Abstract
Genetic diversity is critical for species evolution and their adaptability to global changes, while speciation rate is critical for explaining large-scale patterns of species richness. Exploring correlates of variation in genetic diversity and speciation rates across species is a major interest of evolutionary biologists, but these two questions have mostly been investigated independently. Here, we assess the relationship between intra-specific genetic diversity and speciation rate for 1897 mammal species (~one third of the total diversity) covering all mammalian orders. We find a negative association between mitochondrial genetic diversity and speciation rate across mammalian clades globally. This association is not accounted for by differences in the ecological attributes of species. Our findings suggest a systematic link between micro- and macroevolutionary processes that need to be better understood and considered when investigating determinants of either genetic diversity or speciation rates.
Similar content being viewed by others
Introduction
Genetic diversity and speciation rate play major roles in the evolution of species and clades. Genetic diversity measures the level of polymorphism of DNA sequences among individuals within a species. Understanding how and why genetic diversity varies across species is one of the main questions in population genetics1,2. Research in this area has focused on the role of species ecology (e.g., life-history strategies3,4), demographic changes (e.g., recent bottlenecks or expansions5,6), geographic structure and selection in driving variable levels of genetic diversity7,8, as well as on the consequences of genetic diversity on species adaptation and survival9,10.
Speciation rate measures the frequency at which a given species gives rise to two daughter species. Together with the rate at which species go extinct, it determines how species richness varies in time and across species groups. Understanding how and why speciation rate varies across species is thus one of the main questions in macroevolution11,12,13,14,15,16. Research in this area has focused on the role of species ecologies and the environment they experience (e.g., whether they occur in tropical or temperate biomes) in modulating these rates17,18,19.
In comparison with the wealth of studies investigating the factors that modulate either genetic diversity or speciation rates, and despite the interdependence between genetic and speciation processes20, the genetic diversity-speciation relationship remains poorly characterized. More generally, there is a gap in our understanding of the interrelationship between species ecology, the environment, population genetics, and speciation rates21,22,23,24,25,26. Only a handful of studies have investigated the relationship between speciation rates and population divergence rates27, population structure28,29, substitution rates30,31, or genetic diversity32,33, with contrasting results. The two only studies we are aware of focusing on genetic diversity found an absence32 of or a weak negative association33 between genetic diversity and speciation rates.
The relationship between genetic diversity and speciation rates is crucial, but what type of relationship should we expect? Species-wide genetic diversity is the product of mutation rate and the effective population size of the species (Ne34). In an idealized species constituted of a single Wright-Fisher population with random mating and no selection, Ne is simply the census population size, and genetic diversity is thus higher in more abundant species and/or species with high mutation rates. Given that abundant species with wider ranges are more likely to be hit by isolating mechanisms such as geographic barriers, and that high mutation rates increase the rate at which populations acquire substitutions and thus reproductive isolation20,30, we could expect speciation rates to be higher in lineages with high genetic diversity. This positive association could be bolstered by intraspecific geographic structure (limited dispersal between sub-populations), which is the premise of allopatric speciation and tends to increase Ne (and thus genetic diversity), although the effect of geographic structure on species-wide genetic diversity is highly dependent on the details of the migration process35.
A positive genetic diversity-speciation rate association could also be related to divergent natural selection, which promotes fast speciation according to the theory of adaptive radiations36,37, and is more likely to occur in populations with many polymorphic alleles to act upon38,39,40,41. Speciation itself potentially maintains high genetic diversity within species by increasing the number of interacting species, as proposed by the diversity begets diversity hypothesis42. The proposed mechanism is that biotic interactions increase opportunities for intraspecific divergent selection; this can maintain high genetic diversity within species on genes involved in the interactions or linked loci43. A positive genetic diversity - speciation rate association could also be strengthened by external factors, such as temperature, which induces interspecific variation in mutation rates (e.g. related to latitude) that have either a direct flow-through effect on speciation rates as proposed by the evolutionary speed44 and metabolic45 theories of biodiversity, or are indirectly linked through the effect of temperature on other factors that modulate speciation rates, such as climatic stability and productivity46.
Alternatively, there are potential causes of a negative diversity-speciation rate association. Species with low genetic diversity (reflecting small effective population size) tend to accumulate reproductive incompatibilities faster due to the reduced efficacy of purifying selection, which could lead to higher speciation rates47,48,49. In the other direction, speciation can reduce genetic diversity through bottleneck effects50,51. Previously observed bursts of molecular evolution associated with speciation events52,53 support this inter-relationship between speciation, small effective population sizes and genetic effects. If speciation is instead adaptive, and not limited by standing genetic variation, we can expect positive selection to reduce genetic diversity by increasing heritable variance in fitness among individuals (which reduces Ne) and by fixing beneficial alleles54, while spurring speciation by driving populations towards distinct adaptive peaks. Heterogeneity in the time it takes for dividing populations to complete speciation, which can be related to different ecological, geographic or genomic contexts, could also generate a negative association between genetic diversity and speciation rate, as species for which speciation takes a long time to complete will tend to accumulate more genetic variation (as they encompass increasingly genetically differentiated sub-populations) while having lower speciation rates55 when there are few speciation initiation events and frequent population extinctions: in this case, completing speciation fast is key to induce speciation events56. Finally, if geographic structure spurs speciation but decreases rather than increases species-wide genetic diversity, as expected if some populations contribute much more migrants than others57,58,59, we also expect to find a negative genetic diversity - speciation rate relationship.
In addition to such potential mechanistic links between genetic diversity and speciation rates, an indirect association could also arise from independent factors that correlate with both genetic diversity and speciation rates, as already mentioned for temperature. Another important (somewhat related) example is latitude, which correlates with many biologically important variables: prior work has found lower levels of genetic diversity at temperate latitudes, generally attributed to recent climatic shifts and associated founder events60,61, and a latitudinal gradient in speciation rates, generally attributed to other factors17,26,62, although both the existence and direction of this gradient is debated26,63. Other potentially important factors are intrinsic characteristics of the species, in particular life-history traits that determine species position on the r/K-strategist gradient: it has been proposed that small species with short generations and high fecundity (r-strategists) have higher genetic diversity than large species with long generations and small fecundity (K-strategists), potentially in relation with their differential sensitivity to environmental variations3; on the other hand, r-strategists tend to live in unstable environments, which could either promote speciation (e.g. by inducing divergent selection) or impede it (e.g., by preventing a fine niche partitioning).
Here, to provide a thorough characterization of the genetic diversity - speciation relationship, we assemble a dataset encompassing the whole extant mammalian radiation. To do so, we use a mitochondrial gene (cytochrome b). Compared to nuclear genes, mitochondrial genes are characterized by high mutation rates, low population sizes, strong purifying selection, the absence of recombination, and strong linkage64. While it has been suggested that under this situation natural selection could erase the relationship between genetic diversity and effective population size65, this hypothesis has been disputed66. In mammals in particular, mitochondrial and nuclear polymorphisms have been shown to be correlated67,68. We confirm this finding here by analyzing several nuclear databases. We show that intraspecific genetic diversity and species-specific speciation rates are negatively correlated and that this association is likely not due to differences in species’ ecological characteristics.
