byssus is chemically similar yet physically different from chitin/chitosan (Fig. 2), we experimentally examined the potential involvement of interfacial proteins with chitin-binding domains at the tissue/byssus adhesion interface. Immunofluorescence imaging aimed at confirming the presence of proteins binding to anti-CBD antibodies and their distribution revealed that such proteins are indeed present at the tissue/byssal filament interface (Fig. 3c). Thereafter, proteins from the B. virescens foot were subjected to sodium dodecylsulfate polyacrylamide gel electrophoresis (SDS–PAGE) (Fig. 3a, Lane 1) and western blotting using an anti-CBD antibody (Fig. 3a, Lane 2). One band with strong positive signals for the anti-CBD antibody were detected at approximately 100 kDa. The corresponding proteins were identified by in-gel digestion followed by ESI-LC-MS/MS, which revealed that the band contained the mixture of proteins (bcbp-1 and bcbp-2) (Supplementary Fig. 5). In addition, the full cDNA sequences of bcbp-1 and bcbp-2 were obtained through RNA-seq and rapid amplification of cDNA ends (RACE) sequencing (Supplementary Table 1).
a 12% SDS-PAGE results for B. virescens foot proteins. M: size marker; 1: foot proteins (30 μg) stained with Coomassie brilliant blue (CBB) R‐250; 2: foot proteins (8 μg) stained with anti-CBD antibody. Experiment repeated with three times with similar results. b Schematic of the B. virescens giant byssus. c Image of the B. virescens byssus and the surrounding tissue labelled with anti-chitin-binding-domain antibody (scalebar: 100 μm). Stainings performed 3 times on different tissues blocks. d Predicting the structures of EGF/EGF-like domain-rich proteins present at the tissue/byssus interface (bcbp-1, bcbp-2). The enlarged section reveals the antiparallel β-sheet structure present in the EGF domain. e Functional domains of EGF/EGF-like domain-rich adhesive plaque matrix proteins and (f) taxonomic classification of species that possess EGF/EGF-like domain-rich adhesive plaque matrix proteins. The source data are provided as a Source Data file.
To verify that the strong positive signal observed at the byssus/tissue adhesion interface in immunofluorescence is attributable to bcbp-1 and bcbp-2, we performed peptide-blocking immunofluorescence experiments. When anti-CBD antibody was pre-incubated with an excess of peptides corresponding to sequences of bcbp-1 and bcbp-2, the majority of the positive signal at the byssus/tissue adhesion interface disappeared. This confirmed the specificity of the signal to the bcbp-1 and bcbp-2 proteins (Supplementary Fig. 6).
We next used the SMART programme (http://smart.embl-heidelberg.de/) to analyse the functional domains of the full protein sequence (Supplementary Table 1). SMART confidently predicts the presence of the EGF domain and suggests CBD possibilities, although CBD scores are less significant than the required threshold. Moreover, the protein has a repeating EGF/EGF-like-domain structure throughout its length, with 16 bcbp-1 repetitions and 19 bcbp-2 repetitions (Fig. 3e). In addition, bcbp-1 contains two putative carbohydrate-binding domains (WSC), while bcbp-2 has a transmembrane region (Fig. 3e). Although bcbp-1 and bcbp-2 are more similar to the EGF/EGF-like domain than a CBD, the cDNA-deduced amino acid compositions (Supplementary Table 2) of bcbp-1 and bcbp-2 exhibit characteristics similar to those of CBPs; specifically, they are rich in cysteine (11.84 mol% in bcbp-1 and 13.8 mol% in bcbp-2) and glycine (12.31 mol% in bcbp-1 and 15.84 mol% in bcbp-2)17. Additionally, we used sequences available in the NCBI database (https://www.ncbi.nlm.nih.gov/) to confirm the presence of adhesive plaque matrix proteins derived from an additional 49 species, which were identified as EGF-rich or EGF-containing proteins, such as bcbp-1 and bcbp-2 (Figs. 3e, f; Supplementary Data 2 and 3). Among the additional 49 discovered species, ~2% belong to the Insecta class (insects), ~20% belong to the Arachnida class (spiders), with the remaining ~78% belonging to aquatic organisms. Details of the species that possess EGF-containing adhesive plaque matrix proteins, including their peptide sequences and the number of repeating EGF domains, is provided in Supplementary Data 2.
Bcbp-1, bcbp-2, and all additional EGF/EGF-like-domain-rich adhesive plaque matrix proteins possess six conserved cysteine sites18 and an antiparallel β-sheet protein structure (Supplementary Fig. 7a) that was simulated using ColabFold, which combines AlphaFold2 with MMseqs219,20,21. Previous studies have shown that many chitin-binding and EGF/EGF-like domains have six conserved cysteine sites, and these cysteines are known to play crucial roles in forming the antiparallel β-sheet conformation, which contributes to the activity of the domain22,23,24. With this in mind, we believe that the similarities observed for the EGF/EGF-like and chitin-binding domains may have potentially led to the misclassification of marine-derived chitin-binding domains as containing EGF/EGF-like-rich proteins.
Interactions between EGF/EGF-like-domain-rich proteins and chitosan
To validate our hypothesis that the EGF/EGF-like domains found in adhesive plaque matrix proteins from various species play roles in binding to the GlcNAc units of chitin, chitosan, or glycoproteins, we quantified the interaction forces between two layers of chitosan adsorbed on mica surfaces and Mytilus edulis foot protein-2 (mefp-2), a mussel adhesive plaque matrix protein with repetitive EGF/EGF-like domains, using an SFA. We used native mefp-2 in our adhesive research for two main reasons: (i) attempts to produce recombinant bcbp-1 and -2 were unsuccessful, echoing the challenges encountered with their analogous protein, i.e., mefp-2, despite extensive research, and (ii) mefp-2 not only exhibits repetitive EGF domains such as bcbp-1 and -2, but it is also the most extensively studied analogous protein with significant research interest. One of the distinctive features of an SFA is its ability to offer simultaneous and direct measurements of force (F) as a function of absolute surface separation (D), both in situ and in real time. The apparatus boasts a force sensitivity of <10 nN and a distance resolution of approximately 1 Å. This high precision is achieved using multiple-beam interferometry with fringes of equal chromatic order (FECO)25.
