Abstract
Fluorescent proteins are indispensable molecular tools for visualizing biological structures and processes, but their limited photostability restricts the duration of dynamic imaging experiments. Yellow fluorescent proteins (YFPs), in particular, photobleach rapidly. Here, we introduce mGold2s and mGold2t, YFPs with up to 25-fold greater photostability than mVenus and mCitrine, two commonly used YFPs, while maintaining comparable brightness. These variants were identified using a high-throughput pooled single-cell platform, simultaneously screening for high brightness and photostability. Compared with our previous benchmark, mGold, the mGold2 variants display a ~4-fold increase in photostability without sacrificing brightness. mGold2s and mGold2t extend imaging durations across diverse modalities, including widefield, total internal reflection fluorescence (TIRF), super-resolution, single-molecule, and laser-scanning confocal microscopy. When incorporated into fluorescence resonance energy transfer (FRET)-based biosensors, the proposed YFPs enable more reliable, prolonged imaging of dynamic cellular processes. Overall, the enhanced photostability of mGold2s and mGold2t enables high-sensitivity imaging of subcellular structures and cellular activity over extended periods, broadening the scope and precision of biological imaging.
Similar content being viewed by others
Introduction
Fluorescent proteins (FPs) are essential molecular tools for studying cellular and subcellular structures and functions. Their remarkable versatility has facilitated their widespread adoption in the life sciences to visualize protein localization, intracellular trafficking, and signal dynamics. Engineered FP-based indicators further expand their utility by reporting cellular changes, such as ion fluctuations1,2, membrane potential shifts3,4,5, and pH variations6,7. The broad spectrum of available FPs and indicators allows for multi-spectral imaging, where multiple probes function simultaneously.
Despite the widespread use, many FPs face a critical limitation: rapid photobleaching under repeated or prolonged illumination. This degradation reduces signal-to-noise ratios and constrains experiment duration. Photobleaching is particularly problematic in advanced imaging techniques, such as super-resolution microscopy8, single-molecule imaging9,10, and voltage imaging5, which require intense and/or sustained illumination to achieve high spatial or temporal resolution. These challenges have fueled the demand for FPs that combine brightness with superior photostability.
A breakthrough in photostability came with the development of StayGold, a highly photostable green FP discovered through metagenomic screening of marine organisms11. StayGold and its monomeric versions set a new benchmark, allowing extended imaging with minimal fluorescence loss11,12,13,14. However, this advancement remains limited mainly to the green spectral class, leaving a gap in other color channels. Notably, yellow FPs (YFPs) photobleach more rapidly than FPs in other spectral classes, despite their widespread use in biosensors15,16,17 and multi-spectral imaging18,19,20. Enhancing YFP photostability is crucial for enabling prolonged imaging and expanding their applicability in advanced microscopy.
To facilitate this goal, we previously developed SPOTlight, a high-throughput single-cell screening platform that can evaluate millions of FP variants for brightness and photostability21. Using the platform, we engineered mGold, a YFP variant with a 4- to 5-fold improvement in photostability over its predecessor, mVenus, while maintaining brightness. By selecting for both traits simultaneously, we overcame the typical trade-off between brightness and stability in FP engineering22,23,24.
Building on this work, we here report how we further optimized the SPOTlight platform to increase throughput and applied it to evolve mGold into two new variants: mGold2s and mGold2t. These proteins exhibit a 4-fold improvement in photostability over mGold while preserving brightness. Compared to widely used YFPs such as mVenus and mCitrine, they demonstrate up to a 25-fold increase in photostability with minimal alterations to other biophysical properties.
We tested mGold2s and mGold2t to validate their performance across various imaging modalities, including widefield, total internal reflection fluorescence (TIRF), super-resolution, single-molecule, and laser-scanning confocal microscopy. Additionally, because YFPs are widely used in fluorescence resonance energy transfer (FRET) applications, we incorporated mGold2s and mGold2t into FRET-based biosensors. The superior photostability of the YFPs enhances imaging of subcellular structures and cellular activity over prolonged periods, broadening the utility of biological imaging.
Results
An upgraded single-cell screening platform identifies two bright YFPs with improved photostability
We sought to address the relative photolability of current YFPs via high-throughput protein engineering. Screening was performed using SPOTlight, our pooled single-cell screening platform21 (Fig. 1a & Supplementary Note), which we optimized to shorten single-cell photoactivation, increase screening throughput, and decrease variability brightness measurement variability (Supplementary Note). Enhancements included the use of a stronger photoactivation light (4.9 W vs. 1.7 W), reducing the time to optically label individual cells (Fig. 1a, step 3) from 45–60 to 10–15 s, enabling approximately four times more cells to be tagged in the same period. Switching from phosphate-buffered saline (PBS) to minimal culture media for long-term yeast cell imaging doubled the sorting viability of yeast cells isolated by Fluorescence-Activated Cell Sorting (FACS) post-imaging. Finally, replacing TagBFP with mTurquoise225 as the reference fluorescent protein used for normalizing cell-to-cell expression variation reduced the coefficient of variation of brightness measurements by ~7.5-fold.
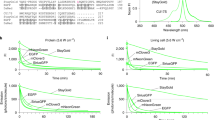
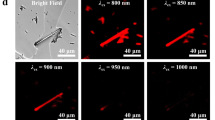
a SPOTlight screening workflow used to develop bright and photostable YFP variants. b mGold2s and mGold2t showed excitation/emission peaks at 517/529 nm and 515/527 nm, respectively. c mGold2s and mGold2t expressed in HEK293A cells retained around half of their fluorescence after continuous illumination of 520/5-nm light at 2.4 mW/mm2 for 1 h. Mean ±95% CI, n = 6 independent transfections. d mGold2s and mGold2t were more photostable than other YFPs and exhibited similar brightness. HEK293A cells expressing the variants were photobleached with 520/5-nm light at 17 mW/mm2. Mean ±95% CI, n = 6 independent transfections. * denotes dimeric YFPs. e mGold2s and mGold2t showed higher photostability than mVenus and mGold at different irradiance levels. Mean ±95% CI, n = 6 independent transfections. f mGold2s did not exhibit sensitivity to chloride in the physiological range (magenta), while mGold2t displayed a slight sensitivity. Mean ±95% CI; For mEYFP, n = 4 replicates from a single purification; For other YFPs, n = 16 replicates from 6 independent purifications. g mGold2s matured at the same rate as mVenus, while mGold2t displayed a faster maturation time. Maturation times were quantified as maturation time delay relative to EBFP2. Red center lines, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range. *P = 0.0298; ns, P > 0.05; Dunnett’s T3 multiple comparisons test after two-sided Welch’s ANOVA test. h mGold2s and mGold2t caused similarly low cytotoxicity as mVenus. Mean ±95% CI, n = 3 independent transfections. **P = 0.0019; ns, P > 0.05; Dunnett’s T3 multiple comparisons test after two-sided Welch’s ANOVA test. i mGold2s successfully performed as a fusion tag in HeLa cells. b–h Statistical analysis is provided in the Supplementary Information. Source data are provided as a Source Data file.
We randomly mutagenized mGold21, a mVenus-derived FP with the highest photostability among existing monomeric YFPs21 (Fig. 1a, step 1). We screened the resulting libraries in yeast cells, evaluating the brightness and photostability of 1,125,438 cells representing 204,987 variants across 7 rounds of screening (Fig. 1a, step 2; Supplementary Fig. 1a–c). In each screening round, we optically labeled 200–400 single variants for retrieval via FACS (Fig. 1a, step 3 → 4). These cells were then grown into colonies on agar plates, picked, expanded, and plated onto multi-well plates (Fig. 1a, step 4). We performed population-level validation by quantifying the mean brightness and photostability of thousands of cells per variant. The top 5–10 variants per round were sequenced.
The two best variants based on brightness and photostability were named mGold2s and mGold2t. mGold2s differs from mGold by 14 mutations: V1A, V22I, H77N, Q80R, K101E, D117G, I123V, S147C, Y151F, K156R, K158Q, A163V, D173V, and G232S (Supplementary Fig. 1d). mGold2t differs from mGold2s by encoding a threonine rather than a serine at position 205, as reflected by the ‘t’ rather than ‘s’ designation (Supplementary Fig. 1e). The impact of these mutations on photostability and brightness is summarized in Supplementary Fig. 1b. The combination of Q80R, S147C, and G232S led to the most significant improvements in brightness (+23%) and photostability (+69%).
Characterization of mGold2s and mGold2t
mGold2s and mGold2t retained similar one- and two-photon spectra to mGold and mVenus (Fig. 1b & Supplementary Fig. 2). When we imaged live mammalian cells expressing mGold2s and mGold2t using continuous widefield illumination of 520/5-nm (central wavelength/spectral width) light at 2.4 mW/mm2 for 1 h, mGold2s and mGold2t retained 50% and 43% of their original fluorescence, corresponding to photobleaching half-lives of 60 and 42 min, respectively (Fig. 1c & Supplementary Fig. 3). In contrast, mVenus and mGold rapidly photobleached with half-lives of 3 and 12 min, respectively. mGold2s and mGold2t showed different photobleaching patterns: mGold2t initially photobleached more rapidly than mGold2s, but this trend reversed at later time points (Fig. 1c & Supplementary Fig. 4a). We attributed the decreased fluorescence under continuous illumination to photobleaching, as fluorescent proteins are generally resistant to degradation. For instance, avGFP, the ancestral protein of mGold and mVenus, has a degradation half-life of 26 h26. Consistent with these previous studies, we observed stable protein levels for all YFPs over 6 h in the presence of the protein synthesis inhibitor cycloheximide (Supplementary Fig. 5).
We expanded our brightness and photostability analysis to include nine additional monomeric or dimeric YFPs. Under continuous widefield illumination of 520 nm light at 17 mW/mm2, mGold2s and mGold2t emerged as the most photostable YFPs, with half-lives of 450 and 445 s, respectively. For example, they demonstrated ~20- and ~25-fold greater photostability than mVenus (half-life: 23 s) and mCitrine (18 s), respectively (Fig. 1d). Compared with their precursor, mGold (half-life: 114 s), their starting template, mGold2s and mGold2t were ~4 times more photostable (Fig. 1d). mGold2s and mGold2t exhibited similar brightness as mGold and mVenus in cells and were 16–17% brighter as purified proteins (Table 1). Both variants displayed superior photostability across all tested irradiance levels in mammalian cells (Fig. 1d, e & Supplementary Fig. 4b) and yeast (Supplementary Fig. 4c–e). mGold2s and mGold2t’s superior photostability was not restricted to widefield imaging; in laser-scanning confocal microscopy using 514-nm light at 14 µW to 57 µW, mGold2s and mGold2t exhibited up to ~8–9 times the photostability of mVenus and up to ~3 times that of mGold (Supplementary Fig. 6). mGold2s and mGold2t are thus the most photostable YFPs to date while maintaining the brightness of the brightest monomeric YFPs.
To confirm our results are robust to variations in measurement approaches, an independent lab (N. Shaner, UCSD) evaluated the relative brightness and photostability of mGold2s and mGold2t in live mammalian cells under widefield microscopy. mGold2s and mGold2t showed similar brightness as mVenus and mGold (Supplementary Fig. 7a), consistent with brightness characterization in the St-Pierre lab (Fig. 1d). Under continuous widefield 510/24-nm illumination at 40 mW/mm², mGold2s was 15 times more photostable than mVenus (Supplementary Fig. 7b), showing similar fold-difference observed in the St-Pierre dataset at a similar power level (38 mW/mm²; Fig. 1e). mGold2t showed a 22-fold improvement over mVenus, exceeding the performance measured in the St-Pierre assay (16-fold), primarily due to the absence of an initial rapid bleaching phase (Supplementary Fig. 7b). The reason for this variation in photobleaching dynamics remains unclear, with potential factors including differences in cell type (U2OS vs. HEK293) and minor differences in illumination wavelength (510/24 vs. 520/5 nm). Overall, this replication experiment confirms that mGold2s and mGold2t are over an order of magnitude more photostable than and similar in brightness to mVenus.