Results and Discussion
Mitochondrial genetic diversity estimates across mammals
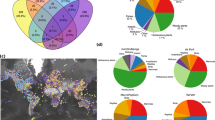
We assembled a mammal dataset using the phylogeny from Upham et al. (2019)69 as our reference. We gathered a database of cytochrome b alignments by matching GenBank sequences to species names from this phylogeny and estimated synonymous genetic diversity for each species as Tajima’s70 \({\theta }_{T{syn}}\) and Watterson’s71 \({\theta }_{W{syn}}\), corrected for gaps in the alignment72 (see Methods). Our database encompassed 90,337 sequences distributed across 1897 species, with a minimum of five sequences per species (Supplementary Fig. 1). These species provided a good representation of the entire extant mammal tree of life (Fig. 1). Consistent with previous analyses of genetic diversity across mammals68, we found that genetic diversity varies by several orders of magnitude across species (from 7.5\(\times\)10−5 to 0.113 for \({\theta }_{T{syn}}\) with a mean of 0.0193, and from 2\(\times\)10−4 to 0.105 for \({\theta }_{W{syn}}\) with a mean of 0.02, Fig. 1, Supplementary Fig. 1). The clade-level distributions of genetic diversity overlap largely (Fig. 1, Supplementary Fig. 2), with Castorimorpha (castors and beavers) showing the highest mean genetic diversity and Carnivora the lowest (Supplementary Fig. 2).
Bars at tips reflect estimated within-species genetic diversity for those species with 5 or more cytochrome b sequences available: Tajima’s \({\theta }_{{Tsyn}}\) (inner circle, red color legend in the top right inset) and Watterson’s \({\theta }_{{Wsyn}}\) (outer circle, blue color legend). Central inset: distribution of tip speciation rates for all mammals (black line, shaded fill) and 14 clades with more than 20 species (coloured lines, no fill); Top-right inset: distribution of genetic diversity, log scaled following the same line colouration. Silhouette figures were contributed by various authors with a public domain license (public domain mark 1.0; CC0 1.0) from PhyloPic (http://phylopic.org). Source data are provided as a source data file.
Tip speciation rate estimates across mammals
We next estimated species-specific (tip) speciation rates using the ClaDS model15,73, fitted to the mammal phylogeny from Upham et al. (2019), using both the consensus Maximum Clade Credibility (MCC) and 100 trees from the posterior distribution69. Consistent with what was found for birds using the same method15,73, we found that speciation rates vary considerably within and across clades (from 0.03 to 1.29 events per lineage per million years with a mean of 0.25, Fig. 1, Supplementary Fig. 3). The speciation rates of the species with genetic diversity estimates spanned the full range found in the group as a whole (from 0.03 to 1.1).
Negative genetic diversity-speciation rate relationship
We found a highly significant negative association (Supplementary Fig. 4; PGLS using \({\theta }_{{Tsyn}}\) with MCC tree: slope estimate −0.431, p-value 2.69\(\times\)10-9, \({R}_{{Resid}}^{2}\) 0.125; \({R}_{{Resid}}^{2}\) computed as in Ives 201874) between intraspecific genetic diversity and speciation rates across all mammals (Fig. 2, Fig. 3). This significant negative relationship was found while accounting for phylogenetic dependence in the data both with phylogenetic generalized least square models (PGLS, Fig. 3) and Bayesian generalized linear multilevel models (BMLM, Fig. 3). This was consistent across clades, most of them showing a negative association (Fig. 2), even though the statistical significance of the relationship depended on the clade, the tree, and the analysis performed (Fig. 3, Supplementary Fig. 5). The results were largely consistent across genetic diversity estimates (i.e. \({\theta }_{{Tsyn}}\) or \({\theta }_{{Wsyn}}\)), with original or mean subsampled estimates, across trees (i.e. the MCC tree and the 100 posterior trees), and when using PGLS or BMLM analyses (Supplementary Fig. 5). The global (all mammals) relationship remained statistically significant when reducing the global dataset from species represented by more than five sequences (1897 species) to species represented by more sequences, up to a threshold of 100 sequences per species (the dataset then encompassed 219 species, Supplementary Fig. 6). The relationship also remained significant when accounting for the geographical extent of sampling (Supplementary Fig. 7). The reliability of speciation rate estimates depends on the reliability of the estimates of phylogenetic branching times; these depend on a variety of factors, including hypotheses made on the molecular clock model75, the birth-death prior76,77, and the potential interaction between the two78. While we acknowledge this uncertainty (and accounted for some of it by running analyses on posterior tree samples), we cannot think of a consistent bias that could artifactually generate a negative correlation between speciation rates (computed from a 31-gene supermatrix phylogeny) and cyt b genetic diversity.
The number of species included in each analysis is indicated. Speciation rates represented by their 95% confidence intervals (CIs) from 100 posterior trees; CIs are very narrow, demonstrating that estimates vary little across posterior trees. Results of the PGLS analyses on the consensus MCC tree are provided and linear regression lines with 95% confidence intervals are shown in purple for visualization purposes. Axes are log scaled. Source data are provided as a source data file.
The grey density plots with median point and 95% confidence intervals in black represent the estimated posterior distribution of slopes obtained with the Bayesian Multilevel Models (BMLM) using 100 phylogenetic trees with approximately 1000 posterior samples per tree. The points below represent the slopes estimated with Phylogenetic Generalized Least Squares analyses conducted on each of the 100 trees and are coloured in red when significant (p-value < 0.05). Source data are provided as a source data file.
No detectable effect of ecological attributes or selection
If some external factors or intrinsic characteristics of the species (i.e. covariates) influence genetic diversity and speciation rates in opposite directions, this could indirectly induce the observed negative genetic diversity - speciation relationship. This would occur, for example, if genetic diversity decreases with latitude60,61 and speciation rates increase with latitude, as suggested by some authors17,26,62. Or if r-strategists sustain higher genetic diversity3 but are less likely to speciate. To investigate these potential effects, we first tested the correlation between genetic diversity and latitudinal midpoint18, mean range temperature18, body-mass79,80, generation length3,7,79, and fecundity (litter size)3,7,79. When these covariates were analyzed one-by-one, we found, as expected, that genetic diversity is significantly higher in mammals inhabiting at low latitudes, in warm climates, and in small species with short generations (Supplementary Fig. 8). The effect of generation length was no longer significant when the covariates were all combined in a single analysis (Supplementary Fig. 9 & Table 1). Genetic diversity was higher in species with small litter sizes (Supplementary Figs. 8, 9 & Table 1), but this relationship was highly sensitive to the set of species included and we therefore do not interpret it biologically (see Methods). The only covariate significantly correlated with speciation rate in at least some analyses was litter size, with a positive association (Supplementary Figs. 8, 9, and Table 1). The absence of a latitudinal gradient in speciation rate is consistent with recent findings across vertebrates63, and suggests that other covariates that correlate with latitude, such as rate of climate change and species richness, would not be strong predictors of speciation rates either. The negative association between genetic diversity and speciation rate could in part be due to an indirect effect of litter size, however this negative association remained highly significant (p-value < 0.01) when accounting for the effect of all traits (including litter size) (Table 1, Supplementary Fig. 9). Hence, although we cannot exclude the potential indirect effect of other covariates not considered here, these results suggest a direct negative association between genetic diversity and speciation rates.
