Abstract
Psilocybin, ketamine, and MDMA are psychoactive compounds that exert behavioral effects with distinguishable but also overlapping features. The growing interest in using these compounds as therapeutics necessitates preclinical assays that can accurately screen psychedelics and related analogs. We posit that a promising approach may be to measure drug action on markers of neural plasticity in native brain tissues. We therefore developed a pipeline for drug classification using light sheet fluorescence microscopy of immediate early gene expression at cellular resolution followed by machine learning. We tested male and female mice with a panel of drugs, including psilocybin, ketamine, 5-MeO-DMT, 6-fluoro-DET, MDMA, acute fluoxetine, chronic fluoxetine, and vehicle. In one-versus-rest classification, the exact drug was identified with 67% accuracy, significantly above the chance level of 12.5%. In one-versus-one classifications, psilocybin was discriminated from 5-MeO-DMT, ketamine, MDMA, or acute fluoxetine with >95% accuracy. We used Shapley additive explanation to pinpoint the brain regions driving the machine learning predictions. Our results suggest a unique approach for characterizing and validating psychoactive drugs with psychedelic properties.
Similar content being viewed by others
Introduction
Psychedelics include classic serotonergic psychedelics, such as psilocybin and 5-methoxy-N,N-dimethyltryptamine (5-MeO-DMT), and related psychoactive compounds, such as ketamine and 3,4-methyl enedioxy methamphetamine (MDMA). These compounds have recently gained widespread interest as potential therapeutics for neuropsychiatric disorders1,2. Psilocybin with psychological support is under active investigation as a treatment for major depressive disorder and treatment-resistant depression3,4,5,6,7. Subanesthetic ketamine has long been studied for its efficacy in treating depression8,9,10 and post-traumatic stress disorder (PTSD)11. The research efforts culminated in the approval of esketamine nasal spray by the FDA in the United States for treatment-resistant depression12,13. Finally, MDMA-assisted psychotherapy has undergone phase III clinical trials for the treatment of moderate to severe PSTD14,15. The clinical relevance has sparked intense interest in understanding the shared and distinct aspects of these compounds’ mechanisms of action.
Beyond the known psychedelics, there is also growing excitement for synthesizing novel psychedelic-inspired analogs that can be new chemical entities for therapeutics16,17,18. Ideally, the novel compounds would retain therapeutic effects while improving pharmacokinetics, minimizing perceptual effects, and eliminating cardiovascular risks. A major roadblock in this pursuit lies in developing screens that can filter thousands of psychedelic-inspired analogs to a manageable list of the most promising compounds for further in-depth characterizations. Currently, most screens operate at the molecular or behavioral level. At the molecular level, candidate compounds can be docked in silico with the structure of the 5-HT2A receptor, followed by biochemical measurements of receptor engagement and activation of downstream G-protein and beta-arrestin pathways. This target-based approach has yielded exciting leads19,20,21,22, but assumes that the 5-HT2A receptor is the key mediator of the therapeutic effect, which has not been proven conclusively. At the behavioral level, candidate compounds may be tested in animals for defined phenotypes. Simple characterizations such as changes in animal movement patterns may be automated to increase throughput and accuracy23,24. However, more complex behavioral assays relevant for depression suffer from limitations, including poor construct validity and weak predictive power for drug efficacy in humans25.
The development of a new screening method may complement current molecular and behavioral approaches to accelerate preclinical drug discovery. Classic psychedelics and ketamine share the ability to enhance neural plasticity in the brain26, as evidenced by the rapid and persistent growth of dendritic spines in the rodent medial frontal cortex after a single dose of ketamine27,28, psilocybin29, and related serotonergic receptor agonists30,31,32,33. A promising approach may thus focus on quantifying indicators of neural plasticity in native brain tissues. To this end, immediate early genes are activated in a cell in response to increased firing activity or an external stimulus34. The immediate early genes are a key part of neural plasticity, because they enable neurons to adapt to stimuli by regulating gene expression, which is crucial for protein synthesis that is needed for synaptic modifications and learning35,36. Taking classic psychedelics as an example, drug administration induces robust increases in the expression of immediate early genes37,38, including c-Fos, that can be detected starting in as few as 30 minutes in multiple brain regions39,40. More recently, technological advances in tissue clearing, light sheet fluorescence microscopy, and automated detection of nuclei have enabled high-throughput mapping of the expression of immediate early genes such as c-Fos in the whole mouse brain41,42. We and others have applied this method to characterize the impact of psilocybin and ketamine43,44,45, joining a rapidly growing number of studies using brain-wide imaging of fluorescence signals to study drugs46,47,48,49,50,51,52,53,54,55,56,57. Although these early studies have provided valuable biological insights, only one or two drugs were typically included in each study thus far. Developing the method as a drug screen requires evaluating its feasibility and accuracy on a larger panel of compounds.
In this study, we measured brain-wide c-Fos expression in male and female mice for 8 drug conditions, including a variety of psychedelics, related psychoactive compounds, and vehicle control. We developed a pipeline for analysis and classification based on explainable machine learning, determining performance in one-versus-rest and one-versus-one classification tasks. We implemented Shapley additive explanation to interpret the machine learning models to identify the brain regions driving the classifications. Collectively the results demonstrate brain-wide imaging of immediate early gene expression as a promising approach for preclinical drug discovery.
Results
Psychedelics and related drugs in the study
For this study, we evaluated 8 drug conditions: psilocybin (PSI, 1 mg/kg, i.p., single dose), ketamine (KET, 10 mg/kg, i.p., single dose), 5‐methoxy‐N,N‐dimethyltryptamine (5-MeO-DMT or 5-MEO, 20 mg/kg, i.p., single dose), 6-fluoro-N,N-diethyltryptamine (6-fluoro-DET or 6-F-DET, 20 mg/kg, i.p., single dose), 3,4-methylenedioxy methamphetamine (MDMA, 7.8 mg/kg, i.p., single dose), acute fluoxetine (A-SSRI, 10 mg/kg, i.p., single dose), chronic fluoxetine (C-SSRI, 10 mg/kg, i.p., one dose every day for 14 days), and saline vehicle (SAL, 10 mL/kg, i.p., single dose) (Fig. 1a).
a Chemical structures for the 8 conditions included in this study: psilocybin (PSI), ketamine (KET), 5-MeO-DMT (5-MEO), 6-fluoro-DMT (6-F-DET), MDMA, acute fluoxetine (A-SSRI), chronic fluoxetine (C-SSRI, daily for 14 days), and saline vehicle (SAL). b Time course of head-twitch response following the administration of 5-MeO-DMT, psilocybin, 6-fluoro-DET, or saline vehicle. Line, mean. Shading, 95% confidence interval based on 1000 bootstraps. 3 males and 3 females for each drug (n = 6), except 4 males and 3 females for saline (n = 7). c Box plot of the total number of head twitches detected within a 2-hour period after drug administration. n = 6 for each drug, n = 7 for saline vehicle. The data centre represents median values, bounds of the box represent the 25th and 75th percentile. Whiskers extend to 0th and 100th percentiles, excluding outliers (points beyond median ±1.5 * interquartile range). Wilcoxon rank-sum test is two-sided. *P < 0.05, **P < 0.01. (SAL vs. 5-MEO: P = 3 × 10−3, 6-F-DET vs. 5-MEO: P = 2 × 10−2, SAL vs. PSI: P = 3 × 10−3, 6-F-DET vs PSI: P = 8 × 10−3). d Experimental timeline. e Box plot of the total number of c-Fos+ cells in the brain for each drug condition. Data center represents median values, bounds of box represent 25th and 75th percentile. Cross, female individual. Circle, male individual. N = 64 mice, including 4 males and 4 females for each condition. f An example of the fluorescence images of c-Fos+ cells in the mouse brain for a psilocybin-treated mouse acquired by light sheet fluorescence microscopy. Inset, magnified view of the dorsal anterior cingulate cortex. Image is comparable to images seen in other 7 samples collected from mice treated with psilocybin. b, c the psilocybin and saline vehicle data had been shown in a prior study33.
We elected to investigate these compounds for several reasons. Psilocybin is a classic psychedelic that acts on the 5-HT2A receptor. Psilocybin stands at the forefront of ongoing late-stage clinical trials evaluating psychedelics’ efficacy for treating depression3,4,5,6,7. Ketamine is primarily an NMDA receptor antagonist58. Despite the distinct molecular targets, ketamine and psilocybin have similarities in their plasticity-promoting action and behavioral effects59,60, making ketamine an intriguing compound to contrast with psilocybin. The doses and route of administration for psilocybin and ketamine were chosen based on prior studies showing behavioral effects in mice29,61.
