Abstract
Spatially-resolved transcriptomics (SRT) technologies now allow exploration of gene expression with spatial context. Recent advances achieving subcellular resolution provide richer data but also introduce challenges, such as aggregating subcellular spots into individual cells, which is a task distinct from traditional deconvolution. Existing methods often grid SRT data into predefined squares, which is unrealistic for accurately capturing cellular boundaries. We propose a method, STP, that integrates subcellular SRT data with nuclei-stained images to partition individual cells. STP first segments nuclei and maps their masks onto the SRT data, then uses a simulated-annealing-inspired approach to expand nuclear boundaries to the full cellular level. Evaluated on subcellular SRT datasets from Drosophila embryos at multiple developmental stages and from mouse embryos with a large field-of-view, STP demonstrated accurate single-cell partitioning, unveiling significant spatial tissue patterns and identifying undetected cell types beyond previous methods.
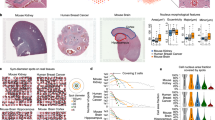
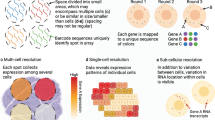
Similar content being viewed by others
Introduction
Spatially-resolved transcriptomics (SRT) technologies, named “Method of the Year 2020” by Nature Methods, opened a new door for simultaneously exploring the spatial and transcriptional context of tissues1,2 They could help biologists study the cell-cell interactions at a fine-grained level and identify spatial patterns in the microenvironment of whole tissues at a broad level3. Through positioning histological sections on glass slides arrayed with DNA oligo spots to capture spatially separated RNA molecules for sequencing, the first sequencing-based (seq-based) SRT technology born in 20164 allowed visualization and quantitative analysis of the transcriptome with spatially localization feature, namely Spatial Transcriptomics (ST), in individual tissue sections. However, every DNA spot of this technology has the diameter of about 100 μm, which involves 10 to 40 cells approximately, providing coarse-grained transcriptional patterns far from the single-cell level. Increasing the resolution of arrayed DNA oligo spots for RNA capture is the technical solution for dissecting genuine single-cell spatial transcriptome. Following the improvement of experimental pipelines, Slide-seq uses the arrays composed of 10 μm-diameter spots, which is nearly the single-cell resolution5,6. In 2022, Stereo-seq achieved 220 nm-diameter spots with a 500 nm center-to-center distance, which leads SRT into the stage of subcellular resolution1. Meanwhile, it could profile the area up to 13.2 cm × 13.2 cm, which allows biologists to explore much larger tissue sections that previous seq-based SRT technologies cannot accomplish. For instance, Stereo-seq profiled the whole mouse embryos in mid- and late-stage gestation with high definition, whereas no other existing technology could1.
The study of single-cell transcriptomes is essential for understanding the functional, developmental, and disease-related mechanisms of living organisms. In the first seq-based SRT4, the spots are low-resolution, containing dozens of cells with several blended cell types, which would obscure the genuine transcriptional pattern7. To overcome this limitation, many computational methods have been developed for the cellular deconvolution task requiring concurrent single-cell sequencing (scRNA-seq) data from the same tissue, enabling the exploration of the microenvironment in a fine-grained level8,9,10. In addition, computational methods, such as Tangram11, were designed to align scRNA-seq data to SRT data, which could reconstruct a genome-wide spatial map at the single-cell resolution. Despite the considerable success of these methods, there are two limitations: (1) systematic variation of gene expression profile between scRNA-seq and SRT data, and (2) potential mismatches of cell types shared between scRNA-seq and SRT data3.
In contrast, subcellular-resolution SRT, such as Stereo-seq, has posed an unprecedented computational challenge, i.e., instead of deconvolution of spots into single cells, it requires the aggregation of spots into single cells. Up to now, researchers have been binning a pre-defined number of spots into a square, e.g., X by X spots (Bin X), which is approximately equivalent to the size of average cell in their specific tissues12,13,1 However, the Bin X strategy combines the spots that do not correlate to true position and shape of a cell, offering gene expression data of an “artificial” cell. This would result in the inclusion of a massive number of incorrect spots for individual cells and profoundly affect downstream analyses. Thus, it is essential to propose a computational method for single-cell partition with genuine RNA distribution relevant to subcellular morphology based on subcellular SRT data.
Here, we proposed a method called STP, which enables the single-cell partition through the integration of subcellular SRT data and a nuclei-stained image. First, the nuclei-stained image is preprocessed and segmented into nuclei masks by a deep learning-based approach. Then, STP maps the masks of nuclei into SRT spots under the same coordinate system. Given all labeled spots, all nuclei would expand their cellular boundaries by our proposed simulated-annealing-inspired (SA-inspired) method until an energy cost is optimized. To evaluate STP, we collected multiple stages and slices of SRT data in Drosophila embryos and the large field-of-view SRT data of mouse embryos. For Drosophila embryos, STP enriches the accurate single-cell partition, reflecting significant and continuous spatial patterns of tissues in the individual slices, even in the stacking embryos via 3D alignment. In the experiments of mouse embryo, STP shows great performance on the large field-of-view SRT data, and the capability to refine the spatial structure of tissues and identify undetected cell types compared with the Bin X strategy.
Results
Overview of STP
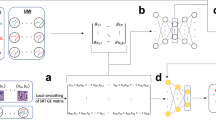
STP was developed as a pipeline that uses multiple data modalities, including nuclei-stained image and subcellular SRT data from the same tissue to enrich single-cell partition (Fig. 1A). In the beginning, the nuclei-stained image, which is performed by nucleic-acid-based staining, is enhanced adaptively14. Then, the noise of the image is removed by automatic thresholding based on maximization of inter-class variance between foreground and background15. After instance segmentation of all masks of nuclei by a deep learning method, STP maps the current masks of all nuclei to the pre-aligned SRT slices under the same coordinate system. Given the labeling of all spots based on instance nuclei, STP performs the cellular expansion of spots based on an SA-inspired strategy for individual cells from the nuclear to the cellular resolution. SA is a classical algorithm for solving an unconstrained optimization problem given an energy function16. For each iteration, randomly selected spots in the next layer outside the previous expansion are used to calculate the current energy function, considering the cellular morphology and expression profile with neighboring cells. The number of selected spots is determined using a predefined parameter \(T\), which controls the extent of spatial expansion, but it does not function as temperature in the classical SA algorithm. Based on the difference of energy between before and after expansion, the current solution of expansion would be kept by the probability of the current \(T\) even if the energy cost of the current solution is higher than the previous one. Once the \(T\) reaches the defined minimum \(T\), the iteration would stop (Fig. 1B and Supplementary Fig 1, Methods).
