Abstract
Many essential proteins require pyridoxal 5’-phosphate, the active form of vitamin B6, as a cofactor for their activity. These include enzymes important for amino acid metabolism, one-carbon metabolism, polyamine synthesis, erythropoiesis, and neurotransmitter metabolism. A third of all mammalian pyridoxal 5’-phosphate-dependent enzymes are localized in the mitochondria; however, the molecular machinery involved in the regulation of mitochondrial pyridoxal 5’-phosphate levels in mammals remains unknown. In this study, we used a genome-wide CRISPR interference screen in erythroleukemia cells and organellar metabolomics to identify the mitochondrial inner membrane protein SLC25A38 as a regulator of mitochondrial pyridoxal 5’-phosphate. Loss of SLC25A38 causes depletion of mitochondrial, but not cellular, pyridoxal 5’-phosphate, and impairs cellular proliferation under both physiological and low vitamin B6 conditions. Metabolic changes associated with SLC25A38 loss suggest impaired mitochondrial pyridoxal 5’-phosphate-dependent enzymatic reactions, including serine to glycine conversion catalyzed by serine hydroxymethyltransferase-2 as well as ornithine aminotransferase. The proliferation defect of SLC25A38-null K562 cells in physiological and low vitamin B6 media can be explained by the loss of serine hydroxymethyltransferase-2-dependent production of one-carbon units and downstream de novo nucleotide synthesis. Our work points to a role for SLC25A38 in mitochondrial pyridoxal 5’-phosphate accumulation and provides insights into the pathology of congenital sideroblastic anemia.
Similar content being viewed by others
Introduction
Pyridoxal 5’-phosphate (PLP), the active form of vitamin B6, is an essential cofactor in cellular metabolism. The human genome encodes nearly 60 PLP-dependent enzymes, which catalyze reactions in diverse pathways (see Supplementary Data 11), including key aspects of amino acid and neurotransmitter metabolism, folate and one-carbon metabolism, polyamine synthesis, and carbohydrate and lipid metabolism, with roles in various mitochondrial-dependent cellular processes and erythropoiesis2. Inherited metabolic diseases leading to PLP deficiency result in a number of pathogenic biochemical anomalies, including severe neurological disorders (e.g. B6-dependent and responsive epilepsy3 and polyneuropathy4) and bone mineralization defects (i.e. hypophosphatasia5,6). Nearly 30 genes encoding PLP-dependent enzymes are mutated in disease (Supplementary Data 1), spanning a wide range of pathologies, including congenital sideroblastic anemia (CSA) (i.e., Δ-Aminolevulinic acid synthase (ALAS2)7 deficiency) and homocystinuria (Cystathionine-β-synthase (CBS) deficiency)8.
Approximately a third of all PLP-dependent enzymes reside in the mitochondria (Supplementary Data 1), including rate-limiting enzymes involved in heme biosynthesis and one-carbon metabolism. Thus, PLP accumulation in the mitochondria, its homeostasis, and the formation of a Schiff base with the active site lysine of its dependent enzymes play key roles in many pathways critical for metabolism, physiology and human disease. Despite its importance, the factors that mediate PLP distribution and maintenance in different cellular compartments in metazoans are poorly understood (Fig. 1A). In an attempt to search for genes that regulate vitamin B6/PLP metabolism, we used whole genome CRISPRi screening to identify conditionally essential genes in low vitamin B6 conditions. We identify SLC25A38 as the most essential gene in vitamin B6-limited conditions and provide data to support a role in mitochondrial PLP accumulation, providing new insight into CSA caused by SLC25A38 deficiency.
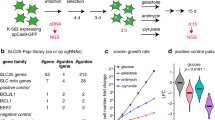
A Cellular uptake of phosphorylated B6 vitamers involves dephosphorylation by phosphatases like TNSALP72. Unphosphorylated forms cross the cell membrane via an unclear mechanism and are phosphorylated by pyridoxal kinase (PDXK). Pyridoxine 5’-phosphate oxidase (PNPO) converts pyridoxine 5’-phosphate (PNP) and pyridoxamine 5’-phosphate (PMP) to pyridoxal 5′-phosphate (PLP) important in the salvage and recycling of intracellular B6. PLPBP (fka PROSC) is key for vitamin B6 homeostasis, possibly a PLP-chaperone73. PLP functions as a cofactor for ~60 human enzymes (Supplementary Data 1), forming Schiff bases with lysine residues (catalytic pocket). B Cells grown in media depleted of vitamin B6 display reduced cumulative growth, magnified by PDXK knockdown. C Mitochondria isolated using the “Mito-IP” method and analyzed by LC-MS alongside with the whole cell lysates (“WC”). Mock IP refers to the exact same Mito IP procedure but executed using cells which do not express Mito-tag. Pyridoxal (PL), PLP and PMP were reliably detected in the mitochondrial fraction above the background (Mock IP) but not pyridoxine (PN). PNP and pyridoxamine (PM) signals were below the detection limits of our instrument. Area ratios are shown as peak areas of the target ion divided by peak areas of internal standards (D3-PN and D3-PLP for unphosphorylated and phosphorylated vitamers respectively) normalized by cell counts; Box plots show min. to max. values, median, and SD for n = 4, each an independent culture of cells. Significance level were indicated as *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 per two-way ANOVA with Šídák post hoc tests. D Diagram describing the genome-wide CRISPRi screening procedure. E Gene scores (GS) for each gene as identified in the B6 depleted (“B6-“) or B6 rich (“B6+”) arms of the CRISPRi screen are shown, highlighting the highest delta gene-scored (dGS) genes. F Top dGS genes are shown following the described color codes. A created in BioRender. Pena, I. (2025) https://BioRender.com/l99m551. D, created in BioRender. Pena, I. (2025) https://BioRender.com/g50p925. Source data are provided as a Source Data file.
Results
A whole-genome genetic screen reveals SLC25A38 as conditionally essential in low vitamin B6 conditions
To identify genes that are specifically required for proliferation in conditions where vitamin B6 is limited, we formulated a culture media that lacks both pyridoxine and albumin-bound PLP. Albumin-bound PLP was depleted from fetal bovine serum added to medium using a hydroxylamine reaction9 and subsequent dialysis. For our screen, we used K562 human erythroleukemia cells, since these cells are dependent on many PLP-dependent pathways for proliferation10,11,12,13,14, and have been widely used in CRISPR-based genetic screens10,11,12,15. K562 cells cultured in B6-depleted media display slower proliferation (Fig. 1B), and this phenotype is exacerbated when a key enzyme in the B6 salvage pathway, pyridoxine kinase (PDXK), is knocked down (Fig. 1B, Supplementary Fig. 1A), suggesting this media condition (“B6-depleted”) is sufficient to cause reductions in cell proliferation mediated by B6 deficiency. Normal plasma pyridoxine (PN) in humans ranges between 7 and 60 nanomolar (nM)16. The concentration of this vitamin B6 vitamer in full RPMI media, 4.8 μM, is supraphysiological compared to blood levels, and a few doublings in B6-depleted media were necessary to acheive maximum reduction in proliferation (Fig. 1B) and low cellular B6 levels (Fig. 1C). Mass spectrometry analysis confirmed low intracellular (whole cell) and intramitochondrial (immunopurified mitochondria using the Mito-IP approach17,18) vitamin B6 vitamers in cell cultures collected after 14 doublings in B6-depleted media (Fig. 1C) compared to full RPMI (“B6-rich”). In the mitochondria, we detected the phosphorylated species PLP and pyridoxamine 5’-phosphate (PMP), whereas PN was only found in the whole cell lysate, consistent with previous studies19. Pyridoxal (PL) was also detected in the mitochondria, which is in line with the presence of an enzyme capable of hydrolyzing PLP in the intermembrane space19 although no hydrolase activity is thought to exist in the mitochondrial matrix where the prevalent species are PLP and PMP19. PMP likely comes from half-transamination in the active site of aminotransferases19. Mitochondria lack vitamin B6 kinase activity19, therefore the phosphorylation of B6 vitamers by PDXK and conversion into PLP occurs in the cytosol and the phosphorylated species are further transported to the mitochondria via an unknown carrier (Fig. 1A). Accordingly, culturing cells in B6-depleted media not only dropped the levels of both unphosphorylated and phosphorylated B6 vitamers (Fig. 1C) but also induced various perturbations to the polar metabolome that may correlate with reduced activity of PLP-dependent enzymes (Supplementary Fig. 1C).
K562 cells expressing endonuclease-deficient Cas9 fused with the Kruppel-associated box (dCas9-KRAB)20 were transduced with a genome-wide lentiviral single guide RNA (sgRNA) library21 (~10 sgRNAs per gene, 209,070 total including 3790 control sgRNAs21, Fig. 1D) that induces transcriptional repression (CRISPRi) with minimal off-target effects. Transduced cells were selected and cultured in B6-depleted or B6-rich RPMI media and, for each gene, we generated a gene score (GS) by calculating the mean log2 fold change of guide representation from the beginning (day 0) to the end of the culture period (14 doublings) for all the sgRNAs targeting that gene (Fig. 1D, E). Most genes displayed similar gene scores in cells cultured in B6-rich or B6-depleted media (Fig. 1E) and known essential genes had a shifted distribution towards negative gene scores in both conditions (Supplementary Fig. 2A-C). As predicted, genes encoding key enzymes in the vitamin B6 salvage pathway (PDXK, PNPO and PROSC) (Fig. 1A) were conditionally essential in the B6-depleted condition (Fig. 1E, F), whereas Pyridoxal Phosphatase (PDXP) loss was beneficial in this media likely due to reduced dephosphorylation of PLP and loss of the active form of the cofactor (Fig. 1E, Supplementary Fig. 2D). A number of genes encoding PLP-dependent enzymes also scored differentially in B6-depleted media such as the cytosolic Ornithine Decarboxylase 1 (ODC1), Selenocysteinyl-tRNA(Sec) synthase (SEPSECS), and the genes encoding the mitochondrially-localized Serine hydroxymethyltransferase 2 (SHMT2) and Glutamic-Oxaloacetic Transaminase 2 (GOT2) (Fig. 1F and Supplementary Fig. 3A). In contrast, SHMT1 and GOT1, encoding for the respective cytosolic isoenzymes that catalyze the same reaction as SHMT2 and GOT2, did not show negative gene scores in the B6-depleted conditions, indicating the mitochondrial branches of these PLP-dependent pathways are particularly essential in low B6 in K562 cells. These results are corroborated by recent CRISPR screens in leukemia cells which identified PDXK as a key gene for proliferation10 emphasizing the importance of vitamin B6 metabolism in these cells. Not all PLP-dependent enzyme genes scored as hits under the conditions tested, likely because they are not expressed in this cell line or belong to pathways that are not essential in K562 cells. PLP depletion may thus have distinct effects on metabolism in different cells, and the essentiality of PLP-dependent enzymes is expected to vary in a cell-type- and environmental condition-dependent context, as seen with neurotransmitter biosynthesis deficiency in B6-dependent epilepsies22,23 and with regard to heme biosynthesis defects in congenital sideroblastic anemia (CSA)24. In addition to the expected gene hits, the membrane protein-encoding gene with the most significant differential gene score observed between B6-rich and B6-depleted conditions was SLC25A38 (Fig. 1F, Supplementary Fig. 3B), an inner mitochondrial membrane carrier protein25,26,27.
Loss of SLC25A38 leads to a specific depletion of mitochondrial PLP and proliferation defects in low vitamin B6 growth conditions
To validate the screen findings across independent experiments, we designed a small CRISPRi-targeted library comprised of sgRNAs for the top and bottom 300 scoring genes (based on delta-gene (dGS) scores with p < 0.05), all PLP-dependent enzymes, and control guides (total: 7500 sgRNAs and 750 non-targeting controls) (Supplementary Data 4). K562 cells were infected and cultured exactly as in the genome-wide CRISPRi screen. This targeted library screen reproduced the main findings of the genome-wide screen (Fig. 2A, B), with SLC25A38 again observed as a top-scoring gene, conditionally essential in vitamin B6-depleted conditions (Fig. 2C, Supplementary Fig. 3B). SLC25A38 was originally identified as a gene defective in a severe form of congenital sideroblastic anemia (CSA)26 and later a role in glycine transport was reported for both the human protein and its yeast ortholog, Hem25p25,26,27. More recently, an additional role in the mitochondrial transport of isopentenyl pyrophosphate (IPP) has been proposed uniquely for the yeast Hem25p protein28. Here, we explored whether SLC25A38 might have an unexpected role, either directly or indirectly, in mitochondrial PLP.
