Abstract
The objective of optical super-resolution imaging is to acquire reliable sub-diffraction information on bioprocesses to facilitate scientific discovery. Structured illumination microscopy (SIM) is acknowledged as the optimal modality for live-cell super-resolution imaging. Although recent deep learning techniques have substantially advanced SIM, their transparency and reliability remain uncertain and under-explored, often resulting in unreliable results and biological misinterpretation. Here, we develop Bayesian deep learning (BayesDL) for SIM, which enhances the reconstruction of densely labeled structures while enabling the quantification of super-resolution uncertainty. With the uncertainty, BayesDL-SIM achieves high-fidelity distribution-informed SIM imaging, allowing for the communication of credibility estimates to users regarding the model outcomes. We also demonstrate that BayesDL-SIM boosts SIM reliability by identifying and preventing erroneous generalizations in various model misuse scenarios. Moreover, the BayesDL uncertainty shows versatile utilities for daily super-resolution imaging, such as error estimation, data acquisition evaluation, etc. Furthermore, we demonstrate the effectiveness and superiority of BayesDL-SIM in live-cell imaging, which reliably reveals F-actin dynamics and the reorganization of the cell cytoskeleton. This work lays the foundation for the reliable implementation of deep learning-based SIM methods in practical applications.
Similar content being viewed by others
Introduction
Recent years have witnessed a vigorous development of super-resolution (SR) fluorescence microscopy, which can surpass the diffraction limit1 and enable access to a wide variety of bioprocesses at unprecedented high resolution2,3,4. Among various SR modalities, structured illumination microscopy (SIM) is recognized as the most promising one for live-cell imaging owing to its fast imaging speed, low phototoxicity, high photon efficiency, and excellent compatibility with fluorophores5,6,7. SIM typically applies a series of spatially patterned illuminations with different phase shifts and orientations on specimens, to render the normally unresolvable high-frequency information encoded into the passband of the microscopic optical transfer function. Apart from the hardware part, as a computational SR technique, SIM necessitates reconstruction algorithms to extract SR information from raw images and then produce a super-resolved SIM (SR-SIM) image. Traditional reconstruction methods5,8,9,10,11 are based on complex frequency-domain workflow or employ hand-crafted analytical models, often prone to artifacts or yielding a compromise between signal preservation and noise suppression (Supplementary Fig. 1). Deep learning-based SIM (DL-SIM) methods6,12,13,14,15, which leverage deep neural networks (DNNs) to capture statistical knowledge from paired raw and SIM data in a data-driven manner, have exhibited impressive advantages over traditional methods, such as improved fidelity and superior noise immunity (Supplementary Fig. 2a, b), thereby becoming a prevailing gold standard in SIM computational imaging.
Despite the prevalence, current DL-SIM methods still suffer from two practical challenges. First, SIM reconstruction in the presence of noise is inherently an ill-posed inverse problem6, suggesting that many latent SR solutions exist for any given set of raw images (Supplementary Note 1a). However, regardless of the input images’ SNR and applicability to the reconstruction model, current DL-SIM methods only output one reconstruction without considering its ill-posedness or the correctness of the SR information (Supplementary Note 1c). Second, the biological specimens/structures are so diverse that even experienced users can hardly tell whether the input images are complied with the knowledge consistency16 (KC) relative to the trained DL-SIM model, hence assessing and quantifying the KC specification is critical for the reliable adoption of DL-SIM models, but it has not been done so far. Note that the model knowledge should encompass all aspects of imaging statistics, and knowledge inconsistency (KIC) in any aspect would cause severe performance drops and notorious artifacts (Supplementary Fig. 2c). Nonetheless, since the model knowledge is implicitly encoded in the pre-trained weights and invisible to users, keeping the KC specification is not trivial, especially for biologists or novice users without much experience in deep learning.
In this work, we develop a Bayesian deep learning (BayesDL) framework for SIM. BayesDL-SIM not only improves the fidelity of intricate structures but also enables the quantification of different types of uncertainty17,18,19 in SIM reconstruction, including aleatoric uncertainty20,21 (AleaU) and epistemic uncertainty22,23 (EpisU), where AleaU characterizes imaging ill-posedness due to SR information incompleteness in raw data while EpisU highlights the knowledge deficiency of the reconstruction model during its generalization (Supplementary Notes 2 and 3). Specifically, by modeling the latent SR manifold as heteroscedastic probability distributions, BayesDL-SIM quantifies AleaU and achieves precise distribution-informed SR imaging for the first time. Moreover, instead of utilizing a deterministic model, we perform Bayesian posterior inference within a stochastic Bayesian neural network (BNN) for EpisU quantification. With the EpisU conferred by BayesDL-SIM, it becomes feasible to identify various KIC-induced erroneous generalizations, thereby helping prevent unreliable imaging and subsequent biological misinterpretation. Additionally, the BayesDL uncertainty also shows versatile utilities as a valuable tool in routine SR imaging, e.g., SR defects estimation, data acquisition evaluation, etc. Furthermore, we demonstrate the superiority of BayesDL-SIM in live-cell imaging experiments, suggesting that BayesDL-SIM enables high-fidelity, reliable, long-term visualization of actin dynamics in live cells.
Results
Development of BayesDL-SIM
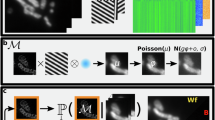
Unlike current DL-SIM methods that aim to learn a one-to-one mapping from raw images to high-quality ground truth SIM (GT-SIM) images with deterministic neural networks (Supplementary Note 1b), we develop BayesDL for SIM, a Bayesian framework capable of inferring SR structures with the highest accuracy while being able to reason about two types of uncertainty (i.e. AleaU and EpisU) in SIM imaging. Concretely, BayesDL-SIM formalizes the AleaU stemming from the SR ill-posedness as probability distributions over the latent SIM images \({{I}}_{SIM}\), while formalizing the EpisU as probability distributions over model parameters θ, given a network parameterized by θ. Then, BayesDL-SIM predicts the distributions of the latent SIM images as:
where \({{I}}_{raw}\) is the input raw images, D is the training dataset, the term \(p({{I}}_{SIM}|{{I}}_{raw},{{\boldsymbol{\theta }}})\) is the likelihood representing a probabilistic function that covers all potentially valid SIM solutions in the SR manifold given input raw images, and the term \(p({{\boldsymbol{\theta }}}|D)\) is the posterior distribution describing all possible model parameterization that can explain the observed training dataset D.