5-MeO-DMT is a classic serotonergic psychedelic in the same tryptamine chemical class as psilocybin16. There is clinical interest in evaluating 5-MeO-DMT as a treatment for depression62,63. At a dose of 20 mg/kg in mice, 5-MeO-DMT induces head-twitch response and evokes structural rewiring in the mouse medial frontal cortex33. Compared to psilocybin, 5-MeO-DMT is shorter-acting and has a higher affinity for the 5-HT1A receptor than for the 5-HT2A receptor. Thus 5-MeO-DMT serves as a useful case of another tryptamine psychedelic with distinct pharmacokinetics and receptor target profile. 6-fluoro-DET is also a tryptamine like psilocybin and 5-MeO-DMT. Although bioavailable in the brain and a 5-HT2A receptor agonist64,65, 6-fluoro-DET induces autonomic effects without causing perceptual changes in humans66. Therefore, it has been used as an active, non-hallucinogenic control in a clinical study67. Concordantly, 6-fluoro-DET provided ineffective as a substitute compound for rats trained to discriminate LSD or 2,5-dimethoxy-4-iodoamphetamine (known as DOI)64,68. To corroborate these prior results, we measured the effect of 6-fluoro-DET on head-twitch response in mice using magnetic ear tags for automated detection of head movements. Our results showed that, unlike 1 mg/kg psilocybin and 20 mg/kg 5-MeO-DMT which elicited robust head-twitch responses33, mice administered with 20 mg/kg 6-fluoro-DET were not statistically different from controls (Fig. 1b, c). Our study adds to other recent studies20,21 that included 6-fluoro-DET as a non-hallucinogenic tryptamine for comparison. The dose of 6-fluoro-DET was chosen to match the dose of 5-MeO-DMT.
MDMA is different from psilocybin: it is a member of the phenethylamine chemical class and has distinct pro-social and euphoric qualities69. MDMA can act on monoamine transporters to enhance release and inhibit reuptake of neuromodulators including serotonin, thus it has been characterized as an entactogen rather than a classic serotonergic psychedelic70. MDMA holds clinical relevance, particularly for PTSD14,15. We selected a dose of 7.8 mg/kg for MDMA based on prior work showing that this dose facilitates fear extinction learning in mice71. Fluoxetine is a commonly prescribed antidepressant that is a selective serotonin reuptake inhibitor (SSRI). The clinical interest lies in understanding the relative efficacies of SSRIs versus psilocybin4 and whether ketamine or psilocybin is suitable for treatment-resistant depression5,12,13. SSRIs require chronic administration to exert therapeutic effects, therefore likely engage a mechanism of action distinct than that of psilocybin and ketamine. For these reasons, we included acute and chronic fluoxetine in this study. We chose a dose of 10 mg/kg, which was used for acute and chronic administration of fluoxetine in mice previously72,73. Control animals received a single injection of saline vehicle.
Light sheet fluorescence imaging of cellular c-Fos expression
For each of the 8 drugs, we tested 4 male and 4 female C57BL/6 J mice, totaling 64 animals for the entire data set. Brains were collected 2 hours after the administration of the single dose or 2 hours after the administration of the last dose for the chronic fluoxetine condition (Fig. 1d). The 2-hour interval was chosen assuming drug penetrance to the brain by 0.5 hours and peak c-Fos expression after an additional 1.5 hours74. Brains were processed for tissue clearing and c-Fos immunohistochemistry (see Methods). Light sheet fluorescence microscopy was used to image each brain at a resolution of 1.8 µm per pixel in the x- and y axis and at 4 µm intervals in the z-axis, which allowed for sampling of all cells in the entire brain without any gap. The images were analyzed using neural nets for automated detection of fluorescent puncta corresponding to c-Fos+ cells (see Methods). The number of c-Fos+ cells detected in each brain for each condition is presented in Fig. 1e. An example image collected from a mouse administered with psilocybin is shown in Fig. 1f.
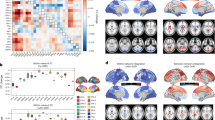
To investigate the regional distribution of c-Fos+ cells, we aligned the images of each brain to the Allen Brain Atlas and segmented the images into summary structures based on the Allen Mouse Brain Common Coordinate Framework75 (see Methods; Supplementary Data 1). The number of c-Fos+ cells in each brain region for all animals is provided as source data. To visualize the entire data set, we normalized the c-Fos+ cell count in each brain region by the total number of c-Fos+ cells of each brain and by the spatial volume of the brain region. Figure 2 is a heatmap of the resulting c-Fos+ cell density for all the samples. We observed that c-Fos+ cell density was generally high in the isocortex, olfactory area, hippocampal area, striatum and pallidum, and thalamus, whereas expression was lower in the midbrain and hindbrain, and cerebellum. There were individual differences across samples from the same drug, but also notable contrasts across different drugs. This begets questions such as: How does the individual variability compare with the differences across drugs? How well can whole-brain c-Fos maps be used to discriminate the different drugs?
The c-Fos+ cell density was defined as the c-Fos+ cell count in each brain region divided by the total number of c-Fos+ cells in each brain and the spatial volume of the brain region. The pixels in the heatmap are positioned by brain region (row) and animal grouped by drug (column). The intensity of the pixel is pseudo-colored by the value of the c-Fos+ cell density. 1st percentile and 99th percentile values are used to define extremes of the colormap. The brain regions, including acronyms and other details, are provided in Supplementary Data 1.
Machine learning pipeline for classifying drugs based on brain-wide c-Fos distribution
To answer these questions, we developed a pipeline for quantitative comparison of the brain-wide c-Fos expression data between different drug conditions. We posited that different compounds may elicit distinct regional distributions of cellular c-Fos expression that can serve as fingerprints for classifying drugs. The pipeline starts with a matrix of c-Fos+ cell counts for different brain regions from different samples (first panel, Fig. 3a). This matrix of c-Fos+ cell counts undergoes preprocessing, starting with normalization (dividing the c-Fos+ cell count in each region by the total c-Fos+ cell count of the brain) (second panel, Fig. 3a). Normalization is important because there may be batch effects across samples. The data were then processed to scale the input data to a standard range such that the values across brain regions are more comparable and amenable to fitting machine learning models (second panel, Fig. 3a), using Yeo-Johnson transformation (monotonic transformation of data using a power function) and robust scaling (median subtraction and interquartile range scaling). We will herein refer to the values after this preprocessing step as the c-Fos scores.
a The pipeline consisted of three steps. First, c-Fos+ cell counts for each brain region undergo normalization, Yeo-Johnson transformation, and robust scaling, into c-Fos scores. Second, the Boruta procedure is used to select the set of informative brain regions. Third, c-Fos scores from this set of brain regions were used to fit a ridge logistic regression model. For each iteration, 75% of the data in each drug condition were used for region selection and training through the three steps, and the remaining 25% of the data were withheld initially, but then processed and tested with the ridge logistic regression model. The entire process was iterated using different splits of the data 100 times. b Linear discriminant analysis of the c-Fos scores to visualize the data in a low-dimensional space. c The confusion matrix shows the mean proportion of predicted labels for each of the true labels across all splits. d The composite precision-recall curves for each drug condition across all splits and the grand average across all drugs. The values in parentheses are the area under the precision-recall curve for each condition.
Next, we adapted the Boruta feature selection procedure19 to determine which brain regions to include for model fitting and testing (third panel, Fig. 3a). The Boruta procedure is a permutation-based method for determining feature importance. It starts by creating “shadow features”: for example, if the data contains 48 c-Fos scores for brain region 1 for various conditions, then the corresponding shadow feature will be those same 48 c-Fos scores with scrambled drug labels. Shadow variants were created for all brain regions to create the expanded Boruta dataset. A random forest classifier was built using this Boruta dataset to determine a feature-importance value for each brain region. If a brain region has a higher feature-importance value than the largest feature-importance value from shadow features, then brain region 1 is a “hit”. This permutation process is iterated 100 times. Given that each brain region can achieve only one of two outcomes (hit or no hit) in each iteration, the distribution of outcomes across all iterations is a binomial distribution, and a brain region is included by the statistical criterion of exceeding the 95th percentile of the binomial distribution. Why Boruta? We used the Boruta procedure in lieu of including all brain regions, because many regions likely contribute little or nothing towards differential drug action and their inclusion in the model would increase noise and lead to overfitting. A distinctive advantage of Boruta is that brain regions do not compete with each other, but rather with the shadows. As a result, the number of brain regions selected by Boruta is not pre-determined but instead dictated by the data as needed.
For the last step, the c-Fos scores from the selected brain regions are used to construct a ridge logistic regression model (fourth panel, Fig. 3a). The entire pipeline is evaluated using fourfold splits, where 75% of the data in each drug condition was used to train and fit the model, while the remaining 25% of the data is used to test the model. Importantly, we emphasize that we used only the training data to optimize the preprocessing parameters, run feature selection, and construct a regression model. The same optimized preprocessing parameters and selected features were then later applied to the test data, ensuring no data leakage. The splits were repeated 100 times to evaluate the prediction accuracy of the pipeline.
One-versus-rest classification shows drug prediction accuracy well above chance
We performed a linear discriminant analysis on the c-Fos scores of all 64 samples, just after the preprocessing step. We plotted the data for the top two linear discriminants (Fig. 3b). This visualization clearly shows that the differences in c-Fos scores across drugs are more separable than the differences in c-Fos scores across samples within the same drug condition. Drugs that alter the serotonergic tone via different mechanisms of action are positioned differently along the first linear discriminant. By contrast, 5-MeO-DMT, 6-fluoro-DET, and psilocybin are separable along the second linear discriminant.
