Abstract
Small-conductance Ca2+-activated K+ (KCa2.1-KCa2.3) channels modulate neuronal and cardiac excitability. We report cryo-electron microscopy structures of the KCa2.2 channel in complex with calmodulin and Ca2+, alone or bound to two small molecule inhibitors, at 3.18, 3.50, 2.99 and 2.97 angstrom resolution, respectively. Extracellular S3-S4 loops in β-hairpin configuration form an outer canopy over the pore with an aromatic box at the canopy’s center. Each S3-S4 β-hairpin is tethered to the selectivity filter in the neighboring subunit by inter-subunit hydrogen bonds. This hydrogen bond network flips the aromatic residue (Tyr362) in the filter’s GYG signature by 180°, causing the outer selectivity filter to widen and water to enter the filter. Disruption of the tether by a mutation narrows the outer selectivity filter, realigns Tyr362 to the position seen in other K+ channels, and significantly increases unitary conductance. UCL1684, a mimetic of the bee venom peptide apamin, sits atop the canopy and occludes the opening in the aromatic box. AP14145, an analogue of a therapeutic for atrial fibrillation, binds in the central cavity below the selectivity filter and induces closure of the inner gate. These structures provide a basis for understanding the small unitary conductance and pharmacology of KCa2.x channels.
Similar content being viewed by others
Introduction
Small-conductance Ca2+-activated K+ (KCa2.x, also called SK) channels are critical modulators of neuronal and cardiac excitability. The three members of this subfamily, KCa2.1-KCa2.3, are characterized by a small unitary conductance (~9.5 pS in symmetrical K+1,2), Ca2+-calmodulin (CaM)-dependent gating3, and a unique pharmacology4 that distinguishes them from other ion channels. They are potently and selectively inhibited by venom peptides (apamin from bee venom; scyllatoxin and tamapin from scorpion venoms) and by small molecule agents (UCL1684, AP14145, and NS8593). In the nervous system, KCa2.x channels underlie the medium afterhyperpolarization, a spike-frequency adaptation required for normal neurotransmission5. Loss-of-function mutations of KCa2.2 channels cause tremors in rats6 and neurodevelopmental disorders including cerebellar ataxia in humans7,8, while gain-of-function mutations of KCa2.3 channels cause Zimmermann-Laband syndrome in humans9,10. In the heart, KCa2.x channels modulate the duration of the action potential11 and genome-wide association12,13, functional14, and clinical15 studies have firmly linked the channels to atrial fibrillation, the most prevalent cardiac arrhythmia. AP30663, an analog of AP14145 and a small molecule inhibitor of KCa2.x channels, was recently shown in a phase II clinical trial to cardiovert chronic atrial fibrillation to sinus rhythm (15). Despite their immense physiological and medical importance, high-resolution structural information does not exist for this class of K+ channels.
We used single-particle cryo-electron microscopy (cryo-EM) to determine structures of full-length KCa2.2_WT and KCa2.2_F244S mutant channels complexed to CaM in the Ca2+-activated state. The cryo-EM structures of the KCa2.2_WT complex bound to UCL1684 (apamin mimetic) and AP14145 (analog of the therapeutic AP30663) were also determined. Site-directed mutagenesis, molecular dynamics simulation (MDS), and electrophysiological studies were used to assess K+ permeation through the pore, measure unitary conductance, and verify binding configurations of pharmacological agents. Our studies provide a framework for understanding the small unitary conductance and the unique pharmacological properties of KCa2.x channels.
Results
Architecture of the Ca2+-bound apo_KCa2.2/CaM complex
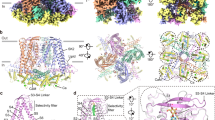
We purified full-length rat KCa2.2 in complex with rat CaM in the presence of 2 mM Ca2+ (Supplementary Fig. 1). Rat KCa2.2 is ~98% identical to human KCa2.2, and rat CaM is 100% identical to human CaM. Through cryo-EM single particle reconstruction, we obtained a 3D cryo-EM map of the Ca2+-bound apo_KCa2.2/CaM complex with a resolution of 3.18 Å (Supplementary Fig. 2, Supplementary Table 1). A total of 361 KCa2.2 residues (Ile118 to Asn478; remaining residues not visible) and 146 CaM residues (Asp2 to Ala147) were fitted into the density map to obtain a model of the Ca2+-bound apo_KCa2.2/CaM complex (referred to henceforth as apo_KCa2.2) (Fig. 1a). This complex, containing four KCa2.2 subunits and four CaM molecules in fourfold symmetry, is 90 Å long and 142 Å wide when viewed from the plane of the membrane (Fig. 1a). Side-chain densities are clearly resolved in the transmembrane S1-S6 helices, the extracellular S3-S4 loop, the inner S45A and S45B helices, the CaM-binding HA and HB C-terminal helices, and CaM (Supplementary Fig. 3a, b). KCa2.2 residues are visibly more resolved and have higher Q-scores16 than CaM residues, suggesting that the KCa2.2 structure is more stable than CaM (Supplementary Fig. 3c, d).
a Cryo-EM density map with fitted model of apo_KCa2.2 viewed from inside the cell (left), from the plane of the membrane (center), and from the extracellular side (right). Four KCa2.2 subunits are shown in blue, yellow, green and salmon, and CaM is shown in gray. Transmembrane helices S5 and S6 form the ion channel pore, which is surrounded by membrane-embedded helices S1 to S4. b Intracellular view of apo_KCa2.2 (blue/green) superimposed on apo_KCa3.1_1 (yellow/pink, PDB 6cnn) shows channel-CaM interactions that are similar in the two channels. In the cytoplasmic domain, the proximal C-termini of the two channels form HA (Lys402-Thr438 in KCa2.2; Lys293-Thr329 in KCa3.1) and HB (His446-Asn478 in KCa2.2; Ser334-Asn366 in KCa3.1) helices that lie almost parallel to the membrane plane. The HC helices, visible in apo_KCa3.1_1, are invisible in apo_KCa2.2 probably because of flexibility. c Side view of one subunit of apo_KCa2.2 (blue/green) superimposed on a subunit of apo_KCa3.1_1 (yellow/pink). The S3-S4 loop forms a β-hairpin in apo_KCa2.2 (highlighted in magenta), while the S3-S4 loop of apo_KCa3.1_1 is not resolved in PDB 6cnn. d Extracellular view of apo_KCa2.2 (blue) superimposed on apo_KCa3.1_1 (yellow). The four S3-S4 β-hairpins (magenta) form a canopy over the outside of the pore. Phe244 residues (shown as sticks) at the tips of four S3-S4 β-hairpins form an aromatic box with a central opening at the outer end of the pore.
Since the small-conductance KCa2.2 and the intermediate-conductance Ca2+-activated KCa3.1 channel share a Ca2+-CaM dependent mechanism of activation4 and ~40% amino acid sequence identity (Supplementary Fig. 4), we superimposed our apo_KCa2.2 structure onto the structure of the Ca2+-bound apo_KCa3.1/CaM activation state I17 (Protein Data Bank [PDB]: 6cnn) (referred to henceforth as apo_KCa3.1_1). In both Ca2+-bound structures, CaM interacts with helices in the S4-S5 loop (S45A) and the C-terminus (HA and HB) (Fig. 1b), suggesting that KCa2.2 and KCa3.1 share a similar Ca2+/CaM gating mechanism, confirming earlier electrophysiological and mutagenesis studies4.
Unique outer pore of the Ca2+-bound apo_KCa2.2/CaM complex
One unique feature of apo_KCa2.2 is the conformation of its unusually long extracellular loop between transmembrane helices S3 and S4 (S3–S4 loop, Supplementary Fig. 4). The S3-S4 loop in apo_KCa2.2 forms a β-hairpin configuration consisting of two antiparallel β-sheet strands with one turn between them (Fig. 1c, Supplementary Fig. 3). The four S3-S4 β-hairpins stretch over the selectivity filter forming an outer canopy (Fig. 1d). At the tips of these four β-hairpins, the side chains of Phe244 form an aromatic box with a central opening18 positioned directly above the selectivity filter (Fig. 1d).