To quantify AleaU, we characterize the likelihood by modeling the latent SR manifold as a heteroscedastic Gaussian distribution, of which both mean and standard deviation (STD) are spatially variant. The BayesDL network is devised to estimate the Gaussian mean and STD simultaneously (Fig. 1a). Note that the estimated mean serves the SR-SIM result while the STD indicates the AleaU. Using maximum likelihood estimation, a heteroscedastic loss function is derived (Supplementary Note 4b):
where N is the number of data items, \({{I}}_{GT}\) is the GT-SIM image, \({{\mu }}_{{{\boldsymbol{\theta }}}}\) and \({{\sigma }}_{{{\boldsymbol{\theta }}}}\) respectively denote the estimated Gaussian mean and STD, whose dependencies on parameters are highlighted via the subscript θ. The BayesDL network can be trained to minimize the heteroscedastic loss in an end-to-end manner with paired raw and GT-SIM images. Admittedly, the heteroscedastic loss allows for AleaU quantification alongside the SIM reconstruction. However, we find that it compromises reconstruction fidelity (Supplementary Note 5). To avoid the adverse impact of heteroscedastic loss on reconstruction quality, we propose a decoupling training (DeT) scheme, which consists of two steps: (1) exert a dual-domain loss24 to guide optimization of the parameters related to SIM reconstruction; (2) freeze the parameters that have been trained in the first step and apply the heteroscedastic loss for AleaU learning (Supplementary Note 4e).
a, The BayesDL network architecture and the schematic of its decoupling training (DeT) scheme. b, Statistical comparisons of DFCAN, RCAN, and BayesDL in terms of reconstruction PSNR (top) and imaging resolution (bottom) on F-actin data, n = 30 cell images. c, Representative single raw SIM images (top left, first column) and corresponding SR images reconstructed by conventional (Conv.) SIM (bottom right, first column), DFCAN (third column), RCAN (fourth column), and BayesDL (fifth column) in the cases of CCPs (first row), MTs (second row), and F-actin (third row). GT-SIM images acquired at ultra-high signal-to-noise ratio (SNR) levels are shown for reference (second column). d, Corresponding Gaussian STD (i.e. AleaU maps) of the three subcellular structures shown in (b), quantified by BayesDL-SIM. The smaller the AleaU, the better. e, Corresponding credibility (cred.) maps of the three subcellular structures with error rate tolerance (δ) of 0.2. The greater the cred., the better. f, Calibration (calib.) diagrams of BayesDL-SIM on CCPs, MTs, and F-actin images, where the calib. errors are also presented, n = 60. g, Intensity profiles of DFCAN (blue), RCAN (green), BayesDL (magenta dotted), and GT-SIM (black dashed) along the lines indicated by the two arrowheads in (b). A credible interval (CI) of three predicted Gaussian STDs centered on BayesDL intensity profile is also shown (light blue shadow). Center line, medians; limits, 75%, and 25%; whiskers, maximum, and minimum. Scale bar, 500 nm (b,d,e). Gamma value, 0.8 for F-actin images in (b). AU, arbitrary units.
To quantify EpisU, we capture the posterior distribution by converting our BayesDL network to a BNN and then performing Bayesian posterior inference (Fig. 2a). Nonetheless, Bayesian posterior inference is generally intractable for DNNs (Supplementary Note 6). Here we adopt an approximate Bayesian inference method called stochastic gradient Langevin dynamics25 (SGLD), whose basic strategy is to inject noise into parameter updates during training, and in such a way the trajectory of model parameters would converge to the true posterior distribution (Supplementary Note 4c).
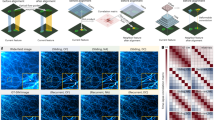
a, Schematic illustration of the EpisU quantification pipeline in BayesDL-SIM. b, Representative SR images of F-actin reconstructed by BayesDL-SIM models that are trained with data of MTs (top left), CCPs (bottom right), and F-actin (middle), respectively. c, Corresponding SIM EpisU of the results in (b). The smaller the EpisU, the better. d, Enlargement of the boxed insets in (b) and (c). Corresponding raw and GT-SIM images are also presented for reference. e-f, Statistical comparison in terms of the reconstruction PSNR metric (e) and FG-EpisU distribution (f) among the three BayesDL-SIM models trained with different subcellular structures, n = 60 cell images. g-h, Representative SR images of fluorescent beads (g) and corresponding EpisU (h) produced by BayesDL-SIM models trained with sparsely (left) and normally (right) distributed fluorescent beads. Experiments were repeated with 150 images, achieving similar results. i, Enlargement of the two boxed insets in (g) and (h). Corresponding raw and GT-SIM images are also presented for reference. Intensity profiles along the lines indicated by two arrowheads are shown at the bottom right of each imaging result. Scale bar, 3 μm (b,c), 1 μm (d,g,h), 200 nm (i).
In the inference phase, we also follow Eq. (1) and use the Monte Carlo (MC) technique to approximate the integral. By sampling θ from the SGLD-inferred posterior, multiple inferences (termed MC samples) can be generated. The predictive mean serves as the final SR-SIM result and is given by:
where\({\mathbb{E}}(\cdot )\) denotes taking the expectation, K denotes the number of MC samples. The two types of uncertainty of the SR-SIM result can also be measured statistically by:
We use constant five MC samples in this study for a best performance-complexity trade-off (Supplementary Fig. 6). We refer the interested reader to Supplementary Note 4 for full details about the development and implementation of BayesDL-SIM.
BayesDL-SIM enables accurate distribution-informed SR imaging
First, we test the reconstruction performance of BayesDL against other state-of-the-art SIM methods on various subcellular structures labeled by fluorescent proteins, including clathrin-coated pits (CCPs), microtubules (MTs), and F-actin. We observe that BayesDL-SIM reconstructs intricate or faint structures more precisely than DFCAN14 and RCAN26, while preventing the generation of undesirable artifacts compared with the conventional SIM (Fig. 1c and Supplementary Movies 1–3). We also demonstrate the superiority of BayesDL-SIM in dealing with noisy raw data, where Hessian-SIM10 features excessive reconstruction artifacts, DFCAN and RCAN cause some detailed structure disarrangement or missing, while BayesDL-SIM exhibits superior noise resilience and still yields faithful SR-SIM results (Supplementary Fig. 7a and Supplementary Movies 1–3). Line-scan profiles along F-actins also indicate the better agreement of BayesDL-SIM results with respect to the GT-SIM ones (Fig. 1g and Supplementary Fig. 7e). Moreover, to further quantitatively evaluate the DL-SIM methods, we calculate two metrics, i.e. peak signal-to-noise ratio (PSNR) and image resolution, where the latter is measured by decorrelation algorithm27 (Supplementary Fig. 8), implying that BayesDL-SIM offers optimal reconstruction fidelity and resolution than alternatives (Fig. 1b and Supplementary Fig. 7d). More comparisons with other advanced SIM methods can refer to Supplementary Fig. 9. More importantly, in addition to being able to precisely predict the SR-SIM intensity (i.e. the mean of the underlying SR distributions), BayesDL-SIM also quantifies pixel-wise AleaU for its reconstructed results (i.e. the STD of the underlying SR distributions) as shown in Fig. 1d and Supplementary Fig. 7b, thereby capturing the entire SR manifold that covers all possible SIM solutions. To validate the reliability, model calibration28,29 (Methods) is performed on each of the three subcellular structures, suggesting that the empirical frequency of the GT-SIM intensities matches the predicted Gaussian distributions with flying colors (Fig. 1f and Supplementary Movies 1–3). That is, BayesDL-SIM precisely captures the underlying SR distributions that, to the best of our knowledge, have not been achieved by previous SIM methods. A related work is CARE30, which also allows for the estimation of SR distributions with a Laplacian likelihood. We comprehensively compare BayesDL-SIM side-by-side with CARE-SIM and demonstrate that BayesDL-SIM outperforms CARE-SIM, highlighted by significantly enhanced fidelity in crisscrossing filaments and ~4-fold lower calibration error (Supplementary Fig. 11).