Selection can potentially affect the genetic diversity - speciation rate relationship, as noted in the Introduction. While we measured genetic diversity at synonymous sites, these neutral sites can be closely linked to nonsynonymous sites under selection, particularly in the mtDNA with limited recombination. Depending on the nature of selection, in particular whether it is purifying or adaptive, and divergent or directional, selection could either generate a negative genetic diversity - speciation rate relationship at linked sites or weaken it. To assess this potential effect, we computed genetic diversity at nonsynonymous sites. Genetic diversity was lower at nonsynonymous than synonymous sites, showing that all sites are not entirely linked despite limited recombination (Supplementary Fig. 10). If selection drives the observed correlation between genetic diversity and speciation rate, we expect the correlation to be stronger with genetic diversity measured at nonsynonymous sites. Instead, the relationship was weaker with nonsynonymous (MCC PGLS slope estimate −0.380, p-value < 0.0001) than synonymous genetic diversity (slope estimate −0.430, p-value < 0.0001) with a significant difference (Pillai’s test p-value 6.37\(\times\)10-8) (Supplementary Fig. 10). These results suggest that the negative genetic diversity - speciation rate relationship is not driven by selection.
Role of mutation rate versus N e
Levels of intraspecific genetic diversity depend on both effective population size (Ne) and mutation rate (\(\mu\)) (\(\theta={N}_{{\rm{e}}}\mu\)). To further investigate the potential mechanisms underlying the observed negative genetic diversity – speciation rates relationship, we used the scaling of phylogenetic branch lengths in units of substitutions at the 3rd codon position versus time (in years) as a proxy for mutation rates (see Methods). As expected, the relationships between genetic diversity and both mutation rate and Ne are positive (Supplementary Fig. 11). We also found a negative, although only marginally significant, correlation between mutation and speciation rates, as well as between Ne (estimated as \(\theta /\mu\)) and speciation rates (Fig. 4). The negative correlation between mutation and speciation rates is unexpected given the evolutionary speed44 and metabolic45 theories, as well as previous empirical results30,31, and could be due to the smoothing of speciation rate differences obtained with ClaDS15 (see Methods). These results nevertheless suggest that fast speciation is not explained by high mutation rates in mammals, and that at least part of the negative genetic diversity – speciation rate relationship arises from a negative association between Ne and speciation rates.
Mutation rates are computed using the scaling of phylogenetic branch lengths in units of substitutions versus time (in years) for 100 trees, and Ne is computed using the ratio of genetic diversity to mutation rates. Left panels: mean \(\lambda\), Ne and \(\mu\) across 100 trees and 95% confidence intervals are shown, with a regression line and log scaled axes. Right panels: slope estimates from BMLM analyses on 100 trees (shaded distributions) and MCC tree (triangles). The black intervals represent the corresponding 95% credibility intervals and the medians. The circles (and triangle) below each of these plots represent PGLS estimates and are coloured red when significant (p-value < 0.05). Source data are provided as a source data file.
Specificities of mammals and mitochondrial markers
Taken together, our results suggest that population genetic processes and the tempo of speciation are tightly linked, although the generality of the negative genetic diversity – speciation rates relationship we observed would need to be tested on other species groups and across genomic data. There is a possibility that mammals experience a particularly high frequency of founder geographic speciation events81, hence a strong reduction of genetic diversity82,83 at speciation, that could generate a negative genetic – diversity speciation rates relationship in this group that wouldn’t necessarily be observed in other species groups dominated by different speciation modes.
It would also be insightful to assess the genetic diversity - speciation rate relationship using nuclear markers. Indeed, nuclear and mitochondrial genetic diversity are expected to show substantial differences, with mitochondrial genetic diversity being in general more strongly influenced by variations in mutation rates, demography, geographic structure and selection than nuclear diversity. It has even been suggested that mitochondrial genetic diversity is relatively constant across species and does not reflect Ne65,68,84 although this remains debated66. Our analyses suggest that, in mammals at least, mitochondrial genetic diversity reflects both Ne and mutation rate, as reported above (Supplementary Fig. 12). Unfortunately, we still lack consistent nuclear data across species at broad taxonomic scales, and as a result, macrogenetic studies with a large phylogenetic scope such as the one conducted here are not yet possible with nuclear markers85,86,87, although we expect that they will soon be. In an effort to assess how much our results may be specific to mitochondrial genetic diversity versus general across genetic markers, we analyzed four nuclear datasets, each with their specific limitations (Supplementary Note 1). We found that mitochondrial genetic diversity is in general positively correlated to nuclear genetic diversity (Supplementary Fig. 12A). We did not recover a negative association between nuclear genetic diversity and speciation rates, but this is likely due to limitations of the nuclear databases (Supplementary Fig. 12B, see the Supplementary Note 1 for a detailed discussion). Our results confirm previous studies that reported several orders of magnitude variation in mitochondrial genetic diversity (and associated Ne estimates) across mammals (e.g. Piganeau and Eyre-Walker 200988) and a good correlation between mitochondrial and nuclear genetic diversity in this group68. This suggests the negative relationship between genetic diversity and speciation rates may hold across markers, although this will need to be tested in the future with a larger number of nuclear datasets.
Processes that lead to a negative association between genetic diversity and speciation rates therefore seem to dominate those that could have generated a positive association, although the exact mechanisms at play cannot easily be disentangled. The relationship could for example be related to the faster genetic divergence between sub-populations in species with small Ne20,48,49, bottleneck effects at speciation50, the preponderance of a geographic mode of speciation with asymmetric dispersal57, heterogeneity in the time it takes to complete speciation, or a combination of such processes. This illustrates the complexity of the micro-macro evolutionary continuum and highlights the need for quantitative models linking population genetics to the tempo of speciation. A first implication of the negative genetic diversity – speciation rates relationship is that the availability of polymorphic alleles does not exert a rate-limiting control on speciation dynamics. A second implication is that speciation does not maintain high genetic diversity below the species level. Quite to the contrary, the frequency of speciation events seems to limit the amount of genetic diversity that a species contains. Given that genetically poor species tend to be more prone to extinction89, we can speculate that fast-speciating lineages, depleted of genetic diversity, are “volatile” lineages with high extinction rates.