A Gene Scores (GS) for the targeted library screen in -B6 or +B6 conditions. B Top differentially essential genes with reduced fitness in B6 stress, with SLC25A38 showing the highest delta Gene Score (dGS: GS in +B6 minus GS in -B6). C Guide scores for SLC25A38 in the genome-wide and targeted CRISPRi screens. D Diagram depicting the PLP-dependent glycine metabolism pathways in mitochondria and cytosol; PLP, pyridoxal 5′-phosphate; CH2-THF, Methylenetetrahydrofolate; 5-ALA, aminolevulinic acid; SHMT, serine hydroxymethyltransferase; GCV, glycine cleavage system (P-protein); ALAS2, Delta-aminolevulinate synthase 2; GCAT, Glycine C-acetyltransferase; PSAT1, phosphoserine aminotransferase 1; SFXN1, sideroflexin 1; ABCB6, ATP-Binding Cassette Sub-Family B Member 6; NFS1, cysteine desulfurase; ISC, iron-sulfur clusters. E Proliferation in B6 rich or B6 poor conditions for clonal knockout cell lines of SLC25A38, SLC25A39, PDXK, and PROSC. Proliferation defects were observed in PDXK, PROSC, and SLC25A38 knockouts in -B6, but not in WT and SLC25A39 knockouts. Cells were plated for proliferation assays after 3 days of conditioning in each respective media. Doublings are shown as mean ± SD; n = 3 independent cell cultures (E), two-way ANOVA followed by Šídák post hoc analysis (E) *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Each n was defined as an independent culture of cells. F K562 cells were conditioned in B6-depleted media supplemented with low B6 (1 nM PN), physiological B6 (10 nM PN), or high B6 (1000 nM PN) for a longer period (6 days, media change every 2 days) and then plated onto 96 well plates at 10.000 cells per well (two replicates) for growth over 4 days in eight media conditions: B6-depleted media without supplementation, +0.5 nM PN, +1 nM PN, +5 nM PN, +10 nM PN, +50 nM PN, +100 nM PN and +1000 nM PN (“PN added to B6-depl.”). Doublings were estimated from absorbance reads using the Presto Blue assay at day 3 (E) or day 4 (F). D Created in BioRender. Pena, I. (2025) https://BioRender.com/m62v944. Source data are provided as a Source Data file.
We generated clonal knockout SLC25A38-(KO) cells by transiently expressing Cas9 and gene-specific sgRNAs followed by single cell expansion and selection of clones with loss-of-function mutations (Supplementary Fig. 3C, Supplementary Fig. 4A). Clonal SLC25A38-KO lines displayed a drastic proliferation defect uniquely in B6-depleted conditions (Fig. 2E). Importantly, this phenotype was observed in physiological B6 conditions resembling normal plasma pyridoxine levels (low nanomolar range) (Fig. 2F). PROSC or PDXK loss, which play roles in B6 homeostasis and the cytosolic salvage pathway (Fig. 1A)4,23,29, also induced proliferation defects in B6-depleted conditions, albeit milder than that seen with SLC25A38 loss (Fig. 2E, F), in line with the gene score results from both genetic screens (Fig. 1E, F and Fig. 2A, B). SLC25A39, encoding an inner mitochondrial membrane protein recently identified as a glutathione carrier30,31, was also knocked out as a control, with no effect on proliferation in B6-rich or B6-depleted conditions (Fig. 2E). The proliferation defect in SLC25A38-KO cells was more pronounced when the cells were pre-conditioned in B6-depleted media for 6 days before assessing proliferation. (Fig. 2F, with media changes every 2 days). Jurkat cells lacking SLC25A38 also displayed reduced proliferation in B6-depleted conditions but not when the mitochondrial serine transporter, SFXN1, was knocked out as a control (Supplementary Fig. 4E).
To test if SLC25A38 loss has an effect on cellular or mitochondrial vitamin B6 levels, we isolated mitochondria using the Mito-IP approach17,18 and measured PLP levels by LC-MS (Supplementary Fig. 3C, E). For these experiments, we cultured cells in B6-depleted media supplemented with 10 nM Pyridoxine (“B6-physiol.”) to match physiologically-relevant plasma levels. Notably, PLP levels were reduced in mitochondria isolated from SLC25A38-KO cells compared to wild-type cells (Supplementary Fig. 3E) even though levels were not depleted at the whole cell level (Supplementary Fig. 3F). The mitochondrial PLP levels of SLC25A38-KO cells cultured in an excess of Pyridoxine (“B6-high”: B6-depleted media supplemented with 1000 nM PN) still showed a mild reduction compared to WT cells (~3.9× reduction in “B6-physiol” versus 1.5x reduction in “B6-high” Supplementary Fig. 3E) indicating that lack of SLC25A38 affects PLP in the mitochondria at physiological B6 levels (e.g. 10 nM PN), but that in excess B6 conditions, SLC25A38-independent mechanisms may partially compensate to support mitochondrial PLP levels. Mitochondrial PLP levels did not decrease in cells lacking SLC25A39 under any conditions. Instead, glutathione (GSH) was specifically depleted in mitochondria but not in the whole cell lysates from SLC25A39-KO cells as expected30,31 (Supplementary Fig. 3G, H).
Expression of SLC25A38 or its yeast homolog Hem25p, but not SLC25A39 or CSA patient variants of SLC25A38, rescues mitochondrial PLP levels in SLC25A38-KO cells
We next cloned and expressed FLAG-tagged human wild-type SLC25A38 and SLC25A39 cDNAs in SLC25A38-KO cells to investigate whether either gene could rescue the cells’ proliferation defect in low B6 and restore mitochondrial PLP levels. We also expressed a cDNA of the yeast ortholog of SLC25A38, hem25, codon-optimized for expression in human cells, to test whether a rescue effect would stem from a conserved function or a specific function of the human protein. SLC25A38-KO cells grew similarly to WT cells upon overexpression of FLAG-tagged SLC25A38 or Myc-tagged Hem25p (Fig. 3A), and a rescue of both mitochondrial PLP and PMP levels was observed (Fig. 3B, C). In fact, overexpression of SLC25A38 beyond WT levels led to higher mitochondrial PLP and PMP levels than those observed in WT cells (Fig. 3B, C). We observed no rescue of proliferation in low B6 nor rescue of mitochondrial PLP levels with an empty vector or upon expression of SLC25A39 cDNA or a codon-optimized version of MTM132 (the closest yeast homolog to SLC25A39) in SLC25A38-null cells (Fig. 3A, C). In contrast, overexpression of SLC25A39 led to a 4-fold increase in mitochondrial GSH levels (Supplementary Fig. 5B), while not affecting mitochondrial PLP levels, confirming that expressed SLC25A39 is active.
A Growth defect of SLC25A38-KO cells in low B6 is rescued by expression of an add-back construct (FLAG-SLC25A38-WT) and the yeast ortholog, Hem25p (3xMyc tagged). Cells were grown for 3 days in B6-depleted media supplemented with 1 nM or 1000 nM pyridoxine prior to plating 10,000 cells/well in 96 well plates of the respective media with growth accessed using the Presto Blue assay. Low Mitochondrial PLP (B) and PMP (C) levels in SLC25A38-KO cells are also rescued by expression of wild-type SLC25A38 and the yeast ortholog Hem25p but not when SLC25A39 or its yeast orthologue Mtm1p was expressed. D Structural model of the human SLC25A38 highlighting residues in predicted substrate contact sites chosen for mutagenesis studies; asymmetric: R134, R278, and R282; symmetric: R187; amino group binding: R187 and D188. E SLC25A38 point mutant constructs were generated to express mutations affecting putative substrate binding sites: R134C, R143H, R187Q, D188H, R278A, and R282A; expressing these constructs in SLC25A38-KO cells do not fully rescue growth in low B6 but expression of wild-type FLAG-SLC25A38 add-backs (“A38-WT”) does. In line with the lack of growth rescue, SLC25A38-KO expressing the above-mentioned point-mutants fail to rescue mitochondrial PLP (F) and PMP (G) with no significant changes to the whole cell levels of PLP (H). For Mito-IP experiments, cells were grown in media supplemented with 10 nM pyridoxine for 3, days n = 3 ± SD; Doublings are shown for n = 3 ± SD; Two-way ANOVA with Šídák (A, E) and one-way ANOVA (B, C, F–H) with Dunnet multiple-comparison tests were used, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Each n was defined as an independent culture of cells. Source data are provided as a Source Data file.
It is possible that SLC25A38 affects mitochondrial PLP levels via a function independent of its transport activity. To test this possibility, we asked whether the rescue of both proliferation and mitochondrial PLP levels in SLC25A38-KO cells can be phenocopied by expression of SLC25A38 variants in which putative substrate contact sites are mutated. Mitochondrial carriers share common structural features, including a single substrate binding site that functions via salt bridge networks between flanking conserved and symmetrical triplets of charged residues (e.g. arginines (R) and aspartic acids (D)) coupled to opening and closing of the transport protein on either side of the membrane. Using computational biology tools, Kunji33 predicted that SLC25A38 has three asymmetric (R134, R278, and R282) and three symmetric (R96, R187, and R281) positively charged and highly conserved residues which may be involved in the substrate binding site33,34,35 (Fig. 3D). R187 and D188 might specifically be involved in binding to a substrate amino group33,35 (Fig. 3D). Interestingly, a number of these residues (R134, D188, R187, R282, and R278) are also found to be mutated in patients suffering from Congenital Sideroblastic Anemia (CSA)36,37. To study the effects of these amino acid alterations on SLC25A38 function in maintaining mitochondrial PLP levels, we generated the following FLAG-tagged SLC25A38 mutants: p. Arg134Cys, p. Arg134His, p. Asp188His, p. Arg187Pro, p. Arg187Gln, p. Arg282Ala, p. Arg278Ala, which were expressed in SLC25A38-KO cells. Importantly, the mutated SLC25A38 constructs still localized to the mitochondria, as evidenced by enrichment in the mitochondrial IP fraction (Supplementary Fig. 5E, with p. Arg134Cys, p. Arg134His, and p. Arg278Ala expressed at similar levels to the add-back WT), and specifically to the inner mitochondrial membrane as shown by indirect immunofluorescence staining (Supplementary Fig. 6). While expression of WT SLC25A38 cDNA fully rescues both proliferation and mitochondrial PLP and PMP levels in B6 physiol and low B6 media, none of the mutated variants rescued either phenotype (Fig. 3E–H).
SLC25A38-null cells fail to accumulate isotopically labeled PLP into the mitochondria
To gain further insights into how the lack of SLC25A38 affects mitochondrial PLP levels, we added 100 nM of isotopically labeled (2’,2’,2’)-2H3-pyridoxine (D3-PN, “M + 3”) to cells in B6-physiol media and rapidly purified mitochondria 1 h later by Mito-IP to trace the 2H atoms into isotopically labeled (2’,2’,2’)-2H3-PLP (M + 3) (Fig. 4A). This would enable testing whether PN to PLP conversion in the cytosol or the accumulation of PLP in the mitochondria is affected by loss of SLC25A38. Our data suggests the latter to be the most likely mechanism. Despite detecting M + 3 PLP in the whole cell lysates of WT cells, as well as in SLC25A38-KO and SLC25A39-KO cell lines, almost no labeled M + 3 PLP was detected in the mitochondria isolated from SLC25A38-KO cells (Fig. 4B). In contrast, M + 3 PLP accumulated similarly in the mitochondria of SLC25A39-KO and WT cells (Fig. 4B). A similar picture is seen for M + 3 PMP (Supplementary Fig. 7A).
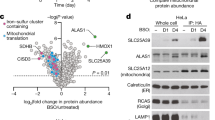
A Cells were supplemented with 100 nM of isotopically labeled (2’,2’,2’)-2H3-pyridoxine (D3-PN, “M + 3”) and the deuterated methyl group was traced into mitochondrial D3-PLP by Mito-IP 1 h later. Corrected ion counts are shown for the endogenous unlabeled isotopologue PLP (M0) and labeled, synthesized from the tracer, (2’,2’,2’)-2H3-PLP is shown as D3-PLP (“M + 3”). B Despite detecting a significant signal of M + 3 PLP in the mitochondria of WT cells and SLC25A39-KOs, SLC25A38-KO mitochondria failed to accumulate M + 3 PLP, despite the presence of labeled M + 3 PLP in the whole cell extracts indicating the conversion of M + 3 PN into M + 3 PLP via PDXK and PNPO occurs. Overexpression of WT SLC25A38 (“AB”, add-back) leads to increased import of M + 3 PLP in the mitochondria compared to SLC25A38-KO transduced with an empty vector (“ev) or overexpressing a mutant form of SLC25A38, R134C both after 1 h (C) or 3 h (D) incubation with D3-PN. In both experiments, K562 cells were grown in B6-physiol. medium (10 nM PN) for 2 days prior to the tracing study. Mass isotopomer distributions were corrected for natural abundance using IsoCorrectoR and shown as “corrected ion counts” (B–D), n = 3 ± SD; Asterisks indicate significance as in two-way ANOVA Dunnett test for multiple comparison to WT levels (B–D), n = 3, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Each n was defined as an independent culture of cells. Synthetic lethal genetic interactions mined from The Cell Map38,39 for S. cerevisiae hem25Δ (SLC25A38 ortholog (E)), bud17Δ (PDXK ortholog (F)) and tpn1Δ (the yeast pyridoxine transporter in the plasma membrane (G)) and mtm1Δ (SLC25A39 ortholog (H) from genome-wide deletion screens. Genetic interaction scores and statistical confidence measures (p-values) were obtained from The Cell Map39,74 A, created in BioRender. Pena, I. (2025) https://BioRender.com/i81d057. Source data are provided as a Source Data file.
