Abstract
To replicate and cause disease, Mycobacterium tuberculosis secretes siderophores called mycobactins to scavenge iron from the human host. Two closely related transporters, MmpL4 and MmpL5, are required for mycobactin secretion and drug efflux. In clinical strains, overproduction of MmpL5 confers resistance towards bedaquiline and clofazimine, key drugs to combat multidrug resistant tuberculosis. Here, we present cryogenic-electron microscopy structures of MmpL4 and identify a mycobactin binding site, which is accessible from the cytosol and also required for bedaquiline efflux. An unusual coiled-coil domain predicted to extend 130 Å into the periplasm is essential for mycobactin and bedaquiline efflux by MmpL4 and MmpL5. The mycobacterial acyl carrier protein MbtL forms a complex with MmpL4, indicating that mycobactin synthesis and export are coupled. Thus, MmpL4 and MmpL5 constitute the core components of a unique multi-subunit machinery required for iron acquisition and drug efflux by M. tuberculosis.
Similar content being viewed by others
Introduction
Iron is essential for growth of M. tuberculosis1, but iron-sequestering proteins in the lung granulomas of tuberculosis (TB) patients establish an iron-deprived environment2. To prevent iron starvation, M. tuberculosis secretes siderophores, small molecules with high iron-binding affinities called mycobactins3. Two small membrane proteins, MmpS4 and MmpS5, interact with the large efflux pumps MmpL4 and MmpL5 to form an inner membrane transporter complex required for siderophore secretion by M. tuberculosis4.
MmpL4 and MmpL5 belong to the superfamily of RND transporters, secondary active transporters that exploit the proton-motive force to pump substrates such as lipids or antibiotics out of the cell5. The genome of M. tuberculosis encodes 13 MmpL proteins, which are further subdivided into two clusters: a large cluster I including MmpL4 and MmpL5 and the smaller cluster II comprising MmpL11 and the essential trehalose monomycolate transporter MmpL3, whose structure has been determined6,7,8,9,10,11,12. Interrupting siderophore secretion by deleting genes encoding MmpL4 and MmpL5 leads to self-poisoning of M. tuberculosis1 and a more than 20,000-fold reduced bacterial burden in mice4, demonstrating that the siderophore secretion system is essential for virulence of M. tuberculosis. Together with the periplasmic protein Rv0455c13 and the MmpS4 and MmpS5 proteins, the closely related and functionally redundant MmpL4 and MmpL5 transporters (64% sequence identity) constitute the central components of a siderophore secretion system unique to mycobacteria. Importantly, the siderophore secretion system of M. tuberculosis is also required for efflux of bedaquiline, the most important drug for treatment of multidrug-resistant tuberculosis14,15,16. Unfortunately, clinical M. tuberculosis strains rapidly become resistant to bedaquiline predominantly by overexpression of the mmpS5-mmpL5 operon17,18,19,20. A recent study showed that overexpression of mmpL4 also confers bedaquiline resistance in M. tuberculosis21. However, no structures of MmpL4 or MmpL5 are known, impeding a molecular understanding of these clinically relevant transporters, which are capable of transporting both siderophores and drugs, two functions critical for M. tuberculosis.
Results
Structure determination of M. tuberculosis MmpL4
To shed light on the molecular mechanism of mycobactin secretion and bedaquiline efflux, we purified MmpL4 upon expression in Escherichia coli from the native mmpS4-mmpL4 operon of M. tuberculosis (see methods). MmpL4 elutes as two broad peaks from the size exclusion chromatography column. The first peak likely corresponds to multimers of variable stoichiometry, while the second peak runs at an elution volume corresponding to a monomer (Fig. 1a). SDS-PAGE analysis revealed that monomeric MmpL4 suffers from protein degradation (Fig. 1b). A MmpL4-specific sybody (Sb09)22,23 was generated, which was used to reconstruct a 6.3 Å cryo-EM map of MmpL4 (Supplementary Figs. 1 and 2, Supplementary Table 1). Docking the AlphaFold224 prediction of MmpL4 into this low-resolution map enabled the assignment of the transmembrane domain and a large proportion of the periplasmic domains. However, no density was present for an unusual 130 Å-long coiled-coil domain predicted by AlphaFold2 (Fig. 1c, Supplementary Fig. 1), likely due to a flexible hinge connecting the coiled-coil domain with the rest of the transporter. Hence, we replaced the coiled-coil domain (residues S491-Y685) with a short glycine-serine linker (MmpL4-T construct). Purified MmpL4-T protein appeared as a single peak on the size exclusion chromatogram with improved sample quality (Fig. 1a, b). This enabled us to determine a 3.0 Å cryo-EM structure of monomeric MmpL4-T in the detergent n-dodecyl-β-D-maltoside (β-DDM) (Fig. 1c, Supplementary Fig. 3, Supplementary Table 1). Density was observed for the transmembrane and periplasmic domains of MmpL4, excluding the N- and C- terminus (residues 1–15, and 940–967) and the glycine-serine linker that replaced the coiled-coil domain. An atomic model was built into the density map, which differs from the AlphaFold2 prediction with a root mean square deviation (RMSD) of 1.45 Å (Supplementary Fig. 4). We also identified a phosphatidylethanolamine lipid bound in a pocket at the inner leaflet of the transmembrane domain of MmpL4 (Supplementary Fig. 5). Comparison of the cryo-EM data of full-length and truncated MmpL4 demonstrates that truncation of the coiled-coil domain does not affect the conformation of MmpL4 (Supplementary Fig. 1). Even though untagged MmpS4 is co-purified with both full-length and truncated MmpL4 (Fig. 1b, Supplementary Fig. 6a), we did not observe it in our cryo-EM maps, suggesting that it either is present at sub-stoichiometric levels or dissociates during grid preparation. However, unaccounted density at the C-terminus of MmpL4 was identified as the E. coli acyl carrier protein (ACP) by mass spectrometry (Fig. 1b, c, Supplementary Fig. 6b), which was unintendedly co-purified from the expression host E. coli.
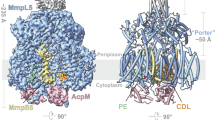
a and b Size exclusion chromatography profile (a) and Coomassie stained SDS-PAGE (b) of detergent-purified full-length and truncated MmpL4. Bands labeled as MmpL4, degradation (degr.), MmpS4 and acyl carrier protein (ACP) were confirmed by mass spectrometry (data shown in Supplementary Fig. 6). Source data are provided as a Source Data file. c Cryo-EM map of truncated MmpL4 (contoured at 7.5 σ). The transmembrane domain (TMD) as well as the two periplasmic porter subdomains PN and PC are marked. ACP of the expression host E. coli was unintendedly co-purified. The AF2 prediction of the coiled-coil domain which was truncated for cryo-EM analyses is shown as cartoon. d Simplified topology representation of the multidrug RND transporter AcrB and the MmpL cluster I and II subfamilies and representative structures shown as cartoon. e Comparison of the C-terminal porter subdomain (PC) of MmpL4 (AF2) and the corresponding PC2 of AcrB shown as cartoon and rotated by 90 degrees compared to (d). f Slice through the transmembrane domain of truncated MmpL4 viewed from the periplasm. The conserved DY pairs are shown as sticks. The two-fold pseudo-symmetry between the N- and C-terminal six helix bundles of the transmembrane domain is indicated by the black oval.
The transmembrane domain of MmpL4 consists of 12 transmembrane helices that form two pseudo-symmetric bundles, the conserved fold of the transmembrane domain of RND transporters (Fig. 1d, Supplementary Fig. 7)5,25. The periplasmic domain is composed of two subdomains adopting an α-β sandwich fold. In analogy to the thoroughly studied RND transporter AcrB responsible for multidrug efflux in Escherichia coli25,26,27,28, we call these subdomains PN and PC, which stands for periplasmic porter subdomain of the N- or C-terminal half of MmpL4, respectively. As for all RND transporters known to date, the PN is placed between TM1-2 and the PC between TM7-85.