A STP consists of two main steps: nuclei segmentation and cellular expansion. Nuclei-stained image is firstly preprocessed and mapped on the pre-aligned SRT data. Then, each nucleus enables cellular expansion until the single-cell resolution based on the proposed simulated-annealing-inspired method. B The detail of cellular expansion (Methods). For each iteration, selected spots in the next layer are used to calculate the current energy function, considering the cellular morphology and gene expression profile with neighboring cells. The current solution of expansion would be kept by the probability of the current \(T\) even if the energy cost of the current solution is higher than that of the previous solution. Once the \(T\) reaches the defined minimum \(T\), the iteration would stop.
STP precisely partitions subcellular SRT data of Drosophila embryos at single-cell resolution
Drosophila is a widely-used model organism for biologists to explore embryogenesis and organogenesis, which has led to key advances in developmental biology including the identification of key developmental pathways and the genetic basis of developmental disorders17,12 The popularity of Drosophila on developmental biology research is benefited from following strengths: (1) Drosophila has cost-effectively convenient cultivation and a shorter life cycle, and (2) Drosophila shares lots of developmental pathways and genes with other organisms, enabling the collected knowledge to be widely applicable17.
In our study, we collected two samples of Drosophila embryos at early and late stages: 4–8 h and 8–10 h (hereafter termed E4-8 and E8-10), containing 17 and 25 7 μm-thick slices, respectively, processed by cryo-sectioning. Each slice in both samples is sequenced via Stereo-seq technology and enabled single-stranded DNA (ssDNA) nuclei staining, where multimodal datasets with spatial pre-alignment were generated: subcellular SRT data and a nuclei-stained image. In these two samples, E4-8 and E8-10 profiles have, on average, 9052 and 8404 genes in 164884 and 126129 spots for each slice respectively, where each spot in E4-8 and E8-10 captures an average of 7.47 and 7.21 counts of unique molecular identifier (UMI), i.e., normalized sequencing reads indicating accurate gene expression.
We applied STP to perform single-cell partition on E4-8 and E8-10 Drosophila embryo datasets. In the two samples from E4-8 and E8-10, the nuclei of all cells were firstly segmented and the cellular boundaries were expanded from nuclei, where different colors in the nuclei segmentation and single-cell partition results indicated individual instances of nuclei and cells (Fig. 2A). According to statistics of E4-8 results, around 50 spots were contained in each segmented nucleus and 100 - 200 spots contained in each partitioned cell after expansion (Fig. 2C). The UMIs of partitioned cells and segmented nuclei were also visualized and the cell sizes of partitioned cells are estimated (Supplementary Fig 2 and 3). The number of spots in E8-10 was slightly higher than that in E4-8, both in terms of the nuclei and the cells. To evaluate the validity and accuracy of STP, we integrated the results of partitioned cells across all slices in the same sample and conducted the downstream analysis to cluster all cells and annotate them to proper cell types according to their marker genes (Methods). From the visualization of tissue annotation from the two stages, tissues were spatially separated, which demonstrated that all cells were partitioned precisely to reflect their transcriptional diversity. We compared several cell types with marker genes proposed in previous studies, including amnioserosa, CNS, and yolk in E4-8 and salivary gland, CNS, and epidermis in E8-10. There were clearly correlated patterns in visualization between the spatial distribution of cell types and marker gene’s expression (Fig. 2B)12. Point-Biserial Correlation Coefficient (PBCC) is specifically designed for cases where one of the variables is binary and the other is continuous, which is mathmetically equivalent with Pearson correlation Coefficient (PCC)18 (Methods). In this case, PBCC is used to measure the correlation between all of pairs of cell types and marker genes, and visualized them as heatmaps, which showed the high PBCCs in both stages and most slices (Fig. 2D and Supplementary Fig. 8). Notably, for amnioserosa from slices 27 to 29 in E4-8 and salivary gland from slice 07 to 16 in E8-10, few of these types of cell appeared in the embryos, where visualization results showed verifiable patterns, and it may have decreased the PBCCs (Supplementary Figs. 4–7).
A Two examples from E4-8 and E8-10 Drosophila embryos, including original nuclei-stained image, nuclei segmentation, and their cell-type annotation results, single-cell partition, and their cell-type annotation results. In the results of nuclei segmentation and single-cell partition, each color represents the different instances of nuclei or cells. In the results of cell-type annotation from nuclei segmentation and single-cell partition, each color represents the annotated cell type. B Visualization of three representative cell types and their marker genes proposed in previous studies from two examples in E4-8 and E8-10 Drosophila embryos. C Several statistics of the results from nuclei segmentation and single-cell partition. The top two violin plots reveal the spot number contained in every slice from the results of nuclei segmentation and single-cell partition in E4-8 and E8-10 Drosophila embryos. D PBCC between three chosen cell types and their marker genes across every slice in E4-8 and E8-10 Drosophila embryos. E. The entropy of the results from nuclei segmentation and single-cell partition across every slice in E4-8 and E8-10 Drosophila embryos. Source data are provided as a Source Data file.