We then asked whether expression of the WT SLC25A38 cDNA in SLC25A38-KO cells (add-back or “AB”) or one of the mutant forms inherited in CSA (p. Arg134Cys or “R134C”) affected accumulation of labeled PLP into the mitochondria (Fig. 4C, D). The same experiment was carried out with mitochondria isolation by Mito-IP performed after cells were incubated for 1 h (Fig. 4C) or 3 h (Fig. 4D) with labeled pyridoxine (D3-PN). Similar to what we previously observed, the signal for PLP M + 3 in the mitochondria of SLC25A38-KO cells was near background levels (Fig. 4C, D). Overexpression of WT SLC25A38 led to much higher accumulation of D3-PLP (M + 3): 3.6 and 4.4-fold higher than in WT cells after 1 h and 3 h incubation with D3-PN, respectively (Fig. 4C, D). Such an increase was not seen upon expression of the CSA mutant form SLC25A38 p. Arg134 Cys (Fig. 4C, D). We observed the same pattern for M + 3 PMP (Supplementary Fig. 7B, C). Together, these data suggest that mitochondrial B6 accumulation is impaired in SLC25A38-KO cells. Such role can be a direct or indirect one in mitochondrial transport, retention, regulation or maintenance of PLP.
Finally, given that both the proliferation defect in low B6 and the low mitochondrial PLP levels could also be rescued by Hem25p expression (Fig. 3), the Saccharomyces cerevisiae ortholog of SLC25A38, we hypothesized that a function for this protein in mitochondrial B6 regulation may be conserved in yeast. If true, yeast genetic coessentiality data may inform which genes are synthetic lethal interactors and thus relevant for hem25 functions. By mining unbiased yeast gene interaction network data from The Cell Map38,39, we identified bud17 (yeast ortholog for PDXK, p = 0) and tpn1 (encoding Tpn1p, the yeast pyridoxine transporter in the plasma membrane40, p = 4.85 × 10−50) as the top and the fourth highest scoring synthetic lethal interactions of hem25Δ (Fig. 4E). hem25 is also the top scoring synthetic lethal interactor in bud17Δ and tpn1Δ (Fig. 4F, G), further indicating that Hem25p is likely linked to B6 maintenance in yeast.
Mitochondrial glycine synthesis via SHMT2 is impaired in low B6 conditions uniquely in SLC25A38-KO cells
To investigate the mechanistic basis of the low proliferation of SLC25A38-null cells, we attempted a number of metabolic rescue experiments in low B6 (PN 1 nM). Formate (1 mM) was the only compound that enabled robust proliferation rescue (Fig. 5A), directly implicating an insufficient supply of one-carbon units in reduced growth of these cells in this media condition. Defective one-carbon metabolism due to low mitochondrial PLP is a plausible explanation for the proliferation defect observed in SLC25A38-KO cells in addition to, or independent from, any defects in mitochondrial glycine import (Fig. 2D). As previously implicated by the high delta gene score upon SHMT2 loss in our screen (Fig. 1F), impairment of serine hydroxymethyltransferase activity due to low mitochondrial PLP is consistent with these data. This PLP-dependent enzyme transfers one-carbon units from serine onto tetrahydrofolate (THF) forming CH2-THF and glycine as part of the folate cycle (Fig. 2D). Importantly, the folate cycle has cytosolic and mitochondrial branches, but most proliferating cells rely primarily on the mitochondrial pathway via SHMT214,41,42. To understand if and how the one-carbon pathway is affected in SLC25A38-KO cells, we used deuterated serine ([2,3,3-2H-Serine, or “D3-Serine”) to trace glycine ([2-2H-glycine, or “D1-Glycine”) synthesis and excretion onto the culture media. In most cells, serine is imported into the mitochondria via SFXN112 to provide substrate for SHMT2 to produce glycine in a PLP-dependent manner. In both B6 rich and low conditions, a reduction in D1-glycine excretion into the media of SFXN1-KO cells is seen whereas SHMT2-KO cells have no D1-glycine excretion (Fig. 5B). SLC25A38-KO cells displayed a drastic reduction of D1-Glycine excretion uniquely in low B6 conditions, which was rescued by expression of WT SLC25A38 (“A38-WT”) but not a CSA point mutant (p. Arg134Cys, here “A38-R134C”, Fig. 5B). We also traced intracellular glutathione (GSH) synthesis (which contains glycine) from D3-Serine (Fig. 5C), and observed similar labeling patterns as observed for excreted glycine. In agreement, whole cell extracts of SLC25A38-KO K562 cells cultured in “B6-physiol.” or B6-depleted conditions displayed high levels of serine (Supplementary Fig. 8A) and lower levels of glycine (Supplementary Fig. 8B), contributing to a large increase in the serine to glycine ratio in SLC25A38-KO cells compared to WT cells and SLC25A39-KO cells (Supplementary Fig. 8C).
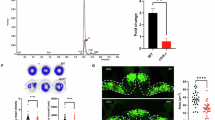
A Growth defect in B6-low can be rescued by addition of 1 mM formate (Form) but not glycine (Gly), serine (Ser), non-essential amino acids mix (NEAA), folate, 5-aminolevulinic acid (5-Ala), pyruvate or uridine. B Loss of SLC25A38 causes a glycine synthesis defect uniquely in low B6. 2,3,3-2H3-serine was used as a tracer to measure de novo synthesis of 2-2H-glycine and export to the culture media after 12 h using LC-MS. The glycine M + 0 species is the endogenous/unlabeled isotopologue and the glycine M + 1 species (2-2H-glycine) is derived from 2,3,3–2 H3-serine. C Glutathione (GSH) synthesized from glycine via glutathione synthetase is another indirect proxy for glycine levels; GSH M + 1 is derived from the de novo synthesized M + 1 glycine. Di Tracing 2,3,3-2H3-serine into thymidine 5’-triphosphate (TTP) can inform the contribution of cytosolic and mitochondrial one-carbon pathways12,14,43. If 2,3,3-2H3-serine is oxidized by mitochondrial SHMT2 and subsequent enzymes, a singly labeled formate species is formed leading to a one mass unit heavier: M + 1 TTP. In contrast, if 2,3,3-2H3-serine is oxidized by cytosolic SHMT1, a doubly labeled M + 2 TTP is formed. Dii SLC25A38-null cells rely more on cytosolic one-carbon pathways to TTP synthesis uniquely in low B6. SHMT2-null cells can only produce M + 2 TTP and SHMT1-null cells can only produce M + 1 TTP. SFXN1-null cells also display reversal of TTP synthesis towards the cytosolic route, independent of B6 status. E Purine synthesis intermediates GAR (5′-phosphoribosyl-glycinamide), SAICAR (phosphoribosylaminoimidazole-succinocarboxamide) and AICAR (5-aminoimidazole-4-carboxamide ribonucleotide) accumulate in SLC25A38-null, a phenotype rescuable by expression of SLC25A38 WT but not R134C. Box plots show min. to max. values with line at the median. Asterisks indicate significance as in Dunnet post-hoc tests, one-way ANOVA (A, comparing to PN 1 nM, and E, comparing to SLC25A38-KO +ev) and two-way ANOVA (B–D, comparisons with WT): *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, mean ± SD are shown for n = 3. Each n represents an independent culture of cells. Di, created in BioRender. Pena, I. (2025) https://BioRender.com/y18c741. Source data are provided as a Source Data file.
To assess compartmentalized glycine synthesis from serine, we used D3-Serine to trace 2H incorporation into thymidine triphosphate (TTP). This enables assessment of the relative contributions of SHMT1 (cytosolic) or SHMT2 (mitochondrial) to the generation of one-carbon units via serine-to-glycine conversion12,14,43. When deuterated serine is catabolized via the cytosolic pathway (SHMT1), it produces D2-TTP (labeled as “M + 2”). In contrast, when processed through the mitochondrial pathway (SHMT2), it results in D1-TTP (labeled as “M + 1”) (Fig. 5Di). In WT cells and SHMT1-KO cells, D1-TTP is the predominant form whereas SFXN1-KO and SHMT2-KO cells produce more D2-TTP as expected (Fig. 5Dii, “B6 low”). Notably, SLC25A38-KO cells exhibit similarly high levels of D1-TTP as WT cells under high B6 conditions (Fig. 5Dii, “B6 high”), suggesting that de novo glycine synthesis predominantly occurs via the mitochondrial SHMT2 pathway. However, under low vitamin B6 conditions, SLC25A38-KO cells show elevated levels of D2-TTP (Fig. 5Dii, “B6 low”), which can be rescued by the expression of WT SLC25A38, but not by the R134 point mutant. Also consistent with impaired one-carbon metabolism in SLC25A38-KO cells in physiological vitamin B6 levels, we observed accumulation of the purine synthesis intermediates 5′-phosphoribosyl-glycinamide (GAR), phosphoribosylaminoimidazole-succinocarboxamide (SAICAR), and 5-aminoimidazole-4-carboxamide ribonucleotide (AICAR) in SLC25A38-KO cells, a phenotype rescued by expression of the WT but not the R134C form of SLC25A38 (Fig. 5E). These results demonstrate impaired SHMT2 activity, likely due to low mitochondrial PLP uniquely in SLC25A38-KOs when cultured in low B6, and are consistent with a role of SLC25A38 in the PLP-dependent mitochondrial synthesis of glycine.
Metabolic changes in SLC25A38-KO cells are consistent with loss of mitochondrial PLP
In order to further distinguish cytosolic and mitochondrial SHMT functions, we knocked down SLC25A38 in WT, SHMT1 KO and SFXN1 KO cells to test whether loss of SLC25A38 induces an additional impairment in de novo D1-glycine synthesis and excretion of when traced from D3-Serine (Fig. 6A). We observed that, in high, supraphysiological B6 media, no major additional effects are seen when SLC25A38 is lost (Fig. 6A, “B6 high”).
A, left Tracing strategy to investigate if mitochondrially synthesized glycine M + 1 species (2-2H-glycine) can be produced and exported in SLC25A38-null cells in an SHMT2-dependent manner (in SHMT1−/− cells) and whether knocking out SLC25A38 in SFXN1-null cells leads to further impairment of glycine export. A, right Glycine production and export defect is driven by SLC25A38 only in low B6 conditions in WT, SFXN1−/− and SHMT1−/− cells without major effects in supraphysiological B6 (“B6 high”). Genetic block of mitochondrial serine uptake in SFXN1−/− cells or one-carbon metabolism in SHMT1−/− and SHMT2−/− cells does not cause mitochondrial PMP (B) or PLP (C) deficiency nor whole cell PLP changes. D Statistical significance as in Dunnet post-hoc tests comparing to WT levels for two way (A) or one way ANOVA (B–D) is shown: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Mean ± SD is shown for n = 3 in (B–D) and n = 2 in (A). HILIC metabolomics reveals a number of amino acid substrates of PLP-dependent enzymes accumulated in the mitochondria of SLC25A38-KO cells compared to WT in K562 (E) and Jurkat (F) (shown here in the log2 of the average fold change; p-value represents two-tailed t-tests, n = 3). PLP was measured using the same mitochondrial IPs by reverse phase chromatography (F5 column) and shown in red, as it cannot be detected using HILIC. G Seahorse analysis using the mitochondrial stress test assay indicates that expression of wild-type FLAG-SLC25A38 (“A38-WT”) but not the R134C point mutant restores oxygen consumption rates (OCR) in B6-low (1 nM PN); n = 6–8, data shown as mean + SEM. Here OCR was normalized by cell count and then fold changes were calculated compared to the WT levels. For all graphs, each n represents an independent culture of cells. A created in BioRender. Pena, I. (2025) https://BioRender.com/a96c224. Source data are provided as a Source Data file.
However, in low B6 media, SLC25A38-KO cells exhibited reduced D1-glycine excretion (Fig. 6A, “B6 low”). Importantly, de novo D1-glycine synthesis was completely abolished in the presence of the SHMT inhibitor Shin1 (2.5 µM) regardless of media and cell type, indicating that the labeling is specific and dependent on SHMT function (Supplementary Fig. 8G). These results suggest that at supraphysiological B6 levels, glycine can be synthesized from serine in the mitochondria via SHMT2, independent of SHMT1, and can be transported out of the organelle even in the absence of SLC25A38. However, at low B6 levels, SHMT2 impairment is uniquely observed in SLC25A38-KO cells, likely due to reduced mitochondrial PLP. This is also reflected in the B6-dependent sensitivity of K562 SLC25A38-KO cells to the SHMT inhibitor Shin1 (IC50s of 90.28 nM in low B6 versus 685 nM in high B6, Ext. Data Fig. 8H). Importantly, mitochondrial PLP and PMP levels did not decrease in SFXN1, SHMT1, or SHMT2 knockout cells grown in physiological B6 medium (10 nM PN) (Fig. 6B-D). This indicates that reduced mitochondrial or cytosolic one-carbon pathway flux (as in SFXN1 KO and SHMT1 KO, respectively) or the absence of flux (as in SHMT2 KO) does not regulate mitochondrial PLP levels. Similarly, no differential short-term proliferation defect was observed in SHMT1 KO, SHMT2 KO, and SFXN1 KO cells when comparing cultures in low or high B6 (Supplementary Fig. 8E), a difference uniquely seen in SLC25A38-KO cells. A glycine import function may not be essential in these cells, and/or alternative glycine import routes may exist in SLC25A38-KO cells, as also suggested to occur in yeast lacking Hem25p28. Supporting this hypothesis, mitochondrial swelling assays suggest that SLC25A38-KO mitochondria remain permeable to glycine compared to WT mitochondria, indicating that the influx of this amino acid still occurs (Supplementary Fig. 9).
