Abstract
Proteins exhibit remarkable molecular recognition by dynamically adjusting their conformations to selectively interact with ligands at specialized binding sites. To bind hydrated ligands, proteins leverage amino acid residues with similar water affinities as the substrate, minimizing the energy required to strip water molecules from the hydrophilic substrates. In synthetic receptor design, replicating this sophisticated adaptability remains a challenge, as most artificial receptors are optimized to bind desolvated substances. Here, we show that proline-based synthetic receptors can mimic the conformational dynamics of proteins to achieve selective binding of hydrophilic and amphiphilic fluoride substances in aqueous environments. This finding highlights the critical role of receptor flexibility and strategic hydrophilicity in enhancing ligand recognition and affinity in water. Moreover, it establishes a new framework for designing versatile synthetic receptors with tunable hydrophobicity and hydrophilicity profiles.
Similar content being viewed by others
Introduction
Recognizing molecular species in water is crucial in biological systems and environmental science1,2. Nature itself has showcased sophisticated mechanisms of molecular recognition3,4,5. For instance, protein receptors fold into various configurations and undergo conformational change, creating binding pockets that can selectively accommodate either hydrophilic or hydrophobic substrates6,7,8,9,10. Mimicking this principle, various synthetic receptors have been made to bind hydrophobic specific substrates through strong and multivalent non-covalent interactions established between the receptor and the substrate11,12,13,14,15,16. However, the molecular recognition of hydrophilic substrates in water presents greater challenges17,18. The complexity arises from the need to overcome the formidable interactions between hydrophilic substrates and their surrounding hydration shells. For hydrophilic substrates such as carbohydrates and anions, the common strategy is to design a multivalent binding receptor with strong non-covalent interactions to compensate for the dehydration enthalpy of substrates19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34. It is also preferable for the receptor to possess some flexibility to maximize these non-covalent interactions between the receptor and substrate. Consequently, fewer examples of receptors demonstrate a high affinity for hydrophilic molecules and anions in aqueous solutions34,35,36,37. One significant achievement is a glucose-binding receptor developed by Davis et al.38, which selectively binds D-glucose with a high affinity of 18,000 M−1, despite glucose’s free energy of hydration of 108.8 kJ/mol39. Another notable example is the design of organic cages for the selective binding of highly hydrated sulfate anions25,34.
Design strategies for synthetic receptors have traditionally emphasized offsetting the dehydration energy for molecular and anion binding through multivalent noncovalent interactions40. However, proteins introduce an alternative methodology for recognizing hydrophilic substrates41,42,43,44,45,46,47. Within their binding pockets, proteins employ amino acid residues whose water-binding affinities match those of the substrates. Hydrophilic substrates that are strongly associated with water are matched with amino acids that also exhibit high hydration levels, thereby minimizing the energy required for the complete removal of water molecules from the hydrophilic substrate. However, the application of this principle to partially or entirely retain the solvation shell of hydrophilic substances in synthetic receptor design has been largely overlooked48. For example, Gibb et al. brought attention to this issue by demonstrating the use of a shape-persistent receptor for the binding of partially hydrated anions, although the specifics of the design principles remain elusive49. This limited understanding raises several key questions in the development of receptors capable of binding hydrophilic substrates along with their associated water molecules: Is shape persistence a crucial factor in receptor design? Is a hydrophobic or hydrophilic binding site preferable for binding these hydrophilic substances and their solvation waters? What approach exists for incorporating such a binding pocket into the design?
In pursuit of advancing receptor designs, we sought to develop organic receptors featuring extensive tunability to facilitate the binding of hydrophilic hydrated substrates and amphiphilic substances, such as fluoride anion and PFASs (Per- and Polyfluoroalkyl Substances) (Fig. 1). The recognition of fluoride by synthetic receptors has garnered significant interest due to its high hydration energy of −429 kJ/mol and its implications in various human pathologies50,51. However, only a few examples have successfully demonstrated high-affinity fluoride binding in an aqueous environment. For instance, Reinaud et al. achieved fluoride recognition by using a pentacationic calix [6] arene-based CuII complex52, leveraging metal-ligand coordination and hydrophobic effect. The authors reported a high affinitive binding of partially hydrated F− at pH = 5.9, but the affinity dropped more than 100 times at pH = 6.7. Additionally, most previously reported receptors rely on favorable enthalpic contributions for fluoride binding53. These reports suggested that receptors need to establish robust interactions with fluoride to compensate for the substantial dehydration enthalpy, thereby increasing the complexity of receptor design.
a Graphical illustration of the design principles (different colors represent different modules, as description in the graph). The modular amino acid building blocks provide a rich library for structural variation. The functional pillar allows for the creation of hydrophilic/hydrophobic micro-environments for guest binding. b The chemical structures of organic cages synthesized in this work.
On the other hand, unlike heavily hydrated hydrophilic substances, amphiphilic substances such as surfactants possess a highly hydrated head and a poorly hydrated tail, making them effective as detergents. One notable group of these amphiphilic substances, PFASs, have recently emerged as organic micropollutants, gaining significant attention due to their environmental impact. Various approaches have been developed to remove them from contaminated water, such as metal-organic frameworks (MOFs)54,55, β-cyclodextrin-based polymers (CDPs)56, and ionic fluorogels57,58. However, these adsorbents exhibit notable drawbacks, such as poor water/chemical stability, low natural organic matter (NOM) selectivity, and structures containing metals or potential pollution sources like fluoride, limiting their practical applications in treating contaminated water59. To advance the design of efficient PFAS adsorbents, it is crucial to establish molecular-level binding mechanisms that selectively target and capture these substances, considering their amphiphilic solvation states and highly fluorinated tails.
In this work, our strategy involves the development of tripodal organic cages, wherein we aim to incorporate amino acid components to introduce a spectrum of structural variations. These variations include hydrophobic, hydrophilic, or bulky functional groups (Fig. 1a), aligning with the principles of precision and versatility found in protein engineering. Specifically, our efforts lead to the design and synthesis of five organic cages utilizing glutamic acid (CGlu), proline (CPro-NH2, CPro-Cy-NH2 and CPro-Cy) and alanine (CAla-Cy-NH2) as key elements (Fig. 1b). Through NMR spectroscopy and X-ray diffraction analysis, we show that the proline-based cages exhibit distinctive structural characteristics, allowing them to adopt flexible conformations with a highly hydrated binding pocket, effectively mimicking the behavior of proteins. Notably, the CPro-NH2 cage display excellent binding specificity and high fluoride binding affinity in pure water, making it one of the most effective neutral organic synthetic receptors for fluoride binding. This feature is characterized by an atypical entropy-driven endothermic binding mechanism. Moreover, this cage exhibits a strong affinity towards amphiphilic fluorinated compounds, such as perfluorooctanoic acid (PFOA) and perfluorooctyl sulfonate (PFOS). The diverse structural capabilities of these peptide cages pave a new way for designing synthetic receptors with varied hydrophobicity and hydrophilicity profiles, enhancing their utility in sensing and biomedical fields.