To ensure the necessity of expanding nuclei to the single cell, we also conducted the same analysis on nuclei segmentation results without expanding to the cell boundary using STP. Compared to the results of nuclei segmentation, the single-cell partition with cytoplasmic expansion annotated more comprehensive cell types (Fig. 2A). In E4-8, amnioserosa was mistakenly annotated as yolk in the nuclei segmentation results, but was more appropriately annotated in the single-cell partition results. Similarly, the CNS from the nuclei segmentation results in E8-10 was also wrongly assigned as muscle, but the results of single cell partition could enrich more fine-grained cell type annotation to identify CNS correctly. To further demonstrate the necessity of cell expansion, the entropy of each slice in E4-8 and E8-10 was quantified to measure the disorder of the spatial distribution of cell types (Methods). Well-partitioned cells should reflect their specific transcriptional patterns to cluster appropriately by cell type, resulting in a more orderly and clear spatial distribution of cell types. The calculated entropy demonstrated that partitioned cells had systematically lower entropy compared to segmented nuclei in both stages. It proved that expanding from nuclei to whole cells transformed the spatial organization of identified cell types to a better ordered arrangement (Fig. 2E).
Moreover, an ablation study was conducted to evaluate the performance of STP under different conditions: \(\alpha=0\), \(\beta=0\), and \(\gamma=0\), which correspond to three assumptions related to the shape of the cell, similarity with neighbor cells, and the size of the cell, respectively, (“Methods”). In the example of E4-8, after partitioning and clustering the cells, we calculated the PBCCs between all of the cell types and their marker genes. Compared to the original STP settings, we found that all three conditions resulted in lower PBCCs (Supplementary Fig.). Notably, the condition where \(\beta=0\) had the worst performance, indicating that the assumption that adjacent cells share similar transcriptional profiles is crucial.
STP reconstructs continuous 3D cell-type patterns of Drosophila embryonic development
In embryonic development of Drosophila, cells of the same cell type tend to have a continuous organization, due to the fact that embryonic tissues differentiate and develop from their respective stem cells, and cells usually migrate and divide in a directed manner based on their fate19. Furthermore, previous studies using genetic lineage tracing have demonstrated that cells of the same origin tend to remain together during development and give rise to clusters of cells in specific regions of the embryo20.
To verify the continuity of identified cell types across multiple slices, we applied STP to all slices in E4-8 and E8-10 of Drosophila embryos, where on average 1660 and 1125 cells per slice were obtained in E4-8 and E8-10, respectively. During the downstream analysis, we used Harmony21 to remove batch effects among slices in the same embryo, which would benefit the accuracy of downstream cell-type clustering and annotation (“Methods”). Given the cell-type annotation of all cells, we also visualized the cell-type distribution of partitioned cells and segmented nuclei across these slices, which revealed visually continuous organization through consecutive slices (Fig. 3A, Supplementary Figs. 10–13). Then, we projected all partitioned cells through Uniform manifold approximation and projection (UMAP)22 and distinguished different cell types and slices with corresponding colors (Fig. 3B). The UMAP results of E4-8 and E8-10 across slices both showed no biased distributions among different slices, which verified successful removal of batch effects (Fig. 3B). Based on this observation, cells were then clustered by the annotations of cell types and all cell types were well-separated with distinct boundaries in UMAP results (Fig. 3B). To observe the cell-type organization at the 3D level, we used a 3D alignment method called PASTE23 to align all partitioned cells from consecutive slices along z-axis. As the visualization of 3D point cloud revealed, individual cell types in 3D also exhibited spatially continuous organizations in E4-8 and E8-10 under different angles (Fig. 3C and Supplementary Fig. 14).
A Cell type annotation of partitioned cells across five consecutive slices in E4-8 and E8-10 of Drosophila embryos. B UMAP results of different cell types and slices in E4-8 and E8-10 Drosophila embryos. C The 3D visualization of all cell types and three representative cell types in E4-8 and E8-10 Drosophila embryos.
To comprehensively evaluate the accuracy of STP, two scRNA-seq datasets of Drosophila embryos from E4-8 and E8-10 were collected24. Tangram was used to match the cell pairs with the highest gene expression similarity between scRNA-seq data and STP-partitioned cells from SRT data11. This way, we transferred the cell type annotation from scRNA-seq data to the paired cell in SRT data. The spatial distributions of STP-partitioned cells were visualized with the cell type annotation transferred from scRNA-seq mapping results in comparison to the clustering results from SRT data only (annotated by clustering results) (Supplementary Figs. 15 and 16). From the visualization, the distributions between mapping and clustering results were highly similar. To quantitatively evaluate the performance, the Adjusted Rand Index (ARI) score was calculated to compare the similarity between clustering cell types and the mapping ones for all cells (Methods and Supplementary Fig. 17). PCCs were also measured between the expression profiles of the identified single cells from STP and the mapped cells from scRNA-seq data (Supplementary Fig. 18). STP achieved high values in terms of both the ARI scores and PCCs, suggesting the correctness of the partitioned cells.
STP refines the tissue identification and organization in mouse embryo
Mouse embryonic development is valuable to study as it is a well-established model system for studying mammalian development. Mouse has a short gestation period of several weeks, and shares extensive genetic and physiological similarities with humans, serving as an important model for studying human developmental and pathological processes25,26 We collected a tissue slice of mouse embryo from embryonic day 14.5 (E14.5), the mid-stage of mouse embryonic development, which was also sequenced by Stereo-seq and stained using ssDNA dye. This large field-of-view subcellular SRT data contained an enormous number of spots (77 million spots) and profiled 27290 genes among all of the spots.
We applied STP to analyze nuclei-stained images and SRT data from an E14.5 mouse embryo, and conducted the downstream analysis to annotate the tissues according to the marker genes of each cluster of partitioned cells (Methods). After the quality control by deleting low-quality cells, STP identified 393,555 cells in total (as described in the methods section). The distribution of cells annotated by corresponding tissues revealed distinct spatial organization of tissues (Fig. 4A). We also visualized the spatial distribution of several tissues and their corresponding marker genes’ expression, including brain, radial glia cell, muscle and choroid plexus (Fig. 4B). The high similarity between spatial distribution of tissues and marker genes’ expressions validated the reliability of STP in partitioning cells based on transcriptional pattern. To comprehensively quantify the PBCCs of identified tissues and corresponding marker genes, we chose three marker genes from each tissue in mouse embryo according to previous study1 and integrated the calculated PBCCs of all tissues into a heatmap (Fig. 4C). From the heatmap, most of tissues exhibited high correlations with their corresponding marker genes, which validated the robustness of STP in single cell partition.
