Abstract
Cell-free gene expression (CFE) systems enable transcription and translation using crude cellular extracts, offering a versatile platform for synthetic biology by eliminating the need to maintain living cells. However, Such systems are constrained by cumbersome composition, high costs, and limited yields due to numerous additional components required to maintain biocatalytic efficiency. Here, we introduce DropAI, a droplet-based, AI-driven screening strategy designed to optimize CFE systems with high throughput and economic efficiency. DropAI employs microfluidics to generate picoliter reactors and utilizes a fluorescent color-coding system to address and screen massive chemical combinations. The in-droplet screening is complemented by in silico optimization, where experimental results train a machine-learning model to estimate the contribution of the components and predict high-yield combinations. By applying DropAI, we significantly simplified the composition of an Escherichia coli-based CFE system, achieving a fourfold reduction in the unit cost of expressed superfolder green fluorescent protein (sfGFP). This optimized formulation was further validated across 12 different proteins. Notably, the established E. coli model is successfully adapted to a Bacillus subtilis-based system through transfer learning, leading to doubled yield through prediction. Beyond CFE, DropAI offers a high-throughput and scalable solution for combinatorial screening and optimization of biochemical systems.
Similar content being viewed by others
Introduction
Cell-free gene expression (CFE) involves the in vitro activation of transcription and translation using crude cellular extracts instead of intact cells1. Compared to in vivo gene expression, CFE offers a more flexible, sustainable, and rapid approach to gene expression. First, by eliminating the necessity of sustaining cellular life, it allows researchers to directly manipulate the molecular environment of the system. This includes the addition of non-native substrates, purified proteins or RNAs, and recombinant DNA templates. Furthermore, CFE circumvents mechanisms essential for cell survival, bypasses limitations on molecular transport across the cell wall, and enables a focused investigation on specific genetic networks or the biosynthesis of a single product. Moreover, CFE can accelerate design-build-test cycles in biotechnological applications by significantly reducing the timeline of cloning steps and gene expression1. Given these advantages, there has been a resurgence of scientific interest in cell-free biotechnology, leading to the development of novel CFE platforms for various compelling applications1,2, such as industrially important chemical production3,4,5,6, metabolic pathway prototyping7, natural product biosynthesis5, clinical therapeutics8,9, and diagnostics10.
However, standardized procedures for optimizing each newly developed CFE system still remain elusive, which are often dependent on empirical approaches. Moreover, current CFE systems are constrained by the complicated formulations and the challenge of balancing cost and yield1, which severely limits their utility in an expanded repertoire of applications. For example, a typical bacteria extract-based CFE system requires around 40 additional components, besides the crude cell extract, to maintain a reasonable level of biocatalytic efficiency. Over half of the cost of a standard bench-scale CFE formulation arises from the expensive energy substrates and multiple additives used to enhance biosynthesis yield1. While some of these additives, such as dNTPs and nucleic acids, are essential, many others can be optimized or minimized. Addressing these challenges calls for comprehensive screening of the CFE system to develop a streamlined formulation with cost-effective, sustainable energy sources and minimal essential additives at optimized concentrations. Such screening not only has the potential to reduce costs but also to provide new insights into CFE as an integrated biochemical system11.
High-throughput screening (HTS) has found widespread use in drug discovery12,13,14,15,16 and biochemical system optimization17. Still, the adoption of HTS in CFE optimization has been hindered by its complexity, compound consumption, and cost1. A typical CFE formulation includes non-native energy sources, gene templates, and various transcription-translation factors (e.g., coenzyme A, NAD, and exogenous tRNAs). Screening these additives can easily generate thousands of combinations. For example, considering five candidate energy source molecules (choose one out of five) and ten different transcription-translation factors (only consider their presence or absence), a primary screening process must evaluate 5120 combinations (5 × 210). Constructing a screening pool at this scale requires approximately 60,000 liquid handling steps and 54 × 96-well plates (without replication). Given the logistical complexity, conventional pipette-based liquid handling techniques are often inefficient or unfeasible. The screening scale can expand exponentially when introducing concentration gradients of the additives. Additionally, CFE formulations vary with the choice of chassis (e.g., Escherichia coli18, Streptomyces19, and yeast20). Consequently, changing the chassis extract necessitates repeating the entire screening process to optimize the CFE system. Given these challenges, adequately screening CFE systems with existing HTS methods is impractical.
Here, we report a droplet-based and AI-guided combinatorial screening strategy (DropAI) designed to optimize CFE systems with high throughput and economic efficiency. A key feature of DropAI is its ability to construct massive combinations in picoliter reactors (80 μm droplets, ~250 pL) and use a fluorescent color-based coding-decoding approach to trace the composition of each combination (Fig. 1a). This is achieved through microfluidics, where one carrier droplet merges with four satellite droplets to form a complete screening unit. The carrier droplet contains the CFE mixture, while each satellite droplet randomly samples a unique set of CFE components. These component sets are labeled with distinct fluorescent colors, and each component within a set is associated with a unique fluorescence intensity. Consequently, the merged droplets are encoded by fluorescent colors and intensities, termed FluoreCode, to identify the contained combinations. The FluoreCode is read in parallel using multi-channel droplet imaging. With our current optical settings, we can resolve nine intensity levels within one color, enabling the satellite droplets to encode 729 (93) combinations. If the carrier droplets are also color-coded, the theoretical combinatorial space expands to 6561 (94). In microfluidics, combinations are created at a rate of approximately 1,000,000/hour. For a screening scale of 500 combinations with 100 replicates each (totaling 50,000 droplets), it takes about three minutes to construct the entire pool (excluding a setup time of around 30 min), consuming approximately 12.5 μL of reagents.
DropAI employs a hybrid strategy of (a) in-droplet screening and (b) in silico optimization. a In-droplet screening: Massively parallel experimental data are generated using picoliter droplet reactors. Each droplet contains a unique combination of cell-free gene expression (CFE) components encoded by fluorescent markers. Multichannel droplet imaging allows for parallel reading of component combinations and resulting CFE yields (sfGFP fluorescence). b In silico optimization: Experimental data are used to develop AI models, which extend predictions beyond experimental scales to enhance screening capacity. Iterations of the screening process may be conducted for further optimization. c Final verification: The optimized screening results are validated in vitro.
After the in-droplet screening, DropAI proceeds with in silico optimization (Fig. 1b). Experimental results obtained from the droplet assays are used to train a machine learning model. The established model explores conditions beyond the experimental scale to predict high-yield combinations, which are then verified in vitro (Fig. 1c). In this study, we first applied DropAI to optimize an E. coli-based CFE system. Using in-droplet screening, we sampled combinations of 12 additives across transcription, translation, and ATP regeneration phases of CFE. We then built a machine learning model based on the experimental data to predict the contribution of each additive to CFE yield. Based on these predictions, we selected only three essential additives and optimized their concentrations. The final simplified and optimized CFE formula was tested by expressing superfolder green fluorescent protein (sfGFP), resulting in a 2.1-fold decrease in unit cost and a 1.9-fold increase in yield compared to the original formula. We further applied the optimized formula to express 12 different proteins ranging from 27 to 370 kDa, with 10 out of 12 proteins exhibiting maintained or increased yields. Furthermore, we demonstrated that the established E. coli model could be tuned and applied to optimize a Bacillus subtilis-based CFE system through transfer learning21. This approach significantly reduced the effort required to optimize CFE systems across different chassis. The transfer model achieved high prediction accuracy with a limited number of input experiments (e.g., 27 and 36 combinatorial sets). The predicted high-yield formulation was verified in vitro by expressing sfGFP (resulting in a two-fold increase in yield) and short antimicrobial peptides.