The coiled-coil domain is essential for MmpL4 and MmpL5 function
The coiled-coil domain predicted by AlphaFold2 sits in the PC at the same location where the funnel domain is inserted in AcrB25 (Fig. 1d, Supplementary Fig. 7). The coiled-coil domain distinguishes MmpL4, MmpL5 and the cluster I MmpL proteins from the trehalose monomycolate exporter MmpL3 (cluster II), which has a periplasmic domain composed of only two porter subdomains (Fig. 1d, Supplementary Fig. 7). In order to investigate the functional role of the coiled-coil domain we used a functional assay based on the self-poisoning phenotype previously described for M. tuberculosis lacking both MmpL4 and MmpL5 (MtbΔmmpL4/mmpL5)21,29. In this strain, externally added ferrated mycobactin is taken up but cannot be exported resulting in toxic intracellular accumulation of mycobactin and growth attenuation. If a mutation of MmpL4 or MmpL5 renders the protein non-functional, siderophore export is disrupted and impairs growth of the MtbΔmmpL4/mmpL5 strain in the presence of mycobactin. In contrast, the MtbΔmmpL4/mmpL5 strain complemented with full-length mmpL4 or mmpL5 grows like wild-type M. tuberculosis at elevated levels of mycobactin (Fig. 2a). Likewise, the MtbΔmmpL4/mmpL5 strain is more sensitive to bedaquiline, a phenotype that is reversed by complementation with mmpL4 or mmpL5 (Supplementary Fig. 8f). Intriguingly, we found that truncated versions of MmpL4 or MmpL5 lacking the coiled-coil domain did not rescue the mycobactin sensitivity of the MtbΔmmpL4/mmpL5 strain, demonstrating that the coiled-coil domain is essential for siderophore export (Fig. 2a, Supplementary Fig. 8). Further, MmpL5 lacking the coiled-coil domain lost its ability to confer bedaquiline resistance (Fig. 2b, Supplementary Fig. 8). Western Blot analysis confirmed that truncated MmpL4 and MmpL5 are produced in M. tuberculosis cells (Supplementary Fig. 9). Since we determined the structure of MmpL4-T, protein misfolding does not explain the lack of transport of truncated MmpL4 and likely also truncated MmpL5.
a Mycobactin toxicity assay. Data were normalized to the viability in the absence of mycobactin (100%). Mtb, wild-type M. tuberculosis; ΔL4L5, M. tuberculosis strain lacking mmpL4 and mmpL5; ::S4L4 or ::S5L5, complementation with mmpS4-mmpL4 or mmpS5-mmpL5, ::L4-T or ::L5-T, complementation with truncated MmpL4 or MmpL5 lacking the coiled-coil domain. b Viability of M. tuberculosis strains determined at increasing bedaquiline concentrations. Data were normalized to the viability in the absence of bedaquiline (100%). ::S5L5, complementation with mmpS5-mmpL5, ::L5-T, complementation with truncated MmpL5 lacking the coiled-coil domain. c Exemplary data of a non-essential residue (Y273A), an intermediate phenotype (T802V) and an essential residue (D272N) of MmpL4 proton-translocation channel variants in the mycobactin toxicity assay. d Left and middle panel: Top view of the transmembrane domain, sliced through as indicated in the right panel, of MmpL4 (this study) and MmpL5 (AF2 prediction) colored according to the results of the mycobactin toxicity assay. The corresponding bar charts for all variants are shown in Supplementary Fig. 8 and source data are provided as a Source Data file. The structures are shown as cartoon with polar and charged residues depicted as sticks. Hydrogen bonds are indicated by dashed lines. Data points shown in the graphs represent technical replicates and representative data of biological duplicates are shown.
Structure of a mycobactin-MmpL4 complex
Initial attempts to determine cryo-EM structures of detergent-purified MmpL4-T in complex with mycobactins were not successful (see methods). Therefore, we reconstituted truncated MmpL4-T into nanodiscs and determined a 3.4 Å cryo-EM structures of MmpL4-T in the absence of desferrated mycobactin (‘apo’) and of MmpL4-T bound to desferrated mycobactin (Fig. 3a, Supplementary Figs. 10 and 11, Supplementary Table 1). Comparison of the two maps showed an unambiguous density for desferrated mycobactin at the interface of the N- and C-terminal six-helix bundles at the cytosolic side of the transmembrane domain when it was added to the sample, but not when it was absent (Supplementary Fig. 12). We observed density for the lipid phosphatidylethanolamine at the same location in the cryo-EM structure determined for detergent-purified MmpL4-T and a less well resolved density resembling a lipid in the ‘apo’ MmpL4-T nanodisc sample (Supplementary Figs. 5 and 12a). Therefore, all MmpL4-T structures presented here are in a ligand bound state, and the structures are very similar with RMSDs of 0.45–0.46 Å across all atoms. The cytosolic access pocket in which mycobactin and phosphatidylethanolamine are bound is formed by Iα1, TM5, TM6, TM8 and TM10 (Fig. 4). The side chains of the three residues, F19, R300 and R791 undergo conformational changes between our PE and MBT bound MmpL4-T structures to accommodate the bulkier MBT substrate (Supplementary Fig. 12c).
a Surface representation of MmpL4 bound to desferrated mycobactin (MBT). b Interaction between MmpL4 and desferrated mycobactin shown from the front (left) and top (right). Contacting residues are shown as sticks and residues investigated in the mycobactin toxicity assay are colored according to the phenotype in the assay. c Mycobactin toxicity assay. Viability of MtbΔmmpL4/mmpL5 complemented with MmpL4 variants of the mycobactin binding site at increasing mycobactin concentration. Data were normalized to the viability in the absence of mycobactin (100%). Data points shown in the graphs represent technical replicates and representative data of biological duplicates are shown. Source data are provided as a Source Data file.
Mycobactin is depicted as gray sphere. The periplasmic cavities calculated with the 3 V tool30 are shown in yellow. Dotted box: DY pairs of the transmembrane domain viewed from the periplasm. Hydrogen bonds are depicted as dashed black lines. Solid box: Close up of the cytosolic access pocket. Iα3 and the C-terminus of MmpL3 are colored in salmon to highlight how they block the cytosolic access pocket in MmpL3.
Functional role of the mycobactin binding pocket in siderophore secretion and bedaquiline efflux
The mycobactin head group is coordinated by Y299, R300, H304, R791 and K854. H304 is within hydrogen bonding distance with mycobactin (Fig. 3b, Supplementary Fig. 12b). Most of these residues are conserved in many MmpL cluster I subfamily members including MmpL5 (Supplementary Fig. 13) and in the three closest MmpL4/5 orthologues of Mycobacterium smegmatis, two of which, MSMEG_0225 and MSMEG_3496 export mycobactins in M. tuberculosis (Supplementary Fig. 14). To investigate the role of the mycobactin coordinating residues of MmpL4, we used the mycobactin toxicity assay. While the variants MmpL4Y299A and MmpL4K854A still export mycobactin, MmpL4R791A partially rescued the MtbΔmmpL4/mmpL5 strain and the MmpL4H304A variant did not rescue it (Fig. 3c, Supplementary Fig. 8). The essential nature of H304 demonstrates the functional importance of the mycobactin binding site for mycobactin export by MmpL4. Next, we examined whether the mycobactin binding site is also required for bedaquiline efflux. Consistent with a previous study21, wild-type MmpL4 increases the resistance of the MtbΔmmpL4/mmpL5 strain to bedaquiline (Fig. 5c). The variants MmpL4Y299A, MmpL4H304A and MmpL4R791A failed to confer bedaquiline resistance (Fig. 5d, Supplementary Fig. 8) in a similar manner as the D276N mutation, which inactivates proton translocation of MmpL4. These results indicate that MmpL4 binds bedaquiline at the same site as mycobactin. However, it should be noted that the effect of MmpL4 on bedaquiline resistance by M. tuberculosis is small compared to that of MmpL521.
a Chemical structure of bedaquiline. b Close-up view of mycobactin binding site. Residues are colored according to the phenotype of mutants in the bedaquiline susceptibility assay. c and d Viability of M. tuberculosis ΔmmpL4mmpL5 (ΔL4L5) complemented with mmpS4-mmpL4 (::S4L4) or the MmpL4 variants (::Y299A, ::H304A, ::R791A and ::K854A) at increasing bedaquiline concentrations. Data were normalized to the viability in the absence of bedaquiline (100%). Data points shown in the graphs represent technical replicates and representative data of biological duplicates are shown. Source data are provided as a Source Data file.
