Abstract
Ataxia-telangiectasia is a rare genetic disorder characterized by neurological defects, immunodeficiency, cancer predisposition, radiosensitivity, decreased blood vessel integrity, and diabetes. ATM, the protein mutated in Ataxia-telangiectasia, responds to DNA damage and oxidative stress, but its functional relationship to the progressive clinical manifestation of this disorder is not understood. CD98HC chaperones cystine/glutamate and cationic/neutral amino acid antiporters to the cell membrane, and CD98HC phosphorylation by ATM accelerates membrane localization to acutely increase amino acid transport. Loss of ATM impacts tissues reliant on heterodimeric amino acid transporters relevant to Ataxia-telangiectasia phenotypes, such as endothelial cells (telangiectasia) and pancreatic α-cells (fatty liver and diabetes), with toxic glutamate accumulation. Bypassing the antiporters restores intracellular metabolic balance in ATM-deficient cells and mouse models. These findings provide insight into the long-known benefits of N-acetyl cysteine in Ataxia-telangiectasia cells beyond oxidative stress through removing glutamate excess by producing glutathione.
Similar content being viewed by others
Introduction
Ataxia-telangiectasia (A-T) is a rare recessive genetic disorder resulting from the mutation and usually loss of protein of ataxia-telangiectasia mutated protein (ATM1). A-T progressively manifests in multiple phenotypes, including immunodeficiency, neurological defects, cancer, telangiectasia, and diabetes without any established treatment2. ATM, a phosphatidylinositol 3-kinase-like kinase, is largely nuclear in location, though a measurable proportion is found in the cytoplasm3,4. Most work to date has outlined a role for ATM in orchestrating cellular response to DNA double-strand breaks and oxidative stress5. In addition to auto-phosphorylation, ATM phosphorylates and activates key damage response proteins including Tumor suppressor p536 and checkpoint kinase CHK27. As such, in response to damage, ATM regulates cell cycle progression, apoptosis, and DNA repair processes. Beyond these functions, ATM regulates peroxisome and mitochondrial function8,9 and impacts glucose import (GLUT1 pS490) and the glycolysis/pentose phosphate pathway (PPP) switch (HSP27:G6PD), particularly in response to exogenous damage10,11.
Tissues of Atm-null mice display increased levels of oxidative stress and damage12. These observations fit prior work in Atm–/– mice demonstrating that treatment with antioxidants N-acetyl cysteine (NAC) or nitroxides (isoindoline nitroxide, CTMIO or Tempol) increased survival, delayed lymphoma onset, and reduced genomic instability13,14,15,16,17,18,19,20. NAC increases intracellular cysteine and augments the production of glutathione (GSH, a tripeptide of glycine, cysteine, and glutamate) which in turn reduces reactive oxygen species (ROS)21. Nitroxides act as weak superoxide dismutases22 and stimulate heme-containing proteins like catalase23. However, contradicting the benefit of NAC and nitroxides in treating Atm-null mice, increasing oxidative stress by co-deleting either SOD1 or SOD2 (superoxide dismutases), or using different antioxidants (EUK-189 (Mn(III) salen) or α-Tocopherol (vitamin E)), had no impact on survival or tumorigenesis24,25, while overexpression of SOD1 negatively impacted Atm-null mouse phenotypes (see Table S1 for summary). These latter findings reduced enthusiasm for antioxidant-based treatments of A-T despite the apparent benefits of NAC or nitroxides. An alternative scenario is that in the absence of ATM, NAC provides benefits beyond or outside of its antioxidant functions.
NAC is a thiol-containing molecule that readily diffuses across the cell membrane21. As one of the two sulfur-containing amino acids, intracellular levels of cysteine are tightly regulated; for example, in some cells the import of cystine (dimer of cysteine) is dependent upon the xc– antiport system, exchanging cystine for export of a glutamate molecule26. xc– is a cell surface heterodimer formed by the covalent linkage of two proteins, a heavy and a light chain27. The heavy chain, 4F2hc/CD98HC, is the product of the SLC3A2 gene, and following its glycosylation, it acts as a cofactor to direct its various light chain partners (SLC7A5-11,13; members of the heteromeric amino acid transporters (HAT) family) to the cell surface28,29,30. The light chain of the HAT transporters is variable and dictates specific amino acid cargos31. For instance, in combination with xCT (SLC7A11), CD98HC forms the glutamate/cystine antiporter (xc–), whereas, with y + LAT1/2 (SLC7A6/SLC7A7), it forms the cationic/neutral amino acid transporter (y+L)32.
Here we worked to identify the basis of NAC benefit when ATM function is lost, considering that it may not be through reducing oxidative stress. We discovered that acute cystine and arginine influx, concomitant with glutamate efflux, is increased following CD98HC phosphorylation by ATM and decreased time for membrane localization of nascent antiporter. Our findings show that NAC treatment rescues a set of A-T phenotypes in cells and mice by rebalancing intracellular glutamate levels in tissues where intracellular glutamate levels depend on xc– and y+L activity and chronic glutamate accumulation is toxic. We propose a mechanistic link where altered regulation of CD98HC-dependent activities in the absence of ATM explains some of the poorly understood clinical manifestations of A-T.
Results
ATM activity impacts mitochondrial function and glutamine oxidation in primary human endothelial cells
ATM is a ubiquitously expressed gene. With the goal of identifying additional physiological roles for ATM, we examined the expression of ATM in correlation with other genes across 40,000 normal tissues available from the ARCHS4 database using Correlation AnalyzeR33. In line with previous reports8,34,35, we observed a strong negative correlation between ATM and gene expression sets involved in oxidative phosphorylation, citric acid (TCA) cycle, and respiratory electron transport (Fig. 1A; Tables S2–4). To investigate a potential metabolic role for ATM relevant to tissues clinically impacted in A-T but also avoiding concerns of the metabolic impacts often observed with transformed cell lines, we used primary endothelial cells (Human Umbilical Vein Endothelial Cells, HUVEC; relevant to telangiectasia). As ATM is known to be activated by oxidative stress36, we carried out this work at tissue-relevant physiological levels of oxygen (3%) and used the well-established pharmacological inhibitors of ATM–KU55933 and KU60019. We confirmed that these ATM inhibitors (ATMi) blocked the H2O2-mediated activation (phosphorylation) of ATM without loss of total protein (Figs. 1B and S1A) as described with other cell types37,38. Under our cell culture conditions, neither ATMi increased intracellular ROS, even after 48 h (Figs. 1C, D and S1B, C; we demonstrated the sensitivity of the ROS assay (CellROX) by treating the cells with increasing doses of H2O2 (Fig. S1D). The same lack of increased basal oxidative stress was recapitulated with shRNA depletion of ATM (Fig. S1E) and further confirmed with an alternate method of ROS detection (ROS-GloTM H2O2 Assay, Fig. S1F). Cell proliferation/viability experiments demonstrated that ATMi induced a cytostatic effect after 18 h (Figs. S1G, H), recapitulating premature senescence observed with primary A-T or Atm null murine fibroblasts39,40,41,42. To avoid any differences caused by changed proliferative state or viability, we conducted all further investigations with ATMi following 8 h of treatment.
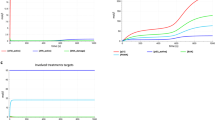
A Correlation analysis between the expression of ATM and gene sets involved in mitochondrial function from a cohort of normal human tissues (RNA-Seq counts provided by ARCHS4 database). B Representative blot showing the efficiency of ATMi in HUVECs treated with H2O2 (n = 6). C Flow cytometry chart showing intracellular ROS levels after KU55933 treatment (histogram shows one representative sample per condition). D Quantification of ROS levels in C. Statistical significance was calculated using the Kruskal-Wallis test with Dunn’s multiple comparison test; data represented as median ± 95% CI, from three independent experiments (n = 3). E Representative graph showing HUVEC’s metabolic potential in response to stressors (Oligomycin and FCCP) after KU55933 and NAC treatments (n = 2 with 5–6 technical replicates per experiment). F Representative graph showing mitochondrial function assay after KU55933 treatment, combined with NAC or Trolox. G Chart showing maximal respiration obtained in F. Statistical significance was calculated using the Kruskal-Wallis test; n = 3 with 5–8 technical replicates per experiment. H Correlation analysis between expression of ATM and genes involved in glutamine deprivation (ARCHS4 database). I Representative graph showing mitochondrial function assay after specific inhibition of glucose, glutamine, or fatty acids oxidation in the presence or absence of ATMi. J Chart showing maximal respiration and spare respiratory capacity after inhibition of glutamine oxidation. Statistical significance was calculated using two-way ANOVA with Šídák’s multiple comparison test; n = 3 with 6–8 technical replicates per experiment. Data in (E-G, I, J) represented as mean ± SD. *p < 0.05, ***p < 0.001. Source data are provided as a Source Data file.
NAC treatment is known to have some physiological benefits in mice and cells with an ATM defect19,20, though neither EUK-189 (a Mn(III) salen) nor α-Tocopherol (vitamin E) antioxidants were able to provide this benefit (Table S1). We therefore asked if ATMi impacts any aspect of metabolism in our primary cells that can be rescued by NAC, but not by Trolox (an analog of vitamin E), another potent ROS scavenger. We first measured the extracellular acidification rate (ECAR) and the oxygen consumption rate (OCR) of HUVECs, with and without ATMi, as indicators of glycolysis and mitochondrial respiration, respectively. Sequential injections to determine the glycolytic flux after ATMi did not show a significant change (Fig. S1I). However, upon injection of stress inducers (Oligomycin and FCCP), HUVECs displayed a strong shift towards glycolysis with ATMi rather than a balanced upregulation of both aerobic respiration and glycolysis as observed in the control (Fig. 1E); this effect was partially rescued by NAC treatment. Examining the OCR results in more detail, we observed a strong impairment in basal respiration, maximal respiration, and spare respiratory capacity following ATMi (Figs. 1F and S1J). The impairment of basal OCR by ATM was not altered by NAC or Trolox treatment. In contrast, NAC, but not Trolox, rescued the maximal respiration and spare respiratory capacity defects (Figs. 1G and S2A), indicating an impact on mitochondrial metabolism. Since we were interested in identifying a NAC-specific Trolox-independent effect, these observations became the focus of our studies.
Given that ATMi reduces the ability to upregulate aerobic respiration, we considered the possibility that this may be the result of altered TCA cycle and electron transport chain (ETC) use. Supporting this concept, examining the ARCHS4 co-expression analysis further revealed a positive correlation between ATM expression and genes upregulated in response to glutamine deprivation (Fig. 1H; Table S5). Consequently, we measured the impact of ATMi on mitochondrial fuel oxidation using specific inhibitors of glucose, glutamine, and fatty acids oxidation (UK5099, BPTES, and etomoxir, respectively). Blocking glucose oxidation clearly impaired mitochondrial function irrespective of treatments (Figs. 1I and S2B left panel); this was expected since endothelial cells are highly glycolytic43. No effect was observed after blocking fatty acids oxidation (Figs. 1I and S2B right panel). Strikingly, blocking glutamine oxidation caused mitochondrial dysfunction only with ATMi, indicating a greater dependency upon glutamine (Figs. 1J and S2C).
Inhibition of ATM alters glycolysis and the pentose phosphate pathway (PPP) activity
To further characterize the metabolic changes induced by ATMi, we performed a stable isotope resolved metabolomics (SIRM) study using [U-13C]-labeled glucose. In line with previous reports10,44,45, we observed reduced glucose-derived label in glycolysis and PPP metabolites after ATMi treatment based on 13C incorporation (Fig. 2A blue and pink panels). For PPP, a clear block in the pathway was noted in the 6PG dehydrogenase step, as 13C labeling was unaltered in 6PG but reduced in R5P with ATMi. To confirm this block, we measured NADPH levels by measuring the NADP+/NADPH ratio (Fig. S2D). No obvious block and metabolite accrual was observed for glycolysis, despite a clear reduction of incorporated label upon ATMi evident from glucose-6-phosphate (G6P) onwards. In fact, our ECAR measurements support that glycolysis activity is intact and can be increased in response to excess glucose or blocking ATP synthesis in ETC (oligomycin) (Figs. S1I, S2A and 1E), demonstrating no block to glycolysis per se. Therefore, our results fit with prior work that ATMi reduces glucose uptake11,44, though the ECAR results indicate no decrease in flux to lactate, thereby providing an explanation for the observed reduction in the amount of labeled glycolysis intermediates. Given these observations, we considered that the flow of glucose through glycolysis into the TCA/Krebs cycle may be altered upon ATMi. An impact on respiration by ATMi was also indicated by the Seahorse assay (Figs. 1E–G, I, J and S1J) and, more importantly to our interest, it can be rescued in a NAC-specific manner.
A13C6-glucose SIRM tracing showing the major significant metabolite changes observed when comparing control (black bars) and 8 h of ATMi (KU55933; red bars) treatment on glycolysis (blue panel), pentose phosphate pathway (pink panel), and TCA cycle (green panel) in HUVECs. Presented are total unlabeled metabolite (0), informative individual 13C isotopologues, and total amount of labeled isotopologues (*Total) presented as either µmol/g protein (absolute amounts) or % 13C of Total (relative amounts of labeled and unlabeled). Statistical significance was calculated using unpaired t-test with two-stage linear step-up procedure of Benjamini, Krieger, and Yekutieli; n = 3 with 2 technical replicates per experiment. B Chart showing intracellular glutamate levels in cells treated with KU55933, NAC, and in combination. Statistical significance was calculated using one-way ANOVA with Tukey’s multiple comparison test (n = 3 with 2 technical replicates per experiment). C Chart showing intracellular GSH levels in cells treated with KU55933, NAC, and in combination. Statistical significance was calculated using one-way ANOVA with Tukey’s multiple comparison test; n = 2 with 2–3 technical replicates per experiment. D Chart showing mRNA levels (n = 2) and blots (n = 3) showing protein levels of GCLc and GCLm after ATMi. Statistical significance was calculated using two-way ANOVA with Šídák’s multiple comparison test; n = 2 with 3 technical replicates per experiment. Data represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Source data are provided as a Source Data file.