We first tested the pipeline with the entire data set and asked the models to predict the exact drug condition. The confusion matrix shows how the predicted drug labels compared with the true drug labels (Fig. 3c). Because there were 8 conditions, the chance-level accuracy was 12.5% (1 out of 8). We found that the model was the most accurate at identifying the MDMA, chronic fluoxetine, and 5-MeO-DMT samples, with 100%, 89%, and 81% accuracy respectively. Performance for other conditions was lower, yielding an overall mean accuracy of 67% for all drugs. Performance was the lowest for saline and acute fluoxetine at 38% and 47%, respectively. Our interpretation of the low-performance conditions is that tradeoffs must be made to solve this 8-way classification problem. The machine learning model uses the cross-entropy loss function, which seeks to maximize the probability of labelling training data correctly across the entire training set rather than drawing boundaries in a one-vs-rest fashion. In this global approach, individual decision boundaries may be placed in a way that performs on one label, such as saline, while leading to a greater improvement in others. In other words, the model was fitted with the goal of maximizing the overall mean classification accuracy, which was not necessarily the most ideal for distinguishing any one specific condition such as saline. Nevertheless, the mean accuracy of 67% was still substantially higher than the chance level of 12.5%.
Confusion matrices are calculated based on a single decision threshold, which may exaggerate the true positive rate for one drug type at the expense of more false positives for another drug type. To understand our model performance from a different perspective, we plotted precision-recall curves (Fig. 3d). These curves consider performance across all possible decision thresholds and summarize the results in terms of precision (true positives relative to false positives) and recall (true positives relative to false negative). The perfect classifier would have an area under the precision-recall curve (precision-recall AUC) of 1. Across all drugs, the pipeline yielded a mean precision-recall AUC value of 0.75. This is well above the theoretical chance level of 0.125 for 1 out of 8 drugs and the empirical chance level of 0.12 calculated with shuffled data. The performance based on precision-recall AUC for predicting different drugs corresponds in rank order to the accuracy in the confusion matrix. Overall, these results provide evidence that brain-wide c-Fos maps can be leveraged to identify the exact drug administered out of a panel of related psychoactive compounds.
A likely use case for the pipeline is to determine how a novel chemical entity may be positioned in the pharmacological space based on the c-Fos expression pattern. To simulate this scenario, we performed a leave-one-drug-out analysis, in which we trained a model using seven conditions (psilocybin, ketamine, 5-MeO-DMT, MDMA, acute fluoxetine, chronic fluoxetine, and saline), but then tested it on all conditions including 6-fluoro-DET. We found that 6-fluoro-DET was most frequently classified as psilocybin at 44% chance but could also be detected as saline at 29% chance (Fig. S1), which is in general agreement with 6-fluoro-DET being a non-hallucinogenic 5-HT2A receptor agonist.
One-versus-one classification suggests a small list of brain regions drives drug prediction
We reasoned that one-versus-one classification, where the machine learning pipeline solves a binary problem of deciding between two drugs (Fig. 4a), may provide deeper insights into the factors that distinguish specific drug classes. Given the prominence of psilocybin in clinical trials and drug discovery, we were particularly interested in comparisons between psilocybin and other conditions that differ in serotonergic receptor affinities (5-MeO-DMT), mechanism of action (MDMA, acute fluoxetine, ketamine), or hallucinogenic potency (6-fluoro-DET). We trained the same machine-learning pipeline using subsets of data involving only two or three drugs. The binary classifiers achieved near-perfect accuracy reflected by precision-recall AUC values at or exceeding 0.90, with the notable exception of psilocybin versus 6-fluoro-DET which had a precision-recall AUC of 0.59 (Fig. 4b). The difficulty in discerning between a classic serotonergic psychedelic and the non-hallucinogenic 5-HT2A receptor agonist extended beyond psilocybin: 5-MeO-DMT versus 6-fluoro-DET as well as psilocybin and 5-MeO-DMT versus 6-fluoro-DET also yielded modest precision-recall AUC values at 0.80 and 0.57 respectively, relative to chance level of 0.5 for one-versus-one classifications. These results suggest that brain-wide cellular c-Fos expression is effective at discriminating between exemplars from different drug classes, such as a classic psychedelic versus an entactogen, a classic psychedelic versus a dissociative, and a classic psychedelic versus SSRI. It also effectively distinguishes between the two classic psychedelics psilocybin and 5-MeO-DMT. However, the prediction is less reliable for the specific problem of predicting a non-hallucinogenic 5-HT2A receptor agonist relative to a classic psychedelic.
a Schematic illustrating the one-versus-one classification problem. b The mean area under the precision-recall curve across all 100 splits for different binary classifiers. Dark gray, real data. Light gray, shuffled data. c Violin plot representing the number of brain regions selected via the Boruta procedure for inclusion in the regression model (n = 100 splits for each classifier). The data centre represents median values, bounds of the inset represent the 25th and 75th percentile. Whiskers extend to 0th and 100th percentiles, excluding outliers (points beyond median ±1.5 * interquartile range). d Heatmaps showing the fraction of splits when a cortical (left) or thalamic (right) region was included in the regression model. The regions are sorted based on usage in all classifiers. Regions that were included in <75% of the splits across all conditions are not shown.
As mentioned, a feature of the Boruta procedure is that a different number of regions may be included depending on the data and the desired classification. Indeed, there were differences in the brain regions chosen for the various drug prediction problems and different training and testing splits of the same data (Fig. 4c). Most classifiers relied on <35 brain regions for drug prediction, except for the two comparisons involving MDMA which included ~40–70 brain regions. Furthermore, we plotted how often various cortical and thalamic regions were selected by the machine learning models (Fig. 4d). Regions such as retrosplenial areas (RSPd, RSPv), somatosensory areas (SSp-m, SSp-tr, SSp-II), and lateral networks (VISC, AId) were included often, but different classifiers relied on them to different extents. We will explore the importance of specific brain regions quantitatively in the next section using Shapley additive explanation. Many thalamic regions were consistently included in comparisons involving MDMA, which contributed to the higher total number of brain regions used by classifiers when MDMA was involved. Overall, the results suggest that one-versus-one drug classifications based on brain-wide c-Fos expression are highly accurate, with the machine learning models only needing data from a small number of brain regions to produce the prediction.
Using Shapley additive explanation to highlight key brain regions driving drug prediction
A brain region selected by Boruta in the pipeline suggests that it is informative, yet it does not communicate the importance of its contribution to the final prediction. To better understand how the c-Fos scores in individual brain regions contribute to decisions in one-versus-one drug classifications we used Shapley additive explanation (SHAP) (Fig. 5a). SHAP uses a game-theoretical approach to determine how the brain regions contribute to driving the machine learning regression model from a starting base value to the final output value for decision21. To illustrate, we present the force plot of two test brain samples in one of our cross-validation splits (Fig. 5b). The top half of the plot shows the c-Fos scores in selected brain regions for the sample of psilocybin and their additive contributions to the decision. In this instance, regions such as posteromedial visual area (VISpm, c-Fos score = 0.44) and lateral habenula (LH, c-Fos score = −0.78) were among the drivers leading to an overall positive SHAP value to predict psilocybin. The posteromedial visual area is located between the primary visual cortex and retrosplenial cortex76 and has been suggested to mediate visual information between the neighboring regions77. Lateral habenula neurons had spiking activity associated with undesirable outcomes78,79, which is consistent with their posited role in mediating depression-related symptoms80 and contributing to antidepressant response81. Intriguingly, another driver was the parafascicular nucleus (PF, c-Fos score = −1.74), which is implicated in arousal and head movements82. By contrast, the c-Fos scores in the same set of selected brain regions sums to an overall negative SHAP value for the 5-MeO-DMT sample, providing the basis for the correct prediction in this case. Across all splits tested for the psilocybin-versus-5-MeO-DMT comparison, we identified regions that were included in >75% of the machine learning models, and then ranked these regions by mean SHAP value difference, which highlight the brain regions most responsible for driving the classification (Fig. 5c, d).
a Diagram illustrating the concept behind SHAP values. The ridge regression model is akin to a black box that takes c-Fos scores as inputs to produce a prediction. SHAP values can be computed to quantitatively assess how strong and in what direction the c-Fos score of each brain region contributes to the prediction. b Example force plots for a psilocybin sample and a 5-MeO-DMT sample from one split, illustrating how actual c-Fos scores of brain regions add to shift the model’s output from the base value to the final value. c Plot relating a region’s c-Fos scores to the SHAP values across individual splits of the 100 iterations for the 5-MeO-DMT-versus-psilocybin classification. Brain regions were shown only if they were used by ≥75% of the splits and listed in rank order by the absolute value of the mean difference in SHAP values between the two drug conditions. The values in parentheses are the absolute value of the mean difference in SHAP values between the two drug conditions. d Visualization of the brain regions included in c, color-coded according to the compound which evoked higher c-Fos score in the region.