Regardless of the mechanisms that underlie the negative genetic diversity – speciation rates relationship we observed, the consistency and robustness of this relationship highlight the importance of microevolutionary processes for understanding the dynamics that shape broad-scale patterns of diversity. Reciprocally, these deep-time dynamics influence the genetic diversity of present-day species, and therefore their capacity to adapt and cope with global change.
Methods
Mammalian phylogeny
We used a recent time-scaled phylogeny built using a “backbone-and-patch” approach to assemble species-level relationships of living mammals69. Using a 31-gene supermatrix, the backbone tree of 28 main groups was estimated, and then the 28 species-level “patch” phylogenies were re-scaled from backbone divergence times and grafted on to form the combined Mammalia-wide trees. We used the phylogeny that contained only extant species with DNA data (“DNA-only” trees) which encompassed 4064 species, representing ~69% of the total mammalian diversity, rather than the trees that additionally contain recently extinct and taxonomically imputed species. In order to account for phylogenetic uncertainty in our analyses, we used both the maximum clade credibility consensus tree (MCC) and a set of 100 trees randomly sampled from the credible set of 10,000 trees from Upham et al. (2019)69.
Estimating intraspecific genetic diversity across mammals
We computed intraspecific genetic diversity across mammals using the mitochondrial DNA locus cytochrome b (cyt b). We downloaded DNA sequence data per mammals family from the NCBI GenBank database on the 13th of December 2019 using as arguments “(Family)[Organism] AND CYTB NOT Homo sapiens[Organism]” and using the R (v.3.5.190) package Reutils (v0.2.391). We excluded sequences with hybrid species names and non-identifiable species names.
We used a list of synonyms to match species names linked to the NCBI GenBank sequences to species names from the mammalian phylogeny, which were from a master taxonomy that includes IUCN (2015) accepted species names and new species names. We updated the synonym list from92 using the R package Taxize (v.0.9.9193) and rotl (v.3.0.1094) and the Integrated Taxonomic Information System (ITIS), IUCN and Open Tree of Life (OTL) databases. We discarded sequences linked to NCBI species names that matched more than one species in the phylogeny. Additionally, subspecies sequence names that were not valid in all species names databases were only kept when the specific epithet but not the generic name with the subspecies name would match the species names in the phylogeny.
We selected one cyt b sequence per family to be used as a reference for the alignments; we chose the longest available sequence (DNA sequence size for most mammal species is 1140 bp). We used the recently developed python toolkit SuperCRUNCH95 to process cyt b sequences and obtain family-level alignments. This consisted of filtering sequences while creating a BLAST database from the reference sequences, adjusting the orientation of the sequences, checking for the presence of stop codons and adjusting accordingly, and aligning them (with adjusted reading frames to improve the alignments) using Mafft96,97 set with the FFT-NS-i algorithm. We then separated the resulting family-level alignments into species-level alignments, trimmed these alignments to remove potential sites with only gaps, and inspected them visually using Geneious v.7.1.998. We then employed the bioseq package (v.0.1.4)99 to identify any not in frame sequences, which we subsequently realigned using macse (v2.06)100. Conveniently, macse takes into account frameshifts in the alignment while incorporating the appropriate genetic code (in this case, the mitochondrial vertebrate genetic code for cyt b). We identified the synonymous sites in all alignments using the bioseq package, and computed genetic diversity at synonymous and nonsynonymous sites across species.
We measured intraspecific molecular genetic diversity using Tajima’s70 θTsyn, which uses the mean pairwise difference among sequences, and Watterson’s71 θWsyn, which is based on the number of segregating sites. We chose these measures of genetic diversity as a) they are well grounded in population genetic theories, being direct estimators of θ in an idealized population following the Wright-Fisher model, b) they are the two mostly widely used measures of genetic diversity in population genetics, and c) they have been modified to account for missing data. We indeed used the modified estimators of Ferretti et al. (2012)72, which account for missing data in the alignments and therefore avoid the need to remove bases or individuals from the analysis. We computed these estimators for all species represented by at least five sequences in the species-level alignments, hereafter referred to as “original” genetic diversity estimates. We initially gathered 124,289 sequences, which reduced to 111,624 after filtering; these sequences spanned 1959 species represented by at least five sequences.
To better account for uncertainty due to the broad variation in the number of sequences per species (median 17 and maximum 2080 sequences, Supplementary Fig. 1), we performed 1000 sub-sampling of each species-level alignment to four sequences and computed genetic diversity estimates for each of these subsamples, hereafter referred to as ‘subsampled’ genetic diversity estimates. We also computed the mean and standard error of these subsampled estimates for each species. The original and subsampled estimates were generally strongly correlated (Supplementary Fig. 1). This suggests that our original genetic diversity estimates were not too sensitive to the number of individuals sampled, and therefore also to the fraction of the full species range represented. We excluded species for which original genetic diversity for either θTsyn or θWsyn was not within the range of subsampled estimates (Supplementary Fig. 1). In the end, the dataset encompassed 98,966 sequences across 1897 species. Nonsynonymous genetic diversity was computed for the 1730 species that had nonsynonymous sites. As another approach to testing the potential effect of the number of sequences per species on the genetic diversity - speciation rate relationship, we restricted the dataset to species represented by at least 10, 20, 50, 75 and 100 sequences (Supplementary Fig. 6). Finally, to test the potential effect of geographic extent of sampling, we obtained geographical coordinates from Theodoridis et al. (2020)61, by matching our sequences with theirs using NCBI IDs. Only a subset of our sequences were georeferenced by Theodoridis et al. 2020, reducing the dataset to 453 species. We assessed the correlation between the mean within-species geographic distance between sequences and genetic diversity, as well as the significance of the genetic diversity - speciation rate association when accounting for mean geographic distance (Supplementary Fig. 7).
Estimating species-specific speciation rates
We estimated branch-specific speciation rates using an updated implementation of the recently developed cladogenetic diversification rate shift (ClaDS) Bayesian model15. This implementation uses data augmentation for faster computation in large phylogenies, and is available in the PANDA.jl (v0.0.2) package73. ClaDS allows for gradual variation in diversification rates by implementing a rate shift at each speciation event. We used the ClaDS2 model, which implements a scenario with a constant turnover rate (i.e extinction rate divided by speciation rate). We fitted ClaDS2 to the full mammalian phylogeny, accounting for missing species by including family-level sampling fraction information. We computed each “patch clade” sampling fraction as the number of species included in the (DNA only) phylogeny for this clade divided by the number of extant species in the “complete” phylogeny of Upham et al. (2019)69 with imputed species. We ran ClaDS2 on both the MCC tree and the 100 trees randomly sampled from the credible set. We extracted speciation rates at the tips to obtain species-specific (tip) speciation rates. Our species-specific rates are positively correlated to those estimated in Upham et al. (2020)101 where rates were estimated with DR102, but only loosely so (R2 = 0.68). We present ClaDS-based results as this method accounts for small rate variations in an explicitly model-based approach15.