Figure 2a shows the cryo-EM density map and the fitted model of the selectivity filter and the S3-S4 β-hairpins of the apo_KCa2.2 structure seen from the plane of the membrane. Both regions are well resolved. In apo_KCa2.2, each S3-S4 β-hairpin is tethered to the selectivity filter in the neighboring subunit by a network of inter-subunit hydrogen bonds (Fig. 2b), which is different from the intra-subunit network that stabilizes apo_KCa3.1_1’s pore (Fig. 2c). In apo_KCa2.2, Arg241, Phe244 and Tyr246 in the S3-S4 β-hairpin are tied by inter-subunit hydrogen bonds and a salt bridge to Asp364 in the selectivity filter (Fig. 2b). Interaction between the S3-S4 β-hairpins and the selectivity filter causes the critical tyrosine (Tyr362) in the G[Y/F]GD signature sequence to flip 180o relative to the position of corresponding aromatic residues in the filters of other Ca2+-activated and voltage-activated K+ channels (Fig. 2d). The 180o flip of Tyr362 relative to the aromatic residues in other K+ channels is also shown in the side view of the selectivity filters (Supplementary Fig. 5). The reorientation of Tyr362 causes the outer selectivity filter of KCa2.2 to be wider (Fig. 3a, b) than the outer selectivity filters of other K+ channels (Supplementary Fig. 5). In this widened outer selectivity filter, K+ is missing from S1 (Fig. 3a) and the entry of water partially hydrates K+ at S2 (Fig. 3b). The inner selectivity filter of apo_KCa2.2 is the same as apo_KCa3.1_1, and in both structures K+ is found at S3 and S4 (Fig. 3a). An additional K+, S0, is found at the opening of the aromatic box of apo_KCa2.2 (Fig. 3a). Analysis using the Hole program19 shows that apo_KCa2.2’s outer filter is wider than other Ca2+-activated and voltage-activated K+ channels, while its outer vestibule is narrower due to the aromatic box (Fig. 3c, d).
a Cryo-EM density map with fitted model of two subunits of apo_KCa2.2 viewed from the plane of the membrane. K+ is seen at positions S0 (at the aromatic box), S2, S3, and S4 within the filter; S1 is missing. Residues in the filter (blue) are highlighted. Extracellular S3-S4 β-hairpins (magenta) from two neighboring subunits are positioned on either side of K+ at S0 in (a, b). Note, the outer filter is widened at the level of Y362-G363. b Each S3–S4 loop is tethered to the filter in the neighboring subunit. Asp364 in the filter of subunit A (blue) forms inter-subunit hydrogen bonds and salt bridge with Arg241, Phe244, and Tyr246 in the S3-S4 loop of neighboring subunit B (green). In addition, an inter-subunit interaction between Trp351 (blue) and Tyr362 (green) in the filter stabilizes the pore. c In apo_KCa3.1_1, Asp255, Val256 and Trp242 (yellow) in the selectivity filter, corresponding to Asp364, Met365 and Trp351 in apo_KCa2.2, form intra-subunit hydrogen bonds. In addition, Tyr253 in apo_KCa3.1_1 (yellow), corresponding to Tyr362 in apo_KCa2.2, forms an inter-subunit hydrogen bond with Thr247 (green). d Extracellular view of the critical tyrosine (Tyr362) in the G[Y/F]G motif of the selectivity filter in apo_KCa2.2 (blue) overlaid on the corresponding aromatic residues in the selectivity filters of the open-conducting conformations of apo_KCa3.1_1 (Tyr253, yellow; PDB 6cnn), apo_KCa3.1_2 (Tyr253, green; PDB 6cno), KVChim (Tyr373, brown, PDB 2r9r), KCa1.1 (Phe279, purple, PDB 5tj6), and AlphaFold_KCa2.2 (Tyr362, gray, https://alphafold.ebi.ac.uk/entry/E9PSQ3). Note, Tyr362 in apo_KCa2.2 is flipped 180° compared to the critical aromatic residues in the other channels.
a Superimposed outer pores of apo_KCa2.2 (blue-magenta) and apo_KCa3.1_1 (yellow, PDB 6cnn) viewed from the plane of the membrane with two subunits shown. The locations of K+ in the selectivity filters of the two channels are compared. Water molecules are shown as small blue spheres. b Dimensions of the outer pore in apo_KCa2.2 are shown together with the locations of K+. Only two opposite subunits are shown (blue, green). The selectivity filter is widened at the level of Tyr362 and Gly361, above K+ at S2. The S3-S4 β-hairpins of two neighboring subunits (yellow and orange) are shown above the outer end of the selectivity filter. c Pore of apo_KCa2.2 generated by the “Hole” program is shown as a dotted surface. Two opposite subunits are shown. Pore radius: red, <1.15 Å; green, 1.15 to 2.30 Å; blue, >2.30 Å. d Pore radii plotted against distance from the extracellular surface for apo_KCa2.2 (blue), apo_KCa3.1_1 (orange, PDB 6cnn), apo_KCa3.1_2 (green, PDB 6cno), KCa1.1 (purple, PDB 5tj6), and KVChim (brown, PDB 2r9r). Apo_KCa2.2’s outer selectivity filter is wider than the other channels, while its outer vestibule is narrower due to the aromatic (Phe244) box.
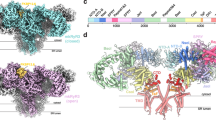
We replaced Phe244, one of the three residues tethering the S3-S4 β-hairpins to the selectivity filter, with the corresponding serine residue (KCa2.2_F244S, Supplementary Fig. 4) in the intermediate-conductance (39 pS unitary conductance in symmetrical K+ 20,21) KCa3.1 channel that lacks such tethers. We then determined the cryo-EM structure of the Ca2+-bound KCa2.2_F244S/CaM mutant complex (referred to henceforth as apo_KCa2.2_F244S) to a resolution of 3.50 Å with C4 symmetry (Supplementary Figs. 6 and 7, Supplementary Table 1). The cryo-EM density corresponding to the S3-S4 β-hairpins is absent from the apo_KCa2.2_F244S map (Fig. 4a). Unlike the S3-S4 β-hairpins tethered to the selectivity filter in apo_KCa2.2, the S3-S4 loops in apo_KCa2.2_F244S are invisible (Fig. 4b). While the selectivity filter of apo_KCa2.2 is stabilized by inter-subunit hydrogen bonds (Fig. 2b), the selectivity filter of apo_KCa2.2_F244S is stabilized by an intra-subunit network (Fig. 4c) similar to that of apo_KCa3.1_1’s pore (Fig. 2c). This rearrangement causes Tyr362 in the selectivity filter of apo_KCa2.2_F244S to flip 180° relative to the same residue in apo_KCa2.2 and align with Tyr253, the corresponding aromatic residue of apo_KCa3.1_1 (Fig. 4d). The apo_KCa2.2_F244S cryo-EM data suggest that the unique conformation of the selectivity filter in apo_KCa2.2 is caused by the tethering of the S3–S4 loops to the selectivity filter.
a Cryo-EM density map with fitted model of two opposite subunits of the apo_KCa2.2_F244S mutant. The cryo-EM density corresponding to the S3-S4 β-hairpins seen in apo_KCa2.2 is missing from apo_KCa2.2_F244S. b In the apo_KCa2.2_F244S mutant structure, the S3-S4 β-hairpins and aromatic box are not visible, and the selectivity filter is rearranged into the conformation of a canonical K+ channel selectivity filter. c In the apo_KCa2.2_F244S mutant structure, the selectivity filter is stabilized by intra-subunit hydrogen bonds. d The tyrosine (Y362, salmon) in the G[Y/F]G motif of the selectivity filter in apo_KCa2.2_F244S is flipped 180° relative to apo_KCa2.2 (Y362, blue), and realigns with KCa3.1_1 (Y253, yellow; PDB 6cnn). e Representative traces of single channel recordings of KCa2.2_WT and KCa2.2_F244S at −100 mV. f Representative amplitude histograms of KCa2.2_WT and KCa2.2_F244S. g Summary statistics of single channel conductance values. Data are presented as mean ± SD of single channel conductance values recorded from cells transfected with KCa2.2_WT (n = 14 cells) versus KCa2.2_F244S (n = 14 cells). ****P < 0.0001 compared with KCa2.2_WT (unpaired two-tailed t-test).
