Abstract
The combination of quantitative phase microscopy (QPM) with imaging flow cytometry (IFC) enables label-free and multi-parameter single-cell analysis. Here, we present a simple yet powerful QPM-IFC platform, the spatial microfluidic holographic integrated (SMHI) platform, which uniquely integrates spatial hydrodynamic focusing microfluidics with digital holographic microscopy (DHM) to achieve high-fidelity single-cell QPM reconstruction without digital refocusing in 0.34 seconds, accounting for only 4.41% of the typical process ( ~ 7.71 seconds). We develop a high-dimensional phase feature hierarchy and implement a maximun-relevance and minimun-redundancy incremental feature selection (MRMR-IFS) strategy, which effectively addresses feature redundancy and constructs the optimal feature set. Consequently, a prediction accuracy of >99.9% is achieved across multiple cancer cell types, breast cancer subtypes, and blood cells, demonstrating its efficacy in analyzing highly heterogeneous cell populations. Notably, this system also exhibits high accuracy in analyzing simulated blood samples, highlighting its great potential in practical applications.
Similar content being viewed by others
Introduction
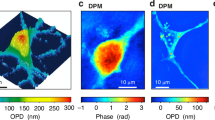
Single-cell research has significantly advanced our understanding of fundamental biological processes and opened new avenues in precision medicine1,2,3. Imaging flow cytometry (IFC), capitalizing on its high-throughput and information-rich advantages, has emerged as a crucial tool for analyzing large-scale single-cell populations4,5. IFC allows for the capture of individual cell images in the detection zone, permitting quantitative assessment of cellular morphology, structure, and multiple biomarkers6, thereby fueling progress in key scientific domains including oncology7,8, immunology9, microbiology10, and metabolomics11. However, IFC’s reliance on diverse labeling presents challenges, such as potential cytotoxicity, non-specificity and interference that may obscure detection results, and complex sample preparation12. Dependency on specific markers is also problematic for cell populations lacking prior knowledge or with frequently changing surface antigens. Therefore, achieving label-free single-cell imaging is a significant and enduring issue, essential for minimizing sample perturbation and simplifying the analytical process. Multiple label-free optical imaging methods have emerged, including quantitative phase microscopy (QPM)13, optical coherence tomography (OCT)14,15, photoacoustic imaging (PAI)16, stimulated Raman scattering (SRS)17, and polarization microscopy18,19, which capture the amplitude, phase, frequency, or polarization of the sample’s inherent properties20. Among them, QPM is distinguished by its non-invasive nature, high contrast, and versatile compatibility. QPM transcends mere visualization of cellular exteriors, such as dimensions and contours21,22,23. By investigating the subtleties of optical path length differences (OPD)24 and integral refractive index (RI) distribution25 through quantifying phase delay, QPM demonstrates more complicated textural characteristics26,27 that cannot be obtained by other imaging methods.
However, the main challenge in combining QPM with IFC is achieving precise single-cell QPM imaging under rapid flow rates, as cells’ optical defocusing are prone to motion blur or elongation distortion28. Early attempts directly merged QPM imaging with commercial flow cytometers29 to utilize their fluidic systems for cell focusing. However, this resulted in bulky and complicated setups. Hence, there is a trend towards compact microfluidic focusing techniques30, with active manipulation of particle focusing through electric31, magnetic32, or acoustic33 fields being efficient but costly and complex due to additional equipment and energy consumption. In contrast, some passive methods, using special channel geometries (such as inertial focusing34,35) or fluid properties (such as hydrodynamic focusing) for focusing, are more economical and biocompatible. Recent advancements in inertial microfluidics36,37,38 demonstrate impressive particle focusing capabilities, yet these approaches generally require complex channel designs (e.g., long bends) and are more dependent on particle physical properties (e.g., size, shape, stiffness, density), which may limit their integration with QPM-IFC systems. By comparison, hydrodynamic focusing stands out for its design simplicity, operational convenience, and broad applicability across various particle sizes and flow rates39,40, along with high compatibility with QPM that facilitates diverse dynamic particle detection. Despite this, a review of current research shows that hydrodynamic focusing is typically conducted laterally to direct cells past the detection area in a single channel. Axially, cells are either unconstrained41,42,43,44 or loosely confined45,46, leading to cell dispersion and potential overlap which can compromise single cell identification precision. While digital refocusing algorithms can help accurately locate cells at different depths47, they are computationally expensive and time-consuming48, slowing down IFC’s rapid workflow. The lack of axial confinement also causes cell oscillation, imaging artifacts and blurriness, and finally loss of key textures in QPM measurements. Therefore, there is a clear need for more precise 3D cell positioning control to improve QPM measurement accuracy and stability for high-quality imaging.
Advances have been made in the label-free classification of biological samples using QPM in conjunction with artificial intelligence technologies such as machine learning and deep learning, encompassing diverse cell types such as blood cells49, immune cells45,50,51, cancer cells52,53,54, sperm55, and bacteria56. Raw QPM images or their quantified features serve as model inputs, with the latter being preferred for its strong interpretability, which can directly correlate QPM phenotypes with biomedical knowledge, thereby avoiding black box issues57,58. Recently, some systematic feature extraction frameworks, which cover morphology, pixel intensity, and textural features, have been built to improve classification accuracy39,59. Nevertheless, the computational and memory burdens associated with high-dimensional QPM feature spaces, as well as the risks of redundancy and overfitting, have not been adequately addressed. Current research frequently neglects the critical process of feature selection60 and does not adequately optimize the combination of features, which is essential for reducing noise, enhancing model generalizability, lowering computational expenses, and ultimately avoiding the curse of dimensionality61. Besides, the scarcity of studies on the classification of extensive and highly heterogeneous cell populations constrains the practical applicability of the technology59. In summary, thorough analysis and selection of features, combined with rigorous testing of machine learning algorithms on complex datasets, are crucial for achieving efficient and precise cell classification, ensuring robust and generalizable outcomes.
In this study, we design a spatial microfluidic holographic integrated (termed SMHI) platform, which combines a compact digital holographic microscopy (DHM) system and a spatial hydrodynamic focusing microfluidic technique for label-free, single-shot QPM imaging. DHM is one of the most representative technologies in the field of QPM due to its advantages of non-contact, high resolution and low cost. By aligning the DHM’s recording focal plane with the sample flow’s hydrodynamic focusing plane, we can directly capture clear, high-quality single-cell holograms. This integration ensures precise confinement of particles/cells within the core of the microchannel, bringing them into the in-focus DHM imaging plane to substantially enhance imaging quality. Therefore, there is no need for digital refocusing anymore, significantly reducing the QPM phase reconstruction time. We then analyze QPM phase features and compare different machine learning algorithms. The incremental feature selection (IFS) strategy based on the Maximun-Relevance and Minimun-Redundancy (MRMR) algorithm62 ranking, previously unused in phase feature training, has finally achieved >99.9% accuracy in predicting multiple cancer cell types, breast cancer subtypes and blood cells, which proves the platform’s efficacy for characterizing heterogeneous cell populations and broad application prospects.
Results
General principle and workflow of the SMHI platform
Our SMHI platform was constructed by systematically integrating three modules, including microfluidics, digital holography, and machine learning (Fig. 1). In the microfluidics module, we defined spatial hydrodynamic focusing to precisely control and strictly constrain the 3D positioning of single cells (Supplementary Note 1). Specifically, two identical vertical sheath flows were first introduced to continuously and symmetrically compress the sample flow in the vertical direction, forming a Sandwich-like structure. As these three laminar flows advanced along the microchannel and reached a designated confluence point, they were enveloped by the other two horizontal sheath flows entering at a 45° angle. This interaction completed the spatial focusing of the sample flow within a single-layer microchannel. Consequently, cells were guided to flow in an orderly fashion along the central axis. Sheath flows’ constraint reduced image blur caused by the rapid movement and particles aggregation, while minimizing sample contamination and channel blockage.
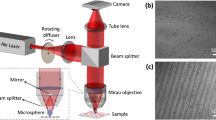
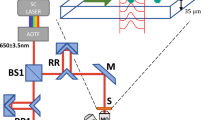
a Schematic of the spatial hydrodynamic focusing digital holographic microscopy (DHM) imaging. In the microfluidic module, two vertical sheath flows (blue) are initially employed to focus the sample flow (gray) vertically, followed by two horizontal sheath flows (yellow) to complete the spatial focusing. By precisely controlling fluid dynamics, each cell is directly guided onto the DHM’s focal plane, finally producing in-focus single-cell holograms, which are recorded by a complementary metal-oxide-semiconductor (CMOS) camera. The main optical components of DHM include: beam splitter (BS), reflector (REFL), microscope objective (MO), tube lens (TL), and neutral density filter (NDF). b Quantitative phase microscopy (QPM) reconstruction process. It is performed on the holograms to obtain precise phase maps of single cells, including frequency filtering, numerical reconstruction, phase extraction, phase unwrapping and phase correction. c Phase feature extraction process. For each QPM phase map, the region of interest (ROI) is segmented by mask, and 81 features are extracted, including 14 bulk features, 17 first-order histogram features, and 50 high-order texture features, which are composed of 20 gray-level co-occurrence matrix (GLCM) features, 15 gray-level gradient co-occurrence matrix (GGCM) features and 15 gray-level run-length matrix (GLRLM) features. d Phase feature analysis process. It includes numerical distribution visualization, significance testing and correlation analysis to comprehensively assess the feature datasets. e Machine learning model construction process. First, the features are scored and ranked for importance using the maximun-relevance and minimun-redundancy (MRMR) algorithm. Machine learning models are then incrementally trained by adding features one by one, with classifier performance finally evaluated using confusion matrices and receiver operating characteristic (ROC) curves.