Structural comparison of MmpL4 and MmpL3
The mycobactin binding pocket of MmpL4 is absent in MmpL3 because of the presence of two short helices, Iα1 and Iα3. In addition, part of the flexible C-terminus of MmpL3 sits in the pocket (Fig. 4, Supplementary Fig. 7). Instead, substrate binding by MmpL3 has been observed at a binding site created by TM7 and TM8 at the outer leaflet of the inner membrane10. We did not observe extra density corresponding to a lipid or substrate molecule in the TM7-8 binding pocket in our detergent or nanodisc-reconstituted MmpL4-T cryo-EM maps.
The porter subdomains of M. tuberculosis MmpL3 are in an open conformation, creating a cavity of 3,128 Å3 which accommodates a detergent molecule7,30 (Fig. 4). Similarly, detergent and lipid molecules including the substrate trehalose monomycolate bind in the cavity created by the porter subdomains of M. smegmatis MmpL3, which is accessible to the periplasm via three openings8,9,10. In contrast, in our MmpL4-T structures, the porter subdomains display a closed conformation surrounding an inaccessible pocket of only 281 Å3 (Fig. 4). This closed conformation is a result of a rigid body rotation of the PC, which coincides with conformational rearrangements within the transmembrane domain, where proton translocation from the periplasm to the cytosol energizes substrate export26,27. We observed notable differences between MmpL3 and MmpL4 in the hydrogen bonding network of the titratable residues at the center of the transmembrane domain (Fig. 4, Supplementary Fig. 15)31. MmpL3 and MmpL4 contain two aspartate-tyrosine (DY) pairs sitting at the center of the channel formed by TM4 and TM10, at the same location where the catalytic key residues D407, D408 and K940 are found in AcrB25,27. In the open conformation of M. tuberculosis MmpL3, both DY pairs form hydrogen bonds. In the closed conformation of MmpL4, D272 forms a hydrogen bond with Y845, but Y273 and D844 have moved apart.
Functional analysis of presumed proton coupling residues in MmpL4 and MmpL5
To investigate the functional role of the conserved DY pairs in MmpL4 and MmpL5 (Fig. 2d), we used the mycobactin toxicity assay. The MmpL4D272N and MmpL5D276N variants were both completely inactive (Fig. 2c, d, Supplementary Fig. 8). However, MmpL5D849N was active, while the corresponding MmpL4D844N variant was inactive. In addition, the MmpL4D272N and MmpL5D276N variants lost their capacity to efflux bedaquiline (Supplementary Fig. 8). The essential nature of the carboxyl group on TM4 (D272 for MmpL4 and D276 for MmpL5) suggests that it becomes protonated/deprotonated during the transport cycle as observed for D407 and D408 of AcrB26,27, and plays a key role for proton translocation. The conserved tyrosines were more tolerant to mutations than the aspartates. Only the variant MmpL4Y845A was functionally impaired, while the variants MmpL4Y273F, MmpL4Y273A, MmpL4Y845F, MmpL5Y277F and MmpL5Y850F export mycobactin (Fig. 2d). From these experiments, a pattern emerged wherein the first DY-pair (D on TM4 and Y on TM10) is functionally more important than the second DY-pair. A similar pattern was reported for the DY pairs of M. tuberculosis MmpL3 (Supplementary Fig. 15)31.
For MmpL4, we extended our mutational analysis to charged and polar residues located in the vicinity of the two DY-pairs (Fig. 2d). The variants MmpL4T877V, MmpL4D912N and MmpL4R917A were found to be inactive and thus essential for MmpL4 function. Intriguingly, these residues are sitting adjacent to the functionally more important DY-pair (D272-Y845). The variants MmpL4T802V and MmpL4N846A exhibited partial mycobactin export activity, while all other variants (MmpL4R280A, MmpL4S309A, MmpL4T312V, MmpL4S806A and MmpL4R852A) showed wild-type activity in our functional assay (Fig. 2d, Supplementary Fig. 8).
MmpL4 forms a complex with MbtL and other acyl carrier proteins
In a final set of experiments, we investigated the serendipitously discovered binding of the E. coli acyl-carrier protein (Ec-ACP) to MmpL4 (Fig. 1c). The complex is formed by extensive electrostatic interactions including six arginine residues of the positively charged cytosolic surface of MmpL4 and four negatively charged residues of Ec-ACP (Fig. 6), a common feature of protein complexes involving ACPs32. Ec-ACP binds to the helix connecting TM6-TM7 and the loop between TM10-TM11 of MmpL4 primarily via its recognition helix (α-2) with additional interactions formed by helices α-1 and α-3 (Fig. 6c). M. tuberculosis has three ACP orthologues: AcpA, AcpM and MbtL. AcpA has a function homologous to the E. coli ACP as a co-factor in fatty acid and polyketide synthesis33. AcpM is an acyl carrier protein required for mycolic acid synthesis and couples this process to arabinogalactan synthesis by interacting with the arabinofuranosyltransferase AftD34. MbtL is encoded in the mbt-2 operon together with other mycobactin synthesis enzymes and catalyzes the transfer of acyl chains onto the iron chelating headgroup of mycobactin35. To investigate potential interactions of MmpL4 with ACPs of M. tuberculosis, biotinylated ACPs were immobilized on magnetic streptavidin beads to capture MmpL4-T, which was purified using a modified protocol to partially remove Ec-ACP (see methods). All three M. tuberculosis ACPs as well as Ec-ACP formed a complex with MmpL4 in this assay, whereas beads functionalized with the unrelated maltose binding protein (MBP) did not pull down MmpL4 (Fig. 6b). The ACPs did not pull down the E. coli RND transporter AcrB, which we included as an additional control (Fig. 6b). The interaction between MmpL4 and mycobacterial ACPs was further confirmed by co-purification of AcpM when MmpL4 was produced in M. smegmatis (Supplementary Fig. 6c). Since transcription of mmpL4 and mbtL is upregulated under iron starvation36, it is plausible to assume that they are functionally connected. To structurally analyze MmpL4 in the absence of a bound ACP, we generated the variant MmpL4R375E/R383E. A 5.1 Å cryo-EM map obtained for MmpL4-TR375E/R383E was indeed devoid of Ec-ACP and exhibited the same closed conformation we observed for all other MmpL4-T structures (Fig. 6d, Supplementary Fig. 16), indicating that binding to the ACP does not affect the transporter’s conformational equilibrium in a discernable manner.
a Electrostatic surface potential of MmpL4 and E. coli ACP. b Pull-down of anion-exchange chromatography purified MmpL4-T or RND transporter AcrB (shown as input on the right panel) using streptavidin-coated magnetic beads functionalized with biotinylated maltose binding protein (MBP, negative control), E. coli acyl carrier protein (Ec-ACP) or mycobacterial ACPs (AcpA – Rv0033, MbtL – Rv1344, AcpM – Rv2244) analyzed by Coomassie-stained SDS-PAGE. After pull-down, the magnetic beads were heated to 95 °C resulting in the release of all proteins. Bands on the SDS-PAGE corresponding to the ACPs and MBP are numbered (1 – MBP, 2 – Ec-ACP, 3 – AcpA, 4 – apo MbtL, 5 – holo MbtL, 6 – AcpM). The pull-down assay has been repeated three times and representative data is shown. Source data are provided as a Source Data file. c Close-up view of MmpL4-ACP interaction interface. Interacting residues on Ec-ACP and MmpL4 are shown as sticks. Salt bridges and hydrogen bonds are depicted as dashed lines. d Superimposition of 5.0 Å cryo-EM map obtained for MmpL4R375E/R383E-T (white, RR-T, devoid of Ec-ACP) with the 3.0 Å map obtained for MmpL4-T (petrol, with Ec-ACP bound).