We examined whether the small unitary conductance of KCa2.2 was related to the interaction between the S3-S4 loop and the selectivity filter (Fig. 2b). In electrophysiological studies on HEK293 cells, we compared the unitary conductance of KCa2.2_WT versus KCa2.2_F244S. The unitary conductance of the KCa2.2_F244S mutant (22.1 ± 2.1 pS, n = 14) was significantly larger than that of KCa2.2_WT (9.9 ± 0.9 pS, n = 14) (Fig. 4e–g), suggesting that the widened outer pore caused by tethering of S3-S4 loop to the selectivity filter contributes to the small unitary conductance of KCa2.2.
We performed all-atom MDS to assess K+ permeation through the pores of apo_KCa2.2 and apo_KCa2.2_F244S (Supplementary Fig. 8). We measured K+ permeation events through both structures embedded in lipid bilayers in the presence of symmetric 300 mM KCl aqueous solution and a transmembrane voltage potential of 300 mV during 1-μs-long simulation runs (Supplementary movie 1 and 2, and Supplementary Fig. 8h, i). Restraints were applied to keep the inner gate open during these runs to determine the contribution of KCa2.2’s outer pore to unitary conductance. Under the same conditions, outward K+ permeation events are more readily observed through apo_KCa2.2_F244S than through apo_KCa2.2, even though the large variability among replicates prevented us from performing statistical analysis (Supplementary Fig. 8j). Large variability exists among replicate simulations probably due to the differences of initial configurations and the inherent stochastic nature of the simulations. Additionally, the complexity of these systems (apo_KCa2.2 and apo_KCa2.2_F244S), with many degrees of freedom, likely contributed to the variability observed in the MDS replicates. Nonetheless, the MDS results are qualitatively consistent with the experimentally measured unitary conductance values (Fig. 4g).
AlphaFold, the artificial intelligence-driven protein structure-prediction tool22, failed to predict the widened outer pore of apo_KCa2.2 resulting from the tethering of the S3-S4 loop to the selectivity filter. We compared the structural model of KCa2.2 predicted by AlphaFold (alphafold_KCa2.2) with our cryo-EM structure (apo_KCa2.2). AlphaFold correctly predicted the β-hairpin configuration of the S3-S4 loops and modeled them extending towards the pore (Supplementary Fig. 9a). However, AlphaFold_KCa2.2 lacks the tether between the S3-S4 β-hairpins and the selectivity filter, its outer selectivity filter is stabilized by an intra-subunit hydrogen bond network resembling that of apo_KCa3.1_1 (Fig. 2c, Supplementary Fig. 9b), and the conformation of its outer pore resembles that of other Ca2+-activated and voltage-activated K+ channels (Supplementary Fig. 9c). Furthermore, the central aromatic (Phe244) box, a key feature of apo_KCa2.2, is absent in AlphaFold_KCa2.2, with the four Phe244 residues positioned too far from each other to form the central aromatic box via cation-π interactions (Supplementary Fig. 9a)18. Our comparison of apo_KCa2.2 and AlphaFold_KCa2.2 highlights the usefulness of experimentally determined cryo-EM structures to understand the small unitary conductance of KCa2.2 channels.
Structure of KCa2.2/CaM bound to blocker UCL1684
To define the structural basis for the pharmacological inhibition of KCa2.x channels, we set out to determine structures of KCa2.2/CaM bound to inhibitors. Apamin, an 18-mer peptide from bee venom, is the archetypal inhibitor of KCa2.x channels23. Since addition of apamin caused aggregation of the KCa2.2/CaM protein complex, we studied UCL1684, a small molecule apamin-mimetic. UCL1684 blocks KCa2.2 at picomolar concentrations and is predicted to interact with the extracellular side of the channel overlapping the apamin binding site24,25,26.
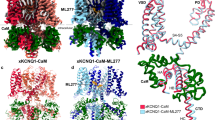
The cryo-EM map of the Ca2+- and UCL1684-bound KCa2.2/CaM complex (henceforth referred to as UCL1684_KCa2.2) was determined to a resolution of 2.99 Å with C4 symmetry (Supplementary Figs. 10 and 11, Supplementary Table 1). The cryo-EM density for UCL1684 is also clearly visible in the refined maps under C1 symmetry (Supplementary Fig. 12). The cryo-EM density map of UCL1684_KCa2.2 with fitted model shows UCL1684 on top of the canopy, close to the opening in the aromatic box (Fig. 5a, b). We successfully built a model for UCL1684 into its cryo-EM density due to the compound’s quaternary chemical structure (Supplementary Fig. 13a). The rigid cyclophane cage made up of two phenyl rings and two quinoline rings fits perfectly into the hydrophobic pocket formed by the aromatic (Phe244) box (Fig. 5a, b, Supplementary Fig. 13b). UCL1684 (500 nM) inhibited the KCa2.2_WT current effectively (Supplementary Fig. 13c, d). We used electrophysiology and mutagenesis to test UCL1684-KCa2.2 interactions predicted by the structure. Since UCL1684 does not inhibit KCa3.1, we replaced Phe244 in KCa2.2 with the corresponding residue from KCa3.1. The KCa2.2_F244S mutation abolished the blockade by UCL1684 without affecting the inhibition by the small molecule inhibitor, AP14145 (Fig. 5c–e). The mutation did not affect Ca2+-dependent channel activation (Supplementary Fig. 14). These results indicate that UCL1684 binds to the outer canopy and occludes the opening in the outer box.
a Cryo-EM density map refined under C4 symmetry with fitted model of UCL1684_KCa2.2 viewed from the plane of the membrane (left), and from the extracellular side (right). Four KCa2.2 subunits are shown in blue, yellow, green and salmon; CaM is shown in gray; and UCL1684 is shown in magenta. The densities for both the protein and UCL1684 are contoured at σ = 6. b One UCL1684 molecule fitted into its cryo-EM density (magenta) is positioned near the aromatic (Phe244) box at the outer end of the pore. c Representative whole-cell current traces of the KCa2.2_F244S mutant elicited by ramps from −120 mV to 40 mV. d The KCa2.2_F244S mutation abolishes blockade by UCL1684 (500 nM) but retains inhibition by AP14145 (30 μM). e Summary statistics show the KCa2.2_F244S mutant’s responses to UCL1684 and AP14145. Data are shown as mean ± SD of current density values recorded from cells transfected with KCa2.2_F244S (n = 8 cells) in response to the indicated treatments. ****P < 0.0001 compared with control (One-way ANOVA and Tukey’s post hoc tests without adjustments).
Guided by the UCL1684_KCa2.2 structure, we generated a docking model of apamin for heuristic purposes (Supplementary Fig. 15). The model shows apamin making contact with residues in the S3-S4 β-hairpins (Phe244, Ser245, Tyr246, Ala247). The essential RXCQ motif of apamin sits close to Asn368 (Supplementary Fig. 15). In support of the model, mutations of Ser245, Tyr246 and Ala247 have been reported to significantly alter the channel’s sensitivity to apamin23,27. Further, the RXCQ motif of Lei-Dab7, a derivative of a scorpion toxin and a potent inhibitor of KCa2.2, was reported to form a close contact with the residue in human KCa2.2 corresponding to Asn36828. These earlier studies provide support for the apamin docking model and suggest that peptide inhibitors from both bee venom and scorpion venom may make similar contacts with the outer canopy and aromatic box of KCa2.2.
