Abstract
DNA folding thermodynamics are central to many biological processes and biotechnological applications involving base-pairing. Current methods for predicting stability from DNA sequence use nearest-neighbor models that struggle to accurately capture the diverse sequence dependence of secondary structural motifs beyond Watson-Crick base pairs, likely due to insufficient experimental data. In this work, we introduce a massively parallel method, Array Melt, that uses fluorescence-based quenching signals to measure the equilibrium stability of millions of DNA hairpins simultaneously on a repurposed Illumina sequencing flow cell. By leveraging this dataset of 27,732 sequences with two-state melting behaviors, we derive a NUPACK-compatible model (dna24), a rich parameter model that exhibits higher accuracy, and a graph neural network (GNN) model that identifies relevant interactions within DNA beyond nearest neighbors. All models show improved accuracy in predicting DNA folding thermodynamics, enabling more effective in silico design of qPCR primers, oligo hybridization probes, and DNA origami.
Similar content being viewed by others
Introduction
The thermodynamics of DNA secondary structure formation is fundamental to understanding diverse biological processes, such as DNA replication1 and repair2. It also underlies many biotechnological applications, including PCR primer design3, in-situ hybridization (ISH) probe engineering4, guide sequence design for genome editing tools5, and DNA nanotechnology6,7. Numerous algorithms have been developed to predict DNA secondary structure thermodynamics, many of which rely on nearest-neighbor models8,9,10.
In brief, nearest-neighbor models assume that the total folding energy of DNA can be calculated by summing up the energies of each two neighboring base pairs. With this assumption, the “folded state” can be defined as the minimum free energy (MFE) base pairing configuration, identified using dynamic programming11. Additionally, by using Boltzmann factors and the partition function, it is possible to calculate the probabilities of DNA folding into alternative base pairing states. The predicted probability of each state is determined based on its computed free energy. These energies contribute to Boltzmann factors that are normalized by the partition function, the sum of the Boltzmann-weighted contributions of all possible states. Thus, nearest-neighbor models form the foundation of both static secondary structure predictions and dynamic ensemble folding predictions. However, nearest-neighbor models often struggle to accurately capture the sheer diversity and complexity of DNA secondary structural motifs, including hairpin loops, mismatches, and bulges. One reason for this limitation is the lack of experimental data spanning those structural motifs upon which these models are built, leading to inaccuracies in prediction. For example, the most widely used parameter set from SantaLucia et al. 2004 used data from 108 sequences to derive 12 parameters for Watson-Crick (WC) base pairs, and 174 sequences to derive 44 parameters for internal single mismatches8.
This data bottleneck has been due to the laborious nature of UV melting12 and differential scanning calorimetry13 experiments, traditionally considered the gold standards of DNA secondary structural thermodynamics. Several groups have attempted to overcome this bottleneck, usually with fluorescent melting in bulk solutions in well plates14,15,16,17, reaching a throughput on the scale of thousands. Still, due to the extremely large combinatorial DNA sequence space, these works are limited to a specific type of DNA structural motif in a fixed sequence scaffold.
To address this data generation bottleneck, we developed a method for systematic, accurate, high-throughput measurements of nucleic acid secondary-structure motif thermal stability, enabling large-scale, quantitative measurements of the thermodynamics of nucleic acid secondary structure. We demonstrate that nearest-neighbor thermodynamic parameters inferred from these data are nearly identical to those observed from bulk thermal melting experiments for WC base pairs but improved for mismatch, bulge, and hairpin loop structural motifs. The improved parameter set is capable of accurately predicting DNA duplex folding energy on fully independent datasets collected using different measurement methods. Furthermore, by deploying advanced computational methods including deep learning to model DNA thermodynamics, we develop a state-of-the-art thermodynamic model for DNA hairpin folding with accuracy comparable to measurement uncertainties.
Results
Array melt: A high-throughput technique for quantifying the melting behavior of nucleic acids
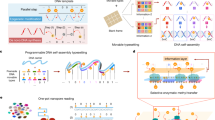
We developed the Array Melt technique, a method that allows for the rapid and comprehensive quantification of DNA folding thermodynamics in high throughput. This technique is based on an Illumina MiSeq chip repurposed for high-throughput measurements18,19. We first designed a DNA library of 41,171 hairpin sequences, referred to as “variants” in 6 major categories, referred to as “classes”. This library was synthesized as an oligo pool, amplified with sequencing adapter sequences, and loaded onto a MiSeq chip for sequencing (Methods). During Illumina sequencing, single DNA molecules on the flow cell surface were amplified into groups of approximately 1000 copies of the same sequence, which we refer to as “clusters”. We mapped each cluster to a sequence variant in the library using the physical locations in sequencing data. The variable region of a variant consists of a DNA hairpin flanked by two “AA” linkers and two oligo binding sites. We engineered a common region for annealing a 3’-fluorophore-labeled oligonucleotide to the 5’-end of the hairpin to be investigated and another for a 5’-quencher-labeled oligonucleotide to the 3’-end (Fig. 1a). During an imaging experiment, we annealed Cy3-labeled and Black Hole Quencher (BHQ)-labeled oligonucleotides to these respective binding sites to assay the melting behavior of the hairpins. These annealed oligos have predicted melting temperatures of 74 °C, much higher than the highest temperature in our experiments (Supplementary Table 1). At room temperature, this fluorophore-quencher pair results in low levels of fluorescence in each cluster, as the bulk of our hairpin library is folded at room temperature. As the sequence-variable hairpin regions are exposed to increasing temperatures between 20 °C to 60 °C, the distance between Cy3 and BHQ increases, leading to brighter fluorescence signals (Fig. 1a, b). Using this measurement scheme, we synthesized a comprehensive library of 41,171 hairpin variants for measurement (Fig. 1c). These variants were designed by integrating diverse “structural motifs” into multiple constant hairpin “scaffolds”. The secondary structural motifs include Watson-Crick pairs, mismatches, bulges, hairpin loops of various lengths, and other control sequences such as super-stable stems, polynucleotide repeats, and single-strand fluorescence controls (see Fig. 1c and Methods). We inserted each structural element into multiple hairpin scaffolds with varying energetic stabilities to increase the likelihood that at least one variant would fall within the dynamic range of our measuring system. It is important to note that some variants share identical sequences. For example, 16 variants in the “tetraloop” and the “Watson Crick” classes have identical sequences, despite originating from different combinatorial enumerations series. The 16 “tetraloop” variants were generated by enumerating the hairpin loop sequence and the two base pairs adjacent to the loop, and the 16 “Watson-Crick” variants by enumerating the hairpin stem while keeping the loop sequence constant. In the library design, we still counted them as separate variants to preserve the completeness of enumerations; in the dataset, we merged variants with identical sequences to remove duplicated data for analysis. In total, there are 40,130 unique sequences in this library of 41,171 variants.
a Schematic representation of DNA molecules in a folded (quenched) and an unfolded (fluorescent) state. b Images of fluorescent DNA clusters immobilized on the sequencing chip surface. Top: Image with only fluorophore-conjugated oligo (Cy3), and with both fluorophore- and quencher-conjugated oligo at increasing temperatures. Bottom: Schematics of DNA molecules in each image. All images are normalized to super-stable stem and repeat control variants for temperature-dependent effects on fluorescence and quenching during the experiment, as shown in Supplementary Fig. 1b. c Design of library variants between the constant-sequence binding sites for fluorophore- and quencher-conjugated oligonucleotides. Red represents scaffold nucleotides, which are constant within each type, and blue represents the structural motif nucleotides, or variable nucleotides systematically permuted (‘N’). Numbers under each class indicate the number of variants in each class. d Fluorescence measurement of control constructs where the single-strand distance between the fluorophore and the quencher increases in single nucleotide steps. The orange line shows the theoretical fit. Fluorescence is measured in arbitrary units (a.u.). e Correlation of ΔG37 derived from library variants under increasing temperatures (melt curve, x-axis) and decreasing temperatures (annealing curve, y-axis). f Representative examples of melt curves for constructs that vary in GC content. Melt curves are calculated from fitted parameters based on data collected from individual clusters (n = 88, 126, and 90 for the three constructs, respectively). Shaded regions represent 95% confidence intervals derived from bootstrapped parameter estimates. Fluorescence is normalized and reported in arbitrary units (a.u.). g Standard error of ∆G37 for various construct classes, colored by ∆G37 values. Standard errors are derived from bootstrapped errors combined across replicates (Methods). Box plots show the median (center line), interquartile range (box bounds: 25th to 75th percentiles), and full range excluding outliers (whiskers extend to the most extreme non-outliers; outliers are hidden). Only series with more than 1000 variants are drawn. h Percentage of variants in each class that fall in different ΔG37 standard error bins. Unless otherwise noted, box plots show the median, 25th–75th percentiles, and range excluding outliers.
We performed thorough quality control on the dataset by requiring clusters and variants to accurately fit a two-state model, and to melt within our temperature measurement range. The two-state model assumes that a given sequence variant exists either in an “unfolded” or a fully “folded” state. This assumption contrasts with an ensemble model, where a variant is modeled to occupy an ensemble of base pairing states, including the unfolded, fully folded, and multiple alternatively folded states, with the probability of each secondary structure state determined by its corresponding free energy. The two-state model is the foundation of nearest-neighbor parameter derivation and serves as the focus of this study. For each variant, we used a two-state model to fit the melt curves to determine ΔH and Tm, then calculated ΔG37 and ΔS from ΔH and Tm (Eqs. (2)–(4), Methods). First, we normalized melt curves of single clusters to the initial fluorescence after Cy3 hybridization to account for cluster size variations and sequence-dependent effects. We further normalized data using control variants to account for temperature dependency and photobleaching (“Fluorescence normalization”, Methods). Next, we fitted the signals to Eq. (2) and estimated the probability distributions of maximum and minimum fluorescence (fmax and fmin) (“Single cluster fitting”, Methods). Then, we refined the fit by bootstrapping single clusters at the variant level, using information from the distribution of fmax or fmin to aid the variants not fully reaching fmax or fmin (“Fit refinement”, Methods). Finally, we combined results from multiple replicates, removing variants not showing enough melting behavior or having high errors (“Filtering and combining replicates”, Methods) and filtering out non-two-state variants (“ΔG line fitting and two-state heuristics”, Methods). Data that met these quality control criteria comprised 6,393,050 individual melt curves of single clusters (including multiple technical replicates), spanning 27,732 sequence variants used in subsequent analysis with a standard two-state melt model (Supplementary Fig. 1a).
Sensitivity and precision of the Array Melt method
To calibrate the amount of fluorescence observed as a function of the distance between Cy3 and BHQ, we evaluated the fluorescence signals of a series of repeat sequence variants. These variants were designed to systematically increase the distance between the fluorophore and quencher by incorporating increasing numbers of mono-, di- or tri-nucleotide repeats. The fluorescent signals, aggregated over repeats of the same length, showed a nearly linear response to distance variations up to approximately 8 nucleotides (Fig. 1d). Our observed length-fluorescence curve closely aligns with a theoretical static quenching curve (Eq. [1], Methods). We normalized the fluorescent signals using unfolded and folded controls designed within the library (Supplementary Fig. 1b). The three example melt curves in Fig. 1f demonstrated increasing observed melting temperatures corresponding to increasing GC content of the variable region. For variants that showed complete melting behavior, such as the variant on the left, we directly inferred the maximum and minimum fluorescence (fmax and fmin) from the melting curves; for variants that did not fully reach fmax or fmin, such as the middle variant, we estimated the distributions of fmax or fmin from an initial round of fitting to aid the refine fit process (Supplementary Fig. 1c; Methods); for variants without enough melting behavior, such as the one on the right, we excluded them from downstream analysis. Technical replicate measurements from Array Melt data showed high correlation, with R > 0.94 (Supplementary Fig. 1d). Melt curves were also highly correlated with anneal curves (R = 0.964), confirming that the DNA molecules were measured at equilibrium (Fig. 1e). Upon analyzing our measurement precision (Methods), we found that variants with \(\Delta {G}_{37}\) values between −1.5 and 0.5 kcal/mol have tight uncertainty levels around 0.1 kcal/mol (Fig. 1g). Additionally, nearly 90% of tested variants have \(\Delta {G}_{37}\) uncertainties within 0.3 kcal/mol, equivalent to approximately 0.5 kBT (Fig. 1h). Overall, these data demonstrate that the equilibrium, aggregate fluorescence signals obtained from the Array Melt method provide highly accurate melting information and can serve as a “molecular ruler” capable of resolving single nucleotide distance variations.