Taken together, the observations that mitochondrially-synthesized glycine can be exported in SLC25A38-KO cells, and supraphysiological vitamin B6 rescues all phenotypes described here support a hypothesis that redundant glycine transport systems and other PLP accumulation mechanisms exist, independent of SLC25A38/Hem25p. As a first step to investigate these potential compensatory mechanisms, we performed proteomics profiling of mitochondria isolated from K562 WT versus SLC25A38-KO cells grown in high, supraphysiological B6 levels (Supplementary Fig. 10A–D) to investigate if other SLC25 family members are upregulated when SLC25A38 is missing. The only SLC25 family members statistically enriched in the SLC25A38-KO mitochondria were SLC25A33 (pyrimidine-nucleotide carrier), SLC25A12 (aralar 1, aspartate/glutamate exchanger) and SLC25A19 (thiamine pyrophosphate carrier) (Supplementary Fig. 10A, C). RNA levels for these proteins appear low in CSA disease-relevant cell types in mouse and humans such as early and proerythroblasts (Supplementary Fig. 10E, F), where both PLP and glycine in the mitochondria are necessary for ALAS2 activity, the first committed enzymatic step of heme biosynthesis. Further research will be needed to investigate the molecular basis of reduncancy and compensation in K562 and whether these targets may be relevant in a CSA disease context.
Given that PLP is crucial for pathways beyond one-carbon metabolism, we investigated whether other mitochondrial PLP-dependent enzymes were expressed in the cells under consideration. Proteomics profiling of mitochondria isolated from K562 WT and SLC25A38-KO cells (Supplementary Fig. 10A) revealed SHMT2, GOT2, OAT, BCAT2, NFS1, CCBL2, and GCAT as the most prominently expressed mitochondrial PLP-dependent enzymes in K562 mitoIPs (Supplementary Fig. 10C). Peptide intensity data indicated that SHMT2, GOT2, and OAT are the highest expressing enzymes which aligns with their high metabolic requirements in this cell type10. To determine if the substrates of these enzymes change in the absence of SLC25A38, we conducted an LC-MS study of mitochondria isolated from WT and SLC25A38-KO K562 (Fig. 6E) and Jurkat (Fig. 6F) cells cultured in B6 physiol. media. In addition to the low levels of PLP detected with our reverse phase LC-MS method, we observed an accumulation of several metabolites in mitochondria from SLC25A38-KO cells, which are suggestive of reduced activity of the PLP-dependent enzymes identified by proteomics (Fig. 6E, F). These metabolite observations include increased arginine and ornithine levels (substrates of ornithine aminotransferase, OAT), as well as of tryptophan, kynurenine and kynurenic acid (substrates of kynureninase (KYNU) and kynurenine aminotransferase (CCBL2)). Such increases are likely due to reduced enzyme activity rather than downregulation of these enzymes in SLC25A38-KO mitochondria (Supplementary Fig. 10C). For example, OAT is almost twofold upregulated in SLC25A38-KO mitochondria according to proteomic data (Supplementary Fig. 10C), yet its substrates ornithine and arginine show more than threefold increases, suggesting that the enzyme’s activity is reduced despite higher enzyme levels. In addition to the characteristic accumulation of ornithine and arginine44, OAT deficiency in humans also leads to low citrulline levels, another biomarker observed in K562 SLC25A38-KO cells (Fig. 6E).
Both mitochondrial iron-sulfur cluster assembly and heme biosynthesis also require PLP in the mitochondria (enzymes NSF1 and ALAS2, respectively, Fig. 2D). Given that both these cofactors are essential for the assembly and activity of the electron transfer complexes we tested whether respiration is affected in SLC25A38-KO. When cultured in B6-low conditions, we observed reduced oxygen consumption rates (OCR), stemming from reduced basal and maximal respiration, proton leak, and oligomycin-sensitive respiration as measured by Seahorse extracellular flux analysis in SLC25A38-KO cells cultured in B6-low, but not in in B6-rich media (Supplementary Fig. 8F). Mitochondrial respiration of SLC25A38-KO cells in low B6 could be rescued by expression of the WT form of SLC25A38 (“A38-WT”, Fig. 6G) but not when the mutant form p. Arg134Cys was expressed (“A38-R134C”, Fig. 6G). In summary, these metabolic changes observed upon SLC25A38 loss are consistent with reduced mitochondrial PLP levels and reduced activity of mitochondrially localized PLP-dependent enzymes. Together, these results may suggest that a number of PLP-dependent mitochondrial pathways are impaired in SLC25A38-KO cells in low, physiologically relevant B6 levels, consistent with a direct or indirect defect on mitochondrial PLP accumulation.
Discussion
Here we performed a CRISPRi-mediated genome-wide genetic screen based on the premise that deficiency in key PLP maintenance genes and PLP-dependent enzymes would slow or halt the proliferation of erythroleukemic cells when B6 levels are limited in the culture media. We found that all known enzymes involved in cytoplasmic B6 salvage (PDXK, PNPO, and PROSC) were conditionally essential in B6-depleted conditions (Figs. 1, 2), as well as known PLP-dependent enzymes necessary for K562 cell proliferation10 such as SHMT2, GOT2 and ODC1. It is important to note that different nutrient conditions and cell types will likely display varying requirements and essentialities for PLP-dependent pathways. Surprisingly, in both the genome-wide and targeted CRISPRi screens, the top conditionally essential gene in the B6-depleted condition was SLC25A38, encoding a mitochondrial glycine carrier. In line with the role of SLC25A38 in mitochondrial vitamin B6 metabolism, clonal K562 cell knockouts for SLC25A38 displayed low mitochondrial PLP levels (but not whole cell depletion) and defective growth in low levels of vitamin B6, matching the nanomolar concentrations seen in human plasma (Figs. 3, 4, Supplementary Fig. 3). These phenotypes could be rescued by the expression of wild-type SLC25A38, but not by point mutant versions found in patients with congenital sideroblastic anemia (CSA) (Fig. 3). Consistent with these observations, we failed to detect the accumulation of isotopically labeled PLP (D3-PLP) in the mitochondria of SLC25A38-KO cells (Fig. 4), which is likely explained by an unanticipated direct or indirect role in the import, accumulation, and/or retention of this vitamin. Taken together, our work highlights an unexpected role of SLC25A38 in the maintenance of mitochondrial PLP at low, physiological levels of vitamin B6.
Literature evidence for the roles of SLC25A38 on mitochondrial glycine transport comes from studies showing low heme levels in morphant zebrafish26,27 and reduced ALA in yeast lacking the ortholog Hem25p25,26,27, as well as transport assays executed using the Hem25p protein reconstituted in proteoliposomes25. Recent evidence suggests that Hem25 may also transport IPP28 in yeast, a function not observed in the human ortholog28. Alternative substrate usage as well as redundancy may explain why only 53 inner mitochondrial transporters (SLC25 family) are present in humans, despite the need to transport thousands of polar molecules into the mitochondrial matrix. These transporters may have additional direct or indirect roles yet to be characterized. Under the conditions tested here, loss of SLC25A38 led to low levels of glycine in K562 (Fig. 5 and Supplementary Fig. 8), but most likely due to impaired mitochondrial de novo synthesis of this amino acid via the PLP-dependent enzyme SHMT2 (Figs. 5, 6). However, SLC25A38 does not seem to be essential in high, supraphysiological B6 conditions, suggesting the presence of other compensatory mechanisms for PLP accumulation. Future studies are needed to identify proteins responsible for these compensatory functions. Our data also suggest the existence of a redundant transport system for glycine in K562/Jurkat cells that is independent of SLC25A38. Mitochondrially-synthesized deuterated glycine (D-glycine, from D3-serine) can still be exported to the culture media in SLC25A38-KO cells grown in supraphysiological B6 levels (Fig. 5A). Mitochondrial swelling assays (Supplementary Fig. 9) suggest that influx of glycine into SLC25A38-KO mitochondria is similar to WT mitochondria. Given that SLC25A38 is a highly erythroid-enriched gene26, a gene playing a redundant role in mitochondrial PLP accumulation must exist in other cell types. This gene is likely relevant for other tissues which are also dependent on vitamin B6, such as the brain, which is susceptible to severe B6-dependent epilepsy disorders when PLP homeostasis is disrupted (e.g. loss-of-function mutations in PLPBP23,29 (fka PROSC), ALDH7A145, and PNPO46). Identifying proteins with redundancy to SLC25A38 either on its glycine transport and/or a PLP accumulation function may lead to the identification of therapeutic targets for CSA, B6-dependent epilepsies, and other diseases related to B6 homeostasis. Future biochemical assays using proteoliposomes will be necessary to test precisely whether these new roles of SLC25A38 in mitochondrial PLP accumulation are due to direct or indirect transport functions and whether it interplay with its glycine carrier function.
Recent research in yeast has implicated Mtm1p32,47 in the mitochondrial binding of PLP. The high affinity of Mtm1p for PLP was suggestive that other proteins may be required to transfer PLP to its target proteins48. In yeast, no genes in the B6 pathway are synthetic lethal with mtm1Δ. Instead, gsh1 and gsh2, which encode the yeast glutamylcysteine ligase and GSH synthetase proteins, respectively, showed the highest synthetic lethality scores (Fig. 4H). The closest homolog of Mtm1p in humans is SLC25A39, recently identified as the mitochondrial glutathione transporter, with secondary effects on iron homeostasis30,31. Shi et al. 202449 and Liu et al.50 found that SLC25A39 acts as a sensor of mitochondrial iron-sulfur cluster levels via its matrix cysteine residues. Glutathione is crucial for mitochondrial iron-sulfur (Fe-S) cluster biosynthesis, making its import essential for iron homeostasis49. These studies did not report any deficiency in PLP import or other PLP-dependent pathways in SLC25A39 knockout50. We tested vitamin B6-dependent phenotypes in SLC25A39-null clonal lines through various experiments (e.g., Figs. 2–4, Supplementary Figs. 3 and 8). Our results showed that clonal K562 cells lacking SLC25A39 grew identically to wild-type cells under low B6 conditions (Fig. 2), with no decrease in mitochondrial PLP concentrations (Supplementary Fig. 3). Additionally, SLC25A39 was not essential in low B6 conditions in our whole-genome or targeted CRISPRi screens. The only condition where we observed a deficiency in mitochondrial PLP was upon the loss of SLC25A38. Expression of SLC25A39 in an SLC25A38-KO context did not rescue low mitochondrial PLP or PMP levels (Fig. 3B, C). Based on these recent studies and our observations, SLC25A39 does not appear to play a role in mitochondrial PLP regulation in K562 cells.
Loss-of-function mutations in SLC25A38 cause a severe form of congenital sideroblastic anemia (CSA)26,51. This hematological disorder is characterized by iron deposits in the mitochondria of erythroid precursors, forming ringed sideroblasts51. Early erythroid precursors significantly upregulate SLC25A3826,52, likely to meet the high demand of the PLP-dependent enzyme ALAS2 for heme biosynthesis. Given that the binding of ALAS2 to PLP is markedly slow52, mechanisms that regulate mitochondrial levels of this cofactor are crucial for its catalytic function. In yeast, loss of the ortholog Hem25p leads to reduced levels of 5-ALA, mitochondrial glycine, and heme, which were initially thought to result from decreased glycine import. In yeast, loss of the ortholog Hem25p leads to reduced levels of 5-ALA, mitochondrial glycine, and heme25, which were hypothesized to result from decreased glycine import. However, these phenotypes could additionally or instead be explained by depleted mitochondrial PLP affecting the activity of various B6-dependent enzymes such as Hem1p (ALAS homolog) and Shm2p (SHMT2 homolog), thereby blocking the synthesis of 5-ALA and mitochondrial glycine, respectively. Our data clearly show that SHMT2 activity is uniquely compromised in low B6 conditions when SLC25A38 is lost, impacting glycine levels (Figs. 5, 6). Low mitochondrial PLP, directly or indirectly due to SLC25A38 loss, may, therefore, affect iron incorporation in erythroblasts. This could be due to reduced de novo glycine synthesis and/or decreased ALAS2 activity leading to low heme synthesis, and potential effects on iron incorporation into mitochondrial iron-sulfur clusters via the PLP-dependent enzyme NFS1 (cysteine desulfurase) (Fig. 2D).
Prolonged pyridoxine deficiency in laboratory animals has long been known to cause reversible hypochromic, microcytic anemia with high serum iron53. A similar condition was observed in a child after months on a pyridoxine-deficient diet54. Recent studies by Ducamp et al.55 add significant in vivo relevance to our research. They developed several mouse models of CSA, including the first viable model for CSA-SLC25A38, by pan-hematopoietic deletion of Slc25a38 using Vav1-Cre (“Slc25a38BMKO”). Unexpectedly, these mice showed extreme hypersensitivity to a pyridoxine-restricted diet (0 and 2 ppm pyridoxine)55. However, when these animals were placed on a control synthetic diet (10 ppm pyridoxine), their anemia and microcytosis promptly improved. This response was not observed in a mouse model for another hereditary anemia dependent on heme (a-thalassemia intermedia (Hbbth3/+))55. These robust animal model findings align closely with our results and hypotheses. In our study, all the described phenotypes in SLC25A38-KO cells were rescuable by supraphysiological concentrations of vitamin B6. Both our work and Ducamp et al’s in vivo model demonstrate the conditional essentiality of vitamin B6 under SLC25A38 loss. Our findings of a role on mitochondrial PLP accumulation and PLP-dependent enzyme function may open therapeutic strategies for CSA-SLC25A38. Future clinical investigations with extended vitamin B6 trials (with or without combinations of glycine, serine, formate, folate, and other supplements) could activate compensatory mechanisms, potentially improving heme synthesis and anemia phenotypes. We believe our findings will have significant clinical implications for CSA diseases and beyond. Understanding how SLC25A38-KO cells adapt to mitochondrial PLP deficiency could uncover strategies for genetic and metabolic interventions to bypass its function, thus treating CSA and other disorders related to mitochondrial PLP-dependent functions. In conclusion, our work suggests a role for SLC25A38 in maintaining mitochondrial PLP, with implications for understanding the pathology of human diseases involving mitochondrial PLP-dependent enzymes.
