Abstract
Based on the expertise of pathologists, the pixelwise manual annotation has provided substantial support for training deep learning models of whole slide images (WSI)-assisted diagnostic. However, the collection of pixelwise annotation demands massive annotation time from pathologists, leading to a high burden of medical manpower resources, hindering to construct larger datasets and more precise diagnostic models. To obtain pathologists’ expertise with minimal pathologist workloads then achieve precise diagnostics, we collect the image review patterns of pathologists by eye-tracking devices. Simultaneously, we design a deep learning system: Pathology Expertise Acquisition Network (PEAN), based on the collected visual patterns, which can decode pathologists’ expertise and then diagnose WSIs. Eye-trackers reduce the time required for annotating WSIs to 4%, of the manual annotation. We evaluate PEAN on 5881 WSIs and 5 categories of skin lesions, achieving a high area under the curve of 0.992 and an accuracy of 96.3% on diagnostic prediction. This study fills the gap in existing models’ inability to learn from the diagnostic processes of pathologists. Its efficient data annotation and precise diagnostics provide assistance in both large-scale data collection and clinical care.
Similar content being viewed by others
Introduction
The pathology diagnosis forms the basis of clinical and pharmaceutical research and is fundamental in determining patient treatment modalities1,2. The quantitative analysis of digital pathology images (whole slide image, WSI) and the development of computer-aided diagnostic systems provide crucial support to pathologists3. This not only saves a significant amount of medical manpower resources but also enables faster and more accurate patient care.
The development of deep learning (DL)-assisted diagnostic systems4,5,6 in the field of WSI classification has garnered widespread attention7. Traditionally, such methods have relied on the manual extraction of pathologists’ professional knowledge, achieved through pixel-wise annotation on ultra-large WSI images8,9,10. A single WSI typically contains billions of pixels, therefore needs to be divided into several 224 × 224-pixel image patches step by step, while manual pixel-wise annotations are usually provided as labels at the patch level10. Through fine guidance based on pathologists’ professional knowledge, DL models have achieved precise diagnostics11. In the Camelyon16 dataset for breast cancer metastasis diagnosis (comprising 400 WSIs with pixel-wise annotations), ResNet achieves an area under the receiver operating characteristic curve (AUC) of 0.916, and VGG-Net achieves an AUC of 90.9, whereas the average AUC for pathologists is only 0.8112. However, conducting manual annotation is very tedious and time-consuming for pathologists, which limits the possibility of building large training datasets (i.e., larger than several hundred WSIs). Due to significant variations in clinical samples, successful results in small datasets are not yet sufficient to confirm practicality in clinical practice11. The substantial workload associated with large-scale images and the high demand for pathological expertise exacerbate the scarcity of large annotated datasets in the field of computational pathology.
Although weakly supervised learning methods13,14,15 may only require the reported diagnostic outcomes as “labels”, which alleviates the issue of high annotation costs16,17,18,19,20, they often exhibit lower performance, particularly the lower robustness due to the lack of guidance from pathologists’ prior knowledge21,22,23,24. Ideally, the predictions of DL models should correspond only to the diagnostically relevant regions in WSIs, which typically represent a small fraction of the gigapixel WSI20,25. Due to the complexity of pathology images, models may incorrectly associate predictions with irrelevant features (e.g., staining variations, while pathologists primarily rely on tissue and cellular morphology for diagnosis). Weakly supervised learning, without manual pixel-wise annotations, struggles to generalize externally for distinguishing diagnostically relevant images from those of irrelevant ones26,27. Thus, these methods struggle to meet clinical requirements, even when trained on large datasets, as our findings also confirm. Furthermore, due to the difficulty of directly associating predictions with the most diagnostically relevant regions, weakly supervised learning lacks interpretability, posing potential safety risks in clinical applications28.
As gaze-tracking data collection methods have matured29, capturing prior knowledge from human visual behavior for use in computer vision has become increasingly popular. This is often applied in fields such as robotic control or autonomous driving30,31,32,33,34,35,36,37. However, current research on the development of WSI diagnostic systems is largely focused on obtaining guidance from traditional manual annotations or reported diagnostic reports. There is insufficient research on extracting professional knowledge from pathologists’ image review processes or collecting visual annotations to replace traditional manual annotations38,39. This situation highlights the significant cost of data annotation, as well as the poor interpretability of the models: the diagnostic process of the model is detached from the pathologists’ diagnostic process. Actually, both time-efficient diagnostic reports and time-consuming manual annotations stem from the visual image review process of pathologists. In other words, collecting visual data from pathologists incurs almost no additional time cost. The absence of this data from existing available datasets represents a high burden on medical resources. We hypothesize that the visual data obtained using eye-tracking devices from pathologists’ image review processes can reflect their areas of interest, thus forming an alternative to traditional pixel-wise annotation. The core issue of this study is to extract pathologists’ professional knowledge from their visual behavior and effectively apply it to DL, surpassing the performance brought about by traditional manual annotations while reducing data annotation costs. Furthermore, this study aims to fill the gap of DL models that learn from pathologists’ diagnostic processes.
This study aims to decode the expertise of pathologists from their visual behavior and utilize it in the development of a DL system that learns from the diagnostic processes of pathologists. The objective is to achieve more accurate and interpretable diagnostic assistance at a lower data annotation cost, ultimately saving medical manpower resources in the construction of diagnostic systems and providing better care for patients. First, we acquired WSIs and pathologists’ slide-reviewing data using custom-developed software and an eye-tracking device and reported the details of their reading behavior, which included the pathologists’ eye movements, zooming or panning the WSIs, and the final diagnoses. Five thousand eight hundred eighty-one WSIs covering five categories of skin lesions were collected from two medical research institutions. We collected slide review data and manual pixel annotations for approximately 25% of the WSIs and used these as the training set. Two testing sets were constituted from the remaining WSIs: an internal testing set (2431) sourced from the same institution as the training data, and an external testing set (1982) sourced from the other institution. The manual pixel-wise annotation was only used for training comparative algorithms and was not involved in the development of our model.
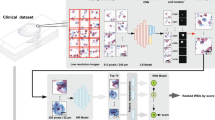
Second, a DL system called Pathology Expertise Acquisition Network (PEAN) was designed to extract the pathologists’ expertise from their slide-reviewing data (as shown in Fig. 1a). We defined the value of this expertise as the “pathologist’s attention level”, with each patch corresponding to an “expertise value”. PEAN computes the “expertise values” for all patches in the WSI, simulating the pathologist’s regions of interest (ROIs) for diagnostic assistance. To validate the correlation between the expertise extracted by PEAN and the actual diagnostic evidence attended to by the pathologist (ground truth), i.e., the ground truth region having higher expertise value fitted by PEAN, we compared the pathologist’s manual pixelwise annotated map, the pathologist’s visual attention map, the expertise value heatmap, and the suspicious region map selected by PEAN to imitate pathologists. We found the overlap among the four types of regions, thus validating the effectiveness of the expertise value.
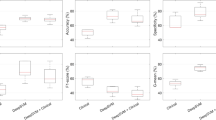
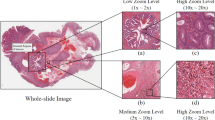
a PEAN model: after training on pathologists’ slide-reviewing data, the model is capable of both performing a multiclassification task and imitating the pathologists’ slide-reviewing behaviors. b Data distribution of the training dataset, internal testing dataset, and external testing dataset. The color legend representing various diseases is utilized in (c, d). c Total number of patients with different skin conditions in the dataset. d Numbers of slide-reviewing operations performed by the different pathologists. The “Overlap” column includes the images listed for each pathologist. e Images at high magnification showing the ROIs (heatmaps, second row) in which the pathologist’s gaze highly overlaps with the actual tumor tissue (marked in blue in the first row). At lower magnifications, the distribution of the pathologist’s observations approximately corresponds with the actual tumor tissue; more examples are illustrated in Fig. 2b. We also observed that areas on which the pathologists focused more attention typically contained chaotic tumor boundaries. Even at high magnification, manual annotation of scattered tumor cells within these areas is challenging, underscoring one of the advantages of using eye tracking for “visual annotation”. BCC basal cell carcinoma, SCC squamous cell carcinoma, SK seborrheic keratosis.
