Abstract
Orphan crops are important sources of nutrition in developing regions and many are tolerant to biotic and abiotic stressors; however, modern crop improvement technologies have not been widely applied to orphan crops due to the lack of resources available. There are orphan crop representatives across major crop types and the conservation of genes between these related species can be used in crop improvement. Machine learning (ML) has emerged as a promising tool for crop improvement. Transferring knowledge from major crops to orphan crops and using machine learning to improve accuracy and efficiency can be used to improve orphan crops.
Similar content being viewed by others
Introduction
Orphan crops, also known as “minor”, “neglected”, “underutilised”, and “understudied” crops are frequently grown in developing countries and are an important source of nutrition for local communities (Table 1)1,2,3,4,5,6. Many of these crops have not benefited from the Green Revolution that improved the productivity of major crops such as wheat and rice. Modern crop improvement techniques, such as marker assisted breeding (MAB) and genome editing, have not been widely applied to orphan crops due to the lack of resources available. However, genomic technologies offer significant potential for orphan crop improvement. The major crops such as wheat, rice, maize and soybean are widely distributed and produced on an industrial scale, whereas the orphan crops vary considerably in their production, from the reasonably wide distribution of sorghum, to crops that are only produced in specific regions such as Ensete. Many orphan crops are tolerant of abiotic and biotic stressors and can be produced in marginal and harsh environments, possessing traits, that if understood, may be transferrable to major crops.
There are currently relatively few genomic resources for orphan crops, though initiatives such as the African Orphan Crops Consortium (AOCC)7 and Crops for the Future (CFF)8 are working towards their improvement. However, there is still much work needed to translate these resources to improve crops. There are orphan crop representatives across major crop types and the relatedness between orphan and major crops can be used to improve crop breeding efforts in both orphan and major crops. Major crops have a wide range of resources available, and this can be transferred to related orphan crop species. Orthologs of agronomically important genes from major crops have been found in orphan crops for traits such as stress tolerance9, and conservation of gene function between major and orphan crops can support their improvement. Similarly, knowledge of novel beneficial genes in orphan crops could be used to enhance traits in the major crops. A major challenge in crop improvement is the continued growth of data. Machine learning based methods are starting to be applied for crop improvement (Fig. 1) and these will have direct applications for orphan crop improvement as well as the translation of knowledge from major crops to orphan crops.
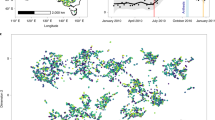
The proportion of articles with the search terms ‘machine learning crops’ (orange) and ‘orphan crops’ (blue) is shown on the Y-axis and the year the papers were published is shown on the X-axis.
In the past 100 years, large gains in crop yield have been made possible by the introduction of statistical methods into plant breeding. R. A. Fisher pioneered statistical methods such as ANOVA and randomised control trials in plant breeding10,11. Since then, statistical approaches have been at the core of plant breeding leading to unprecedented increases in crop yields. These methods include RR (Ridge Regression), BLUP (Best Linear Unbiased Prediction) and its variants such as GBLUP (Genomic Best Linear Unbiased Prediction), all of which fall under the broader category of genomic selection. However, yields are not keeping pace with a growing population and the threat of climate change. To ensure sufficient food production for a warmer world, modern approaches such as CRISPR genome editing and machine learning are needed. Machine learning (ML) is a set of methods that uses large amounts of data to approximate mathematical functions. Deep learning (DL), a subset of ML, utilises deep layers of artificial neural networks to “learn” mathematical functions from training data. ML’s ability to identify complex patterns within large and diverse datasets, from images and genomics to tabular data, makes it a powerful tool for improving trait prediction accuracy and crop breeding efficiency12,13,14,15 (Fig. 2).
Data inputs are shown on the left, including “genome sequencing” at multiple depths from whole-genome sequencing to exome and SNP sequencing. Genomic datasets can be combined with “+ phenotype observations” collected manually or through a wide range of sensors. The “+ complementary ‘OMIC sequencing” refers to transcriptomics, metabolomics, proteomics and other ‘OMIC datasets that can be integrated into the machine learning model to enrich the dataset information. The potential prediction tasks for each input data type are colour-coded on the right.
The success of ML has been facilitated by an explosion of available data, driven by the ever-decreasing costs of genome sequencing ($200 per human genome in sequencing costs only). Other drivers are the increased availability of compute power in the form of accessible and machine learning-specialised GPUs, high performance computing centres, and accessible cloud computing, leading to ML becoming established as a group of tools in genomics and crop improvement.
One area where ML is having an impact in crops is marker-assisted breeding (MAB)16,17, where ML can be used to link phenotypes of agronomic interest with molecular genetic markers so that they can be applied to accelerate breeding. Another is yield prediction, where several studies have evaluated the accuracy of different machine learning architectures across different datasets to predict crop yield18. When ML is combined with CRISPR genome editing19, it can be used both to identify potential favourable modifications and design accurate guide RNAs (sgRNAs) with few off-target effects20.
The knowledge gained by applying these methods to major crops will also assist in the improvement of orphan crops, and vice versa. In this Review, we will discuss the potential of using ML for orphan crop improvement. We will highlight how ML can improve the knowledge available for orphan crops, find similarities between major and orphan crops, and transfer knowledge from major crops to orphan crops.
Machine learning applications for crop improvement
Machine learning has been extensively applied in crop improvement, with hundreds of publications ranging from identifying markers for MAB to using image recognition for accurate phenotyping and disease resistance recognition (Table 2)19,20,21,22,23,24,25,26,27,28,29,30,31,32. Predicting phenotypes has been one of the main applications of machine learning. One of the earliest examples of yield prediction using machine learning is from 2008, where bread wheat field measurements were used in a simple artificial neural network to predict yield over seasons33. Later studies used different variables to predict yield using ML, such as transplanting parameters in rice34, irrigation and evaporation parameters in sugarcane35, or soil and irrigation data in wheat, barley, and canola across years and locations36. All these examples use environmental data, but do not include information about the genetic composition of crops.