UV melting validation
To further validate our method, we compared the results from the Array Melt method to those obtained using conventional UV melting methods on a cloud lab platform. We selected random representative variants from each class within the Array Melt library and sent the synthesized oligonucleotides to Emerald Cloud Lab (ECL) for analysis. Using the variant information and UV melting experiment parameters as input, scripts written in Symbolic Lab Language (SLL) automatically generated experiments as ECL protocol objects. These protocols were then queued for remote execution in a physical laboratory. Raw melting curve data from ECL were automatically analyzed by custom Python code, and samples that failed to pass quality control were resubmitted to ECL for re-measurement (Fig. 2a, b; Methods). We synthesized the variable region of randomly sampled Array Melt variants and measured UV melting curves in the same buffer as in Array Melt experiments (25 mM NaCl, 50 mM HEPES pH8.0). We aimed to ensure good correlations between UV melting and Array Melt, and to correct for any systematic errors in the Array Melt system. As expected, the melting temperature was found to be independent of DNA oligo concentration (Supplementary Fig. 2a), suggesting intramolecular folding. In total, we measured 85 unique sequences, including 66 from the Array Melt library and 19 de novo hairpins designed independently of the Array Melt library. 24 of these sequences had a single peak on HPLC, including 12 from the Array Melt library and 12 de novo hairpins. UV melting data showed slightly worse reproducibility across experimental days than Array Melt (Supplementary Fig. 2b, c). A comparison of the melting profiles of measured library variants showed good agreement between the two methods (Fig. 2c–e, R = 0.85 for ΔG37, or R2 = 0.62 after the linear adjustment described later). Notably, we observed a systematic offset between the UV measurements and our Array Melt results, which could be due to the presence of the hairpins on a flow-cell surface, interaction between the fluorophore and the quencher, or the additional sequence elements and double-stranded areas included to facilitate our quenching-based readout. To account for these systematic differences, we applied a single offset to the melting points observed in our dataset and used the corrected results for subsequent analysis (Methods; on average, a correction of 9.35 °C in Tm or ~0.7 kcal/mol in ΔG37 was applied). Compared to baseline nearest-neighbor model predictions implemented in the NUPACK4 (NUPACK v4.0.0.27) software (“the dna04 model”, an abbreviation for “the model implemented in NUPACK4 using the dna04 parameter set”), the measured values showed moderate improvement for ΔG and Tm, and considerable improvement for ΔH (Fig. 2e, Supplementary Fig. 2d). This UV melting validation demonstrated the ability of the Array Melt method to accurately measure the folding energies of DNA and generalize to in-solution systems.
a Schematic representation of UV melting data collection and analysis using the Emerald Cloud Lab (ECL) platform. Magenta arrows indicate semi-automatic steps, and teal arrows indicate automatic steps. b Example UV melting curve of the variant WC2318. Pink dots represent measured data points of the cooling curve; the green solid line represents the fitted curve; the green dotted line represents the fitted melting temperature. c Mean absolute error (MAE) of ΔG, ΔH, and Tm between Array Melt data and UV melting data before and after systematic error correction. Gray bars represent errors before systematic correction; colored bars after applying correction with a single offset parameter. d Direct comparison of measured ΔG37, ΔH, and Tm between Array Melt and UV melting data. Each datapoint represents one hairpin variant. Error bars indicate Array Melt bootstrapped standard errors (y-axis). Gray dots are before systematic correction; colored dots are after. e Pearson’s correlation coefficient (R) between UV melting data and either Array Melt data or the dna04 model predictions. The colored bars represent the correlation between Array Melt and UV melting data; the black bars represent that between the dna04 model predictions and UV melting data.
Quantifying performance of the dna04 nearest-neighbor model on non-WC hairpins
After validation of our experimental system, we aimed to examine the performance of the baseline model, the dna04 nearest-neighbor model, on hairpins whose stems are not perfect Watson-Crick pairs. The dna04 model, implemented with NUPACK4, uses widely used nearest-neighbor parameters from the SantaLucia 2004 paper8 and incorporated additional parameters for coaxial and dangle stacking. When predicting folding free energies associated with different single mismatch types (Fig. 3a), the dna04 model had predictions with an \({R}^{2}\) of 0.61 with Array Melt data on the average free energy for each of the types (implying it can explain 61% of the observed experimental variance), or \({R}^{2}=0.31\) on all individual variants. However, this model was unable to consistently predict significant variances in folding energies that arise from the same mismatch type with differing flanking sequences (\({R}^{2}=-0.61\) for T > C mismatches, as shown on the right side panel, \({R}^{2}=0.09\) for A > G, \({R}^{2}=-0.48\) for C > T, and \({R}^{2}=-1.66\) for G > A) (Fig. 3b, Supplementary Fig. 3a). Indeed, for three out of the four mismatches we analyzed, the dna04 model introduced extra variance (i.e., negative \({R}^{2}\) value) compared to the class-average prediction, implying that the variance in the dna04 model’s prediction residual was greater than the actual variance present in the data (Fig. 3c, Supplementary Fig. 3b). This limitation of the dna04 model led us to conclude that significant variances in folding energy are not accounted for by the dna04 model, likely due to limitations in its parameterization.
a Example of mismatch type definitions and ∆∆G calculation for single mismatches. b Comparison of single mismatch ∆∆G between the dna04 model prediction and Array Melt measurements. Each variant is colored by the identity of the mismatch (left), and the T > C mismatches are zoomed in on the right. c Variances in data (technical and biological) and variances explained or added by the dna04 model, for each group of mismatches and combined. Green bars, variance explained by the dna04 model; red bars, additional variance introduced. d Nucleotide importance scores for each individual nucleotide in mismatch constructs, grouped by secondary structure. The importance score is calculated as the sum of squares introduced by varying the given nucleotide position. e Nucleotide importance scores for tetraloops and triloops. f Nucleotide importance scores for bulges. g Pairwise interaction scores for each pair of nucleotide locations for Watson-Crick and single mismatch variants. A yellow star indicates the reference position in each subplot, and the color of each position indicates the strength of the interaction with that reference position. The interaction score is calculated as the R2 of a linear model given all combinations of the pair as features, subtracted by that of the two individual nucleotides. h Percentage contribution of single nucleotides, nearest neighbors, and longer-range interactions to the total variance in ∆∆G37 for single mismatches. i Pairwise interaction scores for tetraloops and triloops.
Influence of sequences on energy across different secondary structures
We hypothesized that some of the observed variance in melting behavior not explained by the dna04 model may be due to the influence of next-nearest neighbors of mismatches. To assess this possibility, we calculated the impact of each nucleotide position on folding energy. We first determined a nucleotide importance score at each nucleotide position in different sequence groups with the same target secondary structures by calculating the variance in either ΔG37 or ΔΔG37 values for the four different identities of the nucleotide at the target position (Methods). The importance scores were calculated for every variable nucleotide position but not for the constant ones, such as the bottom base pair and the constant hairpin loop sequence. While it is possible that variants might adopt alternative secondary structures yet maintain two-state behavior, our analysis still provides valuable insights into how sequences affect folding energy. For single mismatches, we used ΔΔG37 values relative to Watson-Crick pairing “parent” variants to regress out the energy expected from Watson-Crick stacks; for other secondary structures, we directly used ΔG37 values. As anticipated, nucleotides along the hairpin stem displayed consistent nucleotide importance scores for Watson-Crick pairing variants, and the two mismatching nucleotides have the highest contribution to this folding energy for single mismatch variants, both in the Array Melt data and the dna04 model predictions (Fig. 3d, Supplementary Fig. 3c). Extending our analysis to hairpin loops and bulges, we observed that specific sequences within these structures critically influence their thermodynamic energies (Fig. 3e, f). In triloops and tetraloops, loop center nucleotides had a more pronounced influence on folding energy in the Array Melt data than dna04 model predictions. We also noted a slight asymmetry between the 5’ and 3’ ends of the loop, regardless of loop size (Fig. 3e, Supplementary Fig. 3d). In bulges, the flanking Watson-Crick pairs had greater influence than the bulge sequence itself. However, double bulges showed stronger sequence dependence than single bulges, suggesting greater sensitivity to bulge sequence in the double-bulge structural motifs (Fig. 3f, Supplementary Fig. 3e).
These observations prompted us to examine the energetic interactions between bases by comparing two linear models: one that considers base interactions (i.e., allowing free parameters for pairwise combinations of bases in the two probed positions) and another that treats each position independently (i.e., only allowing free parameters for each individual position). The “pairwise interaction score” was calculated as the additional proportion of variance explained in the first model than the second. In the figures, we used a star to denote the reference position relative to which the interaction scores are plotted. The location of the star marks all the variable nucleotide positions, which may or may not correspond to a mismatch. As the pairwise interaction scores are symmetric, we only plotted the 5’ starred nucleotide positions to simplify the visualization (Fig. 3g). For comparison, we also calculated pairwise interaction scores from dna04 model predictions (Supplementary Fig. 3f). Notably, the two base pairs flanking a mismatch showed interactions unaccounted for in the dna04 nearest-neighbor model, which only considers interactions between the mismatched base pair and its immediate neighbors. This interaction score decays with increasing distance along the stem (Supplementary Fig. 3g). Our quantification showed that in this specific case of single mismatches, 74% of the variance was contributed by individual nucleotides, 14% by nearest-neighbor interactions, and 12% by longer-range interactions (Fig. 3h). This breakdown highlights that while individual nucleotides and nearest-neighbor interactions account for up to 88% of the variance, longer-range interactions contribute comparably as nearest-neighbor interactions. In hairpin loops, the interactions between the loop-end and the loop-center nucleotides were stronger than expected in tetraloops, but not in triloops (Fig. 3i, Supplementary Fig. 3h, i).
Next, we fitted a series of linear regression models to the Watson-Crick and single-mismatch variants classes in Array Melt data, varying the length of stack features: from 1 (individual base pairs), to 2 (nearest neighbors), and up to 3 (next-nearest neighbors). In the model with only individual base pairs, each individual base pair or mismatch was treated as a feature. In the nearest-neighbor model, each stack of 2 consecutive base pairs or mismatches was treated as a feature. Note that this model was independent of existing nearest-neighbor models, such as the dna04 model, as it was constructed from scratch for comparison purposes. The next-nearest-neighbor model, in contrast, parameterized every stack of 3 consecutive base pairs or mismatches. Although prediction errors were similar for nearest- and next-nearest-neighbor models in Watson-Crick variants, next-nearest-neighbor models outperformed the nearest-neighbor ones in single-mismatch variants (Fig.4a). This observation suggested that while nearest-neighbor models are sufficient to describe folding energies of Watson-Crick variants, incorporating next-nearest neighbor interactions is crucial for accurately modeling variants that contain mismatches.
a Adjusted mean absolute error (adjusted MAE) of linear regression models with features based on single-base-pair, nearest-neighbor, or next-nearest-neighbor stacks (length of stack features = 1, 2, and 3, respectively), applied to Array Melt test set. b Comparison of nearest-neighbor parameters for Watson-Crick pairs fitted from Array Melt data and those reported in the literature8. c Visualization of dna24 and dna04 free energy parameters for single mismatches. “Context” and “mismatch” definitions are consistent with those in Fig. 3a. Note: The flat GT/TG mismatch columns in the dna04 panel result from NUPACK’s use of placeholder (dummy) parameters for these mismatches. This issue is due to a historical artifact in the parameter files, which the NUPACK development team has acknowledged and is currently addressing (Supplementary Discussion; personal communication, N. Pierce, 2025). The notes also apply to Supplementary Fig. 3c. d, e Performance comparison of the dna04, dna24, and rich parameter models on held-out Array Melt data. All calculations were performed at 1 M Na+ concentration. f Variances in data (technical and biological) and variances explained or added by the models, for all Array Melt test data, or Watson Crick, mismatch, and bulges only. g Performance comparison of the dna04, dna24, and rich parameter models on an orthogonal dataset of DNA duplexes with varying numbers of mismatches (Oliveira et al. 15.). h Adjusted MAE on Array Melt data plotted as a function of the percentage of training data used to fit the linear regression models.