We assessed the photostability of mGold2s and mGold2t under laser-scanning two-photon (2P) microscopy, a widely used technique for deep-tissue imaging27. Alongside mGold and mVenus, we included three GFPs as additional controls: EGFP28, mStayGold(J)12, and mBaoJin13. We conducted photobleaching using light at 950 nm, a wavelength at which all FPs displayed similar relative 2P excitation (~80% of the peak; Supplementary Fig. 8a). Unlike our one-photon (1P) results, all YFPs showed comparable brightness and photostability (Supplementary Fig. 8b, c). Two photostable GFPs under 1P illumination —mStayGold(J) and mBaoJin— did not outperform the YFPs in 2P photostability, even though YFPs bleach faster in 1P11. However, mStayGold(J) and mBaoJin were 1.7 and 2.8 times brighter, respectively, consistent with previous findings that lower-wavelength absorption increases 2P brightness29. EGFP, while 1.7-fold brighter than the YFPs, bleached significantly faster. Although mBaoJin displayed reduced 1P photostability than mStayGold(J)13, its photostability under laser-scanning 2P microscopy was similar. These findings highlight the weak correlation between FP performance under one- and two-photon illumination, stressing the importance of screening and evaluating FPs for the intended imaging modality3,30.
mGold2s and mGold2t showed reduced thermostability compared with mVenus and mGold, although their melting temperature of 77 °C remained well above mammalian physiological temperatures (Table 1). At 37 °C, mGold2s and mGold2t displayed a 5–10% brightness reduction compared with their fluorescence at 30 °C, whereas mVenus and mGold showed only a 1-2% decrease (Supplementary Fig. 9). mGold2s and mGold2t nevertheless demonstrated similar in cellulo brightness at 37 °C to mVenus and mGold (Fig. 1d) primarily because their greater molecular brightness (Table 1) compensated for their modest loss in thermostability. These results highlight that in cellulo brightness is influenced not only by cell division rates and FP molecular brightness, expression levels, and maturation times31, but also by FP thermostability.
We determined that mGold2s and mGold2t are monomeric using size exclusion chromatography (Supplementary Fig. 10a) and the OSER cell assay (Table 1, Supplementary Fig. 10b). While mGold2s showed minimal sensitivity to chloride, mGold2t showed 7% and 25% decreases in fluorescence at 0.1 and 1 M Cl-, respectively (Fig. 1f, Table 1). For context, mEYFP, known for its chloride sensitivity, showed 25% and 80% fluorescence reductions under the same chloride concentrations. mGold variants and mVenus exhibited multiple similar characteristics, including a ~3 ns fluorescence lifetime (Table 1), low acid sensitivity (pKa = 5.2–5.4; Supplementary Fig. 11, Table 1), fast maturation times (15 min for mGold2s and 13 min for mGold2t, Fig. 1g & Supplementary Fig. 12), low cytotoxicity (Fig. 1h), and high tolerance to protein fusions (Fig. 1i & Supplementary Figs. 13 and 14).
Evaluation of mGold2s and mGold2t for extended imaging across cell lines and imaging modalities
We next set to determine whether replacing photolabile YFPs with mGold2s and mGold2t would enable prolonged imaging without compromising assay functionality. We focused on a wide range of assays that would benefit from photostable FPs. We began by assessing mGold2s and mGold2t in super-resolution imaging using structure-illuminated microscopy (SIM). We imaged actin filaments in COS-7 cells expressing YFP-LifeAct fusions at a rate of 11 Hz for ~1.5 min, using a 488-nm laser at 95 mW/mm2. Each timepoint comprised nine raw frames reconstructed into a single SIM image. The resolution of SIM images was further enhanced through sparse deconvolution32 (Fig. 2a, b). mGold2s and mGold2t maintained over 80% of their initial fluorescence, significantly outperforming mVenus, which retained only about 25% (Fig. 2c & Supplementary Movie 1). mGold showed greater photostability than mVenus, but lower photostability compared to mGold2s and mGold2t, photobleaching to 50% of its initial fluorescence (Supplementary Fig. 15). mGold2s enabled visualization of actin dynamics similar to the actin blip dynamics described previously33 (Fig. 2d). This dynamic event involves actin punctum formation at the network node followed by disassembly of the punctum and can be detected as a spike in fluorescence intensity. These dynamics could not be observed using photolabile mVenus because of rapid photobleaching.
a YFP-LifeAct fusions were used to visualize actin filaments. TIRF-SIM total internal reflection microscopy structured illumination microscopy. b Imaging workflow. c mGold2s and mGold2t showed greater photostability than mVenus under TIRF-SIM. Traces represent mean values, and shaded areas are 95% CI. n = 40–50 regions across 10 cells. d mGold2s enabled extended visualization of actin dynamics, including an actin blip (arrow). Pseudocolor images are shown. arb. units stand for arbitrary units. e GPHN-FingR-YFP fusions were expressed in dissociated hippocampal neurons to visualize inhibitory post-synapses. KRAB Krüppel-associated box, GPHN Gephyrin. f Imaging workflow. g Representative images of FingR probes. White boxes indicate zoomed-in regions in panel (h). h Superior photostability of FingR-mGold2s enabled sustained imaging of inhibitory synapses. The top two rows show representative time series. The bottom graph displays mean ±95% CI, n = 9 foci across two cells. i Schematics of YR-ICUE, a FRET-based cAMP indicator. FRET fluorescence resonance energy transfer, cAMP cyclic adenosine monophosphate, Epac1 exchange protein directly activated by cAMP 1. j YR-ICUE variants with mGold2s and mGold2t showed enhanced photostability under widefield imaging compared with mVenus or mGold. Excitation: 520-nm light at 17 mW/mm2. Mean ±95% CI, n = 6 independent transfections. k Representative pseudocolor images showing YR-ICUE responses before and after adding Fsk and IBMX. Fsk forskolin, IBMX 3-isobutyl-1-methylxanthine. l Fluorescence responses were comparable for YR-ICUE-mGold2s and YR-ICUE-mVenus, while YR-ICUE-mGold2t showed slightly diminished response. R represents the RFP/YFP emission ratio. R0 is the emission ratio immediately preceding drug addition. Boxplots of 153–225 cells are shown. Center lines, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range. *P = 0.0174; ***P = 0.0008; ns, P > 0.05; Dunnett’s T3 multiple comparisons test after two-sided Welch’s ANOVA test. m Schematics of single-molecule pull-down assay with TIRF imaging. n mGold2s and mGold2t exhibited greater photobleaching lifetimes than mVenus across power levels. Excitation: 488 nm. Mean ±95% CI, n = 3 technical replicates. o The fraction of molecules showing undesirable photoblinking was similar across the YFP variants. Mean ±95% CI, n = 3 technical replicates. ns, P > 0.05; Dunn’s multiple comparisons test after two-sided Kruskal–Wallis test. Source data are provided as a Source Data file.
We also tested whether mGold2s and mGold2t could extend the SIM imaging of mitochondria. We fused mGold2s and mGold2t to an outer mitochondrial membrane localization signal to visualize mitochondria in COS-7 cells. Cells were continuously imaged by SIM for ~2 min under 58 mW/mm2 irradiance at a temporal resolution of 12 Hz. Under these conditions, mGold2s and mGold2t allowed for capturing 26–32 times more usable frames than mVenus and 6.5–8.2 times more than mGold (Supplementary Fig. 16).
Next, we explored the capability of mGold2s and mGold2t to prolong imaging of subcellular structures using laser-scanning confocal microscopy. We fused YFPs to Gephyrin-FingR (GPHN-FingR), an intrabody developed to visualize inhibitory synapses34 (Fig. 2e). We imaged the resulting probes in live dissociated hippocampal neurons, where GPHN-FingR-mGold2s and GPHN-FingR-mGold2t showed the expected punctate structures (Fig. 2g & Supplementary Fig. 17a). To accommodate dendritic movement during imaging, we captured twenty z-stacks at each time point, totaling 3,600 frames for each construct (Fig. 2f). After 30 min of continuous imaging, GPHN-FingR-mGold2s retained around 70% of its original fluorescence while GPHN-FingR-mVenus almost completely lost its fluorescence. GPHN-FingR-mGold2t was slightly less photostable than the mGold2s variant, decaying to around 45% of its original fluorescence. GPHN-FingR-mGold rapidly decayed to around 20% of its original fluorescence over the same timeframe (Supplementary Fig. 17b).
Recognizing the widespread use of YFPs in fluorescence resonance energy transfer (FRET) applications16,17, we evaluated the potential of mGold2s and mGold2t to enhance the photostability of existing FRET-based biosensors. We substituted mVenus in the yellow-red cyclic adenosine monophosphate (cAMP) indicator YR-ICUE35 with mGold, mGold2s, and mGold2t (Fig. 2i). In YR-ICUE, YFPs serve as FRET donors, and mCherry is the FRET acceptor. When continuously photobleached under 520-nm light at 17 mW/mm2 using widefield illumination, YR-ICUE-mGold2s and YR-ICUE-mGold2t produced a 17–20-fold increase in photostability compared with YR-ICUE-mVenus, and a 3–4.7-fold increase compared with YR-ICUE-mGold (Fig. 2j), consistent with previous characterization (Fig. 1c). To assess these biosensors’ dynamic range, cAMP release in HEK293T cells was transiently induced using isoproterenol (iso), a β-adrenergic receptor agonist, followed by maximal stimulation with adenylyl cyclase activator forskolin (Fsk) and phosphodiesterase inhibitor 3-isobutyl-1-methylxanthine (IBMX) (Fig. 2i, k–l & Supplementary Fig. 18). All YR-ICUE variants showed transient responses to β-adrenergic stimulation, confirming the reversibility of the biosensors (Supplementary Fig. 18). While the mGold and mGold2t YR-ICUEs exhibited small reductions in dynamic range, YR-ICUE-mGold2s maintained a response comparable to the original YR-ICUE-mVenus (Fig. 2k–l & Supplementary Fig. 18).
Finally, we assessed mGold2s and mGold2t in single-molecule applications using pull-down assays (Fig. 2m). mEYFP served as a control, given its frequent use in similar assays10,36. YFPs were expressed in HEK293 cells and pulled down post-lysis with an anti-GFP antibody. We then used a total internal reflection fluorescence (TIRF) microscope to monitor YFP fluorescence over several seconds at three excitation power levels: 530 µW, 1 mW, and 2 mW. At 530 µW, mGold2s and mGold2t had photobleaching lifetimes of 6.1 and 8.2 s, respectively —2.5 and 3.4-fold longer than mVenus (2.4 s), 1.2 and 1.7-fold longer than mGold (4.9 s), and 1.8 and 2.4-fold longer than mEYFP (3.4 s) (Fig. 2n). mGold2t consistently outperformed other YFPs at all power levels, while mGold2s extended lifetimes at 530 µW and 1 mW. As power increased, relative lifetime extensions decreased (Fig. 2n). Aside from their enhanced photostability, mGold2 variants could be captured by anti-gfp antibodies with similar specificity to other YFP variants (Supplementary Fig. 19) and we did not observed difference in photoblinking behavior (Fig. 2o).