Third, driven by this expertise, PEAN-C for WSI classification and PEAN-I for imitating pathologists’ visual diagnostic process were developed, respectively. PEAN-C achieved an accuracy of 96.3% and an AUC of 0.992 on the internal testing set, and an accuracy of 93.0% and an AUC of 0.984 on the external testing set. Its classification performance and robustness significantly surpassed existing fully supervised and weakly supervised learning models. For example, PEAN-C outperformed the second-best model by 5.5% in the ACC of the external testing set. Furthermore, learning from multiple pathologists’ experiences concurrently has been proven to enhance the classification ability. Distinct from existing DL models that lack the ability to learn from and imitate human expertise, PEAN-I achieves a “human-like” pathological diagnosis by mimicking the diagnostic process of pathologists. PEAN-I autonomously explores the WSI, capturing an image patch at each step and determining the next interesting position based on the current image. This process imitates the visual trajectory formed by pathologists when reviewing WSIs, ultimately outputs a diagnosis based on the captured images. We observed an overlap between the regions identified by both PEAN-I and the pathologists. Additionally, the images identified by PEAN-I have been proven to assist diagnosis statistically (average accuracy gain of 1.24%, p = 0.0053). These validate the interpretability and effectiveness of the imitator, which fills the gap in human-like diagnosis.
Overall, this study represents the DL model to decode human expertise from visual behavior and apply it to assist in the WSI diagnosis. The integration of pathologists’ diagnostic processes with DL has enhanced model performance and annotation efficiency. Unlike existing fully supervised and weakly supervised learning approaches, this study offers a novel approach to computational pathology.
Results
Specifics of the dataset
The unique retrospective dataset constructed in this study comprised two types of data: hematoxylin and eosin (H&E)-stained pathologic images produced by a whole slide scanner, (the WSIs), and slide-reviewing data generated by eye-tracking devices. A total of 5,881 WSIs representing different skin conditions (benign moles [nevus] and four skin diseases [basal cell carcinoma (BCC), melanoma, squamous cell carcinoma (SCC), and seborrheic keratosis (SK)]) were collected (Fig. 1b, c). of these, 3899 and 1982 WSIs were collected from the First Affiliated Hospital of China Medical University (Hospital F) and the General Hospital of Shenyang Military Region (Hospital G), respectively. All image data were paired with slide-level labels generated from previously recorded diagnostic reports. WSIs collected from Hospital F were divided into the training (1468) and internal testing (2431) datasets, while those collected from Hospital G were used as an external testing dataset. A total of 92 WSIs in the internal testing set and all 1473 WSIs in the training set were reviewed by a group of five dermatopathologists, yielding a total of 1565 reviewed WSIs; of these, 552 were reviewed by all five pathologists (Fig. 1d).
The slide-reviewing data consist of the visual attention patterns of pathologists collected via “EasyPathology”, a self-developed eye-tracking system (detailed in the section “Data acquisition and preprocessing”). The data encompass the pathologists’ eye movements during their reviewing WSIs, the two-dimensional mappings of the corresponding gaze points (example gaze heatmaps are shown in Fig. 1e), the magnification levels employed when viewing WSIs, and the diagnostic results. The external environment (such as light and room temperature) for slide-reviewing data collection was standardized as much as possible to minimize the influence of external disturbances on the pathologists’ review. After conducting fatigue tests on the pathologists, we determined that data could be collected continuously for 50 min at a time (detailed in the section “Data acquisition and preprocessing” and Supplementary Information 1). During this process, the actual labels of WSIs were concealed, and the pathologists were asked to make new diagnoses. This more closely resembles the scenario in which pathologists make an initial diagnosis in a clinical setting. Prior to the commencement of the official data collection, the pathologists underwent thorough training to acclimate to the procedure; the data collected from their first five WSIs, considered a set of pretraining images, were excluded from the final collected dataset.
In addition, all WSIs reviewed by the pathologists using the eye-tracking device, were annotated manually to compare the effort involved in traditional fully supervised learning. The manual annotations were completed by the five pathologists involved in the slide-reviewing data collection process. Collecting manual annotations was shown to be extremely labor-intensive; the monitoring records of our dataset showed that pathologists spent an average of 14.2 min manually annotating a single WSI, while the average time for collecting slide-reviewing data per WSI is 36.5 s. Thus, the pathologists’ workload in viewing one WSI was substantially reduced to less than 5% of that required for manual annotation. This indicates that within the same time frame, a pathologist can “visually annotate” a significantly larger number of WSIs, potentially increasing the power in training more accurate and robust DL models.
For DL model training, the WSIs had their backgrounds removed20, then segmented into 224 × 224-pixel patches at 10 × magnification (the monitoring records showed it is the magnification at which the pathologists most frequently exhibited their gaze behaviors), resulting in approximately 2.8 million patches in our dataset.
Overlap of the pathologist’s manual annotations, the ROIs representing the pathologist’s visual behaviors, and the subregions with high expertise value as recognized by PEAN
We attempted to demonstrate that the pathological expertise decoded by PEAN accurately reflects the pathologists’ own knowledge, including their manual annotations and visual behavior. Figure 2 shows this comparison for four WSIs (corresponding to the four malignant diseases investigated in this study). Specifically, precise lesion area contours (Fig. 2a, directly annotated on the images by the pathologists) are shown alongside the heatmap of pathologists’ ROIs (Fig. 2b) and the heatmap of expertise values output by PEAN (Fig. 2c). In Fig. 2b, the observation points of pathologists, captured at 60 Hz by the eye-tracker, are mapped onto the WSIs. A circular convolution kernel is used to calculate the density of observation points within a certain range around any location in the image, with different colors used to distinguish these densities. Figure 2c shows the “pathology expertise value” computed by PEAN for each patch, with higher values indicating a higher potential diagnostic relevance as predicted by PEAN. The corresponding calculation of PEAN is described in detail in the section “Extraction of pathologists’ expertise and fitting an attention score for each image patch in WSI”. Figure 2d illustrates the positions of interest selected by a variant of PEAN, called PEAN-I. PEAN-I is described in detail in the section “PEAN can imitate the visual behavior of pathologists and maps out review trajectories on WSIs”, and it autonomously selects a series of consecutive positions, resembling the movement of a pathologist’s observation points, to simulate the behavior of pathologists.
Between the pathologists’ pixel-level manual annotations for identifying tumor regions (a), heatmaps representing the pathologists’ selected ROIs (b), heatmaps generated by PEAN as the pathologists’ “expert knowledge” (c), and heatmaps depicting the attention trajectories generated by PEAN imitating the pathologists’ slide-review behavior (d).
These maps show visual overlap among the pathologist’s manually annotated tumor boundaries, the visual ROIs, and the areas with high expertise values output by PEAN. This indicates that PEAN-generated regions of focus tend to match the parts of the WSIs identified by pathologists as the ground truth. Notably, the regions shown in Fig. 2a were manually outlined by the pathologists, and so they, too, are manifestations of the pathologists’ expertise. We have also quantitatively verified that the pathology expertise values corresponding to the ground truth were relatively high across the WSIs with manually pixel-wise annotations in the test dataset. Upon investigation, the average “pathology expertise value” fitted by PEAN for the ground truth was 0.822, while the corresponding mean for non-diagnostic regions was 0.357. Additionally, 87.4% of the observation points captured by the eye-tracker fell within the ground truth. The overlap among the three types of regions suggests that, as a result of learning from the pathologists’ slide-reviewing data, the PEAN-output pathology image features can be considered to represent human expertise well, that is, to effectively capture their pathology knowledge. In the inference process, this “expertise” is manifested by PEAN as higher values. This intuitive map comparison provides evidence for the validity of PEAN and the expertise values it decodes. Leveraging this capability, PEAN can be used to mark diagnostically relevant regions that pathologists are likely to focus on. Since the rationale behind PEAN’s decision-making stems from the pathologist’s visual behavior, it can improve the interpretability of the new DL (PEAN) model and increase the confidence of pathologists to approve or accept classification results generated by the new DL model.
In classification tasks, compared to other models, PEAN has demonstrated superior performance
PEAN was compared with eight other models in the classification task to demonstrate its excellence in the field of pathological diagnostic assistance. The baselines included Fully supervised learning: DLCCP9, SLC10, HSL8; weakly supervised learning: CLAM20, ABMIL16, TransMIL17, DS-MIL40, IB-MIL41. The DL models—DLCCP, SLC, HSL, CLAM, ABMIL, and TransMIL—are representative models in the fields of fully supervised learning and weakly supervised learning, respectively. Each model has its unique model architecture and/or distinctive learning approach. In brief, (1) DLCCP utilizes image encoders (such as a convolutional neural network) to directly classify patches, maintaining low computational complexity, and showed excellent performance in the five-class colon WSI classification task. (2) SLC combines neural networks with the extraction of cellular morphological features, capturing deep image features while integrating morphology that is easily interpretable by humans, offering both strong performance and interpretability. (3) HSL features hybrid supervision, requiring only a small amount of pixel-level annotation for training, thereby significantly reducing labor costs while maintaining high performance. Meanwhile, weakly supervised learning, due to its lower training costs, has gained wide attention in recent studies. Since these models lack detailed supervision, they focus on refining model architectures to improve performance. (4) AB-MIL uses neural networks to fit importance scores for each patch, identifying potential diagnostic regions for classification. (5) CLAM utilizes feature clustering to analyze differences between diseases at the patch level, assisting in DL training. (6) TransMIL employs the Transformer architecture, which is currently the most effective feature aggregator. (7) DS-MIL integrates both high- and low-magnification features of pathology images and analyzes their contextual relationships to improve performance. (8) IB-MIL uses WSI-level feature clustering, addressing challenges posed by contextual variations in data to some extent, making it the current best weakly supervised learning model.