Results
Microfluidic construction of combinatorial droplet library
DropAI relies on high-throughput microfluidics to generate massive combinatorial libraries for in-droplet screening. To realize this concept, we developed a microfluidic device capable of generating arbitrary four-droplet combinations at approximately 300 Hz (Fig. 2a, Supplementary Fig. 1a, and Supplementary Movie 1). In this device, a pool of carrier droplets (~70 μm in diameter; Supplementary Figs. 1b, c) is loaded upstream in a microchannel. The carrier droplets flow downstream sequentially, each meeting three satellite droplets (~36 μm in diameter; Supplementary Figs. 1b, c) before reaching a micro-teeth structure designed for droplet merging. Figure 2b shows micrographs of three junctions where the satellite droplets, depicted in false colors, are being loaded and the micro-teeth structure where the grouped droplets are being merged. This process is achieved by synchronizing the frequency of droplets from different pools, resulting in an overall efficiency of around 90%. We confirmed the high reproducibility of these microfluidic operations through multiple independent trials (N = 3; Supplementary Fig. 2). Each droplet is encoded with a unique fluorescent color and intensity, creating a final merged droplet with a 4-digit nonary FluoreCode (Fig. 2c). To evaluate the coding capacity, we generated a library of 6,561 (94) combinations using fluorescent dye solutions (Fig. 2d). The droplets were imaged under multiple channels (Fig. 2e) to extract the fluorescence of each droplet (n = 206,634; Supplementary Fig. 1d). We confirmed that the intensity profile of each color exhibited nine major peaks (Supplementary Fig. 1e). We then processed the intensity data using a multi-band-pass filter program to eliminate low-quality marginal data points. The remaining data sets (n = 112,156) were binned by the FluoreCodes (see Methods). The binning generated 6,527 distinct clusters, recovering 99.5% of the theoretical combinatorial space. Figure 2f displays these clusters in a 2D space utilizing t-distributed stochastic neighbor embedding (t-SNE).
a DropAI generates massive combinations in droplet reactors with a multi-droplet microfluidic merger. The device loads a pool of carrier droplets into a microchannel at regular intervals. Each carrier droplet meets 3 satellite droplets at downstream junctions, which are all merged under an electrical field at the micro-teeth structure. b Micrographs depict the droplet pairing and merging process. The false coloring depicts carrier droplets (green) and differentiates the satellite droplets (yellow, pink, and blue for satellite drop 1, 2, and 3, respectively). c As every droplet is coded by fluorescent color and intensity, the merged droplets are rendered a 4-color barcode (FluoreCode) indicating the exact combinations. The microfluidic device generates the combinatorial pool at ~300 Hz, and the total coding capacity is 6561 (94). d A stacked micrograph displaying the as-built combinatorial pool. e For decoding, the merged droplets are imaged under 4 fluorescent channels to extract the FluoreCode of each droplet. The micrographs are representative examples from three independent experiments. The droplets are then clustered based on the FluoreCode information. The font colors indicate the fake colors used in the imaging channels. Alexa-488 (green), Alexa-546 (yellow), Alexa-594 (red), Alexa-647 (blue). f A t-distributed stochastic neighbor embedding (T-SNE) projection of 153,302 droplets containing 6527 recognized clusters, ~99.5% of the entire combinatorial space. The colors are used to represent individual clusters. Scale bars: 100 μm.
Validation of CFE in DropAI
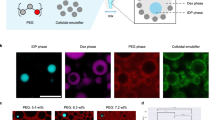
Having established the microfluidics, we next examined the compatibility of CFE in the DropAI workflow. We first asked whether CFE could be carried out in the picoliter droplet reactors. To investigate this, we emulsified an E. coli lysate-based CFE assay into 80-μm droplets using a microfluidic flow-focusing chip (Supplementary Fig. 1a), with fluorinated oil and a biocompatible polyethylene glycol-perfluoro polyether (PEG-PFPE) surfactant (Fig. 2a). The assay was designed to express a reporter protein, sfGFP. After incubation, the droplets exhibited notable fluorescence, indicating successful expression of sfGFP (Supplementary Fig. 3a). However, the emulsions were not mechanically stable, as large collapsed droplets were observed (Supplementary Fig. 3b). To stabilize the emulsions, we added Poloxamer 188 (P-188), a non-ionic triblock-copolymer surfactant, and Polyethylene glycol 6000 (PEG-6000), a biocompatible crowding reagent, to the aqueous phase (CFE mix). These polymers have been verified as effective stabilizers for droplet-based biochemical assays22,23. Indeed, after adding P-188 and PEG-6000, the droplets remained intact throughout the incubation process (Supplementary Fig. 3c). Next, we extracted the aqueous phase from the emulsions and measured the sfGFP fluorescence. The fluorescence level was comparable to that of a parallel bulk CFE assay (standard 15 μL reactions in 1.5 mL test tubes), suggesting that CFE efficiency is identical in both systems (Supplementary Fig. 3d).
We then investigated whether CFE is compatible with droplet fluorescence barcoding. To test this, we first conducted droplet-based CFE in the presence of dye molecules (AlexaFluor 546) and found that adding fluorescent dye did not affect protein expression (Supplementary Fig. 3e). In addition, we confirmed that the dye molecules remained in the aqueous phase without diffusing into the surrounding oil for up to 24 h (Supplementary Fig. 3f). Thus, the fluorescence coding resolution would not be compromised by dye molecules transporting across droplets during the CFE incubation (4 h). Next, we encoded the presence of Mg2+, an essential ion in CFE, to test the fluorescence coding and decoding. As depicted in Fig. 3b, the droplets formed two distinct clusters, and the occurrence of sfGFP fluorescence correlated with the encoded presence of Mg2+. Encouraged by this result, we further encoded a five-grade concentration gradient of Mg2+. As shown in Fig. 3c, five distinct clusters of droplets were clearly resolved. The sfGFP levels in these clusters indicated that sfGFP expression peaked at 12 mM Mg2+, which was confirmed by identical experiments performed in bulk (Fig. 3c inset). Overall, these results validate that DropAI is a feasible and efficient approach for optimizing CFE systems.
a Illustration of the CFE mixture composition and emulsification process. Preliminary experiments encoding the (b) presence and (c) concentrations of Mg2+. The upper panels display the encoding process and post-CFE droplets under the encoding dye (Alexa-647) and sfGFP channels. The scatter plot on the bottom reveals the fluorescence intensity profile of the droplets. The inset column plot in (c) depicts a parallel experiment performed in bulk (15 μL in-tube). The data are presented as mean ± s.d., based on six independent experiments. d–f In-droplet screening of 12 CFE components. d The 3-color coding strategy. e A representative micrograph of the encoding droplets (false colors are used for display purposes). Independent layers are disclosed in Supplementary Fig. 4a. f A 3D scatter plot showing the fluorescence intensity profile of the droplets across the combinatorial space. The color of the dots indicates the sfGFP intensity. g, h In silico optimization. g A 7-layer neural network model used in the optimization. The model interprets the presence of components as binary values and outputs the contribution score of each component. (h) Comparison of the normalized sfGFP yield (fluorescence intensity) obtained from the droplets vs. the model predictions. The linear fitting of the test set exhibits a Y-intercept of 0.09 and a slope of 0.89. i The in silico estimated contribution of the 12 components to the protein yield. The grey shading indicates additives for the transcription (TX) and translation (TL) process. Spermidine, Folinic acid, and PEP are selected for the second-round optimization (highlighted with red fonts). Scale bars: 100 μm.
Combinatorial screening of E. coli-based CFE
After proof-of-principle experiments, we proceeded to use DropAI to optimize a CFE system derived from E. coli, which is well-developed with standardized formulations and widely used in various laboratories18,24. Our goal was to achieve a simplified formulation by identifying key additives essential for protein expression and discarding non-essential ones. Additionally, we aimed to improve the overall yield by optimizing the concentrations of the identified key additives.