We also analyzed other one-versus-one classification problems using Shapley additive explanation. For MDMA versus psilocybin, there was a longer list including 32 brains regions that were used in at least 75% of the cross-validation splits (Fig. 6a, b). Half of these regions (16/32) were in the thalamus. Given the larger number of regions in each model, the SHAP value differences tended to be smaller, because there is redundancy in the information provided by the regions.
For ketamine versus psilocybin, the top 5 regions that were consistently included in >96% of the cross-validation splits and had the highest SHAP value differences were the visceral area (VISC), gustatory area (GU), dorsal agranular insular area (AId), xiphoid thalamic nucleus (Xi), and nucleus of reuniens (RE) (Fig. 6c, d). VISC and GU have direct connections to AId, all of which are part of the lateral subnetworks of the mouse neocortex83,84. The mouse insular cortex contains various cell types that express an abundance of 5-HT2A and 5-HT1A receptors85, which may predispose it to stronger activation by psilocybin. Indeed, the higher c-Fos scores in these lateral cortical regions informed the model to predict psilocybin. Of note, the insular cortex is considered a core region in the mouse homolog of the salience network86,87, which has been implicated in mood regulation and depression in humans88. Xi and RE are part of the midline thalamus, which receives visual inputs to mediate behavioral responses to threat89. Interestingly, higher c-FOS scores in these midline thalamic regions were routinely used by machine learning models to predict ketamine.
Finally, we also plotted SHAP value differences comparing acute fluoxetine and psilocybin (Fig. 6e, f). Here, the strongest differences were detected by in regions involved in somatosensation and motor control, including cortical somatosensory regions (SSs, SSp-m), primary motor cortex (MOp), substantia nigra (SNr), and caudoputamen (CP). These effects may relate to the previously noted effects of psychedelic on the integration of tactile sensory inputs90. Other implicated regions are the interpeduncular nucleus (IPN) and medial mammillary nucleus (MM), which are deep midbrain regions that are components of the limbic midbrain circuitry with long-range connections to the habenula, amygdala, and hippocampus.
Discussion
In this study, we evaluated the possibility of using whole-brain imaging of cellular c-Fos expression for drug classification. We developed a machine learning pipeline with key features including adapting the statistical Boruta procedure to select informative brain regions and using Shapley additive explanation to identify features that drive the classifications. We tested the approach using 64 mice that were administered with a panel of psychedelics and related psychoactive drugs. The results demonstrated high accuracy in various one-versus-rest and one-versus-one classification problems, supporting the utility of the approach for preclinical drug discovery. For dissemination, the data and code are available at a public repository.
Immunohistochemistry can be influenced by factors such as fixation method, incubation time, antibody quality, and antigen retrieval techniques. Consequently, the c-Fos antibody staining can differ from sample to sample. Here, the issue of inter-sample variability was mitigated by not using the absolute c-Fos+ cell counts for analysis, but instead using the proportional distribution in each brain region by dividing c-Fos+ cell counts in each region by the total count in each brain. For instance, if the entire brain was stained poorly and the total c-Fos+ cell count is low, the proportion distribution should remain unchanged. This normalization step is possible when whole-brain data is acquired via light sheet fluorescence microscopy. Experimentally, the variation in antibody staining is also reduced because active electrotransport methods were used for immunolabeling. Although the normalization step is expected to help with inter-sample variability, we note that the 64 samples were processed for imaging over 3 batches (details are provided in “Methods”), and some differences may arise from batch effects.
On average, only a small number of brain regions (~25 brain regions, except for the two comparisons involving MDMA which included ~50 brain regions) out of the >300 summary structures in the brain were included in the machine learning models. From our prior study comparing psilocybin and ketamine43, we know that both compounds induce increases in c-Fos+ expression in numerous brain regions including dorsal and ventral anterior cingulate cortex (ACAd, ACAv), prelimbic area (PL), primary visual cortex (VISp), retrosplenial cortex (RSP), mediodorsal thalamus (MD), locus coeruleus (LC), lateral habenula (LH), claustrum (CLA), basolateral amygdala (BLA), and central amygdala (CEA). These brain regions are likely important for drug action, but shared targets of ketamine and psilocybin are not helpful for distinguishing the compounds. By design, the machine learning pipeline emphasizes brain regions with c-Fos expression changes that can discriminate between drug conditions, for which we found a short list of brain regions.
We anticipate the pipeline to be useful for classifying new chemical entities. For instance, when a novel psychedelic-inspired compound is synthesized, we may predict its action in the brain by its position in the linear discriminant axes (Fig. 3b) and the proximity to existing drug labels (Fig. 3c). We simulated how such a scenario could work by fitting the pipeline with 7 compounds and testing 6-fluoro-DET as if the classifier has never seen it previously (Fig. S1). For the full panel of drugs tested, we show that the exact drug could be identified with mean accuracy of 67%, significantly above the chance level of 12.5%. It is instructive to ask how the pipeline’s performance compared with other approaches to classify drugs. For humans, psilocybin, ketamine, and MDMA exert comparable acute behavioral effects in metrics such as experience of unity, oceanic boundlessness, and changed the meaning of percepts69. However, MDMA preferentially induce blissful state, whereas ketamine evokes disembodiment and psilocybin induces elementary imagery and audio-visual synesthesia69,91. In one study, human participants were asked to guess the administered drug, choosing between mescaline (500 mg and 300 mg), LSD, and psilocybin92. The accuracy for identifying the correct drug ranged from 48% to 58% during the session and 69% to 81% after the study. For animals, there has been recent progress in capturing videos of freely moving mice and analyzing their motion using unsupervised machine learning methods. One study used motion sequencing method to investigate a larger panel of 30 psychoactive compounds and doses from a wide range of drug classes including benzodiazepines, antidepressants, antipsychotics, and stimulants (but not psychedelics and the compounds tested in the current study) to show a F1 precision-recall score of 0.6223. Our pipeline is based on brain-wide cellular c-Fos expression and machine learning, therefore, performed at a level comparable to earlier methods based on human and animal behaviors.
As with any analysis pipeline, there are methodological choices that can affect the outcome, which can plague the interpretation as demonstrated in the field of neuroimaging93. Our codebase is available online for anyone to freely use, adapt, and test. We used a statistical method with the Boruta algorithm, rather than a strict threshold, for region selection. We were careful about data leakage, using only the training data for parameter optimization and feature selection, such that the prediction accuracy for test data would not be inflated. We implemented Shapley additive explanation to decipher the factors driving the decisions, which is a general approach that should find great utility in neuroscience94, and has already seen applications in behavioral classification95 and spike waveform analyses96. There are areas of improvement for the pipeline. While we opted for the simplicity of treating each brain region on its own, regions may have correlated responses to drug administration. There may be biological reasons, such as anatomical proximity or synaptic connectivity, for clustering brain regions prior to region selection, which may outperform our procedure. Network analyses may be used to explore potentially correlated responses to drugs. Furthermore, the pipeline will benefit from testing a larger range of compounds including enantiomers, other drug classes, and different doses. The drugs may be administered in conjunction with a receptor antagonist and a stress or behavioral manipulation, which will all lead to a richer and more refined picture of the ‘drug space’. Finally, c-Fos is one immediate early gene. It is well characterized as an activity-dependent gene and has the advantage of nuclear labeling that permits automated detection. However, there are other immediate early genes and plasticity-related biomarkers that can provide complementary information.
Here we only demonstrated moderate throughput by performing the whole-brain imaging approach for a sample size of 64 brains. This falls short of other current screening methods, which typically involve hundreds of conditions including more compounds, different doses, and additions of antagonists for competitive assays. For whole-brain imaging, the main issue was cost, which precluded us from testing at a larger scale. At the moment, the drug injection and tissue extraction steps are straightforward. The cell counting procedure is mostly automated. However, the cost per brain is high due to tissue processing and imaging, which may drop in the future because of the rapid advances in brain-clearing methods97 and the development of inexpensive light sheet fluorescence microscopes98,99. Thus, there is hope that whole-brain imaging can become a practical method for screening drugs within the next several years.
In summary, there is intense interest in using psychedelics for the treatment of neuropsychiatric disorders. Progress hinges on knowing more about existing psychedelics and finding new psychedelic-inspired drugs with improved characteristics. However, there is currently a paucity of reliable methods to screen psychedelics and related analogs. Here we developed and characterized an approach based on whole-brain imaging of cellular c-Fos expression. We demonstrated high prediction accuracy for drug classifications using a machine-learning pipeline. We expect this and other neuroscience-based approaches to play an important role in accelerating the preclinical development of psychiatric drugs.