Results and discussion
Receptors design
In our design, benzene tricarboxamide was chosen to form the top and bottom floors of the cage. These components are linked together by three pillars, each comprising two amino acid residues and a central 1,3-diaminobenzene derivative (DAB) (Fig. 2). In each pillar, the amino acid building block could be selected from the full spectrum of 20 standard amino acids. This design allows for interchangeability and versatility in the cage structure variability and hydrophilicity/hydrophobicity. Additionally, the DAB unit can be modified with various functional groups to improve their solubility (R3, Fig. 2a) and introduce additional noncovalent interactions (R4, Fig. 2a). These modifications are crucial for adjusting the cage’s internal binding environment, offering precise manipulation of receptor-substrate interactions.
a General synthetic route for tripodal cages. Detailed procedures were included in supplementary information. b, c Partial 1H NMR spectra (600 MHz, 298 K) of CGlu and CPro-NH2 in DMSO-d6. The proton designations were labeled as capital letters (i.e., A, B, C etc.). d Partial NOESY spectrum (600 MHz, 298 K) of CPro-NH2 in DMSO-d6.
Synthesis and Characterizations
To construct these cages, glutamic acid, proline, and alanine were chosen due to their different flexibility/rigidity and hydrophilicity/hydrophobicity. These cages were denoted as CGlu, CPro and CAla based on their amino acid building blocks. Attaching an amino group to the DAB unit leads to the use of a ‘NH2’ suffix in the name, such as CPro-NH2. Additionally, when dicyclohexyl groups are added to the DAB segment, the name of the cage is appended with ‘Cy’, e.g. CPro-Cy-NH2.
Experimentally, amino acid-based tripodal cages were synthesized from starting materials 1, 2, and 4 via the projected route (Fig. 2a). Briefly, intermediate 3 was synthesized by coupling 1 with commercially available Boc-protected amino acids using isobutyl chloroformate as a catalyst. In parallel, compound 4 was activated to form the pentafluorophenyl intermediate 5, which subsequently reacted with 3 to form the C-shaped intermediate 6. The tert-butyl groups of 6 were then removed and the carboxylic groups of the product reacted with pentafluorophenol to afford C-shaped building block 7. In the next step, the macrobicyclization reactions were performed by slowly adding 3 to a dilute solution of 7 in THF. After removing the benzyl-protecting groups, we obtained five tripodal cages (Fig. 2a). The yields of these cages are approximately 8–20% based on compound 7. NMR and mass spectra confirmed the successful macrobicyclizations (Supplementary Fig. 1–39). In DMSO-d6, phenylic protons (B, F, and G) and amide proton (D) of CGlu exhibit as singlets in the 1H NMR spectra (Fig. 2b and Supplementary Fig. 3), indicating that the time-averaged conformation of CGlu is highly symmetric. When a more rigid proline was introduced to the pillar of cages, such as CPro-NH2 and CPro-Cy-NH2, these cages adopted desymmetrized conformations in solution. For example, in the 1H NMR spectrum of CPro-NH2 in DMSO-d6, three singlets B1–3 attributed to the phenyl protons of the benzamides, and three sets of proton signals C1–3, D1–3, and E1–3 were also observed (Fig. 2c and Supplementary Fig. 19). The NOESY spectrum of CPro-NH2 in DMSO-d6 highlighted several NOE correlations between protons B and C (Fig. 2d and Supplementary Fig. 24), indicating the spatial proximity of certain proton groups: proton C1 showed correlations with protons B1 and B2; C2 with B2 and B3; and C3 exhibited correlations with all B-group protons (B1, B2, and B3). These findings imply a closer proximity between two of the pillars and a greater distance from the third. Variable temperature NMR experiments conducted in DMSO-d6 revealed that the distinct conformations of the three pillars in CPro-NH2 gradually converge when the temperature is increased from 298 K to 373 K (Supplementary Fig. 40). Furthermore, the structure of CPro-NH2 is stable across a pH range of 4.85 to 8.8, as evidenced by their consistent retention times during LC-MS analysis within this pH spectrum of HEPES buffer (Supplementary Fig. 41).
X-ray diffraction analysis
Crystals of CGlu•Mg2+ complex, CPro-NH2 and CPro-NH2•NaI complex were obtained using the hanging drop vapor diffusion method. Single crystal X-ray diffraction (SCXRD) analysis showed that the deprotonated CGlu crystallized in the P1 space group with Mg2+ in two conformations, as shown in Fig. 3a and Supplementary Data 1. In these structures, the carbonyl groups of the benzamide nearly align with the plane of the benzene ring, and two benzamide moieties are almost parallel to each other, separated by a distance of 9.0 and 9.4 Å. The conformations of CGlu’s three pillars in the solid state differ from its highly symmetrical averaged conformation in the solution. Specifically, the first pillar features two carbonyl groups oriented towards the interior of the cavity. The other two pillars include one carbonyl group and one NH amide group facing the cavity. These variations in conformations suggest that, in solution, these conformations are likely interconverted rapidly at the NMR timescale. In one conformation of these cages in the solid state, there are two partially included DAB moieties from two neighboring cages (Fig. 3a, Left). In addition, several hydrogen-bonded water molecules were found inside the cavity (Fig. 3a, red), suggesting that the cage included the DAB units with its solvated water and some additional water molecules. In the other conformation, the cavity also included several water molecules (Fig. 3a, Right). The distinct solid-state conformation highlights the cage’s capability for adaptive binding, providing insightful evidence of its structural flexibility.
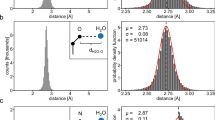
a Crystal structure of CGlu•Mg2+ complex (CCDC number: 2353051). b single-crystal structure of CPro-NH2 with hydrated water molecules (CCDC number: 2347455). The cage and hydrated water molecules are shown separately on the right. c Crystal structure of CPro-NH2•NaI complex with hydrated water molecules (CCDC number: 2347456). The cage and hydrated sodium/water are shown separately on the right. Oxygen atoms are shown in red, nitrogen atoms in blue, carbon atoms in gray, hydrogen atoms in white, magnesium atoms in green, sodium atoms in violet, iodine atoms in purple.
Single crystals of the CPro-NH2 were successfully grown in deionized water for SCXRD analysis at first. The solid-state structure reveals that the cavity of the CPro-NH2 has collapsed, with the benzamide groups oriented orthogonally to each other and positioned merely 4.8 Å apart (Fig. 3b and Supplementary Data 2). Additionally, several hydrated water molecules are incorporated in the solid-state structure of the CPro-NH2 (Fig. 3b). This collapsed conformation of CPro-NH2 is stabilized by a significant number of hydrogen bonds, forming a ‘belt’ of water molecules that connect to cage. An intriguing variation was observed when NaI was introduced into the stock solution, leading to the formation of a hydrated CPro-NH2•NaI complex (Fig. 3c and Supplementary Data 3). Unlike the collapsed conformation observed in the hydrated CPro-NH2, the cavity within the complex retains a highly symmetrical conformation, with the two benzamide groups aligned parallel to each other and separated by 6.7 Å. In this arrangement, the benzamide carbonyl groups are rotated to point toward the cavity, along with the three amino groups of the DAB units (Supplementary Fig. 47). The structure also features three sodium cations that are coordinated with three bridging water molecules inside the cavity to form a distorted hexagonal cluster. In addition, these five-coordinate sodium cations bond with the oxygen atoms of three carbonyl groups and free water molecules within the CPro-NH2 cage. This cluster is further supported by additional water molecules as a hydration shell. Notably, the hydrophobic iodide anions are positioned outside the cavity, maintaining a distance from the cage’s hydrogen-bonding groups. This arrangement is distinct from many cage-halide complexes, where anionic iodine typically engages directly with the hydrogen-bonding sites of the cage. The solid-state structures of the CPro-NH2 indicate its ability to adapt its conformation to accommodate guest molecules. More importantly, the observation of the highly hydrated binding environment of CPro-NH2 suggests that the inclusion of a hydrophilic substrate may not require significant desolvation of water molecules.