Assessing the correlation between species-specific genetic diversity and speciation rate
We assessed the correlation between species-specific speciation rate estimates and species genetic diversity (using both \({\theta }_{{Tsyn}}\) and \({\theta }_{{Wsyn}}\)), at both a global scale for the entire Mammalia class and 14 of the 28 monophyletic patch clades from Upham et al. (2019)69, selected as those with at least 20 species for which genetic diversity could be computed. We analyzed the correlation using two statistical approaches that account for phylogenetic dependence in the data: a frequentist Phylogenetic Generalized Least Squares approach (PGLS) implemented in the R package nlme (v.3.1-162)103, and a Bayesian Multilevel Models approach (BMLM) implemented in the R package BRMS (v.2.11)104. In both cases, analyses were performed for both the MCC tree and each of the 100 trees. We performed the global and per-clade PGLS analyses independently, using either the original estimates of genetic diversity or the mean of subsampled estimates. The analyses used the gls function and included the phylogenetic correlation structure using the corPagel function estimated using maximum likelihood (ML). We also calculated the R-squared values of the PGLS analyses105 using the rr2 package106, which decomposes the total variance of the response variable into variance explained by the predictor variable (R2_lik) and variance explained by other factors (R2_resid). The BMLM approach allows a joint analysis of the entire Mammalia and individual clades while propagating the modelization uncertainty across levels. We performed this joint analysis with clade as a grouping factor, allowing the slope and intercept of the correlation to vary across clades. We ran these analyses with log-transformed genetic diversity as the response variable and log-transformed speciation rate as the predictor variable; preliminary results on a subset of analyses with speciation rate as the response variable suggested similar results. BMLM also allows incorporating the error associated with the response variables, so we additionally performed an analysis using the mean of the subsampled genetic diversity with its standard deviation implemented in an error-in-variables model. The analyses with nonsynonymous genetic diversity were performed with PGLS on the MCC tree and 100 posterior trees using nonsynonymous genetic diversity (\({\theta }_{{Tnonsyn}}\)). Additionally, we performed a Multivariate Analysis of Variance (MANOVA) to compare the association between synonymous and nonsynonymous measures of genetic diversity and speciation rates (MCC tree and rates) using the Pillai’s test implemented in mvMorph (function manova.gls with “LL” method; v.1.1.5107).
Species-specific covariates
To investigate whether the relationship between genetic diversity and speciation rates could be due to indirect effects from relationships of each variable with species-specific characteristics, we gathered data for species’ mean body mass and range area108, mean annual temperature over species range and latitudinal midpoint109, generation length and litter size110. Next, we independently analyzed the correlation between these covariates and both genetic diversity and speciation rate. We also analyzed the correlation between genetic diversity and speciation rate when accounting for the effect of the covariates by including them as predictor variables. These three models were run at the global scale for mammals as a whole, on the MCC tree and each of the 100 posterior trees, using \({\theta }_{{Tsyn}}\) as the estimate of genetic diversity. We ran both “one-by-one” analyses where each covariate was considered independently, and a “combined” analysis where they were all included simultaneously in a single analysis.
Surprisingly, we found that genetic diversity was higher in species with small litter sizes (Supplementary Figs. 8, 9 & Table 1). This was still true when analyzing this relationship without accounting for other covariates (MCC PGLS slope estimate −0.22, p-value = 0.014). This can seem counter-intuitive, and contrasts with Romiguier et al. (2014)3. We also computed fecundity as the product of litter size and frequency of litters per year, as in Welch et al. (2008)111, for the subset of species for which these data were available (1017 species)111. With this subset, we found that the relationship between genetic diversity and fecundity was non-significant with fecundity measured as litter size (MCC PGLS slope estimate 0.0025, p-value = 0.98), and significantly positive with fecundity measured as litter size x litter frequency (MCC PGLS slope estimate 0.12, p-value = 0.039). We also found that the relationship between genetic diversity and litter size was not significant when including all the species for which data were available (1427 species, MCC PGLS slope estimate −0.11, p-value = 0.19). The relationship between genetic diversity and litter size therefore seems highly sensitive to the set of species included, and we do not interpret it biologically in the paper. We include it in our analyses to account for a potential indirect effect of this covariate on the genetic diversity - speciation rate relationship.
Mutation rates and effective population sizes
Under the neutral theory and for a haploid population (the case relevant to cyt b which is a haploid and maternally inherited mtDNA locus) at equilibrium, \(\theta=\,{N}_{{\rm{e}}}\mu\), where Ne is the effective population size and \(\mu\) is the mutation rate per site per generation. We unfortunately do not have direct measures of Ne and \(\mu\); the estimates we use here for \(\mu\) rely on phylogenetic branch lengths (see below), which are also used to estimate speciation rates, and the estimates of Ne are based on our estimates of genetic diversity. This introduces an inevitable circularity in the analyses to keep in mind for cautious interpretation of the results. We approximated species-specific mutation rates by substitution rates at the 3rd codon position of the cyt b alignment from Upham et al. (2019)69. This approach (versus, for example, using codon-based models to estimate synonymous substitution rate) is often done to mitigate computational constraints112. In theory, substitutions at the 3rd position more closely reflect neutral divergence arising from mutations because changes in 3rd codon positions are mostly synonymous, thus limiting the effect of natural selection. We estimated divergence in units of substitutions at third codon positions on the MCC tree and the 100 trees topologies under the GTR substitution model with rate variation among sites (modelled with a gamma distribution with 5 rate categories) using the baseml program in PAML (v.4.9j113). These analyses were run across the same 100 trees used to infer speciation rates but performed per patch clade from Upham et al. (2019)69, with a further split of Carnivora and Muridae clades due to computational limits. We obtained estimates of mutation rates in time units (i.e. per year) for each species by dividing the estimated number of substitutions per site on each terminal branch by branch length in time units (as given by the time-calibrated trees in years). To avoid potential biases, 81 species with estimates of number of substitutions of 4\(\times\)10-6 (default value assigned to branches by PAML when the actual value is zero) were discarded. Finally, we calculated Ne per species by dividing \({\theta }_{{Tsyn}}\) by the estimated mutation rate (per year) for these species. We initially computed the mutation rate per generation by multiplying the mutation rates per year by the length (in years) of a generation, and used this per generation rate to obtain Ne, but we found that variations in mutation rates (and therefore also Ne) were then almost entirely explained by variations in generation length. This led to some spurious results, such as a negative relationship between genetic diversity and mutation rate. Not correcting by generation length is justified by our result that generation length does not explain the negative relationship between speciation rate and genetic diversity (Table 1). It also allows a more direct comparison between mutation rates and speciation rates, which are expressed in Myrs. We explored the relationship between speciation rate (as the explanatory variable) and either \(\mu\) or Ne using PGLS and BMLM analyses on the MCC tree and the 100 posterior trees. We found an unexpected negative correlation between mutation and speciation rates, which we attribute to the smoothing of speciation rates obtained with ClaDS analyses15. This smoothing would lead to speciation rates being underestimated on short branches with high speciation rates, and overestimated on long branches with low speciation rates. On the other hand, mutation rates are likely overestimated on short branches and underestimated on long branches because of the way they are computed (number of substitutions divided by branch length). Combined, these effects would generate the negative correlation we observed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data used in this manuscript are deposited in Figshare (https://doi.org/10.6084/m9.figshare.25627044.v2). The main datasets include cytochrome b sequences from GenBank (more specifically, the accession numbers can be found at https://figshare.com/ndownloader/files/51449870 and in Supplementary Data 1), trait data compiled from published literature, and previously published data from Upham et al. 2019 used for speciation and mutation rate analyses. Additional datasets used in this study for other analyses are also included in the repository. Source data to reproduce figures can be found in files “Figures_v3.Rmd” (code) and “data.tar.xz” in both the Figshare and GitHub repositories provided, as well as in the Source Data file. Source data are provided with this paper.