In the digital holography module, our custom-built DHM system (Supplementary Note 2) featured a 40× objective lens with a numerical aperture (NA) of 0.75 and a light source emitting at a wavelength of 633 nm, thus achieving a sub-micrometer lateral resolution. We used a cage-like structure design which can enhance system stability by dampening external vibrations and maintaining precise alignment of optical components. This design contributed to the high spatiotemporal stability of phase measurement (detailed in Supplementary Note 2). Specifically, the standard deviation of temporal phase noise (0.035 rad) is significantly lower than that of typical Mach-Zehnder configurations in DHM (0.2 rad)63,64,65, highlighting the improved performance of our system. The standard deviation of spatial phase noise (0.007 rad) is much lower than the average phase height of typical cells (3-5 rad), thus ensuring minimal impact on measurements.
The DHM system’s CMOS camera automatically captured images at a resolution of 496 pixels × 496 pixels, which corresponded to a spatial scale of 31 μm × 31 μm. The DHM’s recording focal plane was meticulously aligned with the spatial focusing axis of the microfluidic system, enabling the direct acquisition of an in-focus hologram of single cells and particles, thereby eliminating the need for digital refocusing, which is time-consuming and computationally intensive45 (Supplementary Note 3). After measurement, our system completed the reconstruction of a single-cell hologram in approximately 0.34 s using lab-written MATLAB codes on a laptop equipped with a 2.8 GHz Intel Core i7 CPU and 16 GB of RAM. This was significantly faster compared to the typical process that requires digital refocusing, which take about 7.71 s47 (detailed in Supplementary Note 3). Our reconstruction time is only about 4.41% of the latter.
In the machine learning module, a multi-level feature set (Supplementary Note 4) was established from the phase maps of a single cell or particle. We preliminarily assessed the efficacy of each feature using variance analysis and determined their interrelationships via correlation coefficients. To select the most important features for classification, feature engineering (Supplementary Note 5) was introduced to derive the importance scores of features through specific algorithms, and incrementally adding features in descending order of their scores to train classifiers. The strategy was then applied across various machine learning classifiers, with classification performance being evaluated through confusion matrices and Receiver Operating Characteristic (ROC) curves (Supplementary Note 6, 7).
Theoretical simulation and experimental validation of spatial hydrodynamic focusing
In the spatial hydrodynamic focusing, the velocity ratio between sheath and sample flows is crucial for determining the size of the focused particle stream. To clarify this relationship, we commenced our investigation with theoretical simulations (Supplementary Note 1). Utilizing COMSOL Multiphysics 6.1, we modeled the 3D microchannel with a cross-sectional design (x-z plane) mirroring the actual microfluidic chip dimensions of 100 μm by 35 μm. We configured the simulation for laminar flow physics, assuming incompressible and immiscible fluids with no-slip boundary conditions applied to all channel walls. The sample flow was delineated by velocity streamlines (Supplementary Fig. 1a, 2a).
Firstly, we varied the velocity ratio of the vertical sheath flows to the sample flow (Rv) from 0 to 1.5 in 0.25 increments. The focused sample flow’s height (Hs) in the vertical channel cross-section (y-z plane) was observed to diminish from 35 μm to 5.9 μm (Supplementary Table 1), conforming to a fitted one-phase exponential decay model (Supplementary Fig. 1a, c). Secondly, with Rv stabilized at 0.25 (a deliberate choice as its corresponding Hs is 16.9 μm, representing a mid-range level), we adjusted the velocity ratio of the horizontal sheath flows relative to the combined flow of the upper vertical sheath flow, middle sample flow, and lower vertical sheath flow (Rh) from 0 to 5 in 0.5 increments. The focused sample flow’s width (Ws) in the horizontal channel cross-section (x-y plane) was measured and exhibited a decrease from 100 μm to 6.1 μm (Supplementary Table 2), also adhering to a one-phase exponential decay trend (Supplementary Fig. 2a, c).
To validate our simulations, we employed Rhodamine B, a fluorescent dye, to visualize the sample flow and evaluate the spatial focusing effect within an actual microfluidic chip (Fig. 2a). Using three syringe pumps, we precisely controlled the flow rates of the sample and sheath flows, conducting experiments under conditions paralleling our simulations. The focused sample flow’s size was measured under an inverted fluorescence microscope and quantified by the fluorescence full width at half maximum (FWHM). The experimental measurements (Fig. 2b, c and Supplementary Fig. 1b, c and 2b, c) correlated well with our simulation outcomes (Supplementary Table 1 and 2), confirming that the focusing effect can be accurately manipulated by adjusting the velocity ratios to accommodate cells and particles of various sizes.
a Schematic representation of vertical focusing (y-z plane) and horizontal focusing (x-y plane). b Experimental results of vertical focusing (y-z plane) and horizontal focusing (x-y plane). The red band represents the sample flow containing fluorescent dye Rhodamine B. The dashed lines represent the boundaries of the microfluidic channels. The sample injection speed is 1 μL/min. Rv represents the velocity ratio of vertical sheath flow to sample flow. Rh represents the velocity ratio of horizontal sheath flow to the combined flows of two vertical sheath flows and sample flow. For the horizontal focusing test, Rv is fixed at 0.25. Each focused sample flow height (Hs) and width (Ws) is the mean of 5 independent repeated experiments under the corresponding Rv and Rh conditions. c Statistic results of spatial hydrodynamic focusing. It shows the change in Hs with Rv, and Ws with Rh. Data (red dots) are mean values from 5 independent repeated experiments; error bars are SD. d, e show the flow observation of 10 μm (d) and 20 μm (e) polystyrene microspheres in microfluidic channels under no focusing, planar focusing, and complete spatial focusing conditions in white light illumination. f, g show the DHM reconstructed amplitude (top) and phase (bottom) maps of 10 μm (f) and 20 μm (g) microspheres under the spatial focusing. The color bar denotes the scale of phase. In (d–g), each microsphere constitutes an independent experimental replicate, and finally, amplitude and phase data are collected from 50 independently captured microspheres under each focusing condition for further analysis. h, i show the reconstructed particle size of 10 μm (h) and 20 μm (i) microspheres under different focusing conditions from amplitude. j, k show the reconstructed particle size of 10 μm (j) and 20 μm (k) microspheres under different focusing conditions from phase. l, m show the distance from particle center to channel centerline of 10 μm (l) and 20 μm (m) microspheres under different focusing conditions. In (h–m), data are presented as means ± SD (n = 50 microspheres under each focusing condition); the p-value < 0.05 indicates a significant difference decided by one-way ANOVA followed by Tukey’s multiple comparisons test. Scale bars: 30 μm (b, y-z plane), 100 μm (b, x-y plane), 50 μm (d), 50 μm (e), 10 μm (f), 10 μm (g). Source data are provided as a Source Data file.
It is essential to acknowledge that there exists an upper limit for the flow rates of both the sheath and sample flows. Surpassing these limits may result in pressure disparities within the microchannels and unstable focusing. To investigate this, we fixed the velocity ratios at Rv = 0.25 and Rh = 2, because the corresponding Hs (= 16.9 μm) and Ws (= 13.5 μm) were close to the average cell size (10–20 μm), and measured the consequent fluctuations in the focused flow’s Ws and Hs, as the sample flow rate was incrementally varied from 1 to 20 μL/min (~ 1.46 m/s). The results (Supplementary Fig. 3a–f and Supplementary Table 3 and 4) demonstrated that the spatial hydrodynamic focusing effect remained stable across this range of velocities, with only minor variations observed in Ws and Hs. This finding demonstrated the robustness of our focusing technique even at elevated flow speeds.
Label-free DHM detection of spatial hydrodynamic focusing microparticles
To validate the detection capabilities of our platform for spatial microparticles, we employed 10 μm and 20 μm polystyrene microspheres as proxies for cancer cells, considering their standardized physical properties and consistent dimensions. To begin with, we obstructed the holographic optical path (see Supplementary Note 2 and Supplementary Figs. 4–7 for details) and directly observed the microsphere focusing process under external white light illumination. The microsphere suspension was prepared at a concentration of approximately 1 × 106 particles/mL, and the flow rate was reduced to 1 μL/min to facilitate observation. As shown in Fig. 2d, e, without any focusing, the microspheres traversed the channel with vertical stacking and lateral floating, resulting in inconsistent focal clarity. Upon applying vertical sheath flow constraints, the microspheres were compressed vertically into a single plane, yet they continued to oscillate and float laterally. We refer to this scenario as planar focusing. The addition of horizontal sheath flows subsequently aligned the microspheres onto the channel axis to complete the spatial focusing.