Some studies have used genetic data alone to predict yield directly. One of the earliest examples is DeepGS, a convolutional neural network (CNN) that predicts phenotypes from genotype data, complementing the widely used RR-BLUP26. Other DL architectures have been used successfully to predict mixed phenotypes (binary, ordinal, continuous) from genotypes in bread wheat27 as well as phenotypes while incorporating data from multiple environments37. However, benchmarks reveal that DL on its own usually performed similar to traditional genomic selection approaches, with ensemble-based approaches including several models showing the highest prediction accuracy38. A similar benchmark revealed that in soybean, tree-based machine learning approaches such as XGBoost and Random Forests outperformed deep learning-based approaches in 13 out of 14 phenotypes25, indicating that DL may not be the best machine learning approach in plant phenotype prediction.
Genomic data has been successfully combined with environmental data to improve prediction accuracy. Kick et al.39 utilised genetic data, environmental measurements, and recorded management interventions to predict maize yields, finding that DL models performed similarly to, but with greater consistency than, BLUP models. Måløy et al.40 evaluated the then-novel Performer deep learning architecture using SNPs and environmental data to predict barley yield across locations and years, outperforming other DL architectures and Bayesian approaches. Li et al.41 assessed the accuracy of transfer learning by pre-training DL models using genomic and non-yield phenotypic data in maize, rice, and wheat. The pre-trained layers were then fine-tuned for yield prediction tasks, outperforming established DL and RR BLUP approaches. Image-based phenotyping or drone data is commonly used in conjunction with genetic data to predict yield. In maize, Danilevicz et al.42 combined multispectral imagery with genotyping data to identify high-performing varieties in the field. Later research focuses on multimodal models, as integrating multiple data types has generally shown superior performance compared to single-modality models43.
Once phenotype prediction accuracy has been established, ML can be employed to identify quantitative trait loci (QTLs) or genes underlying traits of interest. An early example used QTL identified by genome-wide association studies and several approaches from RR-BLUP to Random Forest, to predict yield based on genome-wide association study (GWAS)-associated markers in rice and showed that these methods outperformed established pedigree-based approaches29. In soybean, predicting yield from genotypic data using XGBoost led to the identification of SNPs linked with prediction accuracy, and these SNPs overlapped with known markers previously linked with yield25. Liu et al.28 trained a Convolutional Neural Network to predict yield based on soybean SNPs, and then drew saliency maps to identify genomic regions with the strongest impact on phenotype prediction. All identified regions overlapped with GWAS-identified SNPs. Another approach is PlantMine, which identified SNPs associated with prediction accuracy using XGBoost, and then used these ‘core’ SNPs to reduce noise in genomic prediction algorithms44. One interesting approach identified nitrogen-use efficiency genes using RNASeq, and then ranked these genes using an expression-level trained XGBoost to identify candidate genes and transcription factors. These genes were functionally validated and are now available for further nitrogen breeding in maize45. Machine learning is now at the core of crop breeding in companies, leading to improved breeding pipelines and reduced cost, for example the application of an AI assistant for breeders selecting the best breeding candidates46.
For ML to have a significant practical impact on plant breeding, training programs are essential. ML practitioners in plant breeding operate at the intersection of bioinformatics, plant biology, and breeding. They require a unique combination of skills and experience, including computational abilities, domain expertise, and proficiency in experimental design. Similar recommendations have been made previously in the field of plant breeding46. However, training opportunities for this specific skill set are currently limited.
Machine learning applications for orphan crops
There are two main approaches for crop trait prediction using ML, image-based ML models and genomics-based ML models, with some studies combining these in an ensemble approach12,14. While genomic and image data is increasingly abundant for the major crops, it is rarely available for orphan crops. Publicly available orphan crop data sources include online resource for community annotation of eukaryotes (ORCAE)47, a metabolomics database for roots, tubers, and bananas48, and a collection of 26 transcriptomes for orphan crops and their wild relatives49. ORCAE is a database for the genomes and annotations of the orphan crops assembled by AOCC47, and the genomes available through ORCAE could be used for the construction of genomics-based ML models for orphan crop trait prediction where suitable phenotype data is available. These could be complemented by intermediate phenotypes, for example transcriptomic or metabolomic data48,49.
There are currently no public databases hosting orphan crop images that could be used for image-based ML models. However, two studies have applied ML to orphan crops for trait prediction using the limited data available50,51. Nazari and colleagues52 developed a DL model, a type of ML model, to predict the quality traits of protein, tannin and, total phenolic content (TPC) in sorghum. Determining chemical content through conventional laboratory tests is expensive and time consuming, so Nazari et al.52 developed an efficient and cost-effective method to predict chemical composition using images and DL. The grains of ten lines of sorghum were harvested at maturity and the protein, tannin, and TPC content of 100 g of each line was measured using conventional laboratory tests. The remaining sorghum grains were photographed on a black background with consistent lighting, and the colours within each photograph were analysed to determine texture variables. The protein, tannin, and TPC content and the texture variables for all the ten sorghum lines were used as input for a multilayer perceptron (MLP) model for trait prediction. Multilayer perceptron is a type of DL model made up of three layers, the input layer, the output layer and a hidden layer. The hidden layer is where the model identifies patterns within the data and these patterns are then used to predict the output. The model learns through interconnected nodes within each layer that are designed to work in a similar way to the neurons in a human brain. Nazari and colleagues52 found a significant difference in the protein, tannin, and TPC content between each sorghum line and the content measured in the laboratory, and predicted by the DL model had a correlation of greater than 0.9 for each of these traits. Another study used near-infrared reflectance spectroscopy (NIRS) and DL to predict quality traits in the orphan crop Perilla53. The DL models had high prediction accuracy with R2 values of 0.83, 0.92, 0.78, and 0.82 for the biochemical traits ash, protein, total soluble sugar and phenol content respectively. By using NIRS and ML the authors were able to develop a cost-efficient and accurate method for predicting the nutritional content within Perilla germplasm. As the knowledge available for orphan crops grows, more studies could use ML to efficiently predict the traits (Fig. 3). However, the limited quantity of public data highlights the need for establishing and supporting databases of image and genomic data for orphan crops that could be applied for ML based trait prediction.