Revised dna24 parameter set for NUPACK integration
We aimed to develop a refined set of parameters to integrate our findings with the NUPACK4 software18, thereby enabling more accurate DNA thermodynamic predictions that are easily accessible. We divided Array Melt data into training, validation, and test sets, respectively with 24979, 1315, and 1438 variants, stratified by major variant types as shown in Fig. 1c. Initially, we wanted to verify that we could reproduce the established Watson-Crick nearest-neighbor parameters8. We fitted a compact linear regression model with 188 free parameters to the Array Melt data, allowing the 10 Watson-Crick pairing stack parameters to vary freely. While other parts of the parameter set, such as mismatches and bulges, were also adjusted in this model, the WC stacking parameters still remained broadly consistent with the literature8 (Fig. 4b). Some minor differences were expected since they were trained on completely different real-world datasets. Having confirmed the reliability of our approach, we chose to fix the values of the Watson-Crick nearest-neighbor parameters for the remainder of the analysis, as they are widely used, rigorously validated, and offered minimal accuracy improvements when adjusted. With these 10 parameters fixed, we fitted a compact 178-parameter model. Next, we obtained sequence-dependent parameters for tetraloops and triloops by aggregating thermodynamic measurements of relevant Array Melt variants and converting the aggregated ΔΔH and ΔΔG37 values to ΔH and ΔG37 parameters (Methods). This additional curated step overcame the bias of hairpin loop sequence sampling, as most of the variants in the Array Melt library has the same “GAAA” tetraloop sequence. We combined these corrected hairpin loop parameters with the parameters from the previously fitted compact 178-parameter model. This combined NUPACK-compatible model, referred to as “the dna24 model” (the model implemented with NUPACK4 using “the dna24 parameter set”), is easily accessible by pointing NUPACK to our parameter set file (Supplementary Fig. 4a; parameter file available in Supplementary Data19). Note that when using NUPACK to predict free energy, we assumed a two-state model in alignment with the Array Melt dataset, which was filtered based on the two-state criterion. Additionally, the ensemble model yielded inferior results compared to the two-state model, both with the original dna04 and with our new parameter sets (Supplementary Fig. 4b).
We compared the single mismatch parameters of this dna24 model with the original dna04 one and observed similar overall trends. Our new parameters also captured context-dependent variations of GT mismatches (also referred to as wobble pairs) and highlighted the instability of CC mismatches, which are the most thermodynamically unstable type of DNA single mismatch20. The dna24 parameters also agreed with the literature about the context-dependency of GT mismatches, in that CG×CG is the most stable context and AT×AT the least stable20 (Fig. 4c, Supplementary Fig. 4c).
We evaluated the dna04 and the dna24 models on held-out Array Melt data and two orthogonal DNA duplex melting datasets from the literature. To assess model performance, we reported both the raw mean absolute error (MAE) and the adjusted MAE, which specifically accounts for relative ΔΔG errors. As a figure of merit, relative ΔΔG values are often more common than absolute ΔG values for many applications, including minimum free energy structure predictions and sequence design tasks, as they consider the relative energy differences between sequence-structure pairs (Supplementary Discussion). The readers should choose between MAE and adjusted MAE based on the relevant application.
Testing our dna24 model on the held-out Array Melt test data resulted in an adjusted MAE of 0.59 kcal/mol for ΔG37 (Fig. 4d, e, Supplementary Fig. 4f). Consistently, the dna24 model explained a larger fraction of the biological variance in ∆G37 across the test set, especially for mismatch and bulge classes (Fig. 4f). This model also improved predictions for 12 DNA hairpins designed with randomized sequences outside the Array Melt library and measured by UV melting (Supplementary Fig. 4d), alongside the hairpin sequences taken from the Array Melt library and measured using UV melting in Fig. 2. We further asked whether the dna24 model, trained on DNA hairpin data, extends to unseen DNA duplexes. We observed a similar performance for unseen Watson-Crick DNA duplexes with both the dna04 and the dna24 models. However, the dna24 model showed improved performance on more complex variants. Next, we evaluated our parameter set on a UV melting dataset of 384 Watson-Crick pairing DNA duplexes21. This collection contains duplexes used to derive the SantaLucia parameters in dna04, and was split evenly into validation and test sets. Here, our parameter set performed comparably to dna04 (Supplementary Fig. 4e). On a dataset from Oliveira et al. containing DNA duplex melting temperatures with varying numbers of mismatches15 (also split evenly between validation and test sets), both parameter sets showed similar adjusted MAE (~0.6 °C) for Watson-Crick duplexes on the test set. For single mismatches, the dna24 parameter set achieved a much lower adjusted MAE of 1.15 °C (\({R}^{2}=0.68\)) compared to 1.66 °C (\({R}^{2}=0.25\)) with dna04. For double and triple mismatches, the improvement was even greater (Fig. 4g). In conclusion, this updated parameter set, while simple, enhances the prediction accuracy of NUPACK for DNA secondary structures, particularly those with mismatches and hairpin loops. This improvement extends to orthogonal DNA duplex data that the dna24 model was not trained on.
Enhanced linear model with rich parameterization
Building on this NUPACK-compatible dna24 model, we next developed an enhanced linear model that incorporates a rich parameter set (“rich parameter model”), allowing still more accurate and comprehensive prediction of DNA thermodynamics on unseen data. Because this model had more parameters than are implemented in NUPACK, it was not immediately compatible with the software and was implemented in custom Python code. We fitted this rich parameter model with 1338 free parameters to the Array Melt training data, incorporating still more context information. This rich parameter model has a sequence-dependent parameter for each combination of single or double bulge and its two flanking base pairs, while the dna24 model is not aware of the sequence of the bulged nucleotides; it also considers interactions between the two stacks flanking a mismatch, and full hairpin loop sequence for triloops and tetraloops (Methods). This model yielded an even better performance both on the Array Melt test data (adjusted \({MAE}=0.34\,{kcal}/{mol},{R}^{2}=0.81\)) and the external duplex datasets (Fig. 4d–g, Supplementary Fig. 4e). These extended parameters were particularly important for variants with bulges and mismatches, as they allowed the model to better account for the complex context-specific interactions in these cases. Examples of the origins of such interactions include base stacking, as the specific sequence of the bulge or mismatch influences the extent of disruption to base stacking, and some mismatched base pairs can still stack with the flanking base pairs depending on their sequences. Other factors that contribute to the sequence-dependent trends observed here include hydrogen bonding, electrostatic forces, and hydrophobic forces, among others. The rich parameter model explained significantly more variance in mismatches and bulges than dna24, especially for bulges (Fig. 4f). Notably, when comparing the prediction error on the Array Melt validation set as we titrate the amount of training data, the 178-parameter linear regression model used to build the dna24 model plateaued with just a small fraction of the training data. In contrast, the 1338-parameter rich parameter model used more data before plateauing, but still did so after being trained on less than half of the dataset (Fig. 4h). This behavior is expected given the overdetermined nature of these models, suggesting that a more expressive model is needed to capture more variance in the dataset.
Graph neural network (GNN) for DNA energy prediction
Because our rich parameter model likely had insufficient expressiveness to capture the biological variance in our data, we developed a graph neural network model22,23 specifically designed to predict energies of DNA sequence-structure pairs. In this model, a DNA molecule was represented as a graph where nodes represent nucleotides and edges represent chemical bonds. Nucleotides (A, T, C, or G) and chemical bonds (either 5’ to 3’ backbone, 3’ to 5’ backbone, or hydrogen bond) were one-hot encoded as node and edge features, respectively. Taking these graph representations with both sequences and structures as inputs, the GNN iteratively refined node embeddings across graph convolutional layers. Essentially, nodes exchanged information through different edge types, allowing, over multiple layers, for the embedding of one node to encapsulate information from the entire graph. Investigation of various types of graph convolutional layers revealed that the graph transformer layers24 outperformed other architectures, such as graph attention layers25 (Supplementary Fig. 5a). After graph convolution, each node possessed a learned embedded feature, reflecting not only the original node features, but also the edges and overarching graph structures. To embed this graph into a fixed-length vector representation, we used a global pooling layer with the Set2Set algorithm26, which included a Long Short-Term Memory (LSTM) block. This block processed node embeddings and updated its internal state to produce a vector representation of the entire graph invariant to the order of input node features. This processed summary vector was then fed into a small, fully connected neural network, which regressed for ΔH and Tm values (Fig. 5a). Similar to nearest-neighbor models, this GNN model predicted folding energy values from sequence-structure pairs.
a Schematic representation of the graph neural network (GNN) architecture. b Mean absolute error (MAE) of melting temperature on Array Melt data as a function of the number of graph convolution layers in the GNN. c MAE of ∆G37 on Array Melt data as a function of the percentage of training data used to fit the GNN. d Pearson’s R correlation coefficient between GNN predictions and held-out Array Melt data, literature UV melting data, or Oliveira et al. duplex melting data. e Scatter plot comparing GNN predictions with held-out Array Melt test data, both combined and split by variant class. Each dot represents a single sequence variant. f Scatter plot comparing GNN predictions with literature UV melting data or Oliveira et al. duplex melting data. g Pearson’s R between principal components of the learned variant embeddings and measured thermodynamic parameters or variant properties. h Comparison between the most correlated principal components and ∆H or Tm. i UMAP visualization of learned variant embeddings after aggregation, colored by dataset (Array Melt, literature UV melting, or Oliveira et al. duplex melting data, color scheme the same as in (d–f)). j Benchmark of all models on held-out Array Melt data, compared to the measurement error of Array Melt. k Benchmark of all models on the orthogonal Oliveira et al. dataset, plotted by the number of mismatches. Colors correspond to models as in Fig. 5j.
We trained this end-to-end model on Array Melt training set. We first explored the effect of the number of graph convolutional layers on prediction accuracy (Fig. 5b, Supplementary Fig. 5b). Performance plateaued after 4 graph convolutional layers, matching an information-sharing range of four chemical bonds. This observation is consistent with Supplementary Fig. 3g, which shows that pairwise nucleotide interactions span approximately 4 nt. When evaluating model performance over increasing amounts of training data, we observed a slower plateau compared to Fig. 4h, indicating that the GNN model can better make use of this large-scale dataset (Fig. 5c). The trained GNN had a prediction error of 0.18 kcal/mol in \(\Delta {G}_{37}\) (\(R=0.97,{R}^{2}=0.94\), Fig. 5d, e, Supplementary Fig. 5c) compared to 0.34 kcal/mol of a baseline model that simply looked up the k = 8 closest variants in the training set and gave a weighted average of the lookup values as prediction (“the closest sequence lookup model”) (Supplementary Fig. 5d, Table 1). GNN predictions were also consistently accurate across different variant classes. For example, adjusted MAEs for the Watson-Crick and mismatch classes were 0.17 and 0.18 kcal/mol, with R2 values of 0.95 and 0.94 (Fig. 5e). Interestingly, although the GNN was trained entirely on graph representations of DNA hairpins, it was able to extend to DNA duplexes (Fig. 5f). Although on duplexes the GNN performed slightly worse than our rich parameter model (Fig. 5k, Table 1), this was not surprising given that the training data consisted exclusively of hairpins. The GNN model was never explicitly taught the nature of chemical bonds or strand numbers, yet its prediction ability suggested that it grasped the “gist” of the physical system of DNA folding thermodynamics and could extrapolate beyond hairpin structures. Compared to the slow decrease of prediction error on the Array Melt data over the course of training (Supplementary Fig. 5e), for the two external DNA duplex datasets, prediction error rapidly decreased within the first 20 epochs and then plateaued (Supplementary Fig. 5f), likely because of the relatively low complexity of those datasets.
To better understand how the GNN made predictions, we analyzed the intermediate activation from the output of the Set2Set aggregation layer. The GNN projected each DNA molecular graph into a 250-dimensional embedding vector. Many of the top principal components of the embeddings correlated strongly with predicted thermodynamics parameters (\(R=0.8\) between PC1 and ΔH, \(R=-0.77\) between PC5 and Tm, and \(R=0.79\) between PC1 and ΔG37), but none with other qualities of the sequence variants including DNA length or GC content (Fig. 5g, h), underscoring the GNN’s focus on thermodynamics. When examining embeddings from different datasets (Array Melt, literature UV, and Oliveira et al.), each dataset’s distinct design structure mapped to unique areas in a two-dimensional UMAP visualization (Fig. 5i). This observation indicated that the embeddings captured more than just predicted thermodynamics like melting temperatures (Supplementary Fig. 5g), and that the learned embedding space encompassed sequence and structure details beyond mere parameter prediction.