Chitosan and mica, and mefp-2 and mica, primarily interact through electrostatic attraction facilitated by positively charged chitosan molecules or mefp-2 (pI = 9.14) and negatively charged mica surfaces (at pH 3.0). Surfaces were thoroughly washed with 0.1 M sodium acetate (pH 3.0) to eliminate unbound chitosan molecules or mefp-2 prior to any experiment involving the SFA. The adsorption of chitosan or mefp-2 onto each mica substrate and the actual thickness of each sample film were evaluated by observing increases in hard-wall distance (Dhw), which is defined as the asymptotic thickness of the compressed film under increasing normal load26.
Almost no adhesion energy (Wad ≈ −0.57 mJ m−2) was recorded between chitosan and the mefp-2 layer at pH 3.0 (N = 3, Fig. 4a, blue), however a higher adhesion energy of −41.80 mJ m−2 (N = 3, Fig. 4a, red) was determined at pH 5.5. Chitosan has a pKa of ~6.5–7.027, while mefp-2 has a pI of ~9; therefore, these two biopolymers experience strong electrostatic repulsion at both pH 3.0 and 5.5. As a result, strong repulsion arising from the positively charged chitosan and the positively charged protein layers counteract Wad (Fig. 4c), which suggests that the increase in adhesion observed as the pH was increased from 3.0 to 5.5 is likely the result of the realignment of chitosan molecules or mefp-2 in a manner that is conducive to adhesion to surfaces coated with the opposing mefp-2 protein or chitosan. Considering that molecules that bind to sugars, including GlcNAc, rely on pattern recognition to bind to specific sugar, we anticipate that conformational change associated with increasing pH may lead to strong adhesion between the EGF domain and GlcNAc residues of chitosan.
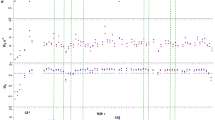
a Force–distance profiles obtained using an SFA. Measured force normalised by the radius of curvature, R (i.e., F/R) is displayed on the left axis, while the corresponding interaction energy per unit area (W) between two flat surfaces is indicated on the right. Here, Fad signifies the adhesion force, and Wad represents the adhesion energy. b Adhesion energies between chitosan and mefp-2 measured before and after blocking with GlcNAc. Adhesion energies of mefp-2 to mefp-2 or mica44, suckerin-12 to mica12, mefp-5 to mica11, Perna viridis foot protein 1 (pvfp-1) to mica or pvfp-139, chitosan to chitosan or mica27, c-dextran to c-dextran or Dad45 are also presented for comparison. c Various mica surfaces were coated to measure the adhesion force between chitosan and mefp-2. Frep signifies the electrostatic repulsive force. d Predicting chitosan mimicking oligosaccharide (GlcN-GlcNAc-GlcN) binding sites to adhesive plaque matrix proteins derived from various species containing EGF domains. The protein-ligand docking PDB files are provided as a Source Data file.
Furthermore, consistent with the predictions provided in Fig. 2, we pre-blocked the protein layer with GlcNAc before evaluating the binding strength to investigate whether or not EGF-domain binding to the GlcNAc residues of chitosan leads to strong adhesion between mefp-2, a protein rich in EGF domains, and chitosan, which contains ~25% GlcNAc units; as a consequence, no adhesion energy was recorded. Furthermore, to determine which of the two monomers comprising chitin, i.e., GlcNAc and GlcN, plays a major role in binding with EGF, the change in adhesion between mefp-2 and chitosan before and after deacetylation was investigated in 0.1 M sodium acetate (pH 5.5). After the degree of acetylation (DA) decreases from 20.0 to 5.34% (representing a 3.73-fold reduction, Supplementary Fig. 8), the absolute value of adhesion energy ( | Wad | ) between mefp-2 and chitosan decreases from 41.80 mJ m−2 to 12.92 ± 0.40 mJ m−2 (mean ± standard deviation, N = 3), indicating a 3.23-fold decrease (Source Data).
Moreover, the value of Wad between Human Epidermal Growth Factor (hEGF), which lacks Dopa (unlike mefp-2), and chitosan was determined to be −20.76 ± 8.74 mJ m−2 (N = 3), which surpasses those of suckerin and mefp-5 (Source Data). This result also suggests that the interaction between mefp-2 and chitosan is not primarily attributable to Dopa-mediated interactions. We subsequently used the Vina-Carb28 protein–ligand docking tool to assess GlcNAc containing oligosaccharides/EGF-domain interaction sites; Vina-Carb, a programme based on AutoDock Vina, incorporates an energy function that accounts for the torsion and angle of glycosidic linkages between carbohydrate molecules, offering improved precision in describing protein-carbohydrate complex systems28. In addition, we simultaneously employed its flexible docking feature to address the flexibility of the receptor protein, which allows flexibility of specified receptor residues.
The results from the Vina-Carb indicate that the fifth cysteines of the six conserved cysteines in the EGF domain, along with the conserved glycine, form critical binding sites for interaction with an oligomer that mimics chitosan (GlcN-GlcNAc-GlcN) in all 10 types of EGF/EGF-like domain rich proteins we tested. Additionally, the remaining binding sites were sequence-specific, and the interaction was mediated by hydrogen bonds and hydrophobic interactions (Fig. 4d). Carbohydrate-binding proteins typically interact with carbohydrates primarily via weak forces such as hydrogen bonding, metal-ion coordination, and hydrophobic interactions29. Nevertheless, strong overall binding should be maintained due to the increased avidity resulting from the multivalency of the EGF/EGF-like domains in various adhesive plaque matrix proteins, aside from bcbp-1 and 2 (Fig. 3e). Additionally, Supplementary Table 3 shows that the binding energy between the EGF/EGF-like domain-rich protein and (GlcNAc)3 is greater than that with (GlcN-GlcNAc-GlcN), which is consistent with the result shown in Fig. 4a, where increased acetylation corresponds to stronger binding.