Anion binding studies
The rich hydrogen bonding sites of these cages made them good receptors for anions in non-aqueous environments, and tetrabutylammonium halogen salts (TBA+F−, TBA+Cl−, TBA+Br−, and TBA+I−) were chosen to probe the receptor-substrate interactions. When these TBA+X− salts are mixed with CPro-NH2 in DMSO-d6, the proton signals of C2 of CPro-NH2 shifted downfield significantly with the addition of anions such as Cl− and Br−, while the remaining protons shifted upfield slightly (Supplementary Figs. 48 and 49). When I− was added, although no significant chemical shift occurred, several new peaks emerged (Supplementary Fig. 50). Conversely, when TBA+ F− was added to CPro-NH2, the 1H NMR spectra exhibited a complex splitting pattern which is attributed to the strong F−•••H‒N hydrogen bonding interactions (Supplementary Fig. 51). A similarly complex splitting pattern was also observed in the ¹H NMR experiment conducted in CD₃CN (Supplementary Fig. 52).
Job plots performed by monitoring the absorption changes in the UV-Vis spectra at 240 nm in dry CH3CN determined the binding stoichiometry of the CPro-NH2•X as 1:1 complex (Supplementary Fig. 53). Isothermal titration calorimetry (ITC) was then used to determine the binding constants (Ka) and thermodynamic parameters (ΔG, ΔH and TΔS) of the CPro-NH2•X complexes in dry CH3CN and water (Table 1). In CH3CN, CPro-NH2 showed a strong binding with TBA+F− with the Ka of 2.90 (±0.39) × 104 M−1 at 25 °C. Its binding toward other halogen anions decreased with the increased anion sizes, with the Ka for Cl− and Br− measured as 7.21 (±0.97) × 103 M−1 and 4.12 (±0.24) × 103 M−1, respectively. In water, CPro-NH2 showed strong binding with F− anion but not binding with other halogen anions. The binding affinities Ka between CPro-NH2 and fluoride anions in water in the forms of NaF, KF, RbF, and CsF are measured as 3.08 (±0.33) × 103 M−1, 2.09 (±0.06) × 103 M−1, 2.28 (±0.25) × 103 M−1, and 3.31 (±0.18) × 103, respectively (Table 1). Clearly, the binding of fluoride anions in water is independent of the counter-cationic species.
The formation of the inclusion complex between NaF and CPro-NH2 in D2O was also confirmed in the 1H NMR titration experiment (Fig. 4a and Supplementary Fig. 54). Upon the addition of NaF, the proton signals B3 and protons D3 of CPro-NH2 shifted downfield, and proton E shifted upfield. Among them, the upfield shift of proton E3 is the most significant, suggesting that the DAB unit rotated upon the inclusion of F−. The average binding affinity is measured as 1498 ± 30 M−1, which is consistent with the ITC measurements. Notably, although CPro-NH2 binds more hydrated SO42− with a weaker binding affinity compared to F− (Ka = 43 ± 7 M−1, Table 1 and Supplementary Fig. 55), this result also underscores the unique capability of CPro-NH2 cage for binding anions with high hydration energy. Moreover, the CPro-NH2 cage exhibited no significant binding to less hydrated anions such as NO3− and ClO4− (Supplementary Figs. 56 and 57). These results render CPro-NH2 as one of the most fluoride-selective binding receptors, and a strong neutral fluoride-binding receptor in pure water (Supplementary Table S5)52,60,61,62,63.
a Partial 1H NMR spectra (top, 600 MHz, D2O, 298 K) and binding analysis curve (bottom) for receptor CPro-NH2 (1 mM) titrated with different amounts of Na+∙F-. The key resonances of protons in CPro-NH2 were labeled in the spectrum, black lines are curved using a 1:1 receptor-substrate binding model. b ITC profiles for the titration of CPro-NH2 with NaF in water. The solid line represents the best non-linear fit of the data to a 1:1 binding model.
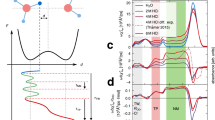
The thermodynamic data, including binding enthalpy and entropy, unveiled crucial insights into the origins of complexation as shown in Fig. 4b. For the CPro-NH2•F− complex in water, the binding process is characterized by an endothermic nature with an unfavorable enthalpy change (+3.47 kJ/mol), while being driven predominately by entropy (−TΔS = −8.23 kJ/mol). This behavior, observed in both CH3CN and water for the CPro-NH2•F− complexation, is indicative of an anti-Hofmeister binding behavior, a notably uncommon phenomenon for anion-binding receptors. Although the single-crystal structures of the inclusion complexes were not obtained, the NMR and ITC investigations of the complexes, and the solid-state structures of CPro-NH2 in the absence and presence of NaI provide a critical understanding of the mechanisms underlying their high affinity and selectivity for fluoride binding. In aqueous environments, CPro-NH2 undergoes extensive hydration, and its structure exhibits sufficient flexibility to accommodate F−. However, the hydrogen bonds formed between the F− anion and the receptor’s binding sites do not fully offset the enthalpic penalty associated with the displacement of hydrogen-bonded water molecules from either CPro-NH2, F−, or both. Meanwhile, the replacement of CPro-NH2’s bound water molecules by an F− anion leads to a positive entropy effect on complexation. Furthermore, the highly hydrated state of CPro-NH2 suggests that the inclusion of F− may not significantly disrupt the hydration shell of the anion. Additionally, the volume of CPro-NH2 was calculated to be 132 ų (Supplementary Fig. 58), which is slightly smaller than that of cucurbituril [6] (142 ų) but much larger than the volume of a single fluoride anion64,65. Due to the numerous polar groups converging towards the binding cavity, fluoride binding by CPro-NH2 is most likely facilitated by the participation of water molecules. Computational modeling using the GFN2-xTB method was also employed to elucidate the possible binding model between CPro-NH2 and hydrated fluoride (Supplementary Fig. 59)66,67. The optimized structure indicates that approximately 10 water molecules engage in hydrogen bonding with CPro-NH2 within the binding pockets. Of these, four water molecules further participate in hydrogen bonding with the fluoride anion, which is encapsulated inside the binding pocket of CPro-NH2. The strong hydrogen bonding between the cavity water molecules and fluoride is further corroborated by independent gradient model (IGM) analysis68,69,70, which displays isosurfaces representing strong attractions (Supplementary Fig. 60). Besides, an isosurface representing weak interaction was found between the fluoride and one of the benzene-1,3,5-tricarboxylate rings. Considering the electron-deficient nature of this aromatic surface, the [F⁻⋯π] interaction could also contribute to the complex formation. These findings have led us to suggest a significantly underexplored path for designing anion receptors. Instead of relying on binding sites to create multivalent directional hydrogen bond interactions, the use of highly hydrated receptors for the binding of hydrophilic anions with their hydration water molecules could markedly lower the energy associated with desolvation. This strategy, although previously observed in protein-substrate interactions, has not been clearly defined as a binding model in synthetic receptor design until this study.