With the distribution-informed SR imaging ability of BayesDL-SIM, the credibility of SIM reconstruction results can be evaluated. Intuitively, pixels with smaller AleaU seem to be more credible. However, we note that the AleaU (i.e. STD) is proportional to the fluorescence intensity, posing challenges in assessing the credibility of regions with varying fluorescence intensities. Here we devise a method for credibility analysis. Specifying an error rate tolerance (δ) beyond which results are deemed to be incredible, we calculate the cumulative probability of all possible intensities within the specified tolerance as the credibility, which ranges from 0 to 1 (Methods and Supplementary Fig. 10). Not surprisingly, non-feature regions in the CCPs images exhibit high credibility since these regions are flat and easily reconstructed, while for MTs and F-actins, the in-focus structures tend to be more credible than the regions with out-of-focus fluorescence background (Fig. 1e and Supplementary Fig. 7c). Users can also get desired credibility maps of other customized tolerance δ (Supplementary Fig. 10). Another valuable aspect of distribution-informed imaging is the ability to provide credible intervals (CIs) for each pixel based on a user-specified credibility probability, instead of a single intensity value (Methods). For instance, given the 3-σ probability of Gaussian distribution (0.9974), the corresponding 3-σ CI can be obtained (light blue shadow, Fig. 1g and Supplementary Fig. 7e), which signifies the probability that the true fluorescence intensity falls within the 3-σ CI is constant 0.9974. Similarly, users can customize the CIs of other credibility probabilities in the same way.
We further isolate the positive effect of the proposed DeT scheme by implementing BayesDL-SIM with (w/) and without (w/o) the DeT scheme, respectively. One can witness that BayesDL-SIM w/o DeT tends to produce over-smoothed results and concurrently the quantified AleaU is moderately calibrated, while the integration of DeT scheme can avoid the performance degradation caused by AleaU learning, boost SIM fidelity, and bring a calibration error decrease of more than 3-fold (Supplementary Fig. 12a–f). Moreover, we corroborate that the DeT scheme can also benefit the probabilistic CARE-SIM model (Supplementary Fig. 12g–j), indicating it is a general-purpose method.
BayesDL-SIM warns unreliable SR imaging due to erroneous generalization
A common pitfall of DL-SIM models in practice is that they can generalize well to unseen data only when complying with the KC specification. However, due to the inaccessibility of large-scale SIM data, DL-SIM models generally extract exclusive knowledge of the specific imaging setting rather than the generic one. The immense variability inherent in biological specimens (e.g. different tissues, cell types) and optical systems (e.g. different microscopy modalities, imaging conditions) exacerbates the risk of SIM model misuse, which violates the KC specification, causes erroneous generalization, and produces unreliable SR results. On the other hand, DL-SIM models are black boxes and their erroneous generalization can usually only be assessed in hindsight with GT-SIM images, which are often unavailable in practice31,32. Hence, it is quite necessary to develop an ex-ante evaluation method to help users identify KIC scenarios, thereby preventing erroneous generalization and promoting SR reliability. In this section, we show that the BayesDL-SIM provides a promising solution to this challenge by quantifying EpisU.
We first examine whether the deep SIM models trained with different subcellular structures could share their knowledge. To handle raw images of F-actin, in contrast to the faithful reconstruction provided by the BayesDL-SIM model trained with F-actin data, models trained with CCPs or MTs are incompetent in resolving F-actin filaments, where plenty of weak signals are removed and some strong actin filaments become discontinuous (Fig. 2b, d). Quantitative measures coincide with our visual observations (Fig. 2e). Such problematic images result from the KIC issue of different subcellular structures and may be imperceptible in daily imaging experiments, especially for biologists who are inexperienced with deep learning. Fortunately, we find that the BayesDL-SIM can be a feasible tool in identifying these KIC cases, by marking the foreground (FG) pixels of the unreliable SR-SIM results with much higher EpisU ( ~400% average increase) when the subcellular structures in training and inference mismatch (Fig. 2c, d, f), thereby raising warnings to users to be careful with the imaging results.
Then, we test the BayesDL-SIM on fluorescent beads to identify the KIC issue arising from the density variation. Green FluoSpheres (Thermofisher, F8803, 100-nm diameter) were dissolved in double-distilled water at 1:1000 and 1:10,000 dilution to produce fluorescence beads of different densities. We programmatically accumulate two datasets, one containing normally distributed beads (density ranges from 0 μm−2 to 4 μm−2) and the other consisting of sparse beads (density ranges from 0 μm−2 to 0.5 μm−2). Two BayesDL networks are separately trained on these two datasets with the same procedure (termed normal- and sparse-density model, respectively). In the inference phase, we notice that the normal-density model produces high-quality reconstruction for both sparse and dense beads as it has seen beads of various densities during training, while the sparse-density model which has only extracted knowledge about sparse beads excels in dealing with sparse beads but tends to over-separate beads that are spaced closely (Fig. 2g, i). It suggests that the KIC of fluorescent bead density can appear at the image patch level and then locally impair model generalization. More importantly, BayesDL-SIM can sensitively detect the local density KIC, labeling the faithfully reconstructed beads with negligible EpisU while labeling the over-separated ones with the 3-4-fold larger EpisU (Fig. 2h, i).