We conducted two rounds of DropAI screening using the E. coli-based PANOx-SP CFE system, following standardized protocols24. The first round aimed to determine the minimal set of essential additives, and the second round aimed to optimize their concentrations. In the first round, we started with 12 widely used components from the PANOx-SP system, including 7 major additives for transcription-translation and 5 energy suppliers for ATP regeneration. We used the suggested concentrations of the components24 (Supplementary Table 1) and only encoded their presence. Note that each combination was set to have no more than one energy substrate. Thus, these components constituted a combinatorial space of 768 [27 × (5 + 1)]. For the in-droplet screening, we applied three fluorescent colors to encode a subset of the combinatorial space (100 combinations) and used sfGFP yield (fluorescence intensity) as the output (Fig. 3d). As the carrier droplets were not coded, we used a simplified microfluidic device (Supplementary Fig. 1a) where the carrier drops were produced directly on-chip.
The in-droplet screening (Fig. 3e) generated 31,598 droplet-based CFE data points (Fig. 3f). The raw in-droplet screening data were binned into 100 clusters according to the FluoreCode (Supplementary Fig. 4a, b) and fed into a neural network model (Fig. 3g). Here, we employed the neural network due to its ability to handle unseen, potentially nonlinear relationships. This approach has recently been adopted for high-dimensional searches in chemical and biological spaces25. The model interpreted the presence of each additive as binary values, with normalized sfGFP fluorescence as the output (Supplementary Fig. 5). Among the input data, 70 combinatorial sets were used to train the model and 30 sets to test its prediction accuracy. We next evaluated nine neural network architectures with varying numbers of trainable linear layers from one to nine (Supplementary Fig. 6a). Considering accuracy and simplicity, the final model consisted of 7 fully connected layers. During the training process, the mean absolute percentage error (MAPE) loss function converged to ~5.6% after 1000 training epochs without obvious overfitting (Supplementary Fig. 5c). Using the 7-layer model, we evaluated the correlation between the predicted and measured yields and obtained a Pearson coefficient of 0.94, a 95% confidence interval of 0.87 to 0.97, and a coefficient of determination (R2) for the test set of 0.88 (Fig. 3h). The P-value was <0.0001. These statistics indicate reasonably good consistency between the predictions and the test sets. The established model was compared with K-nearest neighbors26 (KNN) and support vector regression27 (SVR), and we confirmed that our model outperformed the other two (See Methods and Supplementary Figs. 6b–d). These results verified the establishment and accuracy of the primary screening model.
The established model scanned through the entire combinatorial space and scored the contribution score of the 12 compounds to protein yield by calculating the average ratio between the yields with and without each compound (see Methods), Fig. 3i. Among the transcription-translation factors, spermidine and folinic acid exhibited scores greater than 1.1, suggesting that their presence alone can increase the yield of the CFE system by more than 10%. However, NAD, putrescine, tRNA, K2C2O4, and CoA were deemed non-essential for protein yield. For these additives, we also considered the potential correlations between components that might not be captured when analyzed independently. To address this, we profiled the contributions of all additive pairs. The results (Supplementary Fig. 7) show that pairs containing folinic acid, spermidine, or both consistently exhibit high contribution scores. This agrees with predictions based on individual contributions and suggests no strong promotive or inhibitory correlations between components. Among the energy sources, phosphorylated energy substrates [phosphoenolpyruvate (PEP), fructose 6-phosphate (F6P), and fructose-1,6-diphosphate (FDP)] had significantly higher contribution scores than non-phosphorylated substrates (glucose and pyruvate). This finding aligns with the glycolysis process28, where phosphorylated substrates have higher binding specificity to enzyme molecules, thereby promoting the forward reaction. Based on these prediction results, we decided to retain spermidine and folinic acid while discarding the other transcription-translation factors. For energy substrates, we selected PEP, which has the highest contribution score of 1.4.
Concentration optimization of the simplified CFE formulation
Having established a simplified composition, we proceeded to the second screening round to optimize the concentrations of the selected additives. We used 3 fluorescent colors to encode 53 (125) combinations of PEP, spermidine, and folinic acid concentrations (Fig. 4a). The in-droplet screening produced 49,148 datasets (Fig. 4b), which were subsequently binned into the 125 combinations (Supplementary Fig. 4c). For in silico optimization, we switched to extreme gradient boosting (XGBoost, Fig. 4c), a scalable decision tree-based strategy well-suited for handling continuous input problems through threshold segmentation29. Moreover, XGBoost is robust against overfitting, especially in scenarios where neural networks might produce large output variations from small changes in input29. The XGBoost model was initiated with a training set of 100 experimental combinations. During the training process, the training set was divided into 5 subsets for cross-validation. The final model comprised 180 weak decision trees (Supplementary Fig. 8a) and was verified using the remaining 25 experimental combinations, exhibiting a Pearson coefficient of 0.99, a 95% confidence interval of 0.98 to 1, and an R2 of 0.99 (Supplementary Fig. 8b) for the test set (P < 0.0001). While these statistics validated the capability of the model, it is important to acknowledge the potential risk of data leakage30, which may contribute to the high R2 value observed. We also benchmarked our XGBoost against six other approaches (Supplementary Table 5) and found it to perform the best in our context. These results suggest XGBoost’s particular effectiveness in small-scale regression problems31,32. Similar high R2 values have been reported in recent studies employing XGBoost32,33.
a, b In-droplet screening to optimize the concentrations of the selected components. a The fluorescence coding strategy. b A 3D scatter plot showing the fluorescence intensity profile of the droplets. The color of the dots indicates the sfGFP intensity. c–e In silico optimization. c An extreme gradient boosting model used in the optimization. d Concentration ranges scanned in the model prediction. e The estimation matrix sorted by yield level. f–i In vitro verification. f Fold change in the expression yield of 12 proteins with the simplified and optimized CFE formulation compared to the original recipe. The two sfGFP columns depict the same data because sfGFP was considered fully dissolvable (see “Methods” for fluorescence-based quantification of sfGFP), and the data are shown as mean ± s.d. of three independent experiments. Western blot results of (g) Sfp (27 kDa) and (h) CAR (128 kDa). i SDS-PAGE analysis of Vlm1 (370 kDa). M, T, and S are marker, total fraction, and soluble fraction, respectively. The gel images are representative examples from three independent experiments, all yielding consistent results. The background shadings highlight a representative round of DropAI optimization: in-droplet screening (clear), in silico optimization (grey), and in vitro verification (green).
After validating the model, we used it to span a broader range of concentrations (Fig. 4d) and predict the yields of 3780 combinations (Fig. 4e). The prediction revealed the highest yield at 40 mM PEP, 130 μM spermidine, and 4 mM folinic acid. To validate the prediction, we employed the highest-yield formula to express sfGFP in bulk and found that, compared to the original formula, the yield increased by 1.9-fold, from 0.71 to 1.38 mg/mL (Supplementary Fig. 9a; note that the protein yields are determined using a linear standard curve made in-house). Meanwhile, the total cost of additives was reduced by 2.1-fold, from 1.87 to 0.90 USD per mL of CFE assay (Supplementary Table 1). As a result, the unit cost of expressing sfGFP with the CFE was reduced by 4-fold, from 2.63 to 0.72 USD/mg.