We next captured holograms of a single microsphere within the detection window. Following frequency domain filtering and numerical reconstruction (see Supplementary Note 3 and Supplementary Figs. 8–10 for details), we obtained amplitude maps of the microspheres (Fig. 2f, g). Subsequent phase extraction, unwrapping and distortion compensation processes (Supplementary Fig. 10a–c) led to the final reconstruction of the QPM phase maps of the microspheres (Fig. 2f, g and Supplementary Fig. 10a, b). Through quantitative comparisons, spatial focusing demonstrates three critical advantages over planar/no focusing. First, it enhances the accuracy and efficiency of amplitude and phase reconstruction. The amplitude-based particle sizes were derived from the mean of measured length/width in reconstructed amplitude maps, while phase-based sizes were calculated via Eq. (1) using peak-to-valley phase differences (see Methods section). Statistical analysis of 50 microspheres per condition confirmed spatial focusing’s necessity for both amplitude (Fig. 2h, i) and phase (Fig. 2j, k) reconstruction. For instance, the phase size values of 10 μm and 20 μm microspheres without focusing were 7.13 μm (Fig. 2j) and 16.5 μm (Fig. 2k), respectively. However, with spatial focusing, these values were refined to 10.4 μm (Fig. 2j) and 19.8 μm (Fig. 2k), closely aligning with their true sizes. This accuracy improvement, coupled with a reduction in reconstruction time (0.34 s vs. 7.71 s per hologram) through eliminated digital refocusing needs (Supplementary Table 5), establishes spatial focusing as a precision-efficiency approach. Second, spatial focusing significantly reduced the average distance from particle centers to the microchannel centerline. For 10 µm and 20 µm microspheres, these distances are (1.5 ± 0.8) µm and (1.6 ± 1.0) µm, respectively, largely smaller than the other two focusing models (Fig. 2l, m; see Supplementary Fig. 11a, b and Table 6 for details), thus enabling direct imaging through a fixed acquisition window. This approach eliminates the need for particle tracking and segmentation algorithms typically required in planar or non-focusing processes, which can introduce computational complexity and potential misidentification errors, thus laying the foundation for automated hologram capture. Finally, spatial focusing enhanced measurement consistency through flow stabilization. By constraining the particles’ trajectory both vertically and laterally, the numerical distribution of amplitude and phase of the particles displayed more compact (Fig. 2h–k). The coefficient of variation (CV), a metric indicative of the precision and stability of hydrodynamic focusing and measurement systems in IFC, was employed to evaluate the platform comprehensively. Our calculations (see Supplementary Table 7) indicated that the CVs for amplitude and phase measurements reached 3.3%/4.3% (10 μm microspheres) and 1.9%/4.7% (20 μm microspheres) under spatial focusing. These values align with previous studies on the measurement consistency in flow cytometry, where CVs are typically < 5%66.
QPM phase features extraction and analysis of multiple cancer cell types
An accurate, rapid, stable, and label-free measurement platform facilitates the high-throughput analysis of single-cell images with high-information content. Here, we highlight the platform’s proficiency in capturing single-cell phase. We selected five cell lines representative of highly prevalent cancers: MDA-MB-231 for breast cancer, A549 for lung cancer, HCT116 for colorectal cancer, HeLa for cervical cancer, and THP-1 for human monocytic leukemia. The single-cell suspension was prepared at a concentration of approximately 1 × 106 cells/mL, with a flow rate of 1 μL/min.
From the QPM phase maps (Fig. 3a), it was evident that THP-1 cells were significantly smaller than the other cancer cells. However, given the intrinsic heterogeneity among cells, even those of the same type can display considerable morphological variance. Thus, cell typing based solely on image observation or a single feature is impractical. We noticed that the sub-micrometer resolution of QPM clearly preserved subcellular textures. Consequently, we developed a hierarchical QPM phase feature extraction framework that quantified 14 bulk features, 17 first-order histogram features, and 50 high-order texture features, totally 81 features, to thoroughly characterize single-cell phenotypes (see Supplementary Note 4 and Supplementary Table 8 for details, including feature names and numbers). Violin plots (Fig. 3b) show the distributions of selected cell features, from which we can see differences among the 5 cell lines. Then, one-way analysis of variance (ANOVA) coupled with multiple comparisons (see Supplementary Note 5 for details) was further employed to assess whether these differences were statistically significant in a pairwise manner. We sequentially tested 81 features and concentrated on presenting the normalized mean value differences of each feature across different cancer cell types (Fig. 3c, outer circle), as well as the results of multiple comparisons (Fig. 3c, inner circle), in the circular heatmap following a clockwise arrangement. According to statistics, a total of 22 features showed significant differences in all pairwise group comparisons, suggesting that these features may be more useful than others in distinguishing between different cell types. Finally, a Pearson correlation matrix was computed to examine inter-feature correlations (see Supplementary Note 5 for details), visualized via a heatmap (Fig. 3d). We can see that highly correlated features were mainly clustered within the same category (enclosed by yellow boxes) and less so across different categories, signifying the efficacy of our hierarchical strategy. However, there was a certain degree of autocorrelation and cross-correlation among the subsets of texture features, including gray-level co-occurrence matrix (GLCM), gray-level gradient co-occurrence matrix (GGCM), and gray-level run-length matrix (GLRLM), suggesting that these features encoded closely related or similar cellular information, thereby indicating the existence of redundancy. Regardless, our hierarchical QPM feature extraction scheme has preliminarily achieved a comprehensive and efficient characterization of subcellular feature discrepancies within heterogeneous cancer cell populations in a label-free and at-scale manner.
a Single-cell QPM phase maps of 5 cancer cell lines, including MDA-MB-231 (breast cancer), A549 (lung cancer), HCT116 (colorectal cancer), HeLa (cervical cancer), and THP-1 (monocytic leukemia). The color bar denotes the scale of normalized optical path difference (OPD) to reflect phase information (Scale bar: 10 μm). b Violin plots of 6 QPM phase features (n = 1000 cells for each cancer type), including volume (Bulk), dry mass density (Bulk), kurtosis (First-order histogram), entropy (GLCM), gray asymmetry (GGCM), and run variance (GLRLM). The violins are drawn with the ends at the quartiles and the median as a horizontal line in the violin, where the curve of the violin extends to the minimum and maximum values in the data set. *** indicates p-value < 0.001, ** indicates 0.001 <p-value < 0.01, * indicates 0.01 <p-value < 0.05, ns indicates p-value > 0.05, based on one-way ANOVA followed by Tukey’s multiple comparisons test. In (c), the outer circular heatmap displays the mean value distribution of 81 features (arranged in a clockwise direction) across 5 distinct cancer cell types (ordered from the center out). The color bar denotes the scale of normalized mean values (n = 1000 cells for each cancer type). The inner binary heatmap shows the results of ordinary one-way ANOVA pairwise comparisons (Tukey’s test). Dark blue visualizes whether these differences are statistically significant. The significance level (α) is set to 0.05. d Pearson correlation matrix of 81 features visualized as a heatmap (n = 1000 cells for each cancer type). Blue indicates positive correlation, while red indicates negative correlation. Different feature subsets are divided by yellow boxes. The color bar represents the scale of the Pearson correlation coefficient. Source data are provided as a Source Data file.
Multiple cancer cell types prediction using feature engineering and machine learning
Our feature dataset comprised 5000 cancer cells-1000 from each of the five types-as well as 3000 blank backgrounds, with 81 phase features extracted from each cell. Initially, we trained 9 classic machine learning classifiers on all 81 features (Supplementary Note 5). The results (see Supplementary Table 9 and 10) showed that nearly all methods provided creditable accuracy, validating the efficacy of the extracted QPM phase features. Linear Discriminant Analysis (LDA) achieved 100% prediction accuracy, closely followed by Support Vector Machine (SVM), Neural Networks (NN) and Logistic Regression (LR). Specifically, for the underperforming K-Nearest Neighbors (KNN) and Decision Trees (DT), we enhanced classification accuracy to 98.4% and 98.0%, respectively, by creating an ensemble of KNN using the random subspace method (Subspace KNN) and an ensemble of DT using a bagging strategy (Bagged Trees). The only poor performer was Naive Bayes with an overall accuracy of 87.9%, due to its assumption of feature independence, which is not the case, as revealed by our feature analysis.
However, excessive input features raise the risk of overfitting and computational costs. Therefore, we aimed to reduce feature dimensionality while ensuring high accuracy. Based on our hierarchical extraction strategy, natural feature subsets were formed. We first attempted to train models using each subset individually, but the results (see Supplementary Fig. 12a–f, Supplementary Table 9 and 10) indicated that the accuracy achieved using all features was higher than using features from any single hierarchical level. Obviously, the importance of features for classification cannot be generalized merely based on their categories. To construct a more precise and concise classification model, we introduced the Maximum-Relevance and Minimum-Redundancy (MRMR) algorithm (see Supplementary Note 5, Supplementary Figs. 13 and 14 for details) from feature engineering, which leverages mutual information theory to evaluate the classification efficacy of phase features. The MRMR algorithm can balance redundancy (among feature variables) and relevance (between feature variables and target variables), making it more powerful than traditional feature processing techniques, which we have also verified (Supplementary Fig. 14). It scored all the features and ranked them from high to low (Fig. 4a).
a Importance scores of 81 features for classification given by the MRMR algorithm, sorted from high to low. b Incremental feature selection (IFS) curves for 9 machine learning algorithms, including LR, KNN, LDA, SVM, NN, DT, Naive Bayes, Subspace KNN and Bagged Trees, with the highest predictive accuracy of each model labeled accordingly. The gray shadow represents the MRMR scores of features. The blue curve illustrates the accuracy rates on the ten-fold cross-validation set, while the red curve indicates the accuracy rates on the test set. c Predictive performance metrics, including recall, precision, F1-score, accuracy, and ROC AUC, for the 9 best-trained machine learning models across 5 cancer cell types. Source data are provided as a Source Data file.
