Abstract
Chromatin modifiers often work in concert with transcription factors (TFs) and other complex members, where they can serve both enzymatic and scaffolding functions. Due to this, active site inhibitors targeting chromatin modifiers may perturb both enzymatic and nonenzymatic functions. For instance, the antiproliferative effects of active-site inhibitors targeting lysine-specific histone demethylase 1A (LSD1) are driven by disruption of a protein-protein interaction with growth factor independence 1B (GFI1B) rather than inhibition of demethylase activity. Recently, next-generation precision LSD1 covalent inhibitors have been developed, which selectively block LSD1 enzyme activity by forming a compact N-formyl flavin adenine dinucleotide (FAD) adduct that spares the GFI1B interaction. However, the mechanism accounting for N-formyl-FAD formation remains unclear. Here we clarify the mechanism of these demethylase-specific inhibitors of LSD1, demonstrating that the covalent inhibitor-FAD adduct undergoes a Grob fragmentation. Using inhibitor analogs and structural biology, we identify structure-activity relationships that promote this transformation. Furthermore, we unveil an unusual drug resistance mechanism whereby distal active-site mutations can promote inhibitor-adduct Grob fragmentation even for previous generation compounds. Our study uncovers the unique Grob fragmentation underlying the mechanism of action of precision LSD1 enzyme inhibitors, offering insight into their reactivity with broader implications for drug resistance.
Similar content being viewed by others
Introduction
Dysregulation of chromatin states has been implicated in cancer pathogenesis, prompting the development of ‘epigenetic inhibitors’ that target chromatin modifying enzymes for oncology drug discovery1,2. As the first histone methylation ‘eraser’ to be discovered, LSD1, an FAD-dependent histone H3 lysine 4 (H3K4) demethylase, served as an instructive paradigm for the development of epigenetic inhibitors3,4. Based on mechanistic homology to previous monoamine oxidases, covalent inhibitors targeting LSD1 (e.g., GSK-LSD1 (1)) were rapidly discovered with the intent to antagonize enzyme activity, and subsequently advanced into clinical trials for treating hematological malignancies, including acute myeloid leukemia (AML), and solid tumors (Fig. 1a)5,6,7,8,9,10,11,12. Despite this original intent, it was later discovered that LSD1 inhibitors exert their anti-proliferative effects in AML by disrupting the LSD1-GFI1B interaction to activate GFI1B-target cis-regulatory elements and induce differentiation (Fig. 1b)5,6,7,8,9,13,14. GFI1B contains an N-terminal SNAIL-GFI1 (SNAG) domain, which mimics the histone H3 N-terminus to mediate high-affinity binding with the LSD1 active site, and recruits the broader LSD1-CoREST complex to silence its cognate genomic sites15,16. Consequently, LSD1 inhibitors have pleiotropic on-target effects: they (1) inhibit LSD1’s demethylase activity and (2) disrupt LSD1-TF interactions, the latter of which is responsible for their antiproliferative effects (Fig. 1b).
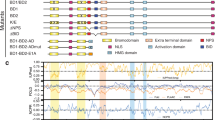
a Chemical structures of LSD1 inhibitors GSK-LSD1 (1), T-448 (2), and TAK-418 (3). b Schematic illustrating the two activities disrupted by active-site covalent LSD1 inhibitors. Upon GSK-LSD1 (1) binding, LSD1 demethylase activity is disrupted (left) and the LSD1-GFI1B complex is dissociated (right). Cartoon adapted from Waterbury et al. 13. c Schematic of T-448 (2) LSD1-inhibition mechanism where the initial covalent inhibitor-FAD adduct (4) fragments to N-formyl-FAD (5). Cartoon adapted from Waterbury et al. 13. d Binding curves showing fluorescence polarization (y-axis) for increasing concentrations of LSD1-CoREST (x-axis) in the presence of a fluorescently labeled GFI1B(2-10) peptide after treatment with DMSO, GSK-LSD1 (1), or T-448 (2) (n = 2 biological replicates). e LC-MS spectra of FAD adducts after extracting recombinant LSD1 treated with d-T448 (6) for 4 h. f Schematic of proposed mechanism of covalent FAD-inhibitor adduct fragmentation to N-formyl-FAD (5) and T-448 styrene (10) via a Grob fragmentation. g Formation of T-448 styrene (10) detected by LC-MS after extracting recombinant LSD1 treated with T-448 at the indicated time points (n = 2 biological replicates). h Kinetics of styrene formation for LSD1 treated with T-448 (2) or p-Br-T448 (11) analog. Styrene formation was quantified relative to an internal standard and standard curves from chemically synthesized styrene standards (n = 3 biological replicates). Results in (d–h) are representative of two independent experiments. Data in (g) are provided in Supplementary Data 1. Source data are provided as a Source Data file for (d–h).
While disruption of LSD1-TF interactions drives the antiproliferative effects of these ‘dual’ LSD1 inhibitors, it also results in dose-limiting hematological toxicities, such as thrombocytopenia, as the LSD1-GFI1B complex is essential for normal hematopoiesis5,15,17. This on-target toxicity presents challenges for the development of LSD1 inhibitors, especially for non-oncology indications where the enzyme activity of LSD1 is the desired function to inhibit and not LSD1-TF interactions. However, given the similar occupancy of the histone H3 substrate and the SNAG domain in the LSD1 active site, the development of ‘pure’ LSD1 enzyme inhibitors seemed highly unlikely.
Remarkably, through a phenotypic screen, Takeda identified precision LSD1 inhibitors that solely inhibit enzyme activity while sparing the GFI1B interaction18,19. These inhibitors, including T-448 (2) and TAK-418 (3) (Fig. 1a), contain a cyclopropylamine core that can in principle form a covalent inhibitor-FAD adduct similar to other related LSD1 inhibitors (Fig. 1c). However, this initial adduct undergoes a transformation to form an N-formyl-FAD adduct. While the N-formyl-FAD adduct is sufficient to inhibit enzyme activity, it is small enough to accommodate the LSD1-GFI1B interaction. Consistent with this notion, T-448 (2) has no antiproliferative activity in AML cell lines18. These unique properties make T-448 (2) effective for neurodevelopmental indications, where it is desirable to target LSD1’s demethylase activity without disrupting the LSD1-GFI1B complex due to the associated hematotoxicity. Indeed, TAK-418 (3) has been well tolerated in human clinical trials20. Despite the remarkable precision and therapeutic index of these demethylase activity-specific inhibitors, the mechanism of N-formyl-FAD formation as well as its broader implications on pharmacologically targeting LSD1 remain to be fully determined.
Here, we resolve the mechanism of LSD1 inhibitor-FAD adduct fragmentation to N-formyl-FAD and other related species. Using a series of T-448 analogs and deuterium labelling studies, we demonstrate that this transformation is consistent with a Grob fragmentation mechanism and further identify the structural features of T-448 (2) that are essential for this activity, especially the meta positioning of the aryl amide substituent. Using X-ray crystallography, we capture the initial inhibitor-FAD adduct before rearrangement has occurred, revealing T-448’s unique positioning in the LSD1 active site. Furthermore, we identify and mechanistically characterize an unusual FAD-distal mutation that confers resistance to LSD1-GFI1B inhibitors by promoting Grob fragmentation to N-formyl-FAD, revealing features of the enzyme active site that alter the stability of covalent FAD-inhibitor adducts. Altogether, our study sheds light on the mechanism of action of clinical-stage LSD1 inhibitors and demonstrates broader possibilities for drug resistance.