ATM inhibition alters TCA cycle while accumulating glutamate
To assess the impact of ATMi on the Krebs cycle, we tracked 13C label from glycolysis into the Krebs cycle (Fig. 2A green panel) and noted a small overall decrease in glucose-derived 13C upon ATMi, ~5% for most metabolites, other than citrate which was reduced about 50%. We also noted the loss of the pyruvate carboxylase (PC; dark green 13C) activity following ATMi as shown by a significant decrease in “reductive” 13C3-malate and 13C3-fumarate, as well as 13C3-Asp production and “oxidative” 13C5-citrate compared to control cells. In contrast, the “oxidative” production of 13C2-glutamate (Glu) (Fig. 2A) via citrate to α-ketoglutarate was increased upon ATMi treatment and, to a lesser extent, so was 13C2-glutathione (GSH). We confirmed the increase in labeled glutamate by NMR analysis (Fig. S2E, F). However, though 13C increased in both Glu and GSH, we noted that the total amount (unlabeled + labeled) of Glu increased (299 to 327 µmol/g) while the total amount of GSH decreased (176 to 150 µmol/g). These data firstly indicate that, in HUVEC cells, the majority of the Krebs cycle metabolites are likely derived from non-glucose sources such as glutamine. Secondly, they indicate that upon ATMi there is an increase in glucose-derived 13C flowing through oxidative TCA cycle to Glu and GSH rather than promoting NADH and FADH2 production corresponding to the decrease in ETC activity indicated by OCR measurement (Fig. 1F). To better understand these observations, we performed an analogous SIRM experiment with [U-13C,15N]-glutamine (Fig. S2G) and found that glutamine indeed contributes significantly to the pool of Krebs cycle metabolites irrespective of ATMi (total 13C enrichment; Fig. S2G *Total). Key points noted from this experiment were (i) oxidative TCA cycle, the main mechanism for 13C-citrate production in HUVEC cells, was increased upon ATMi, while reductive carboxylation decreased, and (ii) ATMi treatment resulted in more de novo GSH production without increasing overall GSH levels. In contrast, following ATMi, it appeared that both unlabeled and labeled glutamate levels increased, with an increased proportion of labeled glutamate, including use of αKG as a source (indicated by either 13C and no 15N, or 15N and no 13C). Overall, these data corroborate the Seahorse results, with ATMi impacting basal TCA cycle function. They also support the 13C-glucose data, with ATMi causing an increase in glutamate levels derived from both glucose-derived αKG and glutamine, but with some impairment in GSH production despite production of de novo labeled GSH. To further confirm the effect of ATMi on glutamate levels, we directly assessed glutamate levels and found that ATMi does in fact induce a significant increase (~10%) in intracellular glutamate over the 8 h of ATMi exposure (Fig. 2B). To independently confirm this observation, we examined primary Atm–/– mouse embryonic fibroblasts (MEFs) and found that these constitutively null cells display a chronic accumulation of intracellular glutamate compared to controls (Fig. S2H).
Having observed that ATMi causes glutamate accumulation in both SIRM experiments as well as increased labeled GSH without an equivalent increase in total amounts of GSH, we decided to investigate the impact of ATMi on GSH levels more directly. We found that GSH levels decreased significantly in HUVEC upon ATMi treatment (Fig. 2C). Given the prior benefits we observed for NAC in our Seahorse assays, we tested if NAC could restore intracellular GSH, which it did (Fig. 2C). This indicates no defect in ability to produce GSH from constituents and in fact we observed no decrease in the expression of GSH producing enzymes GCLc/GCLm (Fig. 2D). NAC relieves the rate-limiting component of GSH production by providing excess cysteine, allowing the production of the tripeptide when combined with glycine and glutamate. Considering this, we examined the effect of NAC on ATMi-increased glutamate levels and noted a significant reduction (Fig. 2B). From this data and the 13C5,15N2-glutamine-based SIRM experiment we reasoned that ATMi alters both GSH synthesis and utilization but with increasing deficit in total GSH. There are many possible fates for GSH beyond oxidation to GSSG, including glutathionylation of proteins46, efflux as conjugates47, or degradation by CHAC1/248, therefore, we focused on identifying the basis of the GSH synthesis defect induced by ATMi. Given no overt defect in the enzymes necessary for GSH production, we considered that the production defect may result from limited substrate availability, most notably cysteine, given that this is the rate-limiting component and would be directly rescued by NAC treatment.
ATM phosphorylates CD98HC to regulate cystine uptake and glutamate export
In response to ATMi or in ATM knockout MEFs, we noted an increase in intracellular glutamate levels and decreased GSH levels, and that these defects were counteracted by NAC; these results indicate a defect in cysteine availability. Intracellular levels of cysteine are tightly controlled, being either produced de novo from methionine and serine (transsulfuration pathway) or, more often, actively imported from outside the cell as cystine via the xc– antiporter with Glu efflux, a process bypassed by NAC. We therefore performed RNA-seq to identify any gene expression changes upon ATMi treatment and noted upregulation of genes involved in de novo cysteine synthesis from serine and methionine (Fig. S3A), suggesting a compensatory response to maintain intracellular cysteine levels. This led us to hypothesize that ATM could be involved in promoting the activity of the cystine/glutamate antiport system, xc– (Fig. 3A). Using radiolabeled cystine, we found that cystine import was significantly impaired by ATMi (Fig. 3B) or shATM knockdown (Fig. S3B,C). Consistent with a role for ATM in stimulating basal xc– activity we also found that the level of extracellular glutamate was lower following ATMi compared to untreated control (Fig. 3C). Sulfasalazine (SAS) and Erastin, two specific inhibitors of the xc– antiport system, were used as controls for the experiments and displayed a more profound effect than ATMi. Overall, inhibition of ATM function results in a partial decrease in xc– activity, leading to an accumulation of intracellular glutamate and insufficient cystine import to maintain basal levels of GSH in the cell.
A Representation of the xc– antiport involved in cystine/glutamate transport (created in Adobe Illustrator, using selected artwork from Servier Medical Art, provided by Servier, licensed under a Creative Commons Attribution 4.0 license, https://creativecommons.org/licenses/by/4.0/). B 14C-L-cystine uptake after treatment with KU55933 or KU60019; SAS and Erastin are used as specific inhibitors of xc–. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparison test; n = 3 with 2–4 technical replicates per experiment. C Fluorometric analysis of extracellular levels of glutamate after ATMi. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparison test; n = 3 with 2–4 technical replicates per experiment. D Chart showing GSH levels measured at different times following ATMi, xc– inhibition (SAS) or the combination (n = 2 with 2 technical replicates per experiment). Data represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; asterisks show significance vs. Control. E Table showing the highly conserved SQ site in SLC3A2. F ATM consensus motif (www.phosphosite.org)125. G Representative images of proximity ligation assay (PLA) showing the interaction between CD98HC and ATM, shRNA-ATM cells were used to show signal specificity. H Dot blot showing binding specificity of the newly synthesized phospho-antibody. I Blots to further confirm antibody specificity in cell lysates treated with alkaline phosphatase (n = 2). J Subcellular fractionation after 2 h of treatment with H2O2. MEK1/2 and LAMIN B1 were used as cytoplasmic and nuclear markers, respectively (n = 3). K Blots showing the detection of phosphorylated CD98HC, its induction by H2O2, and inhibition by KU60019 (n = 3). L Representative images of the photoconversion assay in HEK293 cells transfected with SLC3A2 wild type or phospho-dead. The line plots on the right show average fluorescence intensity from several cell cross-section profiles (three independent experiments, also see Fig. S4B). M Chart showing quantification of fluorescence intensity 24 h after photoconversion. Statistical significance was calculated using two-way ANOVA with Šídák’s multiple comparison test; n = 15, from two independent experiments. Data represented as median ± 95% CI. ****p < 0.0001. Source data are provided as a Source Data file.
Considering that we observed no increase in ROS following ATMi for HUVECs grown at 3% O2, we decided to evaluate the reduced/oxidized state of GSH/GSSG. We observed a decrease in the ratio of GSH/GSSG following 8 h of ATMi treatment, but the change in absolute GSSG levels was not significant (Fig. S3D). This result supports the concept that there is no inherent increase of ROS upon ATMi when cells are maintained at 3% O2, but there is a defect in maintaining GSH levels. This goes in line with the decreased NADPH production due to decreased oxidative PPP activity, as NADPH is used by glutathione reductase to recycle GSH from GSSG. To better understand how ATM impacts intracellular levels of GSH pools, we conducted a time course experiment to monitor GSH changes (Fig. 3D). Interestingly, ATMi alone caused a significant initial decrease (in 1–2 h) of intracellular GSH levels, which then stabilized without returning to control levels in the 8 h monitored. This observation is consistent with our conclusion that the increase in de novo synthesis of cysteine is an attempt to compensate for decreased cystine import. As expected, complete inhibition of xc– by SAS caused a dramatic depletion of GSH. NAC treatment rescued normal GSH levels irrespective of SAS or combined SAS and ATMi. From this data, it appears that ATM augments xc– antiport activity under basal conditions. However, ATM is best understood for activating a pleiotropic cellular response to damage, particularly to ionizing radiation. To explore this possibility, we used primary MEFs and noted that though ionizing radiation induced a strong influx of cystine irrespective of ATM status, the amount of uptake by Atm–/– MEFs was significantly less than that observed for Atm+/+ MEFs (Fig. S3E). Again, these data support the notion that ATM augments xc– antiport activity, in this case in response to ionizing radiation.
Given the concept that ATM regulates xc– antiport activity, the most likely mechanism would involve a direct phosphorylation event by the ATM kinase. Evaluating the sequences of the two proteins that constitute xc– antiporter revealed that CD98HC has a highly conserved SQ site in its intracellular tail (S103, Fig. 3E) that is embedded within a canonical ATM target motif (Fig. 3F). To determine if CD98HC is a substrate of ATM, we first demonstrated their interaction using the proximity ligation assay (PLA, Figs. 3G and S3F) and confirmed it by co-immunoprecipitation (Fig. S3G). We then generated a phospho-specific monoclonal antibody against CD98HC (S103, Fig. 3H, I); this antibody detected the phosphorylation in both glycosylated CD98HC (~100 kDa) and native CD98HC (~50-60 kDa, Fig. 3J). Treatment with the protein glycosylation inhibitor tunicamycin verified reduction of the ~100 kDa glycosylated CD98HC with concomitant increase in ~50–60 kDa unglycosylated CD98HC (Fig. S3H). Detection of phosphorylation of native CD98HC indicates that ATM phosphorylates this protein before it covalently binds to its partner protein and localizes to the cell membrane. Subcellular protein fractionation experiments confirmed that CD98HC phosphorylation at S103 occurs in the cytoplasm (Fig. 3J). CD98HC phosphorylation was increased upon induction of ATM by H2O2, but both basal and induced phosphorylation was abolished by ATMi (Figs. 3K and S3I). To confirm that no other DNA damage-responsive PIKK family member phosphorylates CD98HC S103, at least in response to H2O2, we used inhibitors of DNA-PKcs and ATR (AZD7648 and AZD6738, respectively) and observed no impairment (Fig. S3J). Given that AKT can sometimes mediate an indirect ATM phosphorylation event we also inhibited AKT using MK2206, but again observed no impairment of CD98HC phosphorylation (Fig. S3K).
CD98HC is understood to act as a chaperone for HATs, facilitating their localization to the cell surface28,29,30. We therefore developed an assay to evaluate the rate of CD98HC trafficking to the membrane by fusing CD98HC to the photoconvertible protein mEos3.2. Upon photoconversion, all mEos3.2 present in the cell shifts from green to red fluorescence, while any subsequently produced mEos3.2 will remain green fluorescent. This allows for precise tracking of the red mEos3.2 location over time. With this assay, we demonstrated a clear impairment of the S103A mutant (phospho-dead) CD98HC::mEos3.2 trafficking to the membrane compared to wildtype sequences (Figs. 3L, M andS4A, B). Overall, our findings demonstrate that ATM interacts with and phosphorylates the intracellular 52 kDa monomeric CD98HC in response to oxidative stress to increase the rate of de novo antiport trafficking to the cell surface, providing a mechanistic basis for the impaired cystine import and glutamate accumulation observed with ATM inhibition.