Discussion
This study introduced bright FPs, mGold2s and mGold2t, the most photostable monomeric YFPs to date. This advancement challenges the traditional trade-off seen in previous reports22,23,24,37, where enhanced photostability came at the cost of decreased brightness, and vice versa. Our findings demonstrate that both attributes can be simultaneously improved21.
While photostability mutations are typically found near the chromophore, 10 mutations of the 14-15 present in mGold2s and mGold2t are located on the top or bottom of the beta-barrel, relatively far from the chromophore (Supplementary Fig. 1d, e). Similar to the proposed mechanism for the distal photostability-enhancing mutation G160C in mClover3 GFP38, these mGold2s/t mutations may increase photostability by limiting oxygen access, potentially by closing unidentified conduction pathways or by allosterically reducing inter-strand gaps near the chromophore39. Solving the structures of mGold, mGold2s, and mGold2t will be crucial to evaluate these hypotheses and deepen our understanding of the molecular mechanisms underlying photostability enhancement.
The S205T mutation, distinguishing mGold2t from mGold2s, is responsible for distinct photobleaching kinetics (Fig. 1c) and greater chloride sensitivity (Fig. 1f). This result is consistent with a previous finding that mutations at position 205 influenced photostability and chloride sensitivity in YFP-based biosensors40. However, the mechanisms by which distinct residues at this position differentially modulate photostability and chloride sensitivity remain unclear.
The mGold2 variants exhibit a modest reduction in thermostability. However, they maintain cellular brightness comparable to mVenus and mGold, as their higher molecular brightness offsets the decrease in thermostability (Table 1). We speculate that the A163V mutation contributes to reduced thermostability, as V163A was recently shown to increase thermostability and solubility of a GFP variant at 37 °C (ref. 41). A163V emerged from a screening round with mixed outcomes, enhancing photostability but reducing brightness (Supplementary Fig. 1b). Therefore, reversing A163V could restore the lost thermostability without negatively affecting the YFPs’ overall optical performance.
mGold2s and mGold2t share comparable biophysical properties, making them similarly suitable for a broad range of applications. For experiments involving high chloride concentrations, mGold2s is preferred due to its lower sensitivity to Cl- (Fig. 1f). The relative photostability of these two YFPs varied depending on the assay, and their performance rankings sometimes shifted during the experiment (e.g., Fig. 2j). Therefore, users aiming for maximum photostability should evaluate both variants within their specific experimental conditions.
While some applications presented here could use photostable fluorescent proteins of other colors, developing photostable FPs across the entire color spectrum is critical for experimental flexibility. Selecting a suitable FP depends on the experiment’s specific needs, including the instrument’s optical setup, spectral compatibility with other fluorophores, and intrinsic FP properties such as pKa and maturation time. For example, tri-color experiments often use cyan, yellow, and red FPs, whereas blue/green/red combinations are less common due to the low brightness of blue FPs and the phototoxicity of blue-shifted light used for their illumination42.
mVenus remains one of the most widely used FPs because of its fast maturation and high brightness. Our study demonstrates that substituting mVenus with mGold2 variants significantly enhances photostability without compromising other key performance metrics across diverse imaging methods and conditions. These results highlight the versatility of mGold2s and mGold2t as superior alternatives for upgrading mVenus-based constructs, promoting the adoption of these two YFPs within the research community. We anticipate these variants will set a gold standard for monomeric YFPs, offering extended imaging durations and improved temporal resolution for biological research and biotechnological applications.
Methods
Our research complies with all relevant ethical regulations. All procedures involving experimental animals were conducted in strict accordance with the National Institute of Health guidelines and approved by Animal Care and Use Committee of Baylor College of Medicine.
Cell culture
Bacterial culture
E. coli XL10-Gold (catalog no. 200315, Agilent) was used for general cloning. E. coli BL21(DE3) pLysS strain (catalog no. L1195, Promega) was used for purifying FPs. Miller LB broth (catalog no. BP1426, Fisher Scientific) and Miller LB agar (catalog no. BP1425, Fisher Scientific) supplemented with appropriate antibiotics were used to grow bacteria. They were prepared following the manufacturer’s instructions.
Yeast culture
Experiments with yeast cells were conducted using the S. cerevisiae BY4741 strain. Yeast expression plasmids used the URA3 gene as a selection marker. Cells were grown in yeast synthetic drop-out medium prepared by mixing minimal synthetic defined base (catalog no. 630411, Takara Bio) and uracil drop-out supplement (catalog no. 630416, Takara Bio) following the manufacturer’s instructions. To grow yeast cells at pH 7.0, the synthetic drop-out medium was supplemented with 10 mM HEPES (catalog no. H3375, Sigma-Aldrich) and adjusted to pH 7.0 using NaOH (catalog no. S5881, Sigma-Aldrich). To prepare competent yeast cells, yeast extract, peptone, and dextrose [YPD; catalog no. 242820, Becton Dickinson (BD)] were used. Competent yeast cells were created following a published protocol43. Agar plates were prepared by adding 25 g/L of agar (catalog no. BP1423, Fisher Scientific) to liquid synthetic drop-out and YPD media.
Mammalian cell culture
HEK293A cells (RRID: CVCL_6910, catalog no. R70507, Thermo Fisher Scientific) and HeLa cells (RRID: CVCL_0030, catalog no. CCL-2, ATCC) were used for mammalian experiments. Cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM)–high glucose (catalog no. D1145, Sigma-Aldrich) supplemented with 10% fetal bovine serum (FBS) (catalog no. F2442, Sigma-Aldrich), 2 mM l-glutamine solution (catalog no. G7513, Sigma-Aldrich), and 1% (% v/v) penicillin-streptomycin (catalog no. P4333, Sigma-Aldrich). COS-7 cells (RRID: CVCL_0224, catalog no. CRL-1651, ATCC) were cultured in high-glucose DMEM (catalog no. 21063029, Gibco) supplemented with 10% FBS (catalog no. 10437028, Gibco). HEK293T (RRID: CVCL_0063, catalog no. CRL-3216, ATCC) were cultured in DMEM (catalog no, 11885-084, Gibco) with 1 g/L glucose, 10% FBS (catalog no. 26140-079, Gibco), and 1% penicillin-streptomycin (catalog no. 15140-122, Gibco). All cell lines were maintained in a 5% CO2-controlled humidified incubator at 37 °C. U2-OS cells (HTB-96, ATCC) were grown in a 35 mm glass bottom dish (P35G-1.5-14-C, MatTek corporation) with DMEM (105666-016, Gibco) supplemented with 10% (v/v) FBS (10437-028, Gibco) under 5% humidified CO2 atmosphere at 37 °C. The cell lines were confirmed to be mycoplasma-free using the MycoAlert Mycoplasma Detection Kit (catalog no. LT07-118, Lonza).
Plasmid construction
Expression plasmids were subcloned using standard In-Fusion and Gibson Assembly cloning techniques, following manufacturers’ protocols. Selected plasmids used in this study and their sequences are available from Addgene (Addgene plasmids #231761 - #231790) Other plasmids are available upon request.
For FP purification, we subcloned the FPs in pNCS, a bacterial expression plasmid with a constitutive promoter44, a T7 terminator, and an ampicillin resistance marker. This plasmid also encoded an N-terminal 6xHis-tag for nickel affinity purification.
To express FPs in yeast, we used the pJL1 plasmid described previously21. This plasmid is a multicopy 2-micron plasmid with a URA3 auxotrophic selection marker, a strong constitutive pTDH3 promoter, and the tADH1 terminator. Ribosome-skipping 2A sequences for porcine teschovirus-1 (P2A) and/or Thosea asigna virus (T2A) were used for multi-cistronic expression.
To express constructs in mammalian cells, we utilized several plasmid backbones. For experiments that localized mGold2s and mGold2t to subcellular regions, we replaced the mGold for mGold2s and mGold2t in Addgene plasmids #158001 (Keratin), #158002 (endosome), #158003 (nucleus), #158004 (Endoplasmic reticulum), #158005 (Golgi), #158006 (lysosome), #158007 (mitochondria), #158008 (tubulin), #158009 (Actin). For the FRET biosensor experiment, the green-red (EGFP-stagRFP) indicator of cAMP45 (GR-ICUE2, Addgene plasmid #173017) was used as a template for sensor design. mGold2s and mGold2t replaced EGFP; and stagRFP was replaced with mCherry. For the FingR and two-photon microscopy experiments, we used the pCaggs backbone. The EGFP from GPHN.FingR-EGFP-CCR5TC (Addgene plasmid #46296) was replaced with YFPs. For two-photon spectra characterization, we replaced the mGold from pCaggs-mGold (Addgene plasmid #157996) with mGold2s or mGold2t. For the OSER assay, CytERM-YFP plasmids were constructed by replacing the mGFP from CytERM-mGFP (Addgene plasmid #62237) with YFPs.
To visualize actin filaments using super-resolution imaging, we used the pCMV-mGold-Mito-N-7 (Addgene plasmid #158007), replacing the mitochondrial targeting sequence of COX8 gene with LifeAct and mGold with various YFPs. To visualize mitochondria, the mitochondrial membrane targeting sequence of BCL2L1 and various YFPs were cloned into the pEGFP-N1 backbone, which encodes a cytomegalovirus (CMV) promoter for gene expression.
Other experiments using mammalian cells, including photobleaching, maturation time characterization, two-photon brightness measurements, and protein degradation rate characterization (cycloheximide-chase assay), were conducted using the pJL2 plasmids described previously21. Briefly, YFPs were expressed from the strong constitutive CMV promoter and the bovine growth hormone (bGH) polyadenylation terminator. This plasmid also co-expresses a reference EBFP2 via the P2A sequence. EBFP2 was used to normalize expression level variability.
General hardware setup
For SPOTlight screening and mGold2 characterization experiments, we used a widefield fluorescence microscope (Ti-E, Nikon Instruments) equipped with a motorized XY stage with linear encoders (H139E1N4, Prior Scientific), a hardware autofocus module (Perfect Focus System, Nikon Instruments), a 20 × 0.75–numerical aperture (NA) objective (CFI Plan Apo Lambda, Nikon Instruments), a laser diode illuminator (LDI-WF, 89 North), a scientific complementary metal-oxide-semiconductor (sCMOS) camera (ORCA Flash 4.0 V2, Hamamatsu), and imaging software (NIS-Elements HC version 4.60, Nikon Instruments). This microscope was also equipped with a multi-photon module (A1R-MP, Nikon Instruments), titanium:sapphire Chameleon Ultra II laser (Coherent), galvanometric mirrors, and gallium arsenide phosphide (GaAsP) detectors for 2-P imaging.
Different hardware setup was used for mGold2 application experiments. See the corresponding sections below for their descriptions.
SPOTlight screening to improve photostability and brightness of YFPs
Hardware setup for SPOTlight screening
Cyan, yellow, and red FPs were excited with 445/3-nm (peak/bandwidth), 520/5-nm, and 555/0.6-nm light, respectively, from a laser diode illuminator (LDI-WF, 89 North). Multi-bandpass dichroic mirrors (for CFP and YFP: ZT445/520rpc, Chroma; for RFP: ZT 405/470/555/640rpc, Chroma) were used. Excitation light was filtered with multiband filters (for CFP and YFP: ZET445/520x, Chroma; for RFP: ZET 405/470/555/640x, Chroma). Emission light was filtered with multiband filters (for CFP and YFP: ZET445/520M-OD8, Chroma; for RFP: ZET 405/470/555/640 m, Chroma). For optical tagging cells expressing photoactivatable FPs, light from a high-power 405-nm light-emitting diode (UHP-F3-405, Prizmatix) was patterned using a digital micromirror device (TI-LA-DMD, Nikon Instruments & Polygon1000, Mightex). Note that this light source had a greater wattage output than the light source used in our previous study21. Using a stronger light source increased the photoactivation kinetics and, thereby, the optical tagging throughput (Supplementary Note).