Results
Inhibitor-FAD adduct fragmentation proceeds through a Grob Mechanism
Many covalent LSD1 inhibitors are mechanism-based inactivators, based on the tranylcypromine phenyl cyclopropylamine core. These inhibitors, such as GSK-LSD1 (1), form a carbon-carbon bond with the FAD C4a position, where the resultant inhibitor-adduct may reside in the open aldehyde form or as a 5-membered cyclic hemi-aminal21,22. While T-448 (2) contains the same cyclopropylamine core as GSK-LSD1 and forms a covalent adduct (4), this adduct, unlike the one formed with GSK-LSD1, is unstable and rearranges to N-formyl-FAD — allowing T-448 to selectively target LSD1’s demethylase activity without disrupting the LSD1-GFI1B interaction (Fig. 1c). However, the mechanism for this transformation is unknown. We validated that N-formyl-FAD is detected by LC-MS after treating purified LSD1 with T-448 (Supplementary Fig. S1a). In agreement, treatment of LSD1-CoREST with T-448 leads to time-dependent binding of a fluorescently labelled SNAG peptide (residues 2-10 of GFI1B) as monitored by fluorescence polarization (Fig. 1d), and T-448 has no effect on the proliferation of SET-2 cells, an AML cell line highly sensitive to GSK-LSD1 treatment (Supplementary Fig. S1b,c)14,23. To probe the mechanism for this rearrangement, we synthesized a deuterated T-448 derivative, 2-d-T-448 (6), in which deuterium is incorporated on the C2-carbon position to determine whether this portion of the molecule is incorporated in the formyl group (Fig. 1e). Both the initial full inhibitor-FAD adduct (7) and deuterated N-formyl-FAD (8) were detected by LC-MS after treating LSD1 with 2-d-T-448 (6) (Fig. 1e, Supplementary Fig. S1d). Based on these results we hypothesized that T-448-FAD (9) undergoes a Grob fragmentation via a hemi-aminal intermediate to yield N-formyl-FAD (5) and the corresponding styrene (10) (Fig. 1f)24,25. Consistent with this proposed mechanism, formation of the N-formyl-FAD adduct was accompanied by detection of T-448-derived styrene 10 by LC-MS, with increasing conversions observed over longer time periods (Fig. 1g, Supplementary Fig. S1e). Altogether, our data support the notion that after formation of an initial full T-448-FAD adduct, this species subsequently undergoes a Grob fragmentation where the C2-position of T-448 gives rise to the N-formyl carbon.
After observing the formation of both Grob fragmentation products, we optimized an LC-MS assay to quantify styrene byproduct formation as a proxy for T-448 adduct fragmentation. We quantified T-448 styrene formation over time against an internal standard to obtain a first-order kinetic plot for adduct fragmentation with an estimated rate constant k ≈ 0.11 h-1 (Fig. 1h). Due to poor stereoelectronic alignment of the participating orbitals required for a concerted reaction, the cyclic hemiaminal T-448-FAD adduct likely proceeds stepwise, beginning with C–C bond cleavage between the adduct C11 and FAD C4a (Fig. 1f). This bond cleavage could in theory proceed through a one- or two-electron process to generate either a radical or carbonium intermediate at the benzylic carbon, respectively. To investigate these possibilities, we synthesized bromo-analog p-Br-T-448 (11), which contains an electron-withdrawing bromine substituent at the para position. Treatment of LSD1 with the bromo-substituted analog followed by measurement of styrene formation yielded an estimated rate constant of k ≈ 0.04 h-1 (Fig. 1h). Although the reduced fragmentation rate is consistent with the bromine destabilizing a putative carbocation intermediate at the benzylic carbon, we cannot rule out the possibility that the bromine substituent alters the mechanism more drastically altogether. Collectively, these results are consistent with the T-448-FAD adduct undergoing a stepwise Grob fragmentation.
Structure-activity relationships of T-448 analogs
We next evaluated the chemical features of T-448 necessary to yield efficient N-formyl-FAD formation. We first tested whether moving the 1,3,4-thiadiazole amide from the meta to para position would impact fragmentation. The para-substituted T-448 isomer 12 yielded no detectable N-formyl-FAD, suggesting that substitution at the meta position is required for T-448 fragmentation (Fig. 2a). Based on this result, we next synthesized ‘pared-back’ derivatives of T-448 to probe the necessity of the thiadiazole and amide groups for fragmentation (Fig. 2a). Minor modifications to the thiadiazole heterocycle, such as removal of the methyl group (13) or conversion to a thiazole (14), led to negligible changes in fragmentation, as assessed by fluorescence polarization and LC-MS, leading us to conclude that the thiadiazole of T-448 is not inherently privileged for Grob adduct fragmentation (Fig. 2a, b). In fact, replacement of the thiadiazole altogether with a phenyl group, to yield N-phenyl benzamide 15, still allowed for formation of N-formyl-FAD — albeit 15 still partially inhibited SNAG peptide binding (Fig. 2a, b). LC-MS demonstrated that both the benzamide analog styrene (21) and N-formyl-FAD adduct along with the full inhibitor-FAD adduct (20) are generated upon treatment with 15, consistent with the proposed Grob fragmentation mechanism (Supplementary Fig. S2a). Quantification of FAD inhibition products by LC-MS revealed that the adducts generated by benzamide 15 exhibited attenuated Grob fragmentation compared to T-448, producing only 16% N-formyl-FAD as the minor product compared to T-448 (78% N-formyl-FAD) (Fig. 2a). However, 15 did not inhibit cell proliferation nor disrupt LSD1-GFI1B binding in SET-2 cells, and the corresponding styrene was also detected in cell media following compound treatment, similar to T-448 (Fig. 2c, Supplementary Fig. S1b, c, S2b). Taken together, these data support that while the inhibitor-FAD adduct formed by 15 exhibits slower Grob fragmentation, even low levels of N-formyl-FAD formation are sufficient to maintain the LSD1-GFI1B complex and AML cell proliferation.
a Table summarizing enzyme inhibition (IC50) and fragmentation activity of T-488 (2) analogs where the % of each FAD species is reported along with the relative % N-formyl-FAD formed. IC50 values are reported as mean ± s.d. of n = 3 biological replicates (see Fig. S1b and S2c for full inhibition curves). b Binding curves showing fluorescence polarization (y-axis) for increasing concentrations of LSD1-CoREST (x-axis) in the presence of a labelled SNAG(2-10) peptide after treatment with vehicle or LSD1 inhibitor (n = 3 biological replicates). c Co-IP of FLAG-LSD1 with GFI1B after vehicle or LSD1 inhibitor treatment (500 nM, 48 h) in transiently transfected HEK 293T cells. * denotes skipped lane. Data in (a–c) are representative of two independent experiments. Data in (a) are provided in Supplementary Data 1. Source data are provided as a Source Data file for (b, c).
Given that the N-phenyl benzamide substituent of the 15-inhibitor adduct is insufficient to fully block fragmentation, we next tested more substantial modifications to the T-448 scaffold (Fig. 2a). Reversing the amide group of the N-phenyl benzamide (16) further reduced formation of N-formyl-FAD production to 6%, as did replacement of the N-aryl benzamide with a methyl benzoate (17, 4% N-formyl-FAD) and an N-methyl benzamide (18, 2% N-formyl-FAD). Finally, we observed full inhibition of the chemical fragmentation to N-formyl-FAD upon replacement of the T-448 substituent with an N,N-dimethyl benzamide (19). Of note, although we evaluated these analogs at doses where full inhibition was observed, most modifications to the T-448 scaffold also substantially reduce inhibitor potency, suggesting that some of these modifications may reduce formation of the initial covalent adduct (Supplementary Fig. S2c). Overall, these data indicate that the meta-substituted secondary amide containing aromatic substituents is critical to produce significant levels of N-formyl-FAD.
Structural studies of T-448 and derivatives
To understand how changes in inhibitor structure alter the propensity for FAD-adduct rearrangement, we obtained X-ray crystal structures of LSD1-CoREST bound by T-448 (2) or 15. By soaking the crystals with inhibitor rather than through co-crystallization, we were able to capture the cyclic hemi-aminal T-448-FAD intermediate prior to Grob fragmentation, providing structural insight into this transformation. Specifically, LSD1-CoREST crystals were initially soaked with T-448 (2) or 15 (1 mM), back-washed into solutions lacking the inhibitor, and soaked with 1 mM of the SNAG peptide prior to flash freezing. Structures were obtained at multiple timepoints to determine how the FAD-inhibitor adducts change over time (Fig. 3a, b). Structures obtained after 10 min revealed that T-448- (9) and 15-inhibited FAD adducts (20) exhibited a distinct conformation, where the cyclic hemi-aminal adduct points its aromatic groups away from the ribityl-flavin in a staggered orientation (Fig. 3a, c). This differs from the classic FAD N5-C4a cyclic adduct observed in prior LSD1-inhibitor structures where the aromatic ring is instead positioned on the flavin re- side22,26,27. After 3 days, the T-448-FAD adduct was fully converted to N-formyl-FAD, allowing LSD1 to bind the SNAG peptide in the same conformation as the native protein (Fig. 3a, b, Supplementary Fig. S3a)18,28. The crystallographic analysis also demonstrated that the N-formyl-FAD complex of LSD1 permits the binding of the N-terminal tail of histone H3 (Supplementary Fig. S3b, c). For 15, the conversion to N-formyl-FAD after 3 days of back washing is likely too slow or only partial under the crystallization conditions. Instead, only rotation around the benzylic bond was observed after 3 days, positioning the inhibitor aromatic ring above the re-face of the flavin cofactor (Fig. 3c).