ATM phosphorylation is required for optimal angiogenesis
Given that ATM-dependent phosphorylation increases xc– antiporter activity by increasing its cytoplasm to membrane localization, we asked two key questions: does the impact of ATM phosphorylation of CD98HC extend to other HATs? And can we identify any physiological impact for the reduction in antiporter activity in response to ATMi? To address these questions, we considered another HAT, the y+L system. The antiporter formed between CD98HC and y+LAT1/2 (SLC7A6/SLC7A7) has high affinity for the uptake of arginine49,50,51 (Fig. 4A). Arginine import is essential for nitric oxide synthesis and angiogenesis52,53; a function pertinent to the telangiectasia phenotype of A-T. We found that ATMi (Fig. 4B) or shATM depletion (Fig. S4C) significantly reduced [14C]-L-arginine uptake in line with loss of import by one arginine influx system54,55,56. Given this observation, we went on to test the impact of ATMi on endothelial vessel formation and noted a clear impairment (Fig. 4C,D). The network of capillaries formed in the presence of ATMi had a significantly higher number of meshes and master segments length compared to the control (Fig. 4E, F), resulting in a shorter mesh index (Fig. S4D). These phenotypes indicate an inability of ATM-inhibited HUVECs to migrate and form the same lumen width observed for control cells. Supporting our model that inhibiting ATM decreases both xc– and y+L activity, we found that combining NAC and L-Arginine Ethyl Ester (LAEE) (cell-permeable forms of cysteine and arginine, respectively) restored normal angiogenesis irrespective of ATMi (NAC or LAEE each appear to partially rescue the impact of ATMi, while only the combination of NAC and LAEE significantly reduced the consequence of ATMi on meshes and master segments). Interestingly, using the wound healing scratch assay, we found that ATMi impaired migration, which was restored upon NAC treatment (Fig. S4E,F). These results are further supported by gene expression analysis showing that ATMi modulates the expression of genes involved in migration and vessel formation in endothelial cells (Fig. S4G). Overall, these results demonstrate that ATM is involved in angiogenesis by modulating the activity of both xc– and y+L transporters and that their decreased activity upon ATMi can be bypassed by the addition of both NAC and LAEE.
A Drawing showing one of the two antiport systems involved in arginine uptake in endothelial cells (created in Adobe Illustrator, using selected artwork from Servier Medical Art, provided by Servier, licensed under a Creative Commons Attribution 4.0 license, https://creativecommons.org/licenses/by/4.0/). B Representative chart showing 14C-L-arginine uptake after ATMi. Statistical significance was calculated using one-way ANOVA with Tukey’s multiple comparison test; n = 3 with 2–4 technical replicates per experiment. C Picture showing the parameters evaluated in the vessel formation assay. D Representative images of the angiogenesis assay. E, F Quantification of images using the Angiogenesis Analyzer from Image J. Statistical significance was calculated using the Kruskal-Wallis test with Dunn’s multiple comparison test; n = 3 with 2 technical replicates per experiment, 10 FOV/replicate. Data represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001; when not specified, asterisks show significance vs. Control. Source data are provided as a Source Data file.
ATM phosphorylation of CD98HC impacts alpha and beta pancreatic cells
Having established an acute effect of CD98HC phosphorylation on the activity of associated antiporters, we wanted to determine whether this consequence relates to a progressive phenotype pertinent to A-T, a chronic disease. A-T patients are known to develop abnormalities in glucose homeostasis with 25% of those patients who survive to age 30 developing diabetes mellitus57,58. Similar to neurons, pancreatic α and β cells use glutamate as a signaling molecule59,60 and are highly sensitive to glutamate accumulation61 (Fig. 5A). Thus, we considered the possibility that loss of ATM could lead to a chronic accumulation of glutamate in pancreatic islets with a potential toxic impact. To explore this possibility, we first evaluated single-cell RNA sequencing data from multiple studies of the human pancreas (GSE81076, GSE85241, GSE86469, E-MTAB-5061) and noted heterogeneity in ATM expression across different cell types (Figs. 5B and S5A). Expanding on this, we observed an inverse relationship between ATM and SLC3A2 expression levels (Fig. 5C); indicating that, in this tissue, the absence of ATM-induced activity is compensated by the increased presence of the antiporter. To explore this relationship further, we turned to use murine α TC1 Clone 9 and β TC6 pancreatic cells as an in vitro model. Both cell types were sensitive to ATMi (Fig. 5D), though the effect was clearly stronger in α cells. In addition, both cell types were sensitive to Erastin, substantiating their reliance on the xc– antiport system. We did note that α cells showed a non-significant increase in ROS levels after ATMi that we did not observe in β cells (Fig. S5B, C). Similar to HUVECs, ATMi caused a decrease in intracellular GSH levels and a concomitant increase in intracellular glutamate in both α and β pancreatic cells (Fig. 5E,F). Of note, the relative increase in intracellular glutamate in α cells induced by ATMi is much higher than that observed in β cells, though the absolute basal glutamate levels are significantly higher in β cells compared to α cells (Fig. S5D). If glutamate toxicity is the basis of ATMi-induced death, we would expect that (1) NAC would prevent this by promoting GSH production, while (2) ester-GSH (eGSH) would not confer this benefit despite reducing ROS levels, demonstrating that the rescue is not due to mitigating oxidative stress. This is exactly what we observed (Fig. S5E, F). Comparable with HUVECs, H2O2 treatment induced CD98HC phosphorylation in an ATM-dependent manner in both α and β cells (Fig. 5G). We then asked if islet cells demonstrated an ATM-dependent metabolic change similar to that observed with HUVECs. Here, we found a significant impact on mitochondrial respiration and glycolysis following ATMi in α cells (Fig. 5H, I). In contrast, the decrease observed with β cells was milder and not significant (Figs. 5H and S5G). Finally, considering the functional role of pancreatic islet cells, we asked whether ATMi or depletion impacts hormone secretion and found that indeed glucagon and insulin secretions were impaired in α and β cells, respectively (Fig. 5J). Of note, glutamine-derived αKG followed by reductive carboxylation in the TCA cycle has been shown to be required for insulin secretion62, which reconciles well with the impacts on metabolism and impaired insulin secretion we observe with ATMi. Overall, our findings indicate that pancreatic islet cells, particularly α cells, are impacted by the loss of ATM function, affecting cell viability, glutamate accumulation, GSH production in response to oxidative stress, and hormone secretion.
A Simplified illustration showing TCA cycle and ATP levels modulating insulin and glucagon release (created in Adobe Illustrator, using selected artwork from Servier Medical Art, provided by Servier, licensed under a Creative Commons Attribution 4.0 license, https://creativecommons.org/licenses/by/4.0/). B Ridge plots showing ATM and SLC3A2 heterogeneous expression in human pancreatic cells (data obtained from multiple scRNA-Seq studies). C Box plots showing quantile-normalized ATM and SLC3A2 expression in different human pancreatic cell types. Statistical significance was calculated using the Mann-Whitney test; data showing median, IQR (0.25 and 0.75), whiskers extending to ±1.5*IQR (~95% CI). D Representative confluency charts of α and β cells following ATMi and xc– inhibition (Erastin). Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparison test; n = 3 with 4 technical replicates per experiment. Data represented as mean ± SEM. E, F Intracellular GSH and glutamate levels after ATMi in α and β cells. Statistical significance was calculated using two-way ANOVA with Šídák’s multiple comparison test; n = 3 with 2–3 technical replicates per experiment. G Representative blots showing CD98HC phosphorylation (S103) after H2O2 and ATMi in α and β cells (n = 2). H, I Basal respiration of α and β cells and glycolytic function of a cells after ATMi. Statistical significance was calculated using two-way ANOVA with Šídák’s multiple comparison test; n = 3 with 8–10 technical replicates per experiment. J Glucagon and insulin secretion in α and β cells after ATMi/knockout. Statistical significance was calculated using one-way ANOVA with Šídák’s multiple comparison test; n = 3 with 3 technical replicates per experiment. Data represented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; When not specified, asterisks show significance vs. Control. Source data are provided as a Source Data file.
Atm deficiency results in glucose intolerance and pancreatic islets malfunction
Our observation that loss of ATM activity impacts α and β cell function in culture suggested a tractable model to test in vivo. Others reported an insulin secretion defect in Atm–/– mice63; however, the underlying mechanism was not elucidated. We therefore set out to characterize glucose homeostasis of Atm–/– mice fed with a standard chow diet. Using the glucose tolerance test we found that Atm–/– male mice developed glucose intolerance by 6 months of age, while female mice displayed this defect by 1 year (Figs. 6A, B and S6A). In contrast, insulin sensitivity assessed with an insulin tolerance test was similar between genotypes (Fig. 6C). To further study these responses, we examined the impact of fasting or feeding. When fasted, we observed no difference between the genotypes, but when allowed to feed, blood levels of insulin were significantly lower in Atm–/– mice while glucose levels increased (Fig. S6B, C). Based on our cell culture work, we expected the absence of ATM to impact a cell function by reducing the amount of glucagon, resulting in reducing gluconeogenesis and thus enhancing fatty acid accumulation in the liver. Substantiating our cell-based findings we found that Atm –/– mice develop fatty liver (Fig. 6D), which may be related to lower hepatic fat oxidation from the reduced glucagon. In fact, we observed the same increased accumulation of lipids in liver samples from A-T patients; this finding matches a clinical report using a larger cohort of A-T patients64. To further understand the impact of ATM deficiency on α cells, we evaluated the percentage of glucagon-expressing cells (indicating the presence of α cells) in the pancreatic islets of 6-month-old mice. We observed a significant reduction in the percentage of islet area that was glucagon positive in Atm–/– compared to Atm+/+ mice (Figs. 6E and S6D). We found no differences in pancreatic islet size or CD98HC expression between genotypes (Fig. S6E,F). Strikingly, and in line with our cell culture findings, pancreatic islets of 6-month-old Atm–/– mice accumulated higher levels of glutamate and glutamine compared to Atm+/+ mice (Figs. 6F, G and S6G). To confirm our findings, we performed the same analysis using an independent Atm null model (Atmtm1Fwa, 3–5 month-old)65 and observed similar results, increased glutamate and glutamine levels in the Atm null pancreatic islets compared to wildtype (Fig. S6H). Finally, having observed a strong recapitulation of our cell culture results, we went on to determine if we would also observe the same metabolic defect in islets. For this, we used an ex vivo system, isolating pancreatic islets from Atm+/+ and Atm–/– mice. We found that glucose response was significantly impaired in islets isolated from Atm–/– compared to Atm +/+ mice (Fig. 6H,I). Mimicking our cell culture results, Atm–/– islets showed a similar impairment in the spare respiratory capacity as well as a lower level of basal respiration (Figs. 6I and S6I). Altogether, our findings demonstrate that the absence of ATM results in a progressive chronic metabolic defect in pancreatic islets, with a clear loss of glucagon-expressing α cells and a progressive decline of endocrine functions for both alpha and beta cells.
A, B Intraperitoneal (GTT) and oral glucose tolerance test (OGTT) in Atm+/+ and Atm–/– mice (GTT n = 6 Atm+/+, n = 4 Atm–/–; OGTT n = 3 Atm+/+, n = 4 Atm–/–). C Glucose measurements after intraperitoneal injection of insulin (ITT, n = 6 Atm+/+, n = 4 Atm–/–). Statistical significance in (A–C) was calculated using two-way ANOVA with Šídák’s multiple comparison test; data represented as mean ± SEM. D Representative images and quantification of lipid levels by Oil Red O staining in the liver from 6-month-old Atm+/+ (n = 6) and Atm–/– (n = 5) mice and A-T patients. E Representative images of glucagon staining in pancreas from Atm+/+ (n = 5) and Atm–/– (n = 8) mice and resultant quantification (each dot represents a single islet). F, G Representative images and quantification of glutamate and glutamine staining in pancreatic islets of Atm+/+ (n = 5) and Atm–/– (n = 8) mice (each dot represents a single islet). Statistical significance in (D–G) was calculated using a two-tailed Mann-Whitney test; data represented as median ± 95% CI. H Mitochondrial respiration of pancreatic islets isolated from 6-month-old Atm+/+ (n = 5) and Atm–/– (n = 4) mice. Data represented as mean ± SEM. I Parameters evaluated in (H) show glucose response and mitochondrial performance in Atm+/+ vs. Atm–/– mice (each dot represents an animal). Statistical significance was calculated using two-way ANOVA with Šídák’s multiple comparison test; data represented as mean ± SD. *p < 0.05, **p < 0.01, ****p < 0.0001. Source data are provided as a Source Data file.
N-Acetyl cysteine rescues glucose intolerance, glutamate accumulation, and glucagon production in the pancreatic islets of ATM-deficient mice
Our findings in cells and mice led to the prediction that the underlying consequence of a lack of ATM activity in the pancreas is a chronic accumulation of toxic levels of glutamate through an inability to acutely regulate cysteine levels. If this model is correct, then NAC supplementation should allow GSH production and thereby alleviate toxic glutamate accumulation and prevent the glucose intolerance observed in Atm–/– mice. We, therefore, supplemented the drinking water of our mice with NAC from conception throughout their life. NAC treatment rescued both the glucose intolerance and hepatic lipid accumulation observed in Atm–/– mice (Fig. 7A, C). We went on to show that NAC treatment of Atm–/– mice rescued the percentage of glucagon-positive areas in the pancreatic islets and reduced glutamate accumulation to normal levels (Figs. 7D, E and S6J, K). These results clearly show that in the absence of ATM, NAC supplementation works to reduce the chronic accumulation of toxic levels of intracellular glutamate pools in pancreatic islets, thereby allowing maintenance of glucose homeostasis.