The spatial sampling rate is another vital attribute with considerable variability in optical imaging systems. Even in the same microscope, the spatial sampling rate can vary substantially with different configurations of magnification factors and numerical aperture (NA) of the objective lens. In DL-SIM imaging, on the other hand, the spatial sampling rate also affects the feature representation in neural networks, making it an essential aspect of model knowledge. Thus, KC/KIC issues of spatial sampling rate deserve to be explored. In this work, specimens are captured and digitized by our multimodal SIM system (Methods) with a sampling rate of 31.2 nm per pixel. For simplicity, we simulate imaging of other sampling rates via 2× up- and down-sampling, corresponding to sampling rates of 15.6 and 62.4 nm per pixel, respectively. When applying the BayesDL-SIM model trained on original data (with a sampling rate of 31.2 nm/pixel) to perform reconstruction for data of other sampling rates, severe blur or ghosting artifacts manifest in the reconstructed SR-SIM images, accompanied by the significant deterioration of reconstruction fidelity (Supplementary Fig. 14a, c, d). It indicates that the spatial sampling rate is also an exclusive knowledge of the reconstruction model. Similarly, we observe an obvious increase (more than 250%) of FG-EpisU in the two KIC cases (Supplementary Fig. 14b, c, e), which effectively helps pinpoint the unreliable structures super-resolved by the SIM model that lacks adequate sampling rate knowledge about target data. Besides, we also corroborate that transfer learning33 is an effective approach to mitigate the KIC issue, which can help narrow the knowledge gap between the model and data by fine-tuning the model weights on a small amount of target data (Supplementary Fig. 14f–i).
Additionally, we also conduct other experiments to verify the effectiveness of BayesDL-SIM in identifying common KIC problems caused by SNR variation and engineered perturbation. Interested readers can refer to Supplementary Notes 7 and 8 for more description. These results established the usability and superiority of our BayesDL-SIM, which offers a feasible solution to prevent the KIC-induced erroneous generalization of DL-SIM models and the consequent unreliable SR imaging results.
BayesDL acts as a versatile tool for routine SR imaging
In this section, we showcase the versatility potential of BayesDL, which positions it as a promising tool for routine SR imaging experiments. To begin, we illustrate that the uncertainty quantified by BayesDL provides valuable assessments of SR imaging errors. Owing to the finite model capacity, SR imaging errors are inevitable and hard to assess in practice without the GT images as the gold standard. While methods like SQUIRREL34 have been developed to estimate SR errors using only a diffraction-limited (e.g., wide-field) image, we find that the error estimates of SQUIRREL are distributed differently from the true reconstruction errors (Fig. 3a–c). By contrast, we note that the two types of BayesDL uncertainty better correlate with the true SR errors, where high uncertainty values tend to align with super-resolved structures displaying conspicuous errors. To delve further, we conduct a quantitative analysis of the relationship between SIM reconstruction errors and BayesDL uncertainty through sparsification plots35,36 (Methods). In all three subcellular structures, the sparsification plots of BayesDL uncertainty exhibit impressive concordance beyond SQUIRREL with those of the true SR errors (Fig. 3d–f). These observations demonstrate the potential utility of BayesDL in providing surrogate measures of SR imaging errors in the absence of GT-SIM. Besides, it is worth mentioning that AleaU more accurately reflects the SR error distributions compared with EpisU (Fig. 3d–f).
a–c Top: Representative single raw SIM images (top left, first column), corresponding GT-SIM images (bottom right, first column), and SR-SIM images reconstructed by BayesDL-SIM (second column). Absolute SR errors between SR-SIM and GT-SIM images are also shown (third column); Bottom: Reconstruction AleaU (first column) and EpisU (second column) quantified by BayesDL-SIM and SR error estimates of SQUIRREL (third column); in the cases of CCPs (a), MTs (b), and F-actin (c). d–f Sparsification plots with the sorting criterion of true SR errors, AleaU, EpisU, and the SQUIRREL errors, in the case of CCPs (d), MTs (e), and F-actin (f). Scale bar: 1 μm (a, b, c). Gamma value, 0.7 for F-actin images in c. Experiments were repeated independently for more than 45 cell images, all showing similar results.
Considering that AleaU characterizes the SR information quality in the raw data, BayesDL can be utilized to improve data acquisition in the SR imaging pipeline. For instance, raw images polluted by more serious noise lead to SR-SIM results of lower reconstruction fidelity, meanwhile exacerbating the ambiguity of SR information retrieval and causing increased AleaU (Supplementary Fig. 18). As such, users can optimize their imaging parameters toward the objective of acquiring raw data that yields AleaU as small as possible. In addition, the BayesDL framework can be extended to encompass other SR imaging approaches. We employ the BayesDL for SIM with fewer raw images and single-image SR imaging (SISR) with a diffraction-limited wide-field image (Supplementary Note 4d). Analogously, AleaU quantified by BayesDL can evaluate the physical SR information conveyed by different SR approaches. The SR approach conveying more SR information shows greater capability in resolving densely labeled samples, concomitantly producing smaller AleaU (Supplementary Fig. 19). This effect is most prominent in the case of SISR imaging, as SR information does not physically exist in the wide-field data and can only be generated by pure model guess.
The generative adversarial network37 (GAN) is another popular neural paradigm in fluorescence imaging, which has shown powerful capacity in previous works38,39,40. BayesDL is also applicable to GAN-based SR imaging, by introducing an auxiliary discriminator to provide an additional adversarial loss for the reconstruction model (Supplementary Note 4d, e). While the GAN model enhances the restoration of dense actin meshes and infers high frequencies within wider support in the Fourier domain, the SR fidelity suffers degradation ( ~40% average increase in reconstruction errors), as seen in Supplementary Fig. 20a–d. A more serious issue of GAN-based imaging is that of hallucination, as its explicit training objective is to generate adequate and persuasive details, even under imaging conditions that lack sufficient SR information. The hallucination problem is often too plausible and subtle to detect. BayesDL provides an effective method for hallucination detection by quantifying the EpisU of the GAN model. It can be seen that the unreliable details forged by the GAN are generally marked with ~300-400% larger FG-EpisU, especially under low SNR conditions (Supplementary Fig. 20a, e). Based on the observations, we feel that GANs’ astonishing ability for high-frequency enhancement partially derives from the generation of unreliable hallucinations. When employing GAN-based models to characterize novel cellular processes, users should be cautious about the SR structures with salient EpisU.
