Abstract
Liver transplant recipients (LTRs) are at risk of graft injury, leading to cirrhosis and reduced survival. Liver biopsy, the diagnostic gold standard, is invasive and risky. We developed a hybrid multi-class neural network (NN) model, ‘GraftIQ,’ integrating clinician expertise for non-invasive graft pathology diagnosis. Biopsies from LTRs (1992–2020) were classified into six categories using demographic, clinical, and lab data from 30 days pre-biopsy. The dataset (5217 biopsies) was split 70/30 for training/testing, with external validation at Mayo Clinic, Hannover Medical School, and NUHS Singapore. Bayesian fusion was used to combine clinician-derived probabilities with NN predictions, improving performance. Here we show that GraftIQ (MulticlassNN+clinical insight) achieved an AUC of 0.902 (95% CI:0.884–0.919), up from 0.885 with NN alone. Internal and external validation demonstrated 10–16% higher AUC than conventional ML models. GraftIQ demonstrates high accuracy in identifying graft etiologies and offers a valuable clinical decision support tool for LTRs.
Similar content being viewed by others
Introduction
Liver transplantation (LT) is a life-saving measure for selected patients with end stage liver disease1. Despite tremendous improvement in LT outcomes over recent decades, liver transplant recipients (LTRs) remain at risk of developing graft injury of various etiologies. Graft injury can result in fibrosis and cirrhosis over time, potentially resulting in graft loss in 25% of LTRs2. Graft viability after LT is dependent on the prompt recognition of post-transplant pathologies. Promptly starting treatments like high-dose steroids for rejection or antiviral therapy relies on identifying the cause of graft injury to prevent long-term dysfunction.
Graft injury is often suspected via elevated liver enzymes during routine bloodwork. However, biochemical tests are non-specific and it is difficult to establish a cause of injury based on these alone3. Liver biopsy has therefore remained the gold standard for the diagnosis of graft pathology4. In fact, with improvements in post-transplant survival over the last 2 decades, repeat evaluations of graft function via biopsy have become more frequent5. Liver biopsy is subject to sampling error as well as complications such as bleeding, infection6 and is often unavailable in a timely manner. Time constraints mean that hepatologists often have to make empiric decisions before liver biopsy is carried out. Such decisions are based on clinical data (age, indication for transplant, time after transplant, diabetes, obesity, immunosuppression regimen) and liver biochemical patterns. Thus, there is a clinical need to develop reliable, non-invasive methodologies that can establish or rule out specific etiologies of graft injury prior to biopsy, allowing rational therapeutic decisions to be made as quickly as possible.
Machine learning (ML), specifically neural networks (NN) are efficient in analyzing large, complex, and heterogeneous datasets, generating reproducible predictions and classifications on previously unseen data7. Previous studies have demonstrated the feasibility of convolutional NNs to generate accurate prognostic predictions and fibrosis detection in various chronic liver diseases by leveraging blood test patterns and interrelationships between variables8,9. When applied to liver transplantation, ML has emerged as a promising methodology to stratify patient risk and predict post-transplant outcomes10,11,12,13. Convolutional NNs have been applied to help predict waitlist mortality14, donor–recipient matching15,16 and HCC recurrence17. While ML models have the potential to process and analyze vast amounts of data quickly and efficiently, it is important to recognize that ML models may lack the nuanced understanding, and contextual knowledge that experienced clinicians bring to patient care.
In this work, we hypothesized that combining the extensive, prior knowledge of causal and correlational associations that human experts possess with a machine-learned model would increase model generalizability. To address this challenge, we have developed GraftIQ (Fig. 1a), a hybrid neural network model designed to predict the etiology of graft injury using clinical, demographic, and laboratory data from liver transplant (LT) recipients. Our approach provides a unified, single-step diagnostic solution for six distinct graft injury categories. Beyond multi-class classification, a key innovation of GraftIQ lies in its Bayesian fusion-based framework, which integrates clinician feedback to refine model predictions. By combining data-driven learning with expert knowledge, our hybrid ML tool ‘GraftIQ’ could potentially reduce dependence on longitudinal liver biopsies and lead to earlier therapeutic interventions, improving graft viability and patient survival over time.
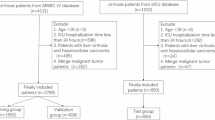
a Schematic representation of our hybrid multi-class neural network, ‘Graft IQ,’ which combines clinician expertise with multiclass neural network capabilities to predict the cause of graft injury. Created in BioRender. S, D. (2025) https://BioRender.com/m3o0749. b Flowchart detailing study design and data distribution.
Results
Patient population
A total of 1791 patients were identified for analysis. Mean recipient age was 52.4 ± 11.0 years, and mean donor age was 43.8 ± 16.5 years. A total of 601 patients (34%) were female, and 1190 (66%) male. Mean recipient weight was 79.2 ± 17.9 kg, and 388 patients (27%) had BMI over 30. Comorbidities included diabetes in 282 patients (16%), hypertension in 237 patients (14%) and dyslipidemia in 59 patients (3%). Mean MELD at time of transplantation was 18.3 ± 9.3. A total of 448 patients (25%) received living donor liver transplant, and 1343 (75%) received deceased donor liver transplant. 90 patients (5%) developed recurrent hepatocellular carcinoma (HCC) and 138 patients (8%) developed cholangitis. The indications for transplant are described in Supplementary Table 1. The most common indication was Hepatitis C with 711 biopsies (40%), followed by immune-mediated liver diseases (AIH, Primary biliary cholangitis, Primary sclerosing cholangitis) at 289 (17%), alcohol-related liver disease at 211 (12%) and MASH at 121 (7%).