Finally, we combined MRMR with an incremental feature selection (IFS) strategy for the model training, adding features one by one based on their MRMR scores, and evaluating their impact on model performance, shown by the IFS curve (Fig. 4b). It can be observed that as the number of important features increased, the accuracy of the 9 machine learning models reached a plateau and no longer increased, and in some cases (such as KNN and Naive Bayes), it even decreased. Besides, more features led to longer training times and slower prediction speeds. After a comprehensive trade-off between accuracy and efficiency, the optimal feature subsets were determined for each model and used to train and test the models (see Supplementary Table 11 for details). Among them, both LDA and SVM (Fig. 4b) achieved 100% overall prediction accuracy for five types of cells with their optimal subsets. Considering LDA required fewer features, had a shorter training time, and faster prediction speeds (Supplementary Table 11), it was identified as the best classification model. Furthermore, the predictive performance for different cancer types was evaluated using recall rate, precision rate, F1 scores, accuracy, and the Receiver Operating Characteristic (ROC) Area Under the Curve (AUC) (Fig. 4c). We can see that overall, after adequate training, all models achieved a prediction accuracy above 90% for all cancer cell types, with the highest accuracy, F1 score, and AUC value for the breast cancer cell line MDA-MB-231. Detailed reports of the confusion matrices (Supplementary Fig. 15a–i), ROC curves (Supplementary Fig. 16a–i), and other metrics (Supplementary Table 11) are provided in the Supplementary Information, proving the effectiveness of our classification strategies in predicting heterogeneous cancer cell populations.
Breast cancer subtypes prediction based on QPM phase features and MRMR-IFS strategy
Due to the model’s notable predictive performance for breast cancer, we established a classification dataset using cell lines from the same organ source that exhibited closely related morphological features, including 1 normal breast cell line (n = 1000 cells), seven subtypes of breast cancer cell lines (n = 1000 cells for each), and a blank background (n = 3000 images). These cell lines were chosen as they cover all major breast cancer subtypes (Luminal A/Luminal B/Her2-enriched/TN). In clinical practice, distinguishing between these subtypes is crucial for guiding treatment plans and predicting prognoses.
From the QPM reconstruction results of the 8 cell types (Fig. 5a), the TN-type cells (including HCC1937 and MDA-MB-231) showed some feature similarity with normal breast cells (MCF-10A). The average sizes of T47D (Luminal A), BT474 (Luminal B), and SK-BR-3 (Her2-enriched) cell groups were similar and smaller than other groups. Violin plots were used to (Fig. 5b) visualize the distribution of some features, highlighting the differences between different breast cancer subtypes and normal breast cells. Figure 5c summarizes the feature mean value distributions of the 8 cell lines and also performs the results of variance analysis and multiple comparison tests. Compared with the feature datasets of different cancer cell lines, the breast cancer multi-subtype dataset was markedly more complicated, reflected in the higher similarity of a single feature between different subtypes (less significant difference in mean values). Statistical analysis indicated that only 1 of the 81 features was significantly different in all pairwise comparisons, revealing the higher difficulty of classifying multi-subtypes of breast cancer. Figure 5d shows the Pearson correlation matrix among features, where high correlations still tend to appear within the same feature categories and higher-order features. The more frequent presence of highly correlated features suggested the increased feature redundancy and, consequently, higher training difficulty for classification models.
a Single-cell QPM phase maps of 7 breast cancer-related cell lines, including 2 LA-type cell lines (MCF-7 and T47D), 2 LB-type cell lines (BT474 and MDA-MB-361), 1 Her2-enriched-type cell lines (SK-BR-3), 2 TN-type cell lines (HCC1937 and MDA-MB-231), and 1 normal breast cell line (MCF-10A). The color bar denotes the scale of normalized optical path difference (OPD) to reflect phase information (Scale bar: 10 μm). b)Violin plots of 6 QPM phase features (n = 1000 cells for each cancer type), including volume (Bulk), dry mass density (Bulk), kurtosis (First-order histogram), entropy (GLCM), gray asymmetry (GGCM), and run variance (GLRLM). The violins are drawn with the ends at the quartiles and the median as a horizontal line in the violin, where the curve of the violin extends to the minimum and maximum values in the data set. *** indicates p-value < 0.001, ** indicates 0.001 <p-value < 0.01, * indicates 0.01 <p-value < 0.05, ns indicates p-value > 0.05, based on one-way ANOVA followed by Tukey’s multiple comparisons test. In (c), the outer circular heatmap displays the mean value distribution of 81 features (arranged in a clockwise direction) across 8 distinct breast cancer-related cell types (ordered from the center out). The color bar denotes the scale of normalized mean values (n = 1000 cells for each cancer type). The inner binary heatmap shows the results of ordinary one-way ANOVA pairwise comparisons (Tukey’s test). Dark blue visualizes whether these differences are statistically significant. The significance level (α) is set to 0.05. d Pearson correlation matrix of 81 QPM phase features visualized as a heatmap (n = 1000 cells for each cancer type). Blue indicates positive correlation, while red indicates negative correlation. Different feature subsets are divided by yellow boxes. The color bar represents the scale of the Pearson correlation coefficient. Source data are provided as a Source Data file.
Consistent with our previous approach, we begin modeling and predicting breast cancer subtypes by using all features and by using different hierarchical subsets separately (see Supplementary Note 7 for details). We found that the accuracy of models trained on feature subsets still fell short of that achieved with the full set (Supplementary Fig. 17a–f and Supplementary Table 12 and 13). Therefore, for feature selection and model training, we again used the combination strategy of MRMR and IFS. Figure 6a illustrates the MRMR scores for each feature in descending order. Training results (Fig. 6b and Supplementary Figs. 18a–i and 19a–i) show that, while more features and longer training times were generally required (Supplementary Table 14), LDA still achieved 100% classification accuracy (Fig. 6b and Supplementary Fig. 18a). The performance of NN and SVM also remained at a high level. The impact of ensemble learning was particularly obvious, with the accuracy of Subspace KNN and Bagged Trees improving by 3.4% and 13.2%, respectively, over their non-ensemble equivalents. Evaluating the predictive performance of each model for different types of cells (Fig. 6c), the prediction accuracy for breast cancer subtypes was generally lower than that for different cancer cell types, which was consistent with expectations, given the closer similarities among cell groups. In conclusion, the strategy of training based on hierarchical QPM phase features in conjunction with MRMR and IFS showed universal applicability across various machine learning models, and these research results also further proved the system’s ability to perform fine classification tasks for multiple groups of cells and particles.
a Importance scores of 81 features for classification given by the MRMR algorithm, sorted from high to low. b Incremental feature selection (IFS) curves for 9 machine learning algorithms, including LR, KNN, LDA, SVM, NN, DT, Naive Bayes, Subspace KNN and Bagged Trees, with the highest predictive accuracy of each model labeled accordingly. The gray shadow represents the MRMR scores of features. The blue curve illustrates the accuracy rates on the ten-fold cross-validation set, while the red curve indicates the accuracy rates on the test set. c Predictive performance metrics, including recall, precision, F1-score, accuracy, and ROC AUC, for the 9 best-trained machine learning models across normal breast cell and breast cancer subtypes. Source data are provided as a Source Data file.
Identification and enumeration of breast cancer cells in human blood
Circulating tumor cells (CTCs), originating from primary tumors and circulating in the bloodstream, are instrumental for monitoring cancer progression and guiding therapeutic strategies, with breast cancer being the most extensively studied in this context67. In our pursuit to advance the practical applications of the SMHI platform, we utilized it to demonstrate label-free detection of simulated CTCs in human blood samples by mixing MDA-MB-231 cells with human blood, following previous reports48,68,69. We procured blood from three healthy volunteers and isolated red blood cells (RBCs) and peripheral blood mononuclear cells (PBMCs). We employed our SMHI platform to capture holograms and reconstruct QPM phase maps for 1000 instances each of RBCs, PBMCs, and MDA-MB-231 cells (Fig. 7a), enabling direct comparison of size and morphology differences among the cell types. Figure 7b presents violin plots illustrating the differentiated distribution of several features. Moreover, we performed t-SNE (t-distributed Stochastic Neighbor Embedding) dimensionality reduction70, transforming the 81 high-dimensional QPM phase features into a two-dimensional space, which illustrated a clear separation between the three cell populations (Fig. 7c). For the comprehensive QPM phase feature characterization and analysis, Fig. 7d concludes the mean distribution of 81 features across the three cell populations, accompanied by variance analysis and multiple comparison tests like before. The multiple comparison tests revealed significant differences in most features, which surpassed those between multiple cancer cells (Fig. 3c) or breast cancer subtypes (Fig. 5c). Figure 7e displays a Pearson correlation matrix among features, revealing a high degree of correlation that indicates feature redundancy due to the consistent information shared among the features. This redundancy highlights the necessity of feature engineering in our classification task.
a Single-cell QPM phase maps of red blood cells (RBCs), peripheral blood mononuclear cells (PBMCs), and MDA-MB-231 cells. Color bar: normalized optical path difference (OPD). Scale bar: 10 μm. b Violin plots of 6 QPM phase features (n = 1000 cells for each type), including volume (Bulk), dry mass density (Bulk), kurtosis (First-order histogram), entropy (GLCM), gray asymmetry (GGCM), and run variance (GLRLM). The violins are drawn with the ends at the quartiles and the median as a horizontal line in the violin, where the curve of the violin extends to the minimum and maximum values in the data set. The p-value < 0.05 indicates a significant difference decided by one-way ANOVA followed by Tukey’s multiple comparisons test. c t-SNE plot of RBCs, PBMCs and MDA-MB-231 cells (n = 1000 cells for each type). In (d), the outer circular heatmap displays the mean value distribution of 81 features across RBCs, PBMCs and MDA-MB-231 cells (n = 1000 cells for each type). Color bar: normalized mean values. The inner binary heatmap shows the results of ordinary one-way ANOVA pairwise comparisons (Tukey’s test). Dark blue visualizes whether these differences are statistically significant. The significance level (α) is set to 0.05. e The Pearson correlation matrix of 81 QPM phase features visualized as a heatmap. Blue indicates positive correlation, while red indicates negative correlation. Different feature subsets are divided by yellow boxes. Color bar: Pearson correlation coefficient. f Incremental feature selection (IFS) curves for the LDA model trained for classifying PBMCs and MDA-MB-231 cells, with the highest predictive accuracy labeled accordingly. A total of PBMCs (n = 5137 cells) and MDA-MB-231 cells (n = 3040 cells) data are used for training. The gray shadow represents the MRMR scores of features. The blue curve illustrates the accuracy rates on the ten-fold cross-validation set. g–j illustrate the predicted total cell numbers, PBMC numbers, and MDA-MB-231 cell numbers in PBMC samples spiked with MDA-MB-231 cells by the LDA model, with the ratios of MDA-MB-231 to PBMC are 1:1 (g), 1:10 (h), 1:100 (i), and 1:1000 (j). Data are presented as means ± SD. The numbers on top of each bar represent the means from 3 independent replicates under each ratio condition. Sample sizes per condition (g–j): 1:1 (n = 1000/1000/1003 cells), 1:10 (n = 1001/1010/1002 cells), 1:100 (n = 1000/1003/1003 cells), 1:1000 (n = 1012/1026/1004 cells). Source data are provided as a Source Data file.