A, B Intraperitoneal (GTT) and oral glucose tolerance test (OGTT) in Atm+/+ and Atm–/– mice supplemented with NAC (GTT n = 9 Atm+/+, n = 7 Atm–/–, n = 8 Atm–/– + NAC; OGTT n = 3 Atm+/+, n = 4 Atm–/–, n = 4 Atm–/– + NAC). Statistical significance was calculated using two-way ANOVA with Dunnett’s multiple comparison test. Data represented as average ± SEM. C Representative images of Oil Red O staining in the liver from 6-month-old Atm+/+ (n = 6) and Atm–/– (n = 5) mice supplemented with NAC (Atm+/+ + NAC n = 3, Atm–/– + NAC n = 4). Representative images and quantification of glucagon (D) and glutamate (E) staining in pancreas from Atm +/+ and Atm –/– mice supplemented with NAC (each dot represents a single islet, n = 5 Atm+/+, n = 8 Atm–/–, n = 3 Atm+/+ + NAC, n = 4 Atm–/– + NAC). Statistical significance was calculated using the Kruskal-Wallis test with Dunn’s multiple comparison test; data represented as median ± 95% CI. *p < 0.05, ***p < 0.001, ****p < 0.0001; when not specified, asterisks show significance vs. control. Source data are provided as a Source Data file.
Discussion
In this study, we aimed to find additional ATM targets/functions that improve our mechanistic understanding of A-T phenotypes and may lead to new treatments for these patients. We started by characterizing the metabolic function of ATM in primary endothelial cells (HUVECs), relevant to the telangiectasia phenotype of A-T. In line with previous reports using A-T fibroblasts, thymoblasts, and cardiomyoblasts8,35, we observed impairment in mitochondrial function of HUVECs after ATMi. However, we did not see an increase in ROS levels upon ATMi as was reported before with ATM-deficient cells34,66; this is most likely due to our use of more physiological levels of oxygen in our cell cultures (3% O2) as opposed to atmospheric (21%) in other studies which is expected to increase oxidative stress levels67. Despite ATMi not increasing oxidative stress in our models, we found HUVECs to be very sensitive to ATMi, suggesting an essential ROS-independent role for ATM in basal maintenance of endothelial cells; under basal conditions, activated ATM is present in the cytoplasm (Fig S1A).
Following these observations, we employed [U-13C]-glucose and [U-13C, 15N]-glutamine in stable isotope tracing to better understand metabolic reprogramming in HUVEC induced by ATMi. Following ATMi we found decreased glucose uptake, compromised PPP, and depleted glycolytic intermediates despite no impact on glycolytic flux and mitochondrial anaplerosis. Glucose-derived label increased in glutamate with an overall increase in glutamate accumulation; critically, this accumulation was rescued by NAC treatment. We went on to find that despite glutamate accumulation, total GSH levels were reduced after ATMi while GSSG level remained largely unchanged. These results indicated that the reduction of GSH is not through oxidation. With no apparent defect in the expression of the enzymes used for GSH production, we proceeded to examine substrate availability. Given that cysteine is the rate-limiting component for GSH production and NAC rescues A-T phenotypes, we pursued the potential role for ATM in controlling intracellular cysteine levels.
We first demonstrated a cystine uptake defect in ATMi-HUVECs as well as a decrease in extracellular export of glutamate compared to control cells. A similar effect was observed with Atm–/– primary MEFs, which were not able to invoke as strong a cystine uptake response following irradiation as wild-type cells, supporting our findings with the pharmaceutical inhibitors. Next, we showed that the interaction of ATM and CD98HC led to the identification of a highly conserved ATM target phosphorylation site in the intracellular tail of CD98HC (S103). After developing a phospho-specific antibody, we showed that H2O2-induced monomeric CD98HC S103 phosphorylation was ATM dependent (Fig. 3K). Though expression of phospho-dead CD98HC increases xc–activity, we did note that phospho-dead CD98HC had a reduced rate of CD98HC trafficking to the cell membrane (Fig. 3L, M). These data provide mechanistic insight into the functional consequence of this phosphorylation that fits with the observed phenotypes.
Our finding that ATM phosphorylates CD98HC is the first demonstration that posttranslational modification of CD98HC can impact overall antiport channel activity, albeit by increasing the rate of trafficking to the cell membrane. Considering this, we examined the y+L channel, a HAT formed by y+LAT1/2 and CD98HC, used by endothelial cells to import arginine32,54,55,56. We found that either ATMi or ATM depletion significantly reduced arginine uptake which corresponded to an angiogenesis defect. In line with our data, Jia et al. showed that ATM haploinsufficiency reduced angiogenesis after myocardial infarction in mice68. Endothelial dysfunction is a hallmark of atherosclerosis progression and blood vessel function69,70. Concordantly, our findings provide insights in regard to the exacerbated atherosclerosis observed in Atm–/–ApoE–/– mice models and the ocular telangiectasia of A-T patients71,72.
One poorly characterized phenotype of A-T patients is diabetes58. Since α and β pancreatic cells respond to glutamate levels and metabolism of this amino acid by reductive TCA cycle secrete glucagon and insulin61,62,73, we considered that the accumulation of intracellular glutamate upon loss of ATM function may impact these cells. We were able to recapitulate the ATMi induced phenotypes we observed in HUVECs in immortalized α and β cells, with the additional consequence of impaired hormone secretion. These results indicate that ATM has a normal role in modulating xc– antiport activity in pancreatic islet cells, be that in response to fluctuations in basal oxidative stress levels, acute stress exposures, or other currently unknown stimuli. Of note, unlike other PI3-like kinases (ATR, PTEN, etc.), ATM displays circadian rhythm-dependent expression changes in both α and β cells74, which suggests an innate regulation of ATM activity in this tissue. To demonstrate the potential impact on the pancreatic islet function, we used Atm–/– mice41,75. Similar to another Atm-deficient mouse model63, our Atm–/– mice developed glucose intolerance, most likely related to the reduction in postprandial insulin secretion. In agreement with an insulin secretion defect, the Atm–/– mice developed fatty livers, a condition characteristic of type 2 diabetes and reported in A-T patients64,76. We also observed a reduced amount of glucagon-producing α cells in Atm–/– mice compared to Atm+/+; this loss of α cells provides an explanation for the observed fatty liver phenotype and may represent a precursor to the acquired peripheral insulin resistance that has been reported for AT patients77,78. Overall, both our animal- and cell-based work demonstrate that without functional ATM there is a defect in endocrine activity in pancreatic islets. Next, we surmise that the phenotypes result from an underlying defect in the release of glucagon and insulin by the α/β pancreatic cells, respectively. Recapitulating our cell-based assays, we demonstrated impaired mitochondrial respiration in pancreatic mouse islets isolated from Atm–/– mice, a function that is essential for pancreatic islet insulin release79.
Last, we supplemented our Atm–/– mice with NAC water. Remarkably, NAC treatment prevented both the glucose response defect and the accumulation of hepatic lipids. Furthermore, NAC treatment rescued the glutamate accumulation in the pancreatic islets, presumably by reducing intracellular glutamate via GSH production. These data strongly support the use of NAC as an effective treatment for A-T patients to deplete chronic accumulation of intracellular glutamate levels. It is tempting to speculate that the previously observed benefit of nitroxides in Atm null mice13,16,17,18 may have been achieving the same benefit indirectly via activation of the heme protein cystathionine beta-synthase, the rate-limiting enzyme catalyzing the first step of transsulfuration to synthesize cysteine from methionine80. It should be noted that depleted GSH pools in the absence of ATM were reported more than 20 years ago in A-T fibroblasts and red blood cells81,82. Similarly, a more recent study, using cerebellar astroglia isolated from ATM mutant mice, noted a GSH-homeostasis defect caused by an impaired ability to import cystine83. It was assumed this decrease was due to a downregulation of xCT levels, a component of the xc– transporter system. It will be interesting to determine if the mechanism we identified is also the basis of the observed defect in that tissue, as this could relate to the ataxia phenotype of A-T.
CD98HC heterodimerizes with several partner proteins, particularly HATs (SLC5-11,13) and facilitates their surface localization84. We showed that ATM impacts the activity of at least two HATs by phosphorylating CD98HC. Based on the impact of ATM status on CD98HC, it is likely that there would be consequences to other HATs as an alternative explanation for at least some of the chronic progressive phenotypes associated with A-T. For instance, (i) lymphocyte proliferation and differentiation, depends on glutamate metabolism and CD98/xc- activity85,86 (A-T patients are immunodeficient due to a lack of T-cells); (ii) ataxia, as mutations in several CD98HC partner proteins result in ataxia (mutation of SLC7A9 or SLC7A10 results in ataxia, while SLC7A8 mutation results in impaired motor performance and the few SLC7A7-null mice that survive have tremors87,88,89,90,91); (iii) ASC-1 (SLC7A10), participates in L-serine uptake and regulates glycine levels in the brain, important for Purkinje cell survival and dendrite growth92, a well-known consequence of A-T, while its loss causes tremors and seizures91; and (iv) growth, as y+L transports thyroid hormone through the blood-brain barrier (A-T patients often have slightly reduced stature)87. Considering these speculations, it would be interesting to assess the impact of ATM-CD98HC interaction in neurons, ideally using primary human or hiPSC-derived neurons, especially considering that SLC3A2 expression is about 3 times as high in mouse cerebellum than human (93, proteinatlas.org). Overall, our findings indicate alternative directions to explore that may lead to effective interventions to treat at least some aspects of this syndrome. Finally, considering that ATM is frequently mutated, deleted, or silenced in various cancers, a defect in amino acid (cysteine/glutamate) metabolism may also offer new therapeutic targets that could be explored in those contexts.
Methods
Experimental model and subject details
Mice
C57BL/6J Atm+/– (Atmtm1Awb) mice were described previously41,94. These mice were backcrossed onto C57BL/6J mice for over fifteen generations94. Genotypes for the Atm allele were determined by PCR amplification as described in ref. 75. The mice were housed in a pathogen-free barrier facility and all animal experiments were conducted as approved by UT Health San Antonio (UTHealth) IACUC policy and outlined in our IACUC protocol number 07005x. The facility is operated in compliance with Public Law 89–544 (Animal Welfare Act) and its amendments, Public Health Services Policy on Humane Care and Use of Laboratory Animals (PHS Policy), using the Guide for the Care and Use of Laboratory Animals (Guide) as the basis of operation. The University has been accredited by the Association for Assessment and Accreditation of Laboratory Animal Care, International. Atm+/– female mice were crossed with Atm+/− males and were given free access to drinking water with or without 40 mM NAC (pH 7.0–7.4, Sigma) throughout pregnancy, lactation, and thence upon weaning. Both, regular and NAC-supplemented water were changed weekly. After weaning, the treatment group received the same 40 mM NAC-supplemented drinking water until 6 months of age when tests were carried out. All mice were maintained on a normal diet, irradiated LM-485 diet (Inotiv, 7912: 19% protein, 5% fat, 5% fiber), food and water provided ad libitum. Animals were euthanized by CO2 inhalation followed by cervical dislocation.
Human tissue
De-identified postmortem liver tissue of a 19- and a 28-year-old A–T patient was obtained from the NICHD Brain and Tissue Bank of Developmental Disorders at the University of Maryland, Baltimore, MD which acquires and distributes patient samples with IRB approvals 5-58 and HM-HP-00042077.
Cell culture
Three different isolates of HUVEC were purchased from GIBCOTM. Cells between passages 3–7 were used for experiments. Cells were grown in Medium 200 (GIBCOTM) supplemented with 2% Low Serum Growth Supplement (GIBCOTM) and 1% PS. Three days before the experiments medium was progressively changed to Medium 199 (Earle’s Salts, GIBCOTM) supplemented with 1 mM sodium pyruvate, 1.82 mM glutamine, 10 mM HEPES, 2% LSGS, 8% FBS, and 1% PS (to keep physiologically relevant glucose and glutamine levels, 5.5 mM, and 2 mM respectively). 16–20 h before experiments, cells were washed twice with HBSS (Corning) and the medium was changed to low serum (2% LSGS only). HUVECs were kept at 3% oxygen and 37 °C in a humidified atmosphere with 5% CO2.
Primary MEFs were obtained by intercrossing Atm heterozygous mice to obtain Atm–/– embryos and littermate controls and isolated as previously described95. MEFs were grown in DMEM (10-013-CV, Corning) supplemented with 10% FBS, and 1% PS and kept at 3% oxygen.
Mouse αTC1 Clone 9 pancreatic cells, Mouse β-TC6 pancreatic cells, and HEK-293 cells were obtained from ATCC and grown in DMEM supplemented with 10% FBS, and 1% PS. Alpha pancreatic cells medium was further supplemented with 0.02% BSA, 100 µM non-essential amino acids, and 10 mM HEPES. Three days before experiments the medium was changed to Basal Medium Eagle (BME, GIBCOTM) containing 10% FBS, 5.5 mM Glucose, 1 mM Sodium Pyruvate, and 2 mM Glutamine. For alpha cells, 0.02% BSA, 100 µM NEAA, and 10 mM HEPES were also added. 16-20 h before experiments, cells were washed twice with HBSS and media changed to 2% FBS-BME. Alpha, beta, and HEK-293 cells were maintained at 37 °C in a humidified atmosphere with 5% CO2 and tested for mycoplasma contamination.