Meanwhile, multiple previous studies have shown that the quality of patch-level image encoders can have an impact on the diagnostic performance of models40,41. Particularly, encoder parameters pre-trained on pathological images usually perform well when transferred to new tasks, as they can obtain prior knowledge outside the original dataset. To comprehensively evaluate the difference in classification performance between PEAN and other baselines, we selected two image encoders that can be decoupled from the subsequent network: ResNet5042 pre-trained on ImageNet43 and “CONCH”44 pre-trained on pathological images. After training the image encoder based on the Vision Transformer structure, CONCH can be transferred to the WSI classification task and help the main model achieve better performance. For the included previous studies, we used the officially released code or model parameters when available. The classification module of PEAN (hereinafter referred to as “PEAN-C”) and eight baselines used the same training conditions to ensure fairness. Each model is used for the five-class classification task at the WSI level: benign (Nevus) and four types of skin diseases (BCC, Melanoma, SCC, SK).
Table 1 shows the ACC and AUC of each model. In addition, we also report the recall of each disease for the models to evaluate the ability of the models to identify different diseases, as shown in Supplementary Tables 1 and 2. The results show that (1) in general, DL models trained fully supervised learning yield higher classification accuracy than DL models trained using weakly supervised learning, (2) using special encoder (CONCH) achieves higher classification performance than using conventional encoder (ResNet50), and (3) our new PEAN-C model achieves the highest performance as compared to all 8 other DL models in both internal and external testing datasets. For example, when using the encoder (CONCH), the ACC and AUC of PEAN increase to 93.0% and 0.984 on the external testing set, respectively. The improvement over the second-best DL model (HSL trained with manual annotation) is 5.5% in ACC and 4.2% in AUC, respectively.
The results indicate that PEAN-C shows excellent performance and strong generalization ability in the WSI classification task. Particularly, its high performance in the external test set makes it promising to maintain high reliability in complex clinical environments. In addition, fully supervised learning methods usually exhibit better performance than weakly supervised learning methods. This shows that although the rich human prior knowledge brought by manual annotation is very costly, it still has great competitiveness in environments with high safety requirements, such as healthcare. Given its lower annotation collection cost and excellent performance, PEAN is expected to be widely applied in clinical work and reduce the pressure on medical resources.
PEAN-C also demonstrates high performance on a small training dataset
In addition to testing on the external dataset, we further evaluated the robustness of PEAN by reducing the training data volume. The models were trained on 5 random samples of 30 WSIs per class from the training set and tested with the complete testing dataset. Even under these conditions, PEAN-C still exhibited the best performance among all the models, as shown in Fig. 3a. In the internal testing set, PEAN-C achieved an average ACC of 89.3% and average AUC of 0.976. In the external testing set, PEAN-C obtained an average ACC of 66.6% and an average AUC of 0.830. In addition to the ACC on the external testing set, the performance gap between PEAN-C and the second-best model, HSL, is larger when trained on a smaller dataset compared to when trained on the original dataset (Table 1). The results indicate that PEAN-C is a superior choice when dealing with small datasets because of its ability to learn from the diagnostic processes of pathologists. Due to its greater consistency and robustness, this approach has the potential to be widely adopted by studies relying on small-sized datasets, thereby avoiding the additional costs and privacy risks associated with collecting large amounts of image data.
a Model performance with a lower amount of training data (30 WSIs per class). PEAN-C achieved the best results for this training sample size. b Comparison of the performance of models trained using the slide-reviewing data from five pathologists. The axes of the radar chart have been normalized. Polygons P1–P5 represent the models trained on the data from Pathologists 1–5, respectively; the size of the polygon represents the performance of the model. c Comparison of results for training on different regions (I internal testing dataset, E external testing dataset). PEAN-I autonomously selected a series of image patches from the WSI. Weakly supervised learning models were trained only on these selected patches, ignoring those not chosen, described as “distilled by PEAN-I”. For comparison, weakly supervised models were also trained on all patches, described as “original images”. Training weakly supervised models using only the patches selected by PEAN-I consistently yielded better performance. DLCCP9, SLC10, HSL8 are fully supervised learning methods, CLAM20, ABMIL16, TransMIL17, DS-MIL40, IB-MIL41 are weakly supervised learning methods.
PEAN integrates the visual behaviors of different pathologists to increase classification performance
The discrepancies in manual annotations due to variations in reviewers’ cognition are a well-known issue. Previous studies have attempted to address this problem by introducing additional and more experienced pathologists as “arbiters” of the annotators’ delineations7. However, we found that PEAN can leverage this cognitive difference, integrating the diverse experiences of multiple pathologists, and thereby improve diagnostic performance. Six PEAN-Cs were constructed, five trained separately using the individual review data from each of the pathologists, and one trained collectively using the mixed data from five pathologists. Due to variations in the volume of data reviewed by each pathologist, all models were trained using the “overlap” WSIs reviewed by all five pathologists (a total of 552 WSIs, as shown in Fig. 1c) to control for confounding variables. The ACC and AUC in the two testing sets were compared, as shown in Fig. 3b. The performance of the individual pathologist models varied; however, the model trained using the data from all the pathologists achieved the best performance, with ACCs of 91.98% and 74.7% and AUCs of 0.984 and 0.903 in the internal and external testing datasets, respectively; with respect to those of the top-performing individual DL model (trained using data from pathologist 1 (P1)), the ACCs were 1.48% and 2.44% greater, and the AUCs were 0.004 and 0.016 greater, respectively.
Unlike manual pixelwise annotations, which serve as the primary ground truth diagnostic standard (hard label) for patches, slide-reviewing data is not used in this manner but rather as a “soft label”. When two different pathologists review the same WSI, although their diagnoses for contentious subregions may differ, as long as these regions have been observed by the two pathologists, PEAN interprets these regions as having a higher “attention level”. Therefore, when training with slide review data from multiple pathologists, PEAN does not experience a performance decline due to label confusion, a common issue that affects traditional supervised learning methods. By learning from a more diverse set of visual behaviors, PEAN can further refine pathologists’ expertise, thereby improving classification performance.
PEAN can imitate the visual behavior of pathologists and map out review trajectories on WSIs
Reinforcement learning (RL)45,46,47,48 was used to develop the imitation module of PEAN (PEAN-I), which is capable of imitating pathologists’ visual behaviors for selecting regions on WSIs (details are discussed in the section “Construction of an RL model to imitate the slide-reviewing behavior of the pathologists”). PEAN-I is an agent capable of autonomously selecting a series of regions on WSIs by scanning the WSI in a manner similar to the gaze patterns used by the pathologists but with a fixed step size and movement direction each time (up, upper-right, right, and so on, eight directions in total), as shown in Fig. 2d. The regions selected by PEAN-I also exhibited high degree of overlap with the ROIs manually annotated by the pathologists and the ground-truth tumor region. This indicates that, in addition to reflecting pathological knowledge learned from the pathologists’ “expertise”, PEAN can imitate the pathologists’ slide-reviewing behavior, truly learning human priors.
Furthermore, PEAN-I can be effectively integrated with existing weakly supervised learning models. The regions selected can be used to select ROIs from the original WSIs, followed by further training of weakly supervised learning models, leading to improved classification performance. As shown in Fig. 3c, when CLAM, ABMIL, and TransMIL were trained with pathology images generated by PEAN-I, both the ACC and AUC were increased in the two testing datasets. This improvement in performance was statistically significant, with p values of 0.0053 and 0.0161, respectively, as determined by paired t tests. This effective enhancement of DL model performance demonstrates the efficacy of imitating pathologist behavior, reflecting its ability to “learn” pathologists’ expertise while providing strong evidence for the validity of this expertise.
Discussion
Practicality
In addition to the high diagnostic performance, another factor affecting the practicality of PEAN is the cost required to collect training data. Expanding WSI data to train more robust artificial intelligence is the development trend in this field. The process of collecting eye-tracking data can be seamlessly integrated into the daily work of pathologists. This avoids repetitive reviews and thus minimizes labor costs. With the maturation of scanners and digital imaging technology, digital pathology review has gradually replaced microscope-based review, providing pathologists with a more optimal working environment11. For example, this shift avoids potential biological contamination and allows for a more relaxed working posture. Our self-developed “EasyPathology” software, combined with an eye-tracking system, forms a new data collection system that integrates seamlessly with digital pathology review, offering a “nearly imperceptible” data collection method. During the review process, pathologists can work in their familiar manner with minimal additional manual intervention. Multiple videos from actual data collection processes have been uploaded as Supplementary Movies 1 and 2 to demonstrate the simplicity and feasibility of this approach.