Genomic and phenotype data is collected from an orphan crop population called a training population. This genomic and phenotype data is used to train ML models. The trained ML models can then be used for trait prediction in orphan crop populations that only have genotype data. These trait predictions are then used to select individuals for crop breeding programs.
Large language model applications for crop improvement
Large language models (LLMs) are a subsection of machine learning, designed to “understand” language and identify patterns from text54. Recently, LLMs have been increasingly applied to analyse biological sequential data, such as gene expression profiles, genomic DNA sequences and protein sequences. In this context, biological language models approach the DNA or amino acid sequence as text strings, splitting the biological sequences into words and finding the relationship between them55. The application of language models to understand plant biological datasets is not a new concept56,57,58,59, but recent technological advances have enabled more powerful LLM architectures to emerge60,61. The application of LLMs can enrich the reduced genomic resources of orphan crops, leading to a better understanding of the diversity in orphan crop genomes.
The large language model’s capacity for transferring knowledge into new domains is particularly valuable in the context of orphan crops, as they can leverage insights from well-studied species to predict gene functions, identify regulatory elements, and uncover genetic patterns in orphan crops. Nucleotide Transformer is a prime example of a collection of foundational LLMs for predicting gene sequence phenotype and function that can be used for transfer learning. The Nucleotide Transformer models were trained using an extensive genomic sequence database with approximately 3202 human genomes and 850 genomes from diverse phyla, which allowed the models to learn context-specific nucleotide sequences and gain a robust understanding of genomic indicators that could be used to support the annotation of orphan plant genomes62. For example, the chia (Salvia hispanica) genome annotation used transcriptome and orthologous gene models from multiple other species, leading to ~94% genes identified according to a BUSCO analysis63. Integrating LLMs into the annotation process could further refine the functional annotation of orphan genomes by identifying the genomic patterns and gene context learned during the LLM training. DNABERT is another foundational LLM that was trained with 135 human genomes for predicting gene function, promoter sites, splice sites and transcription factor binding sites based on DNA sequence58. DNABERT demonstrated a high capacity for transferring learning to other species, effectively detecting transcription factor binding sites in genomes with under 50% non-coding similarity to the human genome58. Since transcription factors regulate gene expression, and their binding sites are often found in non-coding regions at varying distances from target genes, DNABERT’s success in identifying these sites suggests it accurately captures conserved semantic relationships within the DNA sequences. Several studies have leveraged the DNABERT model to advance plant research. A recent study further trained the DNABERT model to identify long non-coding RNA (lncRNA) in six major plant species64. The lncRNAs play an important role in regulating gene expression through interactions with DNA, RNA, and proteins that modulate gene activity being valuable targets for crop improvement65. The LLM identified lncRNA sequences from genomic DNA sequences with up to 83% accuracy in target species and a high average accuracy in identifying lncRNA sequences in previously unseen crop species64. Multiple models leveraging these foundational LLMs were proposed for the prediction of DNA methylation sites in plants due to their importance as gene expression regulators. These LLMs were trained in major plant species and tested on previously unseen plant datasets, showing their capacity to capture the species-specific indicators for methylation sites and an ability to generalise across different species that highlighting the LLMs’ effectiveness in identifying critical regulatory elements in less-studied plant genomes66,67,68.
More recently, a foundational LLM focused on crop genome sequences was released. AgroNT uses a similar structure to DNABERT, but it was trained on 48 crop species genomes, including the orphan crops pigeonpea (Cajanus cajan), cassava (Manihot esculenta) and quinoa (Chenopodium quinoa). The AgroNT model has demonstrated high accuracy in predicting regulatory annotations, promoter/terminator strength, lncRNA prediction and tissue-specific gene expression across species, indicating the model’s versatility and potential uses for identifying sites controlling gene expression in orphan crops69. Being trained exclusively with plant datasets may provide an advantage to AgroNT, as it avoids biases towards genomic structures that are exclusive to other organisms. The foundational LLMs above offer a powerful tool for transferring knowledge from major to orphan crops, as the biological annotations and experimental validation from well-curated plant species can be leveraged to detect gene regulation mechanisms in orphan species.
A major limitation for the genomics-based improvement of orphan crops is the insufficient genome references and annotated genomic resources for these species. This has hindered the identification of causal genes associated with valuable crop phenotypes. Pre-trained LLMs models could be useful to predict gene function from DNA or RNA sequencing datasets58,69. The estimated gene function output could also be applied for prioritising functional variants identified through genome wide association studies (GWAS), RNA sequencing and other genomic analysis69. In addition, the pre-trained LLMs models could be fine-tuned for specific orphan crop prediction using a reduced training dataset, leveraging the model’s learning about the molecular relationships to focus on species specific features. Ultimately, integrating pre-trained LLMs with genomic data and focused fine-tuning could help bridge the gap in understanding and harnessing the unique traits of orphan crops, unlocking their full potential for sustainable agriculture.
Transfer of knowledge between major and orphan crops
The limited knowledge and resources available for orphan crops has slowed their development50,51. However, there are many orphan crops that are closely related to major crops. For example, Solanaceae fruit include tomatoes, a major crop, and ground cherries, an orphan crop70,71. For examples like this, their evolutionary relationship can be used to learn about and improve orphan crops through gene homology. Conservation of orthologs and their functions has been found between orphan crops and related major crops9. These conserved genes allow studies to use genes and knowledge available in major crops to identify candidate genes, edit genomes, and predict traits in orphan crops.
Gene homology with major crops or model species can be used to identify genes associated with a trait of interest. Gene homology with Arabidopsis thaliana, a model species, was used to identify 108 candidate genes for seed mucilage production in chia72. Candidate genes for domestication were identified using gene homology with A. thaliana and rice73. While these studies used model species, the same methods could be applied using major crops. A ML approach for identifying candidate genes from sequences associated with a trait of interest is QTG-Finder274. QTG-Finder2 is a fast and efficient way to identify candidate genes from quantitative trait loci (QTL). The QTG-Finder2 ML model was trained on orthologs of causal genes from major crops and model plant species. Lin et al.74 hypothesised that the QTG-Finder2 model could be applied to species with little to no known causal genes, due to the conservation of orthologs between species. To test this hypothesis, they applied the QTG-Finder2 ML model in sorghum, an orphan crop, to predict causal genes for plant height. QTG-Finder2 correctly identified true plant height causal genes 70% of the time74. QTG-Finder2 improves the efficiency of identifying candidate genes and can be applied to species with few if any known causal genes. Machine learning and gene homology can be used to predict essential genes in species with little knowledge available. Essential genes are required for the reproductive success of a species and are highly conserved75. If ML can identify essential genes using gene homology it could be applied to predict other conserved genes.