Structure of KCa2.2/CaM bound to inhibitor AP14145
AP14145, a negative gating modulator of KCa2.2, inhibits the channel at low micromolar concentrations29. AP14145 effectively terminated atrial fibrillation in goats30 and even terminated vernakalant-resistant atrial fibrillation in pigs31. Its analog, AP3066315, effectively converted chronic atrial fibrillation to sinus rhythm in a phase II clinical trial. AP30663 was not commercially available when we performed the cryo-EM study. We determined the structure of KCa2.2/CaM bound to AP14145 in the presence of 2 mM Ca2+. The cryo-EM map of the Ca2+- and AP14145-bound KCa2.2/CaM complex (henceforth referred to as AP14145_KCa2.2) was determined to a resolution of 2.97 Å under C4 symmetry (Supplementary Figs. 16, 17 and Supplementary Table 1). The cryo-EM density of AP14145 is clearly visible below the selectivity filter in the central cavity near the side chains of both Ser359 and Ala384 (Fig. 6a, b). The cryo-EM density for AP14145 is also apparent in the refined maps under C1 symmetry (Supplementary Fig. 18). We could not delineate the precise orientation of AP14145 and the positioning of its sidechains within its KCa2.2 binding site because of symmetry mismatch arising from the nonsymmetrical AP14145 molecule (Supplementary Fig. 13a) binding to the fourfold symmetrical KCa2.2 channel. We used electrophysiology and mutagenesis to test AP14145’s predicted interactions with Ser359 and Ala384 in KCa2.2. Since AP14145 does not inhibit KCa3.1, we replaced Ser359 and Ala384 in KCa2.2 with the corresponding residues from KCa3.1. Corresponding double mutations in KCa2.3 channels have been used to explore the binding sites of negative gating modulators (NS859332 and AP1414529). AP14145 (30 μM) inhibited the KCa2.2_WT current effectively (Supplementary Fig. 13e, f), while the S359T + A384T double mutant abolished inhibition by AP14145 without affecting blockade by UCL1684 (Fig. 6c–e) or altering Ca2+-dependent channel activation (Supplementary Fig. 14). In summary, our cryo-EM, mutagenesis, and electrophysiology data together show binding of AP14145 near Ser359 and Ala384 in KCa2.2’s central cavity.
a Cryo-EM density map refined under C4 symmetry with fitted model of AP14145_KCa2.2 viewed from the plane of the membrane (left), and from the extracellular side (right). AP14145 density is shown in red. The densities for both the protein and AP14145 are contoured at σ = 6. b Cryo-EM density of AP14145 (red) viewed from the plane of the membrane is shown in the central cavity near Ser359 and Ala384. c Representative whole-cell current traces of the KCa2.2_S359T_A384V mutant elicited by ramps from −120 mV to 40 mV. d The KCa2.2_S359T_A384V double mutations abolish inhibition by AP14145 (30 μM) but retains blockade by UCL1684 (500 nM). e Summary statistics show the KCa2.2_S359T_A384V mutant’s responses to AP14145 and UCL1684. Data are shown as mean ± SD of current density values recorded from cells transfected with KCa2.2_S359T_A384V (n = 6 cells) in response to the indicated treatments. ****P < 0.0001 compared with control (One-way ANOVA and Tukey’s post hoc tests without adjustments).
A clue to how AP14145 inhibits the KCa2.2 channel came from superimposition of AP14145_KCa2.2 onto the apo_KCa2.2 and UCL1684_KCa2.2 structures (Fig. 7a, b). The inner gate defined by Val391 is ~12.4 Å in apo_KCa2.2, while the inner gate in AP14145_KCa2.2 is ~6.5 Å (Fig. 7a, b). In the Supplementary Movies 3 and 4, AP14145 is seen to narrow the inner gate through the movement of the transmembrane S6 helices and the rotation of Val391 side chains. The N-lobe of CaM moves closer toward the pore axis together with S45A helix, as a result of the binding of AP14145 (Supplementary Fig. 19, Supplementary Movies 3 and 4). The narrowing of the inner gate by AP14145 was also shown by comparing pore radii determined with the ‘Hole program’19 (Fig. 7c, d).
a Overlay of apo_KCa2.2 (blue), AP14145_KCa2.2 (orange) and UCL1684_KCa2.2 (green) viewed from the plane of the membrane. AP14145 was docked into the AP14145_KCa2.2 structure guided by the cryo-EM density. The inner gate is defined by Val391 in the transmembrane S6 helices. The inner gate is narrowed in AP14145_KCa2.2 to 6.5 Å compared to 12.3 Å in apo_KCa2.2 and UCL1684_KCa2.2. b In the intracellular view, the inner gate is narrowed in AP14145_KCa2.2 (orange) compared to apo_KCa2.2 (blue) and UCL1684_KCa2.2. c AP14145_KCa2.2’s pore generated by the “Hole” program is shown as a dotted surface with only two opposite subunits shown. Pore radius: red, <1.15 Å; green, 1.15 to 2.30 Å; blue, >2.30 Å. d The pore radii of apo_KCa2.2 (blue), AP14145_KCa2.2 (orange), and UCL1684_KCa2.2 (green) are plotted against distance from the extracellular surface. In this comparison, Phe244 defines the aromatic box formed by the S3-S4 β-hairpins, and Val391 defines the narrowest constriction site of the inner gate. The inner gate is narrowed in AP14145_KCa2.2 compared to the other two structures.
Discussion
Unlike any K+ channel structure determined thus far, apo_KCa2.2’s pore is covered on the outside by a canopy formed by the extracellular S3-S4 loops in a β-hairpin configuration. A network of inter-subunit hydrogen bonds ties the β-hairpins to the selectivity filter. This network flips the aromatic residue (Tyr362) in the G[Y/F]GD signature sequence by 180o, causing the outer selectivity filter to widen and water to enter the widened filter, displacing K+ at S1 and partially hydrating K+ at S2. Disruption of the tether in the apo_KCa2.2_F244S structure narrows the outer selectivity filter and reverts Tyr362 to the position seen in other K+ channels. The unitary conductance of KCa2.2_WT (9.9 ± 0.9 pS) was significantly smaller than that of KCa2.2_F244S (22.1 ± 2.1 pS), suggesting that the widened outer selectivity filter of apo_KCa2.2 contributes to the small unitary conductance of KCa2.2 channels. The structural features reported here are not predicted by the artificial intelligence-driven protein structure-prediction tool, AlphaFold22, highlighting the need for experimentally determined structures to understand the unique features of the KCa2.2 channel, particularly its small unitary conductance and pharmacology.
We determined the cryo-EM structures of KCa2.2 channels bound to two distinct inhibitors (UCL1684 and AP14145). UCL1684, an apamin mimetic, binds to the outer canopy and occludes the opening in the aromatic box. A mutation that removes the aromatic box, F244S, abolishes UCL1684 blockade (Fig. 5). A docking model of apamin based on the UCL1684_KCa2.2 structure shows the peptide sitting atop the canopy, occluding the aromatic box like UCL1684, and making several additional contacts both in the canopy and the outer pore (Supplementary Fig. 15). The docking configuration is consistent with and supported by earlier mutagenesis studies. Scorpion venom peptides (scyllatoxin, tamapin) that block KCa2.x channels may also bind to the canopy and close the opening in the aromatic box28. AP14145, a negative gating modulator, binds in the central cavity below the selectivity filter and induces closure of the inner gate (Fig. 7). AP3066315, its analog in clinical trials for the treatment of atrial fibrillation, likely binds to the same residues in the central cavity and induces a similar closure of the inner gate. It may be possible to use the AP14145_KCa2.2 structure to guide the design of new inhibitors of KCa2.2 that act by the same mechanism.
There is no cryo-EM structure for the other two members of the KCa2 subfamily, KCa2.1 and KCa2.3. Since the three KCa2.x channels exhibit greater than 70% sequence identity and share the Tyr246 residue that ties the S3-S4 β-hairpins to the selectivity filter in apo_KCa2.2 (Supplementary Fig. 4), the outer canopy and unique conformation of KCa2.2’s outer pore may also be a feature of human KCa2.1 and KCa2.3. Rat KCa2.1 may be an outlier. The presence of Leu213 in place of Tyr246 in rat KCa2.1 (Supplementary Fig. 4) may disrupt the critical tether between the S3-S4 β-hairpins and the selectivity filter, resulting in an altered pore conformation. In support, earlier functional and mutagenesis studies predicted that rat KCa2.1 exhibits a very different outer pore shape and sensitivity to inhibitors than other KCa2.x channels23.