Methods
Reagents
Reagents were obtained from the following sources: the antibodies that recognize SLC25A38 (ab133614), anti-ATP5a (Mouse, ab14748) and total OXPHOS Rodent WB Antibody Cocktail (ab110413) from Abcam, SLC25A39 (14963-1-AP), PDXK (15309-1-AP) and PROSC (25154–1-AP) from Proteintech, SHMT2 (HPA020549), SFXN1 (HPA019543) from Atlas Antibodies, GAPDH (2118), VDAC (4661), SHMT1 (12612), Citrate Synthase (CS) (14309), Calreticulin (CalR) (12238), Catalase (12980), Cathepsin B (31718), GOLGIN-97 (13132), myc (2278), DYKDDDDK Tag (14793) and hemagglutinin (HA) epitopes (3724) from Cell Signaling Technology, Tomm20 (HPA011562), FLAG (F1804) from MilliporeSigma and anti-HA raised in rat (11867423001) from Roche. All antibodies were prepared fresh in TBS-T 5% bovine serum albumin. The horseradish peroxidase (HRP)–coupled anti-rabbit and mouse secondary antibody, as well as normal donkey serum were from Cell Signaling Technology (CST); amino acids, glucose, glutamax, oligomycin, carbonyl cyanide p-trifluoromethoxyphenylhydrazone (FCCP), rotenone, antimycin, sodium azide, ADP, sodium formate, uridine, sodium pyruvate, hydroxylamine, biotin, choline, folate, myo-Inositol, niacinamide, p-Aminobenzoate, pantothenate, pyridoxine, riboflavin, thiamine and vitamin B-12 were from Sigma-Aldrich; Alexa 488-, Alexa 568-, and Alexa 642-conjugated secondary antibodies from Invitrogen; anti-HA magnetic beads were from ThermoFisher Scientific; Cell-Tak was from Corning. X-tremeGENE 9, Proteinase K and complete protease cocktail were from Roche; inactivated fetal bovine serum (IFS) from ThermoFisher (10438026); vitamin-free RPMI from US Biologicals; for LC–MS experiments, all solvents (including water) were purchased from ThermoFisher Scientific and were Optima LC–MS grade. Formic acid 99% HPLC–MS mobile phase component was purchased from ThermoFisher Scientific. RPMI 1640, DMEM, glutamax, pluronic, and penicillin–streptomycin solution were from ThermoFisher Scientific. Stellar Competent Cells, E. coli HST08 strain, were purchased from Takara for cloning purposes.
Cell lines and plasmids
K562 and Jurkat cells were obtained from the American Type Culture Collection (ATCC); K562 cells were used for all functional studies in cells unless indicated otherwise. HEK293T cells used in virus production were also obtained from the ATCC. The pMXs-IRES-Bsd backbone vector was obtained from Cell Biolabs. New plasmids generated in this study will be available through Addgene, see “Data Availability”. The CRISPRi pooled plasmid library (whole genome CRISPRi library, “hCRISPRi-v2” 10 guides/gene, #1000000092) used in this study as well as other plasmids were obtained from Addgene: CRISPRi/a v1 plasmid (pU6-sgRNA EF1Alpha-puro-T2A-BFP, #60955), pCRISPRia (#84832), lentiCRISPRv2 (#52961), psPAX2 (#12260), pCMV-VSV-G (#8454), Gag-Pol (#12371), pDONR221-SLC25A38 (#132053), pDONR221-SLC25A39 (#131970). Sequences of S. cerevisiae HEM25, and MTM1 were codon-optimized and cloned into pMXs for expression in human cells. SLC25A38 point mutants were cloned using KAPA HiFi DNA Polymerase in an oligonucleotide-directed mutagenesis protocol using In-Fusion HD Cloning Kit (Takara). The following oligos were used to generate each mutant construct:
Arg134Cys: Gggagtgggatcctgttctgtggcaggcgtgtg and Acaggatcccactcccagcatcacggactccaggg,
Arg134His: Gggagtgggatcccactctgtggcaggcgtgtg and Gtgggatcccactcccagcatcacggactccaggg, Asp188His: Gccacactgctgaggcacgcccctttctctggcatcta and Gtgcctcagcagtgtggcggtcagtccggagaacaggc,
Arg187Gln: Accgccacactgctgcaagacgcccctttctctggcatc and Ttgcagcagtgtggcggtcagtccggagaacaggcctc,
Arg282Ala: Cctcgcgccctgagggccaccctgatggcagcaatg and Ggccctcagggcgcgagggattcctccctgaaagaagc,
Arg278Ala: Gggaggaatccctgccgccctgaggcgcaccctgat and ggcagggattcctccctgaaagaagcccctcaggcc.
Cell culture
Unless otherwise indicated, Jurkat and K562 cells were cultured in RPMI media supplemented with 10% inactivated Fetal Calf Serum (IFS), 2 mM glutamine, and penicillin/streptomycin. HEK293 was cultured in high-glucose DMEM (25 mM) with 10% IFS and supplemented with 2 mM glutamine). Cells were maintained at 37 °C and 5% CO2. For B6-depletion conditions, we used a base medium from US Biologicals (R9010-07): “RPMI 1640 Medium Modified without L-Glutamine, Vitamins and Glucose powder mix”, supplemented with 5% of hydroxylamine-treated and dialyzed IFS, 10 mM glucose, 2 mM glutamax, Penicillin-Streptomycin, and a vitamin mix lacking B6 (prepared in house using biotin, choline, folate, myo-Inositol, niacinamide, p-Aminobenzoate, pantothenate, riboflavin, thiamine and vitamin B-12 (vitamin concentrations follow the 1× RPMI Vitamin mix, see Sigma R7256). Pyridoxine is then added as needed per experimental needs. Hydroxylamine treated serum was prepared following a procedure adapted from ref. 9 and dialyzed against 5 exchanges of PBS (10× volume).
Virus production and transduction of cell lines
HEK-293T cells were co-transfected with the pCRISPRia base plasmids, the vesicular stomatitis virus G (VSV-G) envelope plasmid, and psPAX2 packaging plasmid or with pMXS plasmids and retroviral packaging plasmids Gag-Pol and VSV-G, using X-tremeGENE 9 Transfection Reagent. Media was changed 24 h after transfection. The virus-containing supernatant was collected 48 h after transfection and passed through a 0.45 μm filter to eliminate cells and stored at −80 °C. For transduction of K562 cells, 106 cells were incubated with 50–1000 μl viral supernatant, 8 μg/ml polybrene, and 2 mL culture medium in a 6-well plate before spinning at 1200 g for 45 min at 37 °C. After an 18-h incubation, cells were pelleted to remove virus and were expanded into medium containing the selection agent 36–48 h post-transduction. Selection was maintained for 3–5 days before the use of the cells for experimental procedures. For CRISPRi experiments, we used K562s expressing nuclease-dead Cas9 (dCas9-KRAB) to programmably interfere with gene expression when co-expressing sgRNAs (CRISPRi) (gift from Jonathan Weismann’s lab, described in ref. 20,21).
CRISPRi genome-wide screen
The genome-scale CRISPRi screen was performed as previously described20,21 and as detailed below. In order to achieve >1000-fold library coverage, ~450 million K562s cells expressing nuclease-dead Cas9 (dCas9-KRAB) were transduced with the hCRISPRi-v2 library (whole genome CRISPRi library, 10 guides/gene, 209,070 guides total, Addgene #1000000092, 212,870 gRNA total) at a predetermined multiplicity of infection (MOI) < 1. Virus was removed 16 h after spinfection with the genome-wide library and selection started 48 h post infection, using 2 μg/mL puromycin for 3 days. Cells were grown in 1 L of media containing 0.1% Pluronic in 4 L flasks in agitation in Infors Mammalian cell incubator shaker at CO2 5% and 37 °C) at starting density of 250,000 cells/ml and expanded every 2 days. Following a 36 h recovery post puromycin selection, cells were counted and expanded into the two screen arms (B6-depleted medium (“B6-”) and B6-depleted medium supplemented with 4.8 μM pyridoxine (“B6+” or “B6-rich”), maintaining ~250 million cells each (>1000× coverage). On the same day, aliquots of ~55 million cells were stored for sequencing of the “day 0” time point. Cells were counted and re-seeded at 250,000 cells/ml (total 250 million per condition) in fresh respective media (B6- or B6+) every 2 days for a total of 14 cumulative doublings, after which final cell pellets were collected (~55 million cells/pellet). Each pellet contained at least 250-fold coverage of the library. Frozen samples of cells collected at day 0 and endpoint (day 14) were processed following established procedures21,56. Briefly, genomic DNA was extracted in a solution of 0.5 mg/mL Proteinase K in PBS at 70 °C for 24 h. Subsequently, the Qiagen QIAmp DNA Blood Maxi kit was used according to the manufacturer’s instructions. High-throughput sequencing libraries were prepared using established protocols, sampling enough genomic DNA for at least 250-fold coverage of the library for sgRNA amplification21 and the following primers: “CRISPRa_commonF” and each library using a different barcoded-index primer “Activ_CRISPR_index70”, “Activ_CRISPR_index71” or “Activ_CRISPR_index72”. QIAgen PCR purification kit was used to purify pooled PCR products and sequencing was performed on an Illumina HiSeq. Custom sequencing and indexing primer listed below (“CRISPRa_seq_2” and “CRISPRa_index2”) were used to perform a single-end sequencing run with a 6-base pair indexing read.
Activ_CRISPR_index70: CAAGCAGAAGACGGCATACGAGATCAAGGGAGAAGTTATAAACAGCACAAAAGGAAA
Activ_CRISPR_index71: CAAGCAGAAGACGGCATACGAGATCAAGTATGAAGTTATAAACAGCACAAAAGGAAA
Activ_CRISPR_index72: CAAGCAGAAGACGGCATACGAGATCACATCTGAAGTTATAAACAGCACAAAAGGAAA
CRISPRa_commonF: AATGATACGGCGACCACCGAGATCTACACCTCGGGGACTGTGGGCGATGTG
CRISPRa_seq_2: GTTGATAACGGACTAGCCTTATTTAAACTTGCTATGCTGTTTCCAGCTTAGCTCTTAAAC
CRISPRa_index2: TCAAAACACACAATTACTTTACAGTTAGGGTGAGTTTCCTTTTGTGCTGTTTATAACTTC
Sequencing reads were aligned to the sgRNA library and quantified using Python scripts (Python v3.7.0) from the Weissman lab described in refs. 21,56, Whitehead Institute (https://github.com/mhorlbeck/ScreenProcessing). Guide scores were calculated as log2(fold-change) from T0 (“day 0”) to endpoint samples (14 doublings in “-B6” or “+B6”) of normalized guide counts. Normalization was relative to the total read count. However, if there was a condition where 0 reads were found, a pseudocount of 1 was used (i.e. treat it as 1 read). Gene scores were calculated by taking the average across each gene’s guide scores. Guides were excluded from the gene score if there were fewer than 50 reads initially (“day 0” timepoint). p values were calculated using a Mann-Whitney U test comparing the distributions of guide scores for each condition. The final calculated gene and guide scores are shown in Supplementary Data 2 and Supplementary Data 3, respectively.
CRISPRi-targeted library screen
A targeted sub-library was designed by selecting the top-scoring genes in the B6 depletion screen described here, in order to provide cross-validation of the main genome-wide CRISPRi screen findings. Additionally, a curated list of known PLP-dependent enzyme genes and control sgRNAs were included. Ten sgRNAs per gene were selected from Addgene #1000000092 along with 750 control sgRNAs that were selected randomly from this library. In total, this targeted library contained 7500 sgRNAs and the full list of oligonucleotide sequences is described in Supplementary Data 4. The oligonucleotide pool was synthesized by Agilent Technologies and cloned as described in ref. 21. CRISPRi-targeted library screens were carried out exactly as described for genome-wide screens to achieve 1000× coverage over transduction and B6 depletion experiments and at least 250× coverage for gDNA isolation. Sequencing and analysis were executed as described previously and the table of results for both gene and guide scores are shown in Supplementary Datas 5 and 6, respectively.