Discussion
Our data suggest that MmpL4, and in analogy also MmpL5, constitute the central hub of a mycobactin export machinery that is unique to mycobacteria and works in concert with the mycobactin importer IrtAB37 at the inner mycobacterial membrane (Fig. 7a). Co-production of MmpS4 was essential to obtain high yields of MmpL4, indicating that MmpS4 might play a role in folding and/or multimerization of MmpL4. Supporting this hypothesis, a recent study investigating GFP-tagged MmpL5 in mycobacterial cells using single-molecule TIRF microscopy concluded that MmpL5 forms trimers only in the presence of MmpS538. We identified the predicted coiled-coil domain as crucial for mycobactin and bedaquiline export. To our knowledge it is the first coiled-coil domain described as part of a membrane transporter, and it distinguishes MmpL4 and MmpL5 from the cluster II MmpL proteins such as MmpL3. Size exclusion chromatograms of full-length versus truncated MmpL4 provide strong evidence that the coiled-coil domain is involved in oligomerization, but owing to the complex sample with co-purified MmpS4 and Ec-ACP (Fig. 1b), we were not able to determine the exact oligomer stoichiometry of the early eluting peak 1 (Fig. 1a). We hypothesize that the AlphaFold2 predicted coiled-coil domain assumes a role akin to membrane fusion proteins of tripartite multidrug efflux pump such as AcrA of the AcrAB-TolC complex39,40 by spanning the periplasmic space, as evident by a comparison of the two efflux systems (Supplementary Fig. 17). The coiled-coil domain could thereby act as an anchor to organize periplasmic and outer membrane-associated proteins required for mycobactin and drug efflux. In addition to MmpS4, a potential interaction partner of MmpL4 and MmpL5 could be Rv0455c13, a recently identified periplasmic protein that is required for mycobactin and drug efflux via MmpS4/L4 and MmpS5/L5 (Fig. 7a).
a Simplified schematic of the mycobacterial cell envelope and proteins involved in siderophore biosynthesis, secretion and import. For simplicity only the membrane-bound form of mycobactin is shown. The higher molecular order and the complete composition of the mycobactin secretion machinery including the outer membrane components are unknown. b Mechanistic model of siderophore transport mediated by MmpL proteins. Flipping and extraction of the substrate are likely coupled to proton translocation. Details are provided in the main text.
Our cryo-EM structures of MmpL4 identified a binding site for desferrated mycobactin, which is occupied by lipids in both our detergent-purified and nanodisc-reconstituted ‘apo’ control samples. Several residues of the cytosolic access pocket are repositioned for recognition of mycobactin. Binding of mycobactin to this site, which is accessible from the cytosol, represents the first step of the export mechanism (Fig. 7b). Functional analyses not only confirmed the importance of this binding pocket for siderophore secretion, but also indicated that it plays a role in bedaquiline efflux. To our knowledge, this cytosolic access pocket is unique among RND transporter structures determined thus far.
The periplasmic domains of MmpL4 adopt a closed conformation. By contrast, the multiple structures determined for MmpL3 all feature an open conformation at the periplasmic domains7,8,9,10. Given the resemblance of MmpL3 and MmpL4, it is plausible to assume that their structures represent two alternative conformations in the transport cycle of MmpL proteins to accomplish active transport. We propose that MmpL4 transitions to an open conformation by rigid body rotation of the PC, analogous to the rigid body rotation of the AcrB porter subdomains that open and close the access and deep binding pockets41,42. This process is most likely fueled by proton translocation events at the transmembrane domain as indicated by the essential nature of the aspartates D272 and D844, which are part of an extensive hydrogen bonding network of conserved charged and polar key residues at the center of the transmembrane domain of MmpL4 (Fig. 2d). Similar essential roles of aspartates have been identified in other proton-dependent transporters, whose carboxyl groups are crucial for proton transfer, including AcrB26,27 and MmpL331. Interestingly, the proton networks of MmpL4 and MmpL3 differ slightly (Fig. 4), probably reflecting altered protonation/deprotonation events associated with the different conformations of MmpL3 and MmpL4. As a result of the proton-driven conformational transition of MmpL4, we propose that mycobactin, after entering its binding pocket from the cytosol, is flipped across the cytoplasmic membrane, where it is accommodated at the outer leaflet into the voluminous pocket formed by TM7-TM8. From there, mycobactin could be extracted to the cavity formed between the two periplasmic porter subdomains as suggested for trehalose monomycolate extraction by MmpL3 (Fig. 7b). Further structural and biochemical investigation is required to shed light on how MmpL4 and MmpL5 accomplish these transport steps. This might require assembly of the full efflux machinery including MmpS4/5.
Recently, cryo-EM structures of the MmpL protein MSMEG_1382 of M. smegmatis were reported without any functional data43,44. This protein is annotated as MmpL5, however, three other M. smegmatis MmpL proteins MSMEG_0225, MSMEG_3496 and MSMEG_4383 have a higher sequence identity to MmpL5 and MmpL4 than MSMEG_1382 (Supplementary Fig. 14a). We show that MSMEG_0225 and MSMEG_3496, when produced with their respective MmpS proteins in the context of the native operons, restore mycobactin export in MtbΔmmpL4/mmpL5 in contrast to MSMEG_1382, shedding doubt on the claim43,44 that MSMEG_1382 is the M. smegmatis homolog of the M. tuberculosis MmpL5 protein (Supplementary Fig. 14b).
In conclusion, the structural and functional characterization of MmpL4 and MmpL5 provide insights into the molecular mechanism of mycobactin export, which is essential for virulence and intracellular growth of M. tuberculosis4. In addition, overproduction of MmpL5 is the leading cause of resistance against bedaquiline45, a TB drug that is used worldwide to treat infections with multi-drug and extensively drug-resistant M. tuberculosis strains20. Due to its excellent potency, bedaquiline has been included in several clinical trials aiming at shortening the treatment of TB patients from six to two months16,46. Hence, the structural and functional insights presented here provide a basis for the rational design of drugs that simultaneously block mycobactin secretion and bedaquiline efflux by MmpL4 and MmpL5. This will enable the development of urgently needed novel TB drugs that exploit synergistic effects of mycobactin poisoning1 and increasing the susceptibility of M. tuberculosis to bedaquiline21.
Methods
Strains, media and antibiotics
The Escherichia coli K-12 strain MC1061 was used for cloning and heterologous protein expression. Luria Broth (LB) liquid cultures and agar were used for cloning and Terrific broth (TB) liquid culture for protein expression. In E. coli 100 μg ml−1 ampicillin, 25 μg ml−1 chloramphenicol and 200 μg ml−1 hygromycin were added to the medium when required. M. smegmatis mc2 155 was grown at 37 °C in Middlebrook 7H9 liquid medium (Difco Laboratories) containing 0.05% Tween 80 and OADC or solid medium containing OADC and 4.5 ml/g glycerol. For M. tuberculosis mmpL4 expression in M. smegmatis cells were grown in 7H9 medium supplemented with 0.2% glycerol and 0.05% Tween 80. M. tuberculosis mc26230 strains were grown at 37 °C in Middlebrook 7H9 liquid medium supplemented with 0.2% glycerol, 10% OADC, 0.2% casamino acids, 24 μg/ml pantothenate, or 0.02% Hartmans de Bont (HdB) minimal medium (500 μM MgCl2, 7 μM CaCl2, 1 μM NaMoO4, 2 μM CoCl2, 6 μM MnCl2, 7 μM ZnSO4, 1 μM CuSO4, 15 mM (NH4)2SO4, 12 mM KH2PO4 (pH 6.8) and 1% (v/v) glycerol and was supplemented with 10% OADC, 0.2% casamino acids, 24 μg/ml pantothenate and 0.02% tyloxapol). Hygromycin was used at a concentration of 50 µg ml−1.