PFAS binding and removal
In addition to exploring the strong affinity for hydrophilic hydrated fluoride anion binding in aqueous solution, our research extends to investigating how the cage CPro-NH2 interact with the amphiphilic anionic PFAS bearing large hydrophobic regions. PFAS, including chemicals used in a variety of consumer products, firefighting foams, and surfactants in fluorinated polymer production, present substantial environmental challenges71. Their persistence due to resistance to biodegradation or chemical breakdown, coupled with their association with adverse health outcomes such as significant liver damage, thyroid disorders, and various cancers, underscores the critical need to mitigate their environmental and health impacts72. In this context, our study focused on perfluorooctanoic acid (PFOA), perfluorooctyl sulfonate (PFOS), and perfluoro-2-propoxypropanoic acid (GenX) as model PFAS compounds to elucidate the interactions between these species and our peptide cages. We hypothesize that the ability of these cages to bind both amphiphilic PFAS and hydrophilic fluoride anion could further enhance their utility for practical applications, including environmental remediation.
The stoichiometry of the complex formed between PFOA and CPro-NH2 was first investigated using UV-visible spectroscopy through continuous titration in CH3CN. Job plot analyses confirmed the formation of 1:1 complex (Supplementary Fig. 68). Additional evidence for this 1:1 stoichiometry between PFOA and CPro-NH2 was corroborated by ESI-HRMS, which identified molecular ions corresponding to [PFOA + CPro-NH2]+ (Supplementary Fig. 69). To further quantify the binding affinities of CPro-NH2 with various PFAS compounds, we conducted isothermal titration calorimetry (ITC) in CH3CN (Table 1). The binding constants (Ka) measured for CPro-NH2 with GenX, PFOA, and PFOS were found to be 622 ± 32 M−1, 735 ± 32 M−1, and 1.00 (±0.07) × 103 M−1, respectively. Thermodynamic analyses revealed that the inclusion of PFOA, PFOS, and GenX in CH3CN is largely driven by favorable enthalpic contributions, likely resulting from hydrogen bonding interactions, despite the entropically unfavorable conditions.
In water, the low solubility of PFAS complicates the reliable measurement of binding affinities of CPro-NH2 with GenX, PFOA, and PFOS. However, we successfully obtained single crystals of the CPro-NH2 and PFOA−K+ complex, which were suitable for single-crystal X-ray diffraction (SCXRD) analysis (Fig. 5a and Supplementary Data 4). In the solid state, CPro-NH2 formed a 1:4 complex with PFOA-K⁺, where two PFOA-K⁺ ions were positioned at the portal of the peptide cage, while the remaining two occupied voids within the crystal lattice. Specifically, one PFOA-K⁺, in which the 8-coordinated potassium cation is coordinated with the carboxylate head of PFOA, the carbonyl group of the cage, and water molecules, interacts with the cage through coordination and hydrogen bonding. Meanwhile, the other PFOA-K⁺ involves a 6-coordinated potassium cation that is coordinated with the cage and water molecules, which subsequently interacts with water molecules and the carboxylate head of PFOA. The hydrophobic fluorocarbons of PFOA interacted with the methylene groups of the proline moiety. SCXRD analysis showed some short CH⋯FC contacts of 2.6–2.8 Å73, as well as more distant van der Waals contacts (Supplementary Fig. 70).
a Single crystals of the CPro-NH2 and PFOA-K+ complex (CCDC number: 2382261). Packed PFOA-K+ are shown in the wireframe model. The cage and PFOA- are shown in the capped stick model. Hydrated water (hydrogen atoms omitted) and K+ are shown in the spacefill model. b Batch equilibrium adsorption experiment of CPro-Cy-NH2 with PFOA. Time dependent PFOA sorption by CPro-Cy-NH2 at low ([PFOA]0 = 1 µg/L; [CPro-Cy-NH2] = 0.3 mg/mL, black broken line) and high ([PFOA]0 = 50 µg/L; [CPro-Cy-NH2] = 0.6 mg/mL, red broken line) concentration. Error bars: standard deviation of three experiments. c, Batch equilibrium adsorption experiment of CPro-Cy-NH2 with PFOA ([PFOA]0 = 50 µg/L, black broken line) in the presence of octanoic acid ([octanoic acid]0 = 50 µg/L, red broken line). Error bars: standard deviation of three experiments. d Time dependent PFOA sorption by CPro-Cy-NH2 at groundwater containing 20 ppm HA and extra1µg/L [PFOA]0 ([CPro-Cy-NH2] = 0.3 mg/mL, black broken line). Red broken line represents the removal under the condition of the water without HA. Error bars: standard deviation of three experiments. e Additional cages were synthesized for PFAS removal. The red rectangle highlights the part that was modified. f Time dependent PFOA sorption by CPro-Cy-NH2, CPro-Cy, and CAla-Cy-NH2 at concentration of ([PFOA]0 = 50 µg/L; [Cage] = 0.6 mg/mL). Error bars: standard deviation of two or three experiments.
The binding studies of the inclusion complex between CPro-NH2 and PFOA in CH3CN, as well as their solid-state structural analysis provided critical insights for the removal of PFAS in water. The cage is effective in binding the hydrophilic head at its portal, while the proline residues enhance the van der Waals interactions and some short CH⋯FC contacts. Hence, we attached dicyclohexyl groups on the DAB segment and synthesized cages CPro-Cy-NH2 with increased CH⋯FC contacting surfaces for PFAS binding (see below). Additionally, Maestro MacroModel was then used to carry out NOE-restrained molecular dynamics calculations to predict the architecture of the CPro-Cy-NH2. Based on signal intensities, NOEs observed could be grouped into one of three categories, 2.5–3.5 Å, 3.5–4.5 Å, and 4.5–5.5 Å. The 10 best structures for CPro-Cy-NH2 and their average conformation were generated (Supplementary Fig. 71). This simulated CPro-Cy-NH2 in solution showed a distorted conformation closely mirrors that of the CPro-NH2.