A Cell-type annotation of all partitioned cells from the E14.5 mouse embryo. B Visualization of four representative tissues and their marker genes proposed in previous studies. C The heatmap visualizing the PBCC between all cell types’ distribution and their marker genes’ expression. D The findings on the mouse embryo by STP. The spatial organization of cartilage primordium could be refined, and the spatial organization of radial glia cell could be corrected compared with the Bin X strategy. Source data are provided as a Source Data file.
More remarkably, STP showed a significant improvement in identifying rare tissues and refining the spatial organization of tissues than previous Bin X strategy in the mouse embryo, where X is set as 501 (Fig. 4D). Cartilage primordium refers to the earliest stage of cartilage formation during mouse embryonic development that will eventually differentiate into chondrocytes and form the cartilage matrix27. From the comparison among the spatial expression of marker genes of cartilage primordium, the spatial distribution of cartilage primordium from STP and Bin X strategy, STP identified an obviously fine-grained and accurate distribution of cartilage primordium compared to the results from Bin X in a previous study1. This observation suggests that the Bin X strategy did not fully capture the transcriptional patterns of cells due to its limitations in framing each “cell” with the same size and shape, and it relied heavily on the chosen value of X, making it less robust. In contrast, STP identified the cartilage primordium in a refined manner, demonstrating its superiority in identifying tissue-specific transcriptional patterns.
During mouse embryonic development, radial glia cells act as scaffolds for the migration of neural progenitor cells, facilitating the formation of the cortex and other brain structures. Additionally, radial glia cells themselves can differentiate into neurons and glia, contributing to the development of the nervous system28. However, a previous study1 found that the Bin X strategy mistakenly identified the radial glia cells as a part of the brain in the E14.5, but STP identified radial glia cells successfully with distinct transcriptional patterns. Furthermore, the previous research also confirmed that radial glia cells typically appeared during the development of the mouse embryo between E11.5 and E14.529. These findings also substantiated that STP had the capability to identify correct cell types or discover sub-cell types.
To validate the quality of partitioned cells, STP was benchmarked with other methods, including k-nearest-neighbors (KNN), Voronoi, and the watershed algorithm on the E14.5 mouse embryo dataset. The same nuclei identified by STP were used to expand the cell boundaries using the KNN (\(k=\) 50, 100, 150, and 200) and Voronoi algorithms30. The watershed algorithm was applied as suggested in ref. 1. Since the watershed algorithm used nuclei-stained images to segment all nuclei as “cells”, the number of identified genes per cell using the watershed algorithm (an average of 82.1 genes per cell) was significantly lower than with STP (an average of 187.7 genes per cell) (Supplementary Fig. 19). In addition, KNN algorithm is used to partition the individual cell from segmented nuclei. The number of K from KNN indicates the number of expanded spots, and the neighbors are the nearest spots to the centroid of the segment nuclei. After clustering the identified cells using these three compared methods, spatial distributions of cell types were visualized (Supplementary Fig. 20). From the visualization, KNN with different \(k\) and Voronoi could not identify several cell types, including radial glia cells and the choroid plexus. Using the watershed algorithm, cells from the heart, choroid plexus, and cartilage primordium were not identified. To quantitatively evaluate the performance of the partitioned cells, PBCCs were calculated between the spatial distribution of the identified cell types and their marker genes (Supplementary Fig. 21). Most of the cell types identified by STP had significantly higher PBCCs than those identified by KNN, Voronoi, and watershed algorithms. Cell types with zero PBCC values indicated that they were not identified by the corresponding algorithms.
To compare the quality of partitioned cells with real scRNA-seq datasets, we also calculated the Mean Square Error (MSE) and PCC of gene-gene correlation matrices between the collected scRNA-seq datasets and partitioned cells from all methods. This aims to measure the similarity between partitioned cells obtained by multiple methods and the scRNA-seq datasets (“Methods”). We collected a scRNA-seq dataset from E14.5 mouse embryos, including around 240,000 cells from multiple mouse samples31. Through the MSE and PCC for all methods, we observed that STP achieves a lower MSE and a higher PCC than other methods, indicating that the gene-gene correlation in the partitioned cells from STP showed higher resemblance to scRNA-seq datasets than other methods (Supplementary Fig. 22).
The sizes of partitioned cells are also estimated from different methods and visualized across all cell types, where the y-axis denotes the distribution of the area of partitioned cells (μm²) (Methods, Supplementary Fig. 23). From the visualization, different cell types indeed have different cell sizes. Although there is no rigorous measurement of cell sizes for each cell type in mouse embryos, we found several studies about the general cell sizes in the embryo and adult mouse as references. The general diameter of mouse embryonic cells is around 7 to 17 μm, and the mean cell diameter is 13.82 μm for a 20 g adult mouse32,33. Thus, the areas of mouse embryonic cells are around 38 to 227 μm². From the visualization results, the cell sizes from partitioned cells by STP are similar to the previously described cell sizes from previous studies. On the other hand, the sizes of partitioned cells by Voronoi are the largest among all methods because it expands the boundaries of cells until all spots in the sample are assigned to cells, which is not reasonable in this case. Regarding KNN, different values of k would definitely affect the partitioned cell sizes, which are fairly similar across different cell types.
Discussion
In this article, we proposed a pipeline of single-cell partition called STP, utilizing multimodal data resources: nuclei-stained image and subcellular SRT data. The STP strategy consists of two steps: nuclei segmentation for each cell and SA-inspired expansion of the cellular boundary. STP was applied to multi-stage and multi-slice Drosophila embryos datasets, on which it partitioned individual cells and reconstructed continuous spatial patterns of multiple cell types across consecutive SRT slices. In the large field-of-view dataset of mouse embryo, STP could also reveal distinct cell-type patterns. More excitingly, STP could refine spatial patterns of cell types and even correct misclassified or previously undiscovered cell types by the existing Bin X strategy.