Method details
Metabolic assays
Mitochondrial respiration and glycolytic function were assayed by measuring the oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) in HUVEC, alpha, and beta pancreatic cells using a Seahorse XFe96 Analyzer (Agilent Technologies). Experiments were run using XF Base Medium Minimal DMEM (Agilent Technologies) supplemented with 1 mM Sodium pyruvate, 5.5 mM Glucose, and 2 mM Glutamine (pH 7.4). Injections were prepared following the manufacturer’s protocol for Mito Stress Kit and Glycolysis Stress Kit (Agilent technologies). For the modified Mito Fuel Flex test, the first injection consisted of the drug targeting either glucose, glutamine, or fatty acids oxidation followed by regular injections in the Mito Stress Kit. OCR/ECAR were normalized to confluence using an IncuCyte®ZOOM phase-only processing module (essenBioscience).
ROS levels analysis
Intracellular ROS levels were assayed by ROS-GloTM H2O2 Assay (Promega) and CellROX® oxidative stress assay (Life Technologies). For ROS-GloTM, HUVECs were treated with ATMi (10 µM KU55933/ 5 µM KU60019, Apexbio) for 8 h and luminescence was measured following the manufacturer’s protocol. For CellROX assay, cells were treated with KU60019 for 8, 24, and 48 h. Treatment with stabilized H2O2 (200 µM, Sigma) was used as a positive control of ROS generation. 1 h before collecting cells, they were stained with 5 µM of CellROX® Reagent adding it directly to the media. Cells were then washed with 1X PBS, trypsinized, and fixed with 4% paraformaldehyde (PFA). Propidium iodide (PI) was used to assess viability. Cells were analyzed with the cell analyzer BD LSRFortessa™ X-20. The gating strategy is presented in Fig. S7.
Viability assays
Cells were seeded at 30% confluence in 96- or 384-well plates. The next day, cells were treated with the inhibitors (5 µM KU60019, 10 µM KU55933, and 5 µM Erastin (Sigma)) and cell viability was evaluated after 96 h using CellTiter-Glo. Confluency curves were generated using the IncuCyte®ZOOM phase-only processing module (essenBioscience).
Stable isotope resolved metabolomics (SIRM) experiments
HUVECs were seeded in 100 mm plates using supplemented M199 medium (Mybiosource.com). On the day of the experiment, the medium was changed to one containing either 5.5 mM [U-13C]- glucose and unlabeled Gln or 2 mM [U-13C,15N]-glutamine with 5.5 mM unlabeled glucose and treated with 10 µM KU55933 for 8 h. Isotope-enriched substrates were purchased as dry powders from Cambridge Isotope Laboratories, MA. Polar and non-polar metabolites were extracted from cells and media following metabolic quenching in cold acetonitrile and harvesting for metabolite extraction as described previously96. The polar extracts were reconstituted in nanopure water before analysis on a Dionex ICS-5000+ ion chromatography interfaced to a Thermo Fusion Orbitrap Tribrid mass spectrometer (Thermo Fisher Scientific) as previously described97 using a m/z scan range of 80–700. Peak areas were integrated and exported to Excel via the Thermo TraceFinder (version 3.3) software package before natural abundance correction98. The isotopologue distributions of metabolites were calculated as the mole fractions as previously described99. The number of moles of each metabolite was determined by calibrating the natural abundance-corrected signal against that of authentic external standards. The amount was normalized to the amount of extracted protein and is reported in nmol/mg protein.
Polar extracts reconstituted in D2O (>99.9%, Cambridge Isotope Laboratories, MA) containing 17.5 nmol d6-2,2-dimethyl-2-silapentane-5-sulfonate (DSS) as internal standard were analyzed by 1D 1H and 1H{13C} -HSQC NMR on a 14.1 T DD2 NMR spectrometer (Agilent Technologies, CA). 1D 1H spectra were acquired using the standard PRESAT pulse sequence with 512 transients, 16384 data points, 12 ppm spectral width, an acquisition time of 2 s and a 6 s recycle time with weak irradiation on the residual HOD signal during the relaxation delay. The raw fids were zero filled to 131,072 points and apodized with 1 Hz exponential line broadening prior to Fourier transformation. 1D HSQC spectra were recorded with an acquisition time of 0.25 s with GARP decoupling and recycle time of 2 s over a spectral width of 12 ppm, with 1024 transients. The HSQC spectra were then apodized with unshifted Gaussian function and 4 Hz exponential line broadening and zero filled to 16k data points before Fourier transformation. Metabolites were assigned by comparison with in-house100 and public NMR databases. Metabolite and their 13C isotopomers were quantified using the MesReNova software (Mestrelab, Santiago de Compostela, Spain) by peak deconvolution. The peak intensities of metabolites obtained were converted into nmoles by calibration against the peak intensity of DSS (17.5 nmoles) at 0 ppm for 1H spectra and that of phosphocholine at 3.21 ppm (nmoles determined from 1D 1H spectra) for HSQC spectra before normalization with mg protein in each sample.
Metabolite analysis
Intra- and extra-cellular levels of glutamate were measured with Glutamate assay Kit (Sigma-Aldrich) and Amplex Red Glutamic Acid Assay kit (Molecular probes), respectively. For each experiment, cells were seeded in 6-well plates (HUVECs) and 12-well plates (alpha and beta pancreatic cells) and treated with either 10 µM KU55933 or 5 µM KU60019 for 8 h. Harvesting and measurements were performed following the manufacturer’s protocol. Values were normalized to protein concentration. Both GSH-GloTM Glutathione Assay and GSH/GSSG-GloTM Assay (Promega) were used to measure total, reduced, and oxidized GSH after drug treatments. NADP/NADPH-GloTM Assay (Promega) was performed in HUVECs following the manufacturer’s protocol.
Total RNA isolation and sequencing
HUVECs were treated with either 10 µM KU55933 or 5 µM KU60019 for 8 h. RNA was isolated using the RNeasy Mini Kit (Qiagen). Approximately 500 ng of total RNA was used for RNA-seq library preparation by following the Illumina TruSeq stranded mRNA sample preparation guide. The first step in the workflow involves purifying the poly-A-containing mRNA molecules using poly-T oligo-attached magnetic beads. Following purification, the mRNA is fragmented using divalent cations under elevated temperatures. The cleaved RNA fragments are copied into first strand cDNA using reverse transcriptase and random primers. This is followed by second strand cDNA synthesis using DNA Polymerase I and RNase H. Strand specificity is achieved by replacing dTTP with dUTP in the Second Strand Marking Mix (SMM). These cDNA fragments then go through an end repair process, the addition of a single (A) base, and then ligation of the adapters. The products are then purified and enriched with PCR to create the final RNA-seq library. After RNA-seq libraries were subjected to quantification process, pooled for cBot amplification and subsequent 50 bp single-read sequencing run with Illumina HiSeq 3000 platform. After the sequencing run, demultiplexing with Bcl2fastq2 was employed to generate the fastq file for each sample with about 35–40 million reads per sample.
Lentiviral infection
Lentiviral production was performed following previously published work101. Briefly, HEK-293 cells were co-transfected with packaging plasmids pM2.G and psPAX2 (Addgene) and ATM shRNA or Control shRNA (Santacruz) using Lipofectamine 2000 (Life Technologies). After 8 h, the medium was changed and supplemented with 0.5% bovine serum albumin (Sigma) to improve virus stability. After 60 h, viral supernatants were recollected, centrifuged at 300 g at 4 °C for 10 min, and filtered through a 0.45 µm low protein-binding membrane (Millipore). Single-use aliquots were made and stored at −80 °C. HUVEC, alpha, and beta pancreatic cells were transduced with the Lentiviral particles when they reached 70% confluence. 10 µg/mL Polybrene was added to improve transduction efficiency, and the medium was changed after 8 h. The next day, the medium was supplemented with 1 µg/mL puromycin (GIBCOTM). Cells were growing with puromycin for at least 5 days before running experiments. Successful transduction was confirmed by the detection of ATM (Sigma) by western blot.
Immunofluorescence and proximity ligation assays (PLA)
Cells were plated on coverslips pre-coated with 1% gelatin. HUVECs were treated with 10 µM KU55933 for 8 h and addition of 200 µM H2O2 for the last 2 h. Cells were fixed with 4% PFA and permeabilized with 0.5% Tween in PBS at room temperature (RT). After incubation in blocking buffer (5% bovine serum albumin in PBS), primary antibody (Rabbit anti-ATMphospho S1981) was added for 4 h at RT in a moist chamber. Secondary antibody (Alexa Flour donkey anti-rabbit 488 IgG) and Hoechst 33342, were added for 1 h, and cells were washed and covered with Prolong Gold antifade reagent (Thermo Fisher). For the PLA (Sigma), cells were either treated with 5 µM KU60019 (8 h) or transduced with shATM lentiviral particles (Santa Cruz Biotechnologies) beforehand. The PLA was performed following the manufacturer’s protocol. Primary antibodies used were goat anti-ATM (Sigma) and rabbit anti-CD98 (Cell signaling). Images were recorded on a Confocal Laser Scanning Microscope (Olympus FV3000). Fluorescent images were acquired in a scan format of 1024 × 1024 pixels in a spatial data set (xyz) and were processed with Image J software. Controls without primary antibodies showed no fluorescence labeling.
Immunoblotting and immunoprecipitation
Besides ATMi, HUVECs were treated with ATR (2 µM AZD6738), DNAPKs (1 µM AZD7648), AKT (3 µM MK2206) inhibitors for 8 h and 200 µM H2O2 was added for the last 2 h. Whole-cell lysates were prepared using NaCl Lysis buffer (20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM Na2EDTA, 1 mM EGTA, 1% Triton, 2.5 mM sodium phosphate, 1 mM ß-glycerophosphate) containing a protease inhibitor cocktail (Roche/Sigma) and phosphatase inhibitor (Thermo Fisher) following standard methods. 40–50 µg of total protein was loaded in either precast 3–8% gradient gels (Invitrogen) or laboratory-prepared gels and transferred onto a nitrocellulose membrane. All blots were incubated with primary antibodies overnight and developed using enhanced chemiluminescence (ECL, ThermoFisher). Antibodies used in this study include ATMphosphoS1981 (Abcam), ATM (Sigma and Santa Cruz), CD98phosphoS103, CD98 (Cell signaling), CD98 (H-300 Santa Cruz), ß-ACTIN (Abcam), GAPDH, LAMIN-2, MEK1/2, VINCULIN, AKTphosphoS473, AKT, and CHK1phosphoS347 (Cell signaling), DNA PKcs-phosphoS2056 and DNAPKs (Abcam). Co-immunoprecipitation experiments were done with endogenous proteins as described before102. In brief, cells from nearly confluent 15-cm plates treated with 200 µM H2O2 for 30 min were harvested and lysed in IP buffer (20 mM HEPES, 150 mM NaCl, 2 mM MgCl2, 0.5 mM CaCl2, 1% (v/v) Brij-35, and protease inhibitor cocktail). The lysates were incubated at 4 °C with gentle rotation for 1 h, centrifuged at 12,000 g for 10 min and the supernatants were used for immunoprecipitation. Lysates were pre-cleared using 20 µL of protein A/G dynabeads (Invitrogen) for 30 min at 4 °C. 650 μg total cell lysates were incubated with 4–8 μg of the appropriate primary antibody overnight at 4 °C. The next day, 50 µL of protein A/G dynabeads were added to the lysates and incubated for 2 h at 4 °C. Beads were washed three times in IP buffer and bound proteins were eluted by boiling the beads in NuPAGE sample buffer under reducing conditions. Eluted proteins were evaluated by immunoblotting and compared to inputs (10% of the amount used for immunoprecipitation). Co-immunoprecipitation experiments were repeated with biological replicates at least three times in independent sample preparations.
To assay the glycosylation status of CD98HC, HEK293T cells were treated with or without tunicamycin (Sigma-Aldrich, #T7765) in regular maintenance medium for 24 h (0.5 and 1 µg/mL) or 8 h (5 µg/mL). Cells were then harvested and lysed in IP buffer and western blots were performed as previously described.
[14C]-cystine and -arginine uptake
Amino acid uptake assays were performed as previously described with some minor modifications103,104. Briefly, HUVECs were seeded in 12-well plates. When they reached nearly 90% confluence 8 h treatments were initiated adding DMSO, 10 µM KU55933, 5 µM KU60019, or 400 µM Sulphasalazine (SAS). For ATM+/+ and ATM–/– derived MEFs, cells were seeded in two 24-well plates and left to grow for about 40 h when SAS treatment was started. 4 h later, the medium in both plates was removed and HBSS was added during the ionizing radiation treatment (0.5 Gy) using the Faxitron X-ray 43855F (Bookholt Associates). Cells were fed with fresh medium and returned to the incubator for 1 h before harvesting. At the end of the treatments, cells were washed three times with pre-warmed (37 °C) transport medium (137 mM NaCl, 0.7 mM K2HPO4, 1 mM CaCl2, 1 mM MgCl2, 5 mM glucose, 10 mM HEPES, pH 7.4). Then, the transport medium was replaced with 400 μL transport buffer containing treatments and [14C] L-cystine (PerkinElmer) or [14C] Arginine (PerkinElmer) at a final concentration of 5 μM and 20 μM/well, respectively. Cells were incubated at 37 °C for 10 min. Amino acid transport was stopped by placing the plate on ice. HUVECs were then washed three times with ice-cold transport medium and lysed in 200 µL 0.1 M NaOH. Radioactivity in lysates was measured by liquid scintillation counting and normalized to the quantity of protein in lysates as determined using the Pierce BCA (bicinchoninic acid) Protein Assay Kit (Thermo Fisher).