In summary, we demonstrated that the EGF domain and GlcNAc residues of chitosan bind robustly. Notably, the domains were found to bind mefp-2 and chitosan comparably to, and approximately three-times more strongly than suckerin (Wad ≈ −15 mJ m−2)12 and mefp-5 (Wad ≈ −13.7 mJ m−2)11, which is recognised to be one of the most robust adhesive proteins measured by SFA (Fig. 4b).
Discussion
The increasing interest in natural materials, encompassing areas such as wet adhesives2, bio-inspired hierarchical structures30, animal mucus31, structural colours32, and impact-resistant biomaterials33, is driven by a fascination with their diverse functionalities and structures. This study provides an intriguing contribution to the evolving field of natural materials, highlighting the significance of precision and rigour across multiple scales, from the molecular to the microscopic level, in the exploring these materials. Via advanced bioinformatics and materials analyses, we uncovered the intricate mechanisms governing adhesive interactions.
Dopa, a pivotal component of bioadhesives derived from various marine species, has garnered attention for the role that it plays in wet-surface adhesion3. Nevertheless, challenges such as Dopa reactivity and susceptibility to oxidation are obstacles to practical applications4. In addition, despite the prevalence of EGF/EGF-like domain repetitions in the most abundant adhesive proteins found in mussels, which are regarded as model organisms for underwater adhesion and are also widely observed in adhesive plaque matrix proteins of other marine and terrestrial organisms, such as spiders and insects, their functions have not been completely elucidated (Fig. 3e, f, Supplementary Datas 2 and 3). Specifically, the external secretion of adhesive proteins coupled with EGF/EGF-like domains in locations where they cannot interact with growth factors implies that these domains may have unknown functions. Therefore, we speculate that although Dopa remains crucial, underwater adhesion relies on a complex interplay of mechanisms that transcend Dopa.
Herein, we show that marine organisms establish oxidation-independent and robust adhesion through binding interactions involving the EGF/EGF-like domain and GlcNAc, the chitin/chitosan monomer. This binding involves specific pattern recognition, rendering it oxidation-independent and offering insight into underwater adhesion. Moreover, this adhesion was found to be approximately three-times stronger than that of the well-known suckerin12 or mefp-511 previously regarded as adhering the strongest in marine adhesive-protein research (Fig. 4a, b).
GlcNAc, as an amino sugar, is widely recognised for its vital structural cell-surface functions; it serves as a fundamental component of bacterial-cell-wall peptidoglycans, fungal-cell-wall chitin, and the extracellular matrices of cells and tissue34. Consequently, bacterial fouling (microfouling) generates a diverse array of glycoconjugates, including GlcNAc, at adhesive interfaces involving marine organisms. Moreover, extensive research has established that microfouling plays a pivotal role in macrofouling processes35. According to previous studies, GlcNAc-containing glycoconjugates are found at adhesive interfaces where adhesive proteins with multiple EGF domains are secreted, as observed in species such as sea stars8, limpets6, sea urchins7, and sea anemones9, which suggests that EGF-domain/GlcNAc binding is widely used in the underwater adhesion mechanisms of marine organisms. Hence, understanding the factors responsible for the strong adhesion between EGF/EGF-like-domain-rich proteins and the GlcNAc present at the adhesive interface of organisms is expected to enhance our understanding of biofouling. Furthermore, as GlcNAc is one of the most abundant sugars in biofilms and biological tissue, this knowledge may lead to the development of innovative adhesion or antifouling technologies for bioelectronic, bionic-device, and tissue-engineering applications.
Moreover, chitin and chitosan exhibit considerable potential as versatile functional materials or interfacial substances because they are biodegradable, biocompatible, essentially non-toxic, and have distinctive properties36. However, the limitations of chitin/chitosan-derived materials when providing favourable cell-adhesion interfaces for specific tissue types is significantly challenging for tissue-engineering applications36. Consequently, mimicking the biological adhesion mechanism of EGF/EGF-like-domain-rich proteins to form strong bonds with chitin/chitosan monomers may stimulate cell adhesion on the surface, thereby expanding the utility of chitin/chitosan-based bioionic device platforms. Moreover, given that the tissue regenerating capabilities of EGF is an extensive research focus, its potential to induce adhesion suggests that it may serve as a good adhesive platform with strong adhesion and regenerative capabilities, thereby contributing to the development of tissue-adhesives, haemostatic agents37, multifunctional tough hydrogels, and bioelectronic interfaces. Previous research demonstrated the production of highly durable composites by combining cellulose with recombinant cellulose-binding proteins. Utilising the binding between various GlcNAc residue containing compounds (including the highly abundant, renewable biosourced waste materials chitin and chitosan) and the EGF domains, which commonly occurs in most organisms, can open avenues in sustainable material development with applications across various industries.
Methods
Ethics statement
Under Article 2, Clause 2 of the Laboratory Animal Act of South Korea [https://elaw.klri.re.kr/kor_service/lawView.do?hseq=46430&lang=ENG] and Section 2, Article 10(1) of the Act on Welfare and Management of Animals of Japan [https://www.japaneselawtranslation.go.jp/en/laws/view/3798/en], Mollusca used in this study are classified as invertebrates, and therefore, ethical approval is not required under the relevant laws of South Korea and Japan.
Sample collection
B. virescens ark clams measuring 4.5 to 5 cm were collected from the rocky coast of Moroiso Bay in Misaki, Japan. Collected clams were transferred to the laboratory in a refrigerant-filled container filled with seawater with the temperature maintained at 4 °C. The collected clams were opened by cutting the adductor muscle, and the byssal system, located centrally as shown in Fig. 1c, was carefully dissected to obtain the feet and byssus.