BayesDL-SIM enables reliable visualization of dense F-actin in live cells
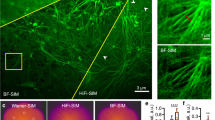
With the exceptional SR performance of BayesDL-SIM, we apply it on probing the dynamic bioprocesses in live cell specimens. Cell adhesion, a fundamental biological process of cell attachment and spreading on a substrate, has been proven to be susceptible to high excitation power adopted in previous SR microscopy41. Here, following the placement of live cells expressing mEmerald-Lifeact on a coverslip, our BayesDL-SIM accomplishes long-term tracking of cell adhesion lasting three hours under low-light imaging conditions, where reorganization of the cell cytoskeleton for cell shape adjustment is clearly observed (Fig. 4a and Supplementary Movie 4). Compared with previous SIM methods including Hifi-SIM5 and DFCAN, BayesDL-SIM improves the reconstruction fidelity of dense actin filaments and enables high-fidelity, long-term, distribution-informed live-cell SR imaging, allowing for the provision of reconstruction credibility maps and intensity CIs (Fig. 4b). Moreover, we also test the live-cell SR imaging in three common KIC scenarios, indicating that BayesDL-SIM can identify KIC-induced unreliable structures consistently over the entire imaging duration by yielding 1.5-2.5-fold larger FG-EpisU (Fig. 4c, d and Supplementary Movie 4).
a Time-lapse BayesDL-SIM SR imaging reveals F-actin dynamics over a three-h cell adhesion process after depositing a COS-7 cell onto a glass coverslip. b Representative SR images of F-actin reconstructed by Hifi-SIM (first column), DFCAN (second column), and BayesDL-SIM (third column) at two different time points. BayesDL AleaU (fourth column) and credibility (fifth column) maps are also shown. Intensity profiles along the lines in DFCAN and BayesDL-SIM reconstructions are presented at the far right, where BayesDL-SIM provides not only intensity values but also 3-σ CI centered on intensity values. c Comparison of SR-SIM images and corresponding EpisU predicted by BayesDL-SIM in the cases of KC (first column), KIC of subcellular structure (second column), KIC of sampling rate (third column), and KIC of fluorescence SNR (fourth column). d The time courses of the FG-EpisU over the three-h imaging duration in the KC and three different KIC scenarios. Scale bar, 8 μm (a), 1 μm (b, c). Gamma value, 0.8 for F-actin images in (b) and (c). Experiments were repeated independently for n = 3 COS-7 cells, all showing similar results.
In live-cell SR imaging, the tradeoff between fluorescence SNR and imaging duration manifests. Maintaining a decent SNR level may cause rapid bleaching while lowering the SNR level may render raw data unusable. To address this tradeoff, we employ BayesDL-SIM for live-cell SIM imaging at extreme low-SNR conditions. Compared with existing Sparse-SIM11 and scUNet-SIM13, BayesDL-SIM shows greater capability in resolving fine details from noisy raw data (Supplementary Fig. 21a and Supplementary Movie 5). More notably, it enables reliable visualization of actin dynamics up to 10,000 SR frames with no apparent fidelity drop (Supplementary Fig. 21b, c and Supplementary Movie 5) and a consistent increase in the average length of reconstructed actin filaments (Supplementary Fig. 21e and Supplementary Movie 5). Besides, we noted that the fluorescence SNR measured by the average photon count14 declines gradually over the course of imaging, and consequently the FG-AleaU of the reconstruction results increases accordingly (Supplementary Fig. 21d, f). Overall, these data illustrate the reliability and superiority of BayesDL-SIM for live-cell SR imaging.
Discussion
The highest priority for scientific SR imaging is reliability. However, previous SR techniques are generally over-confident and assume their results to be reliable, which is not always the case. For instance, conventional model-based SR methods are prone to fixed-pattern artifacts and necessitate meticulous parameter fine-tuning for the most visually pleasant results; current DL-based methods perform end-to-end image transformations blindly without explicit output to inform users of the degree of model confidence. To address the reliability imperative in SR microscopy, in this study, we develop the BayesDL framework for SIM, which combines neural networks and Bayesian learning, aiming to graft the former’s scalability and expressiveness with the latter’s capacity in uncertainty quantification. We emphasize that the implementation of BayesDL-SIM does not impose an additional burden on data acquisition and preparation compared with existing supervised DL-SIM methods. It is also noteworthy that the proposed BayesDL framework has compatibility and extensibility to other network architectures. Extensive experiments on both fixed and live cells demonstrate that the proposed BayesDL-SIM can substantially enhance SIM fidelity beyond state-of-the-art SIM methods even under extremely low-SNR conditions while quantifying two types of well-calibrated uncertainty of its reconstructions, i.e. AleaU and EpisU, thereby promoting the transparency of deep SIM models and helping prevent unreliable SR imaging results.
The AleaU and EpisU quantified by BayesDL convey different information and thus serve different purposes. In routine bioimaging experiments, users can first examine the EpisU map to determine if the model generalization is performed properly in a KC scenario. We have substantiated the consistent effectiveness of BayesDL EpisU in identifying unreliable SR results due to erroneous generalization in diverse KIC cases. Upon successful examination of EpisU, users can resort to AleaU to access the predictive SR distribution rather than a single reconstruction given by current DL-SIM methods. Based on the distribution-informed SR imaging, credibility and intensity CIs can also be customized in combination with statistical analysis. Moreover, the two types of BayesDL uncertainty exhibit additional versatile utilities for routine SR imaging. For instance, both AleaU and EpisU serve as superior surrogate measures for SR imaging errors compared with SQUIRREL; AleaU can be used for data acquisition evaluation thereby guiding the optimization of imaging parameters; EpisU can be used for GAN-based imaging to detect unreliable hallucinations. These findings establish the potential of BayesDL uncertainty for integration into daily SR imaging experiments.
Despite the impressive performance of BayesDL-SIM, its further improvement can be envisaged. A recently published work42 introduces uncertainty into the model training process to regulate pixel-wise attention in the loss function. Inspired by this, we posit that the uncertainty quantified by BayesDL-SIM can be leveraged in turn to further boost SIM reconstruction. Additionally, we have demonstrated the superiority of BayesDL-SIM in 3D-SIM imaging. Under high SNR conditions, BayesDL-SIM outperforms other DL-SIM methods while remaining comparable to conventional Open-3DSIM43 (Supplementary Fig. 22). Even under low SNR conditions, BayesDL-SIM still produces decent SR-SIM results without discernible artifacts (Supplementary Fig. 23). Its applicability for uncertainty quantification in 3D-SIM reconstruction is also validated (Supplementary Fig. 22 and 23). Given the demonstrated effectiveness of BayesDL in SISR and SIM imaging, it is reasonably speculated that its positive impact can extend to other SR microscope types (for example, localization microscopy44, and light-sheet microscopy45). Moreover, to combat the challenge that sufficient high-quality GT data is too laborious or even impractical to acquire in some demanding conditions (for example, rapid-moving and light-sensitive bioprocesses), the integration of unsupervised46,47 or zero-shot48,49 learning techniques with BayesDL should also be valuable.
Moreover, we would like to clarify the difference between BayesDL uncertainty and the similar concepts proposed in NanoJ34 and rFRC50. First, both NanoJ and rFRC are tools for assessing the quality of SR images, while BayesDL aims to quantify the uncertainty in DL-based image reconstruction. Second, NanoJ identifies artifacts by detecting the discrepancy between SR images and diffraction-limited ones, and therefore incapable at the SR scale. The rFRC simply trains two models and needs to sample input data twice for evaluating the uncertainty roughly, which in fact cannot provide the quantitative and wholistic uncertainty information. By contrast, BayesDL is developed on the foundation of Bayesian theory, quantifying AleaU and EpisU by distributional approximation and Bayesian inference. Third, the BayesDL uncertainty is demonstrated to be well-calibrated, which has not been achieved by the other two methods.