CD98 PhosphoS103 monoclonal antibody production and validation
Bioinformatics analysis of CD98HC identified a putative site in its cytoplasmic tail (CGTMSQDTEVDMK), which was previously reported to be phosphorylated in prior phosphoproteome screens for ATM-dependent phosphorylation events105. The project was submitted to BIOMATIK to generate the respective phospho-specific antibody. Validation of CD98phosphoS103 antibody was done by showing the specificity of designed primers (dot bot) and by treating lysates with alkaline phosphatase, Calf Intestinal (CIP, New England Biolabs) followed by immunoblotting. For the first approach, designed phosphopeptides (Biomatik) were diluted into 5, 10, and 20 µg/mL in PBS (pH 7.4). 100 µL per dilution was loaded onto a pre-wet H+ nylon membrane. The membrane was washed twice with dH2O and then left to air dry at room temperature. The membrane was then blocked with 1X TBST containing 5% non-fat dry milk for 1 h and incubated with our customized primary antibody for CD98phosphoS103 (Biomatik) in the blocking buffer for 2.5 h. Anti-mouse IgG-HRP was incubated for 45 min and blots were developed using enhanced chemiluminescence (ECL, ThermoFisher) and the Odyssey® FC imaging system (LI-COR Biosciences).
Subcellular fractionation
Subcellular fractions were purified using NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific). HUVECs were seeded in 10 cm plates and treated with 200 µM H2O2 for 2 h. Cells were harvested following the manufacturer’s procedure. Enriched fractions were blotted against anti-CD98phosphoS103, CD98 (Cell Signaling), pATM (Abcam), ATM (Sigma-Aldrich), xCT (Abcam), LAT-2 (Abcam). Anti-LaminB1 and MEK1/2 (Cell Signaling) were used as nuclear and cytoplasmic markers, respectively. Anti-CD98 (Cell Signaling) was used as a membrane marker. Blots were developed using enhanced chemiluminescence (ECL, ThermoFisher) and the Odyssey® FC imaging system (LI-COR Biosciences). The experiment was performed three times for independent validation.
Photoconversion assay for in vivo protein tracking
First, to create the phospho-dead (PD) mutant, the A and G at positions 307 and 308 of SLC3A2 were mutated to G and C, respectively, which converted the serine to alanine. Mutagenesis was performed in the plasmid CD98 (SLC3A2) (NM_002394) Human Tagged ORF Clone (Origene, RG216640) using the QuickChange II sytem (Qiagen), the primers used were: acctcggtgtcctgggccatggtgcctgtaac and gttacaggcaccatggcccaggacaccgaggt. Positive clones were verified by Sanger sequencing (Genewiz, Azenta Life Sciences).
Next, the photoconvertible protein mEos3.2 (kindly provided by Dr. Lechleiter) was inserted in the N-terminus of both wild-type and phospho-dead SLC3A2 CDS by PCR using the following primers: (i) NM_002394_F: atggagctacagcctcctgaagcctcgatc; (ii)PmeI_Stop_NM_002394_R: cgcggccggccgtttaggccgcgtaggggaagcggagcagcagccc. The fragment coding for mEos3.2 (mEos3.2_1xHis_STOP_pET28a,) was extracted using the following primers: (i) BamHI_mEos32_F: ttcgtcgactggatcatgagtgcgattaagccagacatgaagatc; (ii) NM_002394_mEos32_R: aggctgtagctccattcgtctggcattgtcaggcaatccagaatg. Next, SLC3A2 and GFP were removed from the Origene plasmid using BamHI-HF (NEB, R3136) and PmeI (NEB, R0560) double enzyme digest, and this backbone was used to insert the two PCR products (SLC3A2 and mEos3.2) using the In-fusion HD Cloning Kit (Takara # 638909).
Positive clones were verified by Sanger sequencing (Genewiz, Azenta Life Sciences).
For the transfection and photoconversion, HEK293T cells were cultured on 35 mm glass bottom plates (MatTek # P35GC1.514 C) and transfected with 1 µg of wild-type or PD plasmid using Lipofectamine 3000 (Invitrogen). 24 h after transfection, cells were exposed to a 405 nm laser for 3 min to photoconvert mEos3.2, which normally emits green fluorescence and after exposure will also emit red fluorescence. Cells were treated with CellBrite® Steady Membrane Staining (Biotium) and images from live cells were captured 24 h after the photoconversion on an Olympus FLUOVIEW™ FV3000 confocal microscope.
Angiogenesis and wound healing assays
24-well plates were coated with 92 µL of Geltrex®Matrix and incubated at 37 °C for 30 min. HUVECs were pre-treated with 5 µM KU60019, 2 mM NAC, or 5 mM L-Arginine ethyl ester (LAEE, Sigma) for 4 h. The angiogenesis assay was initiated by seeding 75,000 cells/well in the pre-coated plates and left to form endothelial vessel networks over 16 h maintaining respective drug treatments. Images were taken using the IncuCyte®ZOOM phase-only processing module (essenBioscience). Images were processed with the software Image J and analyzed with the angiogenesis tool. The selected parameters were: Number of meshes; Total master segment length, the sum of the length of the detected master segments; Mesh index, the mean distance separating two master junctions in the trees; Number of nodes; and Number of master segments. 20 images were analyzed per condition. Experiments were performed in duplicate and repeated twice for independent validation. For the wound healing assay, cells were seeded the day before and scratched performed using the WoundMakerTM instrument (essenBiosience). Cells were washed twice with HBSS and treatments were initiated at this point. Images were taken after 12 h using the IncuCyte®ZOOM and quantified using Image J.
Insulin and glucagon ELISA
Alpha and beta pancreatic cells were seeded in 96-well plates and treated with either ATMi. ATM null cells obtained from lentiviral transduction were also used in this experiment. Insulin secretion by beta cells was measured using the Mouse Ultrasensitive Insulin ELISA (ALPCO Diagnostics). Glucagon secretion was measured by using the Quantikine®ELISA-Glucagon (R&D Systems) following the manufacturer’s instructions. Briefly, after treatments supernatant was recollected and centrifuged at 300 g at 4 °C for 10 min to eliminate cell debris. Dilutions of 1:100 and 1:10 were used for insulin and glucagon kits, respectively. The cell lysate was collected in lysis buffer and used to normalize values to protein concentration.
Pancreatic islet isolation and islet bioenergetics
Islet isolation was described previously106. Briefly, islets were isolated from 6-month-old C57BL/6 J Atm+/+ and Atm–/– mice by collagenase XI (Sigma-Aldrich) perfusion and Histopaque (Sigma-Aldrich) separation from acinar and ductal tissues. Islets were then handpicked and cultured overnight in RPMI 1640 plus 10% FBS at 37 °C and 5% CO2 before performing assays. Glucose response and bioenergetic studies were performed with pooled islets (50-80 islets/well), using a Seahorse XF24 Analyzer (Agilent Technologies) as previously described107. At the end of the experiment, islets were lysed in 30 µL of lysis buffer with a protease inhibitor cocktail (Roche/Sigma), and protein concentration was measured using the BCA protein assay (Thermo Fisher). Protein content was used to normalize Seahorse OCR.
Glucose and insulin tolerance test
Mice were kept on a normal chow diet and underwent glucose and insulin tolerance tests at 6 (males) or 12 months (females) of age. Mice were fasted for 14 h and injected intraperitoneally with either glucose (2 g/kg) or insulin (0.75 U/kg). For the oral glucose tolerance test mice were fed 45% glucose solution via oral gavage. Plasma concentrations of insulin were measured using the Mouse Ultrasensitive Insulin ELISA (ALPCO Diagnostics). Tail vein blood glucose was measured using an automated glucometer (Bionime/CVS Glucose Meter) at indicated times.
Immunohistochemistry and Oil Red O staining
Mice pancreas was fixed in 10% formalin and embedded in paraffin. 4µm-thick sections were deparaffinized, rehydrated, and treated either with 1 mM EDTA pH 8 for 40 min at 95 °C followed by a 20-min cool-down step or with citrate pH 6 for 30 min at 98–100 °C followed by a 30-min cool-down step. Slides were then rinsed in 1X Tris-buffered saline (TBS) three times. Following endogenous peroxidase blocking, the slides were incubated with primary antibodies (Glutamate (LS Bio), glutamine (Abcam), glucagon, insulin, and CD98 (Cell signaling)) for 2 h at room temperature in a moist humidity chamber. Biotinylated anti-rabbit secondary antibodies (1:200, Vector Laboratories) were incubated for 60 min at room temperature after slides were washed in 1X TBS three times. Slides were incubated in ABC-HRP complex (Vector Laboratories) for 30 min. To confirm antibody specificity, one slide was incubated with secondary antibody only. Slides were then developed with DAB for 5 min, rinsed with TBS, counterstained with hematoxylin, dehydrated, cleared, and mounted with a synthetic mounting medium. Oil Red O staining was performed in frozen livers from 6-month-old C57BL/6J Atm+/– mice, 4–5-month-old 129Sv/C57B6 Atm+/– (kind gift of Zha Lab), and Ataxia-telangiectasia donors (University of Maryland Brain & Tissue Bank). Images were taken on a Motic Digital Slide Scanning System at 20X and 40X magnification.
Additional information on reagents and resources used to perform experiments are included in Table S6.
Quantification and statistical analysis
Photoconversion image analysis and quantification
CD98HC protein localization to the membrane was measured by taking three cross sections per cell using the straight-line feature in Image J, spanning the length of the cell. CD98HC fluorescence signal corresponding to the peak membrane fluorescence was extracted from the ImageJ plot profiles of each of the three cross sections. The gray values across the pixel distance of each cross-section were binned into 100 equal parts using RStudio. The mean fluorescence intensity from all three cross sections was averaged for each bin for a total of 100 normalized fluorescence intensity values per cell. The CD98HC membrane signal corresponded to bins 1–5 and 95–100, and the intracellular signal to bins 6–94.
Immunohistochemistry quantification
Image J software was used to assess the microscopic analysis. First, the whole tissue section was exported, and hematoxylin staining was separated using color deconvolution. Primary antibody intensity was measured in each pancreatic islet (average intensity/islet area) and normalized to the surrounding background. To measure islet size, insulin staining was used to identify the islets. Glucagon staining was quantified in each islet and expressed as a percentage of positive area/pancreatic islet area. Oil Red O (ORO) staining was quantified and expressed as a percentage of the positive area in the entire pancreatic section.
Statistics
Data analyses were performed using GraphPad Prism 9.4.1 and 10. Flow cytometry data were analyzed using FlowJoTM 10. p-values were provided in the corresponding figure legend and calculated using one- or two-way ANOVA. For non-parametric data, Mann-Whitney or Kruskal-Wallis tests were performed. p < 0.05 was considered significant: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. When not specified, asterisks show significance vs. control. Mice were randomly assigned for the in vivo studies; the sample size is provided in the figure legends. Measurements from all experiments were taken from distinct samples.
Bioinformatics analysis
Correlation analysis
Re-processed RNA-Sequencing counts and metadata were generated by the authors of the ARCHS4 repository as described in their publication108. Counts and metadata were downloaded from the ARCHS4 repository and pre-processed with custom R scripts. Samples were categorized using a manually curated dictionary of disease-related regex terms, allowing isolation of normal tissue samples. Count data filtering followed a five-step procedure: (1) scRNA-Seq samples were identified using a custom regex dictionary and removed because of the demonstrated unsuitability of single-cell data for co-expression network inference by Pearson correlation109. (2) Samples with fewer than 5 million raw read counts were discarded to improve the quality of our gene co-expression calculations by reducing noise from low-quality samples110. (3) Genes with zero raw counts in 10% or more of samples were removed to further reduce noise; an approach based on the recommendations of WGCNA authors111. (4) For each tissue-disease group, studies with only one sample were removed due to the inability to calculate batch effects in these samples112. (5) Tissue-disease groups with fewer than 30 distinct samples or with fewer than four different studies by GEO series accession (GSE) were removed to limit the effects of bias from individual samples or studies and improve the performance of co-expression calculations110. Count data was normalized and batch-corrected following the procedure outlined by the ARCHS4 authors108: Raw counts were log2(x + 1) transformed and subsequently quantile normalized using the normalize.quantiles function from the preprocessCore R package113. Then, batch effects were removed based on a study with the ComBat function of the sva R package112. Then, gene-gene Pearson correlations were calculated using the cor function of the WGCNA package111. Correlations for ATM were used as the ranking criteria for Gene Set Enrichment Analysis (GSEA) implemented via the Cluster Profiler R package’s GSEA function and visualized with their gseaplot function using annotations gathered via the msigdbr package’s msigdbr function114,115.
Single-cell RNA-Seq of pancreas tissue
A comprehensive single-cell RNA-Seq dataset of human pancreas tissue was generated via the analysis methods described by the authors of Seurat in their data integration guide using the data objects conveniently provided for download on the Seurat web page116,117. The data provided for download was derived from three single-cell studies of pancreatic tissue: GSE81076/GSE85241118, GSE86469119, and E-MTAB-5061120. The expression of SLC3A2 and ATM was plotted across cell types (as was determined by the Seurat authors) in the form of ridge plots using the RidgePlot function of Seurat. The expression of ATM and SLC3A2 was quantile normalized using the normalizee.quantiles function of the preprocessCore package113. Then normalized ATM and SLC3A2 expressions were compared using the stat_compare_means function of the ggpubr package across cell types and plotted using the ggboxplot function121.
