Abstract
Organosulfur compounds (OrgS) are fundamental components of life’s biomass, yet the cycling of these compounds in the terrestrial deep subsurface, one of Earth’s largest ecosystems, has gone relatively unexplored. Here, we show that all subsurface microbial genomes reconstructed from Soudan Underground Mine State Park have the capacity to cycle organic sulfur species. Our findings suggest that OrgS degradation may be an integral link between the organic and inorganic sulfur cycle via the production of sulfite and sulfide. Furthermore, despite isolation from surface ecosystems, most Soudan microorganisms retained genes for dimethylsulfoniopropionate and taurine biosynthesis. Metagenomic analyses of an additional 54 deep subsurface sites spanning diverse lithologies revealed the capacity for OrgS cycling to be widespread, occurring in 89% of assembled metagenomes. Our results indicate that consideration of OrgS cycling may be necessary to accurately constrain sulfur fluxes, discern the energetic limits of deep life, and determine the impact of deep subsurface biogeochemical sulfur cycling on greater Earth system processes.
Similar content being viewed by others
Introduction
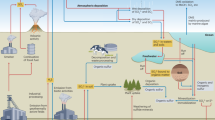
The biogeochemical cycling of organic and inorganic sulfur (S) compounds is tightly intertwined and central to supporting microbial life through S uptake and incorporation into biomass or redox couplings that drive metabolic processes. These microbially-driven sulfur transformations regulate biogeochemical S cycling and couple with other globally important cycles such as Fe, C, and N1,2,3,4,5. Organosulfur compounds (OrgS) have been increasingly recognized as a central aspect of biogeochemical sulfur cycling in terrestrial and marine environments6,7,8. Investigations of OrgS cycling in low-sulfate systems such as Archean oceans9 and modern Lake Superior10 have demonstrated the importance of OrgS in the total sulfur budget and as an inorganic S source that drives chemoautotrophic or dissimilatory metabolisms. Given that microbial assimilation and mineralization of OrgS directly control the availability of inorganic S substrates (Fig. 1), considering these processes in tandem is paramount to obtaining a comprehensive understanding of biogeochemical sulfur cycling and, more broadly, the behavior of Earth’s Critical Zone across modern and deep time.
A simplified diagram showing key transformations in the cycling of inorganic and organic sulfur. A selection of diverse, metabolically and environmentally important organosulfur compounds produced intracellularly is shown. DMS, dimethylsulfide; DMSO, dimethylsulfoxide; DMSP, dimethylsulfoniopropionate; Me, methyl group.
The microbial production of OrgS compounds is ubiquitous in marine11,12,13, freshwater14, and terrestrial systems15, where they serve as building blocks for biomass16,17, osmoprotectants18,19,20, cryoprotectants21, protection against oxidative stress22, or as terminal electron acceptors in dissimilatory processes. Production of many OrgS compounds, such as dimethylsulfoniopropionate (DMSP) and taurine, was historically thought to be constrained to eukaryotes and is dominated by eukaryotes in extant marine ecosystems16,20,23. However, the identification of novel DMSP and taurine biosynthesis pathways in heterotrophic bacteria20,24,25, and evidence of these pathways occurring alongside mineralization pathways26, provide a mechanism for OrgS cycling to persist in prokaryote-dominated communities and expand the environments in which OrgS is expected to play an essential role.
The terrestrial crust is one of Earth’s largest ecosystems, hosting 2–19% of total global biomass27,28,29. This biomass pool requires the synthesis of OrgS compounds, like cysteine, methionine, and coenzyme A, for growth. Likewise, as these organisms die and release OrgS extracellularly, their removal through reincorporation or mineralization is inevitable. However, the importance of OrgS cycling in crustal environments has received little attention despite the potential for OrgS to fuel metabolism in such settings. Several studies have suggested the occurrence of linked organic and inorganic S processes in these vast subsurface environments. Modeling-based approaches by Fakhraee and colleagues6 demonstrate that OrgS mineralization could support deep biosphere communities of inorganic S reducers. Recently, sediments underlying low-sulfate waters (<30 μM) were observed to have increased sulfate concentrations fueled by in situ sulfate production via OrgS degradation10. This mechanism of sulfate production was supported by sediment incubation experiments that demonstrated sulfate production coupled to the degradation of OrgS compounds (i.e., taurine, sodium dodecyl sulfate, cystine, and methionine)10. Finally, the ability of sulfate-reducing bacteria to ferment OrgS compounds like cysteate to acetate, ammonia, sulfide, and sulfate30 suggests these compounds are important substrates capable of fueling life. Although OrgS cycling genes, primarily involved in dimethylsulfoxide (DMSO) and taurine transport, have been observed in terrestrial deep biosphere studies31,32,33, our understanding of the extent and nature of microbial OrgS cycling in these systems is limited.
Inorganic S cycling is well known to play a key role in deep terrestrial subsurface systems34,35,36,37,38,39 and past studies have detected the presence of some OrgS cycling genes in these systems31,32,33. Therefore, we hypothesize that the cycling of OrgS compounds, such as DMSP and taurine, could serve as a key regulator of the inorganic S pool, and thus microbial metabolism, in the deep biosphere. A more extensive understanding of the functional capacity of the deep biosphere to cycle OrgS and the entwined nature of inorganic and organic biogeochemical S cycles encoded in these metagenomes will elucidate the role of OrgS in deep terrestrial subsurface systems. Here, we address this gap using shotgun metagenomic sequencing of anoxic groundwater and suboxic to oxic outflow channel fluids at Soudan Underground Mine State Park, as direct quantification of OrgS is difficult in these highly saline waters. Metagenomic insights into this microbial community reveal the genetic potential for widespread OrgS cycling, including pathways that link organic and inorganic sulfur cycling, and suggest that OrgS could play a significant role in supporting deep life. Further, we examine 54 additional terrestrial subsurface metagenomes to explore the ubiquity and diversity of OrgS cycling genes and demonstrate the widespread genetic potential for deep subsurface OrgS cycling. This work extends our understanding of biogeochemical S cycling in the deep terrestrial subsurface by underscoring the potential role of OrgS as a source and sink of inorganic sulfur and positing that microbial cycling of OrgS is likely globally significant.