Disease cohort characteristics
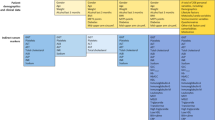
7580 liver biopsies were available from our post-transplant database. After careful review and exclusion of biopsies with missing data and double diagnoses, a total of 5217 biopsies remained and were included in the analysis. This total is higher than the total number of patients, as many patients had more than 1 biopsy over their post-transplant course. From this total, we identified the diagnostic categories of ACR, AIH, BO, congestion, HCV, MASH, and others but focused on the first six for our analysis. A total of 1979 biopsies were consistent with HCV, 1635 with ACR, 383 with BO, 211 with MASH, 163 with AIH, 142 with hepatic congestion, and 704 considered as others. We documented mean values of laboratory variables at time of biopsy and up to 30 days prior for each category. Details about these features for each disease cohort provided in Table 1.
Results of implementation analysis
As shown in Fig. 2, for the 30 cases chosen for the implementation analysis, the ML model exhibited higher predictive accuracies, surpassing hepatologists in every evaluated category. Notably, the ML tool achieved a perfect 100% accuracy for autoimmune hepatitis, BO, HCV, MASH, showcasing its robust diagnostic capabilities. In contrast, hepatologists demonstrated comparatively lower accuracy rates, particularly in predicting ACR, BO, congestion, and MASH. The instances where our ML model misclassified ACR (67%) and congestion (80%) categories shed light on the importance of integrating clinical expertise into our predictive framework. In the case of ACR, cases were misclassified as MASH, despite elevated liver enzymes and blood tests, as the ML model failed to consider the patient’s low age of graft, a key indicator against MASH. Similarly, in congestion misclassified as HCV, the model overlooked the absence of HCV as an indication for transplant. These discrepancies underscore the necessity of incorporating clinical insights to align more closely with the complexities of real-world clinical scenarios.
Comparison of multiclass neural network-based ML model’s prediction using clinical and lab data vs. averaged prediction accuracy provided by 12 hepatologists (medics).
Predictive performance evaluation
Utilizing multiclass neural network model standalone on test set
Firstly, we evaluated the performance of our multiclass NN-based ML model independently, without integrating any clinical expertise for predicting each diagnosis category as shown in Table 2. The best performance was obtained for MASH post-transplant complications with area under the curve (AUC) of 0.929 calculated using the receiver operating characteristic (ROC) curve with a sensitivity, and specificity of 0.89 and 0.92, respectively, followed by AIH and congestion, with AUC of 0.924 and 0.922, respectively. The overall AUC was obtained by averaging the AUC obtained for each individual category (refer to Supplementary Tables 8–10 for detailed results on the CIs, confusion matrix and error rates for each diagnostic category). In our case, the overall AUC for our neural network methodology on the test set was obtained to be 0.885 [95% confidence interval (CI): 0.864, 0.901].
Utilizing “GraftIQ”, hybrid model integrating NN prediction and clinical insight on test set
As shown in Table 2, column 7, the incorporation of clinician-based probabilities (with α = 0.2 and \(\beta\) = 0.8 for fusion after tuning as shown in Supplementary Table 2) into the final layer of our neural network model resulted in improvement in predictive performance for each diagnosis category. Specifically, the AUC values surpassed 0.8 for every category (notably high improvement for ACR prediction), representing a significant improvement compared to predictions made using the ML model alone. The overall AUC based on integrating clinical expertise improved from 0.885 to 0.902.
We then compared our neural network model to other conventional machine learning models (refer Table 3) and found that the neural network performs better in terms of overall AUC for classification with AUC of 0.902, [95% CI: 0.884, 0.919] as compared to the second-best approach Random Forest with an AUC of 0.823 [95% CI: 0.812, 0.839]. Regression approaches performed relatively less accurately in terms of multi-class classification with Logistic Regression with an AUC of 0.767 [95% CI: 0.626, 0.796], Lasso with an AUC of 0.783 [95% CI: 0.769, 0.802] and Ridge regression with an AUC of 0.781 [95% CI: 0.771, 0.811] justifying the prominence of neural networks in understanding the non-linear relationships in the data as well as in assigning subjects accurately to one of the multiple categories of diagnosis. To ensure the representativeness of the modern transplant population, which largely excludes HCV, we compared patients transplanted for HCV-related liver disease to those with non-HCV etiologies. As shown in Table 4, this stratification demonstrated the model’s robustness across graft injury categories, confirming its generalizability to contemporary transplant cohorts.