Aligning with the practical CTCs detection protocols, enrichment is a necessary step due to CTCs’ rarity. According to the previous refs. 68,69, RBCs removal by centrifugation or lysis is a common step in CTC isolation to minimize interference66. Therefore, concentrating our study on PBMCs rather than whole blood aligns with real-world detection protocols and minimizes computational overhead in characterizing easily distinguishable RBCs. We focused on differentiating PBMCs and MDA-MB-231 cells. We processed a dataset comprising 5137 PBMCs and 3040 MDA-MB-231 cells, and constructed an LDA classification model using the MRMR-IFS strategy (Fig. 7f), achieving the highest accuracy rate of 99.9% when using only the top 9 features. Notably, we meticulously considered and analyzed the impact of batch effects, performing cross-validation51 as detailed in Supplementary Note 8. The results (see Supplementary Figs. 20–23 and Supplementary Table 15) demonstrated high cross-validation accuracy, proving the generalizability and broad applicability of our LDA model post-training with the MRMR-IFS strategy. Finally, we prepared PBMC and MDA-MB-231 cell suspensions at a concentration of 1 × 106 cells/mL, to mimic blood samples containing CTCs that have been processed to remove RBCs. We created four gradient concentration samples by mixing MDA-MB-231 cells with PBMCs at ratios of 1:1, 1:10, 1:100, and 1:1000. The lowest spike ratio of 1:1000 was lower than the reported CTC purity ratios after enrichment, which range from 3.64% to over 90%71. Figure 7g–j illustrates the high consistency between the classification ratios predicted by the LDA model and the actual mixing ratios of the samples, providing preliminary evidence of our SMHI platform’s potential to identify and count CTCs in simulated blood samples.
Discussion
In summary, our SMHI platform emphasizes our efforts in integrated innovation and precise system control. We have successfully combined a spatial hydrodynamic focusing microfluidic technique with DHM to accurately capture signals from in-flow single cells and particles. By adjusting the flow rate ratio of sheath flow to sample flow in the microfluidics module, cells or particles of different properties and sizes can be precisely focused in both vertical and horizontal directions within the detection area, achieving high-performance, label-free single-cell analysis. Both theoretical simulations and experimental outcomes have comprehensively validated the flexibility and stability of the spatial hydrodynamic focusing. In addition, we have confirmed the spatiotemporal stability of our cage-structured DHM, with its spatial noise deviation (0.035 rad) significantly lower than that of conventional Mach-Zehnder DHM configurations (0.02 rad), and temporal noise deviation (0.007 rad) virtually negligible. Our SMHI platform can now achieve precise QPM reconstruction without the need for digital refocusing algorithms, completing a single-cell hologram reconstruction in only 0.34 s which accounts for only 4.41% of the typical process (~ 7.71 s). Utilizing polystyrene microspheres as cellular analogs, we assessed the performance of our platform. The results were highly accurate in capturing and reconstructing amplitude and phase information, with a low CV of 4% demonstrating measurement consistency, which is attributed to the stability and flexibility of our system. We then carried out the label-free detection and classification of multi-cancer and heterogeneous cell populations. We established a multi-tiered, high-dimensional QPM phase feature knowledge base to correlate cellular species and subtypes with specific morphological and textural phenotypes (a total of 81), thereby enhancing the interpretability of classification outcomes. Unlike the conventional approach of using all available features, as seen in most studies, we conducted a comprehensive feature analysis, evaluated the significance of each feature for classification tasks and the redundancy between features. We integrated incremental learning and ensemble learning strategies into training machine learning models, which achieved > 99.9% prediction accuracy for 5 high-incidence cancer types and 7 breast cancer subtypes, demonstrating its effectiveness in analyzing highly heterogeneous cell populations. Finally, we classified and modeled the blood cells to simulate the detection of rare cells in human blood samples. By mixing PBMCs and tumor cells in different proportions, our system was able to effectively distinguish and count tumor cells in the samples, demonstrating its generalization ability and significant potential for practical application. This proof-of-concept study marks the first step. Further assessment (such as ROC analysis) will be important in future work to fully uncover the system’s advanced performance. Overall, compared to other state-of-art QPM-IFCs (see Supplementary Note 9 and Supplementary Table 16 for details), our SMHI platform offers advantages such as better system stability, easier setup, lower computational costs, high predictive accuracy, and robust generalization capability.
Here, we have only conducted a proof-of-concept study, and there are still many areas where the performance of our platform needs improvement. First, the detection throughput is currently limited. Referring to the measurement method72, we estimate that our microfluidics module alone has achieved a peak stable throughput of ~ 1600 particles/sec (at a sample flow rate of 10 μL/min and a particle concentration of ~ 1 × 107 particles/mL), which is comparable to the gold-standard commercial IFC systems at 2000 particles/sec72. However, the CMOS sensor’s modest frame rate of 30 frames per second (fps) restricts the maximum recording speed for reliable cell and particle images, necessitating operation at reduced sample concentrations (approximately 1 × 106 cells/mL) and flow rates (1 μL/min). We will upgrade to a high-speed camera with frame rates ranging from thousands to millions, significantly boosting the platform’s detection speed without compromising accuracy or robustness. Second, while our current feature engineering and machine learning strategy has already demonstrated some advantages and intelligent elements, including automating feature selection, enhancing prediction speed and generalization capabilities, as we venture into more complex scenarios, it may be useful to integrate advanced deep learning (DL)/artificial intelligence (AI) models to further improve classification accuracy55,73,74, enhance robustness and generalization75,76, and enable more detailed observation77. In the future, real-time detection on the platform may also be enhanced by accelerating phase reconstruction and feature extraction through the use of Graphics Processing Units (GPUs), Field-Programmable Gate Arrays (FPGAs), or other similar technologies78,79.
Looking ahead, our primary focus will be to continue enhancing the performance of our platform, positioning it at the forefront of label-free single-cell analysis technologies. Additionally, we aim to expand its applicability to a broader range of clinical and biological applications. This includes extending its use to clinical biopsy samples and establishing various patient-derived disease models, transitioning from multi-cancer to pan-cancer analysis, and enhancing its capabilities for analyzing not only human or mammalian cells but also plant, microbial, and algal cells. Such expansions will not only validate the platform’s versatility but also pave the way for its integration into real-world clinical settings. These advancements will facilitate more detailed and boarder researches, aiding in uncovering underlying mechanisms in biology, medicine, chemistry, and environmental science, thereby expanding the technological applications of our approach.
Methods
Ethical statement
The research involving human participants was reviewed by the authorized Institutional Review Board (IRB) of the Peking University Shenzhen Hospital and Shenzhen International Graduate School (Shenzhen, China) with approval IRB No. 2021019. Human whole blood samples were collected from three healthy adult female donors with full informed consent.
Microfluidic chip fabrication and manipulation
We designed the chip’s channel pattern with AutoCAD 2024, which was then printed onto a mask. The chip fabrication proceeded according to the standard soft lithography protocol. Specifically, SU8-2050 photoresist (Microchem) was spin-coated onto a clean, dry silicon wafer at 1000 × g for 60 s. The wafer was then pre-baked at 65 °C for 3 min, followed by incubation at 95 °C for 6 min. The mask pattern was photo-lithographically transferred to the photoresist layer by UV exposures for 3 times, each lasting 4 s. After being baked once more at 95 °C for 10 min, the wafer was developed in SU8 developer for 5 min, subsequently rinsed and dried to finalize the SU8 mold. Next, a mixture of polydimethylsiloxane (PDMS) prepolymer and curing agent (Dow Corning), in a 10:1 ratio, was poured onto the mold and cured at 65 °C for 4 h. The cured PDMS was then demolded, and holes were punched as inlets and outlets. Finally, combine the PDMS layer with a standard glass slide using oxygen plasma treatment, followed by baking them at 65 °C for 20 min to strengthen the adhesion. The completed chip featured microchannels with dimensions of 100 μm in width and 35 μm in height.