a 2Fo-Fc electron density maps (contour level 1.2-1.4 σ) of LSD1-CoREST in complex with T-448 (2) after adding inhibitor for 10 min, crystal back-washing for 3 d, followed by the addition of the SNAG peptide. b Zoomed-in view that illustrates superposition of the FAD adduct with T-448 after 10 min (gray) and 3 d (pink). The SNAG peptide (green) is predicted to sterically clash with the initial T-448-FAD adduct (9) shown in gray. c 2Fo-Fc electron density maps (contour level 1.2 σ) for 15 after 10 min of inhibitor soaking (left; dark purple) followed by 3 d of back washing (right; magenta). The initial cyclic adduct rotates around the benzylic bond to position the aryl substituent of the inhibitor fragment above the re-face of flavin. d The two enantiomers of T-448 form cyclic (T-448-FAD (9); gold) and linear adducts (T-448-FAD (4); green), respectively. The 2Fo-Fc electron density maps are contour level 1.2 σ and were obtained after soaking the crystals for 10 min. e Superposition of the T-448-FAD adducts obtained after soaking LSD1-CoREST crystals with the two enantiomers of T-448. W695 orientation is perturbed with the (1R,2S)-T448 structure, likely due to formation of a branched, non-cyclic C4a-FAD adduct. f Superposition of T-448 (2) (gold), 12 (blue), 15 (purple), 16 (cyan), and 18-FAD (salmon) adducts obtained after soaking LSD1-CoREST crystals with the respective inhibitors for 10 min. FAD adducts formed with 12, 16 and 18 adopt the classic “above the re-face” conformation observed in crystal structures of LSD1 bound to tranylcypromine derived inhibitors. Data are provided in Supplementary Data 3 and 4.
To investigate the effect of inhibitor stereochemistry on N-formyl-FAD formation, we synthesized and tested T-448 prepared as single enantiomers (2)29. Consistent with the literature, the (1S,2R)-eutomer is more active at inhibiting LSD1 demethylase activity compared to the (1R,2S)-distomer, which was 100-fold weaker (Supplementary Fig. S3d)29. Quantification of FAD inhibition products by LC-MS revealed that the (1R,2S)-distomer failed to form the inhibitor-FAD adduct and Grob fragmentation products compared to the (1S,2R)-eutomer, which produced both the full inhibitor-FAD and N-formyl-FAD adducts (10% and 75%, respectively) (Supplementary Fig. S3e). Furthermore, the corresponding styrene was only observed by LC-MS for the (1S,2R)-eutomer (Supplementary Data 1). Neither enantiomer impaired proliferation of SET-2 cells (Supplementary Fig. S3f). This can be explained by efficient fragmentation to N-formyl-FAD upon treatment with the (1S,2R)-eutomer and the weak inhibitory activity of the non-fragmenting (1R,2S)-distomer.
The structures of LSD1-CoREST bound by each of the T-448 enantiomers were collected after 10 min of inhibitor soaking (Fig. 3d). The active T-448 eutomer formed an N5-C4a cyclic hemi-aminal FAD adduct (9) where the inhibitor adopts a staggered orientation relative to FAD as observed with racemic T-448 (Fig. 3a). By contrast, incubation with the distomer resulted in a linear C4a-linked FAD adduct, where its binding pose is slightly rotated compared to the (1S,2R)-eutomer and is accompanied by a distinct shift in the active-site W695 residue (Fig. 3e). Overall, the distomer forms a linear C4a-linked FAD adduct that is likely less stable once released from the protein and thereby difficult to detect by LC-MS (Supplementary Fig. S3e, g).
Next, we crystallized 12, 16, 17, 18 and 19 with LSD1-CoREST to determine if there are any structural differences in the resulting inhibitor-FAD adducts that prevent fragmentation to N-formyl-FAD. Incubation with these compounds produced the classic inhibitor-FAD N5 and C4a cyclic adduct that positions the aromatic ring on the flavin re-face in an eclipsed conformation (Supplementary Fig. S3h). Overlaying these adducts with that of the active T-448 eutomer suggests that the corresponding 5-membered cyclic intermediates of these analogs possess the opposite stereochemistry at the benzylic C2-carbon (Fig. 3f). After 3 days of incubation, 12 produced a linear adduct at N5 of FAD whereas the cyclic adducts formed by 16, 17, 18 and 19 were stable, even after prolonged incubation (3 days) (Supplementary Fig. S3i). Altogether, our findings support the notion that formation of N-formylated FAD arises from the rearrangement of the initial N5-C4a cyclic adduct possessing a staggered as opposed to eclipsed conformation (Supplementary Fig. 3f, S3g). For the non-formylating inhibitors, the N5-C4a cyclic adduct may remain stable over time, open to form a C4a-FAD linked adduct containing a free aldehyde via opening of the hemi-aminal (green arrows) or might alternatively rearrange by forming a carbocation intermediate (black arrows) that then undergoes a 1,3-hydride transfer, as reported in some terpene synthases, to generate the linear N5-FAD adduct (blue arrows) (Supplementary Fig. S3g)22,30. While rearrangement of the non-fragmenting inhibitors would proceed through these mechanisms, the staggered conformation of T-448 and 15 cyclic adducts combined with their meta-amide moiety may instead facilitate Grob fragmentation to N-formyl-FAD (red arrows) (Fig. 3f, Supplementary Fig. S3g). Further experiments will be necessary to fully test this proposed mechanistic model.
LSD1 loop deletion mutant promotes fragmentation and drug resistance
By ultimately forming the N-formyl-FAD adduct that can accommodate GFI1B within the LSD1 active site, T-448 has no effects on the proliferation of AML cell lines despite inhibiting enzyme activity — supporting that LSD1 demethylase activity is not essential in this context18,19,29. Furthermore, our results suggest that T-448 is not uniquely privileged, and that other tranylcypromine derivatives can lead to formation of the N-formyl-FAD adduct, albeit to a lesser extent. Notably, our data with benzamide 15 suggest that even minor formation of the N-formyl-FAD adduct may be sufficient to allow unimpeded AML growth. Together, these findings raise the possibility that mutations in LSD1 could promote Grob fragmentation of antiproliferative tranylcypromine derivatives and subsequent formation of the N-formyl-FAD adduct, and in doing so confer drug resistance to these inhibitors in the context of cancer cell lines.
To explore this idea, we assessed our prior CRISPR-suppressor scan of LSD1, where we identified resistance mutations in LSD1 that were selectively enriched by different analogs of the inhibitor GSK-LSD114. Generally, the most highly enriched single-guide RNAs (sgRNAs) targeted exon 15, leading to mutations that altered the catalytic pocket and disrupted both LSD1 enzyme activity and inhibitor binding. However, distinct mutations were also enriched in exon 16 — most notably by the GSK-LSD1 analog AW4 (32) (Fig. 4a). To characterize exon 16 mutations, sgT684, a highly enriched sgRNA targeting exon 16, was transduced into SET-2 cells, which were then treated with vehicle or AW4. Genotyping exon 16 of these AW4-treated cells revealed that the most prevalent mutation comprises a 4-amino acid deletion of T684–S687 (LSD1 ΔTTAS), in a loop >18 Å from the FAD cofactor (Fig. 4b). A clonal SET-2 cell line harboring LSD1 ΔTTAS displayed partial resistance to AW4 (Fig. 4c). We next tested whether LSD1 ΔTTAS disrupts enzyme activity or inhibitor binding by purifying and testing the mutant. We confirmed that LSD1 ΔTTAS contains FAD, and it was eluted at a similar retention time as wild-type LSD1 by size-exclusion chromatography, suggesting proper folding (Supplementary Fig. S4a-b, Source Data file). Recombinant LSD1 ΔTTAS complexed to CoREST (Δ1-307) was enzymatically active in H3K4me2 demethylation assays and inhibited by both GSK-LSD1 and AW4, similar to wild-type LSD1 (Fig. 4d). Moreover, the ΔTTAS mutation had no impact on the kinetics of demethylation and no demethylase activity was recovered after inhibitor treatment over time, even over the course of days (Supplementary Fig. S4c). Consistent with this, AW4 fully bleached the oxidized flavin peak (458 nm) within the first few minutes of inhibitor incubation with LSD1 ΔTTAS, analogous to the activity of other tranylcypromine LSD1 inhibitors, and the flavin spectra remained bleached over time (Supplementary Fig. S4b)6,26. These results suggest that LSD1 ΔTTAS remains inactivated by AW4. Therefore, LSD1 ΔTTAS does not appreciably block drug binding nor disrupt enzyme activity, differentiating this drug-resistant mutant from those identified in our previous study that alter the active site and inhibitor binding14.