To make our neural network methodology more explainable and clinically relevant, we also computed the variable importance of each clinical feature in the classification task for individual categories. The higher the gradient obtained through the Integrated Gradient methodology detailed in the subsection “Extract important features through neural networks”, the more important the feature is in the classification task. As shown in Fig. 3, ALT, ALP, and hemoglobin were the top 3 features important to the classification of subjects in the ACR category. Similar plots for the rest of the five diagnosis categories are provided in the Supplementary document (Supplementary Figs. 1–5).
The x-axis represents the gradient in neural network learning. The higher the gradient, the more important the feature is.
Results on external validation set
In the UHN dataset, 542 biopsies were reviewed and divided according to the 6 relevant categories along with other biopsies. 233 biopsies were consistent with ACR, 68 with biliary obstruction, 77 with MASH, 18 with congestion, 23 with HCV, 23 with AIH, and 100 were considered as others. We focused on the first six categories for clinical significance. The model performance in the external test set was in line with our main results with the best performance obtained for AIH with a mean AUC of 0.962 followed by MASH and BO (Table 5, Column 2). The overall AUC by averaging the AUCs obtained for each individual category was 0.934 [95% confidence interval (CI): 0.909, 0.959] showing the robustness of our methodology on a completely unseen external validation set. The Mayo dataset (n = 3102) consists of a diverse patient population with key diagnoses distributed as follows: ACR (48.90%), HCV (29.59%), NASH (7.45%), BO (6.45%), Congestion (4.55%), and AH (3.06%). The GraftIQ model demonstrated consistent predictive performance, with AUC values exceeding 0.8 for MASH, ACR, and HCV on this dataset (Table 5, Column 3).
To further establish the model’s robustness, we performed an additional validation on two other international datasets, Hannover dataset (n = 224) which includes biopsies as follows BO (60.3%), HCV (7.1%), MASH (11.2%), AIH (5.8%), and ACR (15.6%) and NUHS, Singapore dataset with BO (9.6%), MASH (14.4%) and ACR (75.9%). The results with AUCs~0.7 from these datasets as shown in Table 5 columns 4 and 5 in Table 5, confirm the model’s reliability, reinforcing its applicability across different medical institutions.
Demonstration of clinical relevance
To demonstrate further clinical relevance of GraftIQ, we randomly chose one patient from each diagnostic category and applied the algorithm to obtain the probability of each possible diagnosis. We then manually reviewed the raw lab values for each patient to determine if the ML output made clinical sense or provided an expedited path to diagnosis that would otherwise have required further investigationGraphs demonstrating the probability of each diagnosis for each selected patient are displayed in Fig. 4a with a threshold of 50% for being classified into a specific category. For example, in patient 1147 with a liver biopsy demonstrating ACR, our algorithm determined an 81% probability of a diagnosis of ACR, followed by a 6% probability of MASH, 5% probability of HCV, 4% probability of BO, 3% probability of AIH and 1% probability of congestion. Subsequent review of the lab parameters for this patient that the algorithm used as part of its analysis demonstrated an ALP of 803, total bilirubin of 147, ALT of 227, and AST of 141.
a Pie chart illustrating the probability of a subject being classified into one of the six diagnosis categories using the hybrid GraftIQ model. Example subjects were selected based on their true diagnosis categories in the dataset, and higher probability in the pie chart represents their predicted class, e.g., for Subject 1501, the true diagnostic category in the test set was MASH and our hybrid model predicted subject being classified as MASH with a probability of 87%. b Illustration of a proposed interactive clinician-facing dashboard for clinical integration of our GraftIQ model.
For recurrent AIH, our hybrid model demonstrated an 88% probability of AIH. Review of the labs demonstrated ALP of 617, ALT of 377, AST of 377, and total bilirubin of 82. For post-transplant congestion, ALP was 331, ALT 66, AST 46, and bilirubin 25, whereas for HCV, ALP was 328, ALT 122, AST 66 and bilirubin was 15. Again, GraftIQ was able to identify these diagnoses with probabilities of 86% and 92%, respectively. Our patient with recurrent MASH had an ALP of 86, ALT 109, and AST 35 and our algorithm identified this diagnosis with a probability of 87%.
As can be seen from these results, the pattern of liver tests between the different diagnoses is not particularly different, and many clinicians would have difficulty distinguishing between the separate diagnoses based on these lab values alone. They would normally request further tests such as imaging or biopsy to clarify the diagnosis. As demonstrated by the high probabilities above, our hybrid algorithm would be able to provide diagnostic confidence much earlier in the care pathway, possibly streamlining the path to appropriate management measures.
Discussion
Liver transplant recipients often develop elevated liver enzymes post-transplant, indicating potential graft issues3. Upon detection, further diagnostic steps like imaging and biopsy are pursued, though they carry risks and may lead to delays in therapy. With no reliable noninvasive tools available, a probabilistic diagnostic ranking system could expedite treatment decisions and mitigate risks, as presented in our study. Our methodology using a multi-class neural network gave the best performance in predicting each individual category of diagnosis as well as in terms of an overall AUC of 0.902 as compared to the conventional machine learning approaches. We also observed no overlap in terms of the 95% confidence interval of our neural network approach [95% CI: 0.884, 0.919] versus the second-best performing approach of Random Forest [95% CI: 0.812, 0.839] validating the improvement provided by our proposed methodology.