Methods
Animals
We used adult, 8-week-old male and female C57BL/6 J mice (#00064, The Jackson Laboratory). Animals were housed in groups with 2—5 mice per cage in a temperature-controlled room, operating on a normal 12 hr light–12 hr dark cycle (8:00 AM to 8:00 PM for light). Food and water were available ad libitum. Tissues were collected and imaged in three batches. The first batch performed in August 2021 included 2 males and 2 females for psilocybin (1 mg/kg, i.p.), 2 males and 2 females for ketamine (10 mg/kg, i.p.), and 2 males and 2 females for saline (10 mL/kg, i.p.). Data from these mice were included in a previous study43. The second batch performed in May 2022 included 2 males and 2 females for psilocybin (1 mg/kg, i.p.), 2 males and 2 females for saline (10 mL/kg, i.p.), 4 males and 4 females for 5-MeO-DMT (20 mg/kg, i.p.), 4 males and 4 females for 6-fluoro-DET (20 mg/kg, i.p.), 4 males and 4 females for acute fluoxetine (10 mg/kg, i.p.), 4 males and 4 females for chronic fluoxetine (10 mg/kg, i.p.; daily for 14 days). The third batch performed in December 2022 included 4 males and 4 females for MDMA (7.8 mg/kg, i.p.) and 2 males and 2 females for ketamine (10 mg/kg, i.p.). All animals were housed and handled according to protocols approved by the Institutional Animal Care and Use Committee (IACUC) at Yale University and Cornell University. Tissue collection for all batches was done at Yale University, except for ketamine in the third batch, which was done at Cornell University. For all batches, the brain samples were shipped for clearing and imaging at LifeCanvas Technologies (Cambridge, MA).
Drugs
Psilocybin, 5-MeO-DMT succinate, and 6-fluoro-DET solids were obtained from Usona Institute’s Investigational Drug & Material Supply Program. We used the succinate salt form of 5-MeO-DMT100 (at equivalent amount to freebase 5-MeO-DMT) because it can be dissolved in saline. Ketamine hydrochloride injection vial (055853, Henry Schein; or Dechra), fluoxetine hydrochloride solid (F132, Millipore-Sigma), 3,4-MDMA hydrochloride (13971, Cayman Chemical), and saline (NDC: 0409-4888-03, Hospira) were purchased from supply vendors. Psilocybin, 5-MeO-DMT succinate, 6-fluoro-DET, MDMA, and fluoxetine were prepared by dissolving powders into saline. Ketamine was prepared by diluting from the injection vial. For ketamine, 5-MeO-DMT succinate, 6-fluoro-DET, MDMA, and acute fluoxetine, the working solutions were prepared fresh on the day of the experiment. For psilocybin, a stock solution was made, and then the working solution was made from the stock solution, with both solutions prepared within 1 month from the day of the experiment. For chronic fluoxetine, the working solution was prepared on the first day of administration and then kept in 4 °C and used for the remainder of the chronic treatment.
Tissue collection and imaging
All the samples underwent the same tissue collection and imaging protocols. Two hours following the single-dose injection or injection of the last dose for chronic fluoxetine, mice were deeply anesthetized with isoflurane and transcardially perfused with phosphate-buffered saline (P4417, Sigma-Aldrich) followed by paraformaldehyde (PFA, 4% in PBS). Brains were fixed in 4% PFA for 24 hours at 4 °C, after which they were transferred to 0.1% sodium azide in PBS for storage until clearing. The SHIELD protocol was used to process the whole mouse brains. A stochastic electrotransport device101 was used to clear samples for 4 days at 42 °C, followed by active immunolabeling using eFLASH technology integrating electrotransport101 and SWITCH102. Each brain sample was stained with 3.5 μg of rabbit anti-c-Fos monoclonal antibody (Abcam, #ab214672), followed by 10 μg of mouse anti-NeuN monoclonal antibody (Encor Biotechnology, #MCA-1B7) and then by fluorescently conjugated secondaries in 1:2 primary:secondary molar ratios (Jackson ImmunoResearch). Following active labeling, refractive index matching (n = 1.52) was done through incubation in EasyIndex (LifeCanvas Technologies). Samples were then imaged at ×3.6 magnification with a SmartSPIM light sheet fluorescence microscope (LifeCanvas Technologies) at a resolution of 1.8 µm/pixel for XY sampling with 4 µm step size for Z sampling over the entire brain. Imaging was done blinded to treatment conditions.
Atlas registration and cell counting
Fluorescence images were tile-corrected, de-striped, and registered to the Allen Brain Atlas using an automated process. For each brain, the image from the NeuN channel was registered to 8–20 atlas-aligned reference samples using SimpleElastix103, which implemented successive rigid, affine, and b-spline warping algorithms. The final atlas alignment value for each sample was determined by taking the average alignment generated across intermediate reference samples. Cell detection was automated by using a custom convolutional neural network design using the TensorFlow Python package. First, a U-Net-based detection network was used to locate fluorescent puncta corresponding to c-Fos-immunolabeled cells. Second, a ResNet-based network was used to filter putative cells to arrive at a final list of cell locations. Each cell location was projected onto the Allen Brain Atlas to identify its anatomical region. We segmented the brain into 316 summary structures based on the Allen Mouse Brain Common Coordinate Framework75. We omitted the ‘fiber tracts’ summary structure in the analysis to focus on grey matter structures. Counts were then generated on a per-region basis for each sample.
Batch effect correction
We observed differences in the total number of c-Fos+ cells in psilocybin samples across batch 1 and 2, saline samples across batch 1 and 2, and ketamine samples across batch 1 and 3. Batch effects are common and, in this study, may arise from differences in antibody quality, microscope condition, and/or subtle changes in the automated cell counting procedure. To correct for these differences, a scaling factor was calculated for the psilocybin, ketamine, and saline conditions individually. This factor was calculated by taking the mean total c-Fos+ cell counts of the batch 2 (psilocybin, saline) or 3 (ketamine) mice belonging to the same drug condition and dividing by mean total c-Fos+ cell counts of the batch 1 (psilocybin, saline, ketamine) mice belonging to the same drug condition. The factor was 2.78 for psilocybin, 4.94 for ketamine, and 3.11 for saline. These factors were applied to the per-region c-Fos+ cell count data in batch 1 to shift the c-Fos+ cell counts to be more comparable to the later batches. All analyses were performed after the batch effect correction. We emphasize that this batch correction step should not affect the machine learning analysis pipeline described below. This is because the first step of the pipeline is to divide per-region count by the total count in each brain, meaning that the absolute values of the cell count should have minimal influence on model fits but instead it is the relative values of the cell count (e.g., the proportion of c-Fos+ cell residing in one brain region over another brain region in a sample) that mattered for analysis and prediction.
Head-twitch response
Head movements were recorded using a magnetic ear tag system as described in detail previously33. Briefly, an ear tag consisted of a neodymium magnet (N45, 3 mm diameter, 0.5 mm thick, #D1005-10, SuperMagnetMan) that was adhered to an aluminium ear tag (La Pias #56780, Stoelting) with cyanoacrylate glue (Super Glue Ultra Gel Control, $1739050, Loctite). The neodymium magnet was coated with a nitrocellulose marker (#7056, ColorTone) and dried for >2 h, which helped to reduce ear irritation for the mice. This magnetic ear tag was applied to the mouse’s ear using an ear tag applicator (#56791, Stoelting). For measurement, the animal was put inside a plastic cube (4” × 4” × 4”). A spool of enameled cooper wire (30 AWG) was used to wind around the cube like a solenoid, with the ends of the wire connected to a current-to-voltage preamplifier (PP444, Pyle) where the voltage was captured with a computer via a data acquisition device (USB-6001, National Instruments). Each mouse was recorded using one cube. Up to four cubes could be used to record from four mice at once inside a soundproof chamber. Data acquisition and analysis were done using custom software written in MATLAB (Mathworks). The voltage signal was sent through a 70–110 Hz bandpass filter because the head twitch response had a characteristic ~90 Hz frequency. The filtered signal was then processed for peak detection to identify individual head-twitch events. A protocol including parts list for the setup and the MATLAB code is available at https://github.com/Kwan-Lab/HTR.
Machine learning pipeline—preprocessing
The analysis pipeline used the Python package sci-kit learn (Version 1.2.1)104. The first step of the pipeline was preprocessing, which entails three steps: normalization, transformation, and scaling. For normalization, we divided each region’s c-Fos+ cell count by the total c-Fos+ cell count across all summary structures used. This was done to mitigate influence of batch effects across samples. For transformation, each brain region’s normalized c-Fos+ cell counts across different drug conditions were transformed using Yeo-Johnson power transformation105. The Yeo-Johnson transformation is a generalized form of the Box-Cox transformation. The transformation leads to data values that more closely approximate a Gaussian distribution. The Yeo-Johnson transformation was implemented in scikit-learn: PowerTransformer(method=’yeo-johnson’, standardize=False). The Yeo-Johnson transformation is parameterized by one variable, lambda. The optimal lambda parameter was calculated for each brain region independently using maximum likelihood estimation to optimize for normality. For scaling, for each brain region, the RobustScaler module in scikit-learn was used to subtract the median value and scales values by the range of the 25th to 75th percentile (quartile scaling). We decided to do this, rather than subtracting mean value and standard-deviation scaling, because it is less sensitive to outliers. The c-Fos+ cell counts of each brain region after undergoing the normalization, transformation, and scaling steps are referred to as the c-Fos scores. To visualize the data, we performed dimensionality reduction on c-Fos scores across all samples using scikit-learn’s LinearDiscriminantAnalysis function and plotted the top two linear discriminants (Fig. 3b).