Results and discussion
Microbial synthesis and utilization of OrgS likely occurred early in Earth’s history40 and has diversified with time, resulting in extant organisms capable of intracellularly cycling OrgS compounds with varying carbon structures and sulfur redox potentials (Fig. 1). Furthermore, the sulfurization of labile organic molecules41 creates a diverse pool of substrates for microorganisms to utilize. Synthesis of microbial biomass, including OrgS compounds such as cysteine, is typically a net-exergonic process under reducing conditions when oxidized S and N sources are available42. OrgS production through biomass creation is expected in deep subsurface systems such as Soudan Mine, where groundwaters are typified by their low reduction potential and the presence of sulfate41,43,44. Indeed, spectroscopic detection of reduced and oxidized OrgS in biofilm-associated mineral particulates from the Soudan Mine provides further evidence of OrgS synthesis (Fig. S2). OrgS catabolism is also expected, as the release of labile carbon backbones, such as the generation of pyruvate via cysteine degradation, provides substrates for central carbon metabolism45. Additionally, OrgS mineralization can generate either sulfite or sulfide, depending on the starting compound and pathway (Fig. 2a). These inorganic S species sourced from organosulfur have been demonstrated to feed dissimilatory redox reactions and drive cellular energy production46,47. Thus, the cyclical metabolism of OrgS is predicted to play a central role in biogeochemical S cycling in diverse terrestrial subsurface systems.
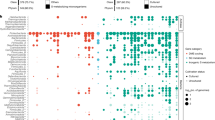
Overview of proposed linked inorganic and organic sulfur cycles at Soudan Mine. a Schematic diagram depicting key intracellular pathways for dissimilation of inorganic sulfur (pink), transformation of taurine (blue), transformation of dimethylsulfide, dimethylsulfoxide, or dimethylsulfoniopropionate (DMS(O)(P), green), and cysteine and methionine production and utilization (yellow). Reactions show only key sulfur species, and not all intermediate steps are shown. Genes encoding the enzyme that catalyzes each reaction are indicated in italics, and the percentage of Soudan Mine metagenome assembled genomes (MAGs) able to carry out that reaction is shown in bold. b Taxonomic distribution of key genes found in the Soudan Mine. Phyla are listed based on evolutionary relationships, with the number of MAGs in that phylum in parentheses. Solid squares indicate the presence of the indicated gene or complete gene complex, while cross-hatched squares represent the presence of some but not all genes in the complex. The bottom row indicates the total number of MAGs in which that gene or group of genes was detected. See Supplementary Data S1 for gene abbreviations and Supplementary Data S2 for detailed reactions. Asterisks indicate genes that were identified using HMSS2 rather than METABOLIC. To see gene hits from METABOLIC, see Supplementary Data S7 (MAGs) and S8 (assemblies).
While pathways generating both sulfide and sulfite appear in the Soudan Mine metagenome (Fig. 2a), the cycling of sulfite-producing compounds is predicted to play a more important ecological role. Sulfide released via mineralization of cysteine or methionine may serve as an endpoint in Soudan groundwaters because it is quickly scavenged by and stably bound to reduced iron48. Conversely, compounds producing sulfite upon microbial decomposition, such as DMSO and taurine, are of interest because sulfite can be biotically oxidized to sulfate to generate ATP. Furthermore, sulfate (0.7 mM) is the most abundant dissolved terminal electron acceptor in Soudan groundwaters, and sulfate reduction coupled to CH4 oxidation is predicted to be a highly energetically favorable metabolism at this site43,49. Here, we focus on aspects of the OrgS cycle that involve DMSO and taurine because they may fuel the sulfate reduction that is predicted to be the dominant process fueling this subsurface community.
In our Soudan shotgun metagenomic datasets, we recovered a total of 65 medium to high-quality MAGs considered in further analyses. Reconstructed genomes are predominantly Bacteria, with two archaeal MAGs identified as Methanolobus. The Bacterial community spans 11 phyla, 16 classes, and 36 orders (Supplementary Data S3 and S6). Many MAGs align with previous diversity assessments at this site44. However, the detection of three previously unobserved classes (Kapabacteria, Spirochaetota, and UBP18) and several taxonomically novel MAGs (Supplementary Data S3) extends our understanding of site diversity and larger subsurface community membership50,51. The potential for OrgS cycling is similar in samples taken directly from anoxic boreholes and those taken from the suboxic to oxic fluid outflow channel. MAGs and bulk assemblies include protein-coding genes for OrgS production, utilization, and mineralization (Fig. 2, Supplemental Data S7 and S8). Additionally, MAGs and metagenomes encode full or partial pathways involved in dissimilatory oxidation or reduction of inorganic S. Genes involved in OrgS catabolism commonly co-occur with dissimilatory S genes, and 25 MAGs (38.5%) have the capacity to degrade OrgS to sulfite and then reduce or oxidize that sulfite (Fig. S3). This includes genes involved in the catabolism of sulfoacetaldehyde, isethionate, or cysteate (xsc, islA, cuyA) appearing alongside genes enabling sulfite reduction to sulfide (cysIJ, dsrABred, asrAB) or sulfite oxidation to sulfate (soeABC, aprABox+satox, sorAB). The co-occurrence of organic and inorganic S cycling genes within MAGs suggests that a single organism has the metabolic potential to couple OrgS catabolism to the dissimilatory oxidation or reduction of sulfite or sulfide. This direct, intracellular link between the inorganic and organic S cycles would provide energy for cells in this nutrient-depleted subsurface system. Furthermore, the established importance of metabolic handoffs in deep subsurface settings35 suggests that even cells in which pathways for sulfite generation and metabolism do not co-occur directly could be integral to the cycling of organic and inorganic S intermediates. This connection highlights the relevance of OrgS degradation as a source and sink of inorganic substrates within the deep terrestrial biosphere.
DMS(O)(P) cycling is ubiquitous across deep subsurface metagenomes
The cycling of DMSP and the related molecules dimethylsulfide (DMS) and DMSO, referred to collectively as DMS(O)(P), has been demonstrated to play an integral role in biogeochemical sulfur cycles in marine and terrestrial systems15,52,53,54. Genes involved in DMSO reduction have also been detected in two deep biosphere settings, suggesting that DMS(O)(P) cycling could be an important process in the deep subsurface31,32. Protein-coding genes involved in DMS(O)(P) cycling were found in all bulk assemblies and 95.4% of Soudan MAGs (Figs. S3, S4). Four MAGs, appearing in 17 of the 21 metagenomes investigated, encode dysB, a canonical gene indicative of DMSP biosynthesis24. A BLAST search of Soudan protein sequences against the related DMSP biosynthesis gene dysGD, which does not currently have an HMM, revealed several hits; however, the low amino acid identity to known DsyGD sequences (<50%) and lack of full coverage suggest these proteins in Soudan may not be functional20. Additionally, 6 MAGs can transport DMSP into the cell via the transporter gene dddT. We detected two pathways for DMSP catabolism: demethylation to methylmercaptopropionate (MMPA) via DMSP demethylase (dmdA) and cleavage to DMS via DMSP lyase (dddL, dddP, or dddD) (Fig. 2). The former was detected in one MAG from the order Rhodobacterales and appears predominantly in outflow channel metagenomes, while DMSP lyases were detected in six MAGs spanning Rhodobacterales, Acetobacterales, Pseudomonadales, and Nevskiales. An additional 3 MAGs encode dmdBCD to degrade MMPA and produce methanethiol (MeSH). Every bulk assembly and 25 MAGs encode methanethiol S-methyltransferase (mddA), which methylates MeSH and generates DMS.