Amino acid analysis
Byssal filaments and plaques were hydrolysed in 6 N HCl containing 5% water-saturated phenol. Vials containing the samples were flame-sealed after being flushed with argon gas, and hydrolysis was carried out at 110 °C for 24 h. Each sample was flash-evaporated at 60 °C and washed with milliQ water (2 × 1 mL) and methanol (2 × 1 mL) until dry. The sample was resuspended in SYKAM sample-dilution buffer (250–500 μL) and analysed using a SYKAM System S4300 amino acid analyser (SYKAM, Germany).
Histology and histochemistry
The byssal system of B. virescens was carefully washed and dissected in fresh artificial seawater, after which it was fixed in paraformaldehyde solution (PAF) with sodium phosphate buffer (PBS solution, pH 7.4), rinsed with PBS solution, dehydrated in ethanol, embedded in paraffin wax, and cut into 5 μm-thick sections with a microtome (Microm HM 340 E, ThermoFisher, Waltham, MA, USA). For examination purposes, sections were stained with a Picro Sirius red (SR) stain kit (#ab150681, Abcam, Cambridge, UK), Alcian blue (pH 2.5), PAS (#ab150680, Abcam, Cambridge, UK), and MT (#25088100, Polysciences, Warrington, PA, USA) according to the manufacturer’s instructions. All sections were observed using a BX53F microscope (Olympus, Tokyo, Japan) equipped with a digital camera (IMT i-Solution, Vancouver, Canada).
Lectin/chitin/chitin binding-domain (CBD) histochemistry
The B. virescens byssal system was examined for the presence of specific carbohydrate moieties using lectins, which are proteins or glycoproteins of non-immune origin that bind carbohydrates without chemical modification. A total of 13 biotinylated lectins (Vector Laboratories, Burlingame, CA, USA) were used: Concanavalin A (Con A), Dolichos biflorus agglutinin (DBA), Griffonia (Bandeiraea) simplicifolia lectin I (GSL I), Lens culinaris agglutinin (LCA), Phaseolus vulgaris Leucoagglutinin (PHA-L), Phaseolus vulgaris Erythroagglutinin (PHA-E), Arachis hypogaea (peanut) agglutinin (PNA), Pisum sativum agglutinin (PSA), Ricinus communis agglutinin (RCA I), Glycine max (soybean) agglutinin (SBA), WGA, succinylated (Succinylated WGA), Ulex europaeus agglutinin I (UEA I), Triticum vulgaris (wheat germ) agglutinin (WGA). Previous research methodology was followed for lectin histochemical analyses14.
To identify the location of chitin-like proteins/chitin-binding domains (CBDs), deparaffinised slides were rinsed with a mixture of 0.01 M PBS at pH 7.4 containing 0.2% TWEEN 20 (TBST). The antigens were retrieved by incubating the slides in a aqueous solution containing 0.05% (w/v) trypsin and 0.1% (w/v) CaCl2 for 15 min at 37 °C. Subsequently, the slides were washed with distilled water for 3 min and treated with a blocking solution (2% (w/v) bovine serum albumin (BSA) in TBST for 1 h, RT). The blocked slides were then treated with the anti-chitin binding domain polyclonal antibody (pAb; 1:300, #PM015, MBLbio, China) in 3% (w/v) BSA-TBS-T for 2 h at 20 °C in the dark. After washing the slides three times with TBST (5 min each), they were treated with Texas-red-conjugated secondary antibodies (1:500 (v/v), #T2767, Invitrogen, Waltham, MA, USA) in 3% (w/v) BSA-TBS-T for 1 h in darkness. The slides were subsequently washed five-times with TBST (10 min each), Milli-Q water, and mounted using Vectashield mounting medium (#H-1200, Vector Laboratories, Newark, CA, USA). For the peptide pre-absorption experiment, peptides from bcbp-1 (TNPGGISKEY) and bcbp-2 (LVGYTGDPYQ) were added to the anti-chitin binding domain polyclonal antibody (pAb; 1:300, #PM015, MBLbio, China) in a 3% (w/v) BSA-TBS-T solution at a concentration of 10 mg/ml. The mixture was then agitated for 1 h at 20 °C to block the antibody’s binding sites. All other experimental conditions were identical to those described for chitin binding-domain (CBD) histochemistry.
Powder X-ray diffractometry and amino acid analysis of deproteinised/depigmented byssus
Incubation of a freeze-dried giant byssus with an alkaline peroxide cocktail (0.25 N NaOH containing 1.5% (w/v) H2O2) for 1 d at 50 °C led to deproteinisation and depigmentation16. Deionised water was used to wash the insoluble fraction, which was then freeze-dried. Samples freeze-dried using liquid nitrogen were subjected to PXRD using a D8 Advance X-ray powder diffractometer (Bruker AXS) equipped a Cu source (Cu Kα = 0.154 nm, 40 kV, 40 mA) in air in the 5–80° 2θ range at two seconds per step. The deproteinised/depigmented byssus was hydrolysed in 6 N HCl containing 5% water-saturated phenol for 24 h at 110 °C, after which its amino acid composition was determined using a ninhydrin-based amino acid analyser (S4300, SYKAM, Germany).
Wide and small angle X-ray scattering (WAXS and SAXS)
WAXS and SAXS data for the byssus were acquired on the 4 °C beam line of the Pohang Accelerator Laboratory (PAL, South Korea) synchrotron (E = 16.9 keV, λ = 0.733635 Å). Precalibrated Ti-SBA-15 (Sigma–Aldrich, Seoul, Korea) was used to calibrate scattering angles. Byssus fibres were placed in a sample holder and secured on both sides using Kapton tape. An MAR-CCD area detector was used to collect scattering data (2048 × 2048 pixels, 0.0796 mm pixel size). Data were analysed using the Nika 2D SAS macros package for IGOR Pro (WaveMetrics, Lake Oswego, OR, USA).
Transmission electron microscopy
The air-dried byssus was embedded in Embed-812 (#14120, Electron Microscopy Science, Hatfield, PA, USA), polymerised for 72 h at 60 °C, and cut into 80-nm sections with a Leica EM UC7 ultramicrotome (Leica Biosystems, Solms, Germany). Sections were rinsed briefly with Milli-Q water and post-stained by applying 2% uranyl acetate for 30 min. TEM images were obtained using a Tecnai G2 transmission electron microscope (FEI, Hillsboro, OR, USA) at 80 kV.