In addition to the current advantages of STP, there are some limitations to be overcome in the future. First, the effectiveness of STP relies on the optimized selection of multiple hyperparameters, such as the setting of initial \(T\), the reduction rate of \(T\) and so on. Actually, these hyperparameters were designed to account for the significant variation in cellular morphology across different species and tissues. For instance, embryonic stem cells, which have the potential to differentiate into various cell types, typically possess larger nuclei with relatively less cytoplasm to maintain their stemness34. On the other hand, one main direction to further improve STP is the ability to handle datasets that contain cells with heterogeneous distributions and various shapes or sizes. In this scenario, not only should the hyperparameters \(\alpha\), \(\beta\), and \(\gamma\) be considered, but also more assumptions about the specific properties of the obtained dataset should be taken into account. Thus, there is a need to further develop a single-cell partition method specifically for heterogeneous datasets that include cells with various shapes and sizes. Second, even if current SRT technologies could achieve subcellular resolution, there is room for improvement in terms of the capture efficiency of each spot. The low sensitivity of existing subcellular SRT technologies leads to a significant dropout of transcriptional data during single-cell partitioning, which could not be remedied by regulation of multiple hyperparameters of STP (Supplementary Figs. 24 and 25). Other than the low sensitivity from SRT data, STP also relies on the quality of ssDNA nuclei-stained images. While STP incorporates several preprocessing strategies, the effectiveness of nucleus staining is impacted by various factors, including the choice and concentration of staining agents, staining time, and pH value. Under certain conditions, distinguishing nuclei through staining becomes challenging, including (1) over-staining or under-staining and cell breakage or damage during the experimental process, and (2) dense organization of cells, where stained nuclei may adjoin or overlap with each other35,36 Advancements in SRT and nuclei-staining technologies in the future are expected to improve data quality, thereby enhancing the outcomes of STP. Third, in this study, we compared STP against several alternatives and demonstrated its superior performance through multiple benchmarks. However, we acknowledge that formal statistical validation, such as hypothesis testing or confidence intervals, was not explicitly conducted to quantify the significance of these improvements. While the observed trends strongly suggest that our method outperforms existing approaches, future work could incorporate rigorous statistical testing to further validate these findings.
By leveraging STP, SRT data can further enhance our understanding of the spatial context and organization of these single-cell-level SRT data. This spatial information provides valuable insights for biologists and clinicians, allowing them to explore the intricate relationships between the higher-dimensional landscape of cells under physiological and pathological conditions. It has the potential to make significant strides in the diagnosis and prognosis of human diseases. The high-dimensional profiling of cells within their spatial context can offer a deeper understanding of disease mechanisms, identify localized cellular alterations associated with pathology, and potentially uncover new therapeutic targets.
Methods
STP pipeline
The main procedure of STP includes nuclei segmentation and single-cell partition. To segment all nuclei in the nuclei-stained image, enhancement and denoising should be done before nuclei segmentation. We used CLAHE (Contrast Limited Adaptive Histogram Equalization) to improve the contrast between foreground (nuclei) and background, which allows for a more localized adjustment of the contrast while avoiding over-amplification of noise as the traditional Histogram Equalization technique does14. The ‘clipLimit’ is set to 2, and ‘tileGridSize’ is set to 8. After image enhancement, the noise from the background was removed by a thresholding method called Otsu, which tries to find an optimal threshold that maximizes the inter-class variance of pixel intensity in the image15. Then, the pixel whose value was less than the optimized threshold would be set to 0.
After the preprocessing procedure, we used an existing method called Cellpose to segment all nuclei from the nuclei-stained image37. Cellpose is a deep learning method based on the architecture U-Net, which could segment all instances of nuclei or cells38. We used the pre-trained model released from Cellpose and chose the ‘model_type’ as ‘nuclei’. After segmentation of nuclei, the bounding boxes and masks for all nuclei were outputted. If the image size is too large to process, such as mouse embryo slices we used (14812 × 21749), the image would be grided as smaller patches that could speed up the segmentation process. In addition, the subcellular SRT data could be mapped on the segmented nuclei, and the spots of SRT would be annotated as the indices of contained nuclei or background. In other words, we finally obtained the transcriptional profiles and masks of all nuclei.
Following the nuclei segmentation, STP expanded the cellular boundaries of all nuclei and enriched single-cell partition based on SA16. Initially, \(X=\{{x}_{1},{x}_{2},\ldots,{x}_{M}\}\) includes the indices of nuclei (\(M\) nuclei totally) and \({G}_{{x}_{i}}\) represents the gene expression vector of nucleus \({x}_{i}\). We initialized the labels of all spots \({L}_{{B}_{j}}\) from SRT data with their indices of contained nuclei. Otherwise, spots outside any of nucleus would be labeled as 0.
To start the cellular expansion, we needed to set some hyperparameters first. \(T\) is the number of expanded bins for a nucleus in each iteration, which does not function as temperature in the classical SA algorithm, and we set the initial \(T\) as \({T}_{0}\) and minimum \(T\) as \({T}_{\min }\). \(r\in ({\mathrm{0,1}})\) refers to the reduction rate of \(T\) (defined below). \({F}_{e}({x}_{i})\) is the energy function of nucleus \({x}_{i}\) in the epoch \(e\), which represents the iteration of the algorithm.
The energy function considers the size, shape of the cell, and transcriptional pattern with adjacent cells, under the following assumptions: (1) Shape assumption: the distances from all of outside spots to the center of the nucleus are similar, which means that the shape of cells should be similar to regular polygons. (2) Non-overlapping assumption: cells should be next to each other without any overlap. 3) Smoothness assumption: adjacent cells should share similar transcriptional profiles. Based on these assumptions, we designed the energy function as three parts: \({F}_{{size}}(x)\), \({F}_{{shape}}(x)\) and \({F}_{{adj}}(x)\). For \({F}_{{size}}\left(x\right)\), we denoted \({C}_{x}\) as the number of spots contained in the cell \(x\) and \(c\) as the expected number of contained spots in each cell. From the following equation, we measured how close the number of spots segmented by STP is to the expected value \(c\), which represents the total number of spots in the slice divided by the number of segmented nuclei, serving as an estimate of the average cell size.