To further test the general applicability of the optimized formulation, we expressed 12 proteins of various molecular weights (27 to 370 kDa) in vitro (Supplementary Fig. 9b). The fold changes in expression levels compared to those obtained with the original formulation are detailed in Fig. 4f. As shown, the expression levels of five proteins, including Sfp (27 kDa, Fig. 4e), BesC (29 kDa), BesD (29 kDa), and BesE (29 kDa), were maintained. Notably, TxtE (45 kDa) experienced a 4.5-fold increase in total expression and a 2.3-fold increase in soluble expression. Additionally, BesA (50 kDa), CAR (128 kDa, Fig. 4f), CAR-Q283P (128 kDa), Vlm2 (284 kDa), and Vlm1 (370 kDa, Fig. 4g) showed noticeable increases in total and/or soluble expression. The expression of TEII (27 kDa) and BesB (54 kDa) was slightly lowered. Interestingly, fusion of sfGFP to three representative proteins (BesB—reduced and no soluble expression, TEII—reduced expression, and BesE—maintained expression) could help enhance their expression levels, yet the terminus of target proteins for fusing sfGFP needs to be determined by experiments, respectively (Supplementary Fig. 10). While these results may suggest that our optimization was biased towards sfGFP, they also indicate a potential approach for optimizing the expression of non-fluorescent proteins: fusing a reporter gene (e.g., sfGFP) with the target gene and applying the standard DropAI workflow. Taken together, our streamlined protocol demonstrates comparable or enhanced expression of various proteins while markedly reducing the costs associated with CFE reactions. These advancements are crucial for leveraging CFE systems in the rapid, economical, and efficient production of therapeutics, chemicals, and materials. The success optimization also showcased the effectiveness of DropAI in combinatorial screening, which significantly reduced the experimental effort and cost via indexed droplet reactions and artificial intelligence.
Optimization of B. subtilis-based CFE by transfer learning
We next questioned if we could adapt the established E. coli model to fit a different chassis, thereby reducing the effort required to optimize CFE systems across various chassis. To achieve this, we employed transfer learning, a machine-learning approach that repurposes a pre-trained model for a different task by learning from a limited set of new data. To examine this concept, we selected B. subtilis, a Gram-positive bacterium widely used as a microbial cell factory for recombinant protein production34, as the new chassis. For the in-droplet screening, we started from the primary screening results from the E. coli system and generated 125 experimental combinations (N = 59,741) by varying the concentrations of spermidine, folinic acid, and PEP (Supplementary Fig. 11a). Next, the pre-trained E. coli model was used to generate baseline predictions based on the B. subtilis experimental data. These baseline predictions were incorporated into the original feature matrix of a XGBoost model. To assess the effectiveness of transfer learning, we varied the experimental set size between 27 (3 × 3 × 3) and 125 (5 × 5 × 5), resulting in 24 distinct datasets (Supplementary Table 4f). For each dataset, we constructed two models: one utilizing transfer learning and the other without (Supplementary Fig. 11b). In all models, 80% of the experimental data was allocated for training, while 20% was reserved for testing. Model performance was benchmarked using R² values. As shown in Fig. 5b, the transfer learning approach exhibited high prediction accuracy (R2 > 0.95) even with a limited experimental scale (e.g., 27 and 36 combinations). In contrast, the direct learning approach required at least 75 experimental sets to achieve similar accuracy. This suggests that transfer learning can achieve a ~ 2- to 3-fold reduction in the experimental inputs. For a 36-combination set (3 × 4 × 3), transfer learning achieved a Pearson coefficient of 0.99, a 95% confidence interval of 0.91 to 1, and an R2 of 0.99 (P < 0.0001). The linear fitting exhibited an intercept of −0.008 and a slope of 1.01. In contrast, the direct learning model produced an R2 of 0.83, an intercept of −0.49, and a slope of 1.50 (Fig. 5c). These results clearly demonstrate the advantage of applying transfer learning, as it significantly reduces the scale of experimental datasets (by a factor of 2 in our case) required to achieve reasonable prediction accuracy.
a Schematic of the transfer learning process. The process initiates with the established E. coli-based model. By learning limited B. subtilis experimental data, the E. coli model is transferred to a B. subtilis model through a feature enhancement approach. b Model accuracy obtained at varying learning set sizes. The transfer learning approach is compared with a model directly built on the B. subtilis experimental data. c The measured sfGFP yield (in-droplet screening) vs. the predicted yield of a direct learning model and the transfer learning model. Both yields are normalized. Both models study 36 (3 × 4 × 3) experimental combinations. d The estimation matrix sorted by yield level. e Bulk sfGFP expression under five notified compositions and the model prediction results. S (spermidine), F (folinic acid), P (phosphoenolpyruvate). f Experimental estimation of the optimized formulation to express sfGFP. The data are presented as mean ± s.d., based on multiple independent experiments (N = 6). NC (negative control without adding plasmid), B.S. (Bacillus subtilis), E.C. (Escherichia coli). Two-sided t-tests were used to analyze the data (see Methods). P values are classified as follows: p < 0.01 (**), p < 0.001 (***), p < 0.0001 (****). For original B.S. vs. rep. 1, p = 0.0002; for rep. 1 vs. rep. 2, p = 0.99, which is considered not significant (ns). g Fold change in the expression yield of three peptides with the simplified and optimized CFE formulation compared to the original recipe.
Next, we utilized a 36-combination transfer-learning model (3 × 4 × 3, R2 ~ 0.99) to predict the yield of 3780 distinct combinations of varying concentrations of spermidine, folinic acid, and PEP (Fig. 5d). To evaluate the predictions, we selected five combinations and performed bulk CFE using the specific concentrations of spermidine, folinic acid, and PEP to express sfGFP. The predicted and experimental yields for these combinations were normalized to the highest value and compared in Fig. 5e. The results indicated a consistent trend in CFE yield between the predictions and bulk experiments, although the exact yield levels did not perfectly match. To further evaluate the optimization, we chose a predicted high-yield combination of 5 mM spermidine, 100 μM folinic acid, and 35 mM PEP to express sfGFP in bulk. This formula led to a sfGFP yield of 0.24 mg/mL, representing an approximate 2-fold increase compared to our previous B. subtilis recipe (Fig. 5f). Compared to B. subtilis CFE using the optimized E. coli formulation (the high-yield combination before model transfer, Fig. 4e), the yield further increased by 14%. Beyond sfGFP, the optimized and simplified formulation also demonstrated maintained or increased yields when expressing three short antimicrobial peptides (Fig. 5g and Supplementary Fig. 12), highlighting the general utility of the optimized formulation. Additionally, our results confirm that transfer learning across different CFE systems is both feasible and effective. This concept could also be extended beyond CFE to optimize other biochemical systems, significantly reducing the need for labor-intensive wet experiments.
Discussion
CFE systems, particularly those based on cell lysates, represent promising platforms for various synthetic biology applications, including therapeutic development8, genetic part/circuit prototyping7,35, artificial cell construction36,37, natural product biosynthesis38,39,40, and biomanufacturing41,42. Central to these applications is the need for consistent and efficient protein expression. In CFE reactions, supplemented components such as amino acids, nucleotides, salts, polyamines, and energy sources are essential alongside cell lysates and gene templates to ensure high-yielding protein production1. However, the cost of these additives can be prohibitive, driving up the overall expense of CFE systems. As a result, CFE has not been utilized for real-world, industrial-scale production. Through DropAI optimization, we successfully streamlined the CFE formulation by eliminating five additives while maintaining or even enhancing protein expression levels compared to unoptimized reactions. Our findings highlight significant opportunities for improving protein expression in cell lysate-based CFE systems. Moreover, we demonstrated the transferability of our optimized recipe to another CFE system derived from B. subtilis using transfer learning. These results underscore the effectiveness and robustness of employing transfer learning to optimize novel CFE systems with minimal experimentation. While we successfully adapted the model from E. coli to B. subtilis, other factors should also be considered when adapting models to different biological systems. Specifically, variations in ribosome abundance, RNA polymerase types, tRNA diversity, and codon usage/bias can differ significantly between organisms, particularly between prokaryotes and eukaryotes. For a real-world setting, the model can be packaged with a user-friendly interface that generates predicted yields when component types and concentrations are entered. Additionally, the model could include real-time adjustment mechanisms, allowing users to fine-tune hyperparameters based on prior outputs.