Transcriptome profiling
Total RNA concentrations were calculated using Quant-IT RiboGreen (#R11490, Invitrogen, Lofer, Austria)). TapeStation RNA screentape (#5067-5576, Agilent Technologies, Palo Alto, CA, USA) was used to assess RNA integrity. RNA libraries were constructed only using RNA integrity numbers (RINs) greater than 7.0. A library was independently prepared with 0.5 mg of total RNA for each sample using an Illumina TruSeq Stranded Total RNA Library Prep Gold Kit (#20020599, Illumina, Inc., San Diego, CA, USA). The rRNA in the total RNA was removed as part of the workflow, and divalent cations were used to fragment the remaining mRNA. The cleaved RNA fragments were copied into first-strand cDNA using SuperScript II reverse transcriptase (#18064014; Invitrogen, Lofer, Austria) and random primers, which was followed by the synthesis of second-strand cDNA using DNA Polymerase I, RNase H, and dUTP. These cDNA fragments were then end-repaired, a single “A” base was added, after which the adapters were ligated. Finally, a polymerase chain reaction (PCR) was used to enrich the cDNA library with purified products. Libraries were quantified using KAPA Library Quantification kits for Illumina Sequencing platforms according to the qPCR Quantification Protocol Guide (#KK4854, KAPA BIOSYSTEMS, Woburn, MA, USA) and qualified using TapeStation D1000 ScreenTape (# 5067-5582, Agilent Technologies, Palo Alto, CA, USA). Indexed libraries were then subjected to Illumina NovaSeq (Illumina, Inc., San Diego, CA, USA) and paired-end (2 × 100 bp) sequencing was performed by Macrogen Inc. After the raw reads from the sequencer had been pre-processed, we assembled the processed reads using Trinity (trinityrnaseq_r20140717) with the -SS_lib_type RF (1) strand-specific option. The Trinity programme was used for de novo transcriptome assembly, combining read sequences with certain overlap lengths to form longer fragments without N gaps referred to as “unigenes”, which were further processed for read alignment and abundance estimation using Bowtie v1.1.2 and RSEM v1.3.1. The expression level of each unigene was calculated using the fragments per kilobase of exon per million mapped fragments (FPKM) method, which excludes sequencing discrepancies when calculating gene(contig) expression and the influence of different gene lengths. Clustered unigenes were annotated using BLASTN, BLASTX, the Kyoto Encyclopedia of Genes and Genomes (KEGG), NCBI Nucleotide (NT), Pfam, Gene Ontology (GO), NCBI non-redundant protein (NR), UniProt, and EggNOG.
Protein extraction
The foot tissue of fresh B. virescens was excised, washed with sodium phosphate buffer (pH 7.0) containing 1 mM KCN, and stored at −80 °C until use. Frozen mussel feet (5.0 g, wet weight) were thawed and minced in protein extraction buffer (30 mL) (5% acetic acid (AcOH) containing 6 M urea, 5 mM KCN, 5 mM dithiothreitol (DTT), and two types of acidic protease (5 mM pepstatin A, 50 mM leupeptin)). A glass Teflon homogeniser was used to homogenise the minced feet. The homogenate was centrifuged (18,620 × g, 10 °C, 30 min), and the supernatant was used in further experiments. Byssal filament proteins were obtained using hydroxylamine extraction method38. In this process, cleaned byssal filaments were ground with liquid nitrogen using a ceramic mortar and pestle. Hydroxylamine extraction buffer (1.0 mL comprising 2.0 M hydroxylamine hydrochloride, 2.0 M guanidine hydrochloride, and 0.2 M potassium carbonate at pH 9.0) was then added to 0.2 g of the powdered byssal filaments, and the mixture was stirred at 45 °C (1500 rpm) for 4 h. The reaction was halted by adding 3.0 mL of 2% (v/v) trifluoroacetic acid and the extracted proteins were concentrated using Amicon Ultra-0.5 filters with cut-off value of a 3-kDa (#ufc500324, MilliporeSigma, Burlington, MA, USA).
Gel electrophoresis
Barbatia foot proteins were separated using 12% SDS–PAGE. Foot protein (30 mg) was loaded in each gel lane, and the gel was stained with Coomassie brilliant blue R-250.
Western blot
Foot proteins (50 mg) were run on 12% SDS–PAGE gel and transferred to a polyvinylidene difluoride membrane for western blotting. Membranes were blocked for 1 h at 20 °C with 5% BSA in TBST, then incubated for 2 h at 20 °C with anti-chitin binding domain pAb (1:1000, #PM015, MBL bio, China) diluted in 5% BSA/TBST, washed five-times with TBST, then incubated for 1 h at 20 °C with anti-IgG pAb-HRP (#AQ132P, 1:5000, Chemicon International, Inc., Temecula, CA, USA) diluted in 5% TBST-BSA. Protein bands were visualised using Clarity Western ECL Substrate (#170–5061, Bio-Rad, Hercules, CA, USA).
In-gel tryptic digestion
The 1DE bands of interest derived from the foot protein extracts were excised from the 12% SDS–PAGE gel and transferred to a 1.5-mL tubes. In addition, the byssal filament proteins were separated by approximately 1 cm on a 10% SDS-PAGE gel to eliminate low molecular weight impurities, such as detergents and buffer components, which could interfere with mass spectral analysis. The bands were washed with distilled water (100 μL) and 50 mM NH4HCO3, (100 μL, pH 7.8), after which acetonitrile (6:4, v/v) was added and shaken for 10 min. The supernatants were decanted and a speed vacuum concentrator (LaBoGeneAps, Lynge, Denmark) was used to dry the bands. The samples were then reduced (45 min, 56 °C) with 10 mM DTT in 25 mM NH4HCO3 (pH 8.0). The solution was decanted and the bands were stored at 20 °C in 55 mM iodoacetamide (IAA) in the dark for 20 min. The solution was decanted and the proteins were digested at 37 °C for 16 h with Pierce Trypsin Protease MS Grade (#90057, Thermo Fisher Pierce, Rockford, IL, USA) with a 1:50 enzyme-to-substrate ratio.