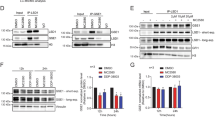
a Homology model of ΔTTAS LSD1 (salmon) overlaid with wild-type LSD1 (gray) is shown to highlight the deletion of the wild-type loop (cyan) compared to the truncated mutant loop (magenta) (PDB: 2hko). Chemical structure of GSK-LSD1 analog AW4 (32) is shown below. Figure adapted from Vinyard et al.14 b Schematic showing the most abundant genotype in exon 16 after introducing sgT684 and treating with AW4 (32) for 6 weeks to generate the ΔTTAS LSD1 clonal cell line. c Bar plot of relative growth (y-axis) of wild-type and ΔTTAS LSD1 SET-2 cells after 10 day treatment with 1 μM of AW4 (32). d Dose-response curves of recombinant wild-type and ΔTTAS LSD1-CoREST treated with GSK-LSD1 or AW4 (32) relative to vehicle control (n = 3 biological replicates). e Binding curves showing fluorescence polarization (y-axis) for increasing concentrations of LSD1-CoREST (x-axis) in the presence of fluorescently labeled GFI1B(2-10) peptide after treatment with DMSO, GSK-LSD1 (1) or AW4 (32) (n = 2 biological replicates). f MS spectra of FAD adducts formed upon ΔTTAS LSD1-CoREST treatment with AW4 (32) for 24 h following FAD extraction and LC-MS analysis. g Formation of AW4-styrene (34) detected by GC-MS after extracting recombinant ΔTTAS LSD1-CoREST treated with AW4 (32) at the indicated time points. Data in (c) represent mean ± standard error of the mean (s.e.m) across three biological replicates where asterisks denote significant p-values. (****p < 0.0001, ***p < 0.001, **p < 0.01); two-tailed unpaired t-test was performed; wild-type vs ΔTTAS p = 2.23 × 10-6 (t = 40.48, df = 4). Results in (c–g) are representative of two independent experiments. Data in g are provided in Supplementary Data 1. Source data are provided as a Source Data file for (c–g).
We next investigated whether AW4 permits LSD1 ΔTTAS-GFI1B binding despite forming a covalent inhibitor-FAD adduct. We monitored binding of LSD1-CoREST complexes to a fluorescently labelled SNAG (GFI1B 2-10) peptide in the presence or absence of GSK-LSD1, AW4, or vehicle control22,25. While GSK-LSD1 and AW4 treatment blocked SNAG peptide binding to wild-type LSD1-CoREST, SNAG peptide binding to ΔTTAS LSD1-CoREST was observed at later timepoints in the presence of AW4 but not GSK-LSD1 suggesting that ΔTTAS LSD1-CoREST binds to GFI1B in the presence of AW4 in a time-dependent manner (Fig. 4e, Supplementary Fig. S4d).
Based on these findings, we next asked whether the ΔTTAS mutation may promote fragmentation of the AW4–FAD adduct to form the N-formyl-FAD adduct and permit GFI1B binding. Consistent with this notion, after treating ΔTTAS LSD1-CoREST with AW4 for 24 h we observed the formation of N-formyl-FAD in addition to the expected inhibitor-FAD adduct (33), similar to our previous experiments using wild-type LSD1 and T-448 analogs, while no N-formyl-FAD was detected with wild-type LSD1-CoREST treated with AW4 under the same conditions nor with ΔTTAS LSD1-CoREST treated with GSK-LSD1 (Fig. 4f, Supplementary Fig. S4e, f, Supplementary Data 1). Furthermore, formation of N-formyl-FAD was accompanied by detection of AW4-derived styrene (34) by GC-MS, with increasing styrene conversion observed over longer time points (Fig. 4g). Moreover, styrene (34) was also detected from extracting the cell media of ΔTTAS SET-2 after AW4 treatment for 24 h (Supplementary Fig. S4g). While the ΔTTAS mutation promotes AW4-FAD fragmentation, it failed to accelerate the fragmentation of T-448 (2) (52% N-formyl-FAD, 20% T-448-FAD adduct, 28% free FAD) nor did it promote GSK-LSD1 and 12 fragmentations to N-formyl-FAD (Supplementary Data 1). Altogether, these results are consistent with the ΔTTAS mutation selectively promoting AW4-FAD adduct fragmentation to N-formyl-FAD. As T-448 is ineffective at blocking AML proliferation, since it permits LSD1-GFI1B complexation (Fig. 2c, Supplementary Fig. S1c), these findings suggest that the ability to fragment the inhibitor-FAD adduct to N-formyl-FAD may be a viable drug-resistance mechanism.
Distal crosstalk between the TTAS loop and the active site can modulate Grob fragmentation
To understand the molecular basis for selective AW4 fragmentation, we next investigated the LSD1-CoREST structure treated with AW4 (Fig. 5a, b). Unfortunately, ΔTTAS LSD1-CoREST was refractory towards crystallization since the TTAS loop mediates key crystal contacts, so only structures with wild-type LSD1 were obtained. These structures revealed that AW4 forms the N5- and C4a-cyclic adducts with the same conformation as 12 and the other non-formyl forming inhibitors, opposite to the staggered conformation observed with the T-448-FAD adduct (Fig. 5c, Supplementary Fig. S5a). These results are consistent with AW4 failing to form N-formyl-FAD with wild-type LSD1. To determine how the AW4-FAD adduct changes over time, crystals were soaked and then back-washed into solution lacking inhibitor and either directly flash frozen or soaked with 1 mM of the SNAG peptide prior to flash freezing. Compared to the initial structure, the AW4-FAD (35) adduct after 3 days undergoes a moderate lateral swing of the inhibitor aromatic group (Fig. 5d). This likely arises from rearrangement of the FAD-inhibitor adduct, where the initially formed 5-membered cyclic adduct rearranges to a linear N5-acylated adduct between the inhibitor and N5, similar to what was also observed with 12 after 3 days of inhibitor soaking (Supplementary Fig. S3g-i). This suggests that the linear N5-acylated adducts are free to rotate about this single bond in a relaxed conformation.
a Crystal structure of LSD1-CoREST bound to AW4 (32) where the TTAS loop is highlighted in red. b Schematic (top) and the corresponding electron density (bottom) of the AW4-FAD adducts formed after 10 min of LSD1-CoREST crystal soaking with AW4 followed by 3 d of back washing. c Overlay of the AW4-FAD adduct (33) (gold) with the T-448-, 12-, and 15-FAD adducts in gray, blue, and green, respectively. d Superposition of the cyclic C4a (gold) and linear acyl AW4-FAD (pink) adducts. e Co-crystal structure of LSD1-CoREST and AW4 (32), highlighting the TTAS loop in salmon and the αD helix in cyan. The AW4-FAD adduct (gold) is overlaid with the T-448-FAD (gray) and GSK-LSD1-FAD adducts (green) (PDB: 2uxx). Residues investigated in the mutagenesis study are shown in sticks (blue) and are labelled. f Table summarizing the % N-formyl-FAD detected by LC-MS after treating LSD1-CoREST mutants for 24 h with AW4 (32). Data in (a–e) are provided in Supplementary Data 3 and 4. Data in (f) are representative of two independent experiments and are provided in Supplementary Data 1. Source data are provided as a Source Data file for (f).
Without structural resolution of the ΔTTAS mutant, we examined the residues in proximity to the TTAS loop in the wild-type LSD1-CoREST structure in more detail. Notably, T685 and R688, are predicted to participate in an H-bonding network with residues in the αD helix of the amine oxidase domain, R526 and D530 (Fig. 5e)31. Critically, the αD helix contacts both the active site and inhibitor adduct. We thus considered if disrupting this H-bonding network with point mutations could recapitulate the effects of the LSD1 TTAS loop deletion and mediate AW4-FAD (33) fragmentation to N-formyl-FAD. Single point mutations of T685, R688, and D530 to alanine were not sufficient to induce AW4-FAD transformation to formylated FAD (Fig. 5f). However, further disruption of the αD helix by introducing double (T685A, R688A) and triple alanine mutants (D530A, T685A, R688A) led to the formation of N-formyl-FAD at nearly comparable levels as LSD1 ΔTTAS upon AW4 treatment (Fig. 5f). With the additional biphenyl aryl substituent, the AW4-FAD adduct projects out of the active site and extends towards the αD helix compared to the more compact inhibitor-FAD adduct observed with GSK-LSD1, potentially explaining why ΔTTAS selectively enhances AW4-FAD adduct fragmentation and permits GFI1B binding (Fig. 5e). Notably, the initial T-448-FAD adduct is already orientated in closer proximity to the αD helix, potentially accounting for its ability to fragment in the absence of the TTAS loop deletion (Fig. 5e, Supplementary Fig. S5b). Altogether, these results demonstrate that the αD helix mediates crosstalk between the TTAS loop and active site and suggest that ΔTTAS may reposition the αD helix to mediate AW4-FAD fragmentation.