Building single-cell yeast libraries
To construct random mutagenesis libraries, we used GeneMorph II Random Mutagenesis Kit (catalog no. 200550, Agilent) following the manufacturer’s instructions. mGold was the initial template for random mutagenesis. The best variant for each screening round became the template for the next round. The PCR fragments generated using random mutagenesis were assembled into the pJL1 yeast expression backbone using the In-Fusion HD Cloning Kit (catalog no. 639650, Takara). This backbone also co-expressed PAmCherry1 (for optical tagging) and mTurquoise2 (for normalizing gene expression).
The In-Fusion product was transformed in yeast using the lithium acetate/single-stranded carrier DNA/polyethylene glycol method46. Transformed yeast cells were plated on 10–15 10-cm uracil drop-out agar plates and incubated at 37 °C. Seventy-two hours post-transformation, all the colonies were collected with a cell scraper (catalog no. 08-771-1A, Thermo Fisher Scientific) in 1 mL of sterile water. The cells were washed three times with water and plated on a 24-well glass-bottom plate (catalog no. P24-1.5H-N, Cellvis) pre-coated with 0.1 mg/mL solution of poly-L-lysine (PLL, catalog no. P1524, Sigma-Aldrich). The attached cells were washed twice with water. After the final wash, leucine drop-out media (catalog no. 630414, Takara Bio, buffered with 10 mM of HEPES, pH 7.0) was loaded to the wells for imaging. Using this imaging media helped increase cell viability after FACS-based cell sorting (Supplementary Note).
Imaging and photobleaching yeast libraries
Single-cell yeast libraries were imaged using the widefield fluorescence microscope described above. 169 nonoverlapping fields of view were sequentially scanned. Cells were photobleached for 45 s per field of view with 520/5-nm light at 90 mW/mm2. Yellow and cyan fluorescence images were captured for each field of view at t = 0, 22.5, and 45 s using 520/5-nm light at 8.1 mW/mm2 with 20 ms exposure time and 445/3-nm light at 8.8 mW/mm2 with 100 ms exposure time respectively.
Analyzing images
Images captured during photobleaching were segmented to identify individual cells. Segmentation was performed on the reference CFP image (at t = 0 s) using a machine learning-based segmentation software ilastik (version 1.3.2)47. To speed up the segmentation process, we developed a custom MATLAB (version r2020b, MathWorks) program that extracted segmentation parameters from ilastik-generated mask and segmented the rest of the images in parallel. With this parallelization, we could achieve a segmentation speed of 3 s per field of view. Because cells were attached to the imaging plate and exhibited minimal movement during imaging, segmentation masks from the initial time points could be used for later time points.
We calculated individual cells’ mean YFP and CFP fluorescence values at each time point. The cell brightness was defined as the ratio of YFP to CFP fluorescence at the initial time point (YFP/CFP). The normalization was performed to reduce the cell-to-cell variability caused by the differences in plasmid copy number among the transformed cells. For SPOTlight screening purposes, photostability was defined as a fraction of final fluorescence over initial fluorescence. Around 200 to 400 cells showing high brightness and photostability were selected for optical tagging and recovery.
Optical tagging and recovery of yeast cells
Each selected target cell’s x and y positions were determined from the images’ metadata. Target cells were placed in the center of the field of view. Photoactivation was conducted for 10–15 s, using the digital micromirror device (DMD) to restrict light at ~260 mW/mm2 to a 4 μm by 4 μm region centered on the target cells.
For each target cell, red fluorescence images were taken before and after photoactivation to monitor the photoactivation level of each cell. After photoactivation, cells were detached with trypsin-EDTA solution for 10 min at 37 °C and resuspended in a leucine drop-out medium supplemented with 10% FBS. Detached cells were washed once with PBS and resuspended in minimal media with 0.5% glycerol.
Fluorescence-activated cell sorting of optically tagged cells
A fluorescence-activated cell sorter (FACSAria II, Becton Dickinson) was used to detect and retrieve optically tagged cells. The cell sorter was equipped with lasers at 405 nm, 488 nm, and 561 nm to detect cyan, yellow, and red fluorescence, respectively. 450/50-nm, 530/30-nm, and 610/20-nm emission filters were used to filter cyan, yellow, and red fluorescence, respectively. We prepared yeast cells expressing single FPs (parental YFP only, mTurquoise2 only, and photoactivated PAmCherry1 only) or empty vector were prepared to compensate for bleed-through between the channels. To determine the optical gating strategy to select optically tagged cells, we prepared three photoactivation standard samples: (1) a sample without photoactivated cells, (2) a sample with ~100 cells photoactivated, and (3) a sample with ~200 cells photoactivated. We placed the sort gating that resulted in the appropriate ratio of cells (0:1:2) among the three samples. Cells were bulk sorted into wells of 96-well plates containing YPD media. The sorted cells were then plated on a URA drop-out agar plate and incubated at 37 °C for 72 h.
Secondary population-level screening
The sorted target cells grew to form colonies. Single colonies were separately picked and cultured overnight at 37 °C in the uracil drop-out medium buffered at pH 7.0 with 10 mM HEPES. Each culture is thus generated from the expansion of one specific variant. Yeast cultures were diluted around 20-fold and regrown until they reached absorbance values at 600 nm (OD600) of 0.3–0.5. The cells were washed three times with water and loaded on 96-well glass-bottom plates (catalog no. P96-1.5H-N, Cellvis), coated with 0.1 mg/mL of PLL. Each well contained one variant. The attached cells were washed twice with water and imaged in a leucine drop-out minimal medium. Cells were photobleached using 520/5-nm light at 90 mW/mm2. During photobleaching, YFP and CFP images were taken every 5 s with 5.1 mW/mm2 LDI power excitation light, 20 ms exposure time for YFP, and 24.7 mW/mm2 LDI power, 30 ms for CFP. Images were taken every 5 s to monitor the fluorescence during photobleaching. For each well, 2–3 fields of view were imaged and segmented using ilastik. The brightness and photostability of individual cells were computed after background subtraction. Brightness was defined as YFP fluorescence/CFP fluorescence. Photostability was determined by computing the photobleaching half-life, the time required for the fluorescence to decrease by half from its initial fluorescence level. Mean brightness and photostability values were determined to obtain a population-level performance of each variant.
Plasmid DNA was prepared from promising variants using a yeast plasmid miniprep kit (catalog no. D2001, Zymo Research) and sequenced to identify mutations.
In cellulo FP characterization
Comparing brightness and photostability of YFPs in yeast
Yeast cells were transformed using the LiAc/ssDNA-carrier/PEG method46. YFP variants were co-expressed with yeast codon-optimized mTurquoise2 using the pJL1 plasmid. A day before imaging, colonies were picked and cultured overnight at 37 °C in the uracil drop-out minimal media buffered at pH 7.0 using 10 mM HEPES. On the imaging day, yeast cultures were diluted around 20-fold and regrown until the OD600 reached 0.3–0.5. The cells were harvested, washed thrice with water, and loaded on 96-well glass-bottom plates coated with PLL. The attached cells were washed twice with water and imaged in leucine drop-out minimal media buffered at pH 7.0 with 10 mM of HEPES. Yeast cells were imaged using the Ti-E microscope described above. Photobleaching was conducted under 520/5-nm illumination at 17 mW/mm2, 38 mW/mm2, 59 mW/mm2, and 80 mW/mm2. During photobleaching, YFP and CFP images were taken every 5 s using 520/5-nm light at 5.1 mW/mm2 with 20 ms exposure and 445/3-nm light at 24.7 mW/mm2 with 30 ms exposure, respectively. We defined the brightness and photostability of the YFP variants as described in the Secondary population-level screening section above. We repeated the experiment on three days, measuring n = 2 independent cultures each day.
Comparing brightness and photostability of YFPs in HEK293A cell by the St-Pierre Lab
HEK239A cells were transfected using FuGENE (catalog no. E2311, Promega) following the manufacturer’s protocol. Briefly, cells were plated in a 96-well glass bottom plate at 30–40% confluency. For each well, 200 ng of plasmid (100 ng of pJL2 plasmid co-expressing YFP and EBFP2 and 100 ng of buffer plasmids) was mixed with 0.6 μL of FuGENE reagent in 13 μL of Opti-MEM (catalog no. 31985070, Thermo Fisher Scientific). The transfection mixture was added to the wells. Two days post-transfection, we replaced the culture media with Hank’s balanced salt solution (HBSS) (catalog no. H8264, Sigma-Aldrich), supplemented with 10 mM HEPES. Photobleaching and analysis were conducted in the same way as yeast photobleaching experiments. One exception was that we took BFP images (instead of CFP images) because EBFP2 was used as a reference protein. BFP was imaged using the 405/0.6-nm light, ZT 405/470/555/640rpc dichroic mirror, ZET 405/470/555/640x excitation filter, and ZET 405/470/555/640m emission filter. During photobleaching, YFP and BFP images were taken every 5 s with 3.5 mW/mm2 of 520/5-nm light with 10 ms exposure for YFP and 4.8 mW/mm2 of 405/0.6-nm light with 30 ms exposure for BFP. Photobleaching was conducted under 520/5-nm illumination at 2.4 mW/mm2, 17 mW/mm2, 38 mW/mm2, 59 mW/mm2, and 80 mW/mm2. We defined the brightness and photostability of the YFP variants as described in the Secondary population-level screening section above. We repeated the experiment on three days, measuring n = 2 independent transfections each day.
Comparing brightness and photostability of YFPs in U2OS cells by the Shaner Lab
U2OS cells were transfected using polyethylenimine (PEI) in ddH2O (1 mg/mL, pH 7.3, catalog no. 23966, Polysciences). The transfection mixture was prepared in Opti-MEM (catalog no. 31985047, Thermo Fisher Scientific) with 4.5 μg PEI and 500 ng plasmid and added to U2OS cells plated in 35 mm glass bottom dish. Two days post-transfection, fresh medium was provided 1–2 h before imaging.
For widefield photobleaching, U2OS cells expressing pCMV-H2B-FP fusions were imaged on a Nikon Ti-E microscope with a 60×/1.2 WI Plan Apo VC objective, Perfect Focus System, a Spectra X light source (Lumencor) set to 510/24 nm bandpass excitation, 514-nm dichroic, 542/27-nm emission filter, and an iXon888 EMCCD camera (Andor). Cells were imaged approximately 48 hours after transfection, with focusing using very low power fluorescence excitation ( ≤1% imaging intensity for ≤5 s) to prevent pre-bleaching of the FPs. Images were collected continuously at 1 s intervals with constant illumination measured at 40 mW/mm2. For photobleaching analysis, image stacks were processed in ImageJ using the TrackMate plugin with Cellpose 3.048 to detect nuclei. After background subtraction, the average nuclear intensity per cell was normalized to its value in the first image, and the normalized curves for all cells were averaged for each fluorescent protein to produce the reported bleaching curves.