The implementation analysis (refer to the section “Methods”) underscored the potential superiority of our ML model over the clinical judgment of clinicians. However, it also revealed an opportunity to refine our misclassification outputs through this analysis. By integrating clinician-based probabilities into our ML model, we imposed logical constraints that reflect the underlying principles of medical diagnosis. These constraints serve as regularization mechanisms, guiding the model to focus on relevant features and preventing it from overfitting to noisy or irrelevant data. As a result, the model’s predictions become more robust and reliable, increasing our overall AUC from 0.885 (ML model only) to 0.902 (ML model + clinical expertise), as they are aligned with established medical knowledge. Ultimately, the synergy between ML models and clinician expertise holds tremendous potential to optimize patient outcomes. Our model performed well across all external validation cohorts, with AUCs exceeding 0.7, reinforcing its robustness and generalizability. The Mayo cohort, the largest validation dataset, achieved AUCs above 0.8 for three categories, demonstrating strong predictive performance. The European cohort (Hannover Medical School) had a higher prevalence of replicative hepatitis E than HCV, reflecting its growing significance in Europe. While HCV is now less clinically relevant, our model’s ability to detect hepatitis E with a mean AUC of 0.768 further supports its adaptability. Additionally, our unseen UHN dataset achieved an overall AUC of 0.934, and validation on an Asian cohort (NUHS Singapore) further confirmed the model’s effectiveness across diverse populations.
Reviews of existing studies show that various ML algorithms, including neural networks, have been used in the context of liver disease and transplantation12. However, studies that focus on using ML to distinguish between various graft-related complications solely from demographic or biochemical parameters are limited12. For example, a study by Hughes et al from 2001 found that an artificial neural network trained on data from 117 patients with biopsies could predict the presence of ACR with an AUC of 0.90218,19,20. This study is limited by its small sample size and the fact that the algorithm can only diagnose one disease state—ACR. Other such examples specific to post-transplant complications include predicting the recurrence of primary disease, patient and graft survival, acute kidney injury, and HCC recurrence21. Most studies examining ML in the context of liver disease are those that automate diagnosis via image analysis of histopathologic slides22,23. This highlights our algorithm’s strength as the first demonstration of a neural network that can distinguish between multiple disease states based on demographic and laboratory data alone.
Neural Networks are usually perceived as black boxes wherein they improve predictive performance but are unable to provide the clinical variables driving the predictive ability. To enable the interpretability of our ML modeling, we explored two avenues: Firstly, through integrated gradient methodology, we were able to identify the most important clinical variables relevant to the diagnosis of each disease state. For example, important clinical variables for ACR included elevation in ALT, AST, and ALP elevation, which are well known to occur in the setting of ACR24. ACR was also associated with recipient age, donor age, and creatinine which are all known to be associated with ACR25. Recurrent AIH was most associated with recipient age, consistent with a recent large study that implicated younger recipient age as a risk factor for recurrence26. There was also a stronger association with cyclosporine than tacrolimus use, consistent with studies of the European liver transplant registry that found cyclosporine use after liver transplant for AIH predicted worse survival when compared to tacrolimus use27. Expectedly, post-LT biliary complications were most associated with ALP and total bilirubin, commonly regarded as the most important biochemical parameters for the diagnosis of biliary obstruction28. Lastly, recurrent MASH was associated with most of the clinical variables used for analysis, including hemoglobin, ALP, CNI use, and creatinine. MASH is often associated with chronic kidney disease before and after transplant, explaining the importance of creatinine in our algorithm29.
Secondly, through probability modeling, we were able to generate the risk of graft etiology for each patient, for example, our randomly chosen patient with ACR had lab values that could be associated with various diagnoses other than ACR, including biliary obstruction or recurrent autoimmune hepatitis. Despite these lab values, our algorithm determined the probability of ACR to be 81%. This shows that our algorithm could allow prompt initiation of ACR treatment, potentially expediting management, reducing resource use, and patient morbidity, and preserving graft and overall survival.
We acknowledge that the sample size for HCV-related graft injury was larger in our primary dataset, reflecting the overall cohort collected from 1992 onwards. However, recognizing that the clinical landscape has evolved and HCV is no longer a predominant cause of graft injury, we have retrained our neural network on stratified samples, differentiating between patients with HCV and non-HCV as the primary transplant indication. Our multiclass NN model demonstrates comparable performance to the original dataset, as presented in Table 4. This finding suggests that the model’s predictive power remains robust, even as the prevalence of HCV in modern transplant populations declines. We also observed that certain etiologies, such as ACR, performed better on the external UHN dataset. Specifically, ACR exhibited a lower AUC on the internal test set compared to the unseen UHN dataset, likely due to greater heterogeneity in clinical features and sample distribution within the internal cohort. While such heterogeneity can introduce noise, it is essential in the main training set to improve model robustness and ensure generalizability across diverse clinical scenarios.