Genome editing using CRISPR can make changes to DNA to improve a trait. To be able to make changes to DNA, information on the gene sequence is needed. The conservation of orthologs between major and orphan crops can be used to identify targets for genome editing. The mutation of tomato orthologs has improved the fruit size and production of ground cherries through genome editing70,71. Lodging resistance in tef, an orphan crop, has been improved by editing a rice ortholog for semi-dwarfism76. Gene conservation between orphan and major crops can be used to identify candidate genes and design genome editing targets when there is no data available for the gene of interest within the orphan crop. Machine learning can also be used to improve the editing efficiency and specificity of genome editing.
One way to find gene orthologs, that can be used for orphan crop studies, is to source it from the literature72,76; however, this information is spread through papers and journals making it challenging to know the extent of gene homology between major and orphan crops and where to find this data. Databases such as NCBI are a source of protein and nucleotide sequences and gene homology for many species77; however, they do not have information specific to orphan crops. Consolidating all major crop orthologs and their presence in orphan crops into a comprehensive database would aid studies identifying candidate genes and improving traits through genome editing in orphan crops.
Transfer learning is a machine learning method that uses pre-trained models and new datasets to fine tune ML models for a new purpose78. Transfer learning can be used to make predictions in a species with little available knowledge by training the model on a species with available data (Fig. 4). Pre-trained models can be transferred from major to related orphan crops due to the conservation of genes and gene functions45. Tomatoes are a major crop with poor quality annotations. Transfer learning was used to improve the prediction accuracy of generalised and specialised metabolism genes in tomatoes79. A model trained on A. thaliana was applied to tomato annotation data, and the prediction accuracy of the transfer learning model was greater than the model trained on the tomato annotation data for generalised metabolism genes. Prediction accuracy did not improve for specialised metabolism genes. The reason the transfer learning model performed better for the generalised metabolism genes is because they are conserved between species while specialised metabolism genes are lineage specific79. While this study focuses on a major crop with poor annotation, the same method can be applied to orphan crops. Transfer learning can be used to link knowledge from resource rich major crops to related orphan crops, for conserved traits. To aid trait prediction in orphan crops a database of trait prediction models trained on major crops should be collated; these pre-trained models could then be applied to related orphan crops using transfer learning.
ML models are trained using data from major crops. These trained ML models can then be used to predict traits in orphan crops, which have limited available data. The trait predictions are used to choose breeding candidates to improve orphan crop varieties.
The limitations of transferring knowledge from major to orphan crops whether it is through gene prediction, genome editing, or transfer learning, is that all these methods rely on conserved genes. Orphan genes are lineage specific genes that have no homologues in other species and make up 10–20% of a genome80. Orphan genes have been found to be associated with agronomically important traits such as disease resistance and abiotic stress tolerance81,82,83. These orphan or novel genes cannot be identified or improved without species specific genomic resources, so, while transferring knowledge from major to orphan crops can be used to improve some traits, we still need orphan crop specific resources to reach the maximum potential for crop improvement. Orphan crop genomic resources can identify these orphan genes that can aid crop improvement in both orphan and major crops.
There are some examples of knowledge transfer between orphan and major crops and vice-versa. For example, abiotic resistance genes not present in the bread wheat genome have been identified in the orphan cereal tef84. The salinity-resistant orphan crop groundnut has been identified as a potential source for salinity resistance in soybean85. Other examples involve transfer of knowledge from wild relatives to major crops. An example is the super-pangenome of Cicer, which included several wild relatives of chickpea and led to the discovery of novel disease resistance genes and genes involved in salt resistance, along with novel mutations in vernalisation genes86,87. A similar super-pangenome in tomato identified a wild-type only cytochrome P450 allele linked with increased yield88. Sequencing of Aegilops accessions has led to the cloning of four novel disease resistance genes not present in bread wheat89. In bread wheat, the Watkins collection of landraces from the 1930s has been a large source of knowledge applied to bread wheat, including novel resistance genes to tan spot, Fusarium head blight90 and eyespot resistance91,92. Sequencing the entire Watkins collection identified and subsequently introgressed 127 QTL alleles from landraces to bread wheat, leading to yield increases of up to 0.91 t ha−193. These examples show that by focusing on wild or landrace relatives, plant breeders can introduce significant yield gains by introgressions and crossbreeding.
Implementation of machine learning-based improvement of orphan crops
Some of the challenges for orphan crop improvement include the lack of genomic resources, limited uptake of modern crop breeding methods, and the lack of local scientists working on these issues. Collaboration between scientists, local communities, smallholder farmers and international collaborators can help bridge the gap between major and orphan crops. The International Maize and Wheat Improvement Centre (CIMMYT) is a non-for-profit organisation that aims to address the challenges faced by smallholder farmers in marginal environments94. CIMMYT develops high yielding, nutritious, and abiotic stress resistant wheat and maize varieties. They work with smallholder farmers in developing countries by providing training, trading knowledge, and exploring market opportunities. With the aid of public and private collaborations CIMMYT has improved the food security of millions of smallholder farmers in Africa, Asia and Latin America. Similar initiatives aim to improve orphan crops. Feed the Future Innovation for Crop Improvement focuses on accelerating the breeding of local roots, tubers, bananas, millets, legumes and sorghum varieties through the collaboration of scientists, global stakeholders, and local communities95. The AOCC uses a network of public and private collaborators from international, non-government, and academic institutes to collect germplasm reserves, sequence genomes and gather local input96. The AOCC aims to sequence a total of 101 orphan crop species, has completed 6 of these genomes and is in the progress of completing an additional 26 genomes. Orphan crop germplasm is held in over 150 gene banks globally, which can be used for sequencing and genotyping by initiatives such as the AOCC97. Important to each of these initiatives is the input of local communities to ensure that the crop varieties are suited to each local environment, the farmers are willing to adopt the technology and that there is a demand for the product within the local marketplace. Another method to increase local involvement and to increase the manpower behind orphan crop improvement is to recruit local farmers as citizen scientists. Triadic comparisons of technologies (TRICOT) is a citizen science method that sends volunteer farmers crop varieties or agronomic technologies to trial98. TRICOT is cost effective and does not require training or specialized skills, making it accessible to farmers in marginal communities. TRICOT has been successfully used to trial the climatic response of crops in marginal environments and to determine consumer preference of orphan crop varieties99,100. Given how important local famer and community input is for orphan crop improvement it is required that these communities benefit from the studies that they take part in. All studies in orphan crops in marginal or regional environments should have the consent of the local community and the results should be accessible by the smallholder farmers that participate. Citizen scientist studies and regional and international collaborations should be supported by policy to ensure funding of initiatives to improve orphan crops. The United Nation’s recommendations for supporting orphan crops includes funding and training for farmers willing to adopt new technologies, funding for smallholder farmers to access markets, and policies encouraging the collaborations of local knowledge and science and technology101,102. Policy frameworks should be developed to train and fund the implementation of ML and modern breeding techniques by local farmers in orphan crops and to encourage further collaborations with local communities when developing new orphan crop varieties.