To conclude, the primary advantage of BayesDL over other DL-SIM methods lies in its ability to enable precise distribution-informed SR imaging in KC scenarios, while also alerting users to erroneous generalizations in KIC scenarios and thereby preventing users from trusting the unreliable outcomes. However, the BayesDL still has a limitation, i.e. it can only identify various KIC scenarios but cannot directly achieve effective generalization to the KIC data. To this end, we consider developing a universal algorithm capable of generalizing effectively in KIC scenarios as a promising avenue for future research. Overall, the proposed BayesDL-SIM facilitates live-cell SR imaging with sustained high fidelity and ensures the reliability of sub-diffraction information conveyed by the algorithmic back-end of modern smart microscopes. Thus, it represents a notable advancement in facilitating reliable employment of DL-SIM models and establishes a groundwork for the evolution of practical DL-based computational SR microscopy.
Methods
Multimodality SIM system
The multimodality SIM system was built based on an invented fluorescence microscope (Ti2E, Nikon). Excitation light from a laser combiner equipped with three laser beams of 488 nm (Genesis-MX-SLM, Coherent), 560 nm (2RU-VFL-P-500-560, MPB Communications) and 640 nm (LBX-640-500, Oxxius) was collimated and passed through an acousto-optic tunable filter (AOTF, AOTFnC-400.650, AA Quanta Tech), which can select the excitation wavelength and control its power and exposure duration flexibly according to imaging demands. Then the output laser light from AOTF was expanded and sent into an illumination pattern generator, which comprises a polarization beam splitter, an achromatic halfwave plate, and a ferroelectric spatial light modulator (SLM, QXGA-3DM, Forth Dimension Display). Different illumination modes could be generated by adjusting the period and orientation of the grating patterns displayed on the SLM, e.g. 1.41 NA TIRF-SIM and 1.35 NA GI-SIM of 3-phase × 3-orientation. To maximize the pattern contrast, a polarization rotator composed of a liquid crystal cell (Meadowlark, LRC200) and a quarter-wave plate was utilized to adjust linear polarization so as to maintain s-polarization.
The high diffraction orders except for ±1 orders were filtered out by a spatial mask. Next, the excitation light was relayed onto the back focal plane of the objectives (1.49 NA, Nikon). Multiple raw images excited by different illumination patterns were collected by the same objectives, separated by a dichroic beam splitter (Chroma, ZT405/488/560/647tpc), and finally captured by a scientific complementary metal oxide semiconductor (sCMOS) camera (Hamamatsu, Orca Flash 4.0 v3).
Cell culture and preparation
The COS-7 cells (in Fig. 4 and Supplementary Fig. 21) were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Gibco) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin and streptomycin at 37 °C and 5% CO2. The cells were infected by the retrovirus system according to the standard procedures (Lipofectamine 3000, Invitrogen) to stably express Lifeact-mEmerald. The transfected cells were seeded onto 50 mg/mL collagen-coated coverslips to achieve 50%-70% confluence prior to imaging.
Data acquisition and pre-processing
All data relevant to this work are acquired using the home-built multimodality SIM system. To validate the superiority of BayesDL-SIM, we utilize our earlier published dataset BioSR14. For each type of specimen, raw images of size 502 × 502 × 9 (height×width×frames) of at least 50 distinct regions of interest (ROIs) are collected. Each ROI is sequentially captured with a constant 1 ms exposure time but at least 9 different levels of excitation power to cover various fluorescence SNR levels (average photon count ranges from 0 to more than 600). The raw images of the highest SNR level are subsequently reconstructed via the conventional SIM algorithm to serve as the corresponding GT-SIM images of size 1004 × 1004. To facilitate model training, the data pre-processing step should be conducted. We first subtract an average camera background from all raw images. Then the rolling ball51 method is adopted to remove the out-of-focus fluorescence and improve contrast of GT-SIM images. Besides, to stretch fluorescence intensity to a common range, percentile-normalization30 is performed for both raw and GT-SIM images:
where \(perc(I,p)\) represents the p-th percentile of all pixel intensities of the image I. We set \({p}_{low}\) and \({p}_{high}\) as 0.01% and 99.9%, respectively.
Additionally, in long-term live-cell imaging of COS-7 cells stably expressing LifeAct-mEmerald (Fig. 4), we acquire the raw images of three-phase×three-orientation for 9720 frames (corresponding to 1080 SR frames) with a constant 1 ms exposure time. The time interval between two consecutive exposures is constant 10 seconds so the entire imaging duration lasts three h. In rapid live-cell imaging of COS-7 cells stably expressing LifeAct-mEmerald (Supplementary Fig. 21), we rapidly acquire raw images for 90,000 frames (corresponding to 10,000 SR frames) with a constant 1 ms exposure time in 260 seconds so that there is 26 ms between two adjacent time points.
Statistical analysis and evaluation methods
Calculation of intensity CIs
Based on the distribution-informed SR imaging ability of BayesDL-SIM, instead of a single intensity value, intensity CIs of different credibility levels can be provided. For raw images \({I}_{raw}\), a conditional heteroscedastic Gaussian distribution \(p({I}_{SIM}|{I}_{raw})\) is predicted by BayesDL-SIM with the mean μ and STD σ. Specifying a credibility level \(\kappa \,(0\le \kappa \le 1)\) which denotes the probability that the true fluorescence intensity falls in a certain interval centered on the predictive Gaussian mean, the corresponding intensity CI of credibility level \(\kappa\) can be calculated as:
where \({\varPhi }^{-1}(\cdot )\) is the quantile function of standard Gaussian distribution.
Credibility assessment
Another utility of distribution-informed SR imaging of BayesDL-SIM is that it enables credibility assessment for reconstructed SR-SIM images. AleaU is a direct measure of credibility but it grows with the intensity. Here we propose a credibility assessment method. First of all, users need to specify an error rate tolerance (δ) for the SR results beyond which the results are considered incredible. That is, reconstructions with the error rate below the tolerance are deemed to be usable to users. Then, the cumulative probability of all possible intensities within the specified tolerance is computed as a proper credibility measure:
Note that the calculated credibility is a pixel-wise estimation that ranges from 0 to 1, with a higher value indicating better credibility of the results.