We are also working towards creating a clinician-facing interactive dashboard for our proposed hybrid ML modeling (as shown in Fig. 4b), where clinicians can load patient covariates such as laboratory data (blood work, liver enzymes, etc.), demographic data, and clinical data (data on cholangitis, diabetes, etc.) directly from the patient’s digital record. Clinicians will then be able to run our ML model alongside their feedback, ensuring that diagnostic decisions are not restricted to the six predefined clinical rules but can be dynamically adjusted based on individual patient scenarios. As an output, the model will return a probability of the patient being classified into one of the six post-LT complications and furthermore, also get a list of the top clinical features instrumental in predicting the post-LT complication in the patient. This dashboard will have the potential to inform the clinician to proactively monitor the most important clinical features in the patient making our ML approach more clinically relevant and useful. To ensure seamless clinical deployment, the model is designed for inference-only use, requiring no retraining in clinical settings. This allows for straightforward integration into existing workflows without additional computational burden. The model was trained on a dedicated dataset and validated on independent test sets to preserve evaluation integrity. Currently, the model runs in 9.8 ms per patient in the test set on standard hardware (~15 s for the full test set), making it feasible for near real-time applications. Further optimizations, including pruning and quantization, are being explored to reduce computational demands while maintaining predictive accuracy.
We acknowledge some limitations of our study such as the exclusion of biopsies with dual diagnoses, potentially limiting generalizability at the current time. Additionally, we did not review the pathologies themselves and used solely the pathology report for diagnosis. Although we might have missed some undiagnosed post-transplant complications, our sample was large and representative enough to offer sound observations. Many patients are treated empirically for mild rejection, without a liver biopsy having been performed. Future research will focus on evaluating the model’s performance in a broader cohort, including cases without biopsy confirmation, to better understand its robustness in real-world clinical settings. Additionally, external validation across multiple independent cohorts strengthens the model’s generalizability, highlighting its potential utility even in diverse healthcare environments. Our model is designed as a decision-support tool, enhancing clinical decision-making by providing probabilistic predictions to complement physician expertise. In cases of disagreement, clinicians should assess the potential for false positives/negatives and consider further testing.
In conclusion, our hybrid multi-class neural network model, GraftIQ, demonstrates the promising potential for non-invasive diagnosis of graft pathology in liver transplant recipients. By combining clinical expertise with efficient deep-learning methodologies, we offer a robust framework for accurate diagnosis, potentially reducing the reliance on invasive procedures and improving patient outcomes. The external validation of our model across 3 international centers in the US, Germany, and Singapore resulted in promising predictive performance, supporting its potential applicability across diverse clinical settings. If validated and implemented clinically, we believe that this method has the potential to decrease the time to diagnosis, and dependency on liver biopsy and lead to earlier therapeutic interventions that will improve graft and patient survival over time.
Methods
Data collection and setting
Demographic, clinical, and laboratory data of all adult LTRs having undergone liver biopsies between January 17, 1992 and June 16, 2020 at the Ajmera transplant center, UHN, Toronto, Canada form the main dataset of our study. This study was approved by the Research Ethics Board at UHN (REB study # 21-6170). Since data was retrieved from medical records, an exemption from informed consent was granted by the REB committee. For the Hannover Medical School dataset, written informed consent was obtained from all patients, and the study was ethically approved (MHH Ethics Committee, Protocol No. 933). The study involving the Mayo Clinic dataset was approved under IRB number 24-002202, titled “Development of AI algorithms for clinical decision support in liver transplant patients.” and written informed consent was obtained from all patients. The NUHS Singapore dataset ethics approval was obtained under ECOS Ref: 2024-4614 and written informed consent was obtained from all patients. For all four datasets, the data were anonymized to protect patient privacy.
Study design
Definition and diagnosis of post-transplant complications
The first part of the study was to establish the most common etiologies for graft injury in LTRs from available biopsies. Biopsies were reviewed by two separate reviewers and labeled according to the appropriate diagnosis from the pathologist’s biopsy report. Biopsies were labeled as normal, acute cellular rejection (ACR), antibody-mediated rejection, biliary obstruction (BO), congestion, autoimmune hepatitis (AIH), viral (Hepatitis C, Hepatitis B, Cytomegalovirus (CMV), Epstein Barr virus (EBV)), metabolic-associated steatohepatitis (MASH) and toxic/drug-induced graft injury. Dual diagnoses were excluded from the analysis. The categories that were considered for statistical and ML analysis included ACR, BO, AIH, Hepatitis C infection (HCV), congestion, and MASH. The biopsies that read as normal as well as all other remaining diagnoses were grouped together as ‘Others’ due to the small number of samples as shown in Fig. 1b.
Demographic and clinical data for each diagnosis
The next step was to allocate demographic and clinical variables measured closest to the biopsy date, up to 30 days before each biopsy. The variables were selected based on relevance to post-transplant outcomes and their availability ensuring a missingness rate of less than 20%. The data included aspartate aminotransferase (AST), alanine aminotransferase (ALT), alkaline phosphatase (ALP), bilirubin, international normalized ratio (INR), white blood cells, hemoglobin, platelets, tacrolimus, and cyclosporine levels. Each biopsy was considered a ‘subject’, and the data was considered for each biopsy separately with an aim to assess which clinical variables triggered the biopsy. Demographic and pre-transplant variables (transplant indication, model for end-stage liver disease [MELD], donor type) for each biopsy were included in the analyses, as well as clinical data prior to the biopsy date (cholangitis, body mass index [BMI], diabetes, hypertension, and dyslipidemia). Missing data if any was imputed using the mean imputation method in the MICE library in R (missing data details in Supplementary Table 6).