The most common pathway for DMS oxidation to DMSO identified in this dataset involves dimethylsulfide dehydrogenase (ddhAB), which was detected in every bulk assembly and 35 MAGs (Figs. 2, S4). Together, MAGs encoding DdhAB comprise 20.7%–65.4% of the total microbial community in each sample (Supplementary Data S4). Pathways using dimethylsulfide S-monooxygenase (dsoBED) and trimethylamine monooxygenase (tmm) for DMS oxidation were also detected in three and one MAG, respectively (Fig. 2). Seven MAGs, spanning all 21 metagenomes, can transform DMSO to methanesulfonate via msuE, and three encode msuD, which produces sulfite via desulfonation of the intermediate methanesulfonate. One MAG encodes the catalytic subunit of DMSO reductase (dmsA), allowing DMSO to serve as a terminal electron acceptor in anaerobic respiration. Four additional MAGs contain the DMSO reductase subunit involved in electron transfer (dmsB) but were missing dmsA. Genes encoding Cym-type DMSO reductase (cymA) and TMAO reductase (torAC), which serve a functionally similar role to dmsAB, were not detected in any samples. The absence of DMSO reduction suggests sulfite could eventually be produced via methanesulfonate, a process which requires molecular oxygen (Fig. 2a). Though the reaction rate of this pathway may be constrained in Soudan’s anoxic groundwater, experimental and modeling-based studies that demonstrate H2 production via the radiolysis of water in similar Precambrian Shield brines suggest that limited oxygen production may be possible in this environment55,56. Additionally, chlorite and nitric oxide dismutation have been suggested as a source of molecular oxygen in ancient groundwaters57, and the detection of the chlorite dismutase gene (cdo) in four Soudan metagenomes supports this as a potential oxygen production mechanism in Soudan Mine.
These findings are well-aligned with previous studies of the terrestrial deep biosphere, which have also detected dmsAB31,32, but widely expand the array of DMS(O)(P)-related genes documented in these settings. The detection of dysB provides evidence of DMSP biosynthesis potential in a deep subsurface environment. The genetic potential to produce DMSP and mineralize it to DMS provides a mechanism to replenish and support a continuous, biotically driven DMS(O)(P) cycle. The ubiquity of genes involved in DMSP synthesis or uptake and high relative abundance of MAGs encoding these genes (9.1%–62.2% of borehole microbial communities) underlines the relevance of these processes in the deep biosphere (Supplementary Data S4). While DMS oxidation to DMSO occurs primarily through tmm in marine environments12,13, our findings indicate that the dominant pathway for DMS oxidation in Soudan relies on ddhAB. The differences between the two systems likely reflect differing redox conditions. The tmm is a dioxygenase enzyme requiring oxygen to drive DMS conversion to DMSO. While the production of molecular oxygen is likely possible in Soudan groundwaters55,56,57, the use of the ddhAB facilitates energy conservation because this complex translocates hydrogen ions and creates a reduced cytochrome. The presence of DMSP production alongside DMS oxidation, and ultimately sulfite production in the presence of oxygen, underscores the importance of continued investigation into organosulfur cycling in the deep subsurface. Furthermore, our results suggest that incorporation of OrgS into thermodynamic and reactive transport models is necessary because OrgS cycling could fuel inorganic S delivery and subsequent S redox.
Pathways to cycle taurine are diverse and prevalent in the deep biosphere
Taurine is ubiquitous in marine settings where it serves as an important source of C, N, S, and energy for marine heterotrophic bacteria16,17. The anaerobic dissimilation of taurine has been proposed as a mechanism to drive sulfite respiration17, and microbial communities in freshwater sediments are known to degrade taurine and accumulate sulfate10. Thus, taurine is of particular interest as a potential energy source in the oligotrophic terrestrial deep biosphere. We identified protein-coding genes involved in the import and degradation of taurine and related intermediates in 64.6% of reconstructed MAGs and all bulk assemblies (Fig. 2, S4). The capacity to import taurine-related compounds was detected in twelve MAGs, including two that encode the taurine importer TauABC and eleven capable of general alkanesulfonate uptake via SsuABC. Additionally, we detected two pathways for taurine degradation to sulfite, either directly or via the intermediate sulfoacetaldehyde (Fig. 2). Three MAGs, all within the phylum Pseudomonadota, encode taurine dioxygenase (tauD) to degrade taurine directly to sulfite. These MAGs are more abundant in outflow channel samples, comprising 3.4–5.4% of the total microbial community, compared to borehole fluids (<0.1% of the community). TauD allows taurine to be used as a sulfur source during aerobic growth and appears in 90.9% of metagenomes from the more oxic outflow channel and only 20.0% of metagenomes from anoxic borehole fluids. Seven MAGs contain genes to transform taurine to sulfoacetaldehyde, including three MAGs encoding taurine:oxoglutarate aminotransferase (toa), two encoding taurine:pyruvate aminotransferase (tpa), and four encoding taurine dehydrogenase (tauXY). The ability to desulfonate sulfoacetaldehyde and produce sulfite via sulfoacetaldehyde acetyltransferase (xsc) appears in eight MAGs (Fig. 2). Five deep terrestrial subsurface MAGs fully encode this pathway for taurine catabolism to sulfite via sulfoacetaldehyde, while six additional MAGs can complete only one of the two necessary steps. In total, 71% of metagenomes, found in every sample except those from DDH951, encode a full pathway for taurine degradation to sulfite. Metagenomes from DDH951 encode only toa, with one also encoding islAB to mineralize the related OrgS compound isethionate to sulfite (Fig. S4). The ability of microorganisms to degrade taurine and related OrgS compounds in oxic and anoxic prokaryote-dominated deep, terrestrial subsurface systems is intriguing, as taurine production is primarily attributed to eukaryotes58. The previous identification of a cysteine sulfinic acid decarboxylase in bacteria25., however, provides direct evidence for prokaryotic taurine biosynthesis and should be further investigated to understand the prevalence of this process under in situ conditions.