One of the greatest challenges for orphan crop improvement and associated improvement in food security in nations that rely on these crops, is the lack of funding. While the majority of orphan crops will remain orphans due to their niche habitats or limited potential, many, with appropriate investment, have the potential to become major crops either regionally or even globally. The rising tide of genomic technologies should lift the performance of all crops, as knowledge can be transferred to closely related species. However, the investment should be focussed on those crops with the greatest potential for improvement considering the use of machine learning to optimise results. Machine learning models can leverage major crop datasets for training, decreasing the amount of data required for trait prediction and identification of genomic features in orphan crops. Nonetheless, strategic data generation from orphan crops is required to ensure strong alignment with the training datasets. As additional datasets are generated from orphan crops, the models can be fine-tuned to improve their accuracy and specificity over time. Additionally, understanding the genomic basis of traits in orphan crops could benefit the major crops through gene introgression and editing, and there is an argument for international seed companies to support orphan crop improvement either directly, or indirectly through technology exchange as many of them currently do. Increased support from breeding and seed companies for orphan crop improvement could substantially accelerate the use of machine learning, as these enterprises have a wealth of data from genome sequencing and field trials. Providing machine learning models with a diverse dataset would allow the model to consider the intra-species genomic variability and other factors impacting trait prediction outcomes.
Investing in single technologies for data generation is unlikely to deliver sufficient results and support across fields, with investment diversification from genomics-based breeding through to agronomy, processing and marketing required to boost orphan crop performance. Moreover, while investing in the countries where orphan crops are predominantly grown can leverage local expertise and has significant social benefits, restricting the investment to a geographic boundary may not always be the most efficient pathway to accelerate crop improvement. A strategy that integrates the best available technologies and expertise on a local and global scale can enhance the effectiveness and impact of such efforts. This is particularly important for developing ML models, that often require high computational resources and specialised skills that can be accessed more cost-effectively on a global scale. Given the limited financial resources available for orphan crop improvement, a balanced approach is important, where the most effective improvements per dollar invested may be through low-cost traditional breeding, education and marketing strategies. Subsequently, machine learning models can exploit the generated knowledge after low-cost approaches have been exploited.
Outlook
Machine learning has emerged as a promising tool for advancing research and breeding efforts in orphan crops. These underutilised plant species, often vital for food security in developing regions, have historically received less scientific attention than major staple crops. Here we have demonstrated how ML techniques are being applied to analyse genomic data, predict crop traits, optimise breeding strategies, and enhance disease resistance in major crops, knowledge which is then transferred to orphan crops. By leveraging large datasets and complex algorithms, ML approaches can accelerate the identification of beneficial genes and help develop improved varieties. This technology shows potential to address challenges specific to orphan crops, such as limited genetic resources and adaptation to local environments, to ensure food for a growing population in a warming climate.
References
Food and Agriculture Organization of the United Nations. Statistics. https://www.fao.org/statistics/en (2024).
Borrell, J. S. et al. Enset‐based agricultural systems in Ethiopia: a systematic review of production trends, agronomy, processing and the wider food security applications of a neglected banana relative. Plants People Planet 2, 212–228 (2020).
Tadele, Z. Orphan crops: their importance and the urgency of improvement. Planta 250, 677–694 (2019).
Sosa, A. Chia crop (Salvia hispanica L.): Its history and importance as a source of polyunsaturated fatty acids omega-3 around the world: A review. JCRF 1, 1–4 (2016).
Zamora-Tavares, P., Vargas-Ponce, O., Sánchez-Martínez, J. & Cabrera-Toledo, D. Diversity and genetic structure of the husk tomato (Physalis philadelphica Lam.) in Western Mexico. Genet. Resour. Crop Evol. 62, 141–153 (2015).
Wasihun, G. & Desu, A. Trend of cereal crops production area and productivity, in Ethiopia. J. of Cereals Oilseeds 12, 9–17 (2021).
African Orphan Crops Consortium. Healthy Africa through nutritious, diverse and local food crops. https://africanorphancrops.org/ (2024).
Crops for the Future. Facilitating the wider use of underutilised crops. https://cropsforthefutureuk.org/ (2024).
Kumar, B., Singh, A. K., Bahuguna, R. N., Pareek, A. & Singla‐Pareek, S. L. Orphan crops: a genetic treasure trove for hunting stress tolerance genes. Food Energy Secur 12, e436 (2023).
Fisher, R. A. Statistical methods for research workers. (1934).
Fisher, R. A. The design of experiments. (1935).
Araújo, S. O., Peres, R. S., Ramalho, J. C., Lidon, F. & Barata, J. Machine learning applications in agriculture: current trends, challenges, and future perspectives. Agronomy 13, 2976 (2023).
Kang, M., Ko, E. & Mersha, T. B. A roadmap for multi-omics data integration using deep learning. Brief Bioinform. 23, (2022).