Discussion
Chromatin modifiers often have pleotropic functions spanning both enzymatic and nonenzymatic functions, and pharmacologically modulating these activities selectively can be challenging32,33,34. These difficulties are exemplified by efforts to target the LSD1 active site, which mediates both histone demethylation and TF binding35. Critically, disruption of the latter can drive on-target, dose-limiting hematological toxicities3. Remarkably, T-448 was recently identified as a demethylase-specific LSD1 inhibitor that preserves LSD1-GFI1B integrity. However, the mechanism underlying its unique chemical fragmentation has remained obscured. Here we shed light on the mechanism of action of T-448, showing that its initial covalent adduct with FAD undergoes a Grob fragmentation to form N-formyl-FAD. These insights provide a deeper understanding of how T-448 and related inhibitors selectively perturb LSD1 demethylase activity, a feature of their pharmacology which is critical for their therapeutic index and effective use as potential treatments for neurodevelopmental disorders.
By profiling the SAR of T-448 analogs, we show that meta substitution of the aryl amide of T-448 is necessary for the chemical fragmentation relative to other tranylcypromine derivatives. While the 1,2,4-thiadiazole of T-448 greatly accelerates fragmentation, its presence is not uniquely required. By comparing the X-ray structures of LSD1 engaged with these T-448 analogs, we reveal that the initial cyclic inhibitor-FAD adduct generated by T-448 and benzamide 15 — both of which are more efficient at fragmentation — are distinct from those generated by the remaining T-448 analogs. Overall, we demonstrate that the amide substituent on the meta position of the aryl ring and the unique staggered conformation of the C4a-N5 inhibitor-FAD adduct appear critical for the Grob fragmentation to proceed with the wild-type enzyme. Moreover, our data with 15 suggests that formation of even a small amount of N-formyl-FAD is sufficient to maintain enough intact LSD1-GFI1B complex in cells to sustain proliferation.
These findings on the mechanism of T-448 analogs have broader ramifications for other LSD1 covalent inhibitors being developed for oncology. Specifically, we demonstrate that a distal site mutation, ΔTTAS, can promote the selective fragmentation of an inhibitor-FAD adduct to yield N-formyl-FAD and confer drug resistance in AML cells despite residing >18 Å from the FAD cofactor. Even though ΔTTAS only leads to a small amount of N-formyl-FAD adduct, this may maintain sufficient LSD1-GFI1B levels to sustain proliferation, consistent with our observations with benzamide 15. The effects of ΔTTAS might be mediated by contacts with the αD helix of the amine oxidase domain, as our structures demonstrate that residues in the TTAS loop H-bond with residues in this α-helix. Double and triple point mutations disrupting this H-bond network promote inhibitor-FAD fragmentation, consistent with the αD helix mediating crosstalk between the TTAS loop and active site. Interestingly, the TTAS loop lies within close spatial proximity (~20 Å) to a previously characterized neuron-specific splicing variant of LSD1, LSD1 + 8a, which contains four additional amino acid residues36. This LSD1 isoform is essential for development and differentiation in the brain and phosphorylation of the insertion regulates LSD1 complex stability36,37,38. Therefore, small changes in this distal region of the amine oxidase domain, like with the TTAS loop mutation or LSD1 + 8a variant, can have significant and functional implications on LSD1 activity, highlighting these regions as potential sites of allosteric control. Regardless of whether these specific resistance mutations might arise in the clinic, they are broadly informative about drug-target pharmacology and reveal thematic possibilities for drug resistance mechanisms39,40.
In summary, we clarify the mechanism of action of T-448 and its broader consequences on the requirements of LSD1-GFI1B in AML cells. We elucidate the structural features of both the inhibitor as well as the protein in promoting a Grob fragmentation in the context of T-448 and LSD1 ΔTTAS, respectively. We expect that these insights will guide future efforts targeting LSD1 for therapeutics discovery and will more broadly inform on thematic mechanisms of drug resistance through covalent-drug adduct fragmentation.
Methods
Reagents
Compounds were stored at –80 °C in 100% DMSO and working aliquots were stored at –20 °C. The 100 mM concentration stocks for soaking experiments were in 100% DMSO while the diluted stocks at 4 mM were in 50% DMSO. The vehicle condition represents 0.1% (v/v) DMSO treatment. GSK-LSD1 was purchased from Sigma-Aldrich (≥98% by HPLC).
Chemical synthesis
LSD1 inhibitors were synthesized as described previously14,18. Please see the Supplementary Information 1 for more details on synthetic procedures.
Cell culture and lentiviral transduction
SET-2 was a gift from Matthew D. Shair; HEK293T was a gift from Bradley E. Bernstein. All cell lines were authenticated by Short Tandem Repeat profiling (Genetica) and routinely tested for mycoplasma (Sigma-Aldrich). All media were supplemented with 100 U/mL penicillin and 100 µg/mL streptomycin (Life Technologies) and fetal bovine serum (FBS, Peak Serum). All cell lines were cultured in a humidified 5% CO2 incubator at 37 °C. SET-2 were cultured in RPMI-1640, 20% FBS. HEK293T were cultured in DMEM (Life Technologies), 10% FBS. Lentivirus was produced by co-transfecting HEK293T cells with pCMV-VSV-G (Addgene plasmid #8454), psPAX2 (Addgene plasmid #12260) and transfer vector plasmid (encoding gene of interest) using Lipofectamine 3000 (Invitrogen L3000) according to the manufacturer’s protocol. Media was exchanged after 6 h and the viral supernatant was collected 48–72 h after transfection and filtered (0.45 µm). Transduction of SET-2 cells was carried out by mixing the lentivirus with cells and 8 µg/ml polybrene followed by centrifugation at 1800 x g for 90 min at 37 °C. After 48 h post-transduction, media was changed and puromycin (Thermo Fisher Scientific) selection was carried out for 5 d at 2 µg/mL.
Immunoblotting
For whole cell extracts, cells were lysed on ice using radioimmunoprecipitation assay (RIPA) buffer (Boston BioProducts) supplemented with fresh HALT™ protease inhibitor cocktail (Thermo Fisher Scientific). The lysate was clarified by centrifugation at 10,000 x g for 10 min. Protein concentration of the lysates was measured using BCA Protein Assay Kit (Thermo Fisher Scientific). Immunoblotting was performed according to standard procedures. The primary antibodies used are as follows: GFI-1B (B-7) X (Santa Cruz Biotechnology sc-28356X, LOT# B2120) for immunopercipitation, Monoclonal Anti-FLAG M2 (Sigma-Aldrich F1804, LOT#0000308215) at 1:1000 in 5% milk TBST, and GFI1B (Santa Cruz Biotechnology sc-28356, E0418) at 1:1000 in 5% milk TBST.
Co-immunoprecipitation
For Co-IPs performed in HEK293T cells, cells were plated at 40-60% confluency and treated with 500 nM of the respective LSD1 inhibitor or vehicle. 24 h after plating, cells were co-transfected with 1.5 µg pcDNA.3 FLAG-LSD1 plasmid and 2 µg pcDNA.3 wild-type GFI1B-FLAG using Lipofectamine 3000 reagent (Invitrogen L3000). 48 h post-transfection, cells were washed with PBS and then flash frozen. Cells were thawed, lysed on ice in N450 buffer (50 mM Tris-HCl pH 7.5, 450 mM NaCl, 1 mM MgCl2, 1% NP-40 alternative, 5% glycerol) supplemented with 1:10,000 benzonase (Sigma-Aldrich), and the lysates were cleared. The protein concentration was quantified as above and diluted to 1 mg/mL in lysis buffer. Supernatants were immunoprecipitated overnight at 4 °C with 4 µg GFI1B antibody and 40 µL Protein G Dynabeads (ThermoFisher Scientific). Beads were washed twice with lysis buffer, eluted in SDS-PAGE loading buffer, and carried forward to immunoblotting as described above.
Generation of clonal drug-resistant LSD1 mutant cell lines
sgRNA sequence targeting LSD1, sg684 (GGCAGTACGACTGCCAGCAG), was cloned into pLentiCRISPR-puro.v2. SET-2 were transduced with the resultant plasmids as indicated above and subsequently treated with 50 nM AW4 for 6 weeks to enrich drug-resistant mutant cells. Surviving cells were FACS-sorted as single cells and then expanded. For library preparation, genomic PCR primers (FWD-CTCATGTGCTTCTTTCTTATGGT, REV-GATTAAAATTAGAAGGCA) with Illumina adapter sequences were used to amplify specified regions of LSD1 as previously described41. Samples were sequenced on a MiSeq genome analyzer (Illumina). The sequencing reads were analyzed using CRISPResso2 (v.2.0.40).