For in cellulo brightness measurements, U2OS cells were grown and transfected as described above with pCMV plasmids encoding full-length mVenus, mEYFP, mGold, mGold2s, or mGold2t fused to a T2A self-cleaving peptide followed by mCherry2, which acted as the standard. Forty eight hours post-transfection, cells were imaged with widefield microscopy as described above with the following modifications: For widefield, single images of fields of cells were imaged using a CFI Super Fluor 10×/0.5 air objective and two sequential acquisition channels, (1) 510/24-nm bandpass excitation, 514-nm dichroic, 542/27-nm emission, and (2) 550/15 nm bandpass excitation, 562-nm dichroic, and 593/46-nm emission. After background subtraction, per-cell average values for the yellow and red channels were plotted, and a linear fit was performed to determine the slope corresponding to the relative in cellulo brightness for each yellow FP.
Evaluating FP cytotoxicity
EGFP, mRuby1, mCherry, mVenus, mGold, mGold2s, and mGold2t were subcloned to the pCaggs backbone and were transfected separately in HeLa cells using Effectene (catalog no. 301425, Qiagen). EGFP, mRuby1, mCherry, mVenus, and mGold were used as controls to assess FP toxicity. HeLa cells were plated in 6-well cell culture plates at 30–40% confluency. A transfection mixture of 2 μg of plasmid, 16 μL of enhancer solution, and 20 μL of Effectene reagent was added for each well. Two days after transfection, cells were detached with the trypsin-EDTA solution. Cells transfected with EGFP plasmids were mixed with cells transfected with other FP-plasmids at a 1:1 ratio. This step generated 6 mixed cultures (i.e., EGFP & mRuby1, EGFP & mCherry, EGFP & mVenus, EGFP & mGold, EGFP & mGold2s, and EGFP & mGold2t). For each mixed culture, a subpopulation was sampled using a flow cytometer (Attune NxT, Thermo Fisher Scientific) to determine the proportion of cells expressing various FPs. This flow cytometer was equipped with 405-nm and 488-nm lasers. The 405-nm laser with a 512/25-nm emission filter was used to detect green fluorescence while the 488-nm laser with a 530/30-nm emission filter was used to detect yellow fluorescence. Red fluorescence was detected using a 561-nm laser with a 620/15-nm emission filter. The remaining mixed cultures were plated on a 6-well plate and cultured for three additional days. 5 days post-transfection, mixed cultures were detached, and the proportions of cells expressing various FPs were analyzed again. Cytotoxicity was calculated using the formula:
Visualizing subcellular localization
HeLa cells were seeded in glass-bottom 96-well plates. We did not coat the wells with PLL because we noticed PLL caused irregular cell morphology for HeLa cells. For each well, a transfection mixture composed of 50 ng FP-localization plasmid, 50 ng of buffer plasmid, 0.3 μL of FuGENE, and 13 μL of Opti-MEM was prepared and added to the well. FP-localization plasmids used for this experiment is mentioned in the Plasmid construction section. In the following morning, the culture media was replaced with fresh media to minimize transfection toxicity. Two days post-transfection, cells were washed twice with PBS and then incubated with 100 μL of 4% (v/v in PBS) paraformaldehyde at room temperature to fix the cells. Cells were gently washed thrice using PBS and resuspended in the HBSS-HEPES imaging solution. Images were acquired using a laser-scanning confocal microscope (LSM880 with Airyscan, Zeiss) equipped with a 40 × 1.1-NA water immersion objective (LD C-Apochromat Korr M27, Zeiss), and 32-channel GaAsP detector (Airyscan, Zeiss). A 488-nm argon laser (LGK7812, Lasos) at 3% power was scanned with a per-pixel dwell time of 2 μs. Emission light was filtered using a multipass beamsplitter (MBS 488/561/633, Zeiss) and acquired with a detector gain of 740 and 1-Airy unit pinhole size.
Characterizing in cellulo maturation time delay
To determine the maturation time delay of each FP. We adopted a protocol described in ref. 49. pJL2 plasmids co-expressing an FP-of-interest, and EBFP2 was used. FPs-of-interest included mVenus, mGold, mGold2s and mGold2t. We also included FPs with known maturation times: mNeonGreen, mClover3, and mOrange as controls. HeLa cells were plated on a 24-well glass bottom plate without PLL coating. Cells were transfected with transfection mixtures of 300 ng of plasmids, 0.9 μL of Fugene, and 13 μL of Opti-MEM. Three hours after transfection, the imaging plate was placed on the microscope. The hardware setup for this microscope was the same as the one used in the photobleaching experiments, except this microscope was equipped with a stage-top incubator (H301, Okolab). The incubator maintained the cells at 37 °C, 5% CO2, and humidity control. YFP, GFP and BFP images were recorded every 30 min for 72 hours (YFP: 520/5-nm light at 2.4 mW/mm2, 10 ms exposure; GFP: 470/2.5-nm light at 6.1 mW/mm2, 10 ms exposure; BFP: 405/0.6-nm light at 4.8 mW/mm2, 30 ms exposure). Cells were segmented and tracked using Cellpose8 and TrackMate7, and their fluorescence intensities were measured over time. Around 300 cells per construct that showed a sigmoidal increase of YFP and BFP intensities were manually selected. We excluded dividing or apoptotic cells that showed discontinuous time traces. The following sigmoidal function was used to fit the fluorescence (F) versus time (t) curve for each cell:
where the initial parameter estimates were defined as the following: a was set to max(F), b was set to 1, and c was set to median(t). These estimates were tuned to fit the sigmoidal curve to actual data. Using the fitted curve, we determined the inflection point of the curve. A linear tangential line that touched the inflection point was used to determine the x-intercept of the tangential line. The x-intercept values for both FP-of-interest and EBFP2 were determined for each cell, and the difference between the two x-intercepts was defined as the maturation delay time.
Estimating in vitro maturation times
We generated a standard curve (Supplementary Fig 9) to convert in cellulo maturation time delay to maturation times. As standards, we used existing FPs with known in vitro maturation times (i.e., mNeonGreen, 10 min; mVenus, 18 min; mClover3, 44 min; and mOrange, 150 min). Their in cellulo maturation time delays were determined using the procedure described above. We plot the log of maturation times versus maturation time delays and fit a linear standard curve. After determining the maturation delay of mGold, mGold2, and mGold2t, we used the standard curve to convert the maturation delay values to in vitro maturation times.
Determining in cellulo protein degradation rates
We conducted cycloheximide (CHX)-chase assay to determine the protein degradation rates of our YFP variants. We used the same microscopy setup described in the Characterizing in cellulo maturation time delay section. HeLa cells were transfected with the pJL2 plasmids in 24-well plates as described above. Forty-eight hours after transfection, 50 ug/mL CHX (catalog no. C7698-1G, Sigma-Aldrich) to inhibit translation. Cells were imaged immediately after CHX addition for 6 h. Images were analyzed using TrackMate7 and Cellpose to track the changes in fluorescence during imaging. We used low excitation power levels and low temporal resolution (imaging once every 15 min) to prevent photobleaching.
Measuring the OSER Score
HeLa cells were transfected with 1 µg of CytERM-FP plasmid using Effectene (catalog no. 301425, Qiagen) in a glass-bottom 6-well plate. A day after transfection, cells were imaged using a confocal microscope (LSM880 with Airyscan, Zeiss) with a 40 × 1.4-NA oil immersion objective (Plan-Apo DIC M27, Zeiss). Images were manually analyzed, and the percentage of transfected cells without whorl structures was defined as the OSER score.
Determining 2P spectra
mVenus, mGold, mGold2s, mGold2t, EGFP, mStayGold(J), and mBaoJin were subcloned to the pCaggs backbone and were transfected separately in HEK293A cells. Two days after transfection, the cells were washed and imaged in an external solution composed of 110 mM NaCl (catalog no. S3014, Sigma-Aldrich), 26 mM sucrose (catalog no. S0389, Sigma-Aldrich), 23 mM glucose (catalog no. G8270, Sigma-Aldrich), 5 mM KCl (catalog no. P9541, Sigma-Aldrich), 2.5 mM CaCl2 dihydrate (catalog no. 223506, Sigma-Aldrich), and 1.3 mM MgSO4 (catalog no. M2643, Sigma-Aldrich), and was adjusted to pH 7.4. Imaging was performed using the microscope described above (see the General hardware setup section). Images were taken using wavelengths from 700 to 1080 nm with 10 nm increments. The laser power was 10 mW for all wavelengths. Images were background-corrected and manually segmented. The mean fluorescence of cells at each wavelength was calculated and plotted. We imaged at 970 nm with regular intervals during the spectral scanning and confirmed minimal photobleaching during the scanning. Small deviations in the actual power from the target power of 10 mW were corrected by assuming a quadratic dependence of fluorescence on illumination power.
Determining 2P photostability
To characterize the photostability of the YFPs under two-photon, mVenus, mGold, mGold2s, mGold2t, EGFP, mStayGold(J) and mBaoJin were subcloned pJL2-YFP/GFP-P2A-EBFP2 plasmid and transfected separately in HEK293A cells. Two days after transfection, the 96-well plate medium was removed, and cells in each well were washed with 200 µL external solution twice. After the wash, 100 µL of external solution was added to each well for imaging. Imaging was performed using the microscope described above (see the General hardware setup section). Photobleaching was conducted under 950 nm excitation because the YFP and GFP variants are excited ~80% of their respective peaks (Supplementary Fig. 8a). We photobleached at different power levels: 24, 48, 72, 96, 120, and 140 mW at the sample plane. FOVs of 512 × 128 pixels were continuously photobleached and imaged at a frame rate of 110 Hz for 90 s. The emission light from the cell was split using a 560-nm dichroic mirror (348958, Chroma), filtered by a 525/50-nm bandpass filter (353716, Chroma) and collected by a gallium arsenide phosphide (GaAsP) photomultipliers tube (PMT). The videos were first background-corrected, and individual cells were segmented. For each cell, fluorescence values at different time points were normalized to the first frame. The mean fluorescence value was determined for each construct.
Determining 2P brightness
To characterize the brightness of the YFPs and GFPs under 2P, HEK293A cells were prepared as described in the section above. An excitation laser of 950 nm (for YFPs) or 800 nm (for EBFP2) was generated from a titanium:sapphire femtosecond laser and directed to the sample plane by the resonant galvanometer scanners through the same 20 × 0.75 NA objective (CFI Plan Apochromat Lambda, Nikon Instruments). The emission light from the cell was split using a 560-nm dichroic mirror (348958, Chroma), filtered by a 525/50-nm (353716, Chroma) and a 440/80-nm (Chroma) bandpass filter for the yellow and blue channel, respectively, and collected by gallium arsenide phosphide (GaAsP) photomultipliers tubes (PMTs). FOVs of 1024×1024 pixels were captured. The power levers for 950 nm and 800 nm lasers were 14.5 mW and 61.4 mW, respectively. For each FOV, images were first background-corrected and segmented to analyze individual cells. The YFP/BFP score was computed for individual cells as the yellow channel’s mean fluorescence divided by the blue channel’s mean fluorescence. The brightness of each well is calculated as the mean YFP/BFP score.
Determining photostability using laser-scanning confocal
HEK293A cells transiently expressing YFPs were prepared as described above. Photobleaching was conducted using a high-speed confocal microscope (LSM880 with Airyscan, Zeiss) using a 40× 1.4-NA oil immersion objective (Plan-Apochromat DIC M27, Zeiss). Photobleaching was conducted for 300 s by unidirectional scanning of a 53.14 μm by 53.14 μm area at 3.18 Hz with a 514 nm argon laser (LGK7812, Lasos) at 5% power (14 μW), 10% power (30 μW) and 20% power (57 μW), and a pixel dwell time of 1.02 μs. Pixels (512 × 512) with an image depth of 12 bits were acquired continuously during photobleaching using a pinhole size of 500 μm. Images were analyzed using ImageJ50 by manually drawing masks around the cells and tracking the fluorescence of individual cells over time.