Liakos, K. G., Busato, P., Moshou, D., Pearson, S. & Bochtis, D. Machine learning in agriculture: a review. Sensors 18, 2674 (2018).
Yoosefzadeh Najafabadi, M., Hesami, M. & Eskandari, M. Machine learning-assisted approaches in modernized plant breeding programs. Genes 14, 777 (2023).
Dudley, J. Molecular markers in plant improvement: manipulation of genes affecting quantitative traits. Crop Sci 33, 660–668 (1993).
Tong, H. & Nikoloski, Z. Machine learning approaches for crop improvement: leveraging phenotypic and genotypic big data. J. Plant Physiol. 257, 153354 (2021).
Van Klompenburg, T., Kassahun, A. & Catal, C. Crop yield prediction using machine learning: a systematic literature review. Comput. Electron. Agric. 177, 105709 (2020).
Scheben, A., Wolter, F., Batley, J., Puchta, H. & Edwards, D. Towards CRISPR/Cas crops–bringing together genomics and genome editing. New Phytol 216, 682–698 (2017).
Chen, K., Wang, Y., Zhang, R., Zhang, H. & Gao, C. CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu. Rev. Plant Biol. 70, 667–697 (2019).
Li, B. et al. Genomic prediction of breeding values using a subset of SNPs identified by three machine learning methods. Front. Genet. 9, 237 (2018).
Li, Y., Raidan, F., Vitezica, Z. & Reverter, A. Using Random Forests as a prescreening tool for genomic prediction: Impact of subsets of SNPs on prediction accuracy of total genetic values. In Proceedings of the World Congress on Genetics Applied to Livestock Production (WCGALP) 11, (2018).
Herr, A. et al. Unoccupied aerial systems imagery for phenotyping in cotton, maize, soybean, and wheat breeding. Crop Sci 63, 1722–1749 (2023).
Singh, A., Ganapathysubramanian, B., Singh, A. K. & Sarkar, S. Machine learning for high-throughput stress phenotyping in plants. Trends Plant Sci 21, 110–124 (2016).
Gill, M. et al. Machine learning models outperform deep learning models, provide interpretation and facilitate feature selection for soybean trait prediction. BMC Plant Biol 22, 180 (2022).
Ma, W. et al. A deep convolutional neural network approach for predicting phenotypes from genotypes. Planta 248, 1307–1318 (2018).
Montesinos-López, O. A. et al. New deep learning genomic-based prediction model for multiple traits with binary, ordinal, and continuous phenotypes. G3: Genes, Genomes, Genet 9, 1545–1556 (2019).
Liu, Y. et al. Phenotype prediction and genome-wide association study using deep convolutional neural network of soybean. Front. Genet. 10, 1091 (2019).
Spindel, J. et al. Genomic selection and association mapping in rice (Oryza sativa): effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines. PLoS Genet 11, e1004982 (2015).
Xu, Y., Laurie, J. D. & Wang, X. CropGBM: An ultra-efficient machine learning toolbox for genomic selection-assisted breeding in crops. Accelerated Breeding of Cereal Crops 133-150 (2022).
Gabur, I., Simioniuc, D. P., Snowdon, R. J. & Cristea, D. Machine learning applied to the search for nonlinear features in breeding populations. Front. Artif. Intell. 5, 876578 (2022).
Parmley, K. A., Higgins, R. H., Ganapathysubramanian, B., Sarkar, S. & Singh, A. K. Machine learning approach for prescriptive plant breeding. Sci. Rep. 9, 17132 (2019).
Ruß, G., Kruse, R., Schneider, M. & Wagner, P. Data mining with neural networks for wheat yield prediction. In: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). 47–56 (2008).
Basir, M. S., Chowdhury, M., Islam, M. N. & Ashik-E-Rabbani, M. Artificial neural network model in predicting yield of mechanically transplanted rice from transplanting parameters in Bangladesh. J. Agric. Food Res. 5, 100186 (2021).
Taherei Ghazvinei, P. et al. Sugarcane growth prediction based on meteorological parameters using extreme learning machine and artificial neural network. Eng. Appl. Comput. Fluid Mech. 12, 738–749 (2018).
Filippi, P. et al. An approach to forecast grain crop yield using multi-layered, multi-farm data sets and machine learning. Precision Agriculture 20, 1015–1029 (2019).
Montesinos-López, A., Montesinos-López, O. A., Gianola, D., Crossa, J. & Hernández-Suárez, C. M. Multi-environment genomic prediction of plant traits using deep learners with dense architecture. G3: Genes, Genomes Genet. 8, 3813–3828 (2018).
Azodi, C. B. et al. Benchmarking parametric and machine learning models for genomic prediction of complex traits. G3: Genes, Genomes Genet. 9, 3691–3702 (2019).
Kick, D. R. et al. Yield prediction through integration of genetic, environment, and management data through deep learning. G3: Genes, Genomes Genet, 13, jkad006 (2023).
Måløy, H., Windju, S., Bergersen, S., Alsheikh, M. & Downing, L. Multimodal performers for genomic selection and crop yield prediction. Smart Agric. Technol. 1, 100017 (2021).
Li, J. et al. TrG2P: A transfer learning-based tool integrating multi-trait data for accurate prediction of crop yield. Plant Commun, (2024).
Danilevicz, M. F., Bayer, P. E., Boussaid, F., Bennamoun, M. & Edwards, D. Maize yield prediction at an early developmental stage using multispectral images and genotype data for preliminary hybrid selection. Remote Sens 13, 3976 (2021).
Togninalli, M. et al. Multi-modal deep learning improves grain yield prediction in wheat breeding by fusing genomics and phenomics. Bioinformatics 39, btad336 (2023).
Tong, K. et al. PlantMine: A machine-learning framework to detect core SNPs in rice genomics. Genes 15, 603 (2024).
Cheng, C.-Y. et al. Evolutionarily informed machine learning enhances the power of predictive gene-to-phenotype relationships. Nature Commun 12, 5627 (2021).
Xu, Y. et al. Enhancing genetic gain in the era of molecular breeding. J. Exp. Bot. 68, 2641–2666 (2017).