The cryo-EM structures for the intermediate-conductance KCa3.1 channel17 and the large-conductance KCa1.133,34,35 have been previously reported. KCa2.x and KCa3.1 channels are activated via a Ca2+-CaM mediated mechanism, which is distinct from the gating mechanism of KCa1.1 channels33,34,35. While apo_KCa2.2 and apo_KCa3.1_1 (PDB 6cnn) are similar in the cytoplasmic domain accounting for their shared mechanism of Ca2+/CaM-dependent gating and sensitivity to pharmacological activators (e.g. 1-EBIO36 and SKA-3137), they exhibit significantly different architectures in the outer pores. This difference may explain why charybdotoxin, a scorpion peptide that blocks both the KCa3.1 channel38,39 and the large-conductance Ca2+-activated KCa1.140 channel by inserting a lysine residue into the selectivity filter, has no effect on KCa2.x channels. The canopy in KCa2.2 would prevent charybdotoxin from reaching its binding site in the outer vestibule. Differences in the outer pore may also contribute to the difference in unitary conductance between KCa2.x channels and KCa3.1 channels.
In conclusion, our structures provide a foundation for understanding the small unitary conductance of KCa2.x channels and define the mechanism of action of two distinct pharmacological inhibitors.
Methods
Protein expression
The cDNA of full-length rat KCa2.2 (accession no. NM_019314) with a C-terminus Strep-II tag was sub-cloned into pEG BacMam (a gift from Eric Gouaux; Addgene plasmid # 160451; http://n2t.net/addgene:160451; RRID:Addgene_160451). The amino acid sequence of the expressed KCa2.2 protein is as follows:
MSSCRYNGGVMRPLSNLSSSRRNLHEMDSEAQPLQPPASVVGGGGGASSPSAAAAASSSAPEIVVSKPEHNNSNNLALYGTGGGGSTGGGGGGGGGGGGSGHGSSSGTKSSKKKNQNIGYKLGHRRALFEKRKRLSDYALIFGMFGIVVMVIETELSWGAYDKASLYSLALKCLISLSTIILLGLIIVYHAREIQLFMVDNGADDWRIAMTYERIFFICLEILVCAIHPIPGNYTFTWTARLAFSYAPSTTTADVDIILSIPMFLRLYLIARVMLLHSKLFTDASSRSIGALNKINFNTRFVMKTLMTICPGTVLLVFSISLWIIAAWTVRACERYHDQQDVTSNFLGAMWLISITFLSIGYGDMVPNTYCGKGVCLLTGIMGAGCTALVVAVVARKLELTKAEKHVHNFMMDTQLTKRVKNAAANVLRETWLIYKNTKLVKKIDHAKVRKHQRKFLQAIHQLRSVKMEQRKLNDQANTLVDLAKTQNIMYDMISDLNERSEDFEKRIVTLETKLETLIGSIHALPGLISQTIRQQQRDFIETQMENYDKHVTYNAERSRSSSRRRRSSSTAPPTSSESSPSTWSHPQFEK
Un-tagged rat calmodulin (CaM) cDNA (accession no. BC063187) was also cloned into pEG BacMam. The amino acid sequence of the expressed CaM protein is as follows:
MADQLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLTMMARKMKDTDSEEEIREAFRVFDKDGNGYISAAELRHVMTNLGEKLTDEEVDEMIREADIDGDGQVNYEEFVQMMTAK
Bacmids for KCa2.2 and CaM were generated separately by transformation of DH10Bac E. coli cells with the pEG BacMam plasmids carrying rat KCa2.2 and rat CaM cDNAs, respectively. Baculoviruses of KCa2.2 and CaM were generated by transfection of Sf9 insect cell with bacmids using CellfectinTM II reagent (Gibco), respectively. Sf9 insect cells were culture in Sf-900TM II SFM medium (Gibco) at 27 °C. P2 baculoviruses after two rounds of amplification were used for cell transduction. KCa2.2 and CaM were co-expressed in HEK293S GnTI− cells (ATCC) using a BacMam method41. Briefly, HEK293S GnTI− cells were cultured in FreeStyle™ 293 expression media (Gibco) at 37 °C under 8% CO2 to a density of ~3 × 106 cells/ml, before transduction with KCa2.2 baculoviruses (10% v/v) and CaM baculoviruses (2% v/v). At 12 h after transduction, 10 mM sodium butyrate was added, and the temperature was changed to 30 °C. The cells were harvested at 72 h post transduction by centrifugation for 20 min at 6200 × g, 4 °C. Expression of the protein complex of KCa2.2_F244S mutant and wildtype CaM was performed using the same technique. The cell pellets were frozen at −80 °C until purification.
Protein purification
Protein purification was performed at 4 °C. Cell pellets from 4 L culture (~40 g) were thawed in hypotonic lysis buffer containing 50 mM Tris pH 8.0, 20 mM KCl, 0.5 mM MgCl2, 2 mM CaCl2, 0.05 mg/ml DNase, Halt™ protease inhibitor cocktail (Thermo Fisher) and 1 mM PMSF. Cell pellets were resuspended using a Dounce homogenizer and then stirred for 30 min at 4 °C. The resulting lysate was clarified by centrifugation at 39,800 × g for 30 min at 4 °C. The crude membrane preparation was resuspended with a Dounce homogenizer in solubilization buffer containing 50 mM Tris pH 8.0, 200 mM KCl, 2 mM CaCl2, 1.2% glyco-diosgenin (GDN), and Halt™ protease inhibitor cocktail for 2 h at 4 °C. The solubilized membranes were clarified by centrifugation at 39,800 × g for 30 min at 4 °C. The resulting supernatant was incubated with Strep-Tactin XT 4Flow resin (IBA LifeSciences) for 2 h. The resin was then washed with 20 column volumes of wash buffer containing 50 mM Tris pH 8, 150 mM KCl, 2 mM CaCl2, and 0.02% GDN. The protein complex was eluted with wash buffer supplemented with 50 mM biotin. The eluted protein was concentrated using an Amicon Ultra centrifugal filter (MWCO 100 kDa cutoff). The concentrated protein was subjected to size-exclusion chromatography (Superdex™ 200 Increase, Cytiva) equilibrated with 20 mM Tris pH 8, 150 mM KCl, 2 mM CaCl2, 0.01% lauryl maltose neopentyl glycol (LMNG), and 0.001% cholesteryl hemisuccinate (CHS) (Supplementary Fig. 1). The peak fractions at ~10.6 ml of elution volume were collected and concentrated to ~3 mg/ml. Purification of the protein complex of KCa2.2_F244S mutant and wildtype CaM was performed using the same technique.
Cryo-EM sample preparation, data collection and processing
The purified protein complexes were brought to the Stanford SLAC Cryo-EM Center (S2C2) for cryo-EM studies. For the determination of AP14145_KCa2.2 and UCL1684_KCa2.2 structures, saturating concentrations of AP14145 and UCL1484 were mixed with the KCa2.2/CaM protein complex 30 min before the grid preparation, respectively.
Cryo-EM grid preparation
Vitrobot Mark IV (ThermoFisher Scientific) was used to prepare grids at 4 °C and 100% humidity. 3.5 μl of purified protein was applied to a glow-discharged Quantifoil R1.2/1.3 300 mesh Copper grid. The grid was then blotted for 4.5 s before being plunged into liquid ethane.
Cryo-EM grid screening and data collection
Grids were screened on a Talos Arctica 200 kV transmission electron microscope (TEM). Dose fractionated movies were subsequently collected using EPU on a Titan Krios G3i (ThermoFisher Scientific) with a K3 detector (Gatan) and a BioQuantum energy filter at the pixel size of 0.86 Å. A total cumulative dose of ∼50 electrons per Å2 was used for recording the movies of 40 frames (1.25 electrons per Å2 per frame) with a defocus range of –1.3 to –2.3 μm. The statistics for data collections are summarized in Supplementary Table 1.