Code availability
All code used to perform the analyses is publicly available at Figshare (https://doi.org/10.6084/m9.figshare.25627044.v2) and GitHub (https://github.com/acafonsosilva/GenDivSpRate). The repository includes scripts for sequence processing, genetic diversity, speciation rate and mutation rate estimation, and statistical analyses. An R Markdown file provides the complete workflow to reproduce all main and Supplementary Figs. in this manuscript.
References
Lewontin, R. C. The Genetic Basis of Evolutionary Change. vol. 560 (Columbia University Press New York, 1974).
Ellegren, H. & Galtier, N. Determinants of genetic diversity. Nat. Rev. Genet. 17, 422–433 (2016).
Romiguier, J. et al. Comparative population genomics in animals uncovers the determinants of genetic diversity. Nature 515, 261–263 (2014).
Leffler, E. M. et al. Revisiting an old riddle: what determines genetic diversity levels within species? PLoS Biol. 10, e1001388 (2012).
Hewitt, G. The genetic legacy of the Quaternary ice ages. Nature 405, 907–913 (2000).
López-Delgado, J. & Meirmans, P. G. History or demography? Determining the drivers of genetic variation in North American plants. Mol. Ecol. 31, 1951–1962 (2022).
Corbett-Detig, R. B., Hartl, D. L. & Sackton, T. B. Natural selection constrains neutral diversity across a wide range of species. PLoS Biol. 13, e1002112 (2015).
Buffalo, V. Quantifying the relationship between genetic diversity and population size suggests natural selection cannot explain lewontin’s paradox. Elife 10, 1–30 (2021).
Laikre, L. et al. Post-2020 goals overlook genetic diversity. Science 367, 1083–1085 (2020).
Wilder, A. P. et al. The contribution of historical processes to contemporary extinction risk in placental mammals. Science 380, eabn5856 (2023).
Alfaro, M. E. et al. Nine exceptional radiations plus high turnover explain species diversity in jawed vertebrates. Proc. Natl Acad. Sci. Usa. 106, 13410–13414 (2009).
Goldberg, E. E. et al. Species selection maintains self-incompatibility. Science 330, 493–495 (2010).
Rabosky, D. L. & McCune, A. R. Reinventing species selection with molecular phylogenies. Trends Ecol. Evol. 25, 68–74 (2010).
Morlon, H. Phylogenetic approaches for studying diversification. Ecol. Lett. 17, 508–525 (2014).
Maliet, O., Hartig, F. & Morlon, H. A model with many small shifts for estimating species-specific diversification rates. Nat. Ecol. Evolution 3, 1086–1092 (2019).
Morlon, H. et al. Phylogenetic Insights into Diversification. Annu. Rev. Ecol. Evol. 55, 1–21 (2024).
Weir, J. T. & Schluter, D. The latitudinal gradient in recent speciation and extinction rates of birds and mammals. Science 315, 1574–1576 (2007).
Rolland, J., Condamine, F. L., Jiguet, F. & Morlon, H. Faster speciation and reduced extinction in the tropics contribute to the Mammalian latitudinal diversity gradient. PLoS Biol. 12, e1001775 (2014).
Rabosky, D. L. et al. An inverse latitudinal gradient in speciation rate for marine fishes. Nature 559, 392–395 (2018).
Hua, X. & Bromham, L. Darwinism for the genomic age: Connecting mutation to diversification. Front. Genet. 8, 1–18 (2017).
Li, J., Huang, J.-P., Sukumaran, J. & Knowles, L. L. Microevolutionary processes impact macroevolutionary patterns. BMC Evol. Biol. 18, 123 (2018).
Harvey, M. G., Singhal, S. & Rabosky, D. L. Beyond Reproductive Isolation: Demographic Controls on the Speciation Process. Annu. Rev. Ecol. Evol. Syst. 50, 75–95 (2019).
Achaz, G. Which Model(s) Explain Biodiversity? in Biodiversity and Evolution (eds. Grandcolas, P. & Maurel, M.-C.) 39–61 (Elsevier, 2018).
Costa, C. L. N. et al. Signatures of Microevolutionary Processes in Phylogenetic Patterns. Syst. Biol. 68, 131–144 (2019).
de Alencar, L. R. V. & Quental, T. B. Linking population-level and microevolutionary processes to understand speciation dynamics at the macroevolutionary scale. Ecol. Evol. 11, 5828–5843 (2021).
Schluter, D. & Pennell, M. W. Speciation gradients and the distribution of biodiversity. Nature 546, 48–55 (2017).
Harvey, M. G. et al. Positive association between population genetic differentiation and speciation rates in New World birds. Proc. Natl Acad. Sci. Usa. 114, 6328–6333 (2017).
Medina, I., Cooke, G. M. & Ord, T. J. Walk, swim or fly? Locomotor mode predicts genetic differentiation in vertebrates. Ecol. Lett. 21, 638–645 (2018).
Singhal, S. et al. Does Population Structure Predict the Rate of Speciation? A Comparative Test across Australia’s Most Diverse Vertebrate Radiation. Am. Nat. 192, 432–447 (2018).
Lanfear, R., Ho, S. Y. W., Love, D. & Bromham, L. Mutation rate is linked to diversification in birds. Proc. Natl Acad. Sci. 107, 20423–20428 (2010).
Goldie, X., Lanfear, R. & Bromham, L. Diversification and the rate of molecular evolution: No evidence of a link in mammals. BMC Evol. Biol. 11, 286 (2011).
Perez-Lamarque, B. et al. Analysing diversification dynamics using barcoding data: The case of an obligate mycorrhizal symbiont. Mol. Ecol. 31, 3496–3512 (2022).
Huang, J.-P., Leavitt, S. D. & Lumbsch, H. T. Testing the impact of effective population size on speciation rates - a negative correlation or lack thereof in lichenized fungi. Sci. Rep. 8, 5729 (2018).
Wright, S. Evolution in Mendelian Populations. Genetics 16, 97–159 (1931).