In the definition of \({F}_{{shape}}(x)\), \({Bound}(x)\) represents the coordinates of spots at the outside layer of cell \(x\) and \({Centriod}(x)\) refers to the coordinates of the central spot of cell \(x\), which could be calculated from the mask of the cell. \(D\) represents the variance of this list of distances mentioned above. Thus, \({F}_{{shape}}\left(x\right)\) calculates the variance of Euclidean distance between the spots at the outside layer and the central spot. As the following equation shows, we expected that the variance of Euclidean distances between all spots from outside the layer to the central spot would be as small as possible.
To quantify \({F}_{{adj}}(x)\), we used KD-tree to identify the \(k\) adjacent expanding cells for each cell, whose transcriptional profile is \({G}_{x}\), where \(k\) is a hyperparameter to regulate39. Then, we extracted the transcriptional profiles from adjacent cells and averaged them as a vector \({G}_{{adj}}(x)\). Following the equation below, we expect that a cell has similar transcriptional profiles to the adjacent cells.
With the definition of \({F}_{{shape}}(x)\), \({F}_{{adj}}(x)\) and \({F}_{{size}}(x)\), we defined the energy function of each cell at each iteration with three hyperparameters \(\alpha\), \(\beta\) and \(\gamma\):
Firstly, after setting the initial \(T\) as \({T}_{0}\), a loop would start with multiple iterations. In each iteration, we set the current \(T\) as \(\lfloor rT\rfloor\). Then, we repeated the following procedures for each segmented nucleus. We randomly chose \(T\) spots of the first layer outside the nucleus and label these spots as \(i\). The identification of the first layer outside each nucleus is based on the binary dilation, a basic operation of mathematical morphology. Subsequently, new \({F}_{e}({x}_{i})\) could be calculated and \(\Delta F=\,{F}_{e-1}\left({x}_{i}\right)-\,{F}_{e}({x}_{i})\). If \(\Delta F < 0\), the current updated labels of \({x}_{i}\) could be kept. Otherwise, the current updated labels of \({x}_{i}\) will be kept by the probability \({e}^{-\Delta F/T}\). Every nucleus would be processed sequentially by the procedures mentioned above until the last nucleus and current iteration is finished. Following the reduction of \(T\) in each iteration, the entire pipeline would stop when current \(T\) reaches \({T}_{\min }\) (Supplementary Fig. 1).
Downstream analysis
To evaluate the performance of STP on the SRT data from Drosophila embryos, we utilized the partitioned single-cell transcriptional profile to conduct downstream analysis. To control the quality of partitioned single-cell data in each slice from E4-8 and E8-10 embryos, we eliminated the cells whose counts were lower than 30 through all genes, and genes whose counts were lower than 10 through all cells by scanpy40. After normalization, we integrated slices from the same embryo and aligned them with a 3D alignment method called PASTE with the \(\alpha\) as 0.123. We set the interval of two adjacent slices as 14 in the relative coordinate system because the thickness of each slice was 7 μm. Then, we selected 1500 highly variable genes from all of cells by scanpy40 and used Harmony21 to remove the batch effects among slices in the embryo. Next, we used Dynamo41 to compute two neighboring graphs based on expression profiles and spatial coordinates, where number of neighbors were set as 30 and 10, respectively. Finally, the Leiden algorithm42 was used for clustering the cells with the resolution of 0.6, and the top 50 variable genes in each group were outputted, which were the reference for annotating the cell type to cluster further.
In the SRT data of mouse embryo, we conducted a similar procedure as downstream analysis without the 3D alignment and batch effect removal. We set 0.8 as the resolution when used Leiden algorithm42 to cluster the partitioned cells. The annotation of cell types were based on the best correlation between variable genes from each cluster and the proposed marker genes in the previous studies1.
Entropy definition
To prove the necessity of cellular expansion, we decided to quantify the entropy of nuclei and partitioned cells under the expectation that the spatial organization would become more structured after cellular expansion. Specifically, we hypothesize that cell types will exhibit increased spatial coherence, with similar cell types tending to cluster together rather than being randomly distributed. We firstly clustered the transcriptional profile of nuclei and partitioned cells through all slices in both E4-8 and E8-10, and labeled each cell with the index of its assigned cluster. Then, we created the graphs for annotated nuclei and cells where nodes were the labeled nuclei/cells, and edges were calculated by K Nearest Neighbors based on the spatial coordinates. Here, we set K as 6. For all \(M\) nuclei/cells in the graph, their entropy could be calculated as \(H\left(M\right)={\sum }_{m}^{M}H(m)\), where \(m\) represents the index of each nucleus/cell. The entropy of \({mth}\) node in the graph is \(H\left(m\right)=-{\sum }_{i=1}^{K}{p}_{i}{\log }_{2}{p}_{{{\rm{i}}}}\), where \({p}_{{{\rm{i}}}}\) represents the probability that the label of \({ith}\) neighbor node occurs.
Calculation of gene-gene correlation matrices
Firstly, we intersected the identified genes between collected scRNA-seq datasets and partitioned cells from all compared methods (STP, nuclei, Voronoi, KNN with k = 50, 100, 150, and 200), resulting in 18,943 genes. With this set of genes, we calculated the gene-gene correlation matrix (18,943 × 18,943) using the PCC for all datasets. Since the calculated matrices shared the same shape and gene order in both rows and columns, we proceeded to calculate the Mean Square Error (MSE) and PCC of the gene-gene correlation matrices between scRNA-seq datasets and partitioned cells from each method, one by one. Specifically, to calculate the MSE, we summed all the squared differences between corresponding elements from the two matrices and divided this sum by the total number of elements in the matrices. To calculate the PCC, we flattened each matrix into a one-dimensional vector and then used the two flattened vectors to calculate the PCC between them.