Cryo-EM image processing
For the apo_KCa2.2 dataset, preprocessing of micrographs (motion correction and CTF estimation) were carried out using CryoSparc42. About 2000 particles were manually picked and subjected to 2D classification to generate picking templates. Template-picked particles were cleaned up by multiple rounds of 2D classification. An ab initio reconstruction was performed, followed by heterogenous and non-uniform refinements to 3.60 Å resolution (C1 symmetry imposed), or 3.18 Å resolution (C4 symmetry imposed) in the CryoSparc42 program (Supplementary Fig. 2). To examine possible conformational heterogeneity, 3D classification using both the RELION program43 and the CryoSparc program was performed on the refined map generated with the CryoSparc42 program. Within the classes generated by 3D classification in the RELION and the CryoSparc programs, only one class contained secondary features of the transmembrane helices and the cytoplasmic domain. Because 3D classification did not improve resolution of the cryo-EM map, the cryo-EM map generated by non-uniform refinement at 3.18 Å resolution described above was used for model building.
The apo_KCa2.2_F244S, AP14145_KCa2.2, and UCL1684_KCa2.2 datasets were similarly processed using the CryoSparc program42. An ab initio reconstruction of apo_KCa2.2_F244S was performed, followed by heterogenous and non-uniform refinements to 3.50 Å resolution (C4 symmetry imposed) in the CryoSparc42 program (Supplementary Fig. 6). An ab initio reconstruction of UCL1684_KCa2.2 was performed, followed by heterogenous and non-uniform refinements to 3.22 Å resolution (C1 symmetry imposed), or 2.99 Å resolution (C4 symmetry imposed) in the CryoSparc42 program (Supplementary Fig. 10). An ab initio reconstruction of AP14145_KCa2.2 was also performed, followed by heterogenous and non-uniform refinements to 3.56 Å resolution (C1 symmetry imposed) or 2.97 Å resolution (C4 symmetry imposed) in the CryoSparc42 program (Supplementary Fig. 16). Resolutions were estimated using the gold standard criterion at the threshold of 0.143. The local resolution was calculated in CryoSparc.
Model building
Coordinates of the Ca2+-bound KCa3.1 activated state I (PDB: 6cnn; apo_KCa3.1_1) were used as the initial model. The initial model was manually docked into the cryo-EM density map and then adjusted in UCSF Chimera. Phenix.real_space_refine44 was used to build and refine the model. Model building was achieved using phenix44 and Coot45 iteratively. Models for Ca2+, K+, and UCL1684 were built by visual inspection of the shape of the density in Coot45 followed by refinement in phenix44. The statistics for model refinements are summarized in Supplementary Table 1. All structural graphics were generated using UCSF ChimeraX46.
Patch-clamp electrophysiology
Human Embryonic Kidney (HEK293) cells transiently transfected with the rat wildtype or mutant KCa2.2 cDNAs were used for manual patch-clamp experiments. Site-directed mutagenesis of KCa2.2_F244S and the KCa2.2_ S359T_A384T was performed on the rat KCa2.2 cDNA through molecular cloning services (Genscript). The wildtype and mutant cDNAs, constructed in the pIRES2-AcGFP1 vector (Clontech), were transfected into HEK293 cells by a calcium–phosphate method. Whole-cell and inside-out KCa2.2 currents were recorded 1–2 days after transfection, with an Axon200B amplifier (Molecular Devices) at room temperature.
The resistance of the patch electrodes ranged from 2 to 3 MΩ. For whole-cell recordings, the intracellular solution containing (in mM): 120 KCl, 10 Hepes (pH 7.2), 1.75 MgCl2, and 10 EGTA/KOH was mixed with CaCl2 to obtain the desired free Ca2+ concentrations, calculated using the MaxChelator software (https://somapp.ucdmc.ucdavis.edu/pharmacology/bers/maxchelator/webmaxc/webmaxcS.htm). The extracellular solution contained (in mM): 145 NaCl, 4 KCl, 10 Hepes (pH 7.4), 1 MgCl2, and 2 CaCl2.
For inside-out recordings, the intracellular solution containing (in mM): 140 KCl, 10 Hepes (pH 7.2), 1 EGTA, 0.1 Dibromo-BAPTA, and 1 HEDTA was mixed with Ca2+ to obtain the desired free Ca2+ concentrations, calculated using the MaxChelator software. The extracellular solution contained (in mM): 140 KCl, 10 Hepes (pH 7.4), 1 MgSO4.
AP14145 (N-[2-[[(1R)-1-[3-(Trifluoromethyl)phenyl]ethyl]amino]-1H-benzimidazol-7-yl]acetamide hydrochloride) was purchased from Tocris Bioscience. AP14145 dilutions were prepared freshly in extracellular solution from 100 mM stock solutions in DMSO.
UCL1684 (6,12,19,20,25,26-Hexahydro-5,27:13,18:21,24-trietheno-11,7-metheno-7H-dibenzo [b,n] [1,5,12,16]tetraazacyclotricosine-5,13-diium dibromide) was purchased from Tocris Bioscience. UCL1684 dilutions were prepared freshly in extracellular solution from 5 mM stock solutions in DMSO. Final DMSO concentrations never exceeded 0.1% (v/v).
pClamp 10.5 (Molecular Devices) was used for data acquisition and analysis. To characterize the responses of the KCa2.2_F244S and the KCa2.2_ S359T_A384T mutants to AP14145 and UCL1684, whole-cell recordings were performed. Seals (>1 GΩ) were formed before the whole-cell configuration was obtained. Currents were elicited by repetitive 1 s voltage ramps from − 120 mV to 40 mV, with a holding potential of −70 mV applied every 5 s. Currents were filtered at 2 kHz and digitized at a sampling frequency of 10 kHz.
To measure the Ca2+ -dependent activation of the KCa2.2_F244S and the KCa2.2_ S359T + A384T mutants, inside-out patch recordings were performed. Seals (>1 GΩ) were formed before the inside-out patch configuration was obtained. The intracellular face was initially exposed to a zero-Ca2+ bath solution, and subsequently to a series of bath solutions with varying Ca2+ concentrations. Currents were recorded by repetitive 1 s voltage ramps from − 100 mV to 100 mV from a holding potential of 0 mV. One minute after switching of bath solutions, ten sweeps with a 1 s interval were recorded. To construct the concentration-dependent positive modulation of channel activities, the current amplitudes at −90 mV in response to various concentrations of Ca2+ were normalized to that obtained at 10 μM of Ca2+. The normalized currents were plotted as a function of the concentrations of Ca2+. EC50 values and Hill coefficients were determined by fitting the data points to a standard concentration–response curve.
Data analysis was performed using pClamp 10.5 (Molecular Devices) in a blinded fashion. Concentration-response curves were analyzed in GraphPad Prism 9.0.2 (GraphPad Software Inc.). All data are shown as mean ± SD unless otherwise indicated. One-way ANOVA and Tukey’s post hoc tests were used for data comparison of three or more groups. The unpaired Student’s t-test (two-sided) was used for data comparison if there were only two groups. Figures were made using GraphPad Prism 9.0.2 (GraphPad Software Inc.).
KCa2.2 single-channel recordings
KCa2.2_WT and KCa2.2_F244S mutant cDNAs were transfected into HEK293T cells using the PEI transfection reagent system, respectively. Single-channel currents were recorded from inside-out patches ~12 h after transfection. The extracellular solution (pipette solution) contained (in mM): 140 KCl, 10 Hepes (pH 7.4), 1 MgSO4. The intracellular solution (bath solution) containing (in mM): 140 KCl, 10 Hepes (pH 7.2), 1 EGTA, and 1 HEDTA was mixed with Ca2+ to obtain free Ca2+ of 0.2 µM, calculated using the MaxChelator software. Borosilicate glass electrodes (2-3 MΩ) were pulled with a Narishige PC-100 pipette puller and polished with a Narishige MF-830 micro forge. Currents were filtered at 100 kHz and acquired using a Tecela Pico 2 system controlled by WinWCP software (Strathclyde Electrophysiology Software) at -100 mV step from a holding potential of -60 mV. Data were digitally filtered at 1 kHz before analysis. Single channel amplitudes were measured directly from the single channel activity recorded in the excised patches containing a single channel during the voltage step. The average amplitudes were estimated from full channel openings.