Finally, we compared prediction errors across various models. The graph neural network (GNN) outperformed other models on the Array Melt test data, while the rich parameter model and the dna24 model showed the best performance on duplex data. Precision of the GNN was notable, approaching the uncertainty range of bootstrapped experimental measurements (mean uncertainty of 0.14 kcal/mol for \(\Delta {G}_{37}\)), demonstrating its effectiveness in capturing complex relationships in DNA sequences and structures (Fig. 5j, Table 1).
When predicting unseen duplex data from Oliveira et al., the GNN model performed well for duplexes with no mismatch (R2 = 0.78) and single mismatch (R2 = 0.56, outperforming R2 = 0.25 of the dna04 model). However, the GNN model struggled with duplexes containing two or more mismatches, yielding R2 = −0.12 for double mismatches and R2 = −1.21 for triple mismatches (Fig. 5k, Supplementary Fig. 5h, Table 1). Our two linear models, dna24 and rich parameter, still improved the prediction quality for double and triple mismatches compared to dna04 despite the low representation of these features in training data. This improvement likely resulted from the incorporation of domain knowledge in human-designed model features. We hypothesize that including more double or triple mismatch variants into the training data would address the existing bias and enhance the GNN model’s performance.
The primary goal of training and evaluating this GNN model was to assess its ability to predict thermodynamic parameters for DNA hairpin variants and its potential to generalize to non-hairpin variants such as duplexes, rather than to provide an immediate utility for all types of DNA secondary structures. Such a general deep learning model would require a large volume of extra data in other types of DNA secondary structures, such as duplexes, external loops, and longer hairpins, which are not yet available. In the future, when such data of sufficient size becomes available, we hope our GNN model could serve as a template for a family of models trained on different types of secondary structures. Eventually, we envision a general, unified deep learning model for predicting folding thermodynamics of arbitrary types of DNA secondary structures.
Discussion
In this work, we developed an accurate, scalable fluorescence-based method for measuring the melting of DNA hairpins (Array Melt) and used it to generate the largest-to-date DNA folding thermodynamics dataset. The Array Melt method, with its high throughput and precise measurement capabilities, allows for a more detailed exploration of the thermodynamics of diverse DNA structures, including those with mismatches, bulges, and hairpin loop motifs. The scale of this dataset revealed the significant role played by sequences not immediately adjacent to mismatches and within hairpin loops to the thermodynamic stability of hairpins. This observation suggests that previous nearest-neighbor models, such as the dna04 model, which parameterized only the energies of immediate nearest neighbors, may lack the complexity needed to fully capture the variance observed in these molecular variants. Our revised dna24 and rich parameter models, while rooted in the same conceptual framework of parameterizing local features for linear additive models, extended previous models by including interactions beyond immediate nearest neighbors, such as those between the two stacks flanking a mismatch. To this end, we developed three quantitative folding energy models of increasing complexity: a revised parameterization of the linear regression model compatible with NUPACK (dna24), a rich parameter linear model with particularly good performance on mismatched DNA duplexes, and a graph neural network (GNN) model that showed the best performance on hairpins. To our knowledge, this GNN model is the first deep learning model applied to DNA folding. These results support the notion that while the dna04 model is accurate in predicting Watson-Crick stack behaviors, it falls short in accurately modeling the thermodynamics of more complex DNA structures, such as hairpins and interior loops. We anticipate that future library designs will be more oriented towards machine learning, enabling a more comprehensive and less biased sampling of the DNA sequencing space.
Overall, by providing a more accurate model for DNA folding thermodynamics, we paved the way for improved design of biotechnological tools and applications. For example, our models will provide a more accurate starting point for in silico design in areas such as qPCR primer design, DNA oligo hybridization probe design, and DNA origami.
A limitation in our model comparison to the baseline arises from how NUPACK handles GT/TG mismatches. The dna04 parameter file in NUPACK currently applies dummy, instead of SantaLucia, parameters for these mismatches, overestimating their energetic contributions by ~3 kcal/mol (Fig. 4c). This discrepancy stemmed from a historical artifact dating back over 20 years, as confirmed by the NUPACK team (Supplementary Discussion; personal communication, N. Pierce, 2025). Given the notable presence of GT/TG mismatches in the Array Melt dataset, this issue may impact predictions by the dna04 baseline model. Predictions in partition function calculations and ensemble properties may be particularly affected, where the over-penalization of GT/TG mismatches could skew folding state distributions. While this limitation does not change the overall trends in our results, we have clarified its impact in relevant figures (Fig. 4c, Supplementary Fig. 4c) and highlighted its potential effects on the dna04 model performance. Future corrections to the dna04 parameter file by the NUPACK team are expected to improve model accuracy for DNA secondary structures involving GT/TG mismatches. Importantly, the provided dna24 parameter file remains fully compatible with both current and future NUPACK releases, ensuring continued usability.
Additionally, the study’s focus on two-state folding presents a notable limitation, although our comparison between the two-state and ensemble methods (Supplementary Fig. 4b) revealed greater accuracy of the former in predicting Array Melt measured thermodynamic parameters. The poorer performance of the ensemble method may be attributed to the parameter estimation methods. Furthermore, the experimental design of the datasets used to derive nearest-neighbor models may also contribute to this limitation. Thus, interpreting nearest-neighbor model predictions and dynamic programming algorithm outputs as “true” energies of single, fixed secondary structures should be done with caution. In this study, the two-state folding energies are better interpreted as an aggregated average across multiple folding states, rather than that of a single folded microstate within this ensemble. We anticipate that future analysis incorporating experimental design, data preprocessing, and parameter inference27 techniques that consider ensemble folding could result in further improvements in prediction accuracy. These approaches are relevant when it is important to predict probabilities of dynamic molecules in microstates.
Another limitation is the dataset-specific implementation of the rich parameter model in custom Python code. While nearest-neighbor implementations other than NUPACK, such as ViennaRNA28, mfold29, and RNAenn30, provide valuable tools, they do not fully address our problem space or support the detailed parameterization in our model. Developing a more general implementation of this model would require significant methodological and software redesign, which is beyond the scope of this study. Additionally, we believe that even richer parameterizations, as seen in RNA models31, could further improve performance.
This work also indicates that cloud labs can provide researchers with practical access to specialized instruments and support, addressing traditional challenges in equipment access and maintenance. These platforms also facilitate streamlined data collection and metadata tracking, improving open data and experiment reproducibility. This study represents an early instance of using commercial cloud labs in research setting32, suggesting potential broader academic use.
Finally, our measurements used a single monovalent salt concentration. Expanding to different concentrations and types of mono- or divalent cations would be straightforward. Furthermore, applying our method to RNA structures offers a promising direction for future research. Given the diverse and critical biological functions of RNA, heavily dependent on its secondary structure, high-throughput, precise thermodynamic measurements could deepen understanding of RNA biology and support the development of RNA-based therapeutics.
Methods
Library assembly and sequencing
A sequence map of the constant and variable regions of the library is available in Supplementary Data19 as LibrarySequenceMaps/array_melt_library.gb. In this map, the “filler sequence” is a 5’ partial segment of “AACAACAACAACATACTAACAACAACATAACAAATCAAAA”. The lengths of this filler sequence and the variable region add up to 40 nt, ensuring all synthesized fragments are of uniform length regardless of the length of the variable region. This map’s reverse complements of all library members were synthesized as a DNA oligo pool by Twist Biosciences. To prevent sequence dropout during synthesis, each unique sequence was included twice in the oligo pool.
This oligo pool was subsequently amplified using internal primers to enrich for full-length library variants. For this PCR reaction, a 1:8 dilution of the oligo pool was used (final concentration of 10 nM, note that the concentration of full-length sequences might be considerably lower). The PCR reaction mixture also included 200 nM of each primer (T7A1library and D-TruSeqR2, as in Supplementary Table 1), and 1x Phire Hot Start II PCR Master Mix (Thermo Fisher Scientific F125L). The thermal cycling protocol was 9 cycles of 98 °C for 10 s, 56 °C for 30 s, and 72 °C for 30 s. The PCR product was purified using the QIAquick PCR Purification Kit (Qiagen, 28104) to remove excess primers and proteins, and was eluted in 20 µL of elution buffer.
Following the initial amplification, a second PCR was performed to introduce terminal sequences compatible with Illumina sequencing. This assembly PCR used two outside primers and two adapter sequences. The reaction mixture consisted of 1 μL of the previous PCR product, 137 nM of outside primers (short_C and short_D as in Supplementary Table 1), 3.84 nM of adapter sequences (C-i7pr-bc-T7A1 and D_TruSeqR2 as in Supplementary Table 1), and 1x Phire Hot Start II PCR Master Mix. The thermal cycling protocol for this PCR was 14 cycles of 98 °C for 10 s, 56 °C for 30 s, and 72 °C for 30 s. The PCR product was again purified using the QIAquick PCR Purification Kit and quantified with Qubit dsDNA HS Assay Kit (Thermo Fisher Q32854). All PCR primers and adapter sequences were synthesized by IDT with standard desalting.
The sequencing was performed using a MiSeq Reagent Kit v3, 150 Cycles (MS-102-3001). The amplified library was diluted and pooled to a final concentration of 1.16 nM (27% of the loaded sample), combined with 2.8 nM PhiX control v3 (70%, Illumina), 0.04 nM fiducial marker (1%, sequence map available as LibrarySequenceMaps/array_melt_fiducial.gb; synthesized by IDT, standard desalting), 0.04 nM Cy3 no RNA control (1%, LibrarySequenceMaps/array_melt_cy3_control.gb; synthesized by IDT, standard desalting), and 0.04 nM single-strand fluorescence control sequences (1%, synthesized by Twist Biosciences), adding up to a total sample concentration of 4 nM. Paired-end sequencing was conducted with a mixture of Illumina’s standard read 1 primer and a custom read 1 primer (stall_R1_primer, as in Supplementary Table 1; synthesized by IDT, standard desalting) for 66 cycles and a standard read 2 primer for 109 cycles to cover the variable region in both read 1 and read 2 directions.
Imaging station setup
An imaging station was used to image the MiSeq flow cell at varying temperatures. This station was built from a combination of custom-designed parts and parts from a disassembled Illumina Genome Analyzer IIx, as previously described33,34. The station had two channels: the “red” channel, which uses a 660 nm laser paired with a 664 nm long pass filter (Semrock BLP01-664R-25), and the “green” channel, which uses a 532 nm laser with a 590 nm band pass filter (Semrock FF01-590/104-25). Images were taken with an exposure time of 600 ms and a laser input power of 150 mW. For each temperature, focusing was achieved by manually adjusting the z-position and re-imaging the four corners of the flow cell; the resulting z-positions were then fitted to a plane to ensure uniform focus across the flow cell.
Following sequencing, the flow cell was washed with Cleavage Buffer (100 mM Tris-HCl, 125 mM NaCl, 0.05% Tween20, 100 mM TCEP, pH 7.4) at 60 °C for 5 min to remove residual fluorescence from the reversible terminators used in the sequencing reaction. The compositions of all buffers used are provided in Supplementary Table 4, and the vendors and catalog numbers of key reagents in Supplementary Table 5. Any DNA strands not covalently attached to the chip surface were removed by washing with 100% formamide at 55 °C. The remaining single-stranded DNA fragments were then incubated with 500 nM of the oligo Biotin_D_Read2 and red_oligo (Supplementary Table 1) in Hybridization Buffer (5x SSC buffer, 5 mM EDTA, 0.05% Tween20) for 15 min at 60 °C, followed by a temperature reduction to 40 °C for another 10 min. The red_oligo, conjugated to Alexa647, binds to a sparse subset of “fiducial” sequences on the chip, providing fluorescence signal for z-plane focusing and for registering macro x and y offsets corresponding to the sequenced tiles. All chemically modified oligos were synthesized by IDT with HPLC purification.
Measuring melt curves on the chip
The Cy3-labeled fluor_oligo was annealed in 1xSSC + MgCl2 (1x SSC, 7 mM MgCl2, 0.01% v/v Tween-20, 0.5 μM labeled oligo) at 37 °C for 12 min, and then washed with 1x SSC buffer. The chip was imaged in the “green” channel to acquire “post Cy3” fluorescence signals, which confirm hybridization and provide data for per-cluster intensity normalization during data processing. Next, the Black Hole Quencher (BHQ)-labeled oligo quench_oligo (Supplementary Table 1) was annealed and washed using the same protocol. Both fluor_oligo and quench_oligo were synthesized by IDT with HPLC purification. Again, the chip was imaged to confirm signal reduction from successful quenching. Subsequently, the chip was rinsed with Melt Buffer (50 mM Na-HEPES pH 8.0, 25 mM NaCl). HEPES buffer was chosen for its low sensitivity to temperature changes (−0.014 ΔpKa/°C), which can help maintain consistent pH during the experiment35.