Droplet microfluidics has recently been integrated with CFE for applications such as high-throughput regulatory part prototyping43, biomolecular circuit optimization44, and synthetic cell construction45. However, using droplet microfluidics to optimize the CFE system itself has remained unexplored prior to this work, largely due to the complexity of CFE systems and the lack of capable platforms. As we have showcased, DropAI is an efficient, rapid, and cost-effective strategy for high-throughput screening (HTS) of CFE systems. During the two-round optimization of the E. coli-based CFE system, approximately 100,000 droplets were measured (Figs. 3f & 4b), corresponding to a total volume of only 25 μL. Considering the liquid lost during microfluidic processes, collection, transfer, and imaging, the overall reagent consumption is around 100 μL. The liquid handling in DropAI is fully streamlined with microfluidics, allowing combinatorial reactions to be prepared at 300 Hz. This means only 6 min are required to prepare 100,000 independent screening reactions. The overall operation time of DropAI is around 4 h per round (15 min reagent preparation, 20 min microfluidic setup, 2 h satellite droplet generation, 25 min merging, and 1 h imaging). If the component-coding droplets are prepared in advance as stock, which is entirely feasible (Supplementary Fig. 2a), the operation time can be reduced to approximately 2 h. In contrast, screening an equivalent scale of reactions using well-plates and liquid-handling robot-based HTS platforms (with a unit reaction volume of around 10 μL) would consume over 1 liter of reagents, and the logistics of liquid handling could take several days15,46,47.
DropAI is a scalable platform. There is further room for improving DropAI’s throughput and combinatorial coding capacity. The microfluidics used in this work include droplet generation and multi-droplet merging. These techniques can be parallelized to enhance throughput48,49,50, allowing approximately 108 combinatorial reactions to be conducted per day. Additionally, the combinatorial coding capacity can be expanded by incorporating more fluorescent colors and engineering the microfluidics to support the grouping and merging of more droplets. For instance, using a commercial 6-color droplet reader, such as the BioRad QX600, the combinatorial coding capacity could reach a scale of 106. The unique scalability and compatibility of DropAI offer potential advantages over other microfluidics-based HTS approaches, such as microwell arrays51. A current limitation of DropAI is that the screening reactions require a fluorescence output, necessitating the design and construction of appropriate fluorescent reporters if the product itself is not inherently fluorescent. Another limitation is the reliance on fluorescent dyes to label specific conditions, which requires ensuring that the dye molecules are compatible with the reaction.
The combinatorial screening of CFE systems presented here uses only part of its capacity. DropAI may also be applied to various other scenarios, such as combinatorial drug screening52,53,54, chemical process optimization25, and catalyst discovery55. For example, massive unique drug combinations can be generated with cancer cells encapsulated in droplets using the microfluidics of DropAI. With appropriate reporter design, cells killed by specific drug combinations produce fluorescence signals that can be measured during in-droplet screening. Using the experimental data, AI can further predict drug combinations and concentrations with optimal therapeutic efficacy. This approach enables high-throughput and cost-efficient screening of anti-cancer drug combinations. This concept can also be extended to the combinatorial screening of enzyme cascades, biomolecular circuits, and catalyst combinations. This versatility and efficiency make DropAI a powerful tool for advancing research and development in synthetic biology and beyond.
Methods
Microfluidic device fabrication
The microfluidic devices (Supplementary Fig. 1a) were designed using AutoCAD and printed as dark-field plastic photomasks. To fabricate the devices, a 3-inch wafer was coated with photoresist SU-8 3025 (Microchem) using a spin coater in a two-step process: 30 seconds at 500 rpm, followed by 60 s at 2500 rpm. The wafer was soft-baked on a hotplate at 95 °C for 20 min, exposed to a 120 mW lamp (M365L2, Thorlabs) for 3 min and 20 s under a photomask, and baked again at 95 °C for 4 min. The repeated process was applied if a second layer was needed. Next, the wafer was incubated in SU-8 developer (Microchem) for 10 min, and excess developer was removed with isopropanol and ethanol, followed by blow-drying with a nitrogen gun. The wafer was then placed in a plastic petri dish, covered with a PDMS precursor (SYLGARD 184, Dow Corning) mixed with a curing agent in a 10:1 (w/w) ratio, degassed in a vacuum chamber, and cured at 60 °C for 8 h. The cured PDMS slab with the pattern was peeled off from the mold, and inlet/outlet ports were created using a 0.7 mm hole puncher. The patterned PDMS slab was bonded to a glass slide using oxygen plasma treatment. Before use, the devices were treated with Aquapel (PPG Industries) to render the channel surfaces hydrophobic.
Strains, media, and plasmids
E. coli DH5α was used for cloning and plasmid propagation. E. coli BL21 Star (DE3) and B. subtilis 164T7P were employed for cell extract preparation, respectively. LB medium (10 g/L sodium chloride, 5 g/L yeast extract, and 10 g/L tryptone) was used for cultivating bacterial cells. For cell extract preparation, cells were grown in 2×YTPG medium, which consists of 10 g/L yeast extract, 16 g/L tryptone, 5 g/L sodium chloride, 7 g/L potassium hydrogen phosphate, 3 g/L potassium dihydrogen phosphate, and 18 g/L glucose, adjusted to pH 7.2. The genes and plasmids used in this work are detailed in Supplementary Table 2.
Preparation of cell extracts
1 L of 2×YTPG medium was inoculated with an overnight culture to an initial OD600 of 0.05. When the OD600 reached 0.6-0.8, cells were induced with 1 mM IPTG and harvested at an OD600 of 3.0. The cells were washed three times with cold S30 Buffer (10 mM Tris-acetate, 14 mM magnesium acetate, and 60 mM potassium acetate). The cell pellet was resuspended in S30 Buffer (1 mL/g of wet cell mass) and lysed by sonication (10 s on/off, 50% amplitude, input energy ~600 Joules). The lysate was centrifuged twice at 12,000 × g for 10 min at 4 °C. The resulting supernatant was flash-frozen in liquid nitrogen and stored at −80 °C until use.
Cell-free gene expression
The CFE reactions were performed in 1.5 mL centrifuge tubes (15 μL) or 80 μm droplets (250 pL) at 30 °C for 4 h. Each reaction contained 27% (v/v) cell extract, 13.3 μg/mL plasmid, an energy substrate (PEP, FDP, F6P, glucose, or pyruvate; initial concentration 33 mM), and 35 other components. The component information for the original and optimized formulations of E. coli and B. subtilis CFE systems is detailed in Supplementary Table 1. The droplet CFE formulation contained additional 0.2% (v/v) P-188 and 1% (v/v) PEG-6000 to stabilize the emulsions. Synthesized proteins were analyzed using SDS-PAGE and Western blotting. The concentration of sfGFP was determined based on fluorescence intensity measured with a microplate reader (SYNETGY H1). For this, 2 μL of the CFE reaction was diluted with 48 μL nuclease-free water and placed in a flat-bottom 96-well plate. Measurements of sfGFP fluorescence were taken with excitation at 485 nm and emission at 528 nm. The fluorescence was converted to concentration (mg/mL) using a linear standard curve created in-house. For other proteins, their Western blot band densities and areas were analyzed by ImageJ, which were used to estimate the fold changes in yield between the optimized and original recipes. For in-droplet screening, the CFE mix was prepared at 1.3-fold of the final concentration, considering dilution in the merging process. The screening components were also removed from the CFE mix and prepared in the satellite droplets.