Bulk RNA-Seq
Raw data were processed into fastq files using the bcl2fastq software program (Illumina). Fastq files were trimmed for adapter sequences using the fastp software program122. Trimmed fastq files were aligned to the GRCh38 transcriptome (Gencode v30) using the salmon aligner software123. Read counts were summarized to the gene level using the tximport R package124. Differential gene expression was calculated using the DESeq2 R package, and log-transformed p-adjusted values (FDR) were used as the ranking criteria for Gene Set Enrichment Analysis implemented via the Cluster Profiler package’s GSEA function and visualized with their gseaplot function using annotations gathered via the msigdbr package’s msigdbr function114,115.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data used for the single-cell RNA-Seq dataset of human pancreas tissue was derived from the following single-cell studies of pancreatic tissue: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE85241; https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE81076; https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE86469; https://www.ebi.ac.uk/biostudies/arrayexpress/studies/E-MTAB-5061. The fastq files and read counts for all bulk RNA-Sequencing runs have been deposited in NCBI GEO under accession GSE140416. Any additional requests for information can be directed to the corresponding author. Source data are provided with this paper.
References
Gatti, R. A. et al. Localization of an Ataxia-telangiectasia gene to chromosome 11q22–23. Nature 336, 577–580 (1988).
Chun, H. H. & Gatti, R. A. Ataxia–telangiectasia, an evolving phenotype. DNA Repair 3, https://doi.org/10.1016/j.dnarep.2004.04.010 (2004).
Alexander, A. et al. ATM signals to TSC2 in the cytoplasm to regulate mTORC1 in response to ROS. Proc. Natl. Acad. Sci. USA 107, https://doi.org/10.1073/pnas.0913860107 (2010).
Kozlov, S. V. et al. Reactive oxygen species (ROS)-activated ATM-dependent phosphorylation of cytoplasmic substrates identified by large-scale phosphoproteomics screen. Mol. Cell. Proteom. 15, https://doi.org/10.1074/mcp.M115.055723 (2016).
Paull, T. T. & Paull, T. T. Mechanisms of ATM activation. Annu. Rev. Biochem. 84, https://doi.org/10.1146/annurev-biochem-060614-034335 (2015).
Khanna, K. K. & Lavin, M. F. Ionizing radiation and UV induction of p53 protein by different pathways in ataxia-telangiectasia cells. Oncogene 8, 3307–3312 (1993).
Matsuoka, S. et al. Ataxia telangiectasia-mutated phosphorylates Chk2 in vivo and in vitro. Proc. Natl. Acad. Sci. USA 97, https://doi.org/10.1073/pnas.190030497 (2000).
Valentin-Vega, Y. A. et al. Mitochondrial dysfunction in ataxia-telangiectasia. Blood 119, 1490–1500 (2012).
Tripathi, D. N., Zhang, J., Jing, J., Dere, R., & Walker, C. L. A new role for ATM in selective autophagy of peroxisomes (pexophagy). Autophagy 12, https://doi.org/10.1080/15548627.2015.1123375 (2016).
Cosentino, C., Grieco, D., Costanzo, V. ATM activates the pentose phosphate pathway promoting anti-oxidant defence and DNA repair. EMBO J. 30, https://doi.org/10.1038/emboj.2010.330 (2011).
Andrisse, S. et al. ATM and GLUT1-S490 phosphorylation regulate GLUT1 mediated transport in skeletal muscle. PloS ONE 8, https://doi.org/10.1371/journal.pone.0066027 (2013).
Kamsler, A. et al. Increased oxidative stress in ataxia telangiectasia evidenced by alterations in redox state of brains from Atm-deficient mice. Cancer Res. 61, 1849–1854 (2001).
Gueven, N. et al. Dramatic extension of tumor latency and correction of neurobehavioral phenotype in Atm-mutant mice with a nitroxide antioxidant. Free Radic. Biol. Med. 41, https://doi.org/10.1016/j.freeradbiomed.2006.06.018 (2006).
Ito, K. et al. Regulation of oxidative stress by ATM is required for self-renewal of haematopoietic stem cells. Nature 431, https://doi.org/10.1038/nature02989 (2004).
Ito, K. et al. Regulation of reactive oxygen species by Atm is essential for proper response to DNA double-strand breaks in lymphocytes. J. Immunol. 178, https://doi.org/10.4049/jimmunol.178.1.103 (2007).
Chen, P. et al. Oxidative stress is responsible for deficient survival and dendritogenesis in Purkinje neurons from ataxia-telangiectasia mutated mutant mice. J. Neurosci. 23, https://doi.org/10.1523/JNEUROSCI.23-36-11453.2003 (2003).
Erker, L. et al. Cancer chemoprevention by the antioxidant tempol acts partially via the p53 tumor suppressor. Hum. Mol. Genet. 14, https://doi.org/10.1093/hmg/ddi181 (2005).
Schubert, R. et al. Cancer chemoprevention by the antioxidant tempol in Atm-deficient mice. Hum. Mol. Genet. 13, https://doi.org/10.1093/hmg/ddh189 (2004).
Reliene, R., Fischer, E. & Schiestl, R. H. Effect of N-acetyl cysteine on oxidative DNA damage and the frequency of DNA deletions in atm-deficient mice. Cancer Res. 64, 5148–5153 (2004).
Reliene, R. & Schiestl, R. H. Antioxidant N-acetyl cysteine reduces incidence and multiplicity of lymphoma in Atm deficient mice. DNA repair 5, 852–859 (2006).
Kelly, G. S. Clinical applications of N-acetylcysteine. Alter. Med. Rev. 3, 114–127 (1998).
Batinić-Haberle, I., Rebouças, J. S., Spasojević, I. Superoxide dismutase mimics: chemistry, pharmacology, and therapeutic potential. Antioxid. Redox Signal. 13, 877–918 (2010).
Krishna, M.C. et al. Stimulation by nitroxides of catalase-like activity of hemeproteins. Kinetics and mechanism. J. Biol. Chem. 271, https://doi.org/10.1074/jbc.271.42.26018 (1996).
Browne, S. E. et al. Treatment with a catalytic antioxidant corrects the neurobehavioral defect in ataxia-telangiectasia mice. Free Radic. Biol. Med. 36, https://doi.org/10.1016/j.freeradbiomed.2004.01.003 (2004).
Erker, L. et al. Effect of the reduction of superoxide dismutase 1 and 2 or treatment with α-tocopherol on tumorigenesis in ATM-deficient mice. Free Radic. Biol. Med. 41, 590–600 (2006).
Lewerenz, J. et al. The cystine/glutamate antiporter system x(c)(-) in health and disease: from molecular mechanisms to novel therapeutic opportunities. Antioxid. Redox Signal. 18, 522–555 (2013).
Bannai, S., Tateishi, N. Role of membrane transport in metabolism and function of glutathione in mammals. J. Membr. Biol. 89, https://doi.org/10.1007/BF01870891 (1986).
Mastroberardino, L. et al. Amino-acid transport by heterodimers of 4F2hc/CD98 and members of a permease family. Nature 395, 288–291 (1998).
Pineda, M. et al. Identification of a membrane protein, LAT-2, that Co-expresses with 4F2 heavy chain, an L-type amino acid transport activity with broad specificity for small and large zwitterionic amino acids. J. Biol. Chem. 274, 19738–19744 (1999).
Yan, R., Zhao, X., Lei, J. & Zhou, Q. Structure of the human LAT1–4F2hc heteromeric amino acid transporter complex. Nature 568, 127–130 (2019).
de la Ballina, L. R. et al. Amino acid transport associated to cluster of differentiation 98 heavy chain (CD98hc) is at the cross-road of oxidative stress and amino acid availability. J. Biol. Chem. 291, 9700–9711 (2016).
Arancibia-Garavilla, Y., Toledo, F., Casanello, P. & Sobrevia, L. Nitric oxide synthesis requires activity of the cationic and neutral amino acid transport system y+L in human umbilical vein endothelium. Exp. Physiol. 88, 699–710 (2003).
Miller, H. E. & Bishop, A. J. R. Correlation AnalyzeR: functional predictions from gene co-expression correlations. BMC Bioinforma. 22, 206 (2021).
Barlow, C. et al. Loss of the ataxia-telangiectasia gene product causes oxidative damage in target organs. Proc. Natl. Acad. Sci. USA 96, 9915–9919 (1999).
Blignaut, M., Loos, B., Botchway, S. W., Parker, A. W. & Huisamen, B. Ataxia-telangiectasia mutated is located in cardiac mitochondria and impacts oxidative phosphorylation. Sci. Rep. 9, 1–11 (2019).
Guo, Z., Kozlov, S., Lavin, M. F., Person, M. D., Paull, T. T. ATM activation by oxidative stress. Science 330, https://doi.org/10.1126/science.1192912 (2010).
Golding, S. E. et al. Improved ATM kinase inhibitor KU-60019 radiosensitizes glioma cells, compromises insulin, AKT and ERK prosurvival signaling, and inhibits migration and invasion. Mol. Cancer Ther. 8, 2894–2902 (2009).
Hickson, I. et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res. 64, 9152–9159 (2004).
Xu, Y. & Baltimore, D. Dual roles of ATM in the cellular response to radiation and in cell growth control. Genes Dev. 10, 2401–2410 (1996).
Elson, A. et al. Pleiotropic defects in ataxia-telangiectasia protein-deficient mice. Proc. Natl. Acad. Sci. USA 93, 13084–13089 (1996).
Barlow, C. et al. Atm-deficient mice: a paradigm of ataxia telangiectasia. Cell 86, 159–171 (1996).
Shiloh, Y. Ataxia-telangiectasia: closer to unraveling the mystery. Eur. J. Hum. Genet 3, 116–138 (1995).
Peters, K. et al. Changes in human endothelial cell energy metabolic capacities during in vitro cultivation. The role of “aerobic glycolysis” and proliferation. Cell. Physiol. Biochem. 24, 483–492 (2009).
Terlizzi, C. et al. ATM inhibition blocks glucose metabolism and amplifies the sensitivity of resistant lung cancer cell lines to oncogene driver inhibitors. Cancer Metab. 11, https://doi.org/10.1186/s40170-023-00320-4 (2023).
Yamamoto, Y. et al. Pentose phosphate pathway activation via HSP27 phosphorylation by ATM kinase: a putative endogenous antioxidant defense mechanism during cerebral ischemia-reperfusion. Brain Res. 1687, https://doi.org/10.1016/j.brainres.2018.03.001 (2018).
Li, X. et al. Defining the S-glutathionylation proteome by biochemical and mass spectrometric approaches. Antioxidants 11, https://doi.org/10.3390/antiox11112272 (2022).
Robertson, H. A., Paul, M. L., Moratalla, R., Graybiel, A. M. Expression of the immediate early gene c-fos in basal ganglia: induction by dopaminergic drugs. Can. J. Neurol. Sci. 18, https://doi.org/10.1017/s0317167100032480 (1991).
Bachhawat, A. K., Yadav, S. The glutathione cycle: glutathione metabolism beyond the γ-glutamyl cycle. IUBMB life 70, https://doi.org/10.1002/iub.1756 (2018).
Bröer, A., Wagner, C. A., Lang, F. & Bröer, S. The heterodimeric amino acid transporter 4F2hc/y+LAT2 mediates arginine efflux in exchange with glutamine. Biochem. J. 349, 787–795 (2000).
Dye, J. F. et al. Characterization of cationic amino acid transporters and expression of endothelial nitric oxide synthase in human placental microvascular endothelial cells. FASEB J. 18, 125–127 (2004).
Zielińska, M. et al. Induction of inducible nitric oxide synthase expression in ammonia-exposed cultured astrocytes is coupled to increased arginine transport by upregulated y(+)LAT2 transporter. J. Neurochem. 135, 1272–1281 (2015).
Murohara, T. et al. Nitric oxide synthase modulates angiogenesis in response to tissue ischemia. J. Clin. Invest. 101, 2567–2578 (1998).
Rudic, R. D. et al. Direct evidence for the importance of endothelium-derived nitric oxide in vascular remodeling. J. Clin. Investig. 101, 731–736 (1998).
Zielińska, M. et al. Induction of inducible nitric oxide synthase expression in ammonia-exposed cultured astrocytes is coupled to increased arginine transport by upregulated y(+)LAT2 transporter. J. Neurochem. 135, https://doi.org/10.1111/jnc.13387 (2015).
Nel, M. J., Woodiwiss, A. J., Candy, G. P. Modeling of cellular arginine uptake by more than one transporter. J. Membr. Biol. 245, https://doi.org/10.1007/s00232-011-9408-0 (2012).
Bae, S. Y., Xu, Q., Hutchinson, D., Colton, C. A. Y+ and y+ L arginine transporters in neuronal cells expressing tyrosine hydroxylase. Biochim. Biophys. Acta. 1745, https://doi.org/10.1016/j.bbamcr.2004.12.006 (2005).
Bar, R. S. et al. Extreme insulin resistance in ataxia telangiectasia. N. Engl. J. Med. 298, https://doi.org/10.1056/NEJM197805252982103 (1978).
Schalch, D. S., McFarlin, D. E., & Barlow, M. H. An unusual form of diabetes mellitus in ataxia telangiectasia. N. Engl. J. Med. 282, https://doi.org/10.1056/NEJM197006182822503 (1970).
Inagaki, N. et al. Expression and role of ionotropic glutamate receptors in pancreatic islet cells. FASEB J. 9, 686–691 (1995).