Cloning of mycobacterial genes
Constructs were generated using the fragment-exchange (FX) cloning platform47 and Gibson assembly48 using the primers listed in Supplementary Table 2. Briefly, open reading frames were amplified by PCR with primers containing a SapI overhang. Fragments were analyzed by agarose gel electrophoresis, extracted, purified and digested, before ligation into the entry vector pINIT (Addgene #46858), Sanger sequencing and further subcloning into expression or complementation vectors. The M. tuberculosis mmpS4-mmpL4 operon was amplified from Bacmid Rv265 and the mmpS5-mmpL5 operon from Bacmid Rv33949. The M. smegmatis homologs msmeg_0225-6, msmeg_1381-2 and msmeg_3495-6 were amplified from M. smegmatis by colony PCR. From pINIT, genes were sub-cloned into the expression vector pBXC3GH (Addgene #47070, containing a C-terminal 3 C cleavage site, GFP and deca-His-tag) by FX cloning for expression in E. coli. M. tuberculosis mmpL4 was subcloned into pACEC3GH50 for expression in M. smegmatis. For biotinylation, the mmpS4-mmpL4 operon was subcloned into pBXCA3GH (Addgene #47071), which contains an additional C-terminal avi-tag for enzymatic biotinylation. The coiled-coil domain of MmpL4 was truncated by backbone amplification of pBXC3GH_mmpS4-mmpL4 with primers excluding the coiled-coil domain followed by blunt end ligation. For the mycobactin toxicity assays the M. tuberculosis ΔmmpL4/mmpL5 mutant was complemented with the pML4324 and the pML4329 vectors, expressing either the wt or mutated mmpS4-mmpL4 or mmpS5-mmpL5 operons, respectively21. Mutations were introduced into the mmpL4 and mmpL5 genes following the QuikChange site-directed mutagenesis protocol using the pBXC3GH_mmpS4-mmpL4, pBXC3GH_mmpS5-mmpL5, pML4324 or pML4329 vectors as templates. The mmpS4-mmpL4 operon, mmpS5-mmpL5 operon and the M. smegmatis homologs were subcloned from the pBXC3GH vector into pML4324 or pML4329 backbone by Gibson assembly if needed. The M. tuberculosis ACPs were amplified from M. tuberculosis gDNA and the E. coli ACP from E. coli MC1061 and cloned into the pBXC3H vector (#47068, containing a C-terminal 3 C cleavage site and deca-His-tag) and pBXCA3H (Addgene #47072, containing an additional C-terminal avi-tag for enzymatic biotinylation) by FX cloning. AcrB was cloned from pBADAcrBAviC51 into pBXC3GH.
MmpL4 purification
MmpL4, MmpL4-T, MmpL4-TR375E/R378E and AcrB were expressed in E. coli and grown in 1.8 L TB supplemented with 100 ug ml−1 ampicillin at 37 °C for 2 h, followed by 40 min at 25 °C. Protein expression was induced with 0.02% L-arabinose and cells were cultured for 16 h for MmpL4 (and derivatives) and 4 h for AcrB. For MmpL4 production in M. smegmatis cells were grown in 10.8 L of 7H9 medium supplemented with 50 ug ml−1 hygromycin to an OD600 of 0.6. Protein production was induced by the addition of 0.02% acetamide and cells were cultured for 16 h. Cells were harvested by centrifugation for 15 min at 4 °C and 8000 g in a F9-6 × 1000 LEX centrifuge rotor (ThermoScientific) and resuspended in TBS (20 mM Tris/HCl pH7.5, 150 mM NaCl). DNase I was added prior to cell lysis by three passages through a Microfluidizer M-110P (Microfluidics) at 25 kPa on ice. All steps following cell lysis were performed at 4 °C. Cell debris was removed by centrifugation for 15 min at 10,000 g in a Sorvall SLA-1500 rotor (ThermoScientific). Membrane vesicles were harvested by ultracentrifugation for 2 h at 170,000 g in a Beckman Colter ultracentrifuge using a Beckman Ti45 rotor and resuspended in 30 ml TBS supplemented with 10% glycerol. Proteins were extracted by solubilization of membranes for 2 h with 1% (w/v) β-DDM (Glycon Biochemicals). Ultracentrifugation at 170,000 g for 30 min was performed to remove any unextracted material. Proteins were purified in TBS, 10% glycerol, 0.03% β-DDM via the C-terminal GFP using 1.5 ml of NHS activated Sepharose 4 Fast Flow (Cytiva) coupled to the anti-GFP nanobody (PDB-ID: 3K1K). Batch binding was performed for 2 h or overnight, after which the resin was washed with 70 ml of TBS, 10% glycerol, 0.03% β-DDM. The resin was incubated with 400 µg 3 C protease for 2 h to elute the protein by cleaving off the GFP-deca-His-tag. The flow through was collected and the resin was washed with 8 ml buffer to collect the cleaved protein. The sample was concentrated with 100 kDa MWCO concentrator (Merck Millipore) to 5–8 mg/ml and flash frozen and stored at -80 °C or immediately purified by size exclusion chromatography on a Superose 6 Increase 10/300 GL column (Cytiva) in TBS, 0.03% β-DDM. For the pulldown assay MmpL4-T was purified by affinity purification via the 3K1K resin as described above, followed by anion-exchange chromatography to obtain a sample only partially bound to ACP. The sample was diluted 25-fold in 50 mM Tris/HCl pH 8.0, 0.03% β-DDM and loaded on a Resource-Q ion-exchange column (Cytiva) followed by elution over a NaCl gradient, the main peak eluted between 100–200 mM NaCl. This peak was concentrated to 5–10 mg/ml using a 100 kDa MWCO concentrator (Merck Millipore).
ACP and MBP purification
The ACPs were expressed in 1.2 L TB and lysed as described for MmpL4 above. Cell debris was removed by centrifugation and the supernatant was incubated with 2 ml Nickle-NTA for batch binding for 2 h at RT. The resin was washed with 20 ml TBS, 30 mM imidazole. Proteins were eluted in 1 ml fractions using TBS, 300 mM imidazole and concentrated to 560 µl using 3 kDa MWCO concentrators (Merck Millipore). The ACPs were further purified by size exclusion chromatography using a Superdex 200 increase 10/300 GL column (Cytiva). Protein concentration of the ACPs was determined using the Pierce™ Coomassie (Bradford) Protein Assay Kit following manufacturer’s instructions (ThermoScientific). MBP was purified as described by Zimmermann et al.22.
Protein identification by mass spectrometry
One gel band was excised for each band of interest. Gel pieces were washed twice in 100 µl of 100 mM NH4HCO3/50% acetonitrile and once with 50 μl acetonitrile. Next 10 μl trypsin (5 ng/μl in 10 mM Tris/2 mM CaCl2, pH 8.2) and 30 μl of digestion buffer (10 mM Tris/2 mM CaCl2, pH 8.2) were added and the protein was digested at 60 °C for 30 min. Peptides were extracted with 150 μl 0.1% TFA/50% acetonitrile from the gel pieces for 15 min in an ultrasonic bath, dried and dissolved in 20 μl ddH2O containing 0.1% formic acid. Data was acquired by Liquid chromatography-mass spectrometry analysis (LC-MS/MS) on a nanoAcquity UPLC coupled to a Q-Exactive mass spectrometer (Thermo). For identification of E. coli ACP and M. smegmatis AcpM, detergent-purified samples of MmpL4-FL expressed in E. coli or M. smegmatis were digested using the iST Kit (PreOmics). The dried peptides were dissolved in 3% acetonitrile with 0.1% formic acid. Peptides were separated on a M-class UPLC and analyzed in a Obritrap mass spectrometer (Thermo). Data was searched against the E. coli database and the mycobacterial sequences of interest using the Mascot search engine (Matrixscience) and PEAKS Studio (Bioinformatic Solutions) or the Maxquant search engine (V2.0.1.0)52. All data was visualized in Scaffold (Proteome Software).
Biotinylation of proteins
Proteins were enzymatically biotinylated in vitro as described53. Briefly, the avi-tagged protein was concentrated to 50–100 µM after affinity purification for MmpL4-FL and to 200–900 µM for ACPs. Biotin was added at a 1:1.5 molar ratio (protein:biotin) as well as 5 mM ATP, 10 mM MgOAc and BirA. The sample was incubated at 4 °C for 16 h. The ACPs were subjected to an IMAC purification step to remove untagged BirA before size exclusion chromatography. MmpL4-FL was directly purified by size exclusion chromatography. MBP was biotinylated as described23.