Western blot
The CFE reaction mixture was centrifuged at 12,000 × g for 10 min at 4 °C, and the supernatant was collected as the soluble fraction. Both 10 μL of total protein and 10 μL of soluble protein were mixed with 10 μL of 2x loading buffer and heated at 98 °C for 10 min. Each sample (10 μL) was then loaded onto an SDS-PAGE gel and transferred to a PVDF membrane (Bio-Rad) using 1x transfer buffer (25 mM Tris-HCl, 192 mM glycine, 20% methanol, pH 8.3). The membrane was blocked with Protein Free Rapid Blocking Buffer (EpiZyme) for 30 min at room temperature, followed by three washes with 1x TBST buffer (10 mM Tris-HCl, 150 mM NaCl, 0.1% Tween 20, pH 7.5). It was then incubated with His-Tag Mouse Monoclonal Antibody (Proteintech) diluted 1:10,000 in TBST buffer for 1 h, washed three more times with 1x TBST buffer, and incubated with HRP-Goat Anti-Mouse IgG (H + L) Antibody (Proteintech) diluted 1:10,000 in TBST buffer for another hour. After a final set of three washes, the membrane was visualized using Omni ECL reagent (EpiZyme) under UVP ChemStudio (analytikjena). Finally, protein band densities were analyzed by Image J, which were subsequently used to evaluate the fold changes of yield (optimized/original).
Microfluidic operations
The microfluidic experiments were conducted at a custom microfluidic station consisting of an inverted microscope equipped with a fast-speed camera and 8 syringe pumps. The oil [HFE-7500 (3 M) and 2.5% wt PEG-PFPE surfactant (Dapu Biotechnology)] and prepared aqueous solutions were stored in 1 mL plastic syringes and connected to the microfluidic chip mounted on the microscope through plastic tubing (SCI PE/2). To produce the fluorescently labeled droplets, stock solutions of each encoded condition were prepared by mixing a fluorescent dye with the encoding component(s) at specified concentrations (see Supplementary Table 4) in 50 μL aliquots. These stock solutions were then emulsified using the microfluidic droplet maker described in Supplementary Fig. 1a. Two different microfluidic designs were employed to produce carrier droplets (only used in experiments of Fig. 2) and satellite droplets to obtain the desired sizes. Different satellite droplets within a single fluorescent color were generated using the same device, with dye concentrations arranged in ascending order. To streamline the process and avoid repeated reagent loading, we sometimes preloaded all solutions into a plastic tubing connected to a syringe. Adjacent aqueous solution segments within the tubing were separated by short oil slugs. The tubing was then attached to the microfluidic chip for droplet generation, and the resulting droplets were collected in a 1.5-mL centrifuge tube, creating a mixed droplet library containing all fluorescence levels. The droplets were gently pipetted to mix them before being loaded onto the droplet merger (Supplementary Fig. 1a).
On the merger, the carrier and satellite droplets formed arbitrary groups. In the main channel, the satellite droplets moved faster than the carrier droplets, allowing them to merge with the carrier droplet before reaching the micro-teeth structure. This increased velocity arose because satellite droplets, being smaller than the channel width, are centered within the channel by the shear-induced lift force. The velocity of a pressure-driven flow follows a parabolic profile, with the highest velocity at the center. In contrast, the larger carrier droplets occupied the entire channel as slugs, moving at the average speed of the pressure-driven flow. At the micro-teeth structure, the grouped droplets were merged under an AC electrical field (1.5 kV Vpp, 50 kHz). The field was generated by an inverter (TDK CXA-L0605-VJL) circuit. The flow rates applied in the droplet generation and merging experiments are detailed in Supplementary Table 3. The stated flow rates allow a slightly lower loading frequency for the satellite droplets compared to the carrier droplets during the synchronization process, minimizing the likelihood of a single carrier droplet merging with two satellite droplets from the same pool.
Merging droplets using an AC electrical field is a widely employed technique in microfluidics. The underlying mechanism involves a combination of dielectrophoresis, electrostatic interaction, capillary instability, and interface destabilization56. Briefly, the merging process relies on polarization and hydrodynamic forces to induce coalescence. The AC field generates temporary dipoles within the droplets, producing electrostatic attraction and dielectrophoretic forces that align and draw the droplets together. Simultaneously, the field deforms the droplet interfaces, reducing the thickness of the separating film. Electrohydrodynamic flows and interfacial instabilities further destabilize this film, ultimately causing its rupture and droplet merging.
In the characterization of the microfluidics of DropAI (Fig. 2), the carrier droplets were color-coded and, therefore, pre-generated as a mixture before being reloaded onto the grouping and merging chip. However, for the in-droplet screening experiments (Figs. 3–5), the carrier droplets (now containing the CFE mixture) were not color-coded. To simplify the process, we utilized a modified grouping and merging device with on-chip generation of carrier droplets (Supplementary Fig. 1a).
In-droplet screening
The screening components, their concentrations, fluorescent dye, and dye concentrations in proof-of-concept experiments and each screening round are detailed in Supplementary Table 4. To obtain the encoded screening conditions in the droplets, the droplet libraries were loaded onto a cell counting slide (Thermal Fisher) for microscopic observation (Nikon, Eclipse Ti2) and to a 4-channel digital PCR droplet reader (Nebula Reader, Dapu Biotechnology) for multi-channel imaging. During imaging, the droplets were simultaneously identified, and the corresponding fluorescent intensities were measured with a built-in droplet analysis software (Nebula-Astrolabe, Supplementary Fig. 1d). For the clustering of 94 combinations, the raw droplet intensity data was filtered to eliminate marginal points with a multi-band-pass filter program built in MATLAB (version R2024a). For other experiments, the raw data was used without filtering. The data was categorized into different combination bins with a custom MATLAB program.
In silico optimization
We used the neural network model57,58 to evaluate the contribution of the 12 additives to sfGFP yield. The data used to train the combinatorial screening model was obtained directly from the in-droplet screening. The droplet experiment tested 100 combinations, each with more than 300 repeats. Before training, the sfGFP yield (fluorescent intensity) was normalized to the maximum value among the data. The normalized yield of each combination was averaged, and the mean values were used. In each combination, we used one-hot encoding to describe the component’s existence, with “1” indicating the presence and “0” indicating the absence (Supplementary Fig. 5a). Among the 100 sets of experimental results, 70 are for training, and 30 are for validation. The dataset was randomly split using the TensorFlow function “train_test_split” after shuffling. The established model traversed the entire combinatorial space and predicted the yield of each combination. Note that every combination was set to have no more than one energy substrate. Thus, the entire combinatorial space contained 768 [27 × (5 + 1)] combinations.
We adhered to the “Occam’s Razor Principle” throughout the modeling and used the simplest model possible to achieve accurate predictions, which fundamentally reduces the risk of overfitting due to the reduction in trainable parameters. We tested on various layers (1-9), determining that the 7-layer model has low prediction loss while the overfitting is within a controllable range. Moreover, we incorporated random pruning into model training, with a dropout rate set at 0.25. We also compared our 7-layer neural network model with KNN and SVR. The KNN algorithm achieved a MAPE of 9.58% on the training set and 11.12% on the validation set, resulting in the highest error among the three models. While SVR’s error was closer to that of neural network, it exhibited more pronounced underfitting (Supplementary Fig. 6).
After prediction, we evaluated the contribution of each additive to the yield by comparing the yields with a certain additive present to the yields with it absent:
where x depicts an additive and i represents one of n combinations. The model was on Python (version 3.10.11) and TensorFlow (version 2.8.0). The training was compiled with the Adam optimizer.
In the second round of optimization, we used the XGBoost algorithm to train the concentration model. The in-droplet screening generated 125 combinations of varying concentrations of spermidine, folinic acid, and PEP. The concentrations of the three components the sfGFP yield (fluorescence intensity) were used as inputs. The input sets were randomly split into a training set (100 combinations) and a test set (25 combinations). The learning set was first standardized with ‘StandardScaler’ in Python’s scikit-learn library, converting the distributions to have a mean of 0 and a variance of 1. Then, the learning started with hyperparameter tuning with the XGBoost regression model using GridSearchCV. The grid search systematically explored candidate values for several key parameters, including maximum tree depth (max_depth), learning rate (learning_rate), subsampling ratio (subsamples), and the number of weak learners (n_estimators). These hyperparameters are provided in Supplementary Table 6. Cross-validation was employed to assess the model’s performance across multiple random data splits, minimizing the risk of overfitting and reducing model bias. The model with the best hyperparameters was finally selected, and its accuracy was estimated using the validation sets. The verified model was used to predict 3,840 combinations and the yield was normalized to the largest value in the prediction (Fig. 4e). The model was compiled in Python (version 3.10.11).