CRISPR/Cas9-mediated generation of knockout cell lines
Human SLC25A38, SLC25A39, PROSC, and PDXK, were depleted using the lentiviral pLentiCRISPRv2 system. The following sense (S) and antisense (AS) oligonucleotides were cloned into pLentiCRISPRv2:
sgSLC25A38 (S): CACCGGCTTTCAGGTCTAGACGTGT
sgSLC25A38 (AS): AAACACACGTCTAGACCTGAAAGCC
sgSLC25A39 (S): CACCGCATGTGTCGTACCGGGAGCT
sgSLC25A39 (AS): AAACAGCTCCCGGTACGACACATGC
sgPROSC (S): CACCGGGACATGGGCAGCGCACTTT
sgPROSC (AS): AAACAAAGTGCGCTGCCCATGTCCC
sgPDXK (S): CACCGCTCCCCCTAAAGGTTATACG
sgPDXK (AS): AAACCGTATAACCTTTAGGGGGAGC
sgEGFP (S): CACCGGGGCGAGGAGCTGTTCACCG
sgEGFP (AS): AAACCGGTGAACAGCTCCTCGCCCC
The cloned plasmids were electroporated for transient expression in K562 cells using one million cells and 1 μg of sgRNA plasmid using an Amaxa Cell Line Nucleofector Kit V and an Amaxa Nucleofector II (Lonza). Cells were selected in 2 μg/mL puromycin for 3 days and then single-cell FACS sorted into 96-well plates. As a control, pLentiCRISPRv2-sgEGFP was electroporated in parallel, selected and also single-cell sorted. Cell clones with the desired knockouts were identified by western blotting (Supplementary Fig. 4A); in addition these cells no longer express the Cas9 protein and puroR cassette. For pooled knockout/knockdown studies shown in Fig. 6B and Supplementary Fig. 4B, we used pLentiCRISPRv2 cloned with the following guides:
SFXN1: GTTCCCATTCTCATCCGTGA
SLC25A38: GAGGAGCATCTATCACAGTG
Non-targeting control (NTC): CACCGTAACCGATACTCCCCACATT
Cell proliferation assays
20,000 cells per well were seeded into 96-well plates in triplicate. Presto Blue (ThermoFisher Scientific) was added to one plate immediately after seeding and absorbance was read at 570 nm using 600 nm as reference wavelength, following the supplier protocol, while a second plate was read-out 3 or 4 days after seeding. Number of doublings in 3 or 4 days was determined by calculating the log2 fold-change in signal between day 0 and 3 or 4. For metabolite rescue experiments, various concentrations of pyridoxine (PN), 1 mM formate, 1 mM sodium folate, 5 mM glycine, 5 mM serine, 5 mM 5-aminolevulinic acid (5-Ala), 1 mM pyruvate, 100 μg/ml uridine or 1× Non-Essential Amino Acids mix (NEAA, Gibco # 11140076) were added to B6-depleted media at the time of seeding unless indicated otherwise. When indicated, cells were pre-cultured in B6-depleted media with or without supplementation with PN prior to seeding.
Immunofluorescence assays
Cells adhered to glass coverslips coated with Poly-D-Lysine (ThermoFisher Scientific NC0672873), then fixed with 4% paraformaldehyde, followed by membrane permeabilization with TBS-T for 10 min and blocked with 2% normal donkey serum and 0.1% fish gelatin in TBS-T for 1 h at room temperature. When indicated, cells were incubated with the following primary antibodies: anti-SLC25A38 (Rabbit, Abcam EPBHMR1) 1:400, anti-Myc (Rabbit CST 2278T) 1:400, anti-HA (Rat, MilliporeSigma 11867423001) 1:400, and anti-ATP5a (Mouse, Abcam ab14748) 1:100 overnight at 4 °C and the following secondary antibodies: anti-rabbit Alexa Fluor 647 plus (ThermoFisher Scientific A32795), anti-rat Alexa Fluor 488 (ThermoFisher Scientific A110006) and anti-mouse Alexa Fluor 555 plus (ThermoFisher Scientific A32773) for 1 h at room temperature. Following incubation, three 10 min washes in TBS-T were performed, with the first including an incubation with DAPI 1:5000 to stain nuclei. Coverslips were mounted using Prolong Gold and imaged using a Leica SP8 LIGHTNING confocal microscope.
Mitochondrial isolation by “MitoIP” for metabolite and immunoblot analyses
Mitochondrial isolations by immunopurification were performed similarly to the described in refs. 17,18. Briefly, 30 × 106 K562 cells expressing the HA-MitoTag or a non-expressing control were washed 1x in PBS, 1x in KPBS. Cells were lysed in 1 ml KPBS with 30 syringe strokes using a 26G needle. Lysates were spun for 2 min at 1100 × g to pellet unbroken cells and nuclei, and 5 μl of the post-nuclear supernatant (input, “whole cell lysate”) was lysed in 1× RIPA Buffer containing protease inhibitors. The remaining whole cell lysate was subsequently incubated with 100 μl pierce anti-HA- magnetic beads for 3.5 min. Beads were washed using a magnet with 3× in KPBS, and mitochondria split into two samples: one lysed in 50 μl RIPA lysis buffer for 10 min and the other half was lysed in ice-cold trichloroacetic acid (TCA) solution (50 g/l) or 80% methanol (MeOH) containing labeled isotope standards for mass spectrometry, vortexing for 10 min in 4 °C. Beads were removed using a magnet, and samples were spun 10 min at 17,000 × g to remove residual beads and insoluble material. Western Blots were performed using NuPAGE™ 4–12%, Bis-Tris, 1.0 mm, Midi Protein Gels in the SureLock™ Tandem Midi Gel and Midi Blot Module following the manufacturer (ThermoFisher Scientific) protocols. Primary antibodies were used at 1:1000 dilution and secondary antibodies were used at 1:4000 dilution.
Hydrophilic interaction liquid chromatography (HILIC) LC–MS
Polar metabolite profiling was conducted on a QExactive benchtop Orbitrap mass spectrometer equipped with an Ion Max source and a HESI II probe, which was coupled to a Dionex UltiMate 3000 HPLC system (ThermoFisher Scientific). External mass calibration was performed using the standard calibration mixture every 7 days. Methods are described in detail elsewhere (27). Briefly, following extraction of whole cell lysate or Mito-IP samples in 80% MeOH spiked with internal standards, 5 μl of the sample was injected onto a SeQuant ZIC-pHILIC 150 × 2.1 mm analytical column equipped with a 2.1 × 20 mm guard column (both with a particle size of 5 µm; MilliporeSigma). Buffer A was 20 mM ammonium carbonate with 0.1% ammonium hydroxide and buffer B was acetonitrile. The column oven and autosampler tray were set at 25 °C and 4 °C, respectively. The chromatographic gradient was run at a flow rate of 0.150 ml min−1 as follows: 0–20 min, linear gradient from 80 to 20% B; 20–20.5 min, linear gradient from 20 to 80% B; and 20.5–28 min, linear gradient was maintained at 80% B. The mass spectrometer was operated in full-scan, polarity-switching mode, with the spray voltage set to 3.0 kV, the heated capillary set at 275 °C and the HESI probe set at 350 °C. The sheath gas flow was set to 40 units, the auxiliary gas flow was set to 15 units and the sweep gas flow was set to 1 unit. MS data acquisition was performed in the m/z range of 70–1000, with the resolution set at 70,000, the AGC target at 1 × 106 and the maximum injection time (Max IT) at 20 ms. Relative quantitation of polar metabolites was performed with XCalibur QuanBrowser 2.2 (Thermo Fisher Scientific) using a 5 p.p.m. mass tolerance and referencing an in-house library of chemical standards.
Reverse-phase LC–MS for B6 vitamer detection
The LC and mass spectrometer general settings were adapted to the Thermo Q Exactive benchtop Orbitrap mass spectrometer instrument from a previously described method18,57. Briefly, whole cell lysate or Mito-IP samples extracted with TCA (50 g/L) spiked with 500 nM internal standards (Metabolomics Amino Acid Mix Standard: Cambridge Isotope Laboratories, Inc.) and, when applicable, 1uM D3-pyridoxine (D3-PN) and/or 1uM D3-Pyridoxal 5’-phosphate (D3-PLP), both from Cambridge Isotope Laboratories, Inc. (DLM-9121-PK and DLM-9793-0.001). Typically, 5 μl of a sample was injected onto a Kinetex F5 - 2.6 u, 100 ×4.6 mm column (Phenomenex) for a 15 min run. This column was first used for successful binding of underivatized PLP enabling detection and quantification by an LC-MSMS method using a triple quadrupole instrument57. We adapted this method into an LC-MS protocol for the Orbitrap instrument. For the liquid chromatography, the mobile phase consisted of 0.1% formic acid in water (A) and 100% acetonitrile (B). The column oven was set to 30 °C, and the flow rate was 0.250 ml min−1. The gradient was as follows: 0–2 min, gradient held at 2% B; 2–7.5 min, linear gradient of 2–60% B; 7.5–8.5 min, linear gradient of 60–100% B; 8.55–10.5 min, wash in 100% B; 10.51–15 min, re-equilibration in 2% B. The data were acquired in positive-ion mode with a scan range of 70–1000 m/z. tSIM was included for endogenous and D3-labeled PLP, centered at m/z values of 248.0318 and 251.0511, respectively. Relative quantification of vitamin B6 metabolites was performed with XCalibur QuanBrowser (ThermoFisher Scientific) using a 5 ppm mass tolerance. The analytes included in this study as well as the experimentally measured exact masses (m/z) were the following: Pyridoxine (PN) (C8H11NO3): 170.0812; Pyridoxal (PL) (C8H9NO3): 168.0655; Pyridoxamine (PM) (C8H12N2O2): 169.0972; pyridoxal 5’-phosphate (PLP) (C8H10NO6P): 248.0318; pyridoxine 5’-phosphate (PNP) (C8H12NO6P): 250.0475; pyridoxamine 5’-phosphate (PMP) (C8H13N2O5P): 249.0635. MSMS validation spectra for the four B6 vitamers detected in our study are shown in Supplementary Fig. 1B. Spiked D3-PN was used as the internal standard to calculate area ratios for the endogenous PN, PL, and PM whereas D3-PLP was used as the internal standard for PLP, PMP, and PNP to control for efficiency of extraction, as well as to correct for any retention time shifts and to quality control the data.
Gas chromatography–mass spectrometry analysis of polar metabolites
Polar metabolites were analyzed by gas chromatography–mass spectrometry (GC–MS) as described previously58. Dried and frozen metabolite extracts were derivatized with 16 µl of MOX reagent (ThermoFisher Scientific, TS-45950) for 60 min at 37 °C, followed by derivatization with 20 µl of N-tert-butyldimethylsilyl-N-methyltrifluoroacetamide with 1% tert-butyldimethylchlorosilane (MilliporeSigma, 375934) for 2 h at 60 °C. Derivatized samples were analyzed by GC–MS, using a DB-35MS column (Agilent Technologies, 122–3832) installed in an Agilent 7890B gas chromatograph coupled to an Agilent 5997B mass spectrometer. Helium was used as the carrier gas at a constant flow rate of 1.2 ml min−1. One microliter of the sample was injected in split mode (1:10) at 270 °C. After injection, the GC oven was held at 100 °C for 1 min, increased to 105 °C at 2.5 °C per min, held at 105 °C for 2 min, increased to 250 °C at 3.5 °C per min, and then ramped to 320 °C at 20 °C per min. The MS system operated under electron impact ionization at 70 eV, and the MS source and quadrupole were held at 230 °C and 150 °C, respectively. The detector was used in scanning mode with an ion range of 100–650 m/z. Total ion counts were determined by integrating appropriate ion fragments for each metabolite59 using EL-MAVEN software (Elucidata). Mass isotopologue distributions were corrected for natural abundance using IsoCorrectoR59. Metabolite data were normalized to the internal standard and cell numbers.
Serine tracing assays
2,3,3-2H3-serine (“D3-serine”) was obtained from Cambridge Isotope Laboratories. Cells were cultured for 2 days in B6 low media (1 nM PN) or B6 rich media (1000 nM PN), then 2,3,3-2H3-serine was spiked to a final concentration of 2 mM. Cells were incubated for 12 h when media and cell extracts were prepared in 80% MeOH as described above for polar metabolite extractions. The fate of the deuteria were traced into glycine (D1-glycine, m/z 77.0455 ESI+ versus unlabeled glycine, m/z 76.0393 ESI+), glutathione (D1-GSH, m/z 309.0873 ESI + versus unlabeled GSH, m/z 308.0911 ESI+) and thymidine 5’-triphosphate (D1-TTP m/z 481.9886 ESI-, D2-TTP m/z 482.8847 ESI- versus unlabeled TTP, m/z 480.9771 ESI-). A diagram illustrating how tracing 2H3 into TTP can distinguish mitochondrial and cytosolic one-carbon pathways is shown in Fig. 5D (top). Briefly, if 2,3,3-2H3-serine is oxidized by mitochondrial SHMT2 and subsequent enzymes, a singly labeled formate species is formed, therefore one mass unit heavier: M + 1 TTP. In contrast, if 2,3,3-2H3-serine is oxidized by cytosolic SHMT1, a doubly labeled (two mass units heavier) M + 2 TTP is formed. By high-resolution LC-MS, the difference between unlabeled (M + 0). M + 1 and M + 2 TTP can be resolved, and the ratio of M + 1 to M + 2 indicates the mitochondria versus cytosol serine hydroxymethyltransferase (SHMT) activities. Positive controls for this experiment are SHMT2-null cells which can only produce M + 2 TTP and SHMT1-null cells which can only produce M + 1 TTP. SFXN1-null cells are also included as control for displaying the reversal of TTP synthesis towards cytosolic route (M + 2 TTP) independent of B6 status. Mass spectrometry data collection was performed as described above (HILIC LC/MS). For culture media analysis we calculated peak areas for labeled glycine following the m/z values described in positive ionization mode. For cell extracts, we detected labeled GSH and TTP following the m/z values described here, as in previous experiments12,14,43. Mass isotopomer distributions were corrected for natural abundance using in-house algorithms and using IsoCorrectoR as previously described59.