Reliability
The core of PEAN is to decode the variable visual patterns of pathologists into a shared feature space to solve the problem that has puzzled previous studies, that is, pathologists are unlikely to always focus on the region most relevant to diagnosis. PEAN can effectively avoid annotation confusion caused by pathologists scanning benign tissues to look for suspicious lesions or being distracted for a moment.
Since the pathologist’s diagnostic conclusions are derived from their observations of the WSI, the following reasonable inferences can be made: (1) the pathologist’s entire slide-reading process can be simplified as a sequence of transitions between observation locations on the WSI; (2) the pathologist obtains diagnostic evidence from at least a subset of these observation locations. Based on these inferences, we posit that the images corresponding to the observation locations (or a subset thereof) possess potential contextual relationships. Then, through an image encoder and attention mechanism, PEAN can capture and analyze their overall connections. This endows PEAN with the following capabilities: (1) it can analyze diseases that require the pathologist to gather different tissue characteristics from multiple locations to make a definitive diagnosis; (2) the multiple gaze locations of the pathologist may correspond to the same underlying lesion, and capturing similarities across these locations can enhance DL model training; and (3) although pathologists may observe locations that are irrelevant to diagnosis, making a correct diagnosis must rely on the observed locations. Viewing the images that were observed as a whole can effectively eliminate the interference of incorrect labels.
A typical example that can reflect the diagnostic logic of pathologists is the examination of melanoma. During the observation process from the epidermis to the deeper layers by pathologists, it may be observed that melanoma has a similar distribution to moles: at the junction of the dermis and epidermis. Subsequently, pathologists need to observe the distribution of melanocytes. A more confused distribution suggests a situ melanoma. Most melanocytes need to be observed closely. For example, immature cells suggest invasive melanoma, while mature cells suggest a nevus. This complex process suggests that pathologists may spend a considerable amount of time observing benign cells, and diagnostic evidence is also difficult to obtain from a single glance alone.
Subsequently, by comparing the image feature differences between the regions pathologists tend to focus on and the regions they have not observed, PEAN identifies the image characteristics of the regions that pathologists are truly inclined to examine. Regions with these characteristics are assigned higher weights, thereby receiving more (not exclusive) attention in subsequent WSI-level classification tasks. In this process, the eye-tracking data from the pathologists serve as “soft labels”, solely guiding PEAN to learn “what types of images pathologists are inclined to focus on”. It is important to note that, at this stage, the “hard labels” used for training the WSI classifier—pathological diagnoses—have not yet been introduced. This approach directly avoids the accumulation of human errors caused by “observing diagnostically irrelevant regions”. In this way, PEAN, having learned human prior knowledge, models the “potential pathologist attention” for subregions within the WSI as the proposed pathology expertise in this study. In the subsequent WSI classifier training, this attention is used as the importance score for each subregion, and after fully integrating the image features of all subregions, a diagnostic prediction for the WSI is made. The detailed architecture and parameters of PEAN are described in the section “Extraction of pathologists’ expertise and fitting an attention score for each image patch in WSI”.
Future applications
As mentioned in the section “Overlap of the pathologist’s manual annotations, the ROIs representing the pathologist’s visual behaviors, and the subregions with high expertise value as recognized by PEAN”, PEAN can fit a “pathology expertise value” for each image patch, which we interpret as the pathologist’s potential level of attention to the patches. The derived downstream task of marking suspicious lesion areas in WSIs to assist pathologists in diagnosis has been validated with higher interpretability. We not only provided multiple examples demonstrating significant overlap between the subregions distilled by PEAN and the ground truth, but also presented quantitative analysis results: the pathological experience value fitted by PEAN for the ground truth regions is 2.3 times higher than that for non-diagnostic (normal tissue) regions. Thus, PEAN can enhance the trust of pathologists in DL-generated recommendations. This increased trust stems not only from the accuracy of PEAN’s diagnostic results but also from its ability to highlight suspicious regions in WSIs by learning from human experience. Collecting slide-viewing data from specific pathologists allows for the customization of DL models to align with their individual work habits, reducing resistance to using DL tools among pathologists. Moreover, ROIs generated through backpropagation based on DL-generated predictions may exhibit significant deviations from the ground truth due to erroneous predictions, whereas PEAN does not suffer from this issue. PEAN follows a forward propagation manner and causal inference process, firstly marking diagnosis-related regions and then classifying WSIs. Compared to weakly supervised learning, PEAN, which aligns more closely with the habits of pathologists and carries less risk in the propagation process, offers greater advantages in assisting pathologists with their work.
Besides developing new DL models, collecting slide-reviewing data also holds great potential for the education of junior pathologists. Pathologists rely on visual observation for diagnosis, and their visual behavior can intuitively reflect their diagnostic reasoning. However, due to the current scarcity of large-scale eye-tracking data collection and analysis in this field, it is difficult to provide timely summaries or guidance on their observational behavior. The accumulation of pathologists’ expertise is still largely driven by “word-of-mouth”, traditional media (such as books), or self-exploration. Large-scale slide-reviewing data collection could offer standardized recommendations on junior pathologists’ visual behavior, such as identifying their raddled working state, or even reducing the risk of potential biases caused by the accumulation of errors by junior pathologists. Collecting the slide-reviewing data from experienced pathologists and decoding their expertise could generate valuable educational materials. By replaying the recorded or modeled key regions, valuable guidance can be provided to train junior pathologists. Enhancing pathologist training through eye-tracking technology represents a novel and practically significant research direction that has yet to be fully explored.
Current limitations
Although this study has been proven to have unique advantages in the WSI-assisted diagnosis task, there are still several aspects that need improvement. For example, when facing out-of-distribution (OOD) WSIs, it may misjudge them as false positives of a certain type of tumor; currently, only slide-reviewing data has been collected from five pathologists, which is not sufficient to represent the whole.
Such problems can inspire the development of further work. Novelty detection has been proven to be an effective way to solve the OOD problem. This module can be added to the existing architecture to detect categories that have not participated in training and avoid false positive detections. At the same time, another important means to address existing deficiencies is to expand diverse training data, including WSIs and slide-reviewing data. Learning to read data representing more pathologists and more disease types can make PEAN more robust and easily transferred to other diagnostic tasks. In particular, it is proved in the section “PEAN can imitate the visual behavior of pathologists and maps out review trajectories on WSIs” that PEAN can be seamlessly combined with existing weak supervised learning methods to achieve more accurate diagnosis. The plug-and-play feature allows researchers to directly use the model weights disclosed by us to further design their own models, which can arouse widespread interest. Collecting diverse data and updating model weights will become our continuous work and provide a basic model with extensive prior knowledge for subsequent research.
Methods
Ethics statement
This research complies with all relevant ethical regulations. The research and dataset don’t contain any personally identifiable information, and have been approved by the Medical Science Research Ethics Committee of the First Affiliated Hospital of China Medical University, with ethical code “kelunshen [2021] 2020-196-2” (number: AF-SOP-07-1.1-01).
Statement of informed consent: the five pathologists who participated in the slide-reviewing phase of this study have consented to the publication. Furthermore, all five individuals are listed as co-authors of the present manuscript, acknowledging their significant contributions to the study.
Data acquisition and preprocessing
Before the structure of the DL model is described, some details of the special eye-tracking data involved in this work are described first. The image dataset used in this study consisted of 5881 H&E-stained, previously diagnosed pathological WSIs, including images of one benign skin condition (nevus) and four diseases: BCC, melanoma, SCC, and SK. These WSIs were collected from 5107 patients between 2016 and 2022 from the First Affiliated Hospital of China Medical University and the Shenyang Military Region General Hospital, respectively. As shown in Supplementary Fig. 2, these WSIs were continuously collected during this time period to maintain a distribution similar to the real world. One WSI representing nevus and one WSI representing BCC are not included because their image quality is low. This involves re-examination by pathologists. The criteria for pathologists to judge data quality in this process are as follows: (1) tissue samples that were too small or had diagnostic areas too limited to adequately represent the entirety of the disease; (2) folded sections; (3) overstaining; (4) high levels of tissue fragmentation; and (5) a significant presence of bubbles. When pathologists believe that the occurrence of the above situations interferes with making a diagnosis, the corresponding WSI will be excluded. Since this process is relatively subjective, two pathologists participated in the screening. Only when they unanimously recognize the image quality will the WSI be included in the data set.