Sterck, L., Billiau, K., Abeel, T., Rouzé, P. & Van De Peer, Y. ORCAE: Online resource for community annotation of eukaryotes. Nat Methods 9, 1041–1041 (2012).
Price, E. J. et al. Metabolite database for root, tuber, and banana crops to facilitate modern breeding in understudied crops. Plant J 101, 1258–1268 (2020).
Sarah, G. et al. A large set of 26 new reference transcriptomes dedicated to comparative population genomics in crops and wild relatives. Mol Ecol Resour 17, 565–580 (2017).
Kumar, B. & Bhalothia, P. Orphan crops for future food security. J. Biosci. 45, 131 (2020).
Mabhaudhi, T. et al. Prospects of orphan crops in climate change. Planta 250, 695–708 (2019).
Nazari, L., Khazaei, A. & Ropelewska, E. Prediction of tannin, protein, and total phenolic content of grain sorghum using image analysis and machine learning. Cereal Chem 99, 843–849 (2022). Image-based ML models accurately and efficiently predicted the protein, tannin, and total phenolic content in grain sorghum, demonstrating the usefulness of these ML models in sorghum improvement.
Kaur, S. et al. NIRS-based prediction modeling for nutritional traits in Perilla germplasm from NEH Region of India: Comparative chemometric analysis using mPLS and deep learning. J. Food Meas. Charact. 18, 9019–9035 (2024). The most accurate ML model for predicting biochemicals using Near-Infrared Reflectance Spectroscopy in Perilla depended on the trait of interest, highlighting the importance of model selection prior to germplasm screening.
Thirunavukarasu, A. J. et al. Large language models in medicine. Nature Medicine 29, 1930–1940 (2023).
Lam, H. Y. I., Ong, X. E. & Mutwil, M. Large language models in plant biology. Trends Plant Sci 29, 1145–1155 (2024).
Meng, J., Chang, Z., Zhang, P., Shi, W. & Luan, Y. (2019). lncRNA-LSTM: Prediction of Plant Long Non-coding RNAs Using Long Short-Term Memory Based on p-nts Encoding. In: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 347–357 (2019).
Lemay, M., de Ronne, M., Bélanger, R. & Belzile, F. k‐mer‐based GWAS enhances the discovery of causal variants and candidate genes in soybean. Plant Genome 16, e20374 (2023).
Ji, Y., Zhou, Z., Liu, H. & Davuluri, R. V. DNABERT: Pre-trained bidirectional encoder representations from transformers model for DNA-language in genome. Bioinformatics 37, 2112–2120 (2021).
Nguyen, V.-N., Ho, T.-T., Doan, T.-D. & Le, N. K. Using a hybrid neural network architecture for DNA sequence representation: A study on N4-methylcytosine sites. Comput. Biol. Med. 178, 108664 (2024).
Brown, T. B. et al. Language models are few-shot learners. https://arxiv.org/abs/2005.14165 (2020).
Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30, (2017).
Dalla-Torre, H. et al. The Nucleotide Transformer: Building and evaluating robust foundation models for human genomics. https://www.biorxiv.org/content/10.1101/2023.01.11.523679v1 (2023).
Gupta, P. et al. Reference genome of the nutrition-rich orphan crop chia (Salvia hispanica) and its implications for future breeding. Front. Plant Sci. 14, 1272966 (2023).
Danilevicz, M. F. et al. DNABERT-based explainable lncRNA identification in plant genome assemblies. Comput. Struct. Biotechnol. J. 21, 5676–5685 (2023).
Urquiaga, M. C. O., Thiebaut, F., Hemerly, A. S. & Ferreira, P. C. G. From trash to luxury: The potential role of plant lncRNA in DNA methylation during abiotic stress. Front. Plant Sci. 11, 603246 (2021).
Shi, H., Li, S. & Su, X. Plant6mA: A predictor for predicting N6-methyladenine sites with lightweight structure in plant genomes. Methods 204, 126–131 (2022).
Yu, Y. et al. iDNA-ABT: advanced deep learning model for detecting DNA methylation with adaptive features and transductive information maximization. Bioinformatics 37, 4603–4610 (2021).
Zeng, W., Gautam, A. & Huson, D. H. MuLan-Methyl—multiple transformer-based language models for accurate DNA methylation prediction. GigaScience 12, giad054 (2022).
Mendoza-Revilla, J. et al. A foundational large language model for edible plant genomes. Commun Biol 7, 835 (2024). The pretrained large language model AgroNT predicted enhancer regions and the effect of promoter-proximal regions in cassava with high and moderate accuracies respectively, demonstraing the ability of AgroNT to predict regulatory features in orphan crops.
Kwon, C. T. et al. Rapid customization of Solanaceae fruit crops for urban agriculture. Nat. Biotechnol. 38, 182–188 (2020).
Lemmon, Z. H. et al. Rapid improvement of domestication traits in an orphan crop by genome editing. Nat. Plants 4, 766–770 (2018). CRISPR-Cas9 was successfully used to mutate tomato orthologues and improve productivity traits in the orphan crop groundcherry.
Alejo-Jacuinde, G. et al. Multi-omic analyses reveal the unique properties of chia (Salvia hispanica) seed metabolism. Commun. Biol. 6, 820–820 (2023).
Li, X. et al. Multi-omics analyses of 398 foxtail millet accessions reveal genomic regions associated with domestication, metabolite traits, and anti-inflammatory effects. Mol. Plant 15, 1367–1383 (2022).
Lin, F., Lazarus, E. Z. & Rhee, S. Y. QTG-Finder2: A generalized machine-learning algorithm for prioritizing QTL causal genes in plants. G3: Genes - Genomes - Genet 10, 2411–2421 (2020). The QTG-Finder2 ML model correctly identified true plant height causal genes in sorghum and can improve the efficiency of candidate gene identification in orphan crops.
Beder, T. et al. Identifying essential genes across eukaryotes by machine learning. NAR Genom. Bioinform. 3, lqab110–lqab110 (2021).
Beyene, G. et al. CRISPR/Cas9‐mediated tetra‐allelic mutation of the ‘Green Revolution’ SEMIDWARF‐1 (SD‐1) gene confers lodging resistance in tef (Eragrostis tef). Plant Biotechnol. J. 20, 1716–1729 (2022).