MD simulations
The protonation states of titratable residues in the apo_KCa2.2 and apo_KCa2.2_F244S structures were determined using the H++ server (http://biophysics.cs.vt.edu/)47. The apo_KCa2.2 and apo_KCa2.2_F244S structures were incorporated into an explicit lipid bilayer consisting of POPC, POPE, POPS, and cholesterol in a molecular ratio of 25:5:5:148, along with a water box with dimensions of 170.5 Å × 170.5 Å × 131.7 Å. This setup was achieved using the CHARMM-GUI Membrane Builder webserver (http://www.charmm-gui.org/?doc=input/membrane)49. Ca2+ ions were included at the two EF-hands of the CaM N-lobe according to the apo_KCa2.2 cryo-EM structure. No Ca2+ ion was included at the CaM C-lobe. To the systems, 300 mM KCl and additional neutralizing counter ions were added. The total number of atoms in the systems were 357,302 and 357,214 for apo_KCa2.2 and apo_KCa2.2_F244S, respectively. Parameter and coordinate files were generated using the tleap (AMBER) program, employing the ff14SB and Lipid17 force fields for proteins and lipids, respectively. MD simulations were conducted using the PMEMD.CUDA program in AMBER 20 with periodic boundary conditions, creating isothermal-isobaric ensembles. A voltage of 0.09 V/nm was added to the systems from the intracellular to the extracellular side (~300 mV membrane potential). Long-range electrostatics were calculated using the particle mesh Ewald (PME) method50 with a 10 Å cutoff. Prior to production runs, energy minimizations were performed. The system was then heated from 0 K to 303 K using Langevin dynamics with a collision frequency of 1 ps−1. During heating, the apo_KCa2.2 and apo_KCa2.2_F244S structures were position-restrained with an initial constant force of 500 kcal/mol/Å2, gradually reduced to 10 kcal/mol/Å2 to allow lipid and water molecules freedom of movement. Subsequently, the systems underwent 5 ns equilibrium MD simulations. Finally, a total of 1 µs production MD simulations were conducted (with a position-restrained force of 50 kcal/mol/Ų applied to residues Val390, Val391, and Ala392 to prevent the inner gate from closing) at Argonne Leadership Computing Facility, saving coordinates every 100 ps for subsequent analysis. GROMCAS analysis tools were employed for trajectory analysis of the MD simulations51.
2D interaction diagram of UCL1684 and the KCa2.2 channel
The 2D diagram depicting the interactions between UCL1684 (Supplementary Fig. 13b) and the KCa2.2 channel was generated using the UCL1684_KCa2.2 structure in the Discovery Studio program (Dassault Systemes Biovia LLC).
Molecular docking of apamin into the outer pore of the KCa2.2 channel
We docked apamin into the KCa2.2 channel structure using the Glide program within the Schrödinger software (Schrödinger Inc.). The grid box was defined based on the apamin mimetic UCL1684’s binding site, with a box size of 32 Å x 32 Å x 32 Å. For docking simulations, we utilized a total of 10 NMR apamin structures from the database (PDB: 7oxf). The top-ranked pose of apamin within the binding site, determined by XP score, was selected. Subsequently, we performed Induced Fit Docking (IFD) docking simulations (Schrödinger Inc.) to account for the flexibility of both the channel and the toxin. The top XP score-ranked pose was then subjected to a 200-step energy minimization using the smart minimizer and the Generalized Born implicit membrane (GBIM) in the Discovery Studio program (Dassault Systemes Biovia LLC).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary information files, and available from the corresponding author upon request. The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 8v2g [https://doi.org/10.2210/pdb8v2g/pdb] (apo_KCa2.2); 9eio [https://doi.org/10.2210/pdb9eio/pdb] (apo_KCa2.2_F244S); 8v3g [https://doi.org/10.2210/pdb8v3g/pdb] (UCL1684_KCa2.2); and 8v2h [https://doi.org/10.2210/pdb8v2h/pdb] (AP14145_KCa2.2). The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-42911 (apo_KCa2.2); EMDB-48088 (apo_KCa2.2_F244S); EMD-42947 (AP14145_KCa2.2); and EMD-42914 (UCL1684_KCa2.2). The following previously published PDB codes are referred to: 6cnn [https://doi.org/10.2210/pdb6cnn/pdb]; 6cno [https://doi.org/10.2210/pdb6cno/pdb]; 2r9r [https://doi.org/10.2210/pdb2r9r/pdb]; and 5tj6 [https://doi.org/10.2210/pdb5tj6/pdb]. The source data underlying Figs. 4g, 5e, 6e, and Supplementary Fig. 8j, 13d, 13f, 14c are provided as a Source Data file. Source data are provided with this paper.
References
Kohler, M. et al. Small-conductance, calcium-activated potassium channels from mammalian brain. Science 273, 1709–1714 (1996).
Li, W. & Aldrich, R. W. Electrostatic influences of charged inner pore residues on the conductance and gating of small conductance Ca2+ activated K+ channels. Proc. Natl Acad. Sci. USA 108, 5946–5953 (2011).
Xia, X. M. et al. Mechanism of calcium gating in small-conductance calcium-activated potassium channels. Nature 395, 503–507 (1998).
Brown, B. M., Shim, H., Christophersen, P. & Wulff, H. Pharmacology of small- and intermediate-conductance calcium-activated potassium channels. Annu. Rev. Pharmacol. Toxicol. 60, 219–240 (2020).
Adelman, J. P., Maylie, J. & Sah, P. Small-conductance Ca2+-activated K+ channels: form and function. Annu. Rev. Physiol. 74, 245–269 (2012).
Kuramoto, T. et al. Tremor dominant Kyoto (Trdk) rats carry a missense mutation in the gene encoding the SK2 subunit of small-conductance Ca(2+)-activated K(+) channel. Brain Res. 1676, 38–45 (2017).
Balint, B. et al. KCNN2 mutation in autosomal-dominant tremulous myoclonus-dystonia. Eur. J. Neurol. 27, 1471–1477 (2020).
Mochel, F. et al. Variants in the SK2 channel gene (KCNN2) lead to dominant neurodevelopmental movement disorders. Brain 143, 3564–3573 (2020).
Bauer, C. K. et al. Gain-of-function mutations in KCNN3 encoding the small-conductance Ca(2+)-activated K(+) channel SK3 Cause Zimmermann-Laband Syndrome. Am. J. Hum. Genet. 104, 1139–1157 (2019).
Orfali, R. et al. Channelopathy-causing mutations in the S(45)A/S(45)B and HA/HB helices of K(Ca)2.3 and K(Ca)3.1 channels alter their apparent Ca(2+) sensitivity. Cell Calcium 102, 102538 (2022).
Zhang, Q. et al. Functional roles of a Ca2+-activated K+ channel in atrioventricular nodes. Circ. Res. 102, 465–471 (2008).
Ellinor, P. T. et al. Common variants in KCNN3 are associated with lone atrial fibrillation. Nat. Genet. 42, 240–244 (2010).
Christophersen, I. E. et al. Large-scale analyses of common and rare variants identify 12 new loci associated with atrial fibrillation. Nat. Genet. 49, 946–952 (2017).
Heijman, J. et al. Enhanced Ca(2+)-dependent SK-channel gating and membrane trafficking in human atrial fibrillation. Circ Res. 132, e116–e133 (2023).
Holst, A. G. et al. Inhibition of the K(Ca)2 potassium channel in atrial fibrillation: a randomized phase 2 trial. Nat Med. 30, 106–111 (2024).
Pintilie, G. & Chiu, W. Validation, analysis and annotation of cryo-EM structures. Acta Crystallogr. D Struct. Biol. 77, 1142–1152 (2021).
Lee, C. H. & MacKinnon, R. Activation mechanism of a human SK-calmodulin channel complex elucidated by cryo-EM structures. Science 360, 508–513 (2018).