We then performed batch adsorption experiments using the peptide cage CPro-Cy-NH2 to assess PFAS removal kinetics. In these experiments, CPro-Cy-NH2 was added to a solution containing a specific concentration of PFAS substrate and the mixture was stirred to allow adsorption of PFAS. At designated intervals, small aliquots of the solution were withdrawn, centrifuged, and the supernatant was analyzed to quantify the remaining PFAS. Initial PFASs removal efficiency of CPro-Cy-NH2 was evaluated by conducting batch equilibrium adsorption experiments in natural water, with an initial PFOA concentration of 1 µg/L ([PFOA]0 = 1 µg/L; [CPro-Cy-NH2] = 0.3 mg/mL). The result showed that CPro-Cy-NH2 removed 70–80% of PFOA within the first 10 min. The adsorption rate subsequently slowed, reaching equilibrium after approximately 6 h, with the final concentration of <50 µg/L within 24 h (Fig. 5b). Additional kinetic adsorption experiments performed at higher PFOA concentration ([PFOA]0 = 50 µg/L; [CPro-Cy-NH2] = 0.6 mg/mL) showed nearly 95% PFOA removal within 4 h (Fig. 5b). ESI-HRMS spectrometry of the precipitate confirmed the existence of 1:1 complex between CPro-Cy-NH2 and PFOA (Supplementary Fig. 75). The adsorption rate at both low and high concentrations of [PFOA]0 was modeled using both linear and nonlinear Ho and McKay’s pseudo-second-order adsorption model. This model was used to derive the observed adsorption rate constant, Kobs, which quantifies how quickly equilibrium is reached (Supplementary Fig. 76). The nonlinear pseudo-second-order model yielded higher correlation coefficients, with Kobs valuates of 1565.56 g mg−1 h−1 for [PFOA]0 = 1 µg/L and [CPro-Cy-NH2] = 0.3 mg/mL, and 44.92 g mg−1 h−1 for [PFOA]0 = 50 µg/L and [CPro-Cy-NH2] = 0.6 mg/mL. To further investigate adsorption capacity, we constructed a PFOA binding isotherm using [CPro-Cy-NH2] = 0.4 mg/mL and [PFOA]0 from 0.18 mg/L to 12 mg/L. Both Freundlich and Langmuir isotherm models were applied to fit the experimental data (Supplementary Fig. 77). From on the Langmuir fit, we calculated an affinity coefficient (KL) of 2.8 × 105 M−1 and an estimated maximum capacity of CPro-Cy-NH2 of 19.99 mg g−1. The Freundlich model provided a Freundlich’s constant (KF) of 5.08 (mg g−1) (L mg−1)1/n and intensity of adsorption (n) of 1.43, respectively. Powder X-ray diffraction (PXRD) analysis showed that the CPro-Cy-NH2 cage remained amorphous before and after PFOA uptake (Supplementary Fig. 78). Interestingly, the adsorbed PFOA showed a broad diffraction at 18° (corresponding to a distance of 5 Å), suggesting that the oleophobic and hydrophobic fluorocarbon tails of PFOA aggregates in an ordered manner similar to the reported perfluorodecanoic salt74.
To evaluate the selectivity of CPro-Cy-NH2 in removing PFOA, we investigated PFOA removal in the presence of octanoic acid, a defluorinated analog. As shown in Fig. 5c, the removal of PFOA remained consistently high, exceeding 97% after 6 h, whereas only 20% of octanoic acid was removed under the same conditions. This result showed that the CH⋯FC interactions between the fluorinated PFOA tail and proline moiety of CPro-Cy-NH2 cage are not simply van der Waals contacts. Instead, they involve specific, albeit weak, CH⋯FC interactions similar to another C(sp3)-H⋯O/X interactions75,76,77. We further investigated the removal of PFOA in the presence of other organic co-contaminants or natural organic matter (NOM), such as humic acid (HA). Settled groundwater from Sweeney Water Treatment Plant (pH = 7.25, TOC ≤ 7.24 mg/L) containing 20 ppm HA was spiked with PFOA at 1 µg/L. The result demonstrated that 0.3 mg/mL CPro-Cy-NH2 was sufficient to reduce the residual concentration of PFOA to below 50 µg/L or 50 ppb, even in the presence of NOM (Fig. 5d).
Following the promising results for PFOA removal, we further evaluated the ability of CPro-Cy-NH2 to absorb GenX and PFOS in water. Batch equilibrium adsorption experiments were conducted in natural water with GenX at a concentration of 100 µg/L ([GenX]0 = 100 µg/L; [CPro-Cy-NH2] = 0.6 mg/mL) and PFOS at a concentration of 100 µg/L ([PFOS]0 = 100 µg/L; [CPro-Cy-NH2] = 0.16 mg/mL), respectively. As shown in Supplementary Fig. 79, CPro-Cy-NH2 effectively removed 80% of GenX within 8 h, ultimately achieving a maximum capacity of 95.7% by the end of the experiment. Notably, CPro-Cy-NH2 demonstrated exceptional efficiency for PFOS, absorbing 62% of PFOS within the first 2 min and achieving virtually complete removal by 8 h. The absorption rate of CPro-Cy-NH2 for both GenX and PFOS were characterized by both linear and nonlinear Ho and McKay’s pseudo-second-order adsorption model, with the results shown in Supplementary Fig. 80.
To gain a analysis of structure-activity relationships between the peptide cages and their removal efficiency, we synthesized two additional cages, namely CPro-Cy and CAla-Cy-NH2 (Fig. 5e). Each of these cages was designed with specific modifications to elucidate the role of amino groups and the conformation of the cages. Specifically, the amino group attached to the DAB unit was removed in the CPro-Cy structure, while in CAla-Cy-NH2, the amino acid was substituted with alanine. As shown in Fig. 5f, the removal efficiency of PFOA was decreased when the amino group was absent (CPro-Cy) or when the proline group was replaced (CAla-Cy-NH2), indicating both amino group and the cage conformation are critical for effective interaction with PFOA.
In summary, we successfully developed a new series of synthetic receptors by incorporating hydrophobic or hydrophilic amino acid residues. Using NMR spectroscopy and X-ray diffraction analysis, we discovered that proline-based cages exhibited unique conformations, characterized by a highly hydrated binding pocket capable of accommodating hydrated guest molecules in aqueous environments. In anion binding studies, proline-based cage CPro-NH2 displays excellent fluoride binding specificity and affinity in aqueous environments via an atypical entropy-driven, endothermic binding mechanism. The ability of this cage to bind hydrophilic F− without requiring significant desolvation of water molecules contributes to its much-enhanced binding selectivity. Furthermore, the cage undergoes conformational changes upon substrate binding, akin to the “induced fit” mechanism observed in protein-ligand interactions. Beyond its affinity for fluoride, the cage also demonstrated strong binding for hydrophobic amphiphilic fluorinated compounds, including PFOA, PFOS, and GenX. Its hydrophobic derivative CPro-Cy-NH2 effectively removed these amphiphilic fluorinated compounds from water, achieving removal efficiencies exceeding 95%. A structure-activity relationship analysis revealed that both the amino group and distinct conformation of the proline-based cage are crucial for its interaction with PFAS compounds and other substrates. This research advances the field by providing valuable insights into the design and synthesis of peptide-based cage structures with theoretically large functional variabilities, offering enhanced recognition of molecular species in aqueous environments.