Model calibration
The model calibration28,29 statistically evaluates the precision of the predictive SR distributions, by measuring the agreement between predicted probabilities and observed frequencies. Given any probability \(\kappa\), we first follow Eq. (6) to calculate its corresponding \(CI(\kappa )\). Then we count the frequency \(\kappa {\prime}\) that the fluorescence intensity of GT-SIM pixels falls within their corresponding \(CI(\kappa )\). By gradually adjusting the probability \(\kappa\) from 0 to 1, the correlation between probability and frequency can be fully depicted and subsequently constitutes the calibration diagram. Intuitively, more accurate predictions of SR distributions will lead to greater agreement with the ideal calibration line \(\kappa=\kappa {\prime}\) in the calibration diagram. Besides, we define the metric of calibration error that is computed as the mean absolute error (MAE) between probability and frequency. The smaller the calibration error, the better the model is calibrated.
Sparsification plot
The sparsification plot35,36 is used to depict the relationship between data entities (e.g. uncertainty vs. error) quantitatively and explicitly. Concretely, it first sorts all pixels of reconstructed SR-SIM images in descending order according to a pre-defined sorting criterion, then gradually removes a fraction of pixels and calculates the MAE of the remained pixels with respect to GT-SIM. We respectively adopt the SQUIRREL error, AleaU, and EpisU as the sorting criteria. Meanwhile, the oracle one with the sorting criterion of the true SR error is also presented for reference. The closer a sparsification plot is to the oracle, the better its corresponding sorting criterion correlates with the true SR error.
FG-AleaU and FG-EpisU
BayesDL quantifies uncertainty for each pixel of its reconstructed SR images. However, what sparks users’ interest is the foreground (FG) pixels instead of the background ones. To this end, we propose two metrics, i.e. FG-AleaU and FG-EpisU, to characterize the uncertainty of FG pixels. For brevity of notation, here we term the reconstructed SR images as \({I}_{SR}\) and term both AleaU and EpisU as σ. FG-AleaU and FG-EpisU are calculated by following three steps: (1) performing percentile-normalization as shown in Eq. (5) on the SR image \({I}_{SR}\); (2) segmenting the normalized SR image using Otsu52 segmentation method and then obtaining its corresponding FG mask (denoted as M); (3) extracting the uncertainty of FG pixels (i.e. FG-AleaU or FG-EpisU) from uncertainty maps σ, according to the index provided by the non-zero elements in the FG mask M.
Other evaluation metrics
To quantitatively assess the fidelity of SR images, we calculate the commonly-used PSNR and SSIM53 metrics between SR images and GT-SIM ones. Besides, we also calculate the normalized root-mean-square error (NRMSE), which is defined as:
where \(B\) is the pixel number, \(\max (\cdot )\) and \(\min (\cdot )\) denote maximum function and minimal function, respectively. In terms of SR resolution evaluation, here we adopt decorrelation analysis27 to estimate the highest SR frequency from the local maxima of the decorrelation function. To quantitatively measure the fluorescence SNR level of raw SIM images, we use the metric of the average photon count14. The calculation of the average photon count metric is detailed in Supplementary Note 7. Additionally, we measure the lengths of different actin filaments by first segmenting actin images with the Otsu52 algorithm and converting the resulting binary segmentation map into the 8-bit type. We then use ImageJ’s plugin AnalyzeSkeleton (2D/3D) to extract various skeleton information including the skeletonized actin lengths.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The published BioSR dataset was used to train the BayesDL models for CCPs, MTs, and F-actin (https://figshare.com/articles/dataset/BioSR/13264793)14. The training data for other biological structures and the raw data underlying this work were also uploaded into the figshare repository, publicly accessible at https://doi.org/10.6084/m9.figshare.27824586.v1. Source data are provided with this paper.
Code availability
The source codes of BayesDL-SIM, representative pre-trained models, as well as some example images for testing are publicly accessible on Github repository54 (https://github.com/HUST-Tan/BayesDL-SIM).
References
Abbe, E. Beiträge zur Theorie des Mikroskops und der mikroskopischen Wahrnehmung. Arch. f.ür. mikroskopische Anat. 9, 413–468 (1873).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Sage, D. et al. Quantitative evaluation of software packages for single-molecule localization microscopy. Nat. Methods 12, 717–724 (2015).
Agarwal, K. & Machan, R. Multiple signal classification algorithm for super-resolution fluorescence microscopy. Nat. Commun. 7, 13752 (2016).
Wen, G. et al. High-fidelity structured illumination microscopy by point-spread-function engineering. Light.: Sci. Appl. 10, 1–12 (2021).
Qiao, C. et al. Rationalized deep learning super-resolution microscopy for sustained live imaging of rapid subcellular processes. Nat. Biotechnol. 41, 367–377 (2023).
Qian, J. et al. Structured illumination microscopy based on principal component analysis. eLight 3, 4 (2023).
Gustafsson, M. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Müller, M., Mönkemöller, V., Hennig, S., Hübner, W. & Huser, T. R. Open-source image reconstruction of super-resolution structured illumination microscopy data in ImageJ. Nat. Commun. 7, 10980 (2016).
Huang, X. et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nat. Biotechnol. 36, 451–459 (2018).
Zhao, W. et al. Sparse deconvolution improves the resolution of live-cell super-resolution fluorescence microscopy. Nat. Biotechnol. 40, 606–617 (2021).
Chen, J. et al. Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes. Nat. Methods 18, 678–687 (2020).
Jin, L. et al. Deep learning enables structured illumination microscopy with low light levels and enhanced speed. Nat. Commun. 11, 1934 (2020).
Qiao, C. et al. Evaluation and development of deep neural networks for image super-resolution in optical microscopy. Nat. Methods 18, 194–202 (2021).
Ward, E. N. et al. Machine learning assisted interferometric structured illumination microscopy for dynamic biological imaging. Nat. Commun. 13, 7836 (2022).
Hoffman, D. P., Slavitt, I. & Fitzpatrick, C. A. The promise and peril of deep learning in microscopy. Nat. Methods 18, 131–132 (2021).
Der Kiureghian, A. & Ditlevsen, O. Aleatory or epistemic? Does it matter?. Struct. Saf. 31, 105–112 (2009).
Kendall, A. & Gal, Y. What uncertainties do we need in bayesian deep learning for computer vision? Proceedings of Advances in Neural Information Processing Systems. 30, 5580–5590 (2017).
Gawlikowski, J. et al. A survey of uncertainty in deep neural networks. Artif. Intell. Rev. 56, 1513–1589 (2023).
Kendall, A., Gal, Y. & Cipolla, R. Multi-task learning using uncertainty to weigh losses for scene geometry and semantics. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 7482–7491, https://doi.org/10.1109/CVPR.2018.00781 (2017).
Chang, J., Lan, Z., Cheng, C. & Wei, Y. Data uncertainty learning in face recognition. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 5709–5718, https://doi.org/10.1109/CVPR42600.2020.00575 (2020).
Kar, A. & Biswas, P. K. Fast Bayesian uncertainty estimation and reduction of batch normalized single image super-resolution network. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 4955–4964, https://doi.org/10.1109/CVPR46437.2021.00492 (2021).