At the same time, the labels of WSIs are also re-examined because previous pathology reports are not absolutely directly usable. For example, pathology reports are more suitable for clinicians to read rather than artificial intelligence researchers. Pathological diagnoses that have already been made may also be incorrect, in the worst possible case. In particular, when two pathologists make inconsistent judgments on the WSI label, at least one more senior pathologist is additionally introduced as a “referee” to jointly discuss and reach the final label.
To collect eye-tracking-based WSI review data from pathologists, we developed the “EasyPathology” slide review software. This software allows pathologists to review WSIs while simultaneously using an external eye tracker to capture their eye movement signals, which are then mapped onto the 2D WSI. A commercialized instrument, “Tobii Pro Spectrum”49 is used as an eye-tracker, which records eye movements at a sampling frequency of 60Hz and uses built-in algorithms to map the pathologist’s gaze position on the screen. Since the screen is often too small to display the full WSI, the review process involves a “sliding window operation”. EasyPathology further maps the recorded screen positions onto the WSI, as illustrated in Supplementary Fig. 3, while their attention heatmaps for WSIs are illustrated in Fig. 1e.
The experimental environment and equipment were carefully evaluated to ensure accurate data collection without disrupting the pathologists’ workflow. The lab was soundproof to minimize distractions, blocked natural light, and used multiple artificial light sources to ensure even lighting. The computer and screen setup matched clinical work conditions, featuring an Intel(R) Core i7-10700F CPU, an Nvidia 1660super GPU, and a screen resolution of 1920 × 1080 with a 60 Hz refresh rate. This hardware setup allowed for clear and smooth WSI reviews. The eye tracker was positioned below the screen, and its accuracy, along with built-in algorithms to reduce potential deviations, was key to ensuring precise data collection. The Tobii Pro Spectrum has a minimum latency of 2.5 ms and maintains an accuracy of less than 0.06° RMS at a sampling frequency of 60 Hz. With an initial viewing distance of 65 cm, the system allowed for a maximum head movement range of 34 cm (without losing eye-tracking accuracy), ensuring that pathologists could comfortably complete the review process.
Before the formal data collection process, five participating pathologists received training on EasyPathology and the use of the eye tracker. Prior to each review session, a 30-s calibration was conducted using Tobii Pro Spectrum’s official software to minimize tracking offset. Additionally, the first 5% of data from each review session was treated as a practice session and excluded from the final dataset to avoid potential noise caused by pathologists adjusting to the system. During the review process, EasyPathology recorded the pathologists’ activities, including (1) zooming in on the WSI; (2) using the sliding window to navigate to different areas; (3) their observation positions (or observation points, representing the area of the WSI being observed in each frame), and (4) logging their final diagnosis (with the true label hidden from the pathologists during the review). After diagnosing a WSI, the system automatically moved to the next WSI for review. As shown in Supplementary Fig. 1b, the pathologists reviewed WSIs for more than two hours. An analysis of visual behavior features over different review durations revealed that both “the first fixation scale” and “searching number” were highly correlated with diagnostic accuracy, and these features showed significant changes after 50 min of review time. Based on this observation, we propose 50 min as the optimal review cycle for pathologists, and this has been incorporated into the data collection protocol.
Finally, all eye-tracking data were re-examined, and data with poor integrity or stability were excluded. Evaluating the integrity of observation points refers to identifying interruptions in the review process caused by autonomous behaviors of the pathologist (e.g., leaving the workstation, taking phone calls), which can result in extended “vacuum periods” where no eye-tracking data is collected. Stability refers to ensuring that the eye movement angular velocity of the pathologist does not remain at a high level for prolonged periods. Eye movement angular velocity greater than 30° per second typically indicates eyelid twitch. If such anomalies occur too frequently in the review data for a particular WSI (set as more than three occurrences), it suggests that the pathologist may not be in an optimal working condition. Additionally, operational errors by the pathologist (e.g., repeated clicking or exiting the software prematurely) could result in missing data. As continuous viewing time increases, a gradual drift in the position of eye-tracking signals may also occur.
Although the eye-tracking device used in the study allows for a significant range of head movement and maintains long-term accuracy, there is a potential risk of drifting after extended work periods. Data cleaning for drift is performed by the pathologists themselves, who can review their recording after data collection (a feature provided by EasyPathology, which allows replay of the pathologist’s observation point movements, as shown in the uploaded videos) to determine whether drift occurred. One important method to prevent drift is to set a reasonable maximum duration for each continuous working session and to recalibrate the eye tracker at the start of each session. This re-examination process resulted in the removal of the slide-reviewing data for 101 WSIs. In the end, a total of 3,978 slide-reviewing data from five pathologists were collected; of these, 542 WSIs were reviewed by all pathologists (Fig. 1c, d).
For a given WSI W, the slide-reviewing data collected by the eye-tracking device contains the observation points captured at a specific frequency, described as a point set \(\left\{Points\right\}\,\in \,{R}^{{\mathrm{points}}{\mathrm{number}}\times \left({\mathbb{1}}{\mathbb{,}}{\mathbb{1}}\right)}\). Each recorded position can be viewed as a specific coordinate point on W (or an approximately circular area centered around this point). W is segmented into an irregular foreground containing human tissue images and the remaining light-colored background, with the boundary of the foreground defined as bW. When reviewing a WSI, pathologists often perform multiple “sliding window” actions, meaning they can only view a portion of the WSI at any given time. To mimic this limited view during slide review and decode pathology expertise from such behavior, we sampled d × d window images from the WSI at magnification M, corresponding to the pathologist’s fixed window position during the review, and denoted these as \(\{{{win}}_{i}^{M}\}_{i}^{K}\in {R}^{K\times \left[d\times d\right]}\). When the pathologist performs multiple “window slides” during the review of W, all “screen images” observed by the pathologist (a total of K) are sampled and used for training PEAN. The size of d is set to 1920 (corresponding to the screen size of 1920 × 1080) with the sampled window images extending both vertically and horizontally to satisfy the typical square image input requirement of convolutional neural networks. The magnification M corresponds to the zoom level selected by the pathologist while viewing the WSI. The set of {Points} is then preprocessed as follows:
-
1)
Eye-tracker can calculate the coordinates of observation points. Pathologists’ visual fields extended beyond the computer screen or the boundaries of the foreground are considered indicative of non-meaningful visual behaviors and are not included.
-
2)
When the angular velocity of the pathologist’s eye movement exceeds 30° per second, the system records it as an eyelid twitch and removes the corresponding points.
The preprocessed point set is then denoted as {Points'}. We classified the pathologists’ visual behavior into two categories—fixation and search, based on the density of observation points. We define fixation locations as image regions where the pathologist spent more time and where the recorded observation points were denser, possibly indicating regions with a relatively higher suspicion of disease. In these areas, the pathologist’s gaze movement was slower, reflecting higher attention. Conversely, the search behavior represents rapid eye movement, where the sampled observation points were sparser. Density-based spatial clustering of applications with noise (DBSCAN)50 is used to classify the pathologists’ observation points into “fixation” and “search”. It is an algorithm that automatically identifies clusters and outliers in a dataset based on the point density. It operates by using two pre-defined parameters—radius and the minimum number of neighboring points—to determine the density around a point. If the number of points within the neighborhood exceeds the pre-set minimum, the point is considered a core point and starts forming a cluster; otherwise, it may be marked as an outlier or noise. This allows DBSCAN to effectively identify dense areas as clusters while isolating sparse points as noise. The underlying principle makes DBSCAN well-suited for distinguishing “clustered” and “dispersed” points in a plane, while also being particularly effective in handling noise. The observation points within the cluster represent fixations, and the points outside the cluster represent searches. That is to say, search can be represented as the migration process of fixation.
Directly locating lesions by using the observation of pathologists is risky because pathologists cannot always focus only on those lesions38. shows that using the images that pathologists have focused on observing as completely equivalent to lesions for training will damage the performance of the model. Obviously, the basis for pathologists to make diagnoses comes at least in part from the images they have focused on observing. Therefore, learning the image features that pathologists tend to focus on rather than the diagnostically evidential features they have observed is a simpler and safer task and is a direct utilization of known information. Taking the images observed by pathologists as a whole and analyzing their internal contextual connections, that is, their similar features, one can infer what kind of images pathologists tend to focus on. A common means to achieve this purpose is attention-based image feature fusion, such as Transformer17. Then, the trained model can provide “potential attention of pathologists” for all regions, used as an assistant in diagnosis. The images from pathologists’ fixation locations are constructed into a sequence, denoted as the expert trajectory \({{\rm{\tau }}}\in {D}_{{demo}}\). This preference of pathologists for different image features is defined as “pathology expertise”. It is specifically manifested as a value of image patch: the “pathology expertise value”. Section “Extraction of pathologists’ expertise and fitting an attention score for each image patch in WSI” discusses in detail the process of learning pathology expertise. Before this, some key parameters and variables are introduced.
