Abstract
Ex vivo normothermic machine perfusion has been proposed to protect deceased donor kidneys. However, its benefits remain ambiguous. We postulate that the use of red blood cells (RBCs) and associated secondary hemolysis may in fact cause renal injury, offsetting potential advantages. During 48-hour normothermic perfusion of seven human donor kidneys, we observed progressive hemolysis, leading to iron accumulation in perfusate and tissue. Untargeted lipidomic profiling revealed significant increases in oxidized phospholipid species in perfused kidneys, pointing towards iron-dependent cell death known as ferroptosis. Next, in twelve additional perfusions, we assessed strategies to mitigate hemolysis-driven injury. Dialysis-based free hemoglobin removal reduced lipid peroxidation, but a ferroptosis gene signature persisted. In contrast, cell-free perfusion at subnormothermia negated iron accumulation, the ferroptosis gene signature, phospholipid peroxidation, and acute kidney injury. Our findings highlight the pathological role of hemolysis and iron on the kidney, urging restraint in the clinical application of RBC-based kidney perfusion.
Similar content being viewed by others
Introduction
Machine perfusion technologies have emerged as an important new modality in the field of organ preservation and transplantation. To address the growing disparity between the available donor organs and the number of patients on the transplant waiting list1, significant efforts have been devoted to developing advanced perfusion platforms. These platforms aim to create a window of opportunity to facilitate the assessment, reconditioning, repair, and regeneration of deceased donor organs2.
While normothermic machine perfusion (NMP) techniques for liver transplantation have become routine clinical practice3,4, successful translation into the field of kidney transplantation, which constitutes the majority of transplant organs, remains elusive. This raises the question how ex vivo NMP of deceased donor livers has been extended up to 12 days5, with successful transplantation after 3 days of perfusion6, whereas studies on normothermic kidney perfusion typically range from 1 to 6 h, and even then, the first randomized controlled trial was discontinued in the absence of a discernible positive effect7. This is concerning given the increasing clinical application of normothermic kidney perfusion by organ procurement organizations and highlights the need to critically evaluate whether all organs are equally suited for each preservation method.
In this study, we sought to elucidate the molecular changes that occur in kidney grafts during NMP, encompassing a detailed analysis of the multi-day perfusion of 19 deceased human donor kidneys. Of these, seven kidneys were perfused up to 48-h. We observed that hemolysis provokes iron accumulation in perfusate and tissue. Through untargeted lipidomic profiling we could confirm the accumulation of oxidized phospholipid species (oxPL), pointing towards iron-dependent cell death known as ferroptosis. Next, we demonstrate that dialysis-based free hemoglobin (fHb) removal reduces lipid peroxidation, but a ferroptosis gene signature persists. Finally, we show that cell-free perfusion at subnormothermia, an approach we recently demonstrated to support four-day human kidney preservation8, negates iron accumulation, the ferroptosis gene signature, phospholipid peroxidation, and acute kidney injury (AKI).
Results
Study design and donor characteristics
In total, 19 human donor kidneys deemed unsuitable for transplantation were perfused. During the first phase, seven kidneys were perfused up to 48-h at normothermia with an RBC-based perfusate. In the second phase, an additional twelve human kidneys were included for either dialysis-based fHb removal during RBC-based perfusion or cell-free perfusion. Donor and RBC characteristics are reported in Table S1.
Kidney perfusion dynamics
A clinical grade perfusion platform was adapted to support prolonged (≥24-h) preservation (Fig. 1A). During the first phase, seven kidneys were perfused up to 48-h. The perfusate was composed of two units of washed red blood cells (RBCs), human serum albumin and DMEM F12 as main components (for full composition see Table S2). Stored RBCs were washed before adding them to the perfusate to remove hemolytic components and the RBC storage solution (Fig. S1). Of the seven kidneys, five maintained a stable renal blood flow (RBF) throughout the 48-h perfusion period (Fig. 1B), whereas Kidney5 and Kidney6 had to be stopped earlier due to graft failure (defined as RBF ≤ 50 mL/min/100 g). After 48 h, all perfusions were stopped due to progressive disturbances in perfusion dynamics (Fig. S2). Blinded histologic scoring of periodic acid-Schiff (PAS) staining was performed by a renal pathologist (Table S3), demonstrating an increase in tubular vacuolization and decrease in tubular dilation beyond 24 h of RBC-based perfusion. Cold preservation method (static cold storage (SCS) or hypothermic machine perfusion (HMP)) prior to warm perfusion had no discernible impact on perfusion dynamics (Fig. S3).
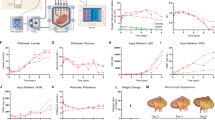
A Schematic overview of the perfusion platform. Kidneys were perfused with an RBC-based perfusate using a pressure-controlled impeller pump set at 75 mmHg following oxygenation at normothermia. Urine was recirculated, and continuous hemofiltration allowed removal of small molecular weight waste products and substitution with fresh perfusate. B Renal blood flow during perfusion. C Perfusate free hemoglobin (fHb) during perfusion. D Association between perfusate fHb and urine fHb levels (Spearman’s rank correlation coefficient, two-sided. Shaded area indicates 95% CI). E Macroscopic appearance of a control kidney, Kidney1 with low fHb accumulation, and Kidney6 that failed after 34-h with significant fHb accumulation. F Immunohistochemistry for heme oxygenase-1 (HO1) in serial biopsies obtained during perfusion (n = 7). Representative images from Kidney3 are shown. Scale bar, 50 μm. G Experimental design comparing RBC-based perfusion with (n = 7) or without (n = 3) a deceased donor kidney present in the platform. H Perfusate fHb concentration during perfusion (Mean ± SD). I Perfusate iron concentration during perfusion (Mean ± SD. Two-way ANOVA). J Association between perfusate fHb and perfusate iron concentrations across both groups (Spearman’s rank