Davis, M. R. & Dougherty, D. A. Cation-pi interactions: computational analyses of the aromatic box motif and the fluorination strategy for experimental evaluation. Phys. Chem. Chem. Phys. 17, 29262–29270 (2015).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996). 376.
Ishii, T. M. et al. A human intermediate conductance calcium-activated potassium channel. Proc. Natl Acad. Sci. USA 94, 11651–11656 (1997).
Grissmer, S., Nguyen, A. N. & Cahalan, M. D. Calcium-activated potassium channels in resting and activated human T lymphocytes. Expression levels, calcium dependence, ion selectivity, and pharmacology. J. Gen. Physiol. 102, 601–630 (1993).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Weatherall, K. L., Seutin, V., Liegeois, J. F. & Marrion, N. V. Crucial role of a shared extracellular loop in apamin sensitivity and maintenance of pore shape of small-conductance calcium-activated potassium (SK) channels. Proc. Natl Acad. Sci. USA 108, 18494–18499 (2011).
Rosa, J. C., Galanakis, D., Ganellin, C. R., Dunn, P. M. & Jenkinson, D. H. Bis-quinolinium cyclophanes: 6,10-diaza-3(1,3),8(1,4)-dibenzena-1,5(1,4)- diquinolinacyclodecaphane (UCL 1684), the first nanomolar, non-peptidic blocker of the apamin-sensitive Ca(2+)-activated K+ channel. J. Med. Chem. 41, 2–5 (1998).
Strobaek, D., Jorgensen, T. D., Christophersen, P., Ahring, P. K. & Olesen, S. P. Pharmacological characterization of small-conductance Ca(2+)-activated K(+) channels stably expressed in HEK 293 cells. Br. J. Pharmacol. 129, 991–999 (2000).
Fanger, C. M. et al. Calcium-activated potassium channels sustain calcium signaling in T lymphocytes. Selective blockers and manipulated channel expression levels. J. Biol. Chem. 276, 12249–12256 (2001).
Nolting, A., Ferraro, T., D’Hoedt, D. & Stocker, M. An amino acid outside the pore region influences apamin sensitivity in small conductance Ca2+-activated K+ channels. J. Biol. Chem. 282, 3478–3486 (2007).
Shakkottai, V. G. et al. Design and characterization of a highly selective peptide inhibitor of the small conductance calcium-activated K+ channel, SkCa2. J. Biol. Chem. 276, 43145–43151 (2001).
Simo-Vicens, R. et al. A new negative allosteric modulator, AP14145, for the study of small conductance calcium-activated potassium (K(Ca) 2) channels. Br. J. Pharmacol. 174, 4396–4408 (2017).
Gatta, G. et al. Effective termination of atrial fibrillation by SK channel inhibition is associated with a sudden organization of fibrillatory conduction. Europace 23, 1847–1859 (2021).
Diness, J. G. et al. Termination of vernakalant-resistant atrial fibrillation by inhibition of small-conductance Ca(2+)-activated K(+) channels in pigs. Circ. Arrhythm. Electrophysiol. 10, e005125 (2017).
Jenkins, D. P. et al. Negative gating modulation by (R)-N-(benzimidazol-2-yl)-1,2,3,4-tetrahydro-1-naphthylamine (NS8593) depends on residues in the inner pore vestibule: pharmacological evidence of deep-pore gating of K(Ca)2 channels. Mol. Pharmacol. 79, 899–909 (2011).
Tao, X., Hite, R. K. & MacKinnon, R. Cryo-EM structure of the open high-conductance Ca(2+)-activated K(+) channel. Nature 541, 46–51 (2017).
Raisch, T. et al. Small molecule modulation of the Drosophila Slo channel elucidated by cryo-EM. Nat. Commun. 12, 7164 (2021).
Tao, X., MacKinnon, R. Molecular structures of the human Slo1 K(+) channel in complex with beta4. Elife 8. https://doi.org/10.7554/eLife.51409 (2019).
Pedarzani, P. et al. Specific enhancement of SK channel activity selectively potentiates the afterhyperpolarizing current I(AHP) and modulates the firing properties of hippocampal pyramidal neurons. J. Biol. Chem. 280, 41404–41411 (2005).
Sankaranarayanan, A. et al. Naphtho[1,2-d]thiazol-2-ylamine (SKA-31), a new activator of KCa2 and KCa3.1 potassium channels, potentiates the endothelium-derived hyperpolarizing factor response and lowers blood pressure. Mol. Pharmacol. 75, 281–295 (2009).
Rauer, H. et al. Structure-guided transformation of charybdotoxin yields an analog that selectively targets Ca(2+)-activated over voltage-gated K(+) channels. J. Biol. Chem. 275, 1201–1208 (2000).
Bychkov, R. et al. Characterization of a charybdotoxin-sensitive intermediate conductance Ca2+-activated K+ channel in porcine coronary endothelium: relevance to EDHF. Br. J. Pharmacol. 137, 1346–1354 (2002).
Miller, C., Moczydlowski, E., Latorre, R. & Phillips, M. Charybdotoxin, a protein inhibitor of single Ca2+-activated K+ channels from mammalian skeletal muscle. Nature 313, 316–318 (1985).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D. Struct. Biol. 74, 531–544 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Gordon, J. C. et al. H++: a server for estimating pKas and adding missing hydrogens to macromolecules. Nucleic Acids Res. 33, W368–371 (2005).
Leal-Pinto, E., London, R. D., Knorr, B. A. & Abramson, R. G. Reconstitution of hepatic uricase in planar lipid bilayer reveals a functional organic anion channel. J. Membr. Biol. 146, 123–132 (1995).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Darden, T. A. & Pedersen, L. G. Molecular modeling: an experimental tool. Environ. Health Perspect. 101, 410–412 (1993).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Acknowledgements
M.Z. was supported by National Institutes of Health (Grant 4R33NS101182-03 and Grant 1R15NS130420), American Heart Association (Grant 23AIREA1039423), Chapman University Institutional FGRSC Grant, and National Ataxia Foundation (YI-SCA) grant. Y.W.N was supported by American Heart Association (24CDA1260237). D.E.L. was supported by National Institutes of Health (Grant R01HL059949-26 and Grant R01NS131467-02). Some of this work was performed at the Stanford-SLAC Cryo-EM Center (S2C2), which is supported by the National Institutes of Health Common Fund Transformative High-Resolution Cryo-Electron Microscopy program (U24 GM129541). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The authors would also like to thank the following S2C2 personnel for their invaluable support and assistance: Patrick Mitchell, Alexandre Cassago, Nathan D. Burrows, Chensong Zhang, and Patrick J. Pascual. The computations were supported by the ITS (Information Technology Services) Research Computing at Northeastern University and the Argonne Leadership Computing Facility (ALCF) at Argonne National Laboratory. We thank Drs. Michael Cahalan, George Augustine, and Vikram Shakkottai for reading the manuscript and providing critical feedback. We thank Drs. Jifang Zhang, Montserrat Samso, and Shashi Bhushan for discussions. We are grateful to Nadeed Naguib, Elyn Lam, Nikita Dave, and Noble Chea for technical assistance.
Author information
Authors and Affiliations
Contributions
M.C., D.E.L., H.W., E.Y., K.G.C., and M.Z conceptualized the project. Y.W.N., A.R., and M.Z. undertook protein expression and purification. Y.W.N., D.I., M.L.T., C.C.S., Y.L., G.P., and M.Z. undertook cryo-EM studies. A.S.C.G., H.M.N., A.R., and R.O. undertook electrophysiology studies. M.C. performed computational studies. K.G.C. and M.Z. wrote the first draft of the manuscript. All authors contributed to the manuscript and the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Koichi Nakajo who co-reviewed with Go Kasuya; Baobin Li, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nam, YW., Im, D., Garcia, A.S.C. et al. Cryo-EM structures of the small-conductance Ca2+-activated KCa2.2 channel. Nat Commun 16, 3690 (2025). https://doi.org/10.1038/s41467-025-59061-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59061-1
This article is cited by
-
Structural basis for the subtype-selectivity of KCa2.2 channel activators
Nature Communications (2026)