The sequence \( < {P}_{i0}^{M},{P}_{i1}^{M},{P}_{i2}^{M}\ldots \ldots > \) formed by fixation locations \({P}_{{ij}}^{M}\in {{win}}_{i}^{M}\). The gaze duration coefficient Etime and gaze point density coefficient Edensity are introduced as indicators for evaluating the differences in importance between different fixation locations:
where β1 and β2 are weight coefficients, both of which are set to 0.8; Pointsnum(i, j) indicates the total number of Points∈{Points'} contained in \({P}_{{ij}}^{M}\); mean({Pointsnum}W) indicates the average value of Points contained in each fixation location in the full slice W; Regiondistance(i, j) is the average distance from \({P}_{{ij}}^{M}\) to other fixation locations \({P}_{{ik}}^{M}\left(i\ne k\right)\) under \({W}_{i}^{M}\); and mean({Regiondistance}W represents the average value of the Regiondistance() function over all fixation locations in the full slice W. Under the action of β1 and β2, Etime (i, j) is positively correlated with the gaze duration of region \({P}_{{ij}}^{M}\), while Edensity (i, j) is positively correlated with the degree of aggregation of \({P}_{{ij}}^{M}\) in W.
When the image is magnified to magnification M, each \({{win}}_{i}^{M}\) is partitioned into patches of dimensions [l × l]. These patches are centered on point \({P}_{{ij}}^{M}\) and recorded as \(\{{x}_{{ij}}^{M}\}\frac{N}{j}\in {R}^{N\times [l\times l]}\). Simultaneously, \({x}_{{ij}}^{2M}\) is also segmented with the same center point. \({x}_{{ij}}^{2M}\) is the patch sampled at the same center point as xij under a magnification of M × 2, with a size of [l × l] (thus, the image covered by this patch has a size of [l/2 × l/2] under magnification M). Images \(\{{{win}}_{i}^{M}, < {x}_{i0}^{M},{x}_{i1}^{M},{x}_{i2}^{M}\ldots > , < {x}_{i0}^{2M},{x}_{i1}^{2M},{x}_{i2}^{2M}\ldots > \}\) are individually input into the image encoder to extract features. Subsequently, the features obtained from images sampled at different magnifications are concatenated to achieve the integration of multi-scale image information. These images serve as the visual input corresponding to the gaze positions of pathologists, with the objective of learning which characteristics of pathological images attract the interest of expert pathologists.
Extraction of pathologists’ expertise and fitting an attention score for each image patch in WSI
The expertise of the pathologists can be described as an attention score based on the pathologists’ manual sampling of WSIs (Fig. 4a), which can also be considered the degree of similarity between any sequence \( < {P}_{i0}^{M},{P}_{i1}^{M},{P}_{i2}^{M}\ldots \ldots > \) in the WSI and the pathologists’ manual sampling at the level of the aggregated image features. We construct an optimal control framework based on the principle of maximum entropy, which essentially posits that the sampled experts’ behavior results from random, nearly optimal responses based on an unknown cost function. Specifically, under the expertise extraction model fexperience, it is assumed that the expert samples the demonstration trajectory τ from a distribution:
a Model for extracting pathologists’ expertise. b Feature distillation classification model using the extracted pathologists’ expertise and c Model imitating pathologists’ slide-reviewing behavior using RL.
\({{{\rm{\tau }}}}_{i}=\{{S}_{0},{S}_{1}\ldots {S}_{T}\}\) can be viewed as the trajectory of the pathologist’s ROIs under \({{win}}_{i}^{M}\). \({C}_{{{\rm{\theta }}}}\left({{\rm{\tau }}}\right)={\sum }_{0}^{T}{S}_{t}\) is an unknown pathology expertise value function parameterized by θ. St represents the set of images \(\{{x}_{{it}}^{M},{x}_{{it}}^{2M},{{win}}_{i}^{M}\}\) at current time t, and \(Z=\exp \int \left(-{C}_{\theta }\left(\tau \right)\right)d\tau\) is a partition function, which is used to keep the integral of the probability distribution function p(τ) always equal to 1. Under this specification, trajectories with higher values have a greater probability of being selected, and while the expert pathologist may select optimal actions, suboptimal actions may also occur.
The randomly sampled trajectory and the subsequent trajectory generated by the imitation model fmimicry are introduced as nonexpert pathologist trajectories \(\tau \in {D}_{{\mathrm{samp}}}\) into this part of the model, which then undergoes adversarial learning with expert trajectories \(\tau \in {D}_{{\mathrm{demo}}}\). In this way, the behavioral trajectories generated by fmimicry are “guided” toward a distribution closer to that of the expert behaviors, and the expertise extraction model fexperience acquires the ability to distinguish between the two types of trajectories. fexperience takes as input τi.
The input images are passed through a pretrained encoder to obtain the feature vectors. The pre-training process of the image encoder involves using image datasets like ImageNet to equip the encoder with the ability to extract image features, serving as the foundational layer of the designed model. During pre-training, the model is trained through supervised learning on labeled images, optimizing its weights using the cross-entropy loss function to gradually learn how to distinguish between different image categories. The main purpose of pre-training is to allow the model to learn image features at various levels, from low-level features such as edges and textures to high-level features such as shapes and complex object characteristics. The output of the image encoder is a feature vector, which is a high-dimensional representation of the image’s features.
\({u}_{i}^{M}\) is considered to represent the window-level information from \({{win}}_{i}^{M}\), and \(\{{\sum }_{t}^{T}{v}_{{ij}}^{M},{\sum }_{t}^{T}{v}_{{ij}}^{2M}\}\) are considered to carry information contained in the transitions among the pathologist’s fixation locations \(\{{\sum }_{t}^{T}{v}_{{ij}}^{M},{\sum }_{t}^{T}{v}_{{ij}}^{2M}\}\). \({v}_{{it}}^{M}\) is concatenated with \({v}_{{it}}^{2M}\) and then passed through a transformer layer, which outputs the second-layer feature vector \({r}_{{it}}^{M}\) at the current time. The \({r}_{{it}}^{M}\) of each moment is concatenated with the global feature \({u}_{i}^{M}\), and the predicted attention scores \({C}_{{{\rm{\theta }}}}^{{\prime} }\left(t\right)\) are obtained through a multilayer perceptron (MLP) using the mean square error loss:
where coefficients λ1 and λ2 are used to balance the importance of gaze duration and gaze point density in the region, satisfying \({\lambda }_{1}+{\lambda }_{2}\equiv 1\). β is set to 0.9.
We combine fexperience based on sampling with fmimicry, which is essentially an RL model. The core idea is to optimize the trajectory distribution for the current cost \({C}_{{{\rm{\theta }}}}\left({{\rm{\tau }}}\right)\) through fmimicry and to assign higher values to trajectories that are closer to expert behavior. This method allows us to make reverse optimal choices in an infinite state space, even without a known system model.
Feature distillation and classification with weakly supervised models
Using the pathology expertise value fitted by PEAN, additional weighted scores can be given to all image patches in the WSI, so that the WSI-level classifier pays more (rather than completely) attention to the locations that pathologists are more inclined to focus on. In addition, the pathological experience value can also be used for feature distillation to eliminate the interference of redundant patches. The architecture of the WSI-level classifier we designed: PEAN-Classification (PEAN-C) is shown in Fig. 4b. For a given WSI W, a series of patches can be obtained through tissue-region and instance-level segmentation. A pretrained image encoder is utilized to extract features from these patches. However, this portion of the model is not involved in the training process; the model designed in this study focuses solely on learning from the extracted features. The patches yield instance features \(X=\{{x}_{1},{x}_{2},\ldots,{x}_{K}\}\), where K represents the total number of patches contained in W. Each individual patch xi possesses a latent label yi that indicates the disease type to which the tissue in xi belongs and that is unknown to the model. The task of feature distillation is to distill the areas on which pathologists are most likely to focus and that best represent a certain disease, i.e., patches with high cθ values representing a high probability of belonging to a specific disease. Specifically, the top-k distilled features \(\{{\hat{o}}_{1},\ldots {\hat{o}}_{k}\}\) satisfy the following relationship:
Here, ci represents the cost value corresponding to xi, and \({\hat{y}}_{i}\) is the probability of xi being predicted as belonging to its disease type. The optimization method for \({\hat{y}}_{i}\) involves guiding the maximum value of the corresponding disease type in \(\{{\hat{y}}_{0},...{\hat{y}}_{K}\}\) with the WSI-level label Y, enabling the model to autonomously learn the “patch most likely to belong to a certain disease” during the optimization process:
loss2 is a cross-entropy loss function that measures the difference between predicted probabilities and true labels. Its goal is to make the model’s predictions closer to the true labels. The features to be distilled, \(\{{\hat{o}}_{1},\ldots {\hat{o}}_{k}\}\), have high “pathologist attentiveness”. We assume these features are relevant to diagnosis, and thus, the predicted categories for these instances should ideally match the category of the WSI. The diagnosis of the WSI is used as a pseudo-label for these instances, and loss2 is effectively used for instance-level classification, aiming to predict the probability that highly attended instances belong to a certain disease category. However, in the real world, it is not possible to pre-know the WSI’s label in order to distill label-related instances. Therefore, representative patches of all disease types covered in this study are grouped together for WSI-level classification:
Additionally, the k features obtained are transformed into WSI-level features through feature fusion ffusion and used for WSI classification. There are numerous suitable feature fusion methods, and so this function can be interchanged with other bag-based MIL approaches. Common methods include feature score weighting16 or self-attention mechanisms17. Transformer17 is selected, which is one of the best-performing network architectures, utilizing a self-attention mechanism. The WSI features are then passed through a fully connected layer to yield the probabilities of belonging to different disease types.
where loss3 is a cross-entropy loss function, aimed at bringing the predictions from the fully connected layer closer to the true labels.