We simulated the spatial hydrodynamic focusing process in the single-phase flow module of the computational fluid dynamics (CFD) software COMSOL Multiphysics 6.1 (see Supplementary Note 1 for details). The actual focusing effect was experimentally assessed using the fluorescent dye Rhodamine B (Aladdin) dissolved in deionized water as the sample flow, with colorless deionized water serving as the sheath flow. These flows were separately drawn into 1 mL medical syringes. Silicone tubing (0.4 mm inner diameter and 0.9 mm outer diameter) was used to connect the syringe with a 23 G steel needle (0.3 mm inner diameter and 0.6 mm outer diameter), which was inserted directly into the sample inlet port (0.6 mm aperture) of the chip. We used three syringe pumps (Longer Pump, LSP01-1A) to control syringes containing vertical sheath flows, horizontal sheath flows and sample flow, respectively. Each pump has 2 parallel channels. Finally, the hydrodynamic focusing phenomenon was observed under the inverted fluorescence microscope (Olympus), and images were captured by the supporting software OlyVIA 4.1. For each fluorescent sample flow focusing experiment under different Rv and Rh conditions, the sample flow was loaded and imaged by 5 independent imaging sessions.
DHM imaging system
The cage-type DHM prototype was built based on an off-axis Mach-Zehnder structure (see Supplementary Note 2 for details). All cage system accessories were provided by RayCage (Zhenjiang) Photoelectric Technology Co., Ltd. A single-frequency solid-state laser (633 nm, LSR-PS-II, Lasever) served as the light source. The laser beam was successively collimated by reflecting mirrors, filtered through a spatial light filter, and directed through a collimating lens to ultimately form a uniform parallel beam. A polarizing beam splitter then diverted the beam into an object beam and a reference beam. The object and the reference beam path were equipped with identical microscope objectives featuring a 40 × magnification and a numerical aperture (NA) of 0.75 (UPlanFL-N, Olympus) to maintain interferometric balance. In addition, a neutral density filter was added to the reference path to correct the sample-induced light intensity attenuation in the object path. Finally, the object beam, including the sample’s information and the reference beam, interfered on the imaging plane, producing a hologram which was captured by an sCMOS camera (5120 pixels × 5120 pixels, 2.5 µm, pco.panda 26, Excelitas Technologies).
Phase reconstruction process
The phase reconstruction from holograms recorded by DHM was realized in MATLAB R2023b (MathWorks) through custom coding. The complete process is detailed in Supplementary Note 3. In short, the hologram was first recorded. Then, the fast Fourier transform (FFT) was applied to obtain spectra (including -1st/0/+ 1st spectrum), followed by spatial filtering to extract the object’s spectrum (+ 1st spectrum), and numerical reconstruction to recover the amplitude and the wrapped phase of the object. The least squares unwrapping algorithm80 was implemented to unwrap the phase. Finally, the distortion-free phase map \(\varphi \left(x,y\right)\) was restored using the quadratic surface fitting and the double-exposure method81,82, which involves background phase subtraction. Phase map further yielded the optical path delay (OPD) map \({OPD}(x,y)\), which is the integral of refractive index (RI) values over object thickness \(d(x,y)\), defined as follows:
λ is the illumination wavelength, \({n}_{1}\) is the RI of the object, \({n}_{0}\) is the RI of the surrounding medium.
Platform validation using polystyrene microspheres
Standard 10 μm and 20 μm polystyrene microspheres were purchased (Yuan Biotech) to serve as samples, which were suspended in 1 × PBS to a concentration of approximately 1 × 106 particles/mL, with the addition of 0.1% Tween 20 (Aladdin) to prevent particle aggregation. The sheath flows were PBS with 0.1% Tween 20 for all focusing experiments. The microfluidic chip was mounted on the DHM prototype. The sample flow and sheath flows were respectively infused into the microchannel at different flow rates. The sample flow rate was fixed at 1 μL/min. For planar focusing, Rv of 10 μm and 20 μm microspheres were 0.75 and 0.125. For spatial focusing, Rv was the same as for planar focusing, and Rh of 10 μm and 20 μm microspheres were 3 and 1.25. 50 holograms of polystyrene microspheres were captured for each focusing status by employing the snapshot shutter mode in the sCMOS camera, with the image size cropped to 496 pixels × 496 pixels, the exposure time of 10 ms, and the frame rate of 30 fps.
The amplitude and phase maps of microspheres were reconstructed. From the amplitude map, we measured the length and width (in pixels) of a single microsphere, calculated their average, which was then multiplied by the pixel size (2.5 µm) and divided by the magnification factor (40) to obtain the actual diameter of the microsphere. From the phase map, we took the difference between the highest and lowest phase values within the microsphere. Given the RI of the microsphere (1.59) and the medium (1.33), the actual height of the microsphere was calculated by Eq. (1). The coefficient of variation (CV) was determined from the amplitude diameters and phase heights of 50 microspheres, calculated as the ratio of the standard deviation to the mean. According to the manufacturer’s specifications, CV < 10% indicates the precision and consistency of the measurements. The statistical analyses and plots (Fig. 2h–k) of the reconstructed amplitude and phase for 50 microspheres under different focusing conditions were done with GraphPad Prism 9.5.0, with p-values determined by one-way ANOVA followed by Tukey’s multiple comparisons test. The level of significance was set at 0.05.
Cell lines culture and preparation of single-cell suspensions
Cell lines were purchased from the Shanghai Cell Bank of the Chinese Academy of Sciences, including A549 (SCSP-503), HCT116 (SCSP-5076), HeLa (SCSP-504), THP-1 (SCSP-567), MCF-10A (SCSP-575), T47D (SCSP-564), MCF-7 (SCSP-531), BT474 (TCHu143), MDA-MB-361 (SCSP-5052), SK-BR-3 (SCSP-5243), HCC1937 (TCHu148), and MDA-MB-231 (SCSP-5043). HCT116 was cultured with McCoy’s 5 A medium (Gibco). THP-1, MCF-10A, HCC1937, and BT474 were cultured with Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco). All other cell lines were cultured with Dulbecco’s modified Eagle’s medium (DMEM, Gibco). Each medium was supplemented with 10% fetal bovine serum (FBS, Gibco) and 1% penicillin-streptomycin (Pen-Strep, Gibco). For BT474 culture, an additional 10 µg/mL insulin (Gibco) and 2 mM L-glutamine (Gibco) were added. For THP-1 culture, 0.05 mM β-mercaptoethanol were added. Cells were incubated in 10 cm culture dishes at 37 °C in 5% CO2 atmosphere at least 1 week in advance of imaging. The adherent cells were then washed 3 times using PBS. After that, 1 mL of trypsin (Gibco) was added to digest cells for 3–5 min until most cells were detached from the dish surface, then the digestion was halted by adding 2 mL of serum-containing medium. The liquid was collected and centrifuged at 500 × g for 5 min, after which the supernatant was discarded. The cell pellet was then resuspended in PBS for imaging. For the sole suspension cell line, THP-1, the process was simplified to merely centrifuging to remove the old medium, followed by the resuspension of the cell pellet in PBS.
Isolation of RBCs and PBMCs from human blood samples
Human blood samples were collected from healthy adult female volunteers after obtaining informed consent and ethical approval. The samples were processed within 2 h to separate RBCs and PBMCs using the density gradient centrifugation method. Specifically, 5–10 mL of whole blood was mixed with an equal volume of 1 × PBS. An equal volume of Ficoll-Paque (GE Healthcare) density gradient medium was added to a centrifuge tube, and the diluted blood was carefully layered onto the surface of the separation solution. The mixture was then centrifuged at 500-1000 × g for 20–30 min at room temperature (18–25 °C). After centrifugation, distinct layers formed in the tube. The white membrane layer was carefully collected into a clean centrifuge tube, and the cells were washed with 1 × PBS. The mixture was centrifuged at 250 × g for 10 min, the supernatant was discarded, and the washing step was repeated three times to isolate PBMCs. The separation solution layer was removed, and the bottom RBC layer was collected into a new tube. The washing process was repeated three times in the same manner. Finally, the cells were resuspended in 1 × PBS to create a single-cell suspension for further use.
Single cell hologram acquisition
1 × PBS was used as sheath flows. Single-cell suspensions were prepared for various cell types to serve as sample flows, which were then filtered using a 40 µm cell strainer (BKMAMLAB) to remove possible cell clumps. Cell concentration and average size were measured using a commercial cell counter (iCytal, JIMBIO), and the suspensions were adjusted to a final concentration of 1 × 106 cells/mL with PBS solution. The average cell size measurements were referenced to determine the appropriate values of Rv and Rh for each cell type, respectively. The sample flow rate was set at 1 µL/min. Sheath and sample flows were introduced into the microfluidic chip channel mounted on the DHM at the determined flow rates for hologram capture of focused cells. The camera capture mode and parameter settings were consistent with those used in the microsphere validation experiments. For each cell type, a minimum of 1000 holograms were captured for subsequent analysis.
Single-cell suspensions of MDA-MB-231 cells and PBMCs, each adjusted to a final concentration of 1 × 106 cells/mL in PBS, were mixed in volumetric ratios of 1:1, 1:10, 1:100, and 1:1000 to prepare the simulated CTCs detection samples. These samples were used as the sample flows for further detection processes. The average flow rate ratio for the two cell types was taken as the initial flow rate ratio for the mixed sample, and the ratio was fine-tuned based on observed results until a satisfactory focusing effect was achieved.
Hologram processing and cell features extraction
A single-cell hologram (496 pixels × 496 pixels) reconstruction was completed within about 0.34 s using lab-written MATLAB codes on our current laptop equipped with a 2.8 GHz Intel Core i7 CPU and 16 GB of RAM. The phase map then underwent essential preprocessing steps, including filling in minor holes and fractures, smoothing boundaries, and converting the images into binary format. The largest connected region was preserved as a mask to delineate the region of interest (ROI) for the cell. Finally, 81 cell features were extracted (see Supplementary Note 4 for details), including 14 bulk features, 17 first-order histogram features and 50 higher-order texture features (20 GLCM features, 15 GGCM features and 15 GLRLM features). The total duration for preprocessing and feature extraction of a single cell was approximately 0.34 s. Finally, at least 1000 single-cell holograms for each cell type were processed and used to build the dataset.