We selected R2 and MAPE as the primary metrics because neural networks and XGBoost are highly flexible models capable of capturing complex relationships. R2 quantified the proportion of variability in the target variable explained by the model. During primary screening, neural networks occasionally exhibited overfitting, which we monitored using MAPE due to its lower sensitivity to extreme variance.
Transfer learning
To transfer the E. coli model to fit B. subtilis-based CFE, we employed the following steps: a) Application of pre-trained model: the pre-trained E. coli model was initially applied to generate baseline predictions based on experimental data from the B. subtilis-based CFE system. This approach allowed us to utilize established model parameters and patterns from E. coli as foundational insights for predicting outcomes in the B. subtilis-based CFE system. b) Feature set augmentation: these baseline predictions were then incorporated into the original feature matrix of the B. subtilis-based CFE system as additional inputs for a XGBoost model. This integration resulted in an enhanced feature set that leveraged both the foundational knowledge from the E. coli model and the unique data characteristics of the B. subtilis-based CFE system. c) Training of transferred model: using the augmented feature set, we trained a semi-quantitative prediction model specifically for yield levels in the B. subtilis-based CF system. We calculated the R² on both the training and testing sets to evaluate model accuracy. To assess transfer capability, the experimental input for transfer learning in the B. subtilis-based CFE system was down-scaled to 27 distinct subsets (Fig. 5b and Supplementary Table 4). For each subset, 80% of the data was used for training and 20% for testing. A transferred model was obtained for each subset, and the R² score was calculated (Supplementary Fig. 11b). For comparison, we also built an XGBoost model trained directly on each experimental subset, following the same training process as with the E. coli model. Among the 27 transfer-learning models, one (Fig. 5c) was selected to predict 3,840 combinations, with the yield normalized to the highest predicted value.
Statistical analysis
The data in Figs. 3c (inset), 5e, and f are presented as mean ± s.d., based on multiple independent experiments (n = 6 for 3c and 5 f). Figure 5f uses two-sided t-tests to assess the significance of changes between the original and optimized B. subtilis CFE systems, as well as between independent sets of optimized B. subtilis CFE experiments. Statistical significance is indicated as follows: p < 0.01 (**), p < 0.001 (***), p < 0.0001 (****). Mean absolute percentage error (MAPE), Pearson coefficient, 95% confidence interval, and coefficient of determination (R2) were used to assess the performance of AI models.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the key findings of this study are available within the article and its Supplementary Information files or from the corresponding author upon request. Data required to generate key results of the machine learning models have been deposited in GitHub (https://github.com/Liulab2023/DropAI/tree/main/Data) and Zenodo59. Source data are provided with this paper.
Code availability
All the codes regarding droplet clustering, AI models and in silico screening are available on GitHub at https://github.com/Liulab2023/DropAI and archived on Zenodo with DOI59.
References
Silverman, A. D., Karim, A. S. & Jewett, M. C. Cell-free gene expression: an expanded repertoire of applications. Nat. Rev. Genet. 21, 151–170 (2020).
Liu, W. Q., Zhang, L. K., Chen, M. Z. & Li, J. Cell-free protein synthesis: recent advances in bacterial extract sources and expanded applications. Biochem. Eng. J. 141, 182–189 (2019).
Lu, H. F., Ouyang, J. P., Liu, W. Q., Wu, C. Z. & Li, J. Enzyme-polymer-conjugate-based pickering emulsions for cell-free expression and cascade biotransformation. Angew. Chem. Int. Ed. 62, e202312906 (2023).
Liu, Y. et al. An in vitro hybrid biocatalytic system enabled by a combination of surface-displayed, purified, and cell-free expressed enzymes. ACS Synth. Biol. 13, 1434–1441 (2024).
Liu, W. Q. et al. Cell-free biosynthesis and engineering of ribosomally synthesized lanthipeptides. Nat. Commun. 15, 4336 (2024).
Liew, F. E. et al. Carbon-negative production of acetone and isopropanol by gas fermentation at industrial pilot scale. Nat. Biotechnol. 40, 335–344 (2022).
Karim, A. S. et al. In vitro prototyping and rapid optimization of biosynthetic enzymes for cell design. Nat. Chem. Biol. 16, 912–919 (2020).
Tan, X., Letendre, J. H., Collins, J. J. & Wong, W. W. Synthetic biology in the clinic: engineering vaccines, diagnostics, and therapeutics. Cell 184, 881–898 (2021).
Stark, J. C. et al. Rapid biosynthesis of glycoprotein therapeutics and vaccines from freeze-dried bacterial cell lysates. Nat. Protoc. 18, 2374–2398 (2023).
Nguyen, P. Q. et al. Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nat. Biotechnol. 39, 1366–1374 (2021).
Rasor, B. J. et al. Mechanistic insights into cell-free gene expression through an integrated-omics analysis of extract processing methods. Acs Synth. Biol. 12, 405–418 (2023).
Bajorath, F. Integration of virtual and high-throughput screening. Nat. Rev. Drug Discov. 1, 882–894 (2002).
Broach, J. R. & Thorner, J. High-throughput screening for drug discovery. Nature 384, 14–16 (1996).
Blay, V., Tolani, B., Ho, S. P. & Arkin, M. R. High-throughput screening: today’s biochemical and cell-based approaches. Drug Discov. Today 25, 1807–1821 (2020).
Sun, X. C., Vilar, S. & Tatonetti, N. P. High-throughput methods for combinatorial drug discovery. Sci. Transl. Med. 5, 205rv1–205rv1 (2013).
Tan, X. et al. Systematic identification of synergistic drug pairs targeting HIV. Nat. Biotechnol. 30, 1125–1130 (2012).
Zeng, W. Z., Guo, L. K., Xu, S., Chen, J. & Zhou, J. W. High-throughput screening technology in industrial biotechnology. Trends Biotechnol. 38, 888–906 (2020).
Kwon, Y. C. & Jewett, M. C. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. Sci. Rep. 5, 8663 (2015).
Li, J., Wang, H., Kwon, Y. C. & Jewett, M. C. Establishing a high yielding streptomyces-based cell-free protein synthesis system. Biotechnol. Bioeng. 114, 1343–1353 (2017).
Hodgman, C. E. & Jewett, M. C. Optimized extract preparation methods and reaction conditions for improved yeast cell-free protein synthesis. Biotechnol. Bioeng. 110, 2643–2654 (2013).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Wagner, O. et al. Biocompatible fluorinated polyglycerols for droplet microfluidics as an alternative to PEG-based copolymer surfactants. Lab Chip 16, 65–69 (2016).
Bollenbach, L., Buske, J., Maeder, K. & Garidel, P. Poloxamer 188 as surfactant in biological formulations—an alternative for polysorbate 20/80? Int. J. Pharm. 620, 121706 (2022).
Jewett, M. C. & Swartz, J. R. Mimicking the Escherichia coli cytoplasmic environment activates long-lived and efficient cell-free protein synthesis. Biotechnol. Bioeng. 86, 19–26 (2004).
Shields, B. J. et al. Bayesian reaction optimization as a tool for chemical synthesis. Nature 590, 89–96 (2021).
Cover, T. & Hart, P. Nearest neighbor pattern classification. IEEE Trans. Inf. Theory 13, 21–27 (1967).
Drucker, H., Burges, C. J., Kaufman, L., Smola, A. & Vapnik, V. Support vector regression machines. Adv. Neural Inf. Process. Syst. 28, 779–784 (1996).