To quantify the melt curves, the image station temperature was initially lowered to 20 °C. For each temperature point, the system was equilibrated to the new temperature (approximately 5 min) before refocusing and imaging. The temperature was increased in 2.5 °C increments up to a maximum of 60 °C. The detailed protocol is available in Supplementary Methods.
Processing sequencing data
Sequencing data from the Illumina MiSeq platform was processed to extract the tile and coordinates for each sequenced cluster. Forward and reverse paired-end reads generated from the sequencing process were aligned using FLASH36 (v1.2.11) with default settings to merge overlapping reads. The consensus merged sequences from FLASH were then aligned to the reverse complement sequences of fluor_oligo and quench_oligo using a Needleman-Wunsch alignment implemented in the package nwalign3 (v0.1.2).
For consensus sequences that successfully aligned to both oligos with a p-value <10−3), evaluated as described in ref.37, the variable region was extracted as the segment between the two aligned flanking regions. This variable region was then aligned to sequence variants in the reference library. A reference sequence was assigned to each cluster based on the best-scoring alignment with a p-value <10−6). These assigned clusters were subsequently used for fluorescence quantification in downstream data analysis.
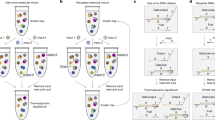
Overview of Array Melt curve processing
In the following sections, we will describe the process of Array Melt curve fitting. As noted in the main text, we refer to a single group of about 1000 DNA molecules on the chip as a “cluster”, and a distinct sequence in the library as a “variant”. Each variant has multiple corresponding clusters measured on the chip.
-
1.
Clusters in the fluorescent images were registered to clusters identified through sequencing to obtain sequence identity information, and raw fluorescence was quantified from the images.
-
2.
Raw fluorescence signals were normalized to the initial signal to account for differences in cluster sizes and sequence-specific effects on fluorescence.
-
3.
Signals were further normalized using controls for folded and unfolded constructs to correct for effects such as temperature and photobleaching during the experiment.
-
4.
Normalized signals from single clusters were fitted to melt curves with minimal constraints.
-
5.
A subset of variants with high-quality single-cluster fits that reached maximum fluorescence (fmax) was selected to estimate probability distributions of fmax. Similarly, another subset was selected to estimate the probability distributions of fmin.
-
6.
At the variant level, we bootstrapped single clusters to refine the fit. For variants that reached fmax or fmin, these values were directly fitted from the data; for those that did not reach fmax or fmin, we drew samples from the distributions fitted in the previous step during bootstrapping.
-
7.
Fitted results from multiple replicates were combined and filtered.
-
8.
An orthogonal fitting method, termed the ΔG line fitting approach, was applied to help filter out variants that did not show two-state behavior.
Fluorescence data processing and image fitting
Fluorescent melt curve images were aligned with sequencing data from the Illumina MiSeq platform. Initially, sequencing data was analyzed to obtain the tile and coordinates of each sequenced cluster. These coordinates were then used to create synthetic cluster images, which were iteratively registered with the fluorescent images to achieve sub-pixel resolution mapping34. After identifying the clusters in the images, each cluster’s fluorescence was quantified by fitting it to a two-dimensional normal distribution30.
Distance sensitivity curve
The repeat controls used to plot the distance sensitivity curve in Fig. 1d are repeats of varied lengths of the following sequences: poly-A, T, AT, AAC, AAG, AAT, AC, ACC, AG, AGG, TC, TG, TTA, TTC, and TTG.
The theoretical curve fitted to the distance-to-fluorescence curve in Fig. 1d is \(f\left(x\right)=\frac{1}{1+({a\cdot }{(x+b)}^{-3})}\), where \(x\) represents the length in nucleotides (nt), \(f\left(x\right)\) is the normalized fluorescence at length \(x\) (dimensionless, normalized to maximum fluorescence), \(b\) is a fitted parameter in nt accounting for systemic distance offset, and \(a\) is another fitted parameter:
where \({K}_{D,{BHQ}}\) is the dissociation constant for the fluorophore-quencher pair, \({N}_{A}\) is Avogadro’s number, and \(k=0.64\times 1{0}^{-9}{m}\cdot n{t}^{-1}\) is a distance conversion factor from nt to physical distance for single-stranded DNA. The fitted parameter values are \(a=71.19\), \(b=0.08\), and \({K}_{D,{BHQ}}=145.6{mol}\cdot {L}^{-1}\). Using the relation \({\Delta G}_{{BHQ}}={RT}\cdot {{\mathrm{ln}}}{K}_{D,{BHQ}}\), we have \(\Delta {G}_{37,{BHQ}}=3{kcal}/{mol}\). This interaction between the fluorophore and the quencher might introduce a systematic offset in the data. We corrected for this and other potential systematic errors using UV melting data (see UV melting section below).
Fluorescence normalization
Intensity normalization
To minimize inter-cluster size variation and sequence-specific effects in fluorescence measurements, we normalized Cy3 fluorescence during melt curve measurements by the total amount of single-stranded DNA in each cluster. This was determined by the Cy3 fluorescence signal immediately after the hybridization of the fluor_oligo but before the hybridization of the quench_oligo. This Cy3 signal was clipped to the 1st and 99th percentile of its total distribution and used as a normalizing factor for all subsequent fluorescent signals from each cluster.
Temperature dependence and photobleaching normalization
Throughout the experiment, fluorescent signals were affected by temperature and photobleaching. To correct for these effects, we further normalized the size-normalized signals of clusters using the median signals of control variants designed to maintain constant distances between the Cy3 fluorophore and the BHQ quencher. These controls include:
-
1.
“Super stable stem” variants with long GC-rich stems that are expected to retain their hairpin secondary structure within the experimental temperature range, showing minimal fluorescence.
-
2.
Within the experimental temperature range, showing minimal fluorescence.
-
3.
“Long repeat” controls that are repeat sequences at least 39 nt long. They are expected to remain linear, not to form hairpins, and to show maximum fluorescence.
For both the “super stable stem” and the “long repeat” control variant groups, we applied the following filtering process: after calculating the median signals of each variant across all of its corresponding clusters, we found the 5th and 95th percentile of these medians within each of the two control groups for each temperature point. Library variants with more than one data point outside this 5th to 95th percentile range were considered outliers and excluded from further analysis.
Afterwards, the median values at each temperature point from both control groups were used to normalize the entire dataset. In total, 5 out of 5 super stable stem control and 40 out of 92 long repeat control variants were used for normalization (Supplementary Fig. 1b).
Direct melt curve fitting
The melt curve fitting pipeline was adapted from previous work38, including the following steps: 1) Directly fit normalized signals of single clusters; 2) Estimate the probability distributions of maximum and minimum fluorescence (\({f}_{\max }\) and \({f}_{\min }\)) from single cluster fit results; 3) Refine the fit at the variant level using bootstrapping. This fitting process targets one of the main challenges in melting curve analysis: \({f}_{\max }\) and \({f}_{\min }\) are not constant and not always reached by all the variants. The full details are described below, and the fitting process is available as a snakemake39 pipeline at https://github.com/keyuxi/array_analysis.
Single cluster fitting
Equation (2) was directly fitted to the normalized signals of single clusters with minimal constraints, where \(T\) and \({T}_{m}\) are temperatures in Kelvin, \({k}_{B}=0.0019872\) is the Boltzmann constant, and \({f}_{\max }\) and \({f}_{\min }\) are the maximum and minimum normalized fluorescence. Initial parameter values were set as follows: Tm was set to the temperature where the normalized signal for a given variant was closest to 0.5, ΔH to −40, \({f}_{\max }\) to 1, and \({f}_{\min }\) to 0. The python package lmfit (v1.0.3)40 was used for least-squares fitting.
Estimating the distributions of \({f}_{\max }\) and \({f}_{\min }\)
To evaluate if a single cluster fit was successful, we applied the following filtering criteria: \({R}^{2}\) of the fit must be at least 0.5, the fitted \({f}_{\min }\) must be between −1 and 2, the estimated standard error of the best-fitted parameter value \({\sigma }_{{f}_{\min}}\) must be less than \({f}_{\min }+1\), the fitted \({f}_{\max }\) must be between 0 and 3, \({\sigma }_{{f}_{\max }}\) must be less than \({f}_{\max }\), \({\sigma }_{Tm}\) must be less than 10 K, and \({\sigma }_{\Delta H}\) must be less than 100 kcal/mol.
Single clusters were then aggregated at the variant level. For each variant, a “success p-value” was calculated assuming a null hypothesis of a 25% rate of success for single cluster fits, as in previous work38. Then, we calculated the medians of the fitted parameters for each variant. A variants was used to estimate the distribution of \({f}_{\max }\) if: 1) its corresponding single melt curves fitted well, defined as having a “success p-value” less than 0.01; 2) it was unfolded at high temperatures, defined as having a fitted \({f}_{\max }\) value of at least 0.5, and at least 97.5% of the molecules are unfolded at 60 °C (\({p}_{{unfolded}}^{60\,^\circ C}\ge 0.975\), this ensures that \({f}_{\max }\) is reached for this variant). Similarly, for estimating the distribution of \({f}_{\min }\), we filtered for variants with a “success p-value” less than 0.01, \({f}_{\min }\) less than 0.05, and \({p}_{{unfolded}}^{20^\circ C}\) at most 0.025.
We then used the selected variants to fit probability distributions for \({f}_{\max }\) and \({f}_{\min }\). For example, \({f}_{\max }\) was modeled as a normal distribution \(N ({{{\upmu }}}_{{f}{{\max}}},{\sigma }_{{f}{{{\rm{max }}}},n})\), where \({\sigma }_{{f}\max,n}\) is a function of the number of clusters per variant, \(n\). This relationship was modeled and fitted as \({\sigma }_{{f}{{{\rm{max}}}}},{n}=\frac{a}{\sqrt{n}}+b\) (Supplementary Fig. 1c). Similarly, \({f}_{\min }\) was modeled as \(N({{{{\rm{\mu }}}}}_{{f}{{\min}}},{{{{\rm{\sigma }}}}}_{{f}{{\min}},n})\).
Fit refinement
A second round of fitting, also using, was performed at the variant level to refine the fit results using parameters estimated from the previous steps. For each variant, \({f}_{\max }\) and \({f}_{\min }\) were either directly fitted from data or drawn from the normal distributions fitted in the previous step. This allows \({f}_{\max }\) and \({f}_{\min }\) some flexibility while improving fitting quality for variants where it is challenging to directly infer these values from melt curves. We independently determined whether to enforce \({f}_{\max }\) or \({f}_{\min }\) distributions to correct for variants that did not fully unfold or fold during the experiment. Specifically, we calculated the lower bound for \({f}_{\max }\) as \({\mu }_{{f}_{\max }}-10\cdot {\sigma }_{{f}_{\max }}\), where \({\mu }_{{f}_{\max }}\) and \({\sigma }_{{f}_{\max }}\) are the mean and standard deviation calculated from the previous step, and the upper bound for \({f}_{\min }\) as \({\mu }_{{f}_{\min }}+{\sigma }_{{f}_{\min }}\). We set a larger margin for \({f}_{\max }\) to enforce it less strictly since \({f}_{\max }\) is more variable than \({f}_{\min }\). If the median signal at the highest temperature was below the lower bound for \({f}_{\max }\), we enforced \({f}_{\max }\); likewise, if the median signal at the lowest temperature was above the upper bound for \({f}_{\min }\), we enforced \({f}_{\min }\). Otherwise, \({f}_{\max }\) and \({f}_{\min }\) were directly fitted. For example, in the first of the four replicates, 21,147 out of the 36,291 remaining variants at the fit refinement step (58.3%) had \({f}_{\max }\) enforced, and 26,799 out of 36,291 variants (73.8%) had \({f}_{\min }\) enforced.
Next, we fitted each variant by bootstrapping the clusters within that variant. At each resampling step, if enforcement was required, we drew an \({f}_{\max }\) or \({f}_{\min }\) value from the global normal distribution; otherwise, we allowed \({f}_{\max }\) or \({f}_{\min }\) to vary freely. We then fitted to the median signal of the drawn clusters, using the single cluster fit results as the initial parameters. Subsequently, we calculated \(\Delta {G}_{37}\) and ΔS for each resampling step:
where \(T\) is the temperature in Kelvin. We repeated this resampling process 100 times and obtained a 95% confidence interval and standard deviation for each parameter: ΔH, Tm, ΔG37 and ΔS.