Construction of an RL model to imitate the slide-reviewing behavior of the pathologists
For a given WSI W, an RL45,46 task can emulate the visual behavior of expert pathologists to conduct a rapid search on a two-dimensional plane and locate areas potentially harboring lesions47,48. As shown in Fig. 4c, the objective is to generate a “human behavior-like” search trajectory.
The trajectory under the window is set to imitate the pathologist’s behavior and is regarded as a Markov decision process (MDP). At time t, the agent acquires patch \({x}_{\left({it}\right)}\) corresponding to a certain position Pt in \({w}_{i}\) (more specifically, the physical location corresponding to a pixel point in the pathological image), and, along with \({w}_{i}\), constitutes the state St at time t. Given the irregular characteristics of pathological images, there can be significant variability between WSIs derived from specimens of the same type of tissue. Therefore, the RL framework in the context of pathological images can be considered to possess an infinite state space. The action at is described as a change in position within wi, namely, starting from Pt, movement occurs in one of the eight preestablished directions (upper-left, up, upper-right, right, etc.) with a fixed step length l, resulting in a new position Pt+1. Based on the expert pathologists’ actions discussed in the section “Specifics of the dataset”, which are both sequential and continuous, we choose to generate a state-action sequence \(\{{S}_{t},{S}_{t+1}\ldots,{S}_{T},{a}_{t},{a}_{t+1}\ldots,{a}_{T-1}\}\) after repeating this pattern multiple times before assigning a reward sequence \(\{{R}_{t},{R}_{t+1}\ldots,{R}_{T-1}\}\). Note that this is not calculated as a reward during the single \({S}_{t}+{a}_{t}\Rightarrow {S}_{t+1}\) process. This approach draws inspiration from the classic RL model: “Deep Reinforcement Q Learning Net” (DRQN)46 and reflects the complete observational information in the process of pathologists reviewing slices, rather than the simplistic model based on a single patch as the basis for action selection. This RL model integrates global observational information with expertise accumulated prior to the current moment t.
The \({c}_{\theta }\left(t+1\right)\) obtained from \({f}_{{\mathrm{experience}}}\) will act as the reward \({R}_{t}\) for the process \(\{{S}_{t},{a}_{t}\}\Rightarrow {S}_{t+1}\), the purpose of which is to determine the reward size for executing action \({a}_{t}\) under the current state \({S}_{t}\) based on the value \({c}_{\theta }\left(t+1\right)\) possessed by the next moment’s state \({S}_{t+1}\). The RL model consists of two networks, \({Q}_{{\mathrm{eval}}}\) and \({Q}_{{\mathrm{target}}}\), both of which use fully connected layers but have different parameters. The input to networks is the image features representing \({S}_{t}\), and the output is \({R}_{t}\) associated with selecting different \({a}_{t}\) in this step. \({Q}_{{\mathrm{eval}}}\) obtains the estimated rewards \(\{{\hat{r}}_{{tn}}\}_{n}^{8}\) for all actions corresponding to \({S}_{t}\) and chooses the action \({a}_{t}={argmax}\left(\hat{{r}_{t}}\right)\) with the highest reward. As shown in Fig. 4.c, \({a}_{t}\) is combined with \({P}_{t}\) to calculate \({P}_{t+1}\) and obtain the state \({S}_{t+1}\) at moment \(t+1\), which is then used by \({f}_{{\mathrm{experience}}}\) to obtain \({R}_{t}\) (the pathology expertise value) and thus guide the learning of the RL model. Additionally, during the execution process, the RL model saves the sequence \(\{{S}_{t},{a}_{t},{R}_{t},{{{\rm{\phi }}}}_{t}\}\) (where \({{{\rm{\phi }}}}_{t}\) indicates whether t = T, i.e., whether it is the last item of the continuous state sequence) to the expertise replay pool \({D}_{{RL}}\). The RL model is then randomly sampled from \({D}_{{RL}}\) for learning. During training, the parameters of \({Q}_{{\mathrm{eval}}}\) are updated continuously using gradient descent after the loss function is calculated. In contrast, \({Q}_{{\mathrm{target}}}\) copies the parameters of \({Q}_{{\mathrm{eval}}}\) only after the completion of an epoch, and this process repeats at the end of each subsequent epoch. This asynchronous dual-network update design effectively prevents oscillations during training and enhances training stability. The optimization of the networks is conducted as follows:
The variable n represents the action with the highest reward value selected by Qeval under the current state St (specifically, the direction of the next move for the image sampling point). rtt is the expected reward that Qeval anticipates after executing the action. rt represents the actual reward received after executing the action, which is used to guide the training of the networks. It consists of two parts: the current return provided by fexperience and the future expected return estimated by Qtarget. The weight γ for the future return is set to 0.9. loss4 is a mean-squared error loss function. Its purpose is to bring the expected reward as predicted by Qeval closer to the actual reward, encouraging actions that maximize both present and future rewards. This allows the pathology expertise decoded from fexperience to be transferred to Qeval.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The 1565 pairs of data (including WSIs, gaze-tracking data, and manual pixel-wise annotations) declared in the section “Specifics of the dataset”, are publicly available. These data are divided into two parts: 150 pairs can be accessed directly, and 1415 pairs require an access application to be submitted to the corresponding author. The data that can be accessed directly has been stored in the GitHub database, and the access link is: https://github.com/MasyerN/PEAN. For access to the remaining data, interested parties need to submit a formal request to Professor Cui, the corresponding author. There are no potential restrictions on access to these additional data. The demonstration of PEAN imitating pathologists’ slide-reviewing behavior has been uploaded to the following link: https://www.tankaai.com/eponline/. We have also recorded the process of collecting pathologists’ slide-reviewing data and the operation of PEAN, which is presented as a demonstration video in the Supplementary Movies 3–10. Source data are provided with this paper.
Code availability
All the code involved in this study, including the open-source version of the “EasyPathology” system (v1.3, for reading data only) and the PEAN model code, will be made publicly available on open-source platforms. The code can be obtained through the following link: https://github.com/MasyerN/PEAN.
References
Cornish, T. C., Swapp, R. E. & Kaplan, K. J. Whole slide imaging: routine pathologic diagnosis. Adv. Anat. Pathol. 19, 152–159 (2012).
Madabhushi, A. Digital pathology image analysis: opportunities and challenges. Imaging Med. 1, 7 (2009).
Pantanowitz, L. et al. Review of the current state of whole slide imaging in pathology. J. Pathol. Inform. 2, 36 (2011).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. nature 521, 436–444 (2015).
Guo, Y. et al. Deep learning for visual understanding: a review. Neurocomputing 187, 27–48 (2016).
Pinckaers, H., Van Ginneken, B. & Litjens, G. Streaming convolutional neural networks for end-to-end learning with multi-megapixel images. IEEE Trans. Pattern Anal. Mach. Intell. 44, 1581–1590 (2020).
Tellez, D. et al. Neural image compression for gigapixel histopathology image analysis. IEEE Trans. Pattern Anal. Mach. Intell. 43, 567–578 (2019).
Li, J. et al. Hybrid supervision learning for pathology whole slide image classification. In International Conference on Medical Image Computing and Computer-Assisted Intervention (Springer International Publishing, Cham, 2021); https://doi.org/10.1007/978-3-030-87237-3_30.
Korbar, B. et al. Deep learning for classification of colorectal polyps on whole-slide images. J. Pathol. Inform. 8, 30 (2017).
Ghosh, A., Singh, S. & Sheet, D. Simultaneous localization and classification of acute lymphoblastic leukemic cells in peripheral blood smears using a deep convolutional network with average pooling layer. In 2017 IEEE International Conference on Industrial and Information Systems (ICIIS) (IEEE, 2017); https://doi.org/10.1109/ICIINFS.2017.8300425.