Genotype determination
Genomic DNA was isolated using QIAamp Blood & Cell Culture DNA Mini Kit (Qiagen) or Lucigen QuickExtract (Lucigen Corporation). For amplicon sequencing, genomic DNA was subjected to two rounds of PCR reactions. PCR1 was conducted using 50 ng of genomic DNA with Q5 HotStart Polymerase (NEB) to amplify the target region. PCR1 products were ran on an agarose gel and gel purified using Zymo Gel Extraction Kit (Zymo). 1 µL of the purified DNA was used as input for PCR2. PCR2 was conducted with PCR primers containing Illumina adaptor sequences. PCR2 products were ran on an agarose gel, gel purified and pooled. The final libraries were quantified using Qubit High Sensitivity Assay Kit (Thermo Fisher Scientific) and sequenced on a MiSeq genome analyzer (Illumina). The sequencing reads were analyzed using CRISPResso2 (v.2.0.40)42.
Cell growth assays
SET-2 were seeded in 96-well plates with 2,000 cells per well in triplicate with drug or vehicle treatments. Cell viability was monitored at day 7 and 10 by measuring end point luminescence using CellTiter-Glo (Promega) on a SpectraMax i3x plate reader. ATP standard curve was prepared using known concentrations of ATP and used to calculate the ATP content of cells. Dose-response curves were determined through interpolation using GraphPad Prism v.10.2.2 nonlinear regression fit ([inhibitor] vs normalized response -- variable slope). All growth assays were performed at least twice and in biological triplicates.
Expression and purification of LSD1(Δ150) from E.coli
The pET15b vector encoding for His6-tagged human LSD1(Δ150, residues 151-852) was a gift from P. A. Cole. Proteins (wild-type and mutants) were expressed in BL21-CodonPlus (DE3)-RIPL competent E. coli and purified at 4 °C according to a literature procedure13. Briefly, cells were disrupted by sonication (10 s on, 20 s off, 60% Amp) after resuspending in lysis buffer (50 mM HEPES pH 7.5, 100 mM KCl, 0.5 mM TCEP) containing a protease inhibitor cocktail (Roche). The lysate was centrifuged at 20,000 x g for 40 min and loaded onto a His60 nickel resin (Takara Bio). After 1 h incubation, the resin was washed with 10 column volumes (CV) of lysis buffer, followed by 10 CV of wash buffer 1 (lysis buffer with 20 mM imidazole), and finally eluted by the addition of 5–10 CV of lysis buffer containing 250 mM imidazole. Combined fractions were dialyzed overnight in 2 L of dialysis buffer (20 mM Tris pH 7.5, 100 mM NaCl, 5 mM β-mercaptoethanol, 2 changes) and concentrated (Amicon Ultra-15 Centrifugal Filter Unit, 30 kDa MWCO) to 5–10 mL. The protein was further purified by using a 5 mL Q Sepharose HP column (Cytiva) with a gradient elution from buffer A (20 mM Tris pH 7.5, 1 mM TCEP) to buffer B (20 mM Tris pH 7.5, 1 M NaCl, 1 mM TCEP). After concentration, the protein was purified on a Superdex 200 Increase 10/300 GL column (Cytiva) in storage buffer (50 mM HEPES pH 7.5, 150 mM NaCl, 0.5 mM TCEP, 5% glycerol). Fractions with >95% purity (coomassie-stained SDS PAGE) were pooled and stored at –80 °C.
Expression and purification of LSD1(Δ150)-CoREST(Δ307) from E.coli
LSD1 (Δ150, residues 151-852)-CoREST (residues 308-485) complexes were purified by tandem affinity chromatography using the LSD1 construct described above while human CoREST (residues 308-485) was expressed from a pGEX vector13. After transformation, BL21-CodonPlus (DE3)-RIPL E.coli cells were grown and a single colony was cultivated in LB media with 100 mg/L ampicillin at 37 °C. Expression was induced at OD600 = 0.8 by adding 0.25 mM isopropyl β-D-thiogalactoside (IPTG). Cells were grown overnight at 17 °C, pelleted at 4000 x g for 30 min, and stored at –80 °C. Pellets were resuspended in lysis buffer (50 mM NaH2PO4 pH 8.0, 300 mM NaCl, 5% glycerol, 7.5 mM imidazole with PMSF, DNAse and EDTA-free Roche protease inhibitor cocktail) in a weight ratio of 1:1.5, respectively. Cells were sonicated, centrifuged, passed through nickel affinity resin, and finally loaded onto GST resin equilibrated in GST affinity buffer (50 mM NaH2PO4 pH 8.0, 300 mM NaCl, 5% glycerol, 1 mM DTT, 1 mM EDTA). The GST-tag was cleaved with GST-PreScission protease (APEXBIO) overnight at 4 °C on the resin. The column was washed with GST affinity buffer and the eluted protein was concentrated and gel-filtered in storage buffer on a Superdex 200 10/300 GL column. Purity was checked by SDS-PAGE and fractions with 90–95% purity were pooled and stored at –80 °C.
LSD1 enzyme assays
LSD1 activities and steady-state kinetics were measured with the Amplex Red Hydrogen Peroxide/Peroxidase Assay (Sigma-Aldrich) using recombinant LSD1 and a synthetic peptide comprising the first 21 amino acids of H3K4me2 (Genscript)26. The final reaction mixture contained 50 mM HEPES pH 8.5, 0.1 mM Amplex Red, 0.3 mM horseradish peroxidase, and 0.3 µM LSD1-CoREST. Each construct was kinetically characterized by testing the enzymatic activities at 8 serially diluted concentrations (from 40 µM to 0.31 µM) of the H3K4me2 (21aa) peptide substrate. For inhibition studies, LSD1 (75 ng/well) and inhibitors at the appropriate concentration were incubated at room temperature for 10 min in reaction buffer with 0.01% BRIJ35 (ThermoFisher Scientific) prior to the addition of the peptide substrate. The residual Kcat,app of inhibited LSD1-CoREST complexes were evaluated by using 4.5 µM of protein with 40 µM H3K4me2 (21aa) peptide. GraphPad Prism v.7 was used to determine KM, Kcat and IC50 values (log(inhibitor) vs. response—variable response) by nonlinear regression analysis.
FAD-adduct detection by LC/Q-TOF MS
Analysis of FAD-adducts after LSD1 protein treatment with inhibitor was adopted from Matsuda et al. 18. Briefly, 80 µM of LSD1 or 50 µM of LSD1-CoREST were treated with 640 µM or 400 µM inhibitor, respectively, at room temperature for the indicated time points along with a vehicle control in 50 mM Tris-HCl (pH 8.0) and then subjected to 8 M urea to a final concentration of 6.4 M for 1 h at 4 °C to denature the sample. The inhibitor-FAD adducts were then extracted at 4 °C with acetonitrile (ACN) for 15 min. The precipitates were removed by centrifugation at 10,000 x g for 10 min. The supernatant was supplemented with 500 mM ammonium acetate to a final concentration of 5 mM and filtered before sample injection. Samples were then injected onto a Phenomenex Kinetix C18 column (2.1 mm, 2.6 micron particle size, 150 mm length) with Orbitrap (Thermo q-Exactive Plus) coupled to an Ultimate 3000 HPLC. Water with 0.1% formic acid and ACN were used as mobile phases A and B, respectively. The mobile phase composition was changed to 100% B over 20 minutes. The experiments were performed at least twice.
Molecular mass determination of FAD inhibition products
The inhibition products were analyzed by MS with 100 µM LSD1-CoREST complex in 25 mM KH2PO4, pH 7.2 and 5% glycerol. Inhibitors were added for a final concentration of 400 µM and the incubation was at 20 °C for 24 h. Inhibited LSD1-CoREST was unfolded by adding a proper volume of 8 M urea solution to have a final concentration of 6.4 M urea. After 1 h on ice, ACN (final concentration 30%) was added to precipitate protein debris. After centrifugation, supernatant was stored at –20 °C until spectrometric analysis. The sample solution was diluted 1:1 with 5 mM ammonium acetate and was injected onto a high resolution QTOF mass spectrometer UHPLC-HRMS/MS- AB Sciex X500B. Chromatographic separation was achieved using a C18 Zorbax extend analytical column (2.1 × 50 mm, 1.8 μm, Agilent). The mobile phase consisted of (A) 5 mM ammonium acetate in water and (B) methanol. The compositions of gradient elution (the mixture of (A) and (B) were as follows designated in percentage of (B), v/v): 5–10% at 0–1 min, 10–80% at 1–10 min, 80% at 10–15 min, 5% at 15–25 min. The flow rate was set at 0.3 mL/min. The mass spectrometer was operated in the negative ionization mode. SCIEX OS software v.3.0.0 was used for data acquisition and processing. The percentage of products represent the percentage of total area of all extracted ion current peaks using GraphPad Prism v.9.3.1. FAD-adducts were usually distributed in different peaks comprising [M-H]-, [M-H-H2O]- (dehydrated adduct), [M-2H+Na]-and [M-H + CH3]- (methylester). The experiments were performed at least twice.