Machine learning pipeline—region selection
Based on Allen Institute definition of summary structures, the brain was divided into 315 regions (316 summary structure and then ‘fiber tracts’ removed). We were concerned that a model involving c-Fos scores from 315 regions may be overfitting due to our limited sample size of 64 brains. Many regions are likely not informative and only contribute noise to the machine learning models. Therefore, we implemented a method to filter out features (i.e., the brain regions) which were not informative for distinguishing the desired drug conditions. Region selection was carried out using the Boruta algorithm, as implemented in the BorutaPy package106. The Boruta algorithm is an ‘all relevant features’ selection method which seeks to identify all the features with information relevant to a task. This was done by creating scrambled versions of each feature, which are called shadow features, and appending them to the original data set. This expanded data set was then used to fit a random forest classifier, as implemented in scikit-learn. We used the BorutaPy package to automatically select the number of trees for the RandomForestClassifier() module based on the size of the feature set. Following this, a threshold was established based on the highest feature importance amongst shadow features. Features exceeding this threshold were considered ‘hits’ and recorded. This procedure was repeated 100 times. The distribution across these 100 iterations created a binomial distribution. The BorutaPy package rejected features based on the cumulative distribution function of a binomial distribution where p = 0.5, alpha = 0.05, and n = number of hits. Features (i.e., brain regions) that were not rejected by this criterion were the features included for the next stage of the pipeline.
Machine learning pipeline—classification
We used the c-Fos scores of the selected brain regions to fit a ridge regression model (L2 normalized logistic regression). The regularization parameter C is a hyperparameter used to modulate the penalty strength. Given the interconnected nature of the exact feature set and hyperparameter, as well as our desire to eventually merge results across many cross-validation splits of the data, we opted to fix this parameter to its default value of 1. The ‘multinomial’ setting was used to generalize from binary classification to multi-class classification.
Cross-validation to determine prediction accuracy
The data were evaluated using the aforementioned pipeline using fourfold splits, where 75% of the data (i.e., 6 brain samples) in each drug condition was used to train and fit the model, while the remaining 25% of the data (i.e., 2 brain samples) was used to test the model. Importantly, preprocessing parameters (e.g., lambda in Yeo-Johnson transformation) and feature selection (brain regions to be included) were chosen using only the training data to ensure no data leakage. Nevertheless, after those stages were fixed, the test data would undergo the same preprocessing and feature selection steps before being inputted into the ridge regression model to generate the prediction of the drug condition. We performed 100 iterations, each time using randomized split for each drug condition generated by sci-kit-learn’s StratifiedShuffleSplit() function. Combining the outcome across the 100 iterations, the predicted classifications were used to generate a mean confusion matrix (Fig. 3c). The probabilities assigned to each label for each test data point were combined to create a composite precision recall curve, generated using scikit-learn’s precision_recall_curve() function (Figs. 3d and 4b). The scikit-learn’s auc function was used to calculate the area under the curve for each composite precision recall curve (legend of Fig. 3d). We used numpy’s random seeds and state objects (numpy.random.RandomState()) to generate reproducible results. The cross-validation splitting function was seeded with an integer, per scikit-learn’s recommendations. The remaining random states were set using a random state object. A null distribution for the area under the precision recall curve was established by shuffling labels during each cross-validation split prior to model fitting and label prediction (Fig. 4b).
Shapley additive explanation
SHAP values were generated by the LinearExplainer object from the SHAP package, which accepted test data points and the fit model. We set the feature perturbation parameter of the LinearExplainer to ‘correlation_dependent’. SHAP values were generated in part by breaking dependencies across features and testing the influence of perturbations on individual features. This ran the risk of creating unrealistic feature combinations, because many brain regions, which would normally change in lockstep may be changed individually by the algorithm to infer feature importance, which would lead to inflated feature importance scores107. By using the “correlation-dependent” intervention, additional measures were taken to address correlations in the feature space and credit was distributed more appropriately. The SHAP values for each test data point were combined across the data splits from the 100 iterations to arrive at composite SHAP summary plots (Figs. 5c and 6a, c, e). We determined which brain regions were included in ≥75% of the cross-validation splits of the data (Fig. 4c, d). Regions meeting this criterion were visualized using the brainrender package108 (Figs. 5d and 6b, d, f).
Leave-one-drug-out analysis
The fitting of the pipeline (pipelineObj.fit) was performed on a reduced dataset of c-Fos scores, excluding all samples in the 6-fluoro-DET condition. That is, for each split, training data were c-Fos+ cell count from 75% of the samples from 7 conditions (psilocybin, ketamine, 5-MeO-DMT, MDMA, acute fluoxetine, chronic fluoxetine, and saline). The test data consists of c-Fos+ cell count from the remaining 25% of the samples from those seven conditions and 25% of the samples drawn from the left-out condition of 6-fluoro-DET. For linear discriminant analysis, the full dataset was transformed (pipelineObj.transform) and plotted using multiple calls to the seaborn scatterplot function (sns.scatterplot).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The behavioral and light sheet data generated in this study are provided in the Supplementary Information and Source Data file. Source data are provided with this paper.
Code availability
Code associated with the study is available at https://github.com/Kwan-Lab/aboharbdavoudian2025 and https://github.com/Kwan-Lab/HTR.
References
Kelmendi, B., Kaye, A. P., Pittenger, C. & Kwan, A. C. Psychedelics. Curr. Biol. 32, R63–R67 (2022).
Vollenweider, F. X. & Preller, K. H. Psychedelic drugs: neurobiology and potential for treatment of psychiatric disorders. Nat. Rev. Neurosci. 21, 611–624 (2020).
Davis, A. K. et al. Effects of psilocybin-assisted therapy on major depressive disorder: a randomized clinical trial. JAMA Psychiatry 78, 481–489 (2021).
Carhart-Harris, R. et al. Trial of psilocybin versus escitalopram for depression. N. Engl. J. Med. 384, 1402–1411 (2021).
Goodwin, G. M. et al. Single-dose psilocybin for a treatment-resistant episode of major depression. N. Engl. J. Med. 387, 1637–1648 (2022).
Raison, C. L. et al. Single-dose psilocybin treatment for major depressive disorder: a randomized clinical trial. JAMA 330, 843–853 (2023).
von Rotz, R. et al. Single-dose psilocybin-assisted therapy in major depressive disorder: a placebo-controlled, double-blind, randomised clinical trial. EClinicalMedicine 56, 101809 (2023).
Berman, R. M. et al. Antidepressant effects of ketamine in depressed patients. Biol. Psychiatry 47, 351–354 (2000).
Murrough, J. W. et al. Antidepressant efficacy of ketamine in treatment-resistant major depression: a two-site randomized controlled trial. Am. J. Psychiatry 170, 1134–1142 (2013).
Zarate, C. A. Jr. et al. A randomized trial of an N-methyl-D-aspartate antagonist in treatment-resistant major depression. Arch. Gen. Psychiatry 63, 856–864 (2006).
Feder, A. et al. Efficacy of intravenous ketamine for treatment of chronic posttraumatic stress disorder: a randomized clinical trial. JAMA Psychiatry 71, 681–688 (2014).
Popova, V. et al. Efficacy and safety of flexibly dosed esketamine nasal spray combined with a newly initiated oral antidepressant in treatment-resistant depression: a randomized double-blind active-controlled study. Am. J. Psychiatry 176, 428–438 (2019).
Daly, E. J. et al. Efficacy and safety of intranasal esketamine adjunctive to oral antidepressant therapy in treatment-resistant depression: a randomized clinical trial. JAMA Psychiatry 75, 139–148 (2018).
Mitchell, J. M. et al. MDMA-assisted therapy for severe PTSD: a randomized, double-blind, placebo-controlled phase 3 study. Nat. Med. 27, 1025–1033 (2021).
Mitchell, J. M. et al. MDMA-assisted therapy for moderate to severe PTSD: a randomized, placebo-controlled phase 3 trial. Nat. Med. 29, 2473–2480 (2023).
Kwan, A. C., Olson, D. E., Preller, K. H. & Roth, B. L. The neural basis of psychedelic action. Nat. Neurosci. 25, 1407–1419 (2022).
McClure-Begley, T. D. & Roth, B. L. The promises and perils of psychedelic pharmacology for psychiatry. Nat. Rev. Drug Discov. 21, 463–473 (2022).
Olson, D. E. Psychoplastogens: a promising class of plasticity-promoting neurotherapeutics. J. Exp. Neurosci. 12, 1179069518800508 (2018).
Kaplan, A. L. et al. Bespoke library docking for 5-HT(2A) receptor agonists with antidepressant activity. Nature 610, 582–591 (2022).
Wallach, J. et al. Identification of 5-HT(2A) receptor signaling pathways associated with psychedelic potential. Nat. Commun. 14, 8221 (2023).