Sybody selection
Sybody selection against full-length MmpL4 was performed as described previously23. Briefly, ribosome display was performed with the convex, concave and loop libraries. The output was cloned into the phagemid vector pDXinit (#110101) and two rounds of phage display were performed. Enrichment after phage display was calculated by qPCR. An enrichment factor over the control protein AcrB of 108 was achieved for the concave library, 12 for the loop library and 7 for the convex library, after the second round of phage display. The output of the selection was analyzed by ELISA, thereby identifying Sb09.
Reconstitution of MmpL4 in nanodiscs
Purified MmpL4-T was reconstituted into MSP2N2 nanodiscs with E. coli polar lipids and PC (3:1, w/w%) at a ratio of 1:5:100. First lipids and protein were mixed and incubated at 4 °C for 30 min under gentle agitation. Then MSP2N2 was added and incubated for another 30 min, resulting in 600 µl of 5 µM protein, 500 µM lipids and 25 µM MSP2N2, followed by addition of 120 mg of biobeads and incubation overnight. The next day, empty nanodiscs were separated from MmpL4-T by size exclusion chromatography on a Superose 6 Increase 10/300 GL column (Cytiva) in TBS.
Mycobactin-S isolation and desferration
Mycobactin was produced in and isolated from M. smegmatis mc2155 as described previously37. Briefly, M. smegmatis cells were grown in minimal medium in glass flasks. Cells were harvested and mycobactin was extracted overnight in 10% EtOH followed by filtration, addition of CHCl3 and saturation with FeCl3. The CHCl3 phase containing mycobactin was washed twice with water, dried and evaporated. For mycobactin desferration, mycobactin was resuspended in 6 ml CHCl3 and 2 ml MeOH was added followed by 2 ml concentrated HCl. Desferrated mycobactin partitions in the CHCl3 phase. The MeOH/HCl phase was discarded and the process was repeated two more times54. CHCl3 was evaporated and desferrated mycobactin was resuspended in EtOH and centrifuged for 5 min at 21,000 g to remove precipitates.
Cryo-EM sample preparation and data acquisition
Full-length MmpL4 in β-DDM was concentrated to 4.2 mg/ml and mixed with Sb09 at a 1:1 molar ratio. MmpL4-T and MmpL4-TR375E/R383E in β-DDM were concentrated to 9 mg/ml. Various substrates were added to MmpL4-T in β-DDM during the screening process including 174 µM clofazimine, 279 µM bedaquiline and 223 µM desferrated mycobactin (15 µM desferrated enterobactin was added to this sample to sequester any free iron present in the sample). We did not observe any of the substrates added to MmpL4-T in β-DDM compared to an apo dataset. The bedaquiline dataset gave the best resolution and was used for this study. The nanodisc samples were concentrated to 1.6 mg/ml for the apo sample and 1.7 mg/ml for the mycobactin sample. Prior to mycobactin addition, 10 µM iron-free enterobactin was added. Then, 50 µM desferrated mycobactin was added and the sample was incubated for 1 min at 25 °C before plunge freezing. The proteins were applied to glow discharged (0.39 mbar, 15 mA, 1 min) holey carbon grids (QuantiFoil R1.2/1.3 Au 200 or 300 mesh), blotted at 4 °C, 100% relative humidity for 4.5–5.5 s with a blotting force of 20 to remove excess sample using a Vitrobot Mark IV (ThermoFisher) and vitrified by plunging into an ethane/propane mixture. Samples were imaged on a Titan Krios G3i (300 kV, 100 μm objective aperture) equipped with a BioQuantum energy filter (20 eV energy filter slit) and a K3 direct electron detector (6k x 4k pixels). EPU 2.9 was used for automated data acquisition of dose-fractionated micrographs in a defocus range of −1.0 to 2.2 μm. Micrographs were recorded in counted super-resolution mode and Fourier cropped by 2. Data was acquired at a magnification of 130,000x, with a pixel size of 0.65 and 1.3 s exposure. 6,616 micrographs were collected for full-length MmpL4 with a dose of 20.7 e−/px/sec resulting in a total dose of 63.7e−/Å2. For truncated MmpL4 in β-DDM 6,759 micrographs were collected with a dose of 20.8 e−/px/sec resulting in a total electron exposure of 64.0 e−/Å2. For MmpL4-TRR a dose of 20.6 e−/px/sec was used, resulting in a total dose of 63.3 e−/Å2. 8,754 micrographs were collected for the apo nanodisc sample with a dose of 20.9 e−/px/sec and a total electron exposure of 64.3 e−/Å2. The mycobactin nanodisc dataset consists of 8,423 micrographs collected with a dose of 21.1 e−/px/sec and total electron exposure of 64.9 e−/Å2.
Cryo-EM image processing, model building and refinement
Data was processed in CryoSPARC v4.1.255. Data quality was monitored and initial processing steps including patch motion correction and patch CTF estimation were performed in CryoSPARC live. Good micrographs were selected based on the resolution of the estimated CTF fit and relative ice thickness. Particles were picked using the blob picker, extracted with a box size of 400 pixel and Fourier cropped to 100 pixels (2.6 Å per pixel). These were subjected to 2D classification, followed by ab initio reconstruction (C1 symmetry) and non-uniform refinement56. See Supplementary Figs. 2, 3, 10, 11 and 16 for specific details on the data processing pipeline. NU refinement using C1 symmetry resulted in a 6.3 Å map of MmpL4-FL in β-DDM, a 3.0 Å map for MmpL4-T in β-DDM, a 3.4 Å map for the apo MmpL4-T nanodisc sample, a 3.5 Å map for nanodisc-reconstituted MmpL4-T in complex with desferrated mycobactin and a 5.1 Å map for MmpL4-TR375E/R383E according to the 0.143 cut-off criterion57. The 3DFSC server was used to estimate the directional resolution58 and the local resolution estimation was carried out in cryoSPARC. The AlphaFold2 prediction of MmpL4 and ACP were docked into the density obtained for MmpL4-T in β-DDM and refined using PHENIX59, Coot60 and ISOLDE61. This model was used as an input for refinement in the nanodisc datasets. The Grade2 webserver v1.5.062 was used to generate restraints for ligands. Figures were made using ChimeraX63. Cavities of MmpL4 and MmpL3 were calculated using the 3 V web server (shell probe radius: 5 Å, solvent-excluded probe radius of 1.5 Å)30. The predicted binding energy of MBT to MmpL4 modeled in the two versions presented in Supplementary Fig. 12 was calculated using the PRODIGY-LIG webserver64,65. The 2D representation of binding was generated using LigPlot+ v.2.266,67.
Pulldown assay
12 µl Dynabeads MyOne Streptavidin C1 beads (Invitrogen) were washed twice by resuspension in 500 µl TBS, 0.5% BSA (w/v), followed by immobilization with a magnet rack and removal of the supernatant. Beads were incubated in the wash buffer for approximately 1 min before collection. The beads were then blocked in 500 µl TBS, 0.5% BSA (w/v). 30 µM of biotinylated ACPs or MBP were incubated with the beads for 5 min. Beads were washed three times with 500 µl TBS supplemented with 0.1% (w/v) β-DDM and 0.5% (w/v) BSA. 4 µM MmpL4-T after anion-exchange chromatography or AcrB were incubated with the beads for 20 minutes, which were subsequently washed three times with 500 µl TBS containing 0.03% β-DDM. For elution from the beads 10 µl SDS loading dye and 40 µl TBS containing 0.03% β-DDM were added and incubated for 5 minutes at 95 °C or 10 min at room temperature. The supernatant was then loaded onto a 4–15% SDS-PAGE gel. The pulldown assay was carried out three times on different days.