Implementation analysis
In this implementation analysis, 12 hepatologists with diverse expertise were chosen to compare their predictive abilities with GraftIQ’s algorithm. Using a dataset of 30 cases covering six pathology categories, diagnostic accuracy was evaluated by comparing the ML model’s predictions to independent diagnoses by hepatologists. Overall accuracy was assessed to compare the ML model’s predictions with the independent diagnoses made by the hepatologists. This analysis helped us obtain insights into how the hepatologists made the prediction and narrow down on 6 simple clinical rules that hepatologists use to distinguish between etiologies namely: for BO, ALP, and bilirubin should be high; for ACR, the age of graft should be low given the increased risk in the early post-transplant phase; ALT > ALP and immune-mediated liver disease as an indication for transplant help to distinguish autoimmune hepatitis. For MASH, the age of the graft should be higher given its progressive nature, in addition to metabolic risk factors, including an elevated BMI in conjunction with more modest elevations of ALT. For HCV, ALT > AST with HCV as an indication for transplant and positive HCV serology are considered. Finally, for congestion, ALP and INR should be high without significant elevations of bilirubin in addition to the age of graft being low. These rules were additionally used in our clinical integration step.
Machine learning analysis
Multiclass neural network model
We propose a neural network model with multiple classes to carry out the classification task (Fig. 1a). Usually, ML classification algorithms restrict the possible outcomes to one of two values (a binary, or two-class model), however, given that our outcome included multiple primary diagnosis categories, we modified the learning function in the neural network to predict multi-class output. In our neural network methodology, we adopt the softmax approach, a multinomial logistic regression extension that directly supports multi-class classification. In our case, with six different diagnosis categories, the output layer consists of six nodes, each representing one of the classes. The softmax activation function is employed for each node, producing a probability distribution across all classes. The model is trained using the categorical cross-entropy loss function, which is well-suited for multi-class scenarios. This methodology fosters an ensemble-like behavior within the neural network, allowing it to collectively predict all classes while maintaining interpretability and computational efficiency.
We divided our dataset into 70% training and 30% test sets for model evaluation. Internal 10 times 10-fold cross-validation was performed to tune hyperparameters and compare our multiclass NN approach with other conventional ML approaches namely, random forest, support vector machines, logistic, lasso, and ridge regression (hyperparameter optimization and ablation study details provided in Supplementary Tables 4, 5, and 7). We conducted external validation on 4 independent datasets: (1) collected an additional 542 liver biopsies and clinical data from the UHN database between July 2020 until June 2024, (2) 3102 liver biopsies from Mayo Clinic collected between 1997 and 2023, (3) 224 biopsies from Hannover Medical School30,31, collected between 2008 and 2024, and (4) 83 biopsies from NUHS, Singapore32 collected between 2008 and 2024 (details of all datasets are provided in the External Validation section in the Supplementary document). To mitigate the potential impact of the relatively small sample size in some of the categories, we employed a repeated bootstrapping approach to assess the robustness and generalizability of the model. Specifically, we generated 1000 bootstrap samples from the external validation set, each consisting of a random sample with replacement. The model’s performance was evaluated using mean AUC and 95% confidence intervals to estimate the model’s stability and ensure it was not overfitting.
Expert-enhanced adaptive integration
To enhance the predictive capabilities of the neural network, we introduced an approach for integrating clinician expertise into the posterior probability calculation. Clinician input was obtained in the form of probability assessments for each graft injury category based on the 6 clinical rules mentioned in the section “Implementation analysis”. In this approach, if a clinical rule is satisfied for a specific diagnostic category, the probability for that category is set to 1. If multiple clinical rules are satisfied, the probabilities for each corresponding diagnostic category are distributed equally, ensuring that their sum equals 1. These clinician-provided probabilities of diagnosis categories were encoded as prior knowledge, reflecting expert assessments of diagnosis likelihoods based on patient data and to guide the inference process. Concurrently, a neural network architecture as illustrated in the section “Multiclass neural network model”, was employed to compute the likelihood of each diagnosis category from observed data. Bayesian inference principles were then applied to fuse the prior knowledge provided by clinicians with the likelihood computed by the neural network. This Bayesian fusion process yielded a posterior probability distribution over-diagnosis categories, capturing the integration of clinical expertise and data-driven predictions. Subsequently, value-based probabilistic inference techniques were employed to make decisions based on the posterior probability distribution in the last layer of the neural network. The integration was achieved through a weighted combination of the probabilities generated by the machine learning model and the clinician. The posterior probability \({P}_{{{\rm {integrated}}}}\left({C}_{i}\right)\) for each diagnosis category \({C}_{i}\) was calculated using the following formula:
where PML (Ci) represents the probability assigned by the multi-class NN ML model, PClinician (Ci) is the probability provided by the clinician for category Ci and α and β are the weight parameters to assign confidence clinician prediction and the ML prediction respectively. This iterative feedback loop not only provides valuable insights into clinicians’ domain expertise but also empowers the ML model to continuously learn from real-world scenarios, potentially resulting in more precise predictions.