Dong, C. et al. Psychedelic-inspired drug discovery using an engineered biosensor. Cell 184, 2779–2792 e2718 (2021).
Cao, D. et al. Structure-based discovery of nonhallucinogenic psychedelic analogs. Science 375, 403–411 (2022).
Wiltschko, A. B. et al. Revealing the structure of pharmacobehavioral space through motion sequencing. Nat. Neurosci. 23, 1433–1443 (2020).
Alexandrov, V., Brunner, D., Hanania, T. & Leahy, E. High-throughput analysis of behavior for drug discovery. Eur. J. Pharm. 750, 82–89 (2015).
Nestler, E. J. & Hyman, S. E. Animal models of neuropsychiatric disorders. Nat. Neurosci. 13, 1161–1169 (2010).
Liao, C., Dua, A. N., Wojtasiewicz, C., Liston, C. & Kwan, A. C. Structural neural plasticity evoked by rapid-acting antidepressant interventions. Nat. Rev. Neurosci., 26, 101–114 (2025).
Li, N. et al. mTOR-dependent synapse formation underlies the rapid antidepressant effects of NMDA antagonists. Science 329, 959–964 (2010).
Phoumthipphavong, V., Barthas, F., Hassett, S. & Kwan, A. C. Longitudinal effects of ketamine on dendritic architecture in vivo in the mouse medial frontal cortex. eNeuro 3, ENEURO.0133–0115.2016 (2016).
Shao, L. X. et al. Psilocybin induces rapid and persistent growth of dendritic spines in frontal cortex in vivo. Neuron 109, 2535–2544.e2534 (2021).
de la Fuente Revenga, M. et al. Prolonged epigenomic and synaptic plasticity alterations following single exposure to a psychedelic in mice. Cell Rep. 37, 109836 (2021).
Cameron, L. P. et al. A non-hallucinogenic psychedelic analogue with therapeutic potential. Nature 589, 474–479 (2021).
Lu, J. et al. An analog of psychedelics restores functional neural circuits disrupted by unpredictable stress. Mol. Psychiatry 26, 6237–6252 (2021).
Jefferson, S. J. et al. 5-MeO-DMT modifies innate behaviors and promotes structural neural plasticity in mice. Neuropsychopharmacology 48, 1257–1266 (2023).
Sheng, M. & Greenberg, M. E. The regulation and function of c-fos and other immediate early genes in the nervous system. Neuron 4, 477–485 (1990).
Yap, E. L. & Greenberg, M. E. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron 100, 330–348 (2018).
Ma, H. et al. Excitation–transcription coupling, neuronal gene expression and synaptic plasticity. Nat. Rev. Neurosci. 24, 672–692 (2023).
Gonzalez-Maeso, J. et al. Transcriptome fingerprints distinguish hallucinogenic and nonhallucinogenic 5-hydroxytryptamine 2A receptor agonist effects in mouse somatosensory cortex. J. Neurosci. 23, 8836–8843 (2003).
Nichols, C. D. Sanders-Bush E. A single dose of lysergic acid diethylamide influences gene expression patterns within the mammalian brain. Neuropsychopharmacology 26, 634–642 (2002).
Leslie, R. A., Moorman, J. M., Coulson, A. & Grahame-Smith, D. G. Serotonin2/1C receptor activation causes a localized expression of the immediate-early gene c-fos in rat brain: evidence for involvement of dorsal raphe nucleus projection fibres. Neuroscience 53, 457–463 (1993).
Grieco, S. F. et al. Psychedelics and neural plasticity: therapeutic implications. J. Neurosci. 42, 8439–8449 (2022).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Kim, Y. et al. Mapping social behavior-induced brain activation at cellular resolution in the mouse. Cell Rep. 10, 292–305 (2015).
Davoudian, P. A., Shao, L. X. & Kwan, A. C. Shared and distinct brain regions targeted for immediate early gene expression by ketamine and psilocybin. ACS Chem. Neurosci. 14, 468–480 (2023).
Rijsketic, D. R. et al. UNRAVELing the synergistic effects of psilocybin and environment on brain-wide immediate early gene expression in mice. Neuropsychopharmacology 48, 1798–1807 (2023).
Datta, M. S. et al. Whole-brain mapping reveals the divergent impact of ketamine on the dopamine system. Cell Rep. 42, 113491 (2023).
Bijoch, L. et al. Whole-brain tracking of cocaine and sugar rewards processing. Transl. Psychiatry 13, 20 (2023).
Kimbrough, A. et al. Characterization of the brain functional architecture of psychostimulant withdrawal using single-cell whole-brain imaging. eNeuro 8, ENEURO.0208–0219.2021 (2021).
Carrette, L. L. G. et al. Hyperconnectivity of two separate long-range cholinergic systems contributes to the reorganization of the brain functional connectivity during nicotine withdrawal in male mice. eNeuro 10, ENEURO.0019–0023.2023 (2023).
Azevedo, H., Ferreira, M., Mascarello, A., Osten, P. & Guimaraes, C. R. W. Brain-wide mapping of c-fos expression in the single prolonged stress model and the effects of pretreatment with ACH-000029 or prazosin. Neurobiol. Stress 13, 100226 (2020).
Cruces-Solis, H., Nissen, W., Ferger, B. & Arban, R. Whole-brain signatures of functional connectivity after bidirectional modulation of the dopaminergic system in mice. Neuropharmacology 178, 108246 (2020).
Hansen, H. H. et al. Whole-brain activation signatures of weight-lowering drugs. Mol. Metab. 47, 101171 (2021).
Keyes, P. C. et al. Orchestrating opiate-associated memories in thalamic circuits. Neuron 107, 1113–1123 e1114 (2020).
Roland, A. V. et al. Alcohol dependence modifies brain networks activated during withdrawal and reaccess: a c-fos-based analysis in mice. Biol. Psychiatry 94, 393–404 (2023).
Skovbjerg, G. et al. Whole-brain mapping of amylin-induced neuronal activity in receptor activity-modifying protein 1/3 knockout mice. Eur. J. Neurosci. 54, 4154–4166 (2021).
Skovbjerg, G. et al. Uncovering CNS access of lipidated exendin-4 analogues by quantitative whole-brain 3D light sheet imaging. Neuropharmacology 238, 109637 (2023).
Stefaniuk, M. et al. Global brain c-Fos profiling reveals major functional brain networks rearrangements after alcohol reexposure. Neurobiol. Dis. 178, 106006 (2023).
Tan, B. et al. Drugs of abuse hijack a mesolimbic pathway that processes homeostatic need. Science 384, eadk6742 (2024).
Zanos, P. et al. Ketamine and ketamine metabolite pharmacology: insights into therapeutic mechanisms. Pharm. Rev. 70, 621–660 (2018).
Savalia, N. K., Shao, L. X. & Kwan, A. C. A dendrite-focused framework for understanding the actions of ketamine and psychedelics. Trends Neurosci. 44, 260–275 (2021).
Aleksandrova, L. R. & Phillips, A. G. Neuroplasticity as a convergent mechanism of ketamine and classical psychedelics. Trends Pharm. Sci. 42, 929–942 (2021).
Ali, F. et al. Ketamine disinhibits dendrites and enhances calcium signals in prefrontal dendritic spines. Nat. Commun. 11, 72 (2020).
Reckweg, J. T. et al. The clinical pharmacology and potential therapeutic applications of 5-methoxy-N,N-dimethyltryptamine (5-MeO-DMT). J. Neurochem. 162, 128–146 (2022).
Davis, A. K., So, S., Lancelotta, R., Barsuglia, J. P. & Griffiths, R. R. 5-methoxy-N,N-dimethyltryptamine (5-MeO-DMT) used in a naturalistic group setting is associated with unintended improvements in depression and anxiety. Am. J. Drug Alcohol Abus. 45, 161–169 (2019).
Blair, J. B. et al. Effect of ring fluorination on the pharmacology of hallucinogenic tryptamines. J. Med. Chem. 43, 4701–4710 (2000).
Rabin, R. A., Regina, M., Doat, M. & Winter, J. C. 5-HT2A receptor-stimulated phosphoinositide hydrolysis in the stimulus effects of hallucinogens. Pharm. Biochem. Behav. 72, 29–37 (2002).
Kalir, A. & Szara, S. Synthesis and pharmacological activity of fluorinated tryptamine derivatives. J. Med. Chem. 6, 716–719 (1963).
Faillace, L. A., Vourlekis, A. & Szara, S. Clinical evaluation of some hallucinogenic tryptamine derivatives. J. Nerv. Ment. Dis. 145, 306–313 (1967).
Helsley, S., Fiorella, D., Rabin, R. A. & Winter, J. C. A comparison of N,N-dimethyltryptamine, harmaline, and selected congeners in rats trained with LSD as a discriminative stimulus. Prog. Neuropsychopharmacol. Biol. Psychiatry 22, 649–663 (1998).
Studerus, E., Gamma, A., Kometer, M. & Vollenweider, F. X. Prediction of psilocybin response in healthy volunteers. PLoS One 7, e30800 (2012).