Methods
General information
Commercial reagents were purchased from Oakwood, Alfa-Aesar, Chem-Impex or Sigma-Aldrich and were used without further purification unless otherwise specified. NMR spectra were recorded on Bruker Neo 600 MHz spectrometer. DMSO-d6 or D2O was used as solvent and chemical shifts were referenced relative to residual solvent. The following abbreviations are used to describe peak patterns where appropriate: br = broad, s = singlet, d = doublet, t = triplet, q = quartet, m = multiplet. Coupling constants (J) are reported in Hertz (Hz). ESI-TOF-MS and TWIM-MS were recorded on a Waters Synapt G2 mass spectrometer and LC-MS was performed on Agilent LC-MS SQ 6120. Isothermal calorimetry titrations were performed on a Microcal ITC 200. Analysis of PFAS concentration was performed using Agilent 1260 HPLC system coupled to an Agilent 6460 triple Quad mass spectrometer system operated in negative ion mode. Analytes were separated on a 2.1 × 50 mm Sunfire C18 3.5 µm column (Waters Corporation, Milford, MA), X-ray diffraction data were measured on Bruker D8 Venture PHOTON III diffractometer equipped with a Cu Kα INCOATEC ImuS 3.0 micro-focus source (λ = 1.54178 Å). The Supplementary Information contains details of syntheses, crystallography, NMR, ITC data and kinetic adsorption data.
Synthesis and characterization of all cages
Detailed synthesis procedures can be found in supplementary information. Notably, the benzyl-protected precursor of CGlu, Cbz-protected precursor of CPro-NH2, CPro-Cy-NH2, and CAla-Cy-NH2 was purified by Waters HPLC system installed with both an analytic module (1 mL/min) and a preparative module (16 mL/min) by employing a method using 5–100% linear gradient of solvent B (0.1% TFA in ACN) in solvent A (0.1% TFA in H2O) over 45 min, followed by 100% solvent B over 15 min. After freeze-drying, each precursor was further purified by column chromatography using dichloromethane /methanol (v/v, 10:1) as the eluent. Subsequently, the benzyl or Cbz protective groups were removed via Pd/C (10%) reduction, and the desired cages were obtained by filtration and concentration. All cages were characterized by NMR spectroscopy and TWIM-MS. All the spectra were included in the supplementary information.
Fluoride captures by cage CPro-NH2
1H-NMR titrations were performed on a Bruker Neo 600 MHz spectrometer. Solutions of different anion (TBAF∙3H2O, TBACl, TBABr, TBAI, NaF, NaNO3, NaClO4 and Na2SO4) in DMSO-d6 or D2O, containing receptor at a known concentration (1 mM) to be used in the experiment, were prepared. The association constant Ka was determined by fitting the data to a 1:1 binding model using the www.supramolecular.org web applet. The detailed procedure was provided in the supplementary information. Isothermal Titration Microcalorimetry (ITC) experiments were performed on a MicroCal iTC200 at 298 K. The guest solution and cage solution were prepared with HPLC-grade extra-dry acetonitrile or HPLC-grade water. Heats of dilution were measured by injecting the same anion solution into HPLC-grade extra-dry acetonitrile or HPLC-grade water, using identical conditions. For every addition, the heat of dilution was subtracted from the heat of binding and the Ka, ∆H, and ∆S were derived by fitting the data to a 1:1 binding model using MicroCal software.
PFASs removed by CPro-Cy-NH2
All materials were purchased from commercial sources and used as received without further purification unless otherwise mentioned. Humic acid and perfluorooctanoic acid (PFOA) were purchased from Sigma-Aldrich. Analytical standards for PFOA, PFOS, and GenX were obtained from Wellington Labs (Guelph, ON, CA). Isotope-labeled analogues (M8PFOA, M3HFPO-DA, M8PFOS) were also purchased from Wellington Labs (Guelph, ON, CA). The settled groundwater was obtained from Sweeney Water Treatment Plant and stored under refrigeration until analysis, the detailed components were described in supplementary information. The quantitative analysis of target compounds was performed using Agilent 1260 HPLC system coupled to an Agilent 6460 triple Quad mass spectrometer system operated in negative ion mode.
Data availability
Data supporting the findings of this study are available from the manuscript and its Supplementary Information. Crystallographic data for the structures reported in this study have been deposited at the Cambridge Crystallographic Data Centre (CCDC), under deposition numbers 2353051 (CGlu•Mg2+), 2347455 (CPro-NH2), 2347456 (CPro-NH2•NaI) and 2382261 (CPro-NH2•PFOA). Copies of the data can be obtained free of charge via https://www.ccdc.cam.ac.uk/structures/. All data is available from the corresponding author upon request. Source data are provided with this paper.
References
Chatterji, D. Basics of Molecular Recognition (CRC Press, 2016).
Gale, P. A. & Caltagirone, C. Anion sensing by small molecules and molecular ensembles. Chem. Soc. Rev. 44, 4212–4227 (2015).
Richard, J. P. Enabling role of ligand-driven conformational changes in enzyme evolution. Biochemistry 61, 1533–1542 (2022).
Persch, E., Dumele, O. & Diederich, F. Molecular recognition in chemical and biological systems. Angew. Chem. Int. Ed. 54, 3290–3327 (2015).
Meyer, E. A., Castellano, R. K. & Diederich, F. Interactions with aromatic rings in chemical and biological recognition. Angew. Chem. Int. Ed. 42, 1210–1250 (2003).
Amaral, M. et al. Protein conformational flexibility modulates kinetics and thermodynamics of drug binding. Nat. Commun. 8, 2276 (2017).
Garvey, P. E. Structural mechanisms of slow-onset, two-step enzyme inhibition. Curr. Chem. Biol. 4, 64–73 (2010).
Monod, J., Wyman, J. & Changeux, J.-P. On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88–118 (1965).
Hammes, G. G., Chang, Y.-C. & Oas, T. G. Conformational selection or induced fit: a flux description of reaction mechanism. Proc. Natl Acad. Sci. USA 106, 13737–13741 (2009).
Greives, N. & Zhou, H.-X. Both protein dynamics and ligand concentration can shift the binding mechanism between conformational selection and induced fit. Proc. Natl. Acad. Sci. USA 111, 10197–10202 (2014).
Wu, Y. et al. A self-assembled cage binding iodide anions over other halide ions in water. Angew. Chem. Int. Ed. 61, e202209078 (2022).
Sigwalt, D. et al. Unraveling the structure–affinity relationship between cucurbit[n]urils (n = 7, 8) and cationic diamondoids. J. Am. Chem. Soc. 139, 3249–3258 (2017).
Wang, Y.-M. et al. Unexpected solvent effect on the binding of positively-charged macrocycles to neutral aromatic hydrocarbons. Chem. Commun. 55, 10924–10927 (2019).
Moghaddam, S. et al. New ultrahigh affinity host−guest complexes of cucurbit[7]uril with bicyclo[2.2.2]octane and adamantane guests: Thermodynamic analysis and evaluation of m2 affinity calculations. J. Am. Chem. Soc. 133, 3570–3581 (2011).