Extract important features through neural networks
To identify the important variables in our predictive modeling, we used the integrated gradient (IG) methodology which is an interpretability technique for deep neural networks that attributes importance to input features by computing the integral of gradients along a path from a baseline input to the actual input33. We calculated gradients to measure the relationship between changes to a variable and corresponding changes in the model’s predictions. The gradient informs which variable has the strongest effect on the model’s predicted class probabilities where the higher the gradient, the more important the feature is considered for the classification task.
Inclusion and ethics statement
This study was conducted using retrospective, de-identified medical record data from the University Health Network (UHN) in Canada and did not involve fieldwork in resource-poor settings. Therefore, considerations regarding collaboration with local researchers, local ethics committee approvals outside UHN, and transfer of biological materials or traditional knowledge were not applicable. Research ethics approval was obtained from the UHN Research Ethics Board (REB), and informed consent was waived due to the retrospective and minimal-risk nature of the study. For all the external validation datasets, written informed consent was obtained. There was no stigmatization, incrimination, discrimination, or personal risk to participants arising from this research. No biological materials, cultural artifacts, or traditional knowledge were transferred, and no benefit-sharing measures were required. Relevant regional and international literature was reviewed and appropriately cited to ensure that the study was built on prior research.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings described in this manuscript are available in the article, in the Supplementary Information, and from the corresponding author upon reasonable request. Source data for each figure are provided with this paper. The raw University Health Network (UHN) dataset is not publicly available at this time due to the presence of sensitive patient information. Access to the UHN dataset may be subject to controlled access, and all research or research-related activities involving an external party may require, at the discretion of UHN, a written research agreement to define obligations and manage associated risks. Requests for access to the UHN dataset should be directed to Dr. Mamatha Bhat (Mamatha.bhat@uhn.ca), with responses provided within two weeks. Any use of the data will be subject to restrictions imposed by UHN through data use agreements. Source data are provided with this paper.
Code availability
Code for pre-processing and prediction is available at https://github.com/divya031090/multiclassNN34.
References
Wiesner, R. et al. Model for end-stage liver disease (MELD) and allocation of donor livers. Gastroenterology 124, 91–96 (2003).
Daugaard, T. R., Pommergaard, H. C., Rostved, A. A. & Rasmussen, A. Postoperative complications as a predictor for survival after liver transplantation-proposition of a prognostic score. HPB (Oxford) 20, 815–822 (2018).
Fedoravicius, A. & Charlton, M. Abnormal liver tests after liver transplantation. Clin. Liver Dis. (Hoboken) 7, 73–79 (2016).
Voigtländer, T. et al. Clinical impact of liver biopsies in liver transplant recipients. Ann. Transpl. 22, 108–114 (2017).
Hübscher, S. G. What is the long-term outcome of the liver allograft? J. Hepatol. 55, 702–717 (2011).
Khalifa, A. & Rockey, D. C. The utility of liver biopsy in 2020. Curr. Opin. Gastroenterol. 36, 184–191 (2020).
Rajkomar, A., Dean, J. & Kohane, I. Machine learning in medicine. N. Engl. J. Med. 380, 1347–1358 (2019).
Wang, D., Wang, Q., Shan, F., Liu, B. & Lu, C. Identification of the risk for liver fibrosis on CHB patients using an artificial neural network based on routine and serum markers. BMC Infect. Dis. 10, 251 (2010).
Wong, G. L. et al. Artificial intelligence in prediction of non-alcoholic fatty liver disease and fibrosis. J. Gastroenterol. Hepatol. 36, 543–550 (2021).
Nitski, O. et al. Long-term mortality risk stratification of liver transplant recipients: real-time application of deep learning algorithms on longitudinal data. Lancet Digit. Health 3, e295–e305 (2021).
Sharma, D. et al. Machine learning approach to classify cardiovascular disease in patients with nonalcoholic fatty liver disease in the UK Biobank Cohort. J. Am. Heart Assoc. 11, e022576 (2022).
Tran, J., Sharma, D., Gotlieb, N., Xu, W. & Bhat, M. Application of machine learning in liver transplantation: a review. Hepatol. Int. 16, 495–508 (2022).
Azhie, A. et al. A deep learning framework for personalised dynamic diagnosis of graft fibrosis after liver transplantation: a retrospective, single Canadian centre, longitudinal study. Lancet Digit. Health 5, e458–e466 (2023).
Nagai, S. et al. Use of neural network models to predict liver transplantation waitlist mortality. Liver Transpl. 28, 1133–1143 (2022).
Ayllon, M. D. et al. Validation of artificial neural networks as a methodology for donor–recipient matching for liver transplantation. Liver Transpl. 24, 192–203 (2018).
Briceno, J. et al. Use of artificial intelligence as an innovative donor-recipient matching model for liver transplantation: results from a multicenter Spanish study. J. Hepatol. 61, 1020–1028 (2014).
Rodriguez-Luna, H., Vargas, H. E., Byrne, T. & Rakela, J. Artificial neural network and tissue genotyping of hepatocellular carcinoma in liver-transplant recipients: prediction of recurrence. Transplantation 79, 1737–1740 (2005).