Liu, T., Cheng, J. & Tan, S. Spectral Bayesian uncertainty for image super-resolution. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 18166–18175, https://doi.org/10.1109/CVPR52729.2023.01742 (2023).
Liu, T., Liu, J., Li, D. & Tan, S. Improving reconstruction of structured illumination microscopy images via dual-domain learning. IEEE J. Sel. Top. Quantum Electron. 29, 1–12 (2023).
Welling, M. & Teh, Y. W. Bayesian learning via stochastic gradient Langevin dynamics. Proceedings of International Conference on Machine Learning, 681–688, https://icml.cc/2011/papers/398_icmlpaper.pdf (2011).
Zhang, Y. et al. Image Super-resolution using very deep residual channel attention networks. Proceedings of the European Conference on Computer Vision, 286–301, https://doi.org/10.1007/978-3-030-01234-2_18 (2018).
Descloux, A. C., Grussmayer, K. S. & Radenovj, A. Parameter-free image resolution estimation based on decorrelation analysis. Nat. Methods 16, 918–924 (2019).
Guo, C., Pleiss, G., Sun, Y. & Weinberger, K. Q. On calibration of modern neural networks. Proceedings of International Conference on Machine Learning, 1321–1330, https://proceedings.mlr.press/v70/guo17a.html (2017).
Kuleshov, V., Fenner, N. & Ermon, S. Accurate uncertainties for deep learning using calibrated regression. Proceedings of International Conference on Machine Learning, 2796–2804, https://proceedings.mlr.press/v80/kuleshov18a.html (2018).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Belthangady, C. & Royer, L. A. Applications, promises, and pitfalls of deep learning for fluorescence image reconstruction. Nat. Methods 16, 1215–1225 (2018).
Laine, R. F., Arganda-Carreras, I., Henriques, R. & Jacquemet, G. Avoiding a replication crisis in deep-learning-based bioimage analysis. Nat. Methods 18, 1136–1144 (2021).
Pan, S. J. & Yang, Q. A Survey on Transfer Learning. IEEE Trans. Knowl. Data Eng. 22, 1345–1359 (2010).
Culley, S. et al. NanoJ-SQUIRREL: quantitative mapping and minimisation of super-resolution optical imaging artefacts. Nat. Methods 15, 263–266 (2018).
Amini, A., Schwarting, W., Soleimany, A. & Rus, D. Deep Evidential Regression. Proc. Adv. Neural Inf. Process. Syst. 33, 14927–14937 (2019).
Gustafsson, F. K., Danelljan, M. & Schön, T. B. Evaluating scalable Bayesian deep learning methods for robust computer vision. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops 1289–1298, https://doi.org/10.1109/CVPRW50498.2020.00167 (2019).
Goodfellow, I. J. et al. Generative adversarial nets. Proceedings of Advances in Neural Information Processing Systems 27, 2672–2680 (2014).
Wang, H. et al. Deep learning enables cross-modality super-resolution in fluorescence microscopy. Nat. Methods 16, 103–110 (2018).
Chen, R. et al. Deep-learning super-resolution microscopy reveals nanometer-scale intracellular dynamics at the millisecond temporal resolution. Preprint at bioRxiv https://doi.org/10.1101/2021.10.08.463746 (2021).
Huang, B. S. et al. Enhancing image resolution of confocal fluorescence microscopy with deep learning. PhotoniX 4, 2 (2023).
Kiepas, A., Voorand, E., Mubaid, F., Siegel, P. M. & Brown, C. M. Optimizing live-cell fluorescence imaging conditions to minimize phototoxicity. J. Cell Sci. 133, jcs242834 (2020).
Ning, Q., Dong, W., Li, X., Wu, J. & Shi, G. Uncertainty-Driven Loss for Single Image Super-Resolution. Proceedings of Advances in Neural Information Processing Systems 34, 16398–16409 (2021).
Cao, R. et al. Open-3DSIM: an open-source three-dimensional structured illumination microscopy reconstruction platform. Nat. Methods 20, 1183–1186 (2023).
Betzig, E. et al. Imaging Intracellular Fluorescent Proteins at Nanometer Resolution. Science 313, 1642–1645 (2006).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. K. Optical Sectioning Deep Inside Live Embryos by Selective Plane Illumination Microscopy. Science 305, 1007–1009 (2004).
Wang, W., Zhang, H., Yuan, Z., Wang, C. & Lab, B. A. Unsupervised real-world super-resolution: a domain adaptation perspective. Proceedings of the IEEE/CVF International Conference on Computer Vision, 4318–4327, https://doi.org/10.1109/ICCV48922.2021.00428 (2021).
Ning, Q. et al. Learning degradation uncertainty for unsupervised real-world image super-resolution. Proceedings of International Joint Conference on Artificial Intelligence, 1261–1267, https://doi.org/10.24963/ijcai.2022/176 (2022).
Shocher, A., Cohen, N. & Irani, M. Zero-shot super-resolution using deep internal learning. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 3118–3126, https://doi.org/10.1109/CVPR.2018.00329 (2017).
Soh, J. W., Cho, S. & Cho, N. I. Meta-Transfer Learning for Zero-Shot Super-Resolution. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 3513-3522 (2020).
Zhao, W. et al. Quantitatively mapping local quality of super-resolution microscopy by rolling Fourier ring correlation. Light.: Sci. Appl. 12, 298 (2023).
Sternberg, S. R. Biomedical image processing. Computer 16, 22–34 (1983).
Otsu, N. A threshold selection method from gray level histograms. IEEE Trans. Syst., Man, Cybern. 9, 62–66 (1979).
Wang, Z., Bovik, A. C., Sheikh, H. R. & Simoncelli, E. P. Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13, 600–612 (2004).
Liu, T. et al. Bayesian deep-learning structured illumination microscopy enables reliable super-resolution imaging with uncertainty quantification. BayesDL-SIM https://doi.org/10.5281/zenodo.15282200 (2025).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (62071197, 62471192 and 32125024, to S.T. and D.L.), the National Key R&D Program of China (2024YFA1307400, to D.L.).
Author information
Authors and Affiliations
Contributions
S.T. and T.L. conceived the project. J.L. prepared samples and performed the imaging, under the supervision of D.L. T.L. developed the reconstruction code. T.L. and J.L. analyzed the data, with conceptual advice from S.T. and D.L. The manuscript was written with input from all authors, starting from a draft provided by T.L. All authors discussed the results and commented on the manuscript. S.T., D.L., and T.L. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
The authors have a pending patent application on the presented framework.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, T., Liu, J., Li, D. et al. Bayesian deep-learning structured illumination microscopy enables reliable super-resolution imaging with uncertainty quantification. Nat Commun 16, 5027 (2025). https://doi.org/10.1038/s41467-025-60093-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60093-w