Yamashina, M., Tsutsui, T., Sei, Y., Akita, M. & Yoshizawa, M. A polyaromatic receptor with high androgen affinity. Sci. Adv. 5, eaav3179 (2019).
Piatnitski, E. L., Flowers, R. A. II & Deshayes, K. Highly organized spherical hosts that bind organic guests in aqueous solution with micromolar affinity: microcalorimetry studies. Chem. Eur. J. 6, 999–1006 (2000).
Ferguson Johns, H. P., Harrison, E. E., Stingley, K. J. & Waters, M. L. Mimicking biological recognition: lessons in binding hydrophilic guests in water. Chem. Eur. J. 27, 6620–6644 (2021).
Dong, J. & Davis, A. P. Molecular recognition mediated by hydrogen bonding in aqueous media. Angew. Chem. Int. Ed. 60, 8035–8048 (2021).
Chen, L., Berry, S. N., Wu, X., Howe, E. N. W. & Gale, P. A. Advances in anion receptor chemistry. Chem 6, 61–141 (2020).
Busschaert, N., Caltagirone, C., Van Rossom, W. & Gale, P. A. Applications of supramolecular anion recognition. Chem. Rev. 115, 8038–8155 (2015).
Lane, J. D. E., Shiels, G., Ramamurthi, P., Müllner, M. & Jolliffe, K. A. Water-soluble squaramide-functionalized copolymers for anion recognition. Macromol. Rapid Commun. 2300406. https://doi.org/10.1002/marc.202300406 (2023).
Elmes, R. B. P. & Jolliffe, K. A. Anion recognition by cyclic peptides. Chem. Commun. 51, 4951–4968 (2015).
Qin, L., Hartley, A., Turner, P., Elmes, R. B. P. & Jolliffe, K. A. Macrocyclic squaramides: anion receptors with high sulfate binding affinity and selectivity in aqueous media. Chem. Sci. 7, 4563–4572 (2016).
Morshedi, M., Thomas, M., Tarzia, A., Doonan, C. J. & White, N. G. Supramolecular anion recognition in water: synthesis of hydrogen-bonded supramolecular frameworks. Chem. Sci. 8, 3019–3025 (2017).
Foyle, E. M. et al. Expedient decagram-scale synthesis of robust organic cages that bind sulfate strongly and selectively in water. J. Am. Chem. Soc. 146, 27127–27137 (2024).
David, A. H. G., Goodwin, R. J. & White, N. G. Supramolecular chemistry of two new bis(1,2,3-triazolyl)pyridine macrocycles: Metal complexation, self-assembly and anion binding. Dalton Trans. 52, 1902–1912 (2023).
Zhang, D., Ronson, T. K., Mosquera, J., Martinez, A. & Nitschke, J. R. Selective anion extraction and recovery using a feii4l4 cage. Angew. Chem. Int. Ed. 57, 3717–3721 (2018).
Zhang, D. et al. Anion binding in water drives structural adaptation in an azaphosphatrane-functionalized feii4l4 tetrahedron. J. Am. Chem. Soc. 139, 6574–6577 (2017).
McTernan, C. T., Ronson, T. K. & Nitschke, J. R. Selective anion binding drives the formation of agi8l6 and agi12l6 six-stranded helicates. J. Am. Chem. Soc. 143, 664–670 (2021).
Langton, M. J., Serpell, C. J. & Beer, P. D. Anion recognition in water: Recent advances from a supramolecular and macromolecular perspective. Angew. Chem. Int. Ed. 55, 1974–1987 (2016).
Lim, J. Y. C. & Beer, P. D. Superior perrhenate anion recognition in water by a halogen bonding acyclic receptor. Chem. Commun. 51, 3686–3688 (2015).
Borissov, A. et al. Anion recognition in water by charge-neutral halogen and chalcogen bonding foldamer receptors. J. Am. Chem. Soc. 141, 4119–4129 (2019).
Liu, Y., Zhao, W., Chen, C.-H. & Flood, A. H. Chloride capture using a c–h hydrogen-bonding cage. Science 365, 159–161 (2019).
Jing, L., Deplazes, E., Clegg, J. K. & Wu, X. A charge-neutral organic cage selectively binds strongly hydrated sulfate anions in water. Nat. Chem. 16, 335–342 (2024).
Yao, H. et al. Molecular recognition of hydrophilic molecules in water by combining the hydrophobic effect with hydrogen bonding. J. Am. Chem. Soc. 140, 13466–13477 (2018).
Huang, G.-B., Wang, S.-H., Ke, H., Yang, L.-P. & Jiang, W. Selective recognition of highly hydrophilic molecules in water by endo-functionalized molecular tubes. J. Am. Chem. Soc. 138, 14550–14553 (2016).
Xiong, S., Nanda Kishore, M. V., Zhou, W. & He, Q. Recent advances in selective recognition of fluoride with macrocyclic receptors. Coord. Chem. Rev. 461, 214480 (2022).
Tromans, R. A. et al. A biomimetic receptor for glucose. Nat. Chem. 11, 52–56 (2019).
Mobley, D. L. & Guthrie, J. P. Freesolv: a database of experimental and calculated hydration free energies, with input files. J. Comput. Aided Mol. Des. 28, 711–720 (2014).
Molina, P., Zapata, F. & Caballero, A. Anion recognition strategies based on combined noncovalent interactions. Chem. Rev. 117, 9907–9972 (2017).
Daniel, R. M., Finney, J. L., Stoneham, M. & Nakasako, M. Water–protein interactions from high–resolution protein crystallography. Philos. Trans. R. Soc. Lond., Ser. B 359, 1191–1206 (2004).
García-Sosa, A. T., Mancera, R. L. & Dean, P. M. Waterscore: a novel method for distinguishing between bound and displaceable water molecules in the crystal structure of the binding site of protein-ligand complexes. J. Mol. Model. 9, 172–182 (2003).
Witschel, M. C. et al. Inhibitors of the herbicidal target ispd: allosteric site binding. Angew. Chem. Int. Ed. 50, 7931–7935 (2011).
Ladbury, J. E. Just add water! The effect of water on the specificity of protein-ligand binding sites and its potential application to drug design. Chem. Biol. 3, 973–980 (1996).
Fox, J. M. et al. Interactions between hofmeister anions and the binding pocket of a protein. J. Am. Chem. Soc. 137, 3859–3866 (2015).
Carey, C., Cheng, Y.-K. & Rossky, P. J. Hydration structure of the α-chymotrypsin substrate binding pocket: The impact of constrained geometry. Chem. Phys. 258, 415–425 (2000).
Lo Nostro, P. & Ninham, B. W. Hofmeister phenomena: an update on ion specificity in biology. Chem. Rev. 112, 2286–2322 (2012).
Patrick, S. C., Beer, P. D. & Davis, J. J. Solvent effects in anion recognition. Nat. Rev. Chem. 8, 256–276 (2024).
Sokkalingam, P., Shraberg, J., Rick, S. W. & Gibb, B. C. Binding hydrated anions with hydrophobic pockets. J. Am. Chem. Soc. 138, 48–51 (2016).