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Bejnordi, B. E. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199–2210 (2017).
Chen, Z. et al. Multi-instance multi-label image classification: a neural approach. Neurocomputing 99, 298–306 (2013).
Chen, R. J. et al. Multimodal co-attention transformer for survival prediction in gigapixel whole slide images. Proc. IEEE/CVF Int. Conf. Comput. Vis. https://doi.org/10.1109/iccv48922.2021.00398 (2021).
Yao, J. et al. Whole slide images based cancer survival prediction using attention guided deep multiple instance learning networks. Med. Image Anal. 65, 101789 (2020).
Ilse, M., Tomczak, J. & Welling, W. Attention-based deep multiple instance learning. International Conference On Machine Learning (PMLR, 2018).
Shao, Z. et al. Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv. Neural Inf. Process. Syst. 34, 2136–2147 (2021).
Zhang, H. et al. Dtfd-mil: Double-tier feature distillation multiple instance learning for histopathology whole slide image classification. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. (2022).
Lin, T. et al. Interventional bag multi-instance learning on whole-slide pathological images. In Proc. the IEEE/CVF Conference on Computer Vision and Pattern Recognition (IEEE, 2023); https://doi.org/10.48550/arXiv.2303.06873.
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Li, S. et al. Multi-instance multi-scale CNN for medical image classification. Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, October 13–17, 2019, Proceedings, Part IV 22 (Springer International Publishing, 2019); https://doi.org/10.1007/978-3-030-32251-9_58.
Hou, L. et al. Patch-based convolutional neural network for whole slide tissue image classification. In Proc. the IEEE Conference on Computer Vision and Pattern Recognition (IEEE, 2016); https://doi.org/10.48550/arXiv.1504.07947.
Kanavati, F. et al. Weakly-supervised learning for lung carcinoma classification using deep learning. Sci. Rep. 10, 9297 (2020).
Lerousseau, M. et al. Weakly supervised multiple instance learning histopathological tumor segmentation. Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, Peru, October 4–8, 2020, Proceedings, Part V 23 (Springer International Publishing, 2020); https://doi.org/10.1007/978-3-030-59722-1_45.
Chen, K., Sun, S. & Zhao, J. Camil: causal multiple instance learning for whole slide image classification. In Proc. the AAAI Conference on Artificial Intelligence, Vol. 38 (AAI, 2024).
Zhang, X., et al. Deep stable learning for out-of-distribution generalization. In Proc. the IEEE/CVF Conference on Computer Vision and Pattern Recognition (IEEE, 2021).
Geirhos, R. et al. Shortcut learning in deep neural networks. Nat. Mach. Intell. 2, 665–673 (2020).
Kaczmarzyk, J. R., Saltz, J. H. & Koo, P. K. Explainable AI for computational pathology identifies model limitations and tissue biomarkers. Preprint at https://arxiv.org/abs/2409.03080 (2024).
Deane, O., Toth, E. & Yeo, S. H. Deep-SAGA: a deep-learning-based system for automatic gaze annotation from eye-tracking data. Behav. Res. 55, 1372–1391 (2023).
Majaranta, P. & Bulling, A. Eye tracking and eye-based human–computer interaction. In Advances in Physiological Computing 39–65 (Springer, 2014); https://doi.org/10.1007/978-1-4471-6392-3_3.
Zhang, X. et al. MPIIGaze: real-world dataset and deep appearance-based gaze estimation. IEEE Trans. Pat. tern. Anal. Mach. Intell. 41, 162–175 (2019). 2778103.
Sugano, Y., Zhang, X. & Bulling, A. Aggregaze: Collective estimation of audience attention on public displays. Proceedings of the 29th annual symposium on user interface software and technology. 821–831 https://doi.org/10.1145/2984511.2984536 (2016).
Sattar, H. et al. Prediction of search targets from fixations in open-world settings. In Proc. the IEEE Conference on Computer Vision and Pattern Recognition 981–990 (2015); https://doi.org/10.48550/arXiv.1502.05137.
Kümmerer, M. Theis, L. & Bethge, M. Deep gaze I: boosting saliency prediction with feature maps trained on ImageNet. In CoRR abs/1411.1045. Preprint at https://arxiv.org/abs/1411.1045 (2014).
Kümmerer, M., Theis, L. & Bethge, M.Deep gaze I: boosting saliency prediction feature maps trained ImageNet. Preprint at https://arxiv.org/abs/1411.1045 (2014).
Kümmerer, M., Wallis, T. S. A. & Bethge, M. DeepGaze II: reading fixations from deep features trained on object recognition. Preprint at https://arxiv.org/abs/1610.01563 (2016).
Kümmerer, M. Wallis, T. S. A. & Bethge, M. DeepGaze III: using deep learning to probe interactions between scene content and scanpath history in fixation selection. In 2019 Conference on Cognitive Computational Neuroscience. (2019).
Mariam, K. et al. On smart gaze based annotation of histopathology images for training of deep convolutional neural networks. IEEE J. Biomed. Health Inform. 26, 3025–3036 (2022).
Brunyé, T. T. et al. Eye tracking reveals expertise-related differences in the time-course of medical image inspection and diagnosis. J. Med. Imaging 7, 051203–051203 (2020).
Li, B., Li, Y. & Eliceiri, K. W. Dual-stream multiple instance learning network for whole slide image classification with self-supervised contrastive learning. In Proc. the IEEE/CVF Conference on Computer Vision and Pattern Recognition (IEEE, 2021).
Lin, T. et al. Interventional bag multi-instance learning on whole-slide pathological images. In Proc. the IEEE/CVF Conference on Computer Vision and Pattern Recognition (IEEE, 2023).
Theckedath, D. & Sedamkar, R. R. Detecting affect states using VGG16, ResNet50 and SE-ResNet50 networks. SN Comput. Sci. 1, 1–7 (2020).
Deng, J. et al. Imagenet: a large-scale hierarchical image database. 2009 IEEE Conference on Computer Vision and Pattern Recognition (IEEE, 2009); https://doi.org/10.1109/CVPR.2009.5206848.
Lu, M. Y. et al. A visual-language foundation model for computational pathology. Nat. Med 30, 863–874 (2024).
Kaelbling, L. P., Littman, M. L. & Moore, A. W. Reinforcement learning: a survey. J. Artif. Intell. Res. 4, 237–285 (1996).
Arulkumaran, K. et al. Deep reinforcement learning: a brief survey. IEEE Signal Process. Mag. 34, 26–38 (2017).
Barata, C. et al. A reinforcement learning model for AI-based decision support in skin cancer. Nat. Med. 29, 1941–1946 (2023).
Zhao, B. et al. RLogist: fast observation strategy on whole-slide images with deep reinforcement learning. In Proc. AAAI Conference on Artificial Intelligence, Vol. 37 (AAAI, 2023); https://doi.org/10.1609/aaai.v37i3.25467.
Nyström, M. et al. The Tobii Pro spectrum: A useful tool for studying microsaccades?. Behav. Res. Methods 53, 335–353 (2021).
Ester, M. et al. Density-based spatial clustering of applications with noise. In Proc. International Conference on Knowledge Discovery and Data Mining 226–231 (ICMLC, 1996).
Acknowledgements
This work was supported by grants from the China Key Research and Development Program (grant no. 2023YFC2508200, Professor X Cui) and the Liaoning Province Medical Engineering Cross Joint Fund (grant no. 2022-YGJC-76, Professor X Cui).
Author information
Authors and Affiliations
Contributions
The authors T.N. and S.Z., have the same contribution. The authors R.Q., Xinghua G and X.C. are corresponding authors. T.N. developed the concept and subject of the paper. S.Z. led the development of the method. S.Q. developed the main software. H.Q. participated in the construction of the software. Xinghua G. participated in the review of manuscripts. J.N., B.Z., C.G., Y. Zhang, X.W., and L.Z. collected the dataset. Z.W., Y.G., X.L., M.Z. and S.N. participated in experimental design. Y. Zhao and H.C. participated in the writing of the manuscript. R.Q., Xinghua G. and X.C. led this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Mehmet Turan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nan, T., Zheng, S., Qiao, S. et al. Deep learning quantifies pathologists’ visual patterns for whole slide image diagnosis. Nat Commun 16, 5493 (2025). https://doi.org/10.1038/s41467-025-60307-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60307-1
This article is cited by
-
Machine learning-based radiopathomics ensemble model for predicting multiple molecular subtypes of adult diffuse glioma: a multicenter retrospective study
BMC Cancer (2026)
-
Quantum-enhanced multimodal prognostic transformer for skin disease progression prediction and visualization
Scientific Reports (2026)
-
Explainable federated transformer framework for joint leukemia classification and stage prediction
Scientific Reports (2026)