Cell features analysis and visualization
A one-way analysis of variance (ANOVA) and multiple comparisons (Tukey’s test) were conducted to compare the mean value distributions of one feature among different cells (see Supplementary Note 5 for details). The significance level α in this method was set as 0.05. Violin plots were used to display the analysis results of several features, while a circular heatmap summarized the analysis results of all 81 features. Beyond univariate analysis, the Pearson correlation coefficient was computed to quantify the linear relationship between pairs of features. The calculated correlation matrix was visualized as a heatmap. All visualizations, including the violin plots and heatmaps, were generated using GraphPad Prism 9.5.0 and Chiplot (https://www.chiplot.online/). In addition, t-SNE (t-distributed Stochastic Neighbor Embedding) method was employed to create a two-dimensional plot that illustrates the unsupervised clustering of three cell types: RBCs, PBMCs, and MDA-MB-231 cells. The t-SNE analysis was conducted using custom code in MATLAB R2023b.
Feature engineering and machine learning
The MATLAB machine learning packages and the Classification Learner toolbox are utilized to facilitate training and prediction tasks. The Maximun-Relevance and Minimun-Redundancy (MRMR) algorithm (see Supplementary Note 5 for details) were used to score and rank 81 features. Using an incremental feature selection (IFS) strategy, features were added sequentially for model training based on their scores. IFS curve based on prediction accuracy were used to identify the best feature combination.
Classical machine learning models were trained, including Logistic Regression (LR), K-Nearest Neighbors (KNN), Linear Discriminant Analysis (LDA), Support Vector Machine (SVM), Neural Network (NN), Decision Trees (DT) and Naive Bayes. Besides, two ensemble models, Subspace KNN and Bagged Trees, were also included to enrich the analytical framework. The dataset was first partitioned into a training set and a test set in a 9:1 ratio. A 10-fold cross-validation approach was then applied on the training set to ensure the model’s generalizability.
The performance of different models was assessed using recall, precision, F1-score, accuracy and area under the curve (AUC) for each cell type. Confusion matrices and Receiver operating characteristic (ROC) curves were plotted for visually demonstration. Model training-related information was also exported from MATLAB, including training duration, prediction speed and so on (see Supplementary Note 6 and 7 for details).
Statistics and reproducibility
Sample sizes and statistical tests have been reported in each figure or figure legend, as well as the number of times that the measurements were repeated for representative results. Data were presented as means ± standard deviation (SD), shown by error bars. Comparisons of more than three groups were performed by one-way ANOVA and Tukey’s multiple comparisons test with GraphPad Prism 9.5.0. p-value < 0.05 was considered as statistically significant. Due to word count limitations, detailed ANOVA statistics (F-values and degrees of freedom) are provided in the Source Data file rather than in the figure legend.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated and analyzed in this study have been deposited in the Zenodo database (https://doi.org/10.5281/zenodo.15455289). Further information regarding to the findings is available from the corresponding authors upon request. Requests will be fulfilled within two weeks. Data underlying the figures are provided in the Supplementary Information or Source Data file. Source data are provided in this paper.
Code availability
Custom code was used in hologram phase reconstruction and feature extraction based on the MATLAB R2023b, which are available at https://github.com/SMHSplatform/SMHI.
References
Gohil, S. H., Iorgulescu, J. B., Braun, D. A., Keskin, D. B. & Livak, K. J. Applying high-dimensional single-cell technologies to the analysis of cancer immunotherapy. Nat. Rev. Clin. Oncol. 18, 244–256 (2021).
Van de Sande, B. et al. Applications of single-cell RNA sequencing in drug discovery and development. Nat. Rev. Drug Discov. 22, 496–520 (2023).
Vandereyken, K., Sifrim, A., Thienpont, B. & Voet, T. Methods and applications for single-cell and spatial multi-omics. Nat. Rev. Genet. 24, 494–515 (2023).
Han, Y. Y., Gu, Y., Zhang, A. C. & Lo, Y. H. Imaging technologies for flow cytometry. Lab. Chip 16, 4639–4647 (2016).
Lei, C. et al. High-throughput imaging flow cytometry by optofluidic time-stretch microscopy. Nat. Protoc. 13, 1603–1631 (2018).
Basiji, D. A., Ortyn, W. E., Liang, L., Venkatachalam, V. & Morrissey, P. Cellular image analysis and imaging by flow cytometry. Clin. Lab. Med. 27, 653–670 (2007).
Görgens, A. et al. Optimisation of imaging flow cytometry for the analysis of single extracellular vesicles by using fluorescence-tagged vesicles as biological reference material. J. Extracell. Vesicles 8, 1587567 (2019).
Kuett, L. et al. Three-dimensional imaging mass cytometry for highly multiplexed molecular and cellular mapping of tissues and the tumor microenvironment. Nat. Cancer 3, 122–133 (2022).
McClelland, R. D., Culp, T. N. & Marchant, D. J. Imaging flow cytometry and confocal immunofluorescence microscopy of virus-host cell interactions. Front. Cell. Infect. Microbiol. 11, 749039 (2021).
Power, A. L. et al. The application of imaging flow cytometry for characterisation and quantification of bacterial phenotypes. Front. Cell. Infect. Microbiol. 11, 716592 (2021).
Gautam, N., Sankaran, S., Yason, J. A., Tan, K. S. W. & Gascoigne, N. R. J. A high content imaging flow cytometry approach to study mitochondria in T cells: MitoTracker Green FM dye concentration optimization. Methods 134-135, 11–19 (2018).
Magidson, V. & Khodjakov, A. Circumventing photodamage in live-cell microscopy. Methods Cell Biol. 114, 545–560 (2013).
Park, Y., Depeursinge, C. & Popescu, G. Quantitative phase imaging in biomedicine. Nat. Photonics 12, 578–589 (2018).
Drexler, W. & Fujimoto J. G. Optical Coherence Tomography: Technology and Applications (Springer, 2008).
Leitgeb, R., Hitzenberger, C. K. & Fercher, A. F. Performance of fourier domain vs. time domain optical coherence tomography. Opt. Express 11, 889–894 (2003).
Wang, L. V. & Hu, S. Photoacoustic tomography: in vivo imaging from organelles to organs. Science 335, 1458–1462 (2012).
Freudiger, C. W. et al. Label-Free Biomedical Imaging with High Sensitivity by Stimulated Raman Scattering Microscopy. Science 322, 1857–1861 (2008).
Oldenbourg, R. Polarized light microscopy of spindles. Methods Cell. Biol. 61, 175–208 (1998).
Zhu, Y., Li, Y., Huang, J. & Lam, E. Y. Smart polarization and spectroscopic holography for real-time microplastics identification. Commun. Eng. 3, 32 (2024).
Shaked, N. T., Boppart, S. A., Wang, L. V. & Popp, J. Label-free biomedical optical imaging. Nat. Photonics 17, 1031–1041 (2023).
Rappaz, B., Charrière, F., Depeursinge, C., Magistretti, P. J. & Marquet, P. Simultaneous cell morphometry and refractive index measurement with dual-wavelength digital holographic microscopy and dye-enhanced dispersion of perfusion medium. Opt. Lett. 33, 744–746 (2008).
Rappaz, B. et al. Measurement of the integral refractive index and dynamic cell morphometry of living cells with digital holographic microscopy. Opt. Express 13, 9361–9373 (2005).
Ahmad, A. et al. Quantitative phase microscopy of red blood cells during planar trapping and propulsion. Lab. Chip 18, 3025–3036 (2018).
Shribak, M., Larkin, K. G. & Biggs, D. Mapping optical path length and image enhancement using quantitative orientation-independent differential interference contrast microscopy. J. Biomed. Opt. 22, 16006 (2017).
Shan, M., Kandel, M. E. & Popescu, G. Refractive index variance of cells and tissues measured by quantitative phase imaging. Opt. Express 25, 1573–1581 (2017).
Khan, S., Jesacher, A., Nussbaumer, W., Bernet, S. & Ritsch-Marte, M. Quantitative analysis of shape and volume changes in activated thrombocytes in real time by single-shot spatial light modulator-based differential interference contrast imaging. J. Biophotonics 4, 600–609 (2011).
Lee, S. et al. Refractive index tomograms and dynamic membrane fluctuations of red blood cells from patients with diabetes mellitus. Sci. Rep. 7, 1039 (2017).
Siu, D. M. D. et al. Optofluidic imaging meets deep learning: from merging to emerging. Lab. Chip 23, 1011–1033 (2023).
Min, J. et al. Quantitative phase imaging of cells in a flow cytometry arrangement utilizing Michelson interferometer-based off-axis digital holographic microscopy. J. Biophotonics 12, e201900085 (2019).
Zhang, T. et al. Focusing of sub-micrometer particles in microfluidic devices. Lab. Chip 20, 35–53 (2020).
Kralj, J. G., Lis, M. T. W., Schmidt, M. A. & Jensen, K. F. Continuous dielectrophoretic size-based particle sorting. Anal. Chem. 78, 5019–5025 (2006).
Afshar, R., Moser, Y., Lehnert, T. & Gijs, M. A. Three-dimensional magnetic focusing of superparamagnetic beads for on-chip agglutination assays. Anal. Chem. 83, 1022–1029 (2011).