Styrene detection assay from LSD1-CoREST complexes
For wild-type LSD1, 80 µM of LSD1 was treated with 640 µM inhibitor or vehicle control at room temperature for the indicated time points in 50 mM Tris-HCl (pH 8.0). Styrene was extracted with 200 µL dichloromethane (DCM) with 100 µM caffeine as an internal standard. The DCM was subsequently removed, and the extract was resuspended in 200 µL of ACN and injected onto a Phenomenex Kinetix C18 column (2.1 mm, 2.6-micron particle size, 150 mm length) with Agilent 6530 Q-TOF Mass Spectrometer. Water with 0.1% formic acid and ACN were used as mobile phases A and B, respectively. The mobile phase composition was changed to 100% B over 20 minutes. The experiments were performed at least twice, and the styrene formation kinetics was done in biological triplicates.
For LSD1-CoREST wild-type and mutant complexes, 50 µM of LSD1 was treated with 400 µM inhibitor (AW4) or vehicle control and 100 µM GFI1B peptide (resides 2-10) at room temperature for the indicated time points in 50 mM Tris-HCl, pH 8.0. Styrene was extracted with 200 µL DCM with 100 µM tridecane as an internal standard. The DCM was subsequently removed, and the extract was resuspended in 200 µL of ACN and analyzed with Waters Quattro micro GC/MS/MS. The experiments were performed at least twice.
Styrene detection assay from cell media
5 million SET-2 cells were resuspended in 1 mL of R-20 and treated with 1 µM of inhibitor or vehicle control for 24 h. The cells were spun down and the media was filtered through a 0.22 µm filter and then extracted 2x with 300 µL DCM. The DCM was subsequently removed, and the extract was resuspended in 100 µL of DCM and analyzed by LC-MS. Samples were injected onto a Phenomenex Kinetix C18 column (2.1 mm, 2.6 micron particle size, 150 mm length) with Orbitrap (Thermo q-Exactive Plus) coupled to an Ultimate 3000 HPLC. Water with 0.1% formic acid and ACN were used as mobile phases A and B, respectively. The mobile phase composition was changed to 100% B over 20 min. The flow rate was kept constant at 0.200 L/min, and a 5 μL volume was injected. AW4 treated samples were injected onto a Waters Quattro micro GC/MS/MS. The experiments were repeated at least twice and in duplicate.
Absorbance spectroscopy
Purified LSD1-CoREST (10 µM) in 25 mM KH2PO4 pH 7.2 and 5% (w/v) glycerol was incubated with 400 µM inhibitor for variable times. Absorbance spectra were measured using a Cary-100 spectrophotometer (Varian). The experiments were performed at least twice.
ThermoFAD assay
ThermoFAD was performed in quintets as previously described with 4 µM of LSD1 in 20 µL of storage buffer43. Thermal curves were obtained using a qRT-PCR detection system (BioRad C1000 Touch Thermal Cycler) with a temperature gradient from 20 to 95 °C performing a fluorescence measurement every 0.5 °C with a 10 s delay for signal stabilization. The Tm was determined by fitting the thermal curves to a Boltzmann sigmoid and calculating the inflection point (GraphPad Prism v.9.3.1).
Fluorescence polarization assay
Binding assays were performed in two independent experiments with n = 2 or 3 biological replicates where specified. The change in fluorescence polarization of fluorescently labelled GFI1B peptide (residues 2-10; labeled at the C-terminus with 5(6)-carboxytetramethylrhodamine) upon binding to LSD1-CoREST was monitored using previously described protocols14,23,44. Briefly, LSD1-CoREST (4 µM) complex was incubated with inhibitor (10 µM) or vehicle control for 10 min on ice and then with the labelled peptides for the indicated time points (2 nM). After incubation, the samples were prepared by a 2-fold serial dilution in the assay buffer (15 mM KH2PO4, pH 7.2, 5% glycerol, and 1 mg/mL BSA) containing 2 nM labeled peptide and the plate was equilibrated to room temperature before data acquisition. Fluorescence polarization was measured using a microplate reader in 384-well black microplates at 25 °C. The G-factor on the microplate reader was adjusted to 35 mP for the reference well containing labelled peptide. The binding curves were fit by nonlinear regression analysis in GraphPad Prism v.9.3.1 with a constant fluorophore concentration, yielding the dissociation constant (Kd) according to the following equation:
where the total recoded signal (Y) depends on that of bound (Ab) and free (Af) fluorescent ligand as a function of total protein concentration (X) and total ligand concentration (c, 2 nM peptide)44.
Crystals soaking and structure determination
Purified LSD1-CoREST complex was crystallized following the previously published conditions with 1.1-1.3 M sodium/potassium tartrate and 100 mM (N-(2-acetamido)-2-iminodiacetic acid), pH 6.545. Crystals were transferred in reservoir drops supplemented with 1 mM inhibitor and 4% DMSO. Once FAD is bleached, crystals were flash cooled for data collection. For a second set of experiments, aimed at investigating changes in FAD-adducts over time, crystals were back washed in a clear reservoir drop and left for 3 days at 20 °C. The third set of experiments were devoted to structurally visualize peptide conformation in presence of different FAD-adducts. Inhibited crystals were soaked in a drop containing 1 mM of SNAG(2-10) or H3(1-21) peptide before flash freezing.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
X-ray crystallography structures have been deposited in the PDB database under the following accession codes: 8F6S, 8FQJ, 8FJ4, 8FSK, 8UL8, 8GJ6, 8FRV, 8FRI, 8FRQ, 9ELA, 8UMQ, 8ULC, 8UL6, 8ULB, 8BOP, 8FDV, 8UOM, 8UNI, 9EL8, and 9EL7. All other data supporting the findings of this study are available within the article and its Supplementary information files, and from the corresponding author(s). Source data are provided with this paper.
References
Zhao, S., Allis, C. D. & Wang, G. G. The language of chromatin modification in human cancers. Nat. Rev. Cancer 21, 413–430 (2021).
Nair, S. S. & Kumar, R. Chromatin remodeling in cancer: a gateway to regulate gene transcription. Mol. Oncol. 6, 611–619 (2012).
Fang, Y., Liao, G. & Yu, B. LSD1/KDM1A inhibitors in clinical trials: advances and prospects. J. Hematol. Oncol. 12, 129 (2019).
Liu, H. et al. LSD1 in drug discovery: From biological function to clinical application. Med. Res. Rev. 44, 833–866 (2024).
Maiques-Diaz, A. et al. Enhancer activation by pharmacologic displacement of LSD1 from GFI1 induces differentiation in acute myeloid leukemia. Cell Rep. 22, 3641–3659 (2018).
Maes, T. et al. ORY-1001, a potent and selective covalent KDM1A inhibitor, for the treatment of acute leukemia. Cancer Cell 33, 495–511.e12. (2018).
Ishikawa, Y. et al. A novel LSD1 inhibitor T-3775440 disrupts GFI1B-containing complex leading to transdifferentiation and impaired growth of AML cells. Mol. Cancer Ther. 16, 273–284 (2017).
Takagi, S. et al. LSD1 inhibitor T-3775440 Inhibits SCLC cell proliferation by disrupting LSD1 interactions with SNAG domain proteins INSM1 and GFI1B. Cancer Res 77, 4652–4662 (2017).
Yamamoto, R. et al. Selective dissociation between LSD1 and GFI1B by a LSD1 inhibitor NCD38 induces the activation of ERG super-enhancer in erythroleukemia cells. Oncotarget 9, 21007–21021 (2018).
Cusan, M. et al. LSD1 inhibition exerts its antileukemic effect by recommissioning PU.1- and C/EBPα-dependent enhancers in AML. Blood 131, 1730–1742 (2018).
Harris, W. J. et al. The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 21, 473–487 (2012).
McGrath, J. P. et al. Pharmacological inhibition of the histone lysine demethylase KDM1A suppresses the growth of multiple acute myeloid leukemia subtypes. Cancer Res. 76, 1975–1988 (2016).
Waterbury, A. L. et al. An autoinhibitory switch of the LSD1 disordered region controls enhancer silencing. Mol. Cell 84, 2238–2254.e11 (2024).