Sayers, E. W. et al. Database resources of the national center for biotechnology information. Nucleic Acids Research 50, D20–D26 (2022).
Yan, J. & Wang, X. Unsupervised and semi‐supervised learning: The next frontier in machine learning for plant systems biology. Plant J 111, 1527–1538 (2022).
Moore, B. M. et al. Within- and cross-species predictions of plant specialized metabolism genes using transfer learning. In Silico Plants 2, diaa005 (2020). A ML model trained on A. thaliana increased the trait prediction accuracy of generalised metabolism genes in tomatoes but did not improve the prediction accuracy of specialised metabolism genes, demonstrating the ability of transfer learning to improve trait prediction of conserved genes.
Khalturin, K., Hemmrich, G., Fraune, S., Augustin, R. & Bosch, T. C. G. More than just orphans: Are taxonomically-restricted genes important in evolution? Trends Genet 25, 404–413 (2009).
Perochon, A. et al. A wheat NAC interacts with an orphan protein and enhances resistance to Fusarium head blight disease. Plant Biotechnol. J. 17, 1892–1904 (2019).
Li, G. et al. Orphan genes are involved in drought adaptations and ecoclimatic-oriented selections in domesticated cowpea. J. Exp. Bot. 70, 3101–3110 (2019).
Ma, D. et al. Identification, characterization and function of orphan genes among the current Cucurbitaceae genomes. Front. Plant Sci. 13, 872137–872137 (2022).
Cannarozzi, G. et al. Genome and transcriptome sequencing identifies breeding targets in the orphan crop tef (Eragrostis tef). BMC genomics 15, 581 (2014).
Mayes, S. et al. Bambara groundnut: an exemplar underutilised legume for resilience under climate change. Planta 250, 803–820 (2019).
Khan, A. W. et al. Super-pangenome by integrating the wild side of a species for accelerated crop improvement. Trends Plant Sci 25, 148–158 (2020).
Khan, A. W. et al. Cicer super-pangenome provides insights into species evolution and agronomic trait loci for crop improvement in chickpea. Nat. Genet. 56, 1225–1234 (2024).
Li, N. et al. Super-pangenome analyses highlight genomic diversity and structural variation across wild and cultivated tomato species. Nat. Genet. 55, 852–860 (2023).
Arora, S. et al. Resistance gene cloning from a wild crop relative by sequence capture and association genetics. Nat. Biotechnol. 37, 139–143 (2019).
Halder, J. et al. Mining and genomic characterization of resistance to tan spot, Stagonospora nodorum blotch (SNB), and Fusarium head blight in Watkins core collection of wheat landraces. BMC Plant Biol 19, 480 (2019).
Burt, C. et al. Mining the Watkins collection of wheat landraces for novel sources of eyespot resistance. Plant Pathol 63, 1241–1250 (2014).
Winfield, M. O. et al. High‐density genotyping of the AE Watkins Collection of hexaploid landraces identifies a large molecular diversity compared to elite bread wheat. Plant Biotechnol. J. 16, 165–175 (2018).
Cheng, S. et al. Harnessing landrace diversity empowers wheat breeding. Nature 632, 823–831 (2024).
International Maize and Wheat Improvement Center. CIMMYT. https://www.cimmyt.org/ (2024).
Cornell University. Feed the Future Innovation Lab for Crop Improvement. https://ilci.cornell.edu/ (2024).
Hendre, P. S. et al. African Orphan Crops Consortium (AOCC): status of developing genomic resources for African orphan crops. Planta 250, 989–1003 (2019).
Genesys. The Global Gateway to Genetic Resources. https://www.genesys-pgr.org (2024).
Van Etten, J. et al. First experiences with a novel farmer citizen science approach: crowdsourcing participatory variety selection through on-farm triadic comparisons of technologies (tricot). Exp. Agric. 55, 275–296 (2019).
van Etten, J. et al. Crop variety management for climate adaptation supported by citizen science. Proc. Nat. Acad. Sci. 116, 4194–4199 (2019).
Moyo, M. et al. Consumer preference testing of boiled sweetpotato using crowdsourced citizen science in Ghana and Uganda. Front. Sustain. Food Syst. 5, 620363 (2021).
United Nations. Agriculture technology for sustainable development: leaving no one behind. https://documents.un.org/doc/undoc/gen/n23/218/53/pdf/n2321853.pdf (2024).
United Nations. Agriculture technology for sustainable development: leaving no one behind the future of food and agriculture: drivers and triggers for achieving sustainable agrifood systems. https://documents.un.org/doc/undoc/gen/n23/216/98/pdf/n2321698.pdf (2024).
Acknowledgements
This research was carried out while the author was in receipt of an Australian Government Research Training Program Stipend at The University of Western Australia. This work was supported by resources provided by the Pawsey Supercomputing Centre with funding from the Australian Government and the Government of Western Australia.
Author information
Authors and Affiliations
Contributions
T.R.M: Conceptualization, Visualization, Writing - Original Draft, Writing - Review & Editing. M.F.D: Conceptualization, Visualization, Writing - Original Draft, Writing - Review & Editing. P.E.B: Conceptualization, Visualization, Writing - Original Draft, Writing - Review & Editing. M.S.B: Writing - Review & Editing. D.E: Conceptualization, Supervision, Funding acquisition, Writing - Original Draft, Writing - Review & Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Carlos Hernández-Suárez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
MacNish, T.R., Danilevicz, M.F., Bayer, P.E. et al. Application of machine learning and genomics for orphan crop improvement. Nat Commun 16, 982 (2025). https://doi.org/10.1038/s41467-025-56330-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56330-x
This article is cited by
-
Printing technologies for monitoring crop health
Nature Communications (2026)
-
Advances in CRISPR/Cas systems for engineering abiotic stress tolerance in plants: mechanisms and future prospects
Planta (2026)
-
Revitalizing orphan crops to combat food insecurity
Nature Communications (2025)
-
The use of web resources for metabolomics in horticultural crops
Horticulture Advances (2025)
-
Towards smart agriculture: AI-driven prediction of key genes for revolutionizing crop breeding
Planta (2025)