The prevalence and diversity of taurine degradation pathways in Soudan Mine metagenomes would suggest that pathways for taurine production are also present in this system; however, we did not detect key genes (hypotaurine monooxygenase, cysteamine dioxygenase, cysteine sulfonic acid decarboxylase) involved in known microbial taurine biosynthesis pathways (Supplementary Data S1). These pathways require oxygen to degrade cysteine to taurine via the intermediates cysteine sulfinic acid or cysteamine, but the highly reducing fluids of Soudan Mine and other subsurface systems may necessitate a pathway that can operate under anoxic conditions. One potential anaerobic taurine synthesis route that has been documented in higher eukaryotes involves the formation and subsequent decarboxylation of cysteate to form taurine59. Cysteate synthesis is one of the first steps in the production of coenzyme-m, and is known to occur in bacteria and archaea60. The cysteate synthase gene (cysS) that facilitates this reaction appears in two archeal and one bacterial MAG, and all bulk assemblies from anoxic borehole fluids. No cysS genes were detected in the more oxic outflow channels, supporting the notion of anaerobic taurine synthesis in the deep subsurface. Additionally, a promiscuous glutamate decarboxylase enzyme (EC. 4.1.1.15) has been documented to use cysteate as a substrate61 and the gene encoding this enzyme (gadAB) was detected in five Soudan MAGs (Fig. 2). While neither of these genes (gadAB or cysS) was found to co-occur in a single MAG, this could be due to MAG incompleteness. Even so, 30.0% of the metagenomes from anoxic borehole fluids have the capacity to complete this pathway at the community scale via the exchange of intermediate substrates (Fig. S4), a common metabolic strategy in the deep terrestrial subsurface35.
The confirmation of multiple taurine degradation pathways strongly suggests a source of taurine in this system, and the protein-coding genes cysS and gadAB provide one possible mechanism for biosynthesis in an anoxic setting. As such, microbes may use taurine as an osmolyte58 to offset the high salinity of Soudan groundwaters. The hypersaline conditions of the Soudan groundwaters may promote the synthesis and uptake of taurine, which would make it abundant and available to the microbial community. The common and diverse nature of genes involved in taurine uptake, transformation, and degradation underscores the potential importance of taurine in supporting microbial growth and diversity in deep subsurface environments.
Dissimilation of inorganic sulfur is a key metabolism in the deep biosphere
While the coupling of OrgS degradation to energy-generating inorganic sulfur redox reactions has received little attention, inorganic redox processes are a well-established cornerstone of metabolism in many subsurface environments, including Soudan Mine44,48,49. Previous work investigating the inorganic sulfur cycle at Soudan has noted genes related to redox and disproportion of intermediate oxidation S species, as well as sulfate and sulfide44,48. Here, we focus on expanding existing knowledge by incorporating the potential for OrgS to impact the biogeochemistry of the system. We focus on sulfate, sulfite, and sulfide because they are common links between the organic and inorganic sulfur cycles. The presence of genes such as phsA suggests that other intermediate forms of sulfur (thiosulfate, S(0)) are cycled here as well, but these pathways are predicted to predominantly add to the sulfite and sulfide pools. Thus, they are not expected to inhibit the processes of mineralization or dissimilatory sulfite redox focused on here.
We detect protein-coding genes involved in reductive or oxidative sulfur metabolisms in every metagenome and 72.3% of MAGs, with four MAGs encoding a complete dissimilatory sulfate reduction pathway and an additional six capable of sulfide oxidation to sulfate (Fig. 2a). These metabolisms are likely coupled to the oxidation of ferric iron and nitrate or the reduction of methane, H2, and more complex organics in this system44,48,49. A total of 26 MAGs encode sulfate adenylyltransferase (sat), with 17 of these identified as the reductive-type satred responsible for the activation of sulfate to adenosine 5’-phosphosulfate (APS). Four MAGs encode the reductive-type APS reductase (aprABred) to reduce APS to sulfite, three of which also contain a complete quinone-modifying oxidoreductase (qmoABC) complex. The QmoABC complex shuttles electrons between AprAB and the quinone pool, coupling APS reduction to energy production62. Dissimilatory sulfite reductase (dsrABred) is present in six MAGs, and anaerobic sulfite reductase (asrAB) and coenzyme F420 sulfite reductase (fsr) are each present in one, indicating the ability of those MAGs to metabolize via sulfite reduction. Every bulk assembly sampled from anoxic borehole fluids and 81.8% of those from outflow channels have the genetic potential (sat, aprAB, qmoABC, and dsrAB all encoded) to fully reduce sulfate to sulfide (Fig. S4). While only four MAGs can fully reduce sulfate to sulfide independently, seventeen others encode a subset of the genes involved. This could be an artifact of missing sequences in incomplete MAGs or could reflect a reliance on metabolic handoffs to complete this pathway as a community35.
Six Soudan MAGs have the capacity to fully oxidize sulfide to sulfate, encoding an oxidative-type dsrABox for sulfide oxidation to sulfite. These and six additional MAGs are capable of sulfite oxidation to sulfate via the oxidative-type aprABox and satox (5 MAGs) or the sulfite dehydrogenases soeABC (11 MAGs) or sorAB (2 MAGs). Genes involved in the SOX pathway for thiosulfate oxidation were identified in 23 MAGs, of which four encode the complete Sox pathway (soxAX, soxZY, soxB, and soxCD) to oxidize sulfide to sulfate63. Five MAGs encode an “incomplete” sox pathway lacking the soxCD gene (Fig. 2). These microorganisms can oxidize thiosulfate to sulfate or oxidize sulfide to elemental sulfur (S(0)), then further oxidize the intermediate-valence S(0) to sulfate using other pathways such as reverse dissimilatory sulfite reductase (dsrABox)64,65. One Gammaproteobacteria MAG in the order Burkholderiales appears capable of using this approach, as it encodes both an incomplete SOX pathway and the oxidative-type DsrAB. MAGs encoding a full sulfur oxidation pathway were identified in all outflow channel bulk assemblies and 70.0% of those from anoxic borehole fluids. Except for DDH951, MAGs capable of sulfur oxidation make up a significant portion of the microbial community in anoxic borehole fluids (8.2–41.0%) and likely support an active inorganic sulfur cycle in this deep subsurface system. Overall, the presence of both oxidative and reductive S metabolisms agrees with findings at other terrestrial subsurface sites and underlines the dynamic nature of sulfur cycling in these settings35,37,44.