Tracing assay for mitochondrial PLP import
100 nM of isotopically labeled (2′,2′,2′)-2H3-pyridoxine (D3-PN, “M + 3”) were added to the cell culture media to trace the deuterated methyl groups into labeled PLP (“M + 3”, “D3-PLP”) in the mitochondrial fraction. 30 million cells were used per replicate, extraction of whole cell lysate and Mito-IP fractions were performed in TCA (50 g/L) spiked only with 500 nM internal standards (Metabolomics Amino Acid Mix Standard: Cambridge Isotope Laboratories, Inc.) and reverse phase LC-MS for detection of endogenous (M0) and heavy (M + 3) B6 vitamers was done as described earlier. Mass isotopomer distributions were corrected for natural abundance using in-house algorithms and using IsoCorrectoR as previously described59.
Mitochondrial swelling assay
Mitochondrial purifications were carried out using an altered previously described protocol60: 1 billion cells were spun down at 800 × g for 10 min, washed with cold pbs and resuspended in cold isolation buffer (250 mM sucrose, 10 mM Tris pH 7.5, 10 mM KCl, 1.5 mM MgCl2, 1 mM potassium EDTA). All further isolation steps were done at 4 °C. After incubating in isolation buffer for 30 min, the cells were lysed with a Dounce homogenizer using 30 strokes with a tight fit pestle. The lysate was centrifuged at 1300 × g for 5 min. Afterward, the supernatant was collected and the previous centrifugation step was repeated. The supernatant was then collected and centrifuged at 7000 × g for 15 min. The pelleted mitochondria were washed with isolation buffer, centrifuged again at 7000 × g for 15 min, and resuspended in isolation buffer. Swelling experiments were performed using an adapted protocol from previous studies61,62. Purified mitochondria were diluted to 2 mg/mL in a swelling buffer (20 mM Tris pH 7.5, 0.5 mM potassium EDTA, 10 μM rotenone) and then incubated at room temperature for 30 min. Mitochondria were then added to a 96-well plate and swelling was induced by adding an equal volume of swelling buffer containing 500 mM amino acid. The swelling response was recorded using a microplate reader measuring Absorbance at 520 nm every 10 s for 5 min. This assay provides a qualitative readout of transport based on the principal that mitochondria swell when suspended in isosmotic solutions of penetrating solutes63. Solute influx, in this case glycine, results in swelling which leads to a decrease in absorbance over time, and the response is proportional to the amount of transport occurring64.
Mitochondrial physiology
Oxygen consumption rates (OCR) of intact cells were measured using a Seahorse XFe96 Analyzer (Agilent Technologies). 250,000 cells were seeded on Seahorse XFe96 cell culture plates coated with Cell-Tak (Corning) following manufacturer protocols (Agilent Technologies) in 180 μl of media per each condition and assayed after incubation at 37 °C for 1 h. Three basal OCR measurements were taken, followed by sequential injections of 1.5 μM oligomycin, 2 μM FCCP, and 1 μM rotenone/antimycin A, taking three measurements following each treatment. After OCR data acquisition, average cell number per each assayed condition was measured for normalyzation.
mRNA quantification by qPCR
RNA was extracted from cells using the RNeasy kit according to the manufacturer’s instructions (Qiagen) and RNA reverse transcribed using the SuperScript VILO Master Mix (ThermoFisher Scientific). The PowerUp SYBR Master Mix (ThermoFisher Scientific) and the following primers were used to assess mRNA levels of SLC25A38 by quantitative PCR (qPCR) by normalizing Ct values to those of β-Actin (ACTB). SLC25A38: gacctgctctaccctccttt and cccaacacgtctagacccat; ACTB: AGGATGGCAAGGGACTTCCTG and AATGTGGCCGAGGACTTTGAT.
Protein digestion for proteomics
Following mitoIP, eluted mitochondrial proteins were separated on 4–20% precast SDS gels (Biorad, USA) followed by staining with Coomassie blue R-250 staining solution (Bio-Rad) for 1 h and de-stained with water for 2–3 h. The gel was further rinsed with ultrapure water three times and gel slices were cut into small pieces and washed with water followed by 20 min wash with 50% acetonitrile (ACN)/50 mM NH4HCO3 (ammonium bicarbonate, ABC). Proteins were reduced by the addition of 100 µL 4.5 mM dithiothreitol (DTT) in 50 mM ABC and incubated at 37 °C for 45 min. The DTT solution was removed, and the samples were cooled to room temperature. The samples were alkylated by the addition of 100 µL 10 mM iodoacetamide (IAA) in 50 mM ABC with incubation at room temperature in the dark for 20 min. The IAA solution was removed, and the gels were washed for 20 min with 50% ACN/50 mM ABC, then washed for 20 min with 50% ACN/25 mM ABC. The gels were resuspended in 100 µL of 50 mM ABC containing 500 ng of digestion-grade trypsin (Promega, V5111), and incubated at 37 °C for 16 h. The supernatants containing the tryptic peptides were transferred to new Eppendorf tubes. The residual peptides in the gel bands were extracted with 250 µL 80% ACN/0.1% formic acid for 15 min, then combined with the original digests, dried in a SpeedVac and desalted using Nest Group C18 microspin columns as per manufacturer’s instructions. The desalted peptides were further dried in a speedvac and used for subsequent MS analysis.
Proteomics mass spectrometry data acquisition and analysis
The desalted lyophilized peptides were dissolved in a loading buffer (2% ACN, 0.2% trifluoroacetic acid) and subjected to LC-MS/MS. The data acquisition was performed on a Thermo Scientific Q Exactive Plus equipped with a Waters nanoAcquity UPLC system utilizing a binary solvent system (A: 100% water, 0.1% formic acid; B: 100% ACN, 0.1% formic acid). The peptides were subsequently separated in a 2–90% ACN gradient using an ACQUITY UPLC M-Class Peptide BEH C18 Column (130 Å, 1.7 µm, 75 µm × 250 mm; Waters, #186007484) at a flow rate of 300 nL/min over a 180 min gradient. Eluting peptides were directly injected into the Q Exactive Plus. Data-dependent acquisition was used in positive ion mode. Full scan MSI data were acquired for mass range 350–1700 m/z, 45 ms maximum injection, and an AGC target of 3e6 at 70,000 resolution. MS2 fragmentation was performed on charge states from 2 to 6 with a 30 s dynamic exclusion, and all MS2 data was collected in centroid mode with an intensity threshold of 1E4. For proteomics analysis, mass spectrometry raw files were searched against a Uniprot human database (6,563,750 entries) using MaxQuant version (2.03.01) and the Andromeda search engine. Carbamidomethylation was set as fixed modification, and N-terminal acetylation and oxidation of methionine were used as variable modifications. A maximum of two Trypsin/P missed cleavages, a tolerance of 10 ppm for precursor mass, a tolerance of 0.5 Da for fragment ions, and match between runs were set for protein search. A maximum of 1% FDR was set for both protein and peptide identification. Protein quantification was performed using the LFQ algorithm and 7 min window for match between runs in MaxQuant software. The protein identifications assigned as contaminants and reverse proteins were removed from the dataset.
Bioinformatic analysis
Python scripts were used as indicated for CRISPRi analysis. The Chopchop online tool was used to design sgRNAs for the generation of knockout lines65. BLAST searches were employed to map orthologous proteins across yeast and human genomes. Yeast genetic interaction scores and p-values shown on Fig. 4E–H were obtained from The Cell Map database38, mined using an online tool, thecellmap.org39. The p-value calculated in ref. 38 is described as a statistical confidence measure assigned to each gene interaction pair based on observed variation over four experimental replicates compared to single gene fitness effects.
SLC25A38 structure was modeled using AlphaFold66. The Avana, Broad, and GeCKO OGEE Database version 367 datasets identified gene essentiality via genetic screens in K562 cells, and a gene was considered essential if at least one of the three datasets labeled it as essential (Supplementary Fig. 2). Subcellular localization data for Supplementary Data 1 was mined using the Gene Ontology database, GeneCards, MitoCarta 3.0 and literature searches.
Confocal image quantification studies were performed using Cell Profiler68,69. Briefly, individual cell nuclei were identified using a three-class Otsu’s method. Cell nuclei objects were then expanded via propagation using the HA and ATP5A channels in succession (Otsu’s two-class method) to delineate the cell boundary. Mitochondrial objects were then identified using the HA and ATP5A channels (Otsu’s two-class method). All object identification was done with global thresholding. All mitochondrial objects within a single cell boundary were then merged to single objects for further analysis. We then measured signal intensity, channel-channel intensity correlations (threshold intensity of 5 percent), and object size across Z-stack planes for all identified cells. We then used a script written in R to calculate overall signal intensity and correlation values for individual cells in each image.
For the proteomics study, the label-free quantitation (LFQ) intensities were log2 transformed in Perseus version 2.0.7.0 and used in subsequent analysis. R scripts using the packages pOmics2 and Tidyverse70 to perform data cleaning, missing data imputation, and normalization. We considered proteins where the ratio between WT MitoIP or SLC25A38-KO MitoIP to MockIP was larger than 2. Both the raw and processed/filtered LFQ are shown in Supplementary Data 7 and Source Data.
Statistical analysis
Differences between groups were tested for statistical significance by parametric and non-parametric tests such as Analysis of Variance (ANOVA) followed by appropriate post-hoc multiple comparisons. Two-tailed t tests were used for comparison between two groups. All comparisons were two-sided, and p values of less than 0.05 were considered to indicate statistical significance. All error bars denote standard deviations between biological replicates unless indicated otherwise. Most plots described here were generated using GraphPad PRISM 10.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data in this study are available within the manuscript, Figures, Supplementary Figs., Supplementary Data, and Source Data files. The full tables reporting our genetic screen (CRISPRi) results as well as primers used for the targeted library construction are shown in Supplementary Data; Source data are provided with this paper. Mass spectrometry proteomics raw data have been deposited to the ProteomeXchange Consortium via the PRIDE71 partner repository with the dataset identifier PXD058312. The following plasmids generated in this study will be available via Addgene: pMXs_FLAG-SLC25A38 (#218412), pMXs_FLAG-SLC25A39 (#218413), pMXs-FLAG-SLC25A38-Arg134Cys (#218416), pMXs-FLAG-SLC25A38-Arg134His (#218417), pMXs-FLAG-SLC25A38-Asp188His (#218418), pMXs-FLAG-SLC25A38-Arg187Gln (#218419), pMXs-FLAG-SLC25A38-Arg282Ala (#218420), pMXs-FLAG-SLC25A38-Arg278Ala (#218421), pMXs-3xMyc-hem25 (#218414) and pMXs-bleoR-MitoTag (#218415). Source data are provided with this paper.
References
Percudani, R. & Peracchi, A. The B6 database: a tool for the description and classification of vitamin B6-dependent enzymatic activities and of the corresponding protein families. BMC Bioinform. 10, 273 (2009).
Wilson, M. P., Plecko, B., Mills, P. B. & Clayton, P. T. Disorders affecting vitamin B6 metabolism. J. Inherit. Metab. Dis. 42, 629–646 (2019).
van Karnebeek, C., Pena, I. A. & Gospe, S. M. Disorders of pyridoxine metabolism. In (Rosenberg, R. N. & Pascual, J. M. eds.) Rosenberg’s Molecular and Genetic Basis of Neurological and Psychiatric Disease (Sixth Edition) 711–728 (Academic Press, 2020).
Chelban, V. et al. PDXK mutations cause polyneuropathy responsive to pyridoxal 5’-phosphate supplementation. Ann. Neurol. 86, 225–240 (2019).
Waymire, K. G. et al. Mice lacking tissue non–specific alkaline phosphatase die from seizures due to defective metabolism of vitamin B–6. Nat. Genet. 11, 45–51 (1995).
Millán, J. L. & Whyte, M. P. Alkaline phosphatase and hypophosphatasia. Calcif. Tissue Int. 98, 398–416 (2016).
Cotter, P. D., Baumann, M. & Bishop, D. F. Enzymatic defect in “X-linked” sideroblastic anemia: molecular evidence for erythroid delta-aminolevulinate synthase deficiency. Proc. Natl. Acad. Sci. USA 89, 4028–4032 (1992).
Skovby, F. Homocystinuria. Clinical, biochemical and genetic aspects of cystathionine beta-synthase and its deficiency in man. Acta Paediatr. Scand. Suppl. 321, 1–21 (1985).
Lipson, M. H., Kraus, J. & Rosenberg, L. E. Affinity of cystathionine beta-synthase for pyridoxal 5’-phosphate in cultured cells. A mechanism for pyridoxine-responsive homocystinuria. J. Clin. Investig. 66, 188–193 (1980).