In vitro characterization of fluorescent proteins
FPs isolation
YFPs and other control RFPs were expressed in E. coli (DE3) pLysS strain (catalog no. L1195, Promega). Single colonies were picked from fresh LB agar and grown in 5 mL LB, supplemented with 100 µg/mL ampicillin (catalog no. BP1760-25, Thermo Fisher Scientific) overnight at 30 °C. Bacteria were large-scale inoculated with 250 mL of LB/ampicillin until OD600 ~ 0.2 and induced with 1 mM isopropyl β-d-1-thiogalactopyranoside (IPTG, catalog no. 11411446001, Roche) for 6 h at 30 °C. Bacterial pellets were lysed and isolated using the Ni-NTA Fast Start Kit (catalog no. 30600, Qiagen) and immediately dialyzed into 5 or 50 mM Tris-HCl, pH 7.5. FP concentrations were determined by the Pierce BCA protein assay kit (catalog no. 23225, Thermo Fisher Scientific), using a bovine serum albumin (BSA) standard as a reference.
Emission and excitation spectra
FPs were diluted in 50 mM Tris-HCl, and 100 µL of samples were loaded into 96-well glass bottom plates with 6 technical replicates. Excitation and emission spectra were calculated on Cytation5 plate reader (BioTek). Briefly, excitation scan data was collected using excitation wavelengths ranging from 400 to 555 nm and emission measured at 580/10 nm. Emission scan data were collected using an excitation wavelength of 475/10 nm and emission wavelengths ranging from 500 to 700 nm. Measurements were also taken for a control well containing only 50 mM Tris buffer without FPs. Excitation and emission curves were generated by subtracting the control measurements from each sample, normalizing the maximum fluorescence intensity of each well to 1, and averaging the resulting curves across the replicates for each FP.
Oligomeric state
Purified and dialyzed YFPs and RFPs were diluted into flow buffer (50 mM Tris-HCl, pH 7.5, and 100 mM NaCl). Size exclusion chromatography (SEC) was conducted to determine the oligomeric state of mGold2s and mGold2t. 100 μL of 10 µM sample was injected separately into the gel filtration column (Superdex 200 10/300 GL, GE Healthcare) at a flow rate of 0.5 mL/min for a total run time of 24 min in the same flow buffer. Fluorescence of the eluted protein was detected with absorbance at 515 nm for YFPs, and 554 nm and 587 nm for tdTomato and mCherry, respectively. tdTomato (tandem dimer), mCherry (monomer), and mVenus (monomer) were used as size standards.
Extinction coefficient
To determine the extinction coefficient, we adapted the protocol from ref. 49. This protocol assumes that alkali-denatured avGFP-type chromophores have a peak absorbance around 430–460 nm with an extinction coefficient of 44,000 M−1cm−1. Purified FPs were diluted in 10 mM Tris-HCl, pH 7.4, in a 5 mL glass bottle. FP solution was loaded into a 1-cm quartz cuvette (catalog no. 1702503, Bio-Rad), and the absorbances of the intact native state were measured by scanning from 400 nm to 600 nm with 1 nm increments. The peak absorbance value was noted. We also measured blank Tris-HCl buffer without FP molecules to subtract background absorbance. We then denatured the FPs by adding 500 μL of 2 M NaOH mixed with 500 μL of FP solution. We then scanned the absorbance between 430 and 460 nm with 1 nm increments. This measurement was done immediately after denaturing the FP. The peak absorbance value of the denatured FP was noted. A blank solution without FP but with NaOH was measured to subtract background absorbance. The extinction coefficients \((\varepsilon )\) of unknown YFPs were determined using the following equation:
where \(\varepsilon\) stands for extinction coefficient, Anative refers to the peak absorbance value of native FP, and Adenatured stands for the peak absorbance value of denatured FP.
Quantum yield
Purified proteins were diluted in 10 mM Tris-HCl, pH 7.4, using 5 mL glass bottles. We used glass bottles because they allowed a more homogenous solution of FPs than plastic vials. We prepared FP solutions with different starting concentrations with absorbance at 490 nm (OD490) of 0.1, 0.09, 0.08, 0.07, and 0.06. FP solutions were serially diluted up to 32-fold to achieve around 20 samples with OD490 values between 0.005 and 0.05. The lower limit was set by the detection limit of the spectrophotometer. The upper limit was set by determining the highest concentration that did not show an inner filter effect. We found that having at least 20 diluted samples for each FP is crucial to determining accurate quantum yield values. The absorbance was recorded with a Biochrom Libra 522 spectrophotometer using 1-cm quartz cuvettes at 490 nm. The fluorescence emission spectra of the diluted samples were measured using the Agilent spectrophotometer (Cary Eclipse Fluorometer). The emission spectrum was recorded using a fixed excitation wavelength at 490 nm. The emission spectrum was scanned from 500 nm to 700 nm at a 200 nm/min scan speed with a step size of 1 nm. The absorbance and emission fluorescence values were background corrected using a black 10 nM Tris-HCl buffer without FPs. After obtaining the emission spectrum, the area under the curve (AUC) was calculated by integrating the emission fluorescence from 500 to 700 nm. The OD490 and AUC values were plotted for each of the diluted samples, and the slope of the best-fit linear line was determined. We repeated the protocol above for a reference fluorophore (Rhodamine 123) with a known quantum yield.
Quantum yields (Q) were calculated using the formula:
where Qreference refers to the quantum yield of Rhodamine 123, which has a value of 0.9 (ref. 51). munkown refers to the slope of the best-fit line for the FP-of-interest, while mreference is the slope of the best-fit line for the reference, Rhodamine 123. nunknown denotes the refractive index of the solvent of the FP solution, with a value of 1.336. nreference is the refractive index of ethanol, the solvent used for rhodamine 123, with a value of 1.366.
pKa measurements
For pH titration, 100 μL, 2 µg of purified YFPs were diluted in 10 mM Tris-HCl, pH 7.5 in wells of 96-well glass-bottom plates containing 100 µL of titration buffer. Titration buffers contained 100 mM citric acid/sodium citrate (pH 3–5.5), 100 mM KH2PO4/Na2HPO4 (pH 6-8), or 100 mM NaOH/Glycine (pH 8.5–10). The Cytation 5 plate reader was then used to determine the emission intensity of each FP at 530/10 nm using 500/10 nm excitation light. Emission intensity versus pH was then normalized to the intensity value at pH 10. The pKa, the pH at which fluorescence is half its maximum, was determined by fitting a sigmoidal curve.
Chloride sensitivity
For the chloride titration assay, 30 μL of purified FPs were diluted to 70 μL MOPS buffer with various chloride concentrations: 0, 0.1, 0.25, 0.4, 0.5, 0.6, 0.8, 0.9, or 1 M KCl. MOPS buffer was prepared by mixing MOPS (20 mM, catalog no. M1254, Sigma-Aldrich), sodium acetate (5 mM, catalog no. S8750, Sigma-Aldrich), and Na2 EDTA (1 mM catalog no. E5134, Sigma-Aldrich) in water; pH was adjusted to 7.0. To keep the ionic strength constant, appropriate amounts of potassium gluconate (catalog no. P1847, Sigma-Aldrich) were added to make the amounts of ions in these solutions the same as that in 1.0 M KCl solution. The Cytation 5 plate reader was used to measure the fluorescence of FPs. The excitation wavelength of 508/10 nm and emission wavelength of 530/10 nm were used. Blank buffers were prepared to subtract the background fluorescence. The corrected fluorescence values were normalized to that of the 0 M KCl sample. Kd for Cl- was calculated as the concentration of Cl-, when fluorescence intensity reached half its initial value.
Fluorescence lifetime
Purified FPs were diluted 20-fold in PBS (at pH 7.4). Fluorescence lifetime measurements were performed using a Nikon A1 Confocal Laser Microscope coupled with PicoQuant (PicoQuant GmbH) and 439 nm (121 mW peak power) and 483 nm (136 mW peak power) lasers. The microscope was operated with SymPhoTime 64 software. Detected photons were counted by a time-corrected single-photon counter (PicoHarp 300, PicoQuant GmbH).
Erythrosine B dye in KI (τ = 0.2 ns) was used as a reference standard to obtain the instrumental response function. The frequency of modulation was 20 MHz. The laser power was adjusted to achieve ~10,000 counts per second at the peak value. YFPs were excited with 488 nm using a 20× Plan-Apo/0.75 NA objective. A FF01-520/35 fluorescence filter was used in the PicoQuant detector. The image resolution was fixed at 512 × 512 pixels and 1.0 airy unit (AU) pinhole size.
Thermostability
To measure the thermostability of the YFP variants, we utilized the Differential Scanning Fluorimetry Thermal Cycler (Roche). We loaded around 4 pmol of purified FP sample per replicate. The temperature was ramped up from 30 °C to 99 °C at 0.9 °C per min, and fluorescence was measured around every 1.7 s. FM filter was used, which had 470/15 nm excitation and 520/15 nm emission filters. Thermostability was determined by calculating the inflection point of the fluorescence versus temperature melting curve.
Application of mGold2s and mGold2t
Super-resolution imaging of actin and mitochondria
COS-7 cells were transfected with plasmids using Lipofectamine 2000 reagent (catalog no. 11668027, Thermo Fisher Scientific). For the TIRF-SIM and 2D-SIM imaging experiments, cells were seeded onto coverslips (H-LAF 10 L glass; reflection index, 1.788; diameter, 26 mm; thickness, 0.15 mm, customized). Before seeding transfected cells, coverslips were coated with 0.01% poly-l-Lysine solution (catalog no. P4707, Sigma-Aldrich) for 10 min and washed twice with sterile water.
Image acquisition of SIM was obtained by HIS-SIM (Guanzhou CSR Biotech Co. Ltd) based on Hessian SIM system52. HIS-SIM was equipped with the objectives (×100/1.7 HI oil, APON, Olympus). Images were captured by an sCMOS camera (Flash 4.0 V3, Hamamatsu). For actin imaging, we utilized TIRF-SIM mode with 95 mW/mm2 488 nm laser power and 5 ms exposure time. For mitochondria imaging, 2D-SIM mode was used with 58 mW/mm2 488 nm laser power and 5 ms exposure time. To further improve the quality of reconstructed images, sparse deconvolution was used to process the SIM images32.
Laser-scanning confocal imaging of gephyrin-FingR in live dissociated neurons
Rat hippocampal cultures were prepared from mixed sex embryonic (E) 21 Long-Evans rats (HsdBlu:LE, Inotiv). Hippocampi were dissected, dissociated with papain (catalog no. LS003127, Worthington Biochemical Corporation) diluted in Hank’s Balanced Solution (catalog no. 14170112, Invitrogen) supplemented with 25 mM MgCl2, 12 mM HEPES, and 1.5 mM Kynurenic acid (catalog no. K3375, Sigma-Aldrich). Cells were then washed with trypsin inhibitor (catalog no. T9253, Sigma-Aldrich), and seeded onto glass bottom plates with high-performance #1.5 cover glass (catalog no. P24-1.5H-N, Cellvis) coated with 20 µg/mL poly-d-lysine (catalog no. 354210, Corning) and 3 µg/mL laminin (catalog no. 354239, Corning) at 2.5 × 105 neurons/mL in Neurobasal medium without Phenol Red (catalog no. 12348017, Invitrogen) supplemented with B27 (catalog no. 17504044, Invitrogen), 2 mM glutamine (catalog no. 25030-081, Thermo Fisher Scientific), and 100 U/mL penicillin/streptomycin (catalog no. 15140122, Thermo Fisher Scientific). The culture medium was changed at a day in vitro (DIV) 1.