Charlesworth, B. Fundamental concepts in genetics: Effective population size and patterns of molecular evolution and variation. Nat. Rev. Genet. 10, 195–205 (2009).
Schluter, D. The Ecology of Adaptive Radiation. (OUP Oxford, 2000).
Simpson, G. G. The Major Features of Evolution. (Columbia University Press, 1953).
Feder, J. L. et al. Allopatric genetic origins for sympatric host-plant shifts and race formation in Rhagoletis. Proc. Natl Acad. Sci. Usa. 100, 10314–10319 (2003).
Hermisson, J. & Pennings, P. S. Soft sweeps: molecular population genetics of adaptation from standing genetic variation. Genetics 169, 2335–2352 (2005).
Jones, F. C. et al. The genomic basis of adaptive evolution in threespine sticklebacks. Nature 484, 55–61 (2012).
Schluter, D. & Conte, G. L. Genetics and ecological speciation. Proc. Natl Acad. Sci. Usa. 106, 9955–9962 (2009).
Whittaker, R. H. Evolution and measurement of species diversity. Taxon 21, 213–251 (1972).
Rennison, D. J., Rudman, S. M. & Schluter, D. Genetics of adaptation: Experimental test of a biotic mechanism driving divergence in traits and genes. Evol. Lett. 3, 513–520 (2019).
Rohde, K. Latitudinal Gradients in Species Diversity: The Search for the Primary Cause. Oikos 65, 514–527 (1992).
Allen, A. P., Gillooly, J. F., Savage, V. M. & Brown, J. H. Kinetic effects of temperature on rates of genetic divergence and speciation. Proc. Natl Acad. Sci. Usa. 103, 9130–9135 (2006).
Condamine, F. L., Rolland, J. & Morlon, H. Assessing the causes of diversification slowdowns: temperature‐dependent and diversity‐dependent models receive equivalent support. Ecol. Lett. 22, 1900–1912 (2019).
Gavrilets, S. & Hastings, A. Founder effect Speciation: A theoretical Reassessment. Am. Nat. 147, 466–491 (1996).
Gavrilets, S. A Dynamical Theory of Speciation on Holey Adaptive Landscapes. Am. Nat. 154, 1–22 (1999).
Maya-Lastra, C. A. & Eaton, D. A. R. Genetic incompatibilities do not snowball in a demographic model of speciation. bioRxiv 2021.02.23.432472 (2021) https://doi.org/10.1101/2021.02.23.432472.
Nei, M., Maruyama, T. & Wu, C. I. Models of evolution of reproductive isolation. Genetics 103, 557–579 (1983).
Nei, M. Bottlenecks, genetic polymorphism and speciation. Genetics 170, 1–4 (2005).
Venditti, C. & Pagel, M. Speciation as an active force in promoting genetic evolution. Trends Ecol. Evol. 25, 14–20 (2010).
Manceau, M., Marin, J., Morlon, H. & Lambert, A. Model-Based Inference of Punctuated Molecular Evolution. Mol. Biol. Evol. 37, 3308–3323 (2020).
Przeworski, M., Coop, G. & Wall, J. D. The signature of positive selection on standing genetic variation. Evolution 59, 2312–2323 (2005).
Bickford, D. et al. Cryptic species as a window on diversity and conservation. Trends Ecol. Evol. 22, 148–155 (2007).
Veron, P., Andréoletti, J., Giraud, T. & Morlon, H. Speciation completion rates have limited impact on macroevolutionary diversification. bioRxiv 2024.07.01.601000 (2024) https://doi.org/10.1101/2024.07.01.601000.
Charlesworth, B., Charlesworth, D. & Barton, N. H. The Effects of Genetic and Geographic Structure on Neutral Variation. Annu. Rev. Ecol. Evol. Syst. 34, 99–125 (2003).
Whitlock, M. C. & Barton, N. H. The effective size of a subdivided population. Genetics 146, 427–441 (1997).
Whitlock, M. C. 7 - Selection and Drift in Metapopulations. in Ecology, Genetics and Evolution of Metapopulations (eds. Hanski, I. & Gaggiotti, O. E.) 153–173 (Academic Press, Burlington, 2004).
Miraldo, A. et al. An Anthropocene map of genetic diversity. Science 353, 1532–1535 (2016).
Theodoridis, S. et al. Evolutionary history and past climate change shape the distribution of genetic diversity in terrestrial mammals. Nat. Commun. 11, 1–11 (2020).
Mittelbach, G. G. et al. Evolution and the latitudinal diversity gradient: speciation, extinction and biogeography. Ecol. Lett. 10, 315–331 (2007).
Quintero, I., Landis, M. J., Jetz, W. & Morlon, H. The build-up of the present-day tropical diversity of tetrapods. Proc. Natl Acad. Sci. Usa. 120, e2220672120 (2023).
Castellana, S., Vicario, S. & Saccone, C. Evolutionary patterns of the mitochondrial genome in Metazoa: Exploring the role of mutation and selection in mitochondrial protein-coding genes. Genome Biol. Evol. 3, 1067–1079 (2011).
Bazin, E., Gle, S. & Galtier, N. Mitochondrial Genetic Diversity in Animals. Science 312, 570–572 (2006).
Allio, R., Donega, S., Galtier, N. & Nabholz, B. Large variation in the ratio of mitochondrial to nuclear mutation rate across animals: implications for genetic diversity and the use of mitochondrial DNA as a molecular marker. Mol. Biol. Evol. 34, 2762–2772 (2017).
Mulligan, C. J., Kitchen, A. & Miyamoto, M. M. Comment on ‘Population size does not influence mitochondrial genetic diversity in animals’. Science 314, 1390 (2006).
Nabholz, B., Mauffrey, J.-F., Bazin, E., Galtier, N. & Glemin, S. Determination of mitochondrial genetic diversity in mammals. Genetics 178, 351–361 (2008).
Upham, N. S., Esselstyn, J. A. & Jetz, W. Inferring the mammal tree: Species-level sets of phylogenies for questions in ecology, evolution, and conservation. PLoS Biol. 17, e3000494 (2019).
Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 105, 437–460 (1983).
Watterson, G. A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975).
Ferretti, L., Raineri, E. & Ramos-Onsins, S. Neutrality tests for sequences with missing data. Genetics 191, 1397–1401 (2012).
Maliet, O. & Morlon, H. Fast and Accurate Estimation of Species-Specific Diversification Rates Using Data Augmentation. Syst. Biol. 71, 353–366 (2022).
Ives, A. R. R$^{2}$s for Correlated Data: Phylogenetic Models, LMMs, and GLMMs. Syst. Biol. 68, 234–251 (2018).
Duchêne, S., Lanfear, R. & Ho, S. Y. W. The impact of calibration and clock-model choice on molecular estimates of divergence times. Mol. Phylogenet. Evol. 78, 277–289 (2014).