Hughes, V. F., Melvin, D. G., Niranjan, M., Alexander, G. A. & Trull, A. K. Clinical validation of an artificial neural network trained to identify acute allograft rejection in liver transplant recipients. Liver Transpl. 7, 496–503 (2001).
Hammann, F., Schöning, V. & Drewe, J. Prediction of clinically relevant drug-induced liver injury from structure using machine learning. J. Appl. Toxicol. 39, 412–419 (2019).
Ahn, J. C. et al. Machine learning techniques differentiate alcohol-associated hepatitis from acute cholangitis in patients with systemic inflammation and elevated liver enzymes. Mayo Clin. Proc. 97, 1326–1336 (2022).
Ferrarese, A. et al. Machine learning in liver transplantation: a tool for some unsolved questions? Transpl. Int. 34, 398–411 (2021).
Jain, D. et al. Evolution of the liver biopsy and its future. Transl. Gastroenterol. Hepatol. 6, 20 (2021).
Nam, D., Chapiro, J., Paradis, V., Seraphin, T. P. & Kather, J. N. Artificial intelligence in liver diseases: Improving diagnostics, prognostics and response prediction. JHEP Rep. 4, 100443 (2022).
Neil, D. A. & Hübscher, S. G. Current views on rejection pathology in liver transplantation. Transpl. Int. 23, 971–983 (2010).
Aloman, C. Acute rejection. In Mount Sinai Expert Guides: Hepatology (eds. Ahmad J., Friedman L. S. & Dancygier H.) 444–452 (John Wiley & Sons, Ltd 2014).
Montano-Loza, A. J. et al. Risk factors and outcomes associated with recurrent autoimmune hepatitis following liver transplantation. J. Hepatol. 77, 84–97 (2022).
Heinemann, M. et al. Longterm survival after liver transplantation for autoimmune hepatitis: results from the European Liver Transplant Registry. Liver Transpl. 26, 866–877 (2020).
Iacob, S. et al. Genetic, immunological and clinical risk factors for biliary strictures following liver transplantation. Liver Int. 32, 1253–1261 (2012).
Fussner, L. A. et al. The impact of gender and NASH on chronic kidney disease before and after liver transplantation. Liver Int. 34, 1259–1266 (2014).
Saunders, E. A. et al. Outcome and safety of a surveillance biopsy guided personalized immunosuppression program after liver transplantation. Am. J. Transplant. 22, 519–531 (2022).
Baumann, A. K. et al. Preferential accumulation of T helper cells but not cytotoxic T cells characterizes benign subclinical rejection of human liver allografts. Liver Transplant. 22, 943–955 (2016).
Tan, E. X.-X. et al. Impact of COVID-19 on liver transplantation in Hong Kong and Singapore: a modelling study. Lancet Reg. Health–West Pac. 16, 100262 (2021).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. Presented at: Proc. 34th International Conference on Machine Learning Research (2017).
Sharma, D. GraftIQ: A Hybrid Multi-class Neural Network for Graft Pathology Prediction (v1.0) https://github.com/divya031090/multiclassNN. https://doi.org/10.5281/zenodo.15225096 (2025).
Acknowledgements
We wish to thank Mary Grace Wong and Shruti Misra for helping to review the pathology reports. Supported by a Canadian Society of Transplantation grant, an American Society of Transplant (AST) grant, and Canadian Institutes of Health Research’s (CIHR) grant to M.B. Grants not specifically for this unfunded study. The content is solely the responsibility of the author. This study was not funded by industry. The work was supported by grants from the German Research Foundation (SFB738 project Z2; E.J.), the Transplantation Center Project 19_02 from Hannover Medical School (R.T.), and the Transplantation Center Project ZN3369 from Hannover Medical School/The Ministry of Science and Culture of the State of Lower Saxony (B.E.). B.E. was supported by the PRACTIS—Clinician Scientist program of Hannover Medical School, funded by the German Research Foundation (DFG, ME 3696/3).
Author information
Authors and Affiliations
Contributions
D.S. conducted statistical analyses, designed the algorithm, and wrote the manuscript. N.G. contributed to the study design, data extraction, pathology review, and editing of the manuscript. D.C. and S.N. wrote, formatted, and edited the manuscript. MN revised and edited the manuscript. A.A. and S.K. extracted clinical data and helped in the pathology review. J.A., B.E., R.T., E.T., and L.K.H. contributed external validation data. A.R., Y.H., S.G., M.S., and S.S. extracted UHN external validation data. B.E., R.T., K.D., L.L., N.S., C.T., and E.J. revised the manuscript. W.X. and M.B. designed the study, provided resources, and mentorship, and edited the manuscript. All authors had full access to all the data in the study and accepted responsibility to submit for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sharma, D., Gotlieb, N., Chahal, D. et al. GraftIQ: Hybrid multi-class neural network integrating clinical insight for multi-outcome prediction in liver transplant recipients. Nat Commun 16, 4943 (2025). https://doi.org/10.1038/s41467-025-59610-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59610-8
This article is cited by
-
Opportunities and challenges of artificial intelligence in hepatology
npj Gut and Liver (2026)