ΔG line fitting and two-state heuristics
In our analysis, we assume two-state melting behaviors, meaning that the hairpins are either in folded or unfolded states. To ensure that the variants used for analysis exhibited such behavior, we used an alternative line fitting approach to derive heuristics for two-state behavior. First, the signal \(f\left[T\right]\), previously normalized to the initial fluorescence after hybridization, was re-normalized using the fitted \({f}_{\max }\) and \({f}_{\min }\) values. This re-normalized signal, \({f}_{{norm}}\left[T\right]=\left(f\left[T\right]-{f}_{\min }\right)/\left({f}_{\max }-{f}_{\min }\right)\), was clipped between 0 and 1 with \(\epsilon=0.01\) to prevent issues in subsequent calculations. Then, we transformed \({f}_{{norm}}\left[T\right]\) to \(\Delta G\left[T\right]\) using:
Error propagation was managed using the uncertainties Python library41 to calculate uncertainties in \(\Delta G\left[T\right]\). Finally, we fitted a linear model to the transformed data with free parameters ΔH and Tm:
If a variant exhibited two-state behavior during the experiment, \(\Delta G\left[T\right]\) should fit well to this linear model, and the fitted parameters should be consistent with those obtained from the direct melt curve fitting approach (details described in the next section); otherwise, deviations would arise. We first used RANSAC42 to identify outliers, then fitted ordinary least squares linear regression to the inliers to determine the final parameters. These fitted parameters were compared to those derived from direct melt curve fitting. Only variants showing agreement between the two methods were considered “two-state”.
This two-step heuristic was inspired by UV melting two-state heuristics in the literature. Traditionally, agreement between parameters from direct melt curve fitting and from van’t Hoff equation analysis (a line fitting approach, not applicable to Array Melt data because it requires multiple strand concentrations) was required to apply the two-state assumption. As SantaLucia stated: “For a given oligonucleotide, agreement of parameters derived by the two methods is a necessary, but not sufficient, criterion to establish the validity of the two-state approximation9.” Thus, a variant meeting our ΔG line fitting two-state heuristics was necessary but not sufficient to confirm its validity of the two-state approximation, and was consistent with historical two-state criterion in the literature.
Filtering and combining data replicates
Each variant, measured and fitted to curves in an individual replicate experiment, was first filtered using the following criteria: measured in at least 5 clusters, a bootstrapped error in ΔG37 of less than 2 kcal/mol, an error in Tm of less than 25 °C, an error in ΔH of less than 25 kcal/mol, and a curve fitting RMSE of less than 0.5. This step keeps 86.43%, 78.88%, 64.31%, and 71.46% variants in the four replicate experiments, respectively. For each replicate, variants were compared to the ΔG line fitting results for two-state behavior assessment. To be considered as passing the two-state criteria, a variant has to have its fitted ΔH values from both curve and line fitting methods agree within 50%, Tm within 10 °C, reduced \({{{{\rm{\chi }}}}}^{2}\) of the line fit below 2, and the number of inlier data points in line fitting greater than 8.
Replicates were then combined across four replicate experiments, including three melting and one colling curve measurements from two different flow cells (Supplementary Table 3). For each parameter (ΔH, Tm, \(\Delta {G}_{37}\), or ΔS), the combined error was \(\sigma=\sqrt{{({\sum }_{i}{\sigma }_{i}^{-2})}^{-1}}\) and the combined parameter was \(X={\sigma }^{2}\cdot {\sum}_{i}{X}_{i}/{\sigma }_{i}^{2}\), where \({X}_{i}\) and \({\sigma }_{i}\) were respectively the measurements and bootstrapped standard errors in each replicate. Missing values were ignored in the summation. This step yields 31,000 variants.
We then applied a filter for two-state behavior that kept 30,091 variants from the previous step, where a variant had to pass the two-state criteria in at least one replicate. Extra control variants were removed to leave 29,864 variants. Lastly, we applied another filter to the combined variants, keeping 27,732 of variants with Tm between 0 and 60 °C. This removed variants not showing enough melting behavior.
UV melting
Designing sequences for UV melting and HPLC quality control
We designed two groups of hairpins for UV melting measurements. The first group consisted of hairpin sequences sampled from the variable regions the Array Melt library for validations in Fig. 2. The second group comprised of 16-mers not in the Array Melt library, designed to fold into hairpins with the target secondary structure ((((((….)))))). These sequences were used in Supplementary Fig. 4d. For this second group, we used NUPACK4 to filter out sequences with misfolded secondary structures. DNA oligo samples were synthesized by Integrated DNA Technologies (IDT) and analyzed using Ion Exchange High Pressure Liquid Chromatography (IE-HPLC) remotely on Emerald Cloud Lab (ECL) with DNAPac PA200, 9 × 250 mm Semi-Prep column (Thermo Fisher Scientific 063421). Fractions were detected by a UV-Vis detector at 260 nm. Only oligos that showed a single peak were selected for further data analysis.
Running cloud lab UV melting experiments
UV melting experiments were run remotely on ECL. Links to the data objects are available in Supplementary Table 2. DNA oligos ordered from IDT were shipped to ECL and resuspended in high-concentration stock solutions. Before each experiment, the DNA stock solutions were snap frozen by heating to 90 °C for 5 min, then cooling to 4 °C for 30 min, before diluted to working concentrations. Initially, the same oligo sequence was tested with a wide range of DNA strand concentrations from 1 to 12 μM to confirm intramolecular hairpin formation rather than intermolecular hybrids (Supplementary Fig. 2a). For method validation in Fig. 2 and model comparison in Supplementary Fig. 4d, hairpin data were all collected at DNA strand concentrations of 6 μM or 9 μM. These concentrations ensured a high signal-to-noise ratio while maintaining intramolecular folding behavior.
The buffer used for Fig. 2 and Supplementary Fig. 4d was the same as in the Array Melt (50 mM Na-HEPES pH 8.0, 25 mM NaCl). Some samples were also measured at higher salt concentrations (50 mM Na-HEPES pH 8.0, 50 mM NaCl; 50 mM Na-HEPES pH 8.0, 1 M NaCl). Measurements were performed on a Cary 3500 UV/Vis spectrophotometer using micro scale black walled UV quartz cuvettes with a 10 mm pathlength. Each sample underwent 2 to 3 melt-cooling cycles, with a temperature ramp rate of 0.8 °C and 15 min equilibrium time between each melt and cool sweep. Temperatures were measured with an immersion probe. Data points were acquired every 1 °C at 260 nm from 5 °C to either 85 or 95 °C.
Analyzing UV melting curves
The UV melting curves were fitted to a modified version of Eq. (2) with an extra term to account for baseline slope:
where f[T] is the absorbance, \(a\) is a free parameter for the slope, and the rest the same as in. We introduced this slope parameter because the raw, unnormalized UV melting data were directly fitted to, unlike the Array Melt data, which were pre-normalized for baseline drifts prior to fitted to.
For each individual melting or cooling curve, the initial value of \({T}_{m}\) was determined as the temperature where the derivative \(\frac{{df}\left[T\right]}{{dT}}\) is maximized, denoted as \({T}_{m}^{{diff}}={argma}{x}_{T}\frac{{df}\left[T\right]}{{dT}}\). In the first round of fitting, we fixed the value of \({T}_{m}\) to \({T}_{m}^{{diff}}\) while allowing parameters \(a\), \({f}_{\max }\), \({f}_{\min }\), and \(\Delta H\) to float. In the second round, we used the best-fitted values from the previous round as the initial values and allowed \({T}_{m}\) to float. This two-step fitting process was employed to prevent the optimization from getting stuck in unlikely local minima.
We selected individual curves based on the following criteria: fitting RMSE less than 0.002, uncertainties of fitted parameters \({\sigma }_{\Delta H} < 10\,{kcal}/{mol}\), \({\sigma }_{{T}_{m}} < 5^\circ C\), \(-100 < \Delta H\le 0\,{kcal}/{mol}\), and \(0 < {T}_{m} < 100\,^\circ C\). After this step, 297 out of 387 individual curves remained. Next, for each sample, we aggregated the fitted \(\Delta H\) and \({T}_{m}\) values from individual melting and cooling curve, using \(X={\sigma }^{2}\cdot {\sum }_{i}{X}_{i}/{\sigma }_{i}^{2}\) and \(\sigma=\sqrt{{\left({\sum}_{i}{\sigma }_{i}^{-2}\right)}^{-1}}\), where \(X\) is the aggregated \(\Delta H\) and \({T}_{m}\) values and \({X}_{i}\) are the fitted values from an individual curve. We then calculated \(\Delta {G}_{37}\) and \(\Delta S\) from the aggregated \(\Delta H\) and \({T}_{m}\) using Eqs. (3) and (4), and used the uncertainty propagation equations \({\sigma }_{\Delta S}=-\Delta S\cdot \sqrt{{\left(\frac{{\sigma }_{\Delta H}}{\Delta H}\right)}^{2}+{\left(\frac{{\sigma }_{{T}_{m}}}{{T}_{m}}\right)}^{2}}\) and \({\sigma }_{\Delta {G}_{T}}=\sqrt{{\sigma }_{\Delta H}^{2}+{\left(T\cdot {\sigma }_{\Delta S}\right)}^{2}}\) to calculate the measurement uncertainties. We then filtered out samples with aggregated \({\sigma }_{\Delta H} > 15\, {kcal}/{mol}\) or \({\sigma }_{{T}_{m}} > 10^\circ C\). After this step, 116 out of 120 samples remained. These 116 samples correspond to 85 different sequence variants, with 87 measured in 25 mM NaCl buffer (the buffer used for Array Melt), 7 in 50 mM NaCl, and 22 in 1 M NaCl.
Variance analysis
Mismatch ΔΔG37 calculation
For each mismatch, we identified its parent sequence variant, which is a hairpin with a Watson-Crick paired stem that can be converted into the mismatch variant by mutating a single nucleotide. The mismatches were categorized by the identities of those nucleotide substitutions. We then calculated the \(\Delta {\Delta G}_{37}\) of each mismatch by subtracting the \(\Delta {G}_{37}\) of the parent variant from that of the mismatch variant. C > G single mismatches were excluded from the variance calculations due to an insufficient number of data points (only 2 variants from the External class).
Expected biological variance and model-explained or -introduced variance
For a dataset of \(n\) sequence variants with \(n\) corresponding ΔΔG values \({y}_{1},\ldots,{y}_{n}\), measurement uncertainty values \({\sigma }_{1},\ldots,{\sigma }_{n}\), and the dna04 model prediction values \({\hat{y}}_{1},\ldots,{\hat{y}}_{n}\), we have:
-
Total variance \({{\sigma }^{2}}_{{tot}}={\sum }_{i=1}^{n}{({y}_{i}-\bar{y})}^{2}/n\)
-
Technical variance \({{\sigma }^{2}}_{{tech}}={\sum }_{i=1}^{n}{{\sigma }_{i}}^{2}/n\)
-
Expected biological variance \({{\sigma }^{2}}_{{bio}}={{\sigma }^{2}}_{{tot}}-{{\sigma }^{2}}_{{tech}}\)
-
Model variance (variance of prediction values) \({{\sigma }^{2}}_{{model}}={\sum }_{i=1}^{n}{(\hat{{y}_{i}}-\bar{\hat{y}})}^{2}/n\)
-
Residual variance \({{\sigma }^{2}}_{{res}}={\sum }_{i=1}^{n}{({y}_{i}-\hat{{y}_{i}})}^{2}/n\).
From these, we calculated the variance explained \({{\sigma }^{2}}_{{explained}}={{\sigma }^{2}}_{{tot}}-{{\sigma }^{2}}_{{res}}\). If \({{\sigma }^{2}}_{{res}} > {{\sigma }^{2}}_{{tot}}\), the model introduced additional variance, calculated as \({{{{\rm{\sigma }}}}}_{{{{\rm{added}}}}}^{2}=-{{{{\rm{\sigma }}}}}_{{{{\rm{explained}}}}}^{2}={{{{\rm{\sigma }}}}}_{{{{\rm{res}}}}}^{2}-{{{{\rm{\sigma }}}}}_{{{{\rm{tot}}}}}^{2}\).