Mycobactin toxicity and bedaquiline susceptibility assays
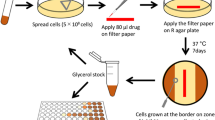
The function of MmpL4 and MmpL5 and their respective variants in siderophore secretion was determined in the MtbΔmmpL4/mmpL5 strain in our mycobactin toxicity assay21. The susceptibility of the MtbΔmmpL4/mmpL5 strain to mycobactin or bedaquiline was determined using a microplate Alamar Blue assay (MABA) as described previously21. Briefly, cultures of the MtbΔmmpL4/mmpL5 strain complemented with pML4324 (mmpS4-mmpL4) or pML4329 (mmpS5-mmpL5) vectors containing the respective mutations were grown in supplemented 7H9 medium to an OD600 of 1, were filtered using a 5 µM filter to remove clumps, and the cells were suspended in Hartmans de Bont (HdB) minimal medium supplemented with 0.2% casamino acids, 24 μg/ml pantothenate, 20 μM hemin and 0.02% tyloxapol. Aliquots of 100 μL of the cultures were incubated in microplates with increasing mycobactin or bedaquiline concentrations for five days at 37 °C and further incubated for six hours after addition of resazurin (90 μM). The fluorescence of the metabolically converted resazurin dye was measured as an indicator of cell viability.
Western Blot analysis of MmpL4 and MmpL5 proteins
To generate polyclonal MmpL4 antiserum, two rabbits were immunized with MmpL4-T purified in TBS containing 0.03% β-DDM (immunization were performed by the company Proteogenix). The polyclonal antibody was validated by performing Western Blotting against purified MmpL4 (Supplementary Fig. 9). M. tuberculosis mc26230 and derivative strains, were grown to an OD600 of ~2 in Middlebrook 7H9 medium (Difco Laboratories) supplemented with 0.2% glycerol, 10% ADS, 0.2% casamino acids, 24 μg/ml pantothenate, 0.02% Tyloxapol and 20 μM hemin. Cultures were harvested and washed twice with 1x PBS and then resuspended in 1x PBS. Bead beating was carried out using a Fast Prep FP120 Homogenizer with intermittent incubations on ice. After the first cycle of bead beating, 1% SDS was added to the cells followed by two cycles of bead beating. Cell lysates were subsequently centrifuged at 3200 x g to remove cell debris, resulting in the isolation of the whole cell lysate (WCL). The protein content in the lysates was determined using the Bradford (Bio-Rad) assay. For gel electrophoresis 5 μg of WCL were mixed with protein loading dye and separated on a 10% sodium dodecyl sulfate (SDS)-polyacrylamide gel. The proteins were then transferred to a polyvinylidene difluoride membrane using the Trans-Blot Turbo Transfer System (Bio-Rad). The blots were blocked with 5% skim milk (BD) in Tris-buffered saline (TBS: 50 mM Tris, pH 7.6, 150 mM NaCl) + 1% Tween 20 (TBST) overnight and incubated with the appropriate primary antibody in the following dilutions: rabbit polyclonal anti-mmpL4 antiserum (1:500, see above), rabbit polyclonal anti-flag (1:5000, Sigma-F7425, 069K4767). After three washes with TBST, the blots were incubated with the goat anti-rabbit IgG-HRP (1:10000, Sigma) antibody. All antibody dilutions were done in TBST. The Western blots were imaged using the Li-Cor Odyssey or Gel Doc XR+ (Bio-Rad).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Structural models have been deposited to the Protein Data Bank (PDB) with the accession codes 9GI0, 9GI2 and 9GI3. The cryo-EM data has been deposited in the Electron Microscopy Data Bank (EMDB) with the accession codes EMD-51370, EMD-51366, EMD-51368, EMD-51369 and EMD-51371. Plasmids and other data that support the findings of this study are available from the corresponding authors upon request. Source data are provided with this paper.
References
Jones, C. M. et al. Self-poisoning of Mycobacterium tuberculosis by interrupting siderophore recycling. Proc. Natl Acad. Sci. USA 111, 1945–1950 (2014).
Kurthkoti, K. et al. The Capacity of Mycobacterium tuberculosis To Survive Iron Starvation Might Enable It To Persist in Iron-Deprived Microenvironments of Human Granulomas. mBio 8, e01092-17 (2017).
Ratledge, C. Iron, mycobacteria and tuberculosis. Tuberculosis 84, 110–130 (2004).
Wells, R. M. et al. Discovery of a Siderophore Export System Essential for Virulence of Mycobacterium tuberculosis. PLoS Pathog. 9, 1–14 (2013).
Nikaido, H. RND transporters in the living world. Res. Microbiol. 169, 363–371 (2018).
Xu, Z., Meshcheryakov, V. A., Poce, G. & Chng, S. S. MmpL3 is the flippase for mycolic acids in mycobacteria. Proc. Natl. Acad. Sci. USA 114, 7993–7998 (2017).
Adams, O. et al. Cryo-EM structure and resistance landscape of M. tuberculosis MmpL3: an emergent therapeutic target. Structure 29, 1182–1191 (2021).
Zhang, B. et al. Crystal Structures of Membrane Transporter MmpL3, an Anti-TB Drug Target. Cell 176, 636–648 (2019).
Su, C. C. et al. MmpL3 is a lipid transporter that binds trehalose monomycolate and phosphatidylethanolamine. Proc. Natl. Acad. Sci. USA 166, 11241–11246 (2019).
Su, C. C. et al. Structures of the mycobacterial membrane protein MmpL3 reveal its mechanism of lipid transport. PLoS Biol. 19, 1–23 (2021).
Chim, N. et al. The Structure and Interactions of Periplasmic Domains of Crucial MmpL Membrane Proteins from Mycobacterium tuberculosis. Chem. Biol. 22, 1098–1107 (2015).
Domenech, P., Reed, M. B. & Barry, C. E. III Contribution of the Mycobacterium tuberculosis MmpL Protein Family to Virulence and Drug Resistance. Infect. Immun. 73, 3492–3501 (2005).
Zhang, L. et al. A periplasmic cinched protein is required for siderophore secretion and virulence of Mycobacterium tuberculosis. Nat. Commun. 13, 1–13 (2022).
Schnippel, K. et al. Effect of bedaquiline on mortality in South African patients with drug-resistant tuberculosis: a retrospective cohort study. Lancet Respir. Med. 6, 699–706 (2018).
Ndjeka, N. et al. High treatment success rate for multidrug-resistant and extensively drug-resistant tuberculosis using a bedaquiline-containing treatment regimen. Eur. Respiratory J. 52, 1–9 (2018).
Conradie, F. et al. Treatment of Highly Drug-Resistant Pulmonary Tuberculosis. N. Engl. J. Med. 382, 893–902 (2020).
Andries, K. et al. Acquired Resistance of Mycobacterium tuberculosis to Bedaquiline. PLoS One 9, 1–11 (2014).
Hartkoorn, R. C., Uplekar, S. & Cole, S. T. Cross-resistance between clofazimine and bedaquiline through upregulation of MmpL5 in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 58, 2979–2981 (2014).
Villellas, C. et al. Unexpected high prevalence of resistance-associated Rv0678 variants in MDR-TB patients without documented prior use of clofazimine or bedaquiline. J. Antimicrobial Chemother. 72, 684–690 (2017).
Derendinger, B. et al. Bedaquiline resistance in patients with drug-resistant tuberculosis in Cape Town, South Africa: a retrospective longitudinal cohort study. Lancet Microbe 4, 972–982 (2023).
Meikle, V., Zhang, L. & Niederweis, M. Intricate link between siderophore secretion and drug efflux in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 67, 1–20 (2023).
Zimmermann, I. et al. Synthetic single domain antibodies for the conformational trapping of membrane proteins. Elife 7, e34317 (2018).
Zimmermann, I. et al. Generation of synthetic nanobodies against delicate proteins. Nat. Protoc. 15, 1707–1741 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Murakami, S., Nakashima, R., Yamashita, E. & Yamaguchi, A. Crystal structure of bacterial multidrug efflux transporter AcrB. Nature 419, 587–593 (2002).
Eicher, T. et al. Coupling of remote alternating-access transport mechanisms for protons and substrates in the multidrug efflux pump AcrB. Elife 3, 1–26 (2014).
Seeger, M. A., Von Ballmoos, C., Verrey, F. & Pos, K. M. Crucial Role of Asp408 in the Proton Translocation Pathway of Multidrug Transporter AcrB: Evidence from Site-Directed Mutagenesis and Carbodiimide Labeling. Biochem 48, 5801–5812 (2009).