Peptide LC–MS/MS
Easy n-LC and LTQ Orbitrap XL mass spectrometers (both from Thermo Fisher, San Jose, CA, USA), each equipped with a nano-electrospray source, were used for nano LC-MS/MS. C18 nanobore columns (150 × 0.1 mm, 3-μm pore size; Agilent) were used to separate samples with mobile phase A comprising 0.1% formic acid and 3% acetonitrile in deionised water containing, and mobile phase B comprising 0.1% formic acid in acetonitrile. The following linear chromatographic gradients were used: 0 to 32% B over 23 min, 32% to 60% B over 3 min, 60% to 95% B over 3 min, returning to 0% B over 6 min (flow rate = 1500 nL/min). A full mass scan (350–1800 m/z) was performed, after which mass spectra were acquired for ten MS/MS scans. An Orbitrap resolution of 15,000 and an automatic gain control (AGC) of 2 × 105 were used for MS1 full scans, while an AGC of 1 × 104 was used for MS/MS in the LTQ.
Database searching
The Mascot algorithm (version 2.1: Matrix Science, USA) was used to identify the peptide sequences present in the transcriptome database of B. virescens foot tissue (190,418 entries, unpublished) with the following parameters: fixed modification, carbamidomethylation at cysteine residues; variable modification, oxidation at methionine residues; maximum allowed missed cleavages, two; MS tolerance, 100 ppm; MS/MS tolerance, 0.1 Da. Only peptides resulting from trypsin digestion were considered. Peptides were filtered using a significance threshold of P < 0.05.
Mass spectrometry data of byssal filament proteins were analyzed using PEAKS Studio version 12 (Bioinformatics Solutions Inc., Canada). The B. virescens foot tissue transcriptome (190,418 entries, unpublished) was used as the database source, with trypsin selected as the cleaving enzyme. Analysis parameters allowed a maximum of three missed cleavages, an MS peptide ion mass tolerance of 1.0 Da, and an MS/MS tolerance of 0.1 Da. Carbamidomethylation of cysteine was set as a fixed modification, while the following were set as variable modifications: oxidation of methionine, DOPA formation, hydroxylation, phosphorylation of serine, threonine, and tyrosine, as well as acetylation at the protein N-terminus. A maximum of three variable PTMs per peptide was allowed. The false discovery rate for peptides and proteins was controlled at a maximum of 0.1% by applying the decoy-fusion database search method in Peaks Studio. Only protein groups with a significance above this threshold are included in Supplementary Data 1. The significance score is calculated as \(-\)10log10 of the p value from significance testing, using a paired t-test.
Simulating the structures of interfacial proteins
A combination of the ColabFold (using AlphaFold2) and MMseqs2 algorithms were used to predict the 3D structures of the building-block proteins19,20,21. We used the structure generated with the highest predicted template modelling (pTM) score to illustrate the protein structure using PyMOL software. The online SMART programme (http://smart.embl-heidelberg.de/) was used to identify protein domains and motifs.
Purifying mefp-2 from mussel feet
Mussel foot protein-2 (mefp-2) was purified using 500-g flash-frozen lots of frozen blue mussel feet (Mytilus edulis; Northeast Transport Inc., ME, USA) according to published procedures39. The purified samples were freeze-dried, resuspended in 50 mM sodium acetate, and then separated into convenient aliquots for storage at 70 °C before testing.
Deacetylation of chitosan
Chitosan (#448877, Sigma–Aldrich, St. Louis, MO, USA) was mixed with a 40 wt.% NaOH solution in a ratio of 1:20, and the mixture was then heated at 100 °C for 6 h in an oven. After the reaction, the precipitate was washed with Milli-Q water until it reached a neutral pH, and it was then air-dried overnight at 60 °C. The chitosan samples were characterised before and after deacetylation using 13C nuclear magnetic resonance spectroscopy (Bruker AV-500 NMR, Bruker, Billerica, MA, USA), with CD3COOH/D2O (v/v, 1:60) used as the solvent. The degrees of deacetylation (DDs) and DAs of chitin and chitosan are calculated using the following equations:
Surface forces apparatus (SFA)
We used an SFA to measure the interaction forces between mefp-2 protein films and chitosan layers in various aqueous solutions. The working principles of the SFA and setup details are as previously reported39. In brief, two 1–5-μm-thick back-silvered thin mica surfaces were glued onto cylindrical silica disks, each with a 2 cm radius (R). The disks were mounted in the SFA chamber in a crossed-cylinder configuration; they interact in a manner equivalent to that of an approaching sphere with radius R to a flat surface when D < < R. Mica surfaces coated with medium molecular-weight chitosan (mw = 190–310 kDa, #448877, Sigma–Aldrich, St. Louis, MO, USA) and mefp-2 were prepared to measure the interaction force between them. Briefly, a 50 μg/mL solution of chitosan or mefp-2 in 0.1 M sodium acetate (50 μL; pH 3.0) was pipetted onto each mica surface and incubated for 1 h in a water-saturated chamber. The surfaces were then rinsed thoroughly with 0.1 M sodium citrate buffer (pH 3.0) to remove unbound or weakly bound molecules. Forces were assessed by injecting buffer solutions with different pH values (3.0, 5.5) between the two surfaces. Strong binding was observed between chitosan and mefp-2 at pH 5.5, suggestive of a potential interaction between the EGF domain of mefp-2 and chitosan. To ascertain this interaction, the mefp-2-coated surface was blocked with 500 μM GlcNAc in a humidity chamber (4 °C, 12 h), after which the surface was thoroughly rinsed with 0.1 M sodium citrate buffer (pH 3.0) to remove unbound GlcNAc. The contact time (tc) between the mica surfaces was 30 min, and the measured adhesion force Fad is correlated to the adhesion energy per unit area of the two flat surfaces (Wad) by:
Measuring the separation distance D in-situ is typically achieved using multiple beam interferometry FECO. Room temperature (23 °C) was maintained in all experiments. A parallel experiment was conducted using the human EGF protein (hEGF, which lacks Dopa) to investigate whether or not the Dopa in mefp-2 affects its binding to chitin. The binding affinity between hEGF (#SRP3027, Sigma–Aldrich, St Louis, MO, USA) and chitosan was examined under the same concentration and time conditions used in the abovementioned experiment involving mefp-2 and chitosan.