Rat hippocampal primary neurons were transfected with mRuby3 (Addgene plasmid # 127808) and Gephyrin-FingR-YFP variants on DIV 4 using the calcium phosphate transfection method53. Confocal live imaging experiments were conducted between DIV 20-22 at 37 °C and with 5% CO2 being supplied. Zeiss LSM 880 microscope was in confocal mode. Objective was plan-Apochomat 63×/1.4 Oil DIC M27. Filter set was MBS 458/514/561/633. Lasers used include an Argon laser for the 514 nm line and a DPSS 561-10 laser for the 561 nm line. Pixel dwell time was 1.45 µs, with a pixel size of 0.09 µm. Images were acquired in line scan mode using pinhole (51 mm), time series, and Z-stacks of 0.38 μm step size. Bleaching of the mVenus, mGold, and mGold variants was achieved using the 514 nm laser set to 2% laser power (11 µW). Time series were analyzed using ImageJ.
Widefield imaging of YR-ICUE FRET indicators
To compare the photostability of the YR-ICUE FRET indicators, we transfected the YR-ICUE plasmids into HEK293A cells using FuGENE (catalog no. E2311, Promega) following the manufacturer’s instructions. Briefly, 100 ng of plasmids (50 ng of YR-ICUE plasmid and 50 ng of buffer plasmid) was mixed with 0.3 µL of FuGENE reagent in 13 µL of Opti-MEM (catalog no. 31985070, Thermo Fisher Scientific) per well of a 96-well plate. Cells were plated in a 96-well plate precoated with PLL. Two days post-transfection, cells were imaged with the 1P widefield setup (Nikon Ti-E) mentioned in the general hardware setting section. YFP photobleaching was conducted under continuous 520/5-nm illumination at 17 mW/mm2 for 10 minutes. YFP images were taken every 5 s using 520/5-nm light at 8.5 mW/mm2 with 20 ms exposure using the 520/60 nm emission filter. RFP images were taken every 5 s using the 555/0.5-nm light at 36 mW/mm2 with 30 ms exposure using the 620/60 nm emission filter. Image analysis was conducted in the same way as the YFP photobleaching characterization experiments mentioned above.
To characterize the response ratio of these indicators, cells were seeded on poly-D-lysine-coated 35 mm glass-bottom imaging dishes (catalog no. D35-14-1.5-N, Cellvis) and grown to 50–70% confluency. Cells were subsequently transfected using Lipofectamine 2000 and sensor plasmid DNA (400 ng) and incubated for an additional 24 hours before imaging.
Cells were pre-incubated for 20 min with 500 nM nuclear acid stain Hoescht 33342 to facilitate automated cell identification during analysis. Cells were washed once, and then imaged with Hank’s balanced salt solution (HBSS, Gibco 14065-056; buffered with 20 mM HEPES, pH 7.4 and supplemented with 2 g/L glucose) with imaging performed at room temperature. Isoproterenol (Iso, 1 μM, catalog no. PHR2722, Sigma-Aldrich), forskolin (Fsk, 50 μM, catalog no. F6886, Sigma-Aldrich), and 3-isobutyl-1-methylxanthine (IBMX, 100 μM, catalog no. I5879, Sigma-Aldrich) were added at indicated times.
Time-lapse epifluorescence images were acquired on a Zeiss AxioObserver Z7 microscope with a 40×/1.4 NA objective and a Photometrics Prime95B sCMOS camera. Images were acquired every 30 s over a 30-min period with an exposure time of 500 ms for YFP direct and YR-FRET channel, 100 ms for RFP direct channel, and 10 ms for DAPI channel. The following excitation/emission filter combinations and dichroic mirrors were used: YFP EX-495/10x, EM-535/25m, DC-T515p; YR-FRET EX-495/10x, EM-645/75m, DC-ZT594rdc; RFP EX-572/35x, EM-645/75m, DC-ZT594rdc; DAPI EX-ET380x, EM-473/24m, DC-T455lpxru. All filter sets were controlled by an external filter exchanger (Ludl Electronic Products, Ltd).
FRET image analysis was performed using Cellpose48 and custom MATLAB scripts similar to those reported previously54. In brief, maximum intensity projections of RFP direct and DAPI channels were used with Cellpose for segmentation and automated selection of transfected cells present for the duration of each time-lapse. Segmentation-based masks were then used on background-subtracted YFP direct and YR-FRET channels to get single-cell ratiometric response curves that were subsequently normalized to the ratio before isoproterenol addition. The response ratio (∆R/R0) was calculated using the equation:
where R is the maximum normalized YFP/YR-FRET value obtained upon isoproterenol addition, and R0 is the ratio before isoproterenol addition. Pseudocolor images were generated using custom macros in ImageJ. In brief, a ratiometric image of the YFP direct channel divided by the YR-FRET channel was used to define hue in an hue, saturation, and value (HSV) image that was intensity-modulated using the RFP direct channel.
Single-molecule pull-down assay
HEK293 cells were transfected individually with mEYFP, mGold, mGold2s, mGold2t and mVenus followed by cell lysed in lysis buffer (40 mM Hepes pH 7.5, 120 mM NaCl, 10 mM sodium pyrophosphate, 10 mM B-glycerolphosphate, 1X cOmpleteTM protease inhibitor mixture and 0.3% CHAPS). Lysates were centrifuged at 12,000 g for 12 min at 4 °C.
We used a prism total internal reflection fluorescence (TIRF) microscope equipped with an electron-multiplying charge-coupled device (EM-CCD) for single-molecule imaging. Quartz slides and glass coverslips passivated with methoxy polyethylene glycol (PEG) containing biotinylated PEG were purchased from Nano Surface Sciences. Rabbit polyclonal biotinylated anti-GFP antibody (Rockland, item #600-406-215) was immobilized at a concentration of ~10 nM on the NeutrAvidin coated PEG surface. Cell lysates were diluted to obtain a surface density optimal for single-molecule analysis using dilution buffer (10 mM Tris-HCl pH 7.5, 200 mM NaCl, 20 mM MgCl2, 10% Glycerol, 0.2 mg/mL BSA), and pulled down over the antibody surface. Immobilized fluorescent proteins were excited at 488 nm and 50 frames were recorded per imaging area. A total of 10 short movies were taken using 100 ms exposure time. Image analysis was conducted using a custom code reported previously50,55. Mean spot-counts per image and relative fluorescence intensity experiments were performed at 1 mW laser power, measured when entering the TIRF prism. Single-molecule peaks were selected by fitting a Gaussian profile on the average intensity of the first ten frames. Relative intensity was calculated by subtracting the midpoint intensity value of each molecule after bleaching. Experiments were performed at room temperature (22-25 °C). Photobleaching lifetime data was acquired from three long movies (100 s) taken using 100 ms exposure time in different regions. The fluorescence time of single-molecule traces was determined by fitting 1−CDF (cumulative distribution function) to a single-exponential decay function. Bleaching experiments were performed at different laser power (530 μW, 1 mW, 2 mW). Each fluorescence trajectory was manually classified into three categories: no photoblinking events, photoblinking and poor-quality trace (low intensity and/or signal-to-noise ratio).
Statistics and reproducibility
Biological replicates were defined as independent transfections for mammalian cells or separate cultures grown in separate culture vessels for yeast. Typically, we used 3–9 biological replicates analyzing approximately hundreds of mammalian cells and thousands of yeast cells per replicate. For experiments requiring measurements of intracellular regions of interest (ROIs, such as foci), biological replicates were defined as individual cells. For experiments involving purified FPs, biological replicates were defined as independently purified FPs obtained from separate bacterial cultures. Experiments were repeated at least once on a separate day to account for day-to-day variability. Sample sizes were determined based on prior studies and established standards in the field. No statistical method was used to predetermine the sample size.
Statistical tests appropriate to the data and their underlying assumptions were applied for analysis. Briefly, for statistical analysis with a sample size greater than 30, parametric tests were performed without assessing normality, as violations of normality have minimal impact on parametric analysis with large sample sizes (65). For sample sizes of 30 or fewer, normality was evaluated using the Shapiro-Wilk test. Parametric analyses were conducted using two-sided analysis of variance (ANOVA) or t-tests, while the Mann-Whitney test was used for nonparametric analyses. The Brown-Forsythe test (for ANOVAs) or the F-test (for t-tests) was used to determine whether Welch’s correction was necessary to address variance differences. If significant variance differences were detected, Welch’s correction was applied. Statistical analyses on trends (e.g., normalized fluorescence versus time) were performed using areas under the curve (AUCs). All analyses were two-sided, with a significant level set at 0.05. For quantitative experiments, blinding was unnecessary because the collection and analysis steps that could be subject to bias were performed using pre-defined automated methods. For the OSER assay that required qualitative manual assessment of OSER structures, we blinded the analysis so that the person evaluating the images did not know which construct the cells were expressing. Data were not omitted unless obvious human errors in sample preparation or data acquisition justified exclusion.
Details of the statistical analyses performed for each figure are provided separately in Supplementary Information. Prism (version 10.3.1, GraphPad) was used to conduct statistical analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
mGold2s and mGold2t DNA and protein sequences are available from GenBank (accession numbers PP826965 and PP826966, respectively). Key characteristics of mGold2s and mGol2t are summarized in FPbase (accession codes CGRMD and HUE6V, respectively). Plasmids used for in vitro characterization of mGold2s and mGold2t are available from Addgene (plasmid #231761 - #231790). Source data are provided with this paper in the Source Data file. Selected raw image data generated in this study is available in Zenodo with the following DOIs: actin dynamics videos (https://zenodo.org/records/14743784), mitochondria dynamics videos (https://zenodo.org/records/14744163), Gephyrin-FingR videos (https://zenodo.org/records/14744239). Other raw image data is available upon request with no restrictions. We cannot publicly store all raw image data due to the absence of repositories capable of cost-effectively handling the 4.2 TB of data generated in this study. Source data are provided with this paper.
Code availability
The data was analyzed using established methods and code as previously reported (see Methods section). Custom scripts used to process the raw data will be made available upon request alongside the raw datasets with no restrictions.
References
Zhang, Y. et al. Fast and sensitive GCaMP calcium indicators for imaging neural populations. Nature 615, 884–891 (2023).
Lodovichi, C., Ratto, G. M., Trevelyan, A. J. & Arosio, D. Genetically encoded sensors for Chloride concentration. J. Neurosci. Methods 368, 109455 (2022).
Liu, Z. et al. Sustained deep-tissue voltage recording using a fast indicator evolved for two-photon microscopy. Cell 185, 3408–3425.e29 (2022).
Bando, Y., Grimm, C., Cornejo, V. H. & Yuste, R. Genetic voltage indicators. BMC Biol 17, 71 (2019).
Yang, H. H. & St-Pierre, F. Genetically Encoded Voltage Indicators: Opportunities and Challenges. J. Neurosci. 36, 9977–9989 (2016).
Mahon, M. J. pHluorin2: an enhanced, ratiometric, pH-sensitive green florescent protein. Adv. Biosci. Biotechnol. 2, 132–137 (2011).
Li, S. A. et al. Progress in pH-Sensitive sensors: essential tools for organelle pH detection, spotlighting mitochondrion and diverse applications. Front. Pharmacol. 14, 1339518 (2024).
Lambert, T. J. & Waters, J. C. Navigating challenges in the application of superresolution microscopy. J. Cell Biol. 216, 53–63 (2016).
Schirripa Spagnolo, C. & Luin, S. Choosing the Probe for Single-Molecule Fluorescence Microscopy. Int. J. Mol. Sci. 23, 14949 (2022).