Guth, S. et al. Toxicity of fluoride: critical evaluation of evidence for human developmental neurotoxicity in epidemiological studies, animal experiments and in vitro analyses. Arch. Toxicol. 94, 1375–1415 (2020).
Tissandier, M. D. et al. The proton’s absolute aqueous enthalpy and gibbs free energy of solvation from cluster-ion solvation data. J. Phys. Chem. A. 102, 7787–7794 (1998).
Brugnara, A. et al. Selective recognition of fluoride anion in water by a copper(ii) center embedded in a hydrophobic cavity. Chem. Sci. 5, 3897–3904 (2014).
Clarke, H. J. et al. Transmembrane fluoride transport: direct measurement and selectivity studies. J. Am. Chem. Soc. 138, 16515–16522 (2016).
Kaur, H. et al. Metal–organic framework-based materials for wastewater treatment: Superior adsorbent materials for the removal of hazardous pollutants. ACS Omega 8, 9004–9030 (2023).
Bayazit, Ş. S. & Şahin, S. Acid-modulated zirconium based metal organic frameworks for removal of organic micropollutants. J. Environ. Chem. Eng. 8, 103901 (2020).
Klemes, M. J. et al. Reduction of a tetrafluoroterephthalonitrile-β-cyclodextrin polymer to remove anionic micropollutants and perfluorinated alkyl substances from water. Angew. Chem. Int. Ed. 58, 12049–12053 (2019).
Kumarasamy, E., Manning, I. M., Collins, L. B., Coronell, O. & Leibfarth, F. A. Ionic fluorogels for remediation of per- and polyfluorinated alkyl substances from water. ACS Cent. Sci. 6, 487–492 (2020).
Manning, I. M. et al. Hydrolytically stable ionic fluorogels for high-performance remediation of per- and polyfluoroalkyl substances (pfas) from natural water. Angew. Chem. Int. Ed. 61, e202208150 (2022).
He, Y. et al. Fluorinated nonporous adaptive cages for the efficient removal of perfluorooctanoic acid from aqueous source phases. J. Am. Chem. Soc. 146, 6225–6230 (2024).
Cametti, M., Dalla Cort, A. & Bartik, K. Fluoride binding in water: A new environment for a known receptor. ChemPhysChem 9, 2168–2171 (2008).
Trembleau, L., Smith, T. A. D. & Abdelrahman, M. H. Receptor conformational change induces fluoride binding despite competitive water binding. Chem. Commun. 49, 5850–5852 (2013).
Kim, Y. & Gabbaï, F. P. Cationic boranes for the complexation of fluoride ions in water below the 4 ppm maximum contaminant level. J. Am. Chem. Soc. 131, 3363–3369 (2009).
Suk, J.-M. & Jeong, K.-S. Indolocarbazole-based foldamers capable of binding halides in water. J. Am. Chem. Soc. 130, 11868–11869 (2008).
Maglic, J. B. & Lavendomme, R. MoloVol: an easy-to-use program for analyzing cavities, volumes and surface areas of chemical structures. J. Appl. Crystallogr. 55, 1033–1044 (2022).
Nau, W. M., Florea, M. & Assaf, K. I. Deep inside cucurbiturils: physical properties and volumes of their inner cavity determine the hydrophobic driving force for host–guest complexation. Isr. J. Chem. 51, 559–577 (2011).
Grimme, S. Exploration of chemical compound, conformer, and reaction space with meta-dynamics simulations based on tight-binding quantum chemical calculations. J. Chem. Theory Comput. 15, 2847–2862 (2019).
Bannwarth, C., Ehlert, S. & Grimme, S. Gfn2-xtb—an accurate and broadly parametrized self-consistent tight-binding quantum chemical method with multipole electrostatics and density-dependent dispersion contributions. J. Chem. Theory Comput. 15, 1652–1671 (2019).
Lu, T. & Chen, Q. Independent gradient model based on hirshfeld partition: A new method for visual study of interactions in chemical systems. J. Comput. Chem. 43, 539–555 (2022).
Lefebvre, C. et al. Accurately extracting the signature of intermolecular interactions present in the nci plot of the reduced density gradient versus electron density. Phys. Chem. Chem. Phys. 19, 17928–17936 (2017).
Lu, T. & Chen, F. Multiwfn: A multifunctional wavefunction analyzer. J. Comput. Chem. 33, 580–592 (2012).
Hu, X. C. et al. Detection of poly- and perfluoroalkyl substances (pfass) in u.S. Drinking water linked to industrial sites, military fire training areas, and wastewater treatment plants. Environ. Sci. Technol. Lett. 3, 344–350 (2016).
Borg, D., Lund, B.-O., Lindquist, N.-G. & Håkansson, H. Cumulative health risk assessment of 17 perfluoroalkylated and polyfluoroalkylated substances (pfass) in the swedish population. Environ. Int. 59, 112–123 (2013).
Kaur, G. & Choudhury, A. R. A comprehensive understanding of the synthons involving c–h⋯f–c hydrogen bond(s) from structural and computational analyses. CrystEngComm 17, 2949–2963 (2015).
O’Neal, K. L., Geib, S. & Weber, S. G. Extraction of pyridines into fluorous solvents based on hydrogen bond complex formation with carboxylic acid receptors. Anal. Chem. 79, 3117–3125 (2007).
Samanta, J. et al. Tripodal organic cages with unconventional ch···o interactions for perchlorate remediation in water. J. Am. Chem. Soc. 145, 21723–21728 (2023).
Chvojka, M., Madea, D., Valkenier, H. & Šindelář, V. Tuning ch hydrogen bond-based receptors toward picomolar anion affinity via the inductive effect of distant substituents. Angew. Chem. Int. Ed. 63, e202318261 (2024).
Abdurakhmanova, E. R. et al. Supramolecular umpolung: converting electron-rich resorcin[4]arenes into potent ch-bonding anion receptors and transporters. Chem 10, 1910–1924 (2024).
Acknowledgements
This work was supported by NIH 5RO1AI52416 (JC) and 5RO1149852 (JC) and the Department of Energy the Basic Energy Sciences DE-SC0022267 (CK). This work also has been supported in part by the Chemical Purification Analysis and Screening Core Facility (CPAS) in the Department of Chemistry at USF.
Author information
Authors and Affiliations
Contributions
J.C. and C.K. conceived and directed the project. B.H. synthesized all the cages and conducted the most of anion binding test and PFAS removal assays. S.L. prepared building blocks for cage synthesis and helped with NMR titration. C.P. prepared the single crystals for X-ray analysis. F.L. helped with the anion binding test. L.W. analyzed the X-ray data, and Q.Q. did the molecular dynamic simulation. T.T helped with crystal optimization. L.C. helped with instruments usage. W.L. helped with the ITC test and computational calculation. B.H., C.K. and J.C. analyzed data and interpreted results. B.H., C.K. and J.C. wrote and revised the manuscript. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Oksana Danylyuk, Qing He and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, B., Li, S., Pan, C. et al. Proline-based tripodal cages with guest-adaptive features for capturing hydrophilic and amphiphilic fluoride substances. Nat Commun 16, 3226 (2025). https://doi.org/10.1038/s41467-025-58589-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58589-6