Vinyard, M. E. et al. CRISPR-suppressor scanning reveals a nonenzymatic role of LSD1 in AML. Nat. Chem. Biol. 15, 529–539 (2019).
Saleque, S., Kim, J., Rooke, H. M. & Orkin, S. H. Epigenetic regulation of hematopoietic differentiation by Gfi-1 and Gfi-1b is mediated by the cofactors CoREST and LSD1. Mol. Cell 27, 562–572 (2007).
Lin, Y. et al. The SNAG domain of Snail1 functions as a molecular hook for recruiting lysine‐specific demethylase 1. Embo J. 29, 1803–1816 (2010).
Sprüssel, A. et al. Lysine-specific demethylase 1 restricts hematopoietic progenitor proliferation and is essential for terminal differentiation. Leukemia 26, 2039–2051 (2012).
Matsuda, S. et al. T-448, a specific inhibitor of LSD1 enzyme activity, improves learning function without causing thrombocytopenia in mice. Neuropsychopharmacol 44, 1505–1512 (2019).
Baba, R. et al. LSD1 enzyme inhibitor TAK-418 unlocks aberrant epigenetic machinery and improves autism symptoms in neurodevelopmental disorder models. Sci. Adv. 7, eaba1187 (2021).
Yin, W. et al. Safety, pharmacokinetics and pharmacodynamics of TAK‐418, a novel inhibitor of the epigenetic modulator lysine‐specific demethylase 1A. Br. J. Clin. Pharmacol. 87, 4756–4768 (2021).
Mohammad, H. P. et al. A DNA hypomethylation signature predicts antitumor activity of LSD1 inhibitors in SCLC. Cancer Cell 28, 57–69 (2015).
Mimasu, S., Sengoku, T., Fukuzawa, S., Umehara, T. & Yokoyama, S. Crystal structure of histone demethylase LSD1 and tranylcypromine at 2.25Å. Biochem. Bioph. Res. Co. 366, 15–22 (2008).
Pilotto, S. et al. LSD1/KDM1A mutations associated to a newly described form of intellectual disability impair demethylase activity and binding to transcription factors. Hum. Mol. Genet. 25, 2578–2587 (2016).
Prantz, K. & Mulzer, J. Synthetic applications of the carbonyl generating grob fragmentation. Chem. Rev. 110, 3741–3766 (2010).
Zhang, Z. et al. Structural and stereoelectronic insights into oxygenase-catalyzed formation of ethylene from 2-oxoglutarate. Proc. Natl Acad. Sci. USA 114, 4667–4672 (2017).
Yang, M. et al. Structural basis for the inhibition of the LSD1 histone demethylase by the antidepressant trans-2-phenylcyclopropylamine †, ‡. Biochemistry-us 46, 8058–8065 (2007).
Mimasu, S. et al. Structurally designed trans-2-phenylcyclopropylamine derivatives potently inhibit histone demethylase LSD1/KDM1. Biochem. US 49, 6494–6503 (2010).
Binda, C. et al. Biochemical, structural, and biological evaluation of tranylcypromine derivatives as inhibitors of histone demethylases LSD1 and LSD2. J. Am. Chem. Soc. 132, 6827–6833 (2010).
Hattori, Y. et al. Design, synthesis, and structure–activity relationship of TAK-418 and its derivatives as a novel series of LSD1 inhibitors with lowered risk of hematological side effects. Eur. J. Med. Chem. 239, 114522 (2022).
Rudolf, J. D. & Chang, C.-Y. Terpene synthases in disguise: enzymology, structure, and opportunities of non-canonical terpene synthases. Nat. Prod. Rep. 37, 425–463 (2019).
Stavropoulos, P., Blobel, G. & Hoelz, A. Crystal structure and mechanism of human lysine-specific demethylase-1. Nat. Struct. Mol. Biol. 13, 626–632 (2006).
Cheng, Y. et al. Targeting epigenetic regulators for cancer therapy: mechanisms and advances in clinical trials. Signal Transduct. Target. Ther. 4, 62 (2019).
Siklos, M. & Kubicek, S. Therapeutic targeting of chromatin: status and opportunities. FEBS J. 289, 1276–1301 (2022).
Ellis, L., Atadja, P. W. & Johnstone, R. W. Epigenetics in cancer: targeting chromatin modifications. Mol. Cancer Ther. 8, 1409–1420 (2009).
Baron, R., Binda, C., Tortorici, M., McCammon, J. A. & Mattevi, A. Molecular mimicry and ligand recognition in binding and catalysis by the histone demethylase LSD1-CoREST complex. Structure 19, 212–220 (2011).
Zibetti, C. et al. Alternative splicing of the histone demethylase LSD1/KDM1 contributes to the modulation of neurite morphogenesis in the mammalian nervous system. J. Neurosci. 30, 2521–2532 (2010).
Rusconi, F., Grillo, B., Toffolo, E., Mattevi, A. & Battaglioli, E. NeuroLSD1: Splicing-generated epigenetic enhancer of neuroplasticity. Trends Neurosci. 40, 28–38 (2017).
Toffolo, E. et al. Phosphorylation of neuronal lysine‐specific demethylase 1LSD1/KDM1A impairs transcriptional repression by regulating interaction with CoREST and histone deacetylases HDAC1/2. J. Neurochem. 128, 603–616 (2014).
Pisa, R. & Kapoor, T. M. Chemical strategies to overcome resistance against targeted anticancer therapeutics. Nat. Chem. Biol. 16, 817–825 (2020).
Pisa, R., Cupido, T., Steinman, J. B., Jones, N. H. & Kapoor, T. M. Analyzing resistance to design selective chemical inhibitors for AAA proteins. Cell Chem. Biol. 26, 1263–1273.e5 (2019).
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843 (2013).
Pinello, L. et al. Analyzing CRISPR genome-editing experiments with CRISPResso. Nat. Biotechnol. 34, 695–697 (2016).
Forneris, F., Orru, R., Bonivento, D., Chiarelli, L. R. & Mattevi, A. ThermoFAD, a Thermofluor®‐adapted flavin ad hoc detection system for protein folding and ligand binding. Febs J. 276, 2833–2840 (2009).
Pilotto, S. et al. Interplay among nucleosomal DNA, histone tails, and corepressor CoREST underlies LSD1-mediated H3 demethylation. Proc. Natl Acad. Sci. 112, 2752–2757 (2015).
Forneris, F., Binda, C., Adamo, A., Battaglioli, E. & Mattevi, A. Structural basis of LSD1-CoREST selectivity in histone H3 recognition*. J. Biol. Chem. 282, 20070–20074 (2007).
Acknowledgements
This work was supported by award no. DP2GM137494 and R01CA274437 from the National Institutes of Health (B.B.L) and from the Italian Ministry for University (MUR) project FISR2019_00374 MeDyCa and the Associazione Italiana per la Ricerca sul Cancro (Italian Association for Cancer Research) IG19808 (A.M.). We would like to thank members of the Liau lab, in particular P. Gosavi and H. Kwok, for helpful advice regarding biochemical and cellular assays. We would like to thank the Balskus lab for their assistance with mass spectrometry. We would like to thank Z. Nizioleck and C. Maesner at the Bauer Core Facility at Harvard University for their assistance with single-cell sorting, and S. Trauger and J. Wang at the Harvard Center for Mass Spectrometry for their assistance with mass spectrometry. The authors thank Dr. Weston Umstead’s team at Chiral Technologies, Inc. for providing the chiral method development and separation services.
Author information
Authors and Affiliations
Contributions
A.L.W., J.C., A.M., and B.B.L. conceived the study, with input from O.Z., P.R.T. and M.B.; A.L.W. and O.Z. designed and performed molecular and cell experiments. A.L.W., J.C., O.Z., and S.M.H. designed and performed protein purification and biochemical assays. O.Z., J.C., P.R.T., and A.L.W. performed and analyzed mass spectrometry experiments. J.C. and M.B. performed X-ray crystallography experiments. P.R.T., C.L., J.L., J.S.P., and A.F. designed and synthesized LSD1 inhibitors. A.LW., J.C., A.M., and B.B.L. wrote and edited the manuscript. B.B.L. and A.M. held overall responsibility for the study.
Corresponding authors
Ethics declarations
Competing interests
B.B.L. is a shareholder and member of the scientific advisory board of Light Horse Therapeutics, and receives research support from Ono Pharmaceuticals. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Waterbury, A.L., Caroli, J., Zhang, O. et al. Covalent adduct Grob fragmentation underlies LSD1 demethylase-specific inhibitor mechanism of action and resistance. Nat Commun 16, 3156 (2025). https://doi.org/10.1038/s41467-025-57477-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57477-3