Chandel, N. S. Glycolysis. Cold Spring Harb. Perspect. Biol. 13, a040535 (2021).
Chen, T. Q. & Guestrin, C. XGBoost: a scalable tree boosting system. Kdd'16: In Proc. of the 22nd Acm Sigkdd International Conference on Knowledge Discovery and Data Mining, 785-794 (2016).
Kapoor, S. & Narayanan, A. Leakage and the reproducibility crisis in machine-learning-based science. Patterns 4, 100804 (2023).
Gong, J., Chu, S., Mehta, R. K. & McGaughey, A. J. H. XGBoost model for electrocaloric temperature change prediction in ceramics. NPJ Comput. Mater. 8, 140 (2022).
Huang, Y.-J., Chen, C.-h & Yang, H.-C. AI-enhanced integration of genetic and medical imaging data for risk assessment of Type 2 diabetes. Nat. Commun. 15, 4230 (2024).
Wang, X., Bouzembrak, Y., Oude Lansink, A. G. J. M. & van der Fels-Klerx, H. J. Designing a monitoring program for aflatoxin B1 in feed products using machine learning. NPJ Sci. Food 6, 40 (2022).
Zhang, K., Su, L. Q. & Wu, J. Recent advances in recombinant protein production by Bacillus subtilis. Annu. Rev. food Sci. Technol. 11, 295–318 (2020).
Liu, W. Q., Wu, C. Z., Jewett, M. C. & Li, J. Cell-free protein synthesis enables one-pot cascade biotransformation in an aqueous-organic biphasic system. Biotechnol. Bioeng. 117, 4001–4008 (2020).
Li, L. et al. Permeability-engineered compartmentalization enables in vitro reconstitution of sustained synthetic biology systems. Adv. Sci. 9, 2203652 (2022).
Rothschild, L. J. et al. Building synthetic cells-from the technology infrastructure to cellular entities. ACS Synth. Biol. 13, 974–997 (2024).
Feng, J. C. et al. Application of cell-free protein synthesis system for the biosynthesis of L-Theanine. ACS Synth. Biol. 10, 620–631 (2021).
Ji, X. Y., Liu, W. Q. & Li, J. Recent advances in applying cell-free systems for high-value and complex natural product biosynthesis. Curr. Opin. Microbiol. 67, 102142 (2022).
Moore, S. J., Lai, H.-E., Li, J. & Freemont, P. S. Streptomyces cell-free systems for natural product discovery and engineering. Nat. Prod. Rep. 40, 228–236 (2023).
Rasor, B. J. et al. Toward sustainable, cell-free biomanufacturing. Curr. Opin. Biotechnol. 69, 136–144 (2021).
Bundy, B. C. et al. Cell-free biomanufacturing. Curr. Opin. Chem. Eng. 22, 177–183 (2018).
Gan, R. et al. High-throughput regulatory part prototyping and analysis by cell-free protein synthesis and droplet microfluidics. ACS Synth. Biol. 11, 2108–2120 (2022).
Hori, Y., Kantak, C., Murray, R. M. & Abate, A. R. Cell-free extract based optimization of biomolecular circuits with droplet microfluidics. Lab Chip 17, 3037–3042 (2017).
Olivi, L. et al. Towards a synthetic cell cycle. Nat. Commun. 12, 4531 (2021).
Dove, A. Screening for content— the evolution of high throughput. Nat. Biotechnol. 21, 859–864 (2003).
Duffy, S. & Avery, V. M. Development and optimization of a novel 384-well anti-malarial imaging assay validated for high-throughput screening. Am. J. Trop. Med. Hyg. 86, 84–92 (2012).
Zhu, P. A. & Wang, L. Q. Passive and active droplet generation with microfluidics: a review. Lab Chip 17, 34–75 (2017).
Zhang, H. et al. An ultra high-efficiency droplet microfluidics platform using automatically synchronized droplet pairing and merging. Lab Chip 20, 3948–3959 (2020).
Lee, S., Kim, H., Won, D. J., Lee, J. & Kim, J. On-demand, parallel droplet merging method with non-contact droplet pairing in droplet-based microfluidics. Microfluid. Nanofluidics 20, 1 (2016).
Kehe, J. et al. Massively parallel screening of synthetic microbial communities. Proc. Natl. Acad. Sci. USA 116, 12804–12809 (2019).
Jaaks, P. et al. Effective drug combinations in breast, colon and pancreatic cancer cells. Nature 603, 166–173 (2022).
Pemovska, T., Bigenzahn, J. W. & Superti-Furga, G. Recent advances in combinatorial drug screening and synergy scoring. Curr. Opin. Pharmacol. 42, 102–110 (2018).
Kulesa, A., Kehe, J., Hurtado, J. E., Tawde, P. & Blainey, P. C. Combinatorial drug discovery in nanoliter droplets. Proc. Natl. Acad. Sci. USA 115, 6685–6690 (2018).
McCullough, K., Williams, T., Mingle, K., Jamshidi, P. & Lauterbach, J. High-throughput experimentation meets artificial intelligence: a new pathway to catalyst discovery. Phys. Chem. Chem. Phys. 22, 11174–11196 (2020).
Teo, A. J. T., Tan, S. H. & Nguyen, N. T. On-demand droplet merging with an AC electric field for multiple-volume droplet generation. Anal. Chem. 92, 1147–1153 (2020).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representations by back-propagating errors. Nature 323, 533–536 (1986).
Abadi, M. et al. TensorFlow: a system for large-scale machine learning. in Proc. of the 12th USENIX conference on Operating Systems Design and Implementation 265–283 (USENIX Association, 2016).
LiuLab_ShanghaiTech. Liulab2023/DropAI: Initial release for manuscript submission. Zenodo https://doi.org/10.5281/zenodo.14878884 (2025).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (grant no. 2023YFA0914000 to J.L.), the National Natural Science Foundation of China (grant nos. 52322305 to S.L., 32171427 to W.Q.L., 52473098 to S.L., and 62374170 to Y.Luo), the Shanghai Science and Technology Committee (grant nos. 23QA1406600 to Y.Liu and 24ZR1451000 to J.L.), and School of Physical Science and Technology of ShanghaiTech University (grant no. SPST-YSFZ-2024-01 to Y.Liu). The authors would also like to acknowledge the ShanghaiTech AI4S Initiative (grant no. SHTAI4S202404 to Y.Liu) and the high-performance computing (HPC) platform of ShanghaiTech University.
Author information
Authors and Affiliations
Contributions
Y.Liu and J.L. designed the experiments. J.Z. and Y.M. performed the experiments. W.G. and S.Y. built the AI models and conducted in silico optimization. W.Z., X.J. and W.Q.L. performed experiments assisted in the experiments. Y.Liu., J.Z., Y.M., Y.Luo and X.Z. analyzed the data and prepared the illustrations. Y.Liu and J.L. wrote the manuscript with input from all authors. Y.Liu, J.L. and S.L. contributed to project conception. Y.Liu supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sriram Chandrasekaran and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, J., Meng, Y., Gao, W. et al. AI-driven high-throughput droplet screening of cell-free gene expression. Nat Commun 16, 2720 (2025). https://doi.org/10.1038/s41467-025-58139-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58139-0
This article is cited by
-
Next-generation biosensing for in situ monitoring
Nature Sensors (2026)
-
High-throughput combinatorial screening of antiplatelet drugs for personalized medicine
Microsystems & Nanoengineering (2026)
-
Integrating the Blood–Brain Barrier and Blood-CSF Barrier Organ-on-Chip Models: Development and Future Directions
BioChip Journal (2026)
-
Sensitive, high-throughput, metabolic analysis by molecular sensors on the membrane surface of mother yeast cells
Nature Communications (2025)
-
Reconstituting transcription–translation-coupled DNA replication within complex in vitro biological systems
Nature Communications (2025)