Nichols, D. E. Differences between the mechanism of action of mdma, mbdb, and the classic hallucinogens - identification of a new therapeutic class - entactogens. J. Psychoact. Drugs 18, 305–313 (1986).
Young, M. B., Andero, R., Ressler, K. J. & Howell, L. L. 3,4-Methylenedioxymethamphetamine facilitates fear extinction learning. Transl. Psychiatry 5, e634 (2015).
Dulawa, S. C., Holick, K. A., Gundersen, B. & Hen, R. Effects of chronic fluoxetine in animal models of anxiety and depression. Neuropsychopharmacology 29, 1321–1330 (2004).
Simard, S. et al. Fibroblast growth factor 2 is necessary for the antidepressant effects of fluoxetine. PLoS One 13, e0204980 (2018).
Morgan, J. I., Cohen, D. R., Hempstead, J. L. & Curran, T. Mapping patterns of c-fos expression in the central nervous system after seizure. Science 237, 192–197 (1987).
Wang, Q. et al. The allen mouse brain common coordinate framework: a 3D reference atlas. Cell 181, 936–953 e920 (2020).
Wang, Q. & Burkhalter, A. Area map of mouse visual cortex. J. Comp. Neurol. 502, 339–357 (2007).
Roth, M. M., Helmchen, F. & Kampa, B. M. Distinct functional properties of primary and posteromedial visual area of mouse neocortex. J. Neurosci. 32, 9716–9726 (2012).
Matsumoto, M. & Hikosaka, O. Lateral habenula as a source of negative reward signals in dopamine neurons. Nature 447, 1111–1115 (2007).
Matsumoto, M. & Hikosaka, O. Representation of negative motivational value in the primate lateral habenula. Nat. Neurosci. 12, 77–84 (2009).
Li, B. et al. Synaptic potentiation onto habenula neurons in the learned helplessness model of depression. Nature 470, 535–539 (2011).
Yang, Y. et al. Ketamine blocks bursting in the lateral habenula to rapidly relieve depression. Nature 554, 317–322 (2018).
Fallon, I. P. et al. The role of the parafascicular thalamic nucleus in action initiation and steering. Curr. Biol. 33, 2941–2951 e2944 (2023).
Zingg, B. et al. Neural networks of the mouse neocortex. Cell 156, 1096–1111 (2014).
Harris, J. A. et al. Hierarchical organization of cortical and thalamic connectivity. Nature 575, 195–202 (2019).
Ju, A., Fernandez-Arroyo, B., Wu, Y., Jacky, D. & Beyeler, A. Expression of serotonin 1A and 2A receptors in molecular- and projection-defined neurons of the mouse insular cortex. Mol. Brain 13, 99 (2020).
Gozzi, A. & Schwarz, A. J. Large-scale functional connectivity networks in the rodent brain. NeuroImage 127, 496–509 (2016).
Mandino, F. et al. A triple-network organization for the mouse brain. Mol. Psychiatry 27, 865–872 (2021).
Lynch, C. J. et al. Frontostriatal salience network expansion in individuals in depression. Nature 633, 624–633 (2024).
Salay, L. D., Ishiko, N. & Huberman, A. D. A midline thalamic circuit determines reactions to visual threat. Nature 557, 183–189 (2018).
Duerler, P. et al. Psilocybin induces aberrant prediction error processing of tactile mismatch responses—a simultaneous EEG–FMRI study. Cereb. Cortex 32, 186–196 (2022).
Vollenweider, F. X. & Kometer, M. The neurobiology of psychedelic drugs: implications for the treatment of mood disorders. Nat. Rev. Neurosci. 11, 642–651 (2010).
Ley, L. et al. Comparative acute effects of mescaline, lysergic acid diethylamide, and psilocybin in a randomized, double-blind, placebo-controlled cross-over study in healthy participants. Neuropsychopharmacology 48, 1659–1667 (2023).
Botvinik-Nezer, R. et al. Variability in the analysis of a single neuroimaging dataset by many teams. Nature 582, 84–88 (2020).
Goodwin, N. L., Nilsson, S. R. O., Choong, J. J. & Golden, S. A. Toward the explainability, transparency, and universality of machine learning for behavioral classification in neuroscience. Curr. Opin. Neurobiol. 73, 102544 (2022).
Goodwin, N. L. et al. Simple Behavioral Analysis (SimBA) as a platform for explainable machine learning in behavioral neuroscience. Nat. Neurosci. 27, 1411–1424 (2024).
Lee, E. K. et al. Non-linear dimensionality reduction on extracellular waveforms reveals cell type diversity in premotor cortex. Elife 10, e67490 (2021).
Lai, H. M. et al. Antibody stabilization for thermally accelerated deep immunostaining. Nat. Methods 19, 1137–1146 (2022).
Vladimirov, N. et al. Benchtop mesoSPIM: a next-generation open-source light-sheet microscope for cleared samples. Nat. Commun. 15, 2679 (2024).
Chen, Y. et al. Low-cost and scalable projected light-sheet microscopy for the high-resolution imaging of cleared tissue and living samples. Nat. Biomed. Eng. 8, 1109–1123 (2024).
Sherwood, A. M., Claveau, R., Lancelotta, R., Kaylo, K. W. & Lenoch, K. Synthesis and characterization of 5-MeO-DMT succinate for clinical use. ACS Omega 5, 32067–32075 (2020).
Kim, S. Y. et al. Stochastic electrotransport selectively enhances the transport of highly electromobile molecules. Proc. Natl Acad. Sci. USA 112, E6274–E6283 (2015).
Murray, E. et al. Simple, scalable proteomic imaging for high-dimensional profiling of intact systems. Cell 163, 1500–1514 (2015).
Marstal K., Berendsen F., Staring M., Klein S. SimpleElastix: a user-friendly, multi-lingual library for medical image registration. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition Workshops (CVPRW)) (2016).
Pedregosa, F. et al. Scikit-learn: machine learning in python. JMLR 12, 2825–2830 (2011).
Yeo, I. K. & Johnson RA. A new family of power transformations to improve normality or symmetry. Biometrika 87, 954–959 (2000).
Kursa, M. B. & Rudnicki, W. R. Feature selection with the Boruta package. J. Stat. Softw. 36, 1–13 (2010).
Mase M., Owen A. B. & Seiler B. Explaining black box decisions by Shapley cohort refinement. arxiv, https://arxiv.org/abs/1911.00467 (2020).
Claudi, F. et al. Visualizing anatomically registered data with brainrender. Elife 10, e65751 (2021).
Acknowledgements
Psilocybin, 5-MeO-DMT, and 6-fluoro-DET were provided by Usona Institute’s Investigational Drug & Material Supply Program; the Usona Institute IDMSP is supported by Alexander Sherwood, Robert Kargbo, and Kristi Kaylo in Madison, WI. This work was supported by NIH grants R01MH121848 (A.C.K.), R01MH128217 (A.C.K.), R01MH137047 (A.C.K.); One Mind—COMPASS Rising Star Award (A.C.K.); Cornell Engineering M.D.-M.Eng. program (F.A.); NIH training grants T32GM152349 (F.A.), T32GM007205 (P.A.D.), T32NS041228 (C.L.); NIH fellowship F30DA059437 (P.A.D.); Source Research Foundation student grant (P.A.D.); VA National Center for PTSD (A.P.K.); Department of Defense HT9425-23-1-0458 (A.P.K.); K08MH122733 (A.P.K.); this work was funded in part by the State of Connecticut, Department of Mental Health and Addiction Services, but this publication does not express the views of the Department of Mental Health and Addiction Services or the State of Connecticut. The views and opinions expressed are those of the authors.
Author information
Authors and Affiliations
Contributions
F.A., P.A.D., and A.C.K. planned the study. P.A.D., L.X.S., C.L., and J.R. performed experiments. G.N.R. and C.W. assisted with tissue processing and imaging. P.A.D. and M.D. measured head-twitch responses. F.A. and P.A.D. analyzed the data, with input from A.C.K. on the pipeline. J.I. assisted with data analysis. A.M.S. and A.P.K. contributed reagents. F.A. and A.C.K. drafted the manuscript. All authors reviewed the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
A.C.K. has been a scientific advisor or consultant for Boehringer Ingelheim, Empyrean Neuroscience, Freedom Biosciences, and Xylo Bio. A.C.K. has received research support from Intra-Cellular Therapies. A.P.K has received research support from Transcend Therapeutics and Freedom Biosciences. A.P.K. has a provisional patent application related to psychedelics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jianjun Cheng, John McCorvy and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aboharb, F., Davoudian, P.A., Shao, LX. et al. Classification of psychedelics and psychoactive drugs based on brain-wide imaging of cellular c-Fos expression. Nat Commun 16, 1590 (2025). https://doi.org/10.1038/s41467-025-56850-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56850-6
This article is cited by
-
The psychoplastogen tabernanthalog induces neuroplasticity without proximate immediate early gene activation
Nature Neuroscience (2025)