Shi, J., Huang, H., Stratton, Z., Huang, Y. & Huang, T. J. Continuous particle separation in a microfluidic channelvia standing surface acoustic waves (SSAW). Lab. Chip 9, 3354–3359 (2009).
Cruz, J. & Hjort, K. High-resolution particle separation by inertial focusing in high aspect ratio curved microfluidics. Sci. Rep. 11, 13959 (2021).
Li, M., van Zee, M., Goda, K. & Di Carlo, D. Size-based sorting of hydrogel droplets using inertial microfluidics. Lab. Chip 18, 2575–2582 (2018).
Zhao, L. et al. Flow-rate and particle-size insensitive inertial focusing in dimension-confined ultra-low aspect ratio spiral microchannel. Sens. Actuat. B Chem. 369, 132284 (2022).
Lee, K. C. et al. Dispersion-free inertial focusing (DIF) for high-yield polydisperse micro-particle filtration and analysis. Lab. Chip 24, 4182–4197 (2024).
Shen, S. et al. Spiral large-dimension microfluidic channel for flow-rate- and particle-size-insensitive focusing by the stabilization and acceleration of secondary flow. Anal. Chem. 96, 1750–1758 (2024).
Sakuma, S., Kasai, Y., Hayakawa, T. & Arai, F. On-chip cell sorting by high-speed local-flow control using dual membrane pumps. Lab. Chip 17, 2760–2767 (2017).
Zhao, L. et al. A plug-and-play 3D hydrodynamic focusing Raman platform for label-free and dynamic single microparticle detection. Sens. Actuat. B Chem. 369, 132273 (2022).
Pirone, D. et al. Identification of drug-resistant cancer cells in flow cytometry combining 3D holographic tomography with machine learning. Sens. Actuat. B Chem. 375, 132963 (2023).
Ugele, M. et al. Label-free high-throughput leukemia detection by holographic microscopy. Adv. Sci. 5, 1800761 (2018).
Merola, F. et al. Tomographic flow cytometry by digital holography. Light Sci. Appl. 6, e16241 (2017).
Vercruysse, D. et al. Three-part differential of unlabeled leukocytes with a compact lens-free imaging flow cytometer. Lab. Chip 15, 1123–1132 (2015).
Hirotsu, A. et al. Artificial intelligence-based classification of peripheral blood nucleated cells using label-free imaging flow cytometry. Lab. Chip 22, 3464–3474 (2022).
Yamada, H. et al. Label-free imaging flow cytometer for analyzing large cell populations by line-field quantitative phase microscopy with digital refocusing. Biomed. Opt. Express 11, 2213–2223 (2020).
Pirone, D. et al. Speeding up reconstruction of 3D tomograms in holographic flow cytometry via deep learning. Lab. Chip 22, 793–804 (2022).
Singh, D. K., Ahrens, C. C., Li, W. & Vanapalli, S. A. Label-free, high-throughput holographic screening and enumeration of tumor cells in blood. Lab. Chip 17, 2920–2932 (2017).
Li, Y. et al. Accurate label-free 3-part leukocyte recognition with single cell lens-free imaging flow cytometry. Comput. Biol. Med. 96, 147–156 (2018).
Lee, K. C. M. et al. Quantitative phase imaging flow cytometry for ultra-large-scale single-cell biophysical phenotyping. Cytom. A 95, 510–520 (2019).
Shu, X. et al. Artificial-intelligence-enabled reagent-free imaging hematology analyzer. Adv. Intell. Syst. 3, 2000277 (2021).
Nissim, N., Dudaie, M., Barnea, I. & Shaked, N. T. Real-time stain-free classification of cancer cells and blood cells using interferometric phase microscopy and machine learning. Cytom. A 99, 511–523 (2021).
Paidi, S. K. et al. Raman and quantitative phase imaging allow morpho-molecular recognition of malignancy and stages of B-cell acute lymphoblastic leukemia. Biosens. Bioelectron. 190, 113403 (2021).
Yip, G. G. K. et al. Multimodal FACED imaging for large-scale single-cell morphological profiling. Apl. Photonics 6, 070801 (2021).
Butola, A. et al. High spatially sensitive quantitative phase imaging assisted with deep neural network for classification of human spermatozoa under stressed condition. Sci. Rep. 10, 13118 (2020).
Kim, G. et al. Rapid species identification of pathogenic bacteria from a minute quantity exploiting three-dimensional quantitative phase imaging and artificial neural network. Light Sci. Appl. 11, 190 (2022).
Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 1, 206–215 (2019).
Park, J. et al. Artificial intelligence-enabled quantitative phase imaging methods for life sciences. Nat. Methods 20, 1645–1660 (2023).
Siu, D. M. D. et al. Deep-learning-assisted biophysical imaging cytometry at massive throughput delineates cell population heterogeneity. Lab. Chip 20, 3696–3708 (2020).
Cai, J., Luo, J., Wang, S. & Yang, S. Feature selection in machine learning: A new perspective. Neurocomputing 300, 70–79 (2018).
Poggio, T., Mhaskar, H., Rosasco, L., Miranda, B. & Liao, Q. Why and when can deep-but not shallow-networks avoid the curse of dimensionality: a review. Int. J. Autom. Comput. 14, 503–519 (2017).
Peng, H., Long, F. & Ding, C. Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans. Pattern Anal. Mach. Intell. 27, 1226–1238 (2005).
Hsu, W. C., Su, J. W., Tseng, T. Y. & Sung, K. B. Tomographic diffractive microscopy of living cells based on a common-path configuration. Opt. Lett. 39, 2210–2213 (2014).
Mehta, D. S., Butola, A. & Singh, V. Quantitative Phase Microscopy and Tomography: Techniques Using Partially Spatially Coherent Monochromatic Light.(IOP Publishing, Bristol, 2022).
Singh, V., Tayal, S. & Mehta, D. S. Highly stable wide-field common path digital holographic microscope based on a Fresnel biprism interferometer. OSA Contin. 1, 48–55 (2018).
Wang, C. et al. Sheathless microflow cytometer utilizing two bulk standing acoustic waves. Cytom. A 99, 987–998 (2021).
Ring, A., Nguyen-Sträuli, B. D., Wicki, A. & Aceto, N. Biology, vulnerabilities and clinical applications of circulating tumour cells. Nat. Rev. Cancer 23, 95–111 (2023).
Suzuki, Y. et al. Label-free chemical imaging flow cytometry by high-speed multicolor stimulated Raman scattering. Proc. Natl. Acad. Sci. USA 116, 15842–15848 (2019).
Zhu, J. et al. Microfluidic impedance cytometry enabled one-step sample preparation for efficient single-cell mass spectrometry. Small 20, 2310700 (2024).
Maaten, L. V. D. & Hinton, G. E. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Shen, Z., Wu, A. & Chen, X. Current detection technologies for circulating tumor cells. Chem. Soc. Rev. 46, 2038–2056 (2017).
Hua, X. et al. Light-field flow cytometry for high-resolution, volumetric and multiparametric 3D single-cell analysis. Nat. Commun. 15, 1975 (2024).
O’Connor, T., Anand, A., Andemariam, B. & Javidi, B. Deep learning-based cell identification and disease diagnosis using spatio-temporal cellular dynamics in compact digital holographic microscopy. Biomed. Opt. Express 11, 4491–4508 (2020).
Zhang, J. K., He, Y., Sobh, N. & Popescu, G. Label-free colorectal cancer screening using deep learning and spatial light interference microscopy (SLIM). APL Photonics 5, 040805 (2020).
Lo, M. C. K. et al. Information-distilled generative label-free morphological profiling encodes cellular heterogeneity. Adv. Sci. 11, 2307591 (2024).
Sypetkowski, M. et al. RxRx1: A dataset for evaluating experimental batch correction methods. In IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW). 4285–4294 (2023).
Bhatt, S. et al. Characterizing the consistency of motion of spermatozoa through nanoscale motion tracing. F. S. Sci. 5, 215–224 (2024).
He, W. et al. Neuromorphic-enabled video-activated cell sorting. Nat. Commun. 15, 10792 (2024).
Nitta, N. et al. Intelligent image-activated cell sorting. Cell 175, 266–276 (2018).
Ghiglia, D. C. Two-Dimentional Phase Unwrapping: Theory, Algorithms, and Software (Wiley, New York, 1998).
Běhal, J. Quantitative phase imaging in common-path cross-referenced holographic microscopy using double-exposure method. Sci. Rep. 9, 9801 (2019).
Colomb, T. et al. Total aberrations compensation in digital holographic microscopy with a reference conjugated hologram. Opt. Express 14, 4300–4306 (2006).
Acknowledgements
The authors acknowledge the financial support from the Natural Science Foundation of Guangdong Province 2024A1515011906 (to Y.T.) and the National Key R&D Program of China, Synthetic Biology Research 2023YFA0913600 (to Y.T.).
Author information
Authors and Affiliations
Contributions
J.S., Y.T. and P.S. conceived the idea and initiated the study. Y. T., P.S., D.G. and J.M. directed the project. J.S. constructed the spatial microfluidic holographic integrated platform. Y. L. assisted with the preparation of breast, lung, colorectal and cervical cancer cells. Z.Y. facilitated the acquisition of blood samples. H.T. and Z.Z. helped with the blood cell separation. J.S., H.T. and Z.Z. performed imaging experiments. J.S., J.D., P.S. and Y.T. contributed to the data analysis and writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, J., Teng, H., Zhang, Z. et al. Spatial microfluidic holographic integrated platform for label-free and high-dimensional analysis of cancer heterogeneity. Nat Commun 16, 5890 (2025). https://doi.org/10.1038/s41467-025-60897-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60897-w