Chen, C.-C. et al. Vitamin B6 addiction in acute myeloid leukemia. Cancer Cell 37, 71–84.e7 (2020).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Kory, N. et al. SFXN1 is a mitochondrial serine transporter required for one-carbon metabolism. Science 362, (2018).
Pikman, Y. et al. Targeting MTHFD2 in acute myeloid leukemia. J. Exp. Med. 213, 1285–1306 (2016).
Ducker, G. S. et al. Reversal of cytosolic one-carbon flux compensates for loss of the mitochondrial folate pathway. Cell Metab. 24, 640–641 (2016).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Gori, A. M. et al. Predictors of vitamin B6 and folate concentrations in older persons: the InCHIANTI study. Clin. Chem. 52, 1318–1324 (2006).
Chen, W. W., Freinkman, E. & Sabatini, D. M. Rapid immunopurification of mitochondria for metabolite profiling and absolute quantification of matrix metabolites. Nat. Protoc. 12, 2215–2231 (2017).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337.e11 (2016).
Lui, A., Lumeng, L. & Li, T. K. Metabolism of vitamin B6 in rat liver mitochondria. J. Biol. Chem. 256, 6041–6046 (1981).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. Elife 5, (2016).
Pena, I. A. et al. Pyridoxine-dependent epilepsy in zebrafish caused by Aldh7a1 deficiency. Genetics 207, 1501–1518 (2017).
Johnstone, D. L. et al. PLPHP deficiency: clinical, genetic, biochemical, and mechanistic insights. Brain 142, 542–559 (2019).
Cotter, P. D., Rucknagel, D. L. & Bishop, D. F. X-linked sideroblastic anemia: identification of the mutation in the erythroid-specific delta-aminolevulinate synthase gene (ALAS2) in the original family described by Cooley. Blood 84, 3915–3924 (1994).
Lunetti, P. et al. Characterization of human and yeast mitochondrial glycine carriers with implications for heme biosynthesis and anemia. J. Biol. Chem. 291, 19746–19759 (2016).
Guernsey, D. L. et al. Mutations in mitochondrial carrier family gene SLC25A38 cause nonsyndromic autosomal recessive congenital sideroblastic anemia. Nat. Genet. 41, 651–653 (2009).
Fernández-Murray, J. P. et al. Glycine and folate ameliorate models of congenital sideroblastic anemia. PLoS Genet. 12, e1005783 (2016).
Tai, J. et al. Hem25p is required for mitochondrial IPP transport in fungi. Nat. Cell Biol. 25, 1616–1624 (2023).
Darin, N. et al. Mutations in PROSC disrupt cellular pyridoxal phosphate homeostasis and cause vitamin-B6-dependent epilepsy. Am. J. Hum. Genet. 99, 1325–1337 (2016).
Wang, Y. et al. SLC25A39 is necessary for mitochondrial glutathione import in mammalian cells. Nature 599, 136–140 (2021).
Shi, X. et al. Combinatorial GxGxE CRISPR screen identifies SLC25A39 in mitochondrial glutathione transport linking iron homeostasis to OXPHOS. Nat. Commun. 13, 2483 (2022).
Whittaker, M. M., Penmatsa, A. & Whittaker, J. W. The Mtm1p carrier and pyridoxal 5’-phosphate cofactor trafficking in yeast mitochondria. Arch. Biochem. Biophys. 568, 64–70 (2015).
Kunji, E. R. S. Structural and mechanistic aspects of mitochondrial transport proteins. In (Egelman, E. H. ed.) Comprehensive Biophysics 174–205 (Elsevier, 2012).
Kunji, E. R. S. & Robinson, A. J. The conserved substrate binding site of mitochondrial carriers. Biochim. Biophys. Acta 1757, 1237–1248 (2006).
Robinson, A. J. & Kunji, E. R. S. Mitochondrial carriers in the cytoplasmic state have a common substrate binding site. Proc. Natl. Acad. Sci. USA 103, 2617–2622 (2006).
Heeney, M. M. et al. SLC25A38 congenital sideroblastic anemia: phenotypes and genotypes of 31 individuals from 24 families, including 11 novel mutations, and a review of the literature. Hum. Mutat. 42, 1367–1383 (2021).
Kannengiesser, C. et al. Missense SLC25A38 variations play an important role in autosomal recessive inherited sideroblastic anemia. Haematologica 96, 808–813 (2011).
Costanzo, M. et al. A global genetic interaction network maps a wiring diagram of cellular function. Science 353 (2016).
Usaj, M. et al. Thecellmap.org: a web-accessible database for visualizing and mining the global yeast genetic interaction network. G3 7, 1539–1549 (2017).
Stolz, J. & Vielreicher, M. Tpn1p, the plasma membrane vitamin B6 transporter of Saccharomyces cerevisiae. J. Biol. Chem. 278, 18990–18996 (2003).
Tibbetts, A. S. & Appling, D. R. Compartmentalization of mammalian folate-mediated one-carbon metabolism. Annu. Rev. Nutr. 30, 57–81 (2010).
Lewis, C. A. et al. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell 55, 253–263 (2014).
Herbig, K. et al. Cytoplasmic serine hydroxymethyltransferase mediates competition between folate-dependent deoxyribonucleotide and S-adenosylmethionine biosyntheses. J. Biol. Chem. 277, 38381–38389 (2002).
de Sain-van der Velden, M. G. M. et al. The proline/citrulline ratio as a biomarker for OAT deficiency in early infancy. JIMD Rep. 6, 95–99 (2012).
Mills, P. B. et al. Mutations in antiquitin in individuals with pyridoxine-dependent seizures. Nat. Med. 12, 307–309 (2006).
Mills, P. B. et al. Neonatal epileptic encephalopathy caused by mutations in the PNPO gene encoding pyridox(am)ine 5’-phosphate oxidase. Hum. Mol. Genet. 14, 1077–1086 (2005).
Whittaker, M. M. & Whittaker, J. W. Expression and purification of recombinant Saccharomyces cerevisiae mitochondrial carrier protein YGR257Cp (Mtm1p). Protein Expr. Purif. 93, 77–86 (2014).
Yien, Y. Y. & Perfetto, M. Regulation of heme synthesis by mitochondrial homeostasis proteins. Front. Cell Dev. Biol. 10, 895521 (2022).
Shi, X. et al. Dual regulation of SLC25A39 by AFG3L2 and iron controls mitochondrial glutathione homeostasis. Mol. Cell 84, 802–810.e6 (2024).
Liu, Y. et al. Autoregulatory control of mitochondrial glutathione homeostasis. Science 382, 820–828 (2023).
Bottomley, S. S. Sideroblastic anaemia. Clin. Haematol. 11, 389–409 (1982).
Kardon, J. R. et al. Mitochondrial ClpX activates a key enzyme for heme biosynthesis and erythropoiesis. Cell 161, 858–867 (2015).
Cartwright, G. E. Dietary factors concerned in erythropoiesis. Blood 2, 256–298 (1947).
Snyderman, S. E., Holt, L. E. Jr, Carretero, R. & Jacobs, K. Pyridoxine deficiency in the human infant. J. Clin. Nutr. 1, 200–207 (1953).
Ducamp, S. et al. Murine models of erythroid 5ALA synthesis disorders and their conditional synthetic lethal dependency on pyridoxine. Blood https://doi.org/10.1182/blood.2023023078 (2024).
Adelmann, C. H., Wang, T., Sabatini, D. M. & Lander, E. S. Genome-wide CRISPR/Cas9 screening for identification of cancer genes in cell lines. Methods Mol. Biol. 1907, 125–136 (2019).
Pena, I. A. et al. Simultaneous detection of lysine metabolites by a single LC-MS/MS method: monitoring lysine degradation in mouse plasma. Springerplus 5, 172 (2016).
Metallo, C. M. et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 481, 380–384 (2011).
Heinrich, P. et al. Correcting for natural isotope abundance and tracer impurity in MS-, MS/MS- and high-resolution-multiple-tracer-data from stable isotope labeling experiments with IsoCorrectoR. Sci. Rep. 8, 17910 (2018).
Li, W., Zhang, C. & Sun, X. Mitochondrial Ca2+ retention capacity assay and Ca2+-triggered mitochondrial swelling assay. J. Vis. Exp. https://doi.org/10.3791/56236 (2018).
Meyer, J. Proline transport in rat liver mitochondria. Arch. Biochem. Biophys. 178, 387–395 (1977).
Beavis, A. D., Brannan, R. D. & Garlid, K. D. Swelling and contraction of the mitochondrial matrix. I. A structural interpretation of the relationship between light scattering and matrix volume. J. Biol. Chem. 260, 13424–13433 (1985).
LaNoue, K. F. & Schoolwerth, A. C. Metabolite transport in mitochondria. Annu. Rev. Biochem. 48, 871–922 (1979).
Benavides, J., Garcia, M. L., Lopez-Lahoya, J., Ugarte, M. & Valdivieso, F. Glycine transport in rat brain and liver mitochondria. Biochim. Biophys. Acta 598, 588–594 (1980).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Gurumayum, S. et al. OGEE v3: online GEne essentiality database with increased coverage of organisms and human cell lines. Nucleic Acids Res. 49, D998–D1003 (2021).
Stirling, D. R., Carpenter, A. E. & Cimini, B. A. CellProfiler Analyst 3.0: accessible data exploration and machine learning for image analysis. Bioinformatics 37, 3992–3994 (2021).
Stirling, D. R. et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinform. 22, 433 (2021).
Wickham, H., Grolemund, G. R for Data Science: Import, Tidy, Transform, Visualize, and Model Data (“O’Reilly Media, Inc., 2016).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Whyte, M. P. et al. Alkaline phosphatase: placental and tissue-nonspecific isoenzymes hydrolyze phosphoethanolamine, inorganic pyrophosphate, and pyridoxal 5’-phosphate. Substrate accumulation in carriers of hypophosphatasia corrects during pregnancy. J. Clin. Investig. 95, 1440–1445 (1995).
Ciapaite, J. et al. Maintenance of cellular vitamin B6 levels and mitochondrial oxidative function depend on pyridoxal 5’-phosphate homeostasis protein. J. Biol. Chem. 299, 105047 (2023).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010).
Acknowledgements
We thank all the members of the Heiman lab for helpful insights and support; Prof. Jackie Lees for productive discussions and feedback; Dr. Fiona Fitzpatrick for insightful discussions on the SLC25 family of proteins; the Weissman lab at the Whitehead Institute for kindly sharing their K562 CRISPRi cell line expressing dCas9-KRAB; the Kory lab at Harvard School of Public Health for kindly sharing Jurkat clonal cell lines knockout for SLC25A38 and SFXN1; Drs Mark Fleming, Sarah Ducamp and Naama Kanarek for feedback and discussions. Drs. Genya Frenkel and Raghu Chivukula for support on the genome-wide CRISPRi screening; We thank T. Kunchok from the Whitehead Institute Metabolite Profiling Facility for HILIC metabolomics sample runs and support with our reverse phase method. We thank the Whitehead Institute FACS facility and the Whitehead Institute Genome Technology Core for services, instrumentation, and technical assistance. Dr Michael DeMott and the Bioimaging and Chemical Analysis Facilities Core at MIT for technical support. This work was made possible due to the generous support from numerous research funding agencies: Picower Institute Innovation Fund award from the Freedom Together Foundation (M.H.), a Pew Latin American Fellowship from the Pew Charitable Trust (I.A.P.), and a postdoctoral fellowship from the Hereditary Disease Foundation (I.A.P.). The authors also acknowledge support from T32GM007287 (S.D.B.), F30CA268633 (S.M.C.), a Medical Scientist Training Program grant (T32GM007753 and T32GM144273) from the National Institute of General Medical Sciences (P.G.), and R35CA242379, P30 CA014051, the Ludwig Center at MIT, MIT Center for Precision Cancer Medicine (M.G.V.H.), NIH F31 CA228241-01 (C.H.A.), NIH DA018343 (A.C.N.), NINDS F31 NS127458 (P.G.), Damon Runyon-Rachleff Innovation Award (73S-22, N.K.), and Damon Runyon Cancer Research Fellowship (DG-2454-22, C.H.A.).
Author information
Authors and Affiliations
Contributions
Conceptualization: I.A..P., M.H. Methodology: I.A.P., M.H., S.M.C., J.S., C.A.L., A.N. Investigation: I.A.P., J.S., S.M.C., S.B., H.K., J.Y., C.H.A., C.A.L., P.G., I.W., G.S., S.B., N.K. Visualization: I.A.P., J.S., S.M.C., P.G. Funding acquisition: I.A.P., M.H., D.S. Project administration: I.A.P., M.H. Supervision: I.A.P., M.H., N.K., M.G.V.H. Writing – original draft: I.A.P., M.H., M.G.V.H. Writing – review & editing: all authors.
Corresponding authors
Ethics declarations
Competing interests
M.G.V.H. discloses that he is an advisor for Agios Pharmaceuticals, iTeos Therapeutics, Sage Therapeutics, Pretzel Therapeutics, Droia Ventures, MPM Capital, and Auron Therapeutics. G.S. is currently an employee and shareholder of AstraZeneca. All other authors declare that they have no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pena, I.A., Shi, J.S., Chang, S.M. et al. SLC25A38 is required for mitochondrial pyridoxal 5’-phosphate (PLP) accumulation. Nat Commun 16, 978 (2025). https://doi.org/10.1038/s41467-025-56130-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56130-3