Nucleotide importance score
We calculated nucleotide importance score (NIS) using \(\Delta \Delta {G}_{37}\) for mismatches, and \(\Delta {G}_{37}\) for triloop, tetraloop, and bulges. For a dataset of \(n\) sequence variants with \(n\) corresponding energy values \({y}_{1},\ldots,{y}_{n}\) (either \(\Delta {G}_{37}\) or \(\Delta {\Delta G}_{37}\)), we grouped the \(n\) data points into four groups based on the identity of the nucleotide of interest. For each of the four groups, we calculated the mean values, \(\bar{{y}_{A}}\), \(\bar{{y}_{T}}\), \(\bar{{y}_{C}}\), and \(\bar{{y}_{G}}\). The grand mean, \(\bar{{y}_{{tot}}}\), is the mean of all \({y}_{1},\ldots,{y}_{n}\). The NIS is simply the between group sum of squares (SSB):
Pairwise interaction score
For a pair of nucleotides of interest, \(M\) and \(N\), we compared two linear models that predicted the energy value \(Y\), model A with an independence assumption and model B without. Model A was in the form of \(M+N \sim Y\), and model B \({MN} \sim Y\). The pairwise interaction score (PIS) was calculated as the difference of \({R}^{2}\) between the two linear models:
Salt correction
All salt corrections used the equation from ref.43:
where \({T}_{m}\left(1\right)\) and \({T}_{m}\left(2\right)\) are the melting temperatures before and after salt correction in Kelvin, \({\left[N{a}^{+}\right]}_{1}\) and \({\left[N{a}^{+}\right]}_{2}\) the corresponding sodium concentrations, and \(f\left({GC}\right)\) the fraction of GC in the sequence. This salt correction approach was applied to all places where NUPACK predictions were used at a salt concentration lower than 1 M, or when correcting Array Melt data to 1 M for parameter estimation. This includes the NUPACK-compatible NN model and all comparisons to NUPACK (Figs. 4d–g, 5j, k). The Na+ concentration of the Array Melt buffer (25 mM NaCl, 50 mM HEPES pH8.0) should be approximately 75 mM. We initially set sodium concentration to 88 mM when converting Array Melt data to 1 M sodium, but the difference is minimal (about 0.3 °C in Tm at 1 M sodium, not considering the errors from the salt correction equation itself) compared to 1.28 °C, the mean of Array Melt bootstrapped measurement uncertainties (Table 1). We used this salt correction equation instead of the built-in one of NUPACK because the latter doesn’t interpolate well to low sodium concentrations (equation S6 in ref. 18): \(\Delta {G}^{{salt}}=-0.114\log \left(\left[N{a}^{+}\right]\right)\frac{T}{{T}_{37}}\).
Calculating melting temperature with NUPACK
For DNA duplexes, melting temperatures were calculated from the fraction of molecules unpaired. We first calculated structure free energies of the paired state at temperature T, \(\Delta {G}_{{paired},T}\), then got the equilibrium constant
where \(\beta=1/{RT}\) is the thermodynamic beta, \(\left[{dsDNA}\right]\) is the concentration of double-strand DNA, and [ssDNA] that of the single-strand DNA. At the melting temperature, \(\left[{{{\rm{dsDNA}}}}\right]=\left[{{{\rm{ssDNA}}}}\right]=\left[{{{\rm{DNA}}}}\right]/2\) for distinguishable strands and \(\left[{{{\rm{dsDNA}}}}\right]=\left[{{{\rm{DNA}}}}\right]/4\) for indistinguishable strands, where \(\left[{{{\rm{DNA}}}}\right]\) is the total concentration of DNA strand species. This gives \(K=\frac{2}{\left[{{{\rm{DNA}}}}\right]}\) or \(K=\frac{1}{\left[{{{\rm{DNA}}}}\right]}\) for distinguishable and indistinguishable strands respectively. Solving this equation with bisection gave us the melting temperature.
For DNA hairpins, melting temperature is independent of DNA strand concentration. The Boltzmann factor becomes:
\({T}_{m}\) was the temperature when \(d{G}_{{folded}}=0\). Therefore, we calculated structure free energies \(\Delta {G}_{1}\) and \(\Delta {G}_{2}\) at a lower temperature \({T}_{1}\) and a higher one \({T}_{2}\), then calculated \(\Delta S=-\left(\Delta {G}_{2}-\Delta {G}_{1}\right)/\left({T}_{2}-{T}_{1}\right),\Delta H=\Delta {G}_{1}+{T}_{1}\cdot \Delta S\), and \({T}_{m}=\Delta H/\Delta S\).
Data splitting for machine learning models
After filtering with the two-state criteria as described in “ΔG line fitting and two-state heuristics”, we split the clean Array Melt data of 27,732 variants into train, validation, and test sets, stratified by variant type. We first split 5% of the data as the test set (1438 variants), then split 5% of the remaining data as the validation set (1315 variants), and 95% of the rest as the training set (24,979 variants). We split the external 384-duplex dataset and Oliveira et al. dataset to a 1:1 split of validation and test data (192 each). Cloud lab UV melting data used for correction between Array Melt and bulk solution were not used for model testing to prevent circularity.
Linear models
Weighted linear regression model
We used a singular value decomposition (SVD) solver to solve the linear regression solutions, taking measurement error of each sequence into account. For a training dataset of \(n\) sequence variants with \(n\) corresponding \(\Delta H\) or \({\Delta G}_{37}\) values \({y}_{1},\ldots,{y}_{n}\), and measurement uncertainty values \({\vec{{{{\boldsymbol{\sigma }}}}}}=[\sigma_{1},\ldots,{\sigma }_{n}]\), we had a \(n\) by \({n}_{{feature}}\) feature count matrix \(X\) derived from the sequences. We then did SVD on matrix \(A=X/\vec{{{{\boldsymbol{\sigma }}}}}\) and got \(U,S,V\). Any diagonal value in \(S\) that was smaller than a relative numeric threshold \(1{0}^{-15}\) was set to zero. Then, the estimated parameter values for the features were \({V}^{T}\cdot {S}_{{inv}}\cdot {U}^{T}\cdot \vec{{{{\bf{b}}}}}\), and the estimated errors for the features were \(\sqrt{\sum {V}^{T}\cdot {{S}^{2}}_{{inv}}}\), where \(\vec{{{{\bf{b}}}}}=\vec{{{{\bf{y}}}}}/\vec{{{{\boldsymbol{\sigma }}}}}\), \({S}_{{inv}}\) was \(S\) but taking \(\frac{1}{s}\) for all diagonal elements, and \({{S}^{2}}_{{inv}}\) was \(S\) but taking \(\frac{1}{{s}^{2}}\) for all elements.
The dna24 model
We trained the improved NUPACK-compatible linear regression model on the Array Melt training dataset, containing 24,971 of the 27,732 variants that passed quality control and two-state criteria, and tuned the model on a validation set of 1315 variants. We first adjusted salt concentration to 1 M Na+ using Eq. (10) to be consistent with NUPACK and other software and then corrected systematic errors by linearly shifting Tm and ΔG, such that the medians of each construct type matched those of NUPACK predictions. The underlying assumption was that NUPACK predictions have minimal bias but could be improved by explaining more variance. Improved parameters for NUPACK were fitted with weighted linear regression, as described above. We allowed an extra intercept parameter to adjust mean absolute error. From sequence-structure pairs in the training set, we extracted features of stacks and loops. Stacks include nearest-neighbor Watson-Crick pairs and terminal stack penalty, as in NUPACK; loops include the following:
-
1)
Hairpin loops, defined as a subsequence \({\phi }_{i:j}\) where \({i\cdot j}\) is the closing base pair. We extracted “hairpin_mismatch” features as in NUPACK, \({\varDelta G}_{{\phi }_{j-1},{\phi }_{j},{\phi }_{i},{\phi }_{i+1}}^{{hairpinmm}}\), and introduced an intermediate “hairpin_loop_mid” feature, \({\varDelta G}_{{\phi }_{\left[i+2:j-2\right]}}^{{hairpinmid}}\) to account for sequence variations in the middle of hairpin loops for triloops and tetraloops. The final energy of hairpin is \({\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpin}}={\varDelta G}_{n}^{{hairpinsize}}+{\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpinseq}}\) as in ref. 18, where
$${\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpinseq}}={\varDelta G}_{{\phi }_{j-1},{\phi }_{j},{\phi }_{i},{\phi }_{i+1}}^{{hairpinmm}}+{\varDelta G}_{{\phi }_{\left[i+2:j-2\right]}}^{{hairpinmid}}\left(+{\varDelta G}_{{\phi }_{j},{\phi }_{i}}^{{terminalbp}}{{{\rm{if\; triloop}}}}\right)$$(14) -
2)
Interior loops, defined from 5’ to 3’ as two subsequences \({\phi }_{\left[i:d\right]}\) and \({\phi }_{\left[e:j\right]}\). Any other interior loop types were defined as in ref. 18, apart from the \({\varDelta G}_{{\phi }_{j-1},{\phi }_{j},{\phi }_{i},{\phi }_{i+1}}^{{interiormm}}\), which were fitted separately instead of shared with terminal mismatches as in NUPACK. For single mismatches, we also introduced an intermediate parameter for the interaction between the two base pairs surrounding 1 × 1 mismatches \({\varDelta G}_{{\phi }_{i},{\phi }_{j},{\phi }_{i+2},{\phi }_{j-2}}^{{interiormmstacks}}\). For longer mismatches, we did not include this interiormmstacks parameter.
This resulted in 178 features for \(\varDelta H\) and \(\varDelta G\) respectively. Parameters for hairpin and interior loop sizes were taken from NUPACK and clamped during fitting. Lookup tables of \({\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpinseq}}\) and \({\varDelta G}_{{\phi }_{\left[i:d\right]},{\phi }_{\left[e:j\right]}}^{{interiorseq}}\) for “interior_1_1”, “interior_1_2”, and “interior_2_2” were calculated from the fitted parameters. All updated parameters were then written to a NUPACK-compatible json file dna24.json.
Afterwards, we obtained parameters for \({\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpinseq}}\) for tetraloops and triloops directly from the ‘2nd flanking’ construct types of either tetraloop or triloop parts of the library (Fig. 1c). These variants covered all possible sequences for hairpin loops of length 3 or 4, as well as the nearest-neighbor and next-nearest-neighbor closing pair sequences. We first regressed out the next-nearest-neighbor closing pair of the loops from measured ΔH and ΔG values, then aggregated the mean values for variants with identical hairpin loop and nearest-neighbor closing pair sequences. These aggregated values have the correct ΔΔH and ΔΔG relationships between each other but need a constant to be added to get the ΔH and ΔG values needed for the dna24 model. To solve this problem, we fitted two correction values \(\varDelta \Delta H\) and \(\Delta \varDelta G\) in kcal/mol. We searched for values that minimize the equation
where \({w}_{{Tm}}=4,{w}_{\Delta H}=0.3,{w}_{\Delta G}=3.4\) and \(\alpha=0.0001\) are weights determined by validation on 100 sampled variants in the training set. Calculating this equation is computationally expensive, as it involves directly calling NUPACK in each iteration. We minimized it on 25 randomly sampled variants from the training set, repeating the process 4 times on different subsets using the BFGS algorithm implemented in the scipy.optimize.minimize function, with parameters tol=1e-1 and maxiter=20. The resulting \(\varDelta \Delta H\) and \(\Delta \varDelta G\) values were mostly consistent between the runs (Supplementary Fig. 4g). We took the median values from the 4 runs (\(\Delta \Delta H=-28\,{{{\rm{kcal}}}}/{{{\rm{mol}}}}\), \(\Delta \Delta G=-4\,{{{\rm{kcal}}}}/{{{\rm{mol}}}}\)) and added them to the aggregated \(\Delta \Delta H\) and \(\varDelta \Delta G\) values of hairpin loops.
Augmenting the compact 178-parameter model with these curated hairpin loop parameters improved that model performance on the validation set of 1315 variants. This improvement might be due to the predominant overrepresentation of ‘GAAA’ hairpin loops in Array Melt data, which resulted in a highly skewed data distribution and reduced the accuracy of directly regressed hairpin loop parameters.