Estimation of size of partitioned cells
Since the spots are arranged hexagonally, we assume the partitioned cells are hexagons to calculate the area of the cell size conviently. From the previous study, the diameter of the spot is 220 nm, and the spot-to-spot distance is 500 nm. Firstly, we determine the number of layers \(L\) based on the total number of spots \(k\). Since the number of spots in a hexagonal lattice with \(L\) layers is \(3L(L-1)+1\), we compute the layers as \(L=\lceil \frac{3+\sqrt{12k-3}}{6}\rceil\). Here, ⌈⋅⌉ denotes the ceiling function, which rounds up to the nearest integer. Secondly, the side length \(S\) of the hexagon, the distance from the center of the hexagon to the outermost layer of spots, would be \(500\times \left(L-1\right){nm}\). Finally, the area of a regular hexagon with side length \(S\) would be \(\frac{3\sqrt{3}}{2}{S}^{2}\).
Settings of hyperparameter
There are four hyperparameters to set: \(T\), \(\alpha\) (shape of cell), \(\beta\) (similarity with neighbor cells) and \(\gamma\) (size of cell), and we set the same hyperparamters in the two Drosophila embryos datasets and mouse embryo datasets: T = 100, \(\alpha=0.5\), \(\beta=0.1\), and \(\gamma=3\). These parameter were chosen based on a grid search. We chose hyperparapmers following these ranges: T ( = 50, 100, and 150), \(\alpha\) (=0.1, 0.5, and 0.9), \(\beta\) (= 0.1, 0.5, and 0.9), and \(\gamma\) (= 1, 3, and 5). Following these multiple combinations of hyerparameters, we selected the hyperparameter combination by visual inspection to confirm alignment between spatial distributions of cell types and known biological spatial patterns.
The mathematical equivalence between PBCC and PCC
PBCC is often used when one variable is binary and the other is continuous, which is a special case of the PCC43. Since the mathematical structure of both coefficients is the same when the binary variable (0 or 1) is treated as a continuous variable in the PCC formula, PBCC and PCC are mathematically equivalent18.
The formula of PBCC is as follows44,45
where \(\bar{{Y}_{1}}\) and \(\bar{{Y}_{0}}\) are the corresponding mean values of the continuous variable for the group where the binary variables are 1 and 0. \({s}_{Y}\) denotes the standard deviation of the continuous variable across both groups. \({n}_{1}\) and \({n}_{0}\) are the numbers of observations where the binary variables are 1 and 0. \(n\) is the total number of observations \(n={n}_{1}+{n}_{0}\).
The formula of PCC is as follows:
where \({x}_{i}\) and \({y}_{i}\) are the sample values of the two variables, and \(\bar{x}\) and \(\bar{y}\) are the means of the two variables. \(n\) is the total number of observations.
From these two formulas, we can observe that in PBCC, the formula uses \(\bar{{Y}_{1}}-\bar{{Y}_{0}}\), which calculates the difference between the means of the two groups. When one of the variables is binary, PCC’s covariance calculation is mathematically equivalent to the standardized mean difference used in PBCC. In addition, PBCC uses \({s}_{Y}\), the standard deviation of the continuous variable, while PCC also normalizes the covariance by the standard deviation of both variables. The standardization steps are also equivalent in both cases. Therefore, PBCC is a special case of PCC when one variable is binary, and the two are mathematically equivalent in this situation.
Adjusted Rand Index (ARI)
To evaluate the accuracy of mapping scRNA-seq data to partitioned cells in the spatial transcriptomics data, we use the Adjusted Rand Index (ARI) to quantify the agreement between predicted and true cell type annotations. Given that each partitioned cell in the spatial transcriptomics and scRNA-seq data has a known cell type annotation, we assign predicted labels based on the scRNA-seq-derived mapping. The ARI is computed as:
where Rand Index (RI) is \(\frac{{Number\; of\; agreeing\; pairs}}{{Total\; possible\; pairs}}\), which is the proportion of correctly assigned cell type pairs among all possible pairs, and \({\mathbb{E}}[{RI}]\) is the expected Rand Index under a random assignment. The denominator ensures that ARI is normalized between − 1 and 1.
Statistics & reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The SRT data of Drosophila embryos in E4-8 and E8-10 is from ref. 24. The SRT data of the mouse embryo in E14.5 is from ref. 1. Source data are provided in this paper.
Code availability
The code is released on the github: https://github.com/leihouyeung/STP46
References
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Marx, V. Method of the Year 2020: spatially resolved transcriptomics,” Nat. Methods 18, 9–14 (2021).
Li, H. “A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics. https://doi.org/10.5281/zenodo.7674290 (2023).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Rodriques, S. G. et al. Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Williams, C. G., Lee, H. J., Asatsuma, T., Vento-Tormo, R. & Haque, A. An introduction to spatial transcriptomics for biomedical research. Genome Med. 14, 68 (2022).
Li, H., Li, H., Zhou, J. & Gao, X. SD2: Spatially resolved transcriptomics deconvolution through integration of dropout and spatial information. Bioinformatics 38, 4878–4884 (2022).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomic. Nat. Biotechnol. 40, 517–526 (2021).
Ma, Y. & Zhou, X. Spatially informed cell-type deconvolution for spatial transcriptomics. Nat. Biotechnol. 40, 1349–1359 (2022).
Elosua-Bayes, M., Nieto, P., Mereu, E., Gut, I. & Heyn, H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab043 (2021).
Biancalani, T. et al. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat. Methods 18, 1352–1362 (2021).
Wang, M. et al. High-resolution 3D spatiotemporal transcriptomic maps of developing Drosophila embryos and larvae. Dev. Cell 57, 1271–1283 (2022).
Liu, C. et al. Spatiotemporal mapping of gene expression landscapes and developmental trajectories during zebrafish embryogenesis. Dev. Cell 57, 1284–1298 (2022).
Reza, A. M. Realization of the contrast limited adaptive histogram equalization (CLAHE) for real-time image enhancement. J. VLSI Signal Process. Syst. Signal Image Video Technol. 38, 35–44 (2004).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man. Cybern. 9, 62–66 (1979).
Chibante, R. Simulated Annealing: Theory with Applications. (2010).
Jennings, B. H. Drosophila – a versatile model in biology & medicine. Mater. Today 14, 190–195 (2011).
MacCallum, R. C., Zhang, S., Preacher, K. J. & Rucker, D. D. On the practice of dichotomization of quantitative variables. Psychol. Methods 7, 19 (2002).