Jain, A. et al. Probing cellular protein complexes using single-molecule pull-down. Nature 473, 484–488 (2011).
Hirano, M. et al. A highly photostable and bright green fluorescent protein. Nat. Biotechnol. 40, 1132–1142 (2022).
Ando, R. et al. StayGold variants for molecular fusion and membrane-targeting applications. Nat. Methods 21, 648–656 (2024).
Zhang, H. et al. Bright and stable monomeric green fluorescent protein derived from StayGold. Nat. Methods 21, 657–665 (2024).
Ivorra-Molla, E. et al. A monomeric StayGold fluorescent protein. Nat. Biotechnol. 42, 1368–1371 (2024).
Tutol, J. N. et al. Engineering the ChlorON Series: Turn-On Fluorescent Protein Sensors for Imaging Labile Chloride in Living Cells. ACS Cent. Sci. 10, 77–86 (2024).
Ni, Q. et al. Signaling diversity of PKA achieved via a Ca2 + -cAMP-PKA oscillatory circuit. Nat. Chem. Biol. 7, 34–40 (2011).
Ponsioen, B. et al. Detecting cAMP-induced Epac activation by fluorescence resonance energy transfer: Epac as a novel cAMP indicator. EMBO Rep. 5, 1176–1180 (2004).
Cohen, S., Valm, A. M. & Lippincott-Schwartz, J. Multispectral live-cell imaging. Curr. Protoc. Cell Biol. 79, e46 (2018).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Cai, D., Cohen, K. B., Luo, T., Lichtman, J. W. & Sanes, J. R. Improved tools for the Brainbow toolbox. Nat. Methods 10, 540–547 (2013).
Lee, J. et al. Versatile phenotype-activated cell sorting. Sci. Adv 6, eabb7438 (2020).
Bindels, D. S. et al. mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 14, 53–56 (2017).
Shaner, N. C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat. Methods 5, 545–551 (2008).
Campbell, B. C. et al. mGreenLantern: a bright monomeric fluorescent protein with rapid expression and cell filling properties for neuronal imaging. Proc. Natl. Acad. Sci. 117, 30710–30721 (2020).
Goedhart, J. et al. Structure-guided evolution of cyan fluorescent proteins towards a quantum yield of 93%. Nat. Commun. 3, 751 (2012).
Corish, P. & Tyler-Smith, C. Attenuation of green fluorescent protein half-life in mammalian cells. Protein Eng 12, 1035–1040 (1999).
So, P. T., Dong, C. Y., Masters, B. R. & Berland, K. M. Two-photon excitation fluorescence microscopy. Annu. Rev. Biomed. Eng. 2, 399–429 (2000).
Cormack, B. P., Valdivia, R. H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Molina, R. S. et al. Blue-Shifted Green Fluorescent Protein Homologues Are Brighter than Enhanced Green Fluorescent Protein under Two-Photon Excitation. J. Phys. Chem. Lett. 8, 2548–2554 (2017).
Drobizhev, M., Makarov, N. S., Tillo, S. E., Hughes, T. E. & Rebane, A. Two-photon absorption properties of fluorescent proteins. Nat. Methods 8, 393–399 (2011).
Balleza, E., Kim, J. M. & Cluzel, P. Systematic characterization of maturation time of fluorescent proteins in living cells. Nat. Methods 15, 47–51 (2018).
Zhao, W. et al. Sparse deconvolution improves the resolution of live-cell super-resolution fluorescence microscopy. Nat. Biotechnol. 40, 606–617 (2022).
Mo, Y. et al. Quantitative structured illumination microscopy via a physical model-based background filtering algorithm reveals actin dynamics. Nat. Commun. 14, 3089 (2023).
Gross, G. G. et al. Recombinant probes for visualizing endogenous synaptic proteins in living neurons. Neuron 78, 971–985 (2013).
Aye-Han, N.-N., Allen, M. D., Ni, Q. & Zhangy, J. Parallel tracking of cAMP and PKA signaling dynamics in living cells with FRET-based fluorescent biosensors. Mol. Biosyst. 8, 1435–1440 (2012).
Jain, A., Liu, R., Xiang, Y. K. & Ha, T. Single-molecule pull-down for studying protein interactions. Nat. Protoc. 7, 445–452 (2012).
Dean, K. M. et al. Microfluidics-Based Selection of Red-Fluorescent Proteins with Decreased Rates of Photobleaching. Integr. Biol. Quant. Biosci. Nano Macro 7, 263–273 (2015).
Bajar, B. T. et al. Improving brightness and photostability of green and red fluorescent proteins for live cell imaging and FRET reporting. Sci. Rep. 6, 20889 (2016).
Chapagain, P. P., Regmi, C. K. & Castillo, W. Fluorescent protein barrel fluctuations and oxygen diffusion pathways in mCherry. J. Chem. Phys. 135, 235101 (2011).
Zhong, S., Navaratnam, D. & Santos-Sacchi, J. A Genetically-Encoded YFP Sensor with Enhanced Chloride Sensitivity, Photostability and Reduced pH Interference Demonstrates Augmented Transmembrane Chloride Movement by Gerbil Prestin (SLC26a5). PLoS ONE 9, e99095 (2014).
Zheng, J., Guo, N., Huang, Y., Guo, X. & Wagner, A. High temperature delays and low temperature accelerates evolution of a new protein phenotype. Nat. Commun. 15, 2495 (2024).
Icha, J., Weber, M., Waters, J. C. & Norden, C. Phototoxicity in live fluorescence microscopy, and how to avoid it. Bioessays News Rev. Mol. Cell. Dev. Biol. 39, 1700003 (2017).
Gietz, R. D. & Schiestl, R. H. Frozen competent yeast cells that can be transformed with high efficiency using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 1–4 (2007).
Bajar, B. T., Wang, E. S., Zhang, S., Lin, M. Z. & Chu, J. A guide to fluorescent protein FRET pairs. Sensors 16, 1488 (2016).
Mo, G. C., Posner, C., Rodriguez, E. A., Sun, T. & Zhang, J. A rationally enhanced red fluorescent protein expands the utility of FRET biosensors. Nat. Commun. 11, 1848 (2020).
Gietz, R. D. & Schiestl, R. H. Quick and easy yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 35–37 (2007).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Gadella, T. W. J. et al. mScarlet3: a brilliant and fast-maturing red fluorescent protein. Nat. Methods 20, 541–545 (2023).
Liao, T.-W. et al. Linking folding dynamics and function of SAM/SAH riboswitches at the single molecule level. Nucleic Acids Res. 51, 8957–8969 (2023).
Kubin, R. F. & Fletcher, A. N. Fluorescence quantum yields of some rhodamine dyes. J. Lumin. 27, 455–462 (1982).
Huang, X. et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nat. Biotechnol. 36, 451–459 (2018).
Duman, J. G. et al. The adhesion-GPCR BAI1 regulates synaptogenesis by controlling the recruitment of the Par3/Tiam1 polarity complex to synaptic sites. J. Neurosci. 33, 6964–6978 (2013).
Schmitt, D. L. et al. Spatial regulation of AMPK signaling revealed by a sensitive kinase activity reporter. Nat. Commun. 13, 3856 (2022).
Singh, N. et al. Redefining the specificity of phosphoinositide-binding by human PH domain-containing proteins. Nat. Commun. 12, 4339 (2021).
Acknowledgements
We thank the following cores at Baylor College of Medicine for equipment and expertise: the Cytometry and Cell Sorting Core (CCSC) for FACS sorting, the Optical Imaging & Vital Microscopy Core for confocal microscopy, the Integrated Microscopy Core at BCM for fluorescence measurements with plate readers, the Recombinant Protein Production and Characterization Core for conducting the thermostability assay. We thank the Center of Advanced Microscopy Core at the University of Texas Health Science Center for fluorescence lifetime microscopy and the Shared Equipment Authority at Rice University for fluorometry. We also thank Paul Leonard at MD Anderson for conducting the size exclusion chromatography; Carlos Llanos and Dr. Laura Segatori at Rice University for providing a spectrophotometer; Michelle Lee at Rice University for assisting with SPOTlight screening. F.S.-P. and N.C.S. were supported by the National Institute of Health (NIH) grant R01EB032854. F.S.-P. was also supported by a Vivian L. Smith Endowed Professorship, Welch Foundation grant (Q-2016-20220331), NIH grants (U01NS133971, U01NS133657, U01NS118288), National Science Foundation (NSF) grants (1935265 and 1935265), the McNair Medical Foundation, and the Klingenstein-Simons Fellowship Award in Neuroscience. J.Z. was supported by NIH grants (R35 CA197622, R01 DK073368). A.C.L was supported by the NSF Graduate Research Fellowship (DGE-2038238) and the American Heart Association Predoctoral Fellowship (24PRE1186687). L.C. was supported by grants from the National Natural Science Foundation of China (32227802, T2288102, and 8195022) and New Cornerstone Science Foundation. T.H. was supported by NSF grant EF1934864 and NIH grant R35GM122569. K.F.T.’s lab was supported by NIH grant RF1NS062829. F.A.B. was supported by NIH fellowship F31 NS122427. The CCSC is supported by the Cancer Prevention and Research Institute of Texas (CPRIT-RP180672) and the NIH (P30 CA125123 and S10 RR024574). Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the NSF.
Author information
Authors and Affiliations
Contributions
S.Y., S.Z., F.A.B., A.C.L., and R.M.U. contributed equally to this work as co-second authors. Lab-level contributions: This study was a collaborative project among six laboratories. F.S.-P.’s lab conducted SPOTlight screening and FP characterization. N.C.S.’s lab conducted FP characterization cross-lab validation experiments. L.C.’s lab conducted super-resolution imaging. K.F.T.’s lab conducted confocal imaging of FingR constructs. J.Z.’s and F.S.-P.’s labs conducted YR-ICUE characterization experiments. T.H.’s lab conducted single-molecule pull-down assays.
Individual contributions: F.S.-P. conceived, managed and directed the overall project with the help from J.L. and S.L. J.F.A, S.L. and J.L. conducted SPOTlight screening, including experimental design and analysis. S.L., J.L., N.A.N and S.Y. conducted mGold2 characterization experiments, including experimental design and analysis. H.L. and Z.L. developed the image analysis software for SPOTlight screening and mGold2 characterization. S.Z., F.A.B., A.C.L., and R.M.U conducted their respective mGold2 application experiments, including experimental design and analysis. G.G.L. conducted the cross-lab validation experiments, which included experimental design and analysis. Senior authors N.C.S., L.C., K.F.T, J.Z., T.H., F.S.-P. helped design experiments and guided the project. J.L., S.L., S.Y., and F.S.-P wrote the manuscript with significant input from S.Z., F.A.B., A.C.L., R.M.U., G.G.L., J.F.A, H.L., Z.L., L.C., K.F.T, J.Z., N.C.S., and T.H. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interest: and F.S.-P, J.L., and Z.L. have filed a US patent application encompassing the sequence of mGold, the parental protein of mGold2s and mGold2t, the two YFPs reported here. F.S.-P., J.L., and Z.L. have filed a provisional patent for the SPOTlight technology used to screen mGold2s and mGold2t. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Masato Maesako and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, J., Lai, S., Yang, S. et al. Bright and photostable yellow fluorescent proteins for extended imaging. Nat Commun 16, 3241 (2025). https://doi.org/10.1038/s41467-025-58223-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58223-5
This article is cited by
-
A multiplexed assay by self-assembled dual-target responsive DNA hydrogels for efficacy evaluation of immunotherapy
Nature Communications (2025)