The coupling of organosulfur mineralization to dissimilatory redox reactions can fuel deep life
Organosulfur degradation may produce reduced forms of inorganic sulfur (e.g., HS-) that can be oxidized or intermediate forms of inorganic S (e.g., SO32-) that can be either further oxidized or reduced. Thus, OrgS mineralization can generate redox-reactive S species under oxidizing and reducing conditions66. This process extends the metabolic framework of the sulfur cycle and calls attention to a previously overlooked pool of S in organic compounds. It also bolsters our understanding of the extensive hidden, or cryptic, S cycle uncovered by metagenomic approaches when geochemical approaches alone may not adequately reveal the full extent of cycling2. The deep terrestrial biosphere at Soudan Mine has the genetic potential to acquire, create, transform, and mineralize a wide range of OrgS compounds (Fig. 2). Only a fraction of this OrgS is expected to be used in biosynthesis based on the rarity of sulfur-containing amino acids, with cysteine and methionine previously reported to comprise just ~1.1% and ~2.9% of total cellular proteins, respectively17,67. Thus, the majority of S in these compounds remains available for energy production via respiratory processes. DMSO can serve directly as a terminal electron acceptor68, but most other OrgS compounds must first be mineralized. The fate of sulfur in DMSP, taurine, and other OrgS is often sulfite, which can then fuel energy production46.
As an example, taurine degradation by TauD frees sulfite to act as an electron acceptor through the interaction with the dissimilatory sulfite reductase (DsrAB), a process that could occur cryptically (Fig. 3). Given the highly reactive and toxic nature of sulfite, OrgS may provide a safer mechanism to carry sulfur at intermediate oxidation states until it is mineralized and used for energy production. Protein-coding genes involved in DMS(O)(P) and taurine cycling co-occur with inorganic sulfur cycling genes in 57 MAGs (87.7%), with bulk assemblies from every outflow channel sample and 70% of borehole fluid samples encoding complete pathways for OrgS degradation to sulfite as well as sulfite dissimilation (Fig. S4).
Example of coupled organic and inorganic sulfur cycles, with catabolism of taurine providing the substrate used in dissimilatory sulfite reduction.
The Gibbs energy yield of several potential sulfite metabolisms under in situ conditions was evaluated to determine the feasibility of these strategies in Soudan Mine (Fig. S6, Supplementary Data S10). These metabolisms included the oxidation and reduction of sulfite coupled to known electron acceptors (NO3-) and donors (CH4, H2). Thermodynamic calculations suggest that all investigated reactions are exergonic, with the coupling of nitrate reduction to ammonia with sulfite oxidation to sulfate providing the highest energy yield per electron. Despite being less exergonic per mole of electron, energy density calculations suggest that the reactions coupling sulfite reduction with hydrogen or methane oxidation have the potential to provide a similar amount of energy per kg of water due to the limited availability of nitrate in Soudan borehole fluids. Thus, linking OrgS mineralization to dissimilatory oxidation or reduction of the inorganic byproduct would allow a diverse array of OrgS compounds, including those lacking reductases, to serve indirectly as electron donors or acceptors and to play a cryptic but central role in deep terrestrial subsurface energy production.
Organosulfur cycling is important across the terrestrial subsurface
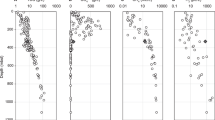
While the microbial community at Soudan is clearly capable of OrgS cycling, the iron-rich formation is not representative of all deep terrestrial environments. To investigate whether OrgS serves a central role in other terrestrial subsurface systems, we searched 286 MAGs generated from 54 metagenomes obtained through the Census of Deep Life, a program within the Deep Carbon Observatory that sampled the deep biosphere around the planet69. We observe patterns similar to those identified in Soudan MAGs across diverse lithologies (Fig. 4 and Supplementary Data S5). Genes involved in S assimilation were detected in 283 MAGs (99.0%). Genes related to the synthesis, utilization, and degradation of DMS and taurine are present across all sites, identified in 241 (84.3%) and 118 (41.3%) MAGs, respectively. Of note, 42 MAGs (14.7%) have the capacity to produce sulfite via mineralization of organosulfur, with 16 (5.6%) of these MAGs also encoding pathways for either dissimilatory reduction or oxidation of that sulfite. The gene ddhAB is the most commonly occurring OrgS gene outside of those involved in sulfur assimilation, suggesting that 120 MAGs (42.0%) are capable of oxidizing DMS to DMSO. These results support the ubiquity of OrgS cycling genes in deep subsurface metagenomes and the possibility that OrgS compounds could act as an energy source for the deep biosphere, especially considering the availability of new and ancient sulfurized organic matter in these settings10,70.
Number of Census of Deep Life (CoDL) metagenome assembled genomes (MAGs) of the total 286 analyzed that contain each gene or complex of interest. Pink bars indicate genes involved in the dissimilation of organic sulfur. Blue bars indicate genes involved in taurine transformations. Green bars indicate genes involved in the transformation of dimethylsulfide, dimethylsulfoxide, or dimethylsulfoniopropionate (DMS(O)(P)). Yellow bars indicate genes involved in the production and utilization of cysteine and methionine. The inset map depicts sample locations (black dots) of the 54 CoDL samples assembled into MAGs. See Supplementary Data S1 for gene abbreviations and Supplementary Data S9 for all gene hits with METABOLIC.
This study demonstrates the diverse, cosmopolitan genetic potential for OrgS cycling throughout the terrestrial deep subsurface, highlighting pathways encoded in deep biosphere metagenomes that link the organic and inorganic sulfur cycle and have the potential to fuel the energy-limited deep biosphere. The biosynthesis and subsequent cycling of OrgS can directly impact the availability of redox-reactive inorganic S species that act as electron donors or acceptors, and so OrgS cycling could provide an underappreciated control on terrestrial deep subsurface S fluxes. The growing recognition of OrgS as an integral aspect of biogeochemical sulfur cycling across diverse environments6,7,8,10 underscores the need for a fundamental revision to our conceptualization of S cycling to acknowledge the potential impact of both the inorganic and organic aspects of this cycle. These efforts will expand our understanding of microbial community functioning and inform other avenues of research, including the search for novel biomarkers and attempts to culture the uncultured microbial majority. Continued investigation into the activity of microbial OrgS transformations is necessary to understand the full extent of deep subsurface OrgS cycling and its larger impact on Earth’s Critical Zone.
Methods
Detailed descriptions of these abbreviated methods are provided in the Supplementary Information.
Site description
Soudan Underground Mine State Park lies within the Superior Province of the Canadian Shield in Northeastern Minnesota (47.82333 N, 92.23722 W) and transects the Soudan Formation, a Neoarchean (~2.7 Ga) massive hematite iron formation. The lowest level of the mine is 715 meters below land surface and has flowing artesian boreholes that tap a fractured rock aquifer underlying the mine. The four boreholes sampled in this study (DDH932, DDH942, DDH944, DDH951) are angled downward and are 102–144 m long, intersecting chlorite schists, massive hematite, and banded iron formation with minor pyrite and chalcopyrite inclusions. Low-temperature (~10 °C), anoxic, and reducing (~−200 mV) groundwaters continuously flow at an average rate of 10–20 ml min-143. These waters have long residence times and are the most isolated from meteoric waters as compared to other boreholes at the site43. The borehole waters (dominated by calcium, sodium, and chloride ions) are circumneutral pH and have total dissolved solids concentrations between 76,000–116,000 mg kg−1. Geochemical and metagenomic analyses have identified ferric iron (~1.7 mM), nitrate (~3 µM), and sulfate (~0.7 mM) as potential anaerobic electron acceptors in this system, and methane (~4.8 mM), H2 (~0.2 mM), and more complex organics as key electron donors43,44,49. Thermodynamic calculations support CH4 oxidation by SO42- as a key metabolism fueling the microbial community in Soudan Mine49. Both reduced and oxidized forms of organic and inorganic S, as well as zero-valent S, have been observed in this system via sulfur XANES spectroscopy (Fig. S2).