Lecuit, T. & Lenne, P.-F. Cell surface mechanics and the control of cell shape, tissue patterns and morphogenesis. Nat. Rev. Mol. Cell Biol. 8, 633–644 (2007).
Ochoa-Espinosa, A. et al. The role of binding site cluster strength in Bicoid-dependent patterning in Drosophila. Proc. Natl. Acad. Sci. USA 102, 4960–4965 (2005).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Healy, J. & McInnes, L. Uniform manifold approximation and projection. Nat. Rev. Methods Prim. 4, 82 (2024).
Zeira, R., Land, M., Strzalkowski, A. & Raphael, B. J. Alignment and integration of spatial transcriptomics data. Nat. Methods 19, 567–575 (2022).
Wang, M. et al. A single-cell 3D spatiotemporal multi-omics atlas from Drosophila embryogenesis to metamorphosis. Preprint at https://doi.org/10.1101/2024.02.06.577903 (2024).
Rossant, J. & Tam, P. P. L. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Behringer, R. Gertsenstein, M. Nagy, K. V. & Nagy, A. Manipulating the Mouse Embryo: A Laboratory Manual. (2014).
Hall, B. K. & Newman, S. A. Cartilage Molecular Aspects. (1991).
Noctor, S. C. et al. Dividing precursor cells of the embryonic cortical ventricular zone have morphological and molecular characteristics of radial glia. J. Neurosci. 22, 3161–3173 (2002).
Hartfuss, E., Galli, R., Heins, N. & Götz, M. Characterization of CNS precursor subtypes and radial glia. Dev. Biol. 229, 15–30 (2001).
Aurenhammer, F. Voronoi diagrams—a survey of a fundamental geometric data structure. ACM Comput. Surv. 23, 345–405 (1991).
Qiu, C. et al. A single-cell time-lapse of mouse prenatal development from gastrula to birth. Nature 626, 1084–1093 (2024).
Ocqueteau, C. et al. Three-dimensional morphometry of mammalian cells. I. Diameters. Arch. Biol. Med. Exp. 22, 89–95 (1989).
Pillarisetti, A., Ladjal, H., Ferreira, A., Keefer, C. & Desai, J. P. Mechanical characterization of mouse embryonic stem cells. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2009, 1176–1179 (2009).
Pera, M. F. & Tam, P. P. L. Extrinsic regulation of pluripotent stem cells. Nature 465, 713–720 (2010).
Puchtler, H. & Meloan, S. N. On the chemistry of formaldehyde fixation and its effects on immunohistochemical reactions. Histochemistry 82, 201–204 (1985).
Darzynkiewicz, Z., Halicka, H. D. & Zhao, H. Analysis of cellular DNA content by flow and laser scanning cytometry. Adv. Exp. Med. Biol. 676, 137–147 (2010).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Ronneberger, O., Fischer, P. & Brox, T. in Medical Image Computing and Computer-Assisted Intervention–MICCAI (2015).
Bentley, J. L. Multidimensional binary search trees used for associative searching. Commun. ACM 18, 509–517 (1975).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Qiu, X. et al. Mapping transcriptomic vector fields of single cells. Cell 185, 690–711 (2022).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Kornbrot, D. Point Biserial Correlation. https://doi.org/10.1002/9781118445112.stat06227 (2014).
Linacre, J. M. & Rasch, G. The expected value of a point-biserial (or similar) correlation. Rasch Meas. Trans. 22, 1154 (2008).
Gene, V. Glass & Kenneth, D. Hopkins, Statistical Methods in Education and Psychology. (1995).
Li, H. Leihouyeung/STP v0.0.1 https://doi.org/10.5281/zenodo.15202403 (2025).
Acknowledgements
H.L. and X.G. were supported by the King Abdullah University of Science and Technology (KAUST) Office of Research Administration (ORA) under Award No REI/1/5234-01-01, REI/1/5414-01-01, REI/1/5289-01-01, REI/1/5404-01-01, REI/1/5992-01-01, URF/1/4663-01-01, Center of Excellence for Smart Health (KCSH), under award number 5932, and Center of Excellence on Generative AI, under award number 5940. Q.H. and Y.H. were supported by the Shenzhen Science and Technology Innovation Program (Grant No. KQTD20180411143432337, China) and Shenzhen Key Laboratory of Gene Regulation and Systems Biology (Grant No. ZDSYS20200811144002008) (to Y.H. and Q.H.), Shenzhen Medical Research Fund (Grant No. D2401016 to Y.H.), Guangdong Basic and Applied Basic Research Foundation (Grant No. 2024A1515012343 to Q.H.), and the National Natural Science Foundation of China (Grant No. 32100684 to Q.H.). Computational resources and experimental facilities were supported by the Center for Computational Science and Engineering and the Core Research Facilities at Southern University of Science and Technology. Z.Q. was supported by Heilongjiang Provincial Key Research and Development Plan 2023ZX02C10, 2023ZXJ02C03, and Jiangsu Provincial Key Research and Development Plan BE2023081. H.X. was supported in part by the National Key R&D Program of China (Grant No.2023YFF0725001), in part by the National Natural Science Foundation of China (Grant No.92370204), in part by the guangdong Basic and Applied Basic Research Foundation (Grant No.2023B1515120057) in part by Guangzhou-HKUST(GZ) Joint Funding Program (Grant No.2023A03J0008), Education Bureau of Guangzhou Municipality.
Author information
Authors and Affiliations
Contributions
Y.H., X.G., H.L., and Q.H. conceived and initiated this study. H.L. and Q.H. designed the methodology. H.L. and Q.H. conducted all the experiments. H.L. outputted the figure and tables. H.L. wrote the manuscript under the supervision of X.G. and Y.H. X.H. and Q.W. polished the writing of manuscript. All authors are involved in the discussion and finalization of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Honda Naoki, Young Tae Kim, Daniel Tward, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, H., Hu, Q., Qiu, Z. et al. STP: single-cell partition for subcellular spatially-resolved transcriptomics. Nat Commun 16, 4665 (2025). https://doi.org/10.1038/s41467-025-59782-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59782-3