Molnár, E., Váradi, A., McIlhinney, R. A. & Ashcroft, S. J. Identification of functional ionotropic glutamate receptor proteins in pancreatic beta-cells and in islets of Langerhans. FEBS Lett. 371, 253–257 (1995).
Huang, X.-T. et al. An excessive increase in glutamate contributes to glucose-toxicity in β-cells via activation of pancreatic NMDA receptors in rodent diabetes. Sci. Rep. 7, 44120 (2017).
Zhang, G.-F. et al. Reductive TCA cycle metabolism fuels glutamine- and glucose-stimulated insulin secretion. Cell Metab. 33, 804–817.e805 (2021).
Miles, P. D., Treuner, K., Latronica, M., Olefsky, J. M. & Barlow, C. Impaired insulin secretion in a mouse model of ataxia telangiectasia. Am. J. Physiol. 293, https://doi.org/10.1152/ajpendo.00259.2006 (2007).
Donath, H. et al. Progressive liver disease in patients with ataxia telangiectasia. Front Pediatr. 7, 458 (2019).
Zha, S., Sekiguchi, J., Brush, J. W., Bassing, C. H. & Alt, F. W. Complementary functions of ATM and H2AX in development and suppression of genomic instability. Proc. Natl. Acad. Sci. USA 105, 9302–9306 (2008).
Ambrose, M., Goldstine, J. V. & Gatti, R. A. Intrinsic mitochondrial dysfunction in ATM-deficient lymphoblastoid cells. Hum. Mol. Genet. 16, 2154–2164 (2007).
Ravi, D. et al. 14-3-3 σ expression effects G2/M response to oxygen and correlates with ovarian cancer metastasis. PloS ONE 6, e15864 (2011).
Jia, L. et al. Haplodeficiency of ataxia telangiectasia mutated accelerates heart failure after myocardial infarction. J. Am. Heart Assoc. 6, e006349 (2017).
Rajendran, P. et al. The vascular endothelium and human diseases. Int. J. Biol. Sci. 9, 1057–1069 (2013).
Xu, S. et al. Endothelial dysfunction in atherosclerotic cardiovascular diseases and beyond: from mechanism to pharmacotherapies. Pharm. Rev. 73, 924–967 (2021).
McKinnon, P. J. ATM and ataxia telangiectasia. EMBO Rep. 5, 772–776 (2004).
Schneider, J. G. et al. ATM-dependent suppression of stress signaling reduces vascular disease in metabolic syndrome. Cell Metab. 4, https://doi.org/10.1016/j.cmet.2006.10.002 (2006).
Jenstad, M. & Chaudhry, F. A. The amino acid transporters of the glutamate/GABA-glutamine cycle and their impact on insulin and glucagon secretion. Front. Endocrinol. 4, https://doi.org/10.3389/fendo.2013.00199 (2013).
Petrenko, V. et al. Pancreatic α- and β-cellular clocks have distinct molecular properties and impact on islet hormone secretion and gene expression. Genes Dev. 31, https://doi.org/10.1101/gad.290379.116 (2017).
Bishop, A. J. R., Barlow, C., Wynshaw-Boris, A. J. & Schiestl, R. H. Atm deficiency causes an increased frequency of intrachromosomal homologous recombination in mice. Cancer Res. 60, 395–399 (2000).
Angulo, P. Nonalcoholic fatty liver disease. N. Engl. J. Med. 346, https://doi.org/10.1056/NEJMra011775 (2002).
Connelly, P. J. et al. Recessive mutations in the cancer gene ataxia telangiectasia mutated (ATM), at a locus previously associated with metformin response, cause dysglycaemia and insulin resistance. Diabet. Med. 33, 371–375 (2016).
Schneider, J. G. et al. ATM-dependent suppression of stress signaling reduces vascular disease in metabolic syndrome. Cell Metab. 4, 377–389 (2006).
Schuit, F. et al. Metabolic fate of glucose in purified islet cells glucose-regulated anaplerosis in β cells. J. Biol. Chem. 272, 18572–18579 (1997).
Kery, V., Bukovska, G. & Kraus, J. P. Transsulfuration depends on heme in addition to pyridoxal 5’-phosphate. Cystathionine beta-synthase is a heme protein. J. Biol. Chem. 269, 25283–25288 (1994).
Meredith, M. J. & Dodson, M. L. Impaired glutathione biosynthesis in cultured human ataxia-telangiectasia cells. Cancer Res. 47, 4576–4581 (1987).
Rybczyńska, M., Pawlak, A. L., Sikorska, E. & Ignatowicz, R. Ataxia telangiectasia heterozygotes and patients display increased fluidity and decrease in contents of sulfhydryl groups in red blood cell membranes. Biochim. Biophys. Acta 1302, 231–235 (1996).
Campbell, A. et al. Mutation of ataxia-telangiectasia mutated is associated with dysfunctional glutathione homeostasis in cerebellar astroglia. Glia 64, 227–239 (2016).
Palacín, M. & Kanai, Y. The ancillary proteins of HATs: SLC3 family of amino acid transporters. Pflug. Arch. 447, 490–494 (2004).
Cantor, J. et al. CD98hc facilitates B cell proliferation and adaptive humoral immunity. Nat. Immunol. 10, https://doi.org/10.1038/ni.1712 (2009).
Cantor, J. Slepak, M., Ege, N., Chang, J. T., Ginsberg, M. H. Loss of T cell CD98 H chain specifically ablates T cell clonal expansion and protects from autoimmunity. J. Immunol. 187, https://doi.org/10.4049/jimmunol.1100002 (2011).
Braun, D. et al. Aminoaciduria, but normal thyroid hormone levels and signalling, in mice lacking the amino acid and thyroid hormone transporter Slc7a8. Biochem. J. 439, 249–255 (2011).
Ehmsen, J. T. et al. The astrocytic transporter SLC7A10 (Asc-1) mediates glycinergic inhibition of spinal cord motor neurons. Sci. Rep. 6, 6 (2016). 10-19.
Lee, E. H., Kim, Y. H., Hwang, J. S. & Kim, S. H. Non-type i cystinuria associated with mental retardation and ataxia in a korean boy with a new missence mutation(G173R) in the SLC7A9 gene. J. Korean Med. Sci. 25, https://doi.org/10.3346/jkms.2010.25.1.172 (2010).
Sperandeo, M. P. et al. SLC7A7 disruption causes fetal growth retardation by downregulating Igf1 in the mouse model of lysinuric protein intolerance. Am. J. Physiol. 293, https://doi.org/10.1152/ajpcell.00583.2006 (2007).
Xie, X. et al. Lack of the alanine–serine–cysteine transporter 1 causes tremors, seizures, and early postnatal death in mice. Brain Res. 1052, 212–221 https://doi.org/10.1016/j.brainres.2005.06.039 (2005).
Matsuo, H. et al. High affinity D- and L-serine transporter Asc-1: cloning and dendritic localization in the rat cerebral and cerebellar cortices. Neurosci. lett. 358, https://doi.org/10.1016/j.neulet.2004.01.014 (2004).
Sjostedt, E. et al. An atlas of the protein-coding genes in the human, pig, and mouse brain. Science 367, https://doi.org/10.1126/science.aay5947 (2020).
Bishop, A. J. R. et al. Atm-, p53-, and Gadd45a-deficient mice show an increased frequency of homologous recombination at different stages during development. Cancer Res. 63, 5335–5343 (2003).
Claybon, A., Karia, B., Bruce, C. & Bishop, A. J. R. PARP1 suppresses homologous recombination events in mice in vivo. Nucleic Acids Res. 38, 7538–7545 (2010).
Fan, T. W. M. Considerations of sample preparation for metabolomics investigation. In Fan, TM., Lane, A., Higashi, R. (eds) The Handbook of Metabolomics. Methods in Pharmacology and Toxicology. https://doi.org/10.1007/978-1-61779-618-0_2 (Humana Press, 2012).
Fan, T. W. M. et al. Distinctly perturbed metabolic networks underlie differential tumor tissue damages induced by immune modulator β-glucan in a two-case ex vivo non-small-cell lung cancer study. Mol. Case Stud. 2, https://doi.org/10.1101/mcs.a000893 (2016).
Moseley, H. N. & Moseley, H. N. Correcting for the effects of natural abundance in stable isotope resolved metabolomics experiments involving ultra-high resolution mass spectrometry. BMC Bioinform. 11, https://doi.org/10.1186/1471-2105-11-139 (2010).
Lane, A. N., Fan, T. W. M. & Higashi, R. M. Isotopomer-based metabolomic analysis by NMR and mass spectrometry. Methods Cell Biol. 84, https://doi.org/10.1016/S0091-679X(07)84018-0 (2008).
Fan, T. W.-M. & Lane, A. N. Structure-based profiling of metabolites and isotopomers by NMR. Prog. Nucl. Magn. Reson. Spectrosc. 52, https://doi.org/10.1016/j.pnmrs.2007.03.002 (2008).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Cai, S. et al. CD98 modulates integrin β1 function in polarized epithelial cells. J. Cell Sci. 118, 889–899 (2005).
D’Angelo, J. A. et al. The cystine/glutamate antiporter regulates dendritic cell differentiation and antigen presentation. J. Immunol. 185, https://doi.org/10.4049/jimmunol.1001199 (2010).
Thomas, A. G. et al. High-throughput assay development for cystine-glutamate antiporter (xc-) highlights faster cystine uptake than glutamate release in glioma cells. PLoS ONE 10, https://doi.org/10.1371/journal.pone.0127785 (2015).
Olsen, J. V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, https://doi.org/10.1016/j.cell.2006.09.026 (2006).
Nie, J. et al. Synapses of amphids defective (SAD-A) kinase promotes glucose-stimulated insulin secretion through activation of p21-activated kinase (PAK1) in pancreatic β-cells. J. Biol. Chem. 287, 26435–26444 (2012).
Wikstrom, J. D. et al. A novel high-throughput assay for islet respiration reveals uncoupling of rodent and human islets. PLoS ONE 7, e33023 (2012).
Lachmann, A. et al. Massive mining of publicly available RNA-seq data from human and mouse. Nat. Commun. 9, 1–10 (2018).
Chen, S. & Mar, J. C. Evaluating methods of inferring gene regulatory networks highlights their lack of performance for single cell gene expression data. BMC Bioinform. 19, 232 (2018).
Ballouz, S., Verleyen, W. & Gillis, J. Guidance for RNA-seq co-expression network construction and analysis: safety in numbers. Bioinformatics 31, 2123–2130 (2015).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
preprocessCore: A collection of pre-processing functions (Bioconductor version: Release (3.10), 2020).
msigdbr: MSigDB Gene Sets for Multiple Organisms in a Tidy Data Format (2018).
Yu, G., Wang, L.-G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Muraro, M. J. et al. A Single-cell transcriptome atlas of the human pancreas. Cell Syst. 3, 385–394.e383 (2016).
Lawlor, N. et al. Single-cell transcriptomes identify human islet cell signatures and reveal cell-type-specific expression changes in type 2 diabetes. Genome Res. 27, 208–222 (2017).
Segerstolpe, Å et al. Single-cell transcriptome profiling of human pancreatic islets in health and type 2 diabetes. Cell Metab. 24, 593–607 (2016).
Kassambara A. ggpubr: 'ggplot2' Based Publication Ready Plots. R package version 0.6.0, https://rpkgs.datanovia.com/ggpubr/ (2023).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Research 4, 1521 (2015).
Hornbeck, P. V. et al. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 43, D512–D520 (2015).
Acknowledgments
We are grateful to the UTH-SA/Cancer Center Sequencing core. Human tissue was obtained from the NIH NeuroBioBank at the University of Maryland, USA. NMR and MS were recorded using the Metabolism Shared Resources supported in part by P30CA177558 (to B.M.E.). This work was funded by the NIH (K22ES012264, R01CA152063 R01CA241554), a Voelcker Fund Young Investigator Award, CPRIT (RP150445), Stand Up 2 Cancer-Cancer Research UK (RT6187) and GCCRI pilot funds to A.J.R.B.; CPRIT (RP140105) to J.C.R.; NIH T32 (5T32CA148724-3) to S.S.T; NIH T32 (AG021890) to J.N.; DoD grant (W81XWH-19-1-0180) and TL1 (TL1TR002647) to L.A.L.; 2018 AACR-AstraZeneca START grant (18-40-12-GORT) to A.G.; Greehey Family Foundation and NIH (F31AG072902) to H.E.M.; Greehey Family Foundation and GSBS-T32 (CA291696) to A.S.; MCC-T32 (CA148724) to K.K.; and NIH (P30CA054174) to MCC support sequencing core.
Author information
Authors and Affiliations
Contributions
A.J.R.B. and S.S.T. conceived the study. A.J.R.B. and J.C.R. designed the study and wrote the manuscript. J.C.R. conducted the majority of the research and formal analysis. S.S.T., M.P., N.B., L.A.L., J.N., K.K., N.G., T.WM.F., T.A.C., A.G., and A.S. performed experiments. E.L., D.G.R., and L.C. provided technical support. H.E.M. conducted the bioinformatic analysis. A.N.L., B.J.S., S.Z., J.D.L., L.Q.D., and N.M. provided reagents/insights.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Stefan Bröer and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Romero, J.C., Tonapi, S.S., Parihar, M. et al. Loss of CD98HC phosphorylation by ATM impairs antiporter trafficking and drives glutamate toxicity in Ataxia telangiectasia. Nat Commun 16, 5109 (2025). https://doi.org/10.1038/s41467-025-60304-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60304-4