Sample collection
Samples for metagenomics were collected between January 2019 and October 2021. Ten anoxic borehole fluid samples were collected using a peristaltic pump to filter water from a single depth through a 0.22 µm Sterivex filter to collect biomass. Additionally, 11 fluid outflow samples were taken from the drainage channel that moves borehole fluid outflow to a water collection area in the basement of the mine. This channel represents an oxic-anoxic transition zone where anoxic borehole waters are exposed to the oxygenated surface air. A single ditch connects all boreholes, so these samples reflect a mixture of borehole waters. These samples were collected by manually pushing groundwater through a Sterivex filter using a sterile 60 ml syringe. All filters were stored in sterile WHIRL-PAK bags, immediately frozen on dry ice, and maintained at −80 °C until being processed.
DNA extraction and sequencing
DNA was extracted directly from the filters using an MP Biomedicals FastDNA Spin Kit for Soil following the manufacturer’s protocol with minor modifications, as described in the Supplementary Information. Samples from unused Sterivex filters were processed and extracted following the same protocols to serve as a negative control. Genomic libraries were prepared and sequenced in one lane of an Illumina NovaSeq 6000 S1 by the University of Minnesota Genomics Center, St Paul, MN.
Metagenome assembly and metabolic reconstruction
Shotgun metagenomic sequencing yielded 9.47 gigabases of raw sequencing data. Reads were processed, assembled into 2.7 million contigs, binned into metagenome assembled genomes (MAGs), and deduplicated and aggregated to create an optimized set of non-redundant bins as described in the Supplementary Information. Quality of the resulting 81 bins was assessed using MIMAG guidelines71, with 52 high-quality MAGs of >90% completeness and <5% contamination and 14 medium-quality MAGs of ≥50% completeness and <10% contamination. These MAGs were retained for further analysis, while the 15 low-quality bins falling below these thresholds were excluded. One additional MAG was removed after being identified as a likely subsurface contaminant following the workflow outlined in Sheik et al.72. All bioinformatic analyses were performed using computational resources from the Minnesota Supercomputing Institute.
Key (organo)sulfur cycling enzymes were manually curated using literature searches and the Kyoto Encyclopedia of Genes and Genomes (KEGG)73 to identify relevant metabolic pathways.
The genetic potential of MAGs and metagenomes was reconstructed using the program METABOLIC v. 474, and supplemented with analyses using HMSS2 v. 1.0.575.
Processing of census of deep life datasets
Fifty-four additional deep subsurface metagenomes were obtained through the Census of Deep Life (CoDL)69. The collection, processing, and assembly protocols employed by CoDL to generate these MAGs are described in detail in the Supplementary Information.
Thermodynamic modeling of sulfite metabolisms
The feasibility of several candidate dissimilatory sulfite metabolisms at Soudan Mine was determined using geochemical data to calculate the Gibbs energy yields of seven candidate microbial reactions under in situ conditions (Supplementary Data S10). The equations used and assumptions underlying these calculations are described in detail in Supplementary Information.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The sequence reads and MAGs generated in this study have been deposited in the NCBI Sequence Read Archive under BioProject accession code PRJNA248749 [https://www.ncbi.nlm.nih.gov/bioproject/248749].
References
Jørgensen, B. B. Mineralization of organic matter in the sea bed—the role of sulphate reduction. Nature 296, 643–645 (1982).
Canfield, D. E. et al. A cryptic sulfur cycle in oxygen-minimum–zone waters off the Chilean coast. Science 330, 1375–1378 (2010).
Osorio, H. et al. Anaerobic sulfur metabolism coupled to dissimilatory iron reduction in the extremophile Acidithiobacillus ferrooxidans. Appl. Environ. Microbiol 79, 2172–2181 (2013).
Fike, D. A., Bradley, A. S. & Rose, C. V. Rethinking the ancient sulfur cycle. Annu. Rev. Earth Planet. Sci. 43, 593–622 (2015).
Guo, W., Wen, Y., Chen, Y. & Zhou, Q. Sulfur cycle as an electron mediator between carbon and nitrate in a constructed wetland microcosm. Front. Environ. Sci. Eng. 14, 57 (2020).
Fakhraee, M., Li, J. & Katsev, S. Significant role of organic sulfur in supporting sedimentary sulfate reduction in low-sulfate environments. Geochim. Cosmochim. Acta 213, 502–516 (2017).
Tang, K. & Liu, L. Bacteria are driving the ocean’s organosulfur cycle. Trends Microbiol. 31, 772–775 (2023).
Landa, M. et al. Sulfur metabolites that facilitate oceanic phytoplankton–bacteria carbon flux. ISME J. 13, 2536–2550 (2019).
Fakhraee, M. & Katsev, S. Organic sulfur was integral to the Archean sulfur cycle. Nat. Commun. 10, 4556 (2019).
Phillips, A. A. et al. Organic sulfur from source to sink in low-sulfate Lake Superior. Limnol. Oceanogr. 68, 2716–2732 (2023).
Howard, E. C., Sun, S., Biers, E. J. & Moran, M. A. Abundant and diverse bacteria involved in DMSP degradation in marine surface waters. Environ. Microbiol. 10, 2397–2410 (2008).
Teng, Z.-J. et al. Biogeographic traits of dimethyl sulfide and dimethylsulfoniopropionate cycling in polar oceans. Microbiome 9, 1–17 (2021).
O’Brien, J. et al. The microbiological drivers of temporally dynamic dimethylsulfoniopropionate cycling processes in Australian Coastal Shelf waters. Front. Microbiol. 13, 894026 (2022).
Lomans, B. P., van der Drift, C., Pol, A. & Op den Camp, H. J. M. Microbial cycling of volatile organic sulfur compounds. CMLS, Cell. Mol. Life Sci. 59, 575–588 (2002).
Tebbe, D. A. et al. Microbial drivers of DMSO reduction and DMS-dependent methanogenesis in saltmarsh sediments. ISME J. https://doi.org/10.1038/s41396-023-01539-1 (2023).