Protein–ligand docking
Vina-Carb28 which is based on AutoDock Vina40 was used to evaluate the binding energy between the EGF domain and oligosaccharides. The EGF domain structures of EGF-rich adhesive plaque matrix proteins found in various species were predicted using ColabFold, which combines AlphaFold2 with MMseqs219,20,21. Additionally, the initial structures of oligosaccharides (GlcNAc3 and GlcN-GlcNAc-GlcN) were generated using the tLEaP from the AmberTools programme suite41. Then, the residue names and atom types from the GLYCAM-06j force field42, as well as the glucosamine parameters developed by Arunima Singh et al. were utilised for the Vina-Carb application42,43. To determine the binding site, docking calculations were performed iteratively along the receptor protein’s amino acid sequence, with the grid box containing the ligand oligosaccharide gradually shifted and the flexible docking feature of Vina-Carb activated for the centred amino acid residue. From each calculation, 9 docking poses were obtained and the one with the strongest binding energy was selected. This process yielded a binding energy and pose for every amino acid residue. Lastly, among all results along the sequence, the pose exhibiting the strongest binding energy was chosen as the final result. The resulting binding energies and optimal docking poses are provided in the Supplementary Table 3 and Source Data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The ColabFold models generated in this study have been deposited in Zenodo (https://doi.org/10.5281/zenodo.13994402). The authors declare that all other data supporting the findings of this study are available within the article and its Supplementary Data files or from the authors upon request. The source data are provided with this article. Accession codes: The Barbatia chitin binding protein-1 (bcbp-1) and -2 (bcbp-2) datasets were deposited in Genbank of the National Center for Biotechnology Information under the accession numbers PP475025 and PP475026. The raw transcriptomic sequencing data of B. virescens foot tissue from this study are available via NCBI BioProject accession number PRJNA1194323; BioSample accession number SAMN45177264 [https://www.ncbi.nlm.nih.gov/biosample/45177264]; and SRA accession numbers SRP549524 and SRX26980243. In addition, the mass spectral proteomics data were deposited in the MassIVE repository (massive.ucsd.edu) with identifier MSV000094301 [https://massive.ucsd.edu/ProteoSAFe/dataset.jsp?task=3a043a8e07184f75941823365e7c2964] and MSV000094783 [https://massive.ucsd.edu/ProteoSAFe/dataset.jsp?task=ebe13a27f86441a2ab6b1f2d517746db]. Source data are provided with this paper.
References
Amini, S. et al. Preventing mussel adhesion using lubricant-infused materials. Science 357, 668–673 (2017).
Lee, B. P., Messersmith, P. B., Israelachvili, J. N. & Waite, J. H. Mussel-inspired adhesives and coatings. Annu. Rev. Mater. Res. 41, 99–132 (2011).
Waite, J. H. & Tanzer, M. L. Polyphenolic substance of Mytilus edulis: novel adhesive containing L-dopa and hydroxyproline. Science 212, 1038–1040 (1981).
Yu, J. et al. Mussel protein adhesion depends on interprotein thiol-mediated redox modulation. Nat. Chem. Biol. 7, 588–590 (2011).
Harrington, M. J., Masic, A., Holten-Andersen, N., Waite, J. H. & Fratzl, P. Iron-clad fibers: a metal-based biological strategy for hard flexible coatings. Science 328, 216–220 (2010).
Kang, V., Lengerer, B., Wattiez, R. & Flammang, P. Molecular insights into the powerful mucus-based adhesion of limpets (Patella vulgata L). Open Biol. 10, 200019 (2020).
Gaspar, L. et al. Interspecific analysis of sea urchin adhesive composition emphasizes variability of glycans conjugated with putative adhesive proteins. Front. Mar. Sci. 8, 737886 (2021).
Algrain, M. et al. In the footsteps of sea stars: Deciphering the catalogue of proteins involved in underwater temporary adhesion. Open Biol. 12, 220103 (2022).
Clarke, J. L., Davey, P. A. & Aldred, N. Sea anemones (Exaiptasia pallida) use a secreted adhesive and complex pedal disc morphology for surface attachment. BMC Zool. 5, 1–13 (2020).
Silverman, H. G. & Roberto, F. F. Understanding marine mussel adhesion. Mar. Biotechnol. 9, 661–681 (2007).
Danner, E. W., Kan, Y., Hammer, M. U., Israelachvili, J. N. & Waite, J. H. Adhesion of mussel foot protein Mefp-5 to mica: an underwater superglue. Biochemistry 51, 6511–6518 (2012).
Deepankumar, K. et al. Supramolecular β‐sheet suckerin–based underwater adhesives. Adv. Funct. Mater. 30, 1907534 (2020).
Coyne, K. J., Qin, X.-X. & Waite, J. H. Extensible collagen in mussel byssus: a natural block copolymer. Science 277, 1830–1832 (1997).
Choi, J., Hennebert, E., Flammang, P. & Hwang, D. S. A sugar–lectin rich interface between soft tissue and the stiff byssus of Atrina pectinata. Biomater. Sci. 8, 3751–3759 (2020).
Minke, R. & Blackwell, J. The structure of α-chitin. J. Mol. Biol. 120, 167–181 (1978).
Hwang, D. S., Masic, A., Prajatelistia, E., Iordachescu, M. & Waite, J. H. Marine hydroid perisarc: a chitin- and melanin-reinforced composite with DOPA-iron(III) complexes. Acta Biomater. 9, 8110–8117 (2013).