The rich parameter model
We trained the rich parameter model on the same Array Melt training dataset as for the dna24 model. For feature extraction, we used nearest-neighbor Watson-Crick pairs and terminal stack penalty as in NUPACK, and the following parameters for loops:
-
1)
Hairpin loops. For triloops and tetraloops, \({\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpinseq}}={\varDelta G}_{{\phi }_{j-1},{\phi }_{j},{\phi }_{i},{\phi }_{i+1}}^{{hairpinmm}}+{\varDelta G}_{{\phi }_{\left[i+1:j-1\right]}}^{{hairpinloop}}\), where \({\varDelta G}_{{\phi }_{\left[i+1:j-1\right]}}^{{hairpinloop}}\) was the full sequence of the hairpin loop, in contrast to only the middle of loops in the dna24 model. Hairpin loops longer than tetraloop are rare in our dataset, so we fitted a single loop sequence parameter \({\varDelta G}_{{long}}^{{hairpinloop}}\). The final energy of hairpin was also \({\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpin}}={\varDelta G}_{n}^{{hairpinsize}}+{\varDelta G}_{{\phi }_{\left[i:j\right]}}^{{hairpinseq}}\).
-
2)
Interior loops. For single and double bulges or single mismatches, a sequence-dependent parameter, \({\varDelta G}_{{\phi }_{\left[i:d\right]},{\phi }_{\left[e:j\right]}}^{{interiorseq}}\), was fitted for each sequence.
This rich parameter model was implemented in custom Python code without interactions with the NUPACK software.
Deep learning models
The closest sequence lookup baseline model
We defined the distance between any two sequence-structure pairs as the sum of the edit distance between the sequences and the dot-bracket secondary structure representations. For every query sequence-structure pair, we found the k closest sequence-structure pairs in the training set, whose distance-weighted sum became the predicted value for the query using Python package scipy44. We found the best value for k using the held-out Array Melt validation data.
Graph neural network model
We implemented the graph neural network using Python packages PyTorch and PyTorch Geometric45. The final model had 4 graph transformer layers with a 125-vec node attribute at each layer, stacked with dropout of 0.01 at each layer, a Set2Set module with 10 processing steps, and one fully connected hidden layer with 128 channels and a dropout ratio of 0.49. We normalized ΔH and Tm of the training data to fall within 0 and 1 and used the sum of the root mean square error (RMSE) of both ΔH and Tm as the loss function. We trained with Adam optimizer (learning rate = 0.002379; betas = 0.9, 0.999; weight decay = 0) for 200 epochs. The model had 287,136 parameters. The validation model registry was monitored and stored on weight and biases (wandb.ai).
Using the NUPACK-compatible model
The parameter file for the NUPACK-compatible dna24 model is available in Supplementary Data19 as dna24.json. To use this file, download and point to it when using the python package NUPACK (v4.0.0.27)46:
my_model = nupack.Model(material = ’/path/to/parameter/file/dna24.json’, …)
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The Array Melt dataset generated in this study, including processed fluorescence data, thermodynamic parameters, imaging scripts, and trained model files, has been deposited in Zenodo under accession code https://doi.org/10.5281/zenodo.1529957819. Source data supporting the figures and tables in the main text and Supplementary Information are provided with this paper in the Source Data file. No restrictions apply to data access.
Code availability
The code used to develop the model, perform the analyses, and generate results in this study is publicly available and has been deposited in the repository nnn_paper at https://github.com/GreenleafLab/nnn_paper/ under MIT license. The specific version of the code associated with this publication is archived in Zenodo and is accessible via: https://doi.org/10.5281/zenodo.1529968047. Additional code, not needed for figure reproduction, specifically for earlier Array Melt data preprocessing, is available at https://github.com/GreenleafLab/array_analysis.
References
Kunkel, T. A. DNA replication fidelity. J. Biol. Chem. 279, 16895–16898 (2004).
Rajski, S. R., Jackson, B. A. & Barton, J. K. DNA repair: models for damage and mismatch recognition. Mutat. Res. Mol. Mech. Mutagen. 447, 49–72 (2000).
Untergasser, A. et al. Primer3—new capabilities and interfaces. Nucleic Acids Res.40, e115 (2012).
Kibbe, W. A. OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Majikes, J. M. et al. Revealing thermodynamics of DNA origami folding via affine transformations. Nucleic Acids Res. 48, 5268–5280 (2020).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
SantaLucia, J. & Hicks, D. The thermodynamics of DNA structural motifs. Annu. Rev. Biophys. Biomol. Struct. 33, 415–440 (2004).
SantaLucia, J., Allawi, H. T. & Seneviratne, P. A. ImprOved Nearest-neighbor Parameters for Predicting DNA duplex stability. Biochemistry 35, 3555–3562 (1996).
SantaLucia, J. A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics. Proc. Natl. Acad. Sci. USA 95, 1460–1465 (1998).
Zuker, M. On finding all suboptimal foldings of an RNA molecule. Science 244, 48–52 (1989).
Schroeder, S. J. & Turner, D. H. Optical melting measurements of nucleic acid thermodynamics. Methods Enzymol. 468, 371–387 (2009).
Spink, C. H. Differential Scanning Calorimetry. in Methods in Cell Biology vol. 84 115–141 (Academic Press, 2008).
Hadiwikarta, W. W., Walter, J.-C., Hooyberghs, J. & Carlon, E. Probing hybridization parameters from microarray experiments: nearest-neighbor model and beyond. Nucleic Acids Res. 40, e138 (2012).
Oliveira, L. M., Long, A. S., Brown, T., Fox, K. R. & Weber, G. Melting temperature measurement and mesoscopic evaluation of single, double and triple DNA mismatches. Chem. Sci. 11, 8273–8287 (2020).
Bae, J. H., Fang, J. Z. & Zhang, D. Y. High-throughput methods for measuring DNA thermodynamics. Nucleic Acids Res. 48, e89–e89 (2020).
You, Y., Tataurov, A. V. & Owczarzy, R. Measuring thermodynamic details of DNA hybridization using fluorescence. Biopolymers 95, 472–486 (2011).
Fornace, M. E., Porubsky, N. J. & Pierce, N. A. A unified dynamic programming framework for the analysis of interacting nucleic acid strands: enhanced models, scalability, and speed. ACS Synth. Biol. 9, 2665–2678 (2020).
Ke, Y., Sharma, E. & Wayment-Steele, H. Dataset for High-throughput DNA melt measurements enable improved models of DNA folding thermodynamics. Zenodo https://doi.org/10.5281/zenodo.15299578 (2025).
Tikhomirova, A., Beletskaya, I. V. & Chalikian, T. V. Stability of DNA duplexes containing GG, CC, AA, and TT mismatches. Biochemistry 45, 10563–10571 (2006).
Khandelwal, G. & Bhyravabhotla, J. A phenomenological model for predicting melting temperatures of DNA sequences. PLOS ONE 5, e12433 (2010).
Yan, Z., Hamilton, W. L. & Blanchette, M. Graph neural representational learning of RNA secondary structures for predicting RNA-protein interactions. Bioinformatics 36, i276–i284 (2020).
Zhao, Q. et al. Review of machine learning methods for RNA secondary structure prediction. PLOS Comput. Biol. 17, e1009291 (2021).
Shi, Y. et al. Masked Label Prediction: Unified Message Passing Model for Semi-Supervised Classification. In Proc. IJCAI 1548–1554. https://doi.org/10.24963/ijcai.2021/214 (2021).
Veličković, P. et al. Graph Attention Networks. In Proc. ICLR https://openreview.net/forum?id=rJXMpikCZ (2018).
Vinyals, O., Bengio, S. & Kudlur, M. Order Matters: Sequence to sequence for sets. In Proc. ICLR https://doi.org/10.48550/arXiv.1511.06391 (2016).
Spasic, A. et al. Improving RNA nearest neighbor parameters for helices by going beyond the two-state model. Nucleic Acids Res. 46, 4883–4892 (2018).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Mathews, D. H., Sabina, J., Zuker, M. & Turner, D. H. Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J. Mol. Biol. 288, 911–940 (1999).
Dotu, I., Mechery, V. & Clote, P. Energy parameters and novel algorithms for an extended nearest neighbor energy model of RNA. PLOS ONE 9, e85412 (2014).
Zakov, S., Goldberg, Y., Elhadad, M. & Ziv-ukelson, M. Rich parameterization improves RNA structure prediction. J. Comput. Biol. 18, 1525–1542 (2011).
Boiko, D. A., MacKnight, R., Kline, B. & Gomes, G. Autonomous chemical research with large language models. Nature 624, 570–578 (2023).
Buenrostro, J. D. et al. Quantitative analysis of RNA-protein interactions on a massively parallel array reveals biophysical and evolutionary landscapes. Nat. Biotechnol. 32, 562–568 (2014).
She, R. et al. Comprehensive and quantitative mapping of RNA–protein interactions across a transcribed eukaryotic genome. Proc. Natl. Acad. Sci. USA 114, 3619–3624 (2017).
Good, N. E. et al. Hydrogen ion buffers for biological research. Biochemistry 5, 467–477 (1966).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Altschul, S. F. & Gish, W. [27] Local alignment statistics. in Methods in Enzymology vol. 266 460–480 (Academic Press, 1996).
Denny, S. K. et al. High-throughput investigation of diverse junction elements in RNA tertiary folding. Cell 174, 377–390.e20 (2018).
Köster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Newville, M., Stensitzki, T., Allen, D. B. & Ingargiola, A. LMFIT: Non-Linear Least-Square Minimization and Curve-Fitting for Python. Zenodo https://doi.org/10.5281/zenodo.11813 (2014).
Lebigot, E. O. Uncertainties: a Python package for calculations with uncertainties. Version 3.1.7, http://pythonhosted.org/uncertainties/ v3.1.7. (2022).
Fischler, M. A. & Bolles, R. C. Random sample consensus: a paradigm for model fitting with applications to image analysis and automated cartography. Commun. ACM 24, 381–395 (1981).
Owczarzy, R. et al. Effects of sodium ions on DNA duplex oligomers: improved predictions of melting temperatures. Biochemistry 43, 3537–3554 (2004).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Fey, M. & Lenssen, J. E. Fast Graph Representation Learning with PyTorch Geometric. In Proc. ICLR-W. https://doi.org/10.48550/arXiv.1903.02428 (2019).
Fornace, M. E. et al. NUPACK: Analysis and Design of Nucleic Acid Structures, Devices, and Systems. ChemRxiv https://doi.org/10.26434/chemrxiv-2022-xv98l (2022).
Ke, Y. GreenleafLab/nnn_paper: v1.0. Zenodo https://doi.org/10.5281/zenodo.15299680 (2025).
Acknowledgements
We thank Dr. Rhiju Das (Stanford University) and Owen Dunkley (Stanford University) for comments on the manuscript; Dr. Mark Schnitzer (Stanford University) for supporting the project; Dr. Niles Pierce and Dr. Mark Fornace (Caltech) for answering NUPACK-related questions. We thank Emerald Cloud Lab (ECL) for the UV melting experiments, with special attention to Malav Desai and Ben Kline for tutorials and technical support of Symbolic Lab Language (SLL) programming, and Brian Frezza for funding. This work was supported in part by NIH grants R01GM111990, P50HG007735, R01HG009909, P01GM066275, UM1HG009436, DP1HG013599, and R01GM121487 to W.J.G. W.J.G. acknowledges support as a Chan Zuckerberg Investigator. E.M. acknowledges support from the Swedish Research Council (grant 2020-06459), the Foundation Blanceflor, and the Science for Life Laboratory (SciLifeLab).
Author information
Authors and Affiliations
Contributions
Conceptualization, W.J.G., Y.K., E.S., and H.K.W.; Methodology, Y.K., E.S.; Software, Y.K. and H.K.W.; Validation, Y.K.; Formal Analysis, Y.K.; Investigation, Y.K., E.S., H.K.W., W.R.B., and A.H.; Data Curation, Y.K.; Writing—Original Draft, Y.K. and W.J.G.; Writing—Review and Editing, Y.K., E.S., H.K.W., E.M., and W.J.G.; Visualization, Y.K.; Supervision, E.M. and W.J.G.; Funding Acquisition, W.J.G.
Corresponding author
Ethics declarations
Competing interests
Stanford filed a patent application on aspects of this work with H.K.W., E.S., R.D., W.J.G., A.H., W.B., and Y.K. as inventors (WO2023028618). W.J.G. is a consultant and equity holder for 10x Genomics, Guardant Health, Quantapore, and Ultima Genomics, and cofounder of Protillion Biosciences. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hashim Al-Hashimi, Stefan Badelt, and Constantine Evans for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ke, Y., Sharma, E., Wayment-Steele, H.K. et al. High-throughput DNA melt measurements enable improved models of DNA folding thermodynamics. Nat Commun 16, 5572 (2025). https://doi.org/10.1038/s41467-025-60455-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60455-4