Clifford, E. L. et al. Crustacean zooplankton release copious amounts of dissolved organic matter as taurine in the ocean. Limnol. Oceanogr. 62, 2745–2758 (2017).
Cook, A. M. & Denger, K. Metabolism of Taurine in Microorganisms. In Taurine 6 Vol. 583 (eds. Oja, S. S. & Saransaari, P.) 3–13 (Springer US, 2006).
Wittek, B., Carnat, G., Tison, J.-L. & Gypens, N. Response of dimethylsulfoniopropionate (DMSP) and dimethylsulfoxide (DMSO) cell quotas to salinity and temperature shifts in the sea-ice diatom Fragilariopsis cylindrus. Polar Biol. 43, 483–494 (2020).
Lyon, B. R., Bennett-Mintz, J. M., Lee, P. A., Janech, M. G. & DiTullio, G. R. Role of dimethylsulfoniopropionate as an osmoprotectant following gradual salinity shifts in the sea-ice diatom Fragilariopsis cylindrus. Environ. Chem. 13, 181–194 (2016).
Wang, J. et al. Alternative dimethylsulfoniopropionate biosynthesis enzymes in diverse and abundant microorganisms. Nat. Microbiol. https://doi.org/10.1038/s41564-024-01715-9 (2024).
Kirst, G. O. et al. Dimethylsulfoniopropionate (DMSP) in icealgae and its possible biological role. Mar. Chem. 35, 381–388 (1991).
Sunda, W., Kieber, D. J., Kiene, R. P. & Huntsman, S. An antioxidant function for DMSP and DMS in marine algae. Nature 418, 317–320 (2002).
Clifford, E. L. et al. Taurine Is a major carbon and energy source for marine prokaryotes in the North Atlantic Ocean off the Iberian Peninsula. Micro. Ecol. 78, 299–312 (2019).
Curson, A. R. J. et al. Dimethylsulfoniopropionate biosynthesis in marine bacteria and identification of the key gene in this process. Nat. Microbiol 2, 1–9 (2017).
Agnello, G., Chang, L. L., Lamb, C. M., Georgiou, G. & Stone, E. M. Discovery of a substrate selectivity motif in amino acid decarboxylases unveils a taurine biosynthesis pathway in prokaryotes. ACS Chem. Biol. 8, 2264–2271 (2013).
McParland, E. L., Alexander, H. & Johnson, W. M. The osmolyte ties that bind: genomic insights into synthesis and breakdown of organic osmolytes in marine microbes. Front. Mar. Sci. 8, 689306 (2021).
McMahon, S. & Parnell, J. Weighing the deep continental biosphere. FEMS Microbiol Ecol. 87, 113–120 (2014).
Bar-On, Y. M., Phillips, R. & Milo, R. The biomass distribution on Earth. Proc. Natl Acad. Sci. USA 115, 6506–6511 (2018).
Magnabosco, C. et al. The biomass and biodiversity of the continental subsurface. Nat. Geosci. 11, 707–717 (2018).
Laue, H., Denger, K. & Cook, A. M. Fermentation of cysteate by a sulfate-reducing bacterium. Arch. Microbiol. 168, 210–214 (1997).
Mateos, G. et al. Shewanella sp. T2.3D-1.1 a novel microorganism sustaining the iron cycle in the deep subsurface of the Iberian Pyrite Belt. Microorganisms 10, 1585 (2022).
Nuppunen-Puputti, M. et al. Epilithic microbial community functionality in deep oligotrophic continental bedrock. Front. Microbiol. 13, 826048 (2022).
Itävaara, M., Salavirta, H., Marjamaa, K. & Ruskeeniemi, T. Geomicrobiology and metagenomics of terrestrial deep subsurface microbiomes. In Advances in Applied Microbiology Vol. 94 1–77 (Elsevier, 2016).
Ulrich, G. A. et al. Sulfur cycling in the terrestrial subsurface: commensal interactions, spatial scales, and microbial heterogeneity. Microb. Ecol. 36, 141–151 (1998).
Anantharaman, K. et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 7, 13219 (2016).
Lau, M. C. Y. et al. An oligotrophic deep-subsurface community dependent on syntrophy is dominated by sulfur-driven autotrophic denitrifiers. Proc. Natl Acad. Sci. USA 113, E7927–E7936 (2016).
Bell, E. et al. Active sulfur cycling in the terrestrial deep subsurface. ISME J. 14, 1260–1272 (2020).
Momper, L., Casar, C. P. & Osburn, M. R. A metagenomic view of novel microbial and metabolic diversity found within the deep terrestrial biosphere at DeMMO: a microbial observatory in South Dakota, USA. Environ. Microbiol. 25, 3719–3737 (2023).
Krumholz, L. R., McKinley, J. P., Ulrich, G. A. & Suflita, J. M. Confined subsurface microbial communities in Cretaceous rock. Nature 386, 64–66 (1997).
Mateos, K. et al. The evolution and spread of sulfur cycling enzymes reflect the redox state of the early Earth. Sci. Adv. 9, eade4847 (2023).
Raven, M. R. et al. Organic matter sulfurization and organic carbon burial in the Mesoproterozoic. Geochim. Cosmochim. Acta 347, 102–115 (2023).
LaRowe, D. E. & Amend, J. P. The energetics of anabolism in natural settings. ISME J. 10, 1285–1295 (2016).
Schuler, C. J. et al. Water and rock chemistry inform our understanding of the deep biosphere: case study in an Archaean banded iron formation. Front. Earth Sci. 10, 803250 (2022).
Sheik, C. S. et al. Novel microbial groups drive productivity in an Archean iron formation. Front. Microbiol. 12, 627595 (2021).
Sheik, C. S. et al. Abiotic and biotic processes that drive carboxylation and decarboxylation reactions. Am. Mineral. 105, 609–615 (2020).
Cook, A. M., Smits, T. H. M. & Denger, K. Sulfonates and Organotrophic Sulfite Metabolism. In Microbial Sulfur Metabolism (eds. Dahl, C. & Friedrich, C. G.) 170–183 (Springer, Berlin, 2008).
Ye, H. et al. Ecophysiology and interactions of a taurine-respiring bacterium in the mouse gut. Nat. Commun. 14, 5533 (2023).
Schuler, C. J. et al. Densely populated biofilms and linked iron and sulfur cycles in the fractured-rock continental subsurface. Geochim. Cosmochim. Acta https://doi.org/10.1016/j.gca.2024.04.019 (2024).
Dowd, W. S. et al. Potential energy sources for the deep continental biosphere in isolated anoxic brines. Earth Planet. Sci. Lett. 595, 117720 (2022).
Magnabosco, C. et al. Comparisons of the composition and biogeographic distribution of the bacterial communities occupying South African thermal springs with those inhabiting deep subsurface fracture water. Front. Microbiol. 5, 679 (2014).