Seeger, M. A. et al. Structural Asymmetry of AcrB Trimer Suggests a Peristaltic Pump Mechanism. Science 313, 1295–1298 (2006).
Zhang, L. et al. Comprehensive analysis of iron utilization by Mycobacterium tuberculosis. PLoS Pathog. 16, 1–27 (2020).
Voss, N. R. & Gerstein, M. 3V: Cavity, channel and cleft volume calculator and extractor. Nucleic Acids Res. 38, 555–562 (2010).
Bernut, A. et al. Insights into the smooth-to-rough transitioning in Mycobacterium bolletii unravels a functional Tyr residue conserved in all mycobacterial MmpL family members. Mol. Microbiol. 99, 866–883 (2016).
Byers, D. M. & Gong, H. Acyl carrier protein: structure-function relationships in a conserved multifunctional protein family. Biochem. Cell Biol. 85, 649–662 (2007).
Kapopoulou, A., Lew, J. M. & Cole, S. T. The MycoBrowser portal: a comprehensive and manually annotated resource for mycobacterial genomes. Tuberculosis 91, 8–13 (2011).
Tan, Y. Z. et al. Cryo-EM Structures and Regulation of Arabinofuranosyltransferase AftD from Mycobacteria. Mol. Cell 78, 683–699 (2020).
Krithika, R. et al. A genetic locus required for iron acquisition in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. 103, 2069–2074 (2006).
Rodriguez, G. M., Voskuil, M. I., Gold, B., Schoolnik, G. K. & Smith, I. ideR, An essential gene in mycobacterium tuberculosis: role of IdeR in iron-dependent gene expression, iron metabolism, and oxidative stress response. Infect. Immun. 70, 3371–3381 (2002).
Arnold, F. M. et al. The ABC exporter IrtAB imports and reduces mycobacterial siderophores. Nature 580, 413–417 (2020).
Yamamoto, K., Nakata, N., Mukai, T., Kawagishi, I. & Ato, M. Coexpression of MmpS5 and MmpL5 Contributes to Both Efflux Transporter MmpL5 Trimerization and Drug Resistance in Mycobacterium tuberculosis. mSphere 6, e00518–e00520 (2021).
Du, D. et al. Structure of the AcrAB-TolC multidrug efflux pump. Nature 509, 512–515 (2014).
Chen, M. et al. In situ structure of the AcrAB-TolC efflux pump at subnanometer resolution. Structure 30, 107–113 (2022).
Seeger, M. A. et al. Engineered disulfide bonds support the functional rotation mechanism of multidrug efflux pump AcrB. Nat. Struct. Mol. Biol. 15, 199–205 (2008).
Kobylka, J., Kuth, M. S., Müller, R. T., Geertsma, E. R. & Pos, K. M. AcrB: a mean, keen, drug efflux machine. Ann. N. Y. Acad. Sci. 1459, 38–68 (2020).
Maharjan, R. et al. Cryo-EM structure of the Mycobacterium smegmatis MmpL5-AcpM complex. mBio 0, e03035–24 (2024).
Maharjan, R. et al. Structures of the mycobacterial MmpL4 and MmpL5 transporters provide insights into their role in siderophore export and iron acquisition. PLoS Biol. 22, 1–19 (2024).
Nimmo, C. et al. Opportunities and limitations of genomics for diagnosing bedaquiline-resistant tuberculosis: a systematic review and individual isolate meta-analysis. Lancet Microbe 5, 164–172 (2024).
Rosu, L. et al. Economic evaluation of shortened, bedaquiline-containing treatment regimens for rifampicin-resistant tuberculosis (STREAM stage 2): a within-trial analysis of a randomised controlled trial. Lancet Glob. Health 11, e265–e277 (2023).
Geertsma, E. R. & Dutzler, R. A Versatile and Efficient High-Throughput Cloning Tool for Structural Biology. Biochemistry 50, 3272–3278 (2011).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Brosch, R. et al. Use of a Mycobacterium tuberculosis H37Rv bacterial artificial chromosome library for genome mapping, sequencing, and comparative genomics. Infect. Immun. 66, 2221–2229 (1998).
Arnold, F. M. et al. A uniform cloning platform for mycobacterial genetics and protein production. Sci. Rep. 8, 9539 (2018).
Seeger, M. A. et al. Tuning the Drug Efflux Activity of an ABC Transporter in vivo by in vitro Selected DARPin Binders. PLoS One 7, e37845 (2012).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Kuhn, B. T. et al. Biotinylation of Membrane Proteins for Binder Selections. Methods Mol. Biol. 2127, 151–165 (2020).
Snow, G. A. The Structure of Mycobactin P, a Growth Factor for Mycobacterium Johnei, and The Significance of its Iron Complex. Biochem. J. 94, 160–165 (1965).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: Algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Rosenthal, P. B. & Henderson, R. Optimal Determination of Particle Orientation, Absolute Hand, and Contrast Loss in Single-particle Electron Cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Tan, Y. Z. et al. Addressing Preferred Specimen Orientation in Single-Particle Cryo-EM through Tilting. Nat. Methods 14, 793–796 (2017).
Adams, P. D. et al. PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr. D. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. 66, 486–501 (2010).
Croll, T. I. ISOLDE: A physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D. 74, 519–530 (2018).
Smart, O. S. et al. Grade2 Version 1.5.0. (Global Phasing Ltd, Cambridge, United Kingdom, 2021).
Meng, E. C. et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 32, 1–13 (2023).
Kurkcuoglu, Z. et al. Performance of HADDOCK and a simple contact-based protein–ligand binding affinity predictor in the D3R Grand Challenge 2. J. Comput. Aided Mol. Des. 32, 175–185 (2018).
Vangone, A. et al. Large-scale prediction of binding affinity in protein-small ligand complexes: The PRODIGY-LIG web server. Bioinformatics 35, 1585–1587 (2019).
Laskowski, R. A. & Swindells, M. B. LigPlot+: Multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011).
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995).
Acknowledgements
We thank all members of the Seeger lab for stimulating discussions and are grateful to Dr. Imre Gonda for sharing his deferrated mycobactin stock with us. Work in the laboratory of M.N. was supported by the National Institutes of Health grants R21 AI151239 and R01 AI137338. Work in the laboratory of M.A.S. was funded by the European Research Council (ERC) (consolidator grant n° 772190) and the Swiss National Science Foundation (PP00P3_170625). A.A.G. received support from the Forschungskredit of the University of Zurich (FK-21-041). The Center for Microscopy and Image Analysis (ZMB) of the University of Zurich is acknowledged for access to the electron microscope and we thank Dr. Simona Sorrentino and Dr. Piotr Szwedziak for their technical support. We also thank Petra Selchow and Tizian Griesser for sharing M. tuberculosis Bacmids and genomic DNA. We acknowledge the Functional Genomics Center Zurich (FGCZ) for the mass spectrometry analyses and technical support.
Author information
Authors and Affiliations
Contributions
J.C.E., A.A.G., M.N., and M.A.S. conceived the project. J.C.E. cloned the mmpS-mmpL genes, designed variants with input from A.A.G. and M.A.S. and generated variants, established the protein purification protocols, and performed sybody selection. A.A.G. established the nanodisc reconstitution. J.C.E. and A.A.G. prepared samples for cryo-EM, collected and analyzed the data. A.A.G. cloned, purified, and biotinylated ACPs, and J.C.E. performed the pulldown assay. J.C.E. built the MmpL4 and ACP models and refined them. V.M. performed all mycobactin toxicity and bedaquiline susceptibility assays and did the Western blot analyses. J.C.E., A.A.G., V.M., M.N., and M.A.S. interpreted the data. J.C.E. prepared the figures, and J.C.E, M.N., and M.A.S. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Earp, J.C., Garaeva, A.A., Meikle, V. et al. Structural basis of siderophore export and drug efflux by Mycobacterium tuberculosis. Nat Commun 16, 1934 (2025). https://doi.org/10.1038/s41467-025-56888-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56888-6
This article is cited by
-
Cryo-EM structures of mycobacteria MmpL5-AcpM complex
Science China Life Sciences (2025)