Huang, X., Xie, W.-j & Gong, Z.-z Characteristics and antifungal activity of a chitin binding protein from Ginkgo biloba. FEBS Lett. 478, 123–126 (2000).
Tjoelker, L. W. et al. Structural and functional definition of the human chitinase chitin-binding domain. J. Biol. Chem. 275, 514–520 (2000).
Mirdita M. et al ColabFold-Making protein folding accessible to all. Nat. Method 19, 679–682 (2022).
Hassabis D. & Jumper J. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Ikegami, T. et al. Solution structure of the chitin-binding domain of Bacillus circulans WL-12 chitinase A1. J. Biol. Chem. 275, 13654–13661 (2000).
Suetake, T. et al. Chitin-binding proteins in invertebrates and plants comprise a common chitin-binding structural motif. J. Biol. Chem. 275, 17929–17932 (2000).
Schrijver, I., Liu, W., Brenn, T., Furthmayr, H. & Francke, U. Cysteine substitutions in epidermal growth factor–like domains of fibrillin-1: distinct effects on biochemical and clinical phenotypes. Am. J. Hum. Genet. 65, 1007–1020 (1999).
Bhushan, B., Israelachvili, J. N. & Landman, U. Nanotribology: friction, wear and lubrication at the atomic scale. Nature 374, 607–616 (1995).
Benz, M., Gutsmann, T., Chen, N., Tadmor, R. & Israelachvili, J. Correlation of AFM and SFA measurements concerning the stability of supported lipid bilayers. Biophys. J. 86, 870–879 (2004).
Lee, D. W., Lim, C., Israelachvili, J. N. & Hwang, D. S. Strong adhesion and cohesion of chitosan in aqueous solutions. Langmuir 29, 14222–14229 (2013).
Nivedha, A. K., Thieker, D. F., Makeneni, S., Hu, H. & Woods, R. J. Vina-Carb: improving glycosidic angles during carbohydrate docking. J. Chem. Theory Comput. 12, 892–901 (2016).
Zeng, X., Andrade, C. A., Oliveira, M. D. & Sun, X.-L. Carbohydrate–protein interactions and their biosensing applications. Anal. Bioanal. Chem. 402, 3161–3176 (2012).
Hu, H. et al. Bioinspired hierarchical structures for contact‐sensible adhesives. Adv. Funct. Mater. 32, 2109076 (2022).
Cerullo, A. R. et al. Comparative mucomic analysis of three functionally distinct Cornu aspersum Secretions. Nat. Commun. 14, 5361 (2023).
Xiao, M. et al. Bio-inspired structural colors produced via self-assembly of synthetic melanin nanoparticles. ACS nano 9, 5454–5460 (2015).
Huang, W. et al. Nanoarchitected tough biological composites from assembled Chitinous Scaffolds. Acc. Chem. Res. 55, 1360–1371 (2022).
Naseem, S. & Konopka, J. B. N-acetylglucosamine regulates virulence properties in microbial pathogens. PLoS Pathog. 11, e1004947 (2015).
Ank, G., Porto, T. F., Pereira, R. C. & da Gama, B. A. Effects of different biotic substrata on mussel attachment. Biofouling 25, 173–180 (2009).
Yang, T.-L. Chitin-based materials in tissue engineering: applications in soft tissue and epithelial organ. Int. J. Mol. Sci. 12, 1936–1963 (2011).
Doan, P. L. et al. Epidermal growth factor regulates hematopoietic regeneration after radiation injury. Nat. Med. 19, 295–304 (2013).
Walker J. M. The protein protocols handbook (Springer Science & Business Media, 1996).
Hwang, D. S., Zeng, H., Lu, Q., Israelachvili, J. & Waite, J. H. Adhesion mechanism in a DOPA-deficient foot protein from green mussels. Soft Matter 8, 5640–5648 (2012).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Case, D. A. et al. AmberTools. J. Chem. Inf. Modeling 63, 6183–6191 (2023).
Kirschner, K. N. et al. GLYCAM06: a generalizable biomolecular force field. Carbohydrates. J. Comput. Chem. 29, 622–655 (2008).
Singh, A. et al. Extension and validation of the GLYCAM force field parameters for modeling glycosaminoglycans. Can. J. Chem. 94, 927–935 (2016).
Hwang, D. S. et al. Protein-and metal-dependent interactions of a prominent protein in mussel adhesive plaques. J. Biol. Chem. 285, 25850–25858 (2010).
Park, J., Park, J., Lee, J., Lim, C. & Lee, D. W. Size compatibility and concentration dependent supramolecular host–guest interactions at interfaces. Nat. Commun. 13, 112 (2022).
Acknowledgements
We thank Kousaku Ohkawa (Institute for Fiber Engineering, ICCER, Shinshu University) for collecting Barbatia virescens. This work was performed with the support of the Pohang Accelerator Laboratory (PAL). This study was supported by the National Research Foundation of Korea (NRF) grant funded by the Ministry of Education (grant number 2022R1I1A1A01064177 to J.C.) and the Korean government (MSIT) (No.2022R1A2C2007874, RS-2024-00408795 to DSH, RS-2024-00340746 to J.C.).
Author information
Authors and Affiliations
Contributions
Conceptualisation: J.C.; methodology: J.C., S.L. (SFA measurements), Y.L. (Vina-Carb simulation); investigation: J.C.; visualisation: J.C.; funding acquisition: D.S.H. and J.C.; project administration: J.C.; supervision: D.S.H.; writing – original draft: J.C.; writing – review & editing: J.C. and D.S.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Olga Makshakova and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Choi, J., Lee, S., Lee, Y. et al. Sticky organisms create underwater biological adhesives driven by interactions between EGF- and GlcNAc- containing polysaccharides. Nat Commun 16, 233 (2025). https://doi.org/10.1038/s41467-024-55476-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-55476-4
This article is cited by
-
Reversible biobased adhesives enable closed-loop engineered composites
Nature Communications (2025)