Momper, L. et al. Major phylum-level differences between porefluid and host rock bacterial communities in the terrestrial deep subsurface. Environ. Microbiol. Rep. 9, 501–511 (2017).
Carrión, O. et al. DMSOP-cleaving enzymes are diverse and widely distributed in marine microorganisms. Nat. Microbiol 8, 2326–2337 (2023).
Williams, B. T. et al. Bacteria are important dimethylsulfoniopropionate producers in coastal sediments. Nat. Microbiol 4, 1815–1825 (2019).
Curson, A. R. J., Todd, J. D., Sullivan, M. J. & Johnston, A. W. B. Catabolism of dimethylsulphoniopropionate: microorganisms, enzymes and genes. Nat. Rev. Microbiol 9, 849–859 (2011).
Lin, L.-H., Slater, G. F., Sherwood Lollar, B., Lacrampe-Couloume, G. & Onstott, T. C. The yield and isotopic composition of radiolytic H2, a potential energy source for the deep subsurface biosphere. Geochim. Cosmochim. Acta 69, 893–903 (2005).
Warr, O., Giunta, T., Ballentine, C. J. & Sherwood Lollar, B. Mechanisms and rates of 4He, 40Ar, and H2 production and accumulation in fracture fluids in Precambrian Shield environments. Chem. Geol. 530, 119322 (2019).
Ruff, S. E. et al. Hydrogen and dark oxygen drive microbial productivity in diverse groundwater ecosystems. Nat. Commun. 14, 3194 (2023).
Tevatia, R. et al. The taurine biosynthetic pathway of microalgae. Algal Res. 9, 21–26 (2015).
Malatesta, M. et al. Birth of a pathway for sulfur metabolism in early amniote evolution. Nat. Ecol. Evol. 4, 1239–1246 (2020).
Wu, H.-H. et al. The pathway for coenzyme M biosynthesis in bacteria. Proc. Natl Acad. Sci. USA 119, e2207190119 (2022).
Kim, H.-W., Kashima, Y., Ishikawa, K. & Yamano, N. Purification and characterization of the first archaeal glutamate decarboxylase from Pyrococcus horikoshii. Biosci. Biotechnol. Biochem. 73, 224–227 (2009).
Ramos, A., Keller, K., Wall, J. & Pereira, I. A. The membrane QmoABC complex interacts directly with the dissimilatory adenosine 5´-phosphosulfate reductase in sulfate reducing bacteria. Front. Microbiol. 3, 137 (2012).
Friedrich, C. G., Rother, D., Bardischewsky, F., Quentmeier, A. & Fischer, J. Oxidation of reduced inorganic sulfur compounds by bacteria: emergence of a common mechanism? Appl Environ. Microbiol. 67, 2873–2882 (2001).
Schedel, M., Vanselow, M. & Trüper, H. G. Siroheme sulfite reductase isolated from Chromatium vinosum. Purification and investigation of some of its molecular and catalytic properties. Arch. Microbiol. 121, 29–36 (1979).
Frigaard, N.-U. & Dahl, C. Sulfur metabolism in phototrophic sulfur bacteria. In Advances in Microbial Physiology Vol. 54 (ed. Poole, R. K.) 103–200 (Academic Press, 2008).
Wasmund, K., Mußmann, M. & Loy, A. The life sulfuric: microbial ecology of sulfur cycling in marine sediments. Environ. Microbiol. Rep. 9, 323–344 (2017).
Miseta, A. & Csutora, P. Relationship between the occurrence of cysteine in proteins and the complexity of organisms. Mol. Biol. 17, 1232–1239 (2000).
McCrindle, S. L., Kappler, U. & McEwan, A. G. Microbial dimethylsulfoxide and trimethylamine-N-oxide respiration. In Advances in Microbial Physiology Vol. 50 (ed. Poole, R. K.) 147–201e (Academic Press, 2005).
Ruff, S. E. et al. A global comparison of surface and subsurface microbiomes reveals large-scale biodiversity gradients, and a marine-terrestrial divide. Sci. Adv. 10, eadq0645 (2024).
Osterholz, H. et al. Terrigenous dissolved organic matter persists in the energy-limited deep groundwaters of the Fennoscandian Shield. Nat. Commun. 13, 4837 (2022).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Sheik, C. S. et al. Identification and removal of contaminant sequences from ribosomal gene databases: Lessons from the Census of Deep Life. Front. Microbiol. 9, 840 (2018).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Zhou, Z. et al. METABOLIC: high-throughput profiling of microbial genomes for functional traits, metabolism, biogeochemistry, and community-scale functional networks. Microbiome 10, 33 (2022).
Tanabe, T. S. & Dahl, C. HMSS2: An advanced tool for the analysis of sulphur metabolism, including organosulphur compound transformation, in genome and metagenome assemblies. Mol. Ecol. Resour. 23, 1930–1945 (2023).
Acknowledgements
This research was supported by funding from the National Science Foundation Integrated Earth Systems Program (EAR−1813526) to B.M.T., C.M.S., C.S.S., and a Graduate Assistantship from the University of Minnesota Informatics Institute and Minnesota’s Discovery, Research, and Innovation Economy to A.C.P. The authors thank the Minnesota Department of Natural Resources, particularly Jim Essig, for safe access to Soudan Mine and the Census of Deep Life and Deep Carbon Observatory for providing additional metagenomes. We thank Caroline Pierce for XANES sample preparation and data collection. Part of the research described in this paper was performed at the Canadian Light Source, a national research facility of the University of Saskatchewan, which is supported by the Canada Foundation for Innovation (CFI), the Natural Sciences and Engineering Research Council (NSERC), the Canadian Institutes of Health Research (CIHR), the Government of Saskatchewan, and the University of Saskatchewan. We thank Scott Alexander, Calvin Alexander, Daniel Bond, Jeff Gralnick, David Hsu, Peter Kang, Woonghee Lee, Billy Dowd, Jill McDermott, Tucker Ely, Anders Noren, and the Continental Scientific Drilling Facility for contributions to field sampling and collaboration on the Banded Together project.
Author information
Authors and Affiliations
Contributions
C.S.S., C.M.S., A.C.P., and B.M.T. contributed to study conception; A.C.P., C.J.S., C.S.S., C.M.S., and B.M.T. performed research; A.C.P., C.J.S., C.S.S., and B.M.T. analyzed data; A.C.P., C.S.S., and C.M.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Patsis, A.C., Schuler, C.J., Toner, B.M. et al. The potential for coupled organic and inorganic sulfur cycles across the terrestrial deep subsurface biosphere. Nat Commun 16, 3827 (2025). https://doi.org/10.1038/s41467-025-59241-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59241-z