Condamine, F. L., Nagalingum, N. S., Marshall, C. R. & Morlon, H. Origin and diversification of living cycads: a cautionary tale on the impact of the branching process prior in Bayesian molecular dating. BMC Evol. Biol. 15, 65 (2015).
Barido-Sottani, J. & Morlon, H. The ClaDS rate-heterogeneous birth-death prior for full phylogenetic inference in BEAST2. Syst. Biol. 72, 1180–1187 (2023).
Ritchie, A. M., Hua, X. & Bromham, L. Investigating the reliability of molecular estimates of evolutionary time when substitution rates and speciation rates vary. BMC Ecol. Evol. 22, 61 (2022).
Bielby, J. et al. The fast-slow continuum in mammalian life history: an empirical reevaluation. Am. Nat. 169, 748–757 (2007).
Gardezi, T. & da Silva, J. Diversity in Relation to Body Size in Mammals: A Comparative Study. Am. Nat. 153, 110–123 (1999).
Skeels, A. & Cardillo, M. Reconstructing the Geography of Speciation from Contemporary Biodiversity Data. Am. Nat. 193, 240–255 (2019).
Mayr, E. Change of genetic environment and evolution. (1954).
Gavrilets, S., Li, H. & Vose, M. D. Patterns of parapatric speciation. Evolution 54, 1126–1134 (2000).
Nabholz, B., Glémin, S. & Galtier, N. The erratic mitochondrial clock: variations of mutation rate, not population size, affect mtDNA diversity across birds and mammals. BMC Evol. Biol. 9, 54 (2009).
Pelletier, T. A. & Carstens, B. C. Geographical range size and latitude predict population genetic structure in a global survey. Biol. Lett. 14, 20170566 (2018).
Lawrence, E. R. & Fraser, D. J. Latitudinal biodiversity gradients at three levels: Linking species richness, population richness and genetic diversity. Glob. Ecol. Biogeogr. 29, 770–788 (2020).
Paz-Vinas, I. et al. Macrogenetic studies must not ignore limitations of genetic markers and scale. Ecol. Lett. 24, 1282–1284 (2021).
Piganeau, G. & Eyre-Walker, A. Evidence for variation in the effective population size of animal mitochondrial DNA. PLoS One 4, 2–9 (2009).
Spielman, D., Brook, B. W. & Frankham, R. Most species are not driven to extinction before genetic factors impact them. Proc. Natl Acad. Sci. Usa. 101, 15261–15264 (2004).
R Core Team. R: A Language and Environment for Statistical Computing. (2018).
Schöfl, G. reutils: Talk to the NCBI EUtils. R package version 0.2.3. https://CRAN.R-project.org/package=reutils. 520 (2016).
Meyer, C., Kreft, H., Guralnick, R. P. & Jetz, W. Global priorities for an effective information basis of biodiversity distributions. Nat. Commun. 6, 8221 (2015).
Chamberlain, S. A. & Szöcs, E. taxize: taxonomic search and retrieval in R. F1000Res. 2, 191 (2013).
Michonneau, F., Brown, J. W. & Winter, D. J. rotl: an R package to interact with the Open Tree of Life data. Methods Ecol. Evol. 7, 1476–1481 (2016).
Portik, D. M. & Wiens, J. J. SuperCRUNCH: A bioinformatics toolkit for creating and manipulating supermatrices and other large phylogenetic datasets. Methods Ecol. Evol. 11, 763–772 (2020).
Katoh, K., Misawa, K., Kuma, K.-I. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Keck, F. Handling biological sequences in R with the bioseq package. Methods Ecol. Evol. 11, 1728–1732 (2020).
Ranwez, V., Douzery, E. J. P., Cambon, C., Chantret, N. & Delsuc, F. MACSE v2: Toolkit for the Alignment of Coding Sequences Accounting for Frameshifts and Stop Codons. Mol. Biol. Evol. 35, 2582–2584 (2018).
Upham, N. S., Esselstyn, J. A. & Jetz, W. Molecules and fossils tell distinct yet complementary stories of mammal diversification. Curr. Biol. 31, 4195–4206.e3 (2021).
Jetz, W., Thomas, G. H., Joy, J. B., Hartmann, K. & Mooers, A. O. The global diversity of birds in space and time. Nature 491, 444–448 (2012).
Pinheiro, J. Bates, D. DebRoy, S. Sarkar, D. nlme: linear and nonlinear mixed effects models. R package version 3.1-153. RCore Team. 2021. (2023).
Bürkner, P.-C. brms: An R package for Bayesian multilevel models using Stan. J. Stat. Softw. 80, 1–28 (2017).
Ives, A. R. R$^{2}$s for Correlated Data: Phylogenetic Models, LMMs, and GLMMs. Syst. Biol. 68, 234–251 (2019).
Ives, A. & Li, D. rr2: An R package to calculate R2s for regression models. J. Open Source Softw. 3, 1028 (2018).
Clavel, J. & Morlon, H. Reliable Phylogenetic Regressions for Multivariate Comparative Data: Illustration with the MANOVA and Application to the Effect of Diet on Mandible Morphology in Phyllostomid Bats. Syst. Biol. 69, 927–943 (2020).
Upham, N. S., Esselstyn, J. A. & Jetz, W. Ecological causes of uneven speciation and species richness in mammals. bioRxiv 504803 (2020) https://doi.org/10.1101/504803.
Rolland, J. et al. The impact of endothermy on the climatic niche evolution and the distribution of vertebrate diversity. Nat. Ecol. Evol. 2, 459–464 (2018).
Myhrvold, N. P. et al. An amniote life-history database to perform comparative analyses with birds, mammals, and reptiles. Ecology. 96, 3109–3000 (2015).
Welch, J. J., Bininda-Emonds, O. R. P. & Bromham, L. Correlates of substitution rate variation in mammalian protein-coding sequences. BMC Evol. Biol. 8, 53 (2008).
Yang, Z. Molecular Evolution: A Statistical Approach. (Oxford University Press, 2014).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Acknowledgements
We thank Anne Chenuil, Sylvain Glémin, and Bert Van Bocxlaer for thoughtful and helpful discussions, as well as the members of the BioDiv team at IBENS, particularly Julien Clavel, Benoît Perez-Lamarque, Ignacio Quintero, and Isaac Overcast for further technical advice. HM acknowledges support from the European Research Council (grant CoG-PANDA).
Author information
Authors and Affiliations
Contributions
A.A.S. and H.M. conceived the study. A.A.S. analyzed the data, with help from O.M. and L.A. D.N.B., N.U. and W.J. contributed data and advice. A.O.S. and H.M. wrote the first draft of the manuscript, and all authors contributed substantially to the writing and revisions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Michael Harvey and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Afonso Silva, A.C., Maliet, O., Aristide, L. et al. Negative global-scale association between genetic diversity and speciation rates in mammals. Nat Commun 16, 1796 (2025). https://doi.org/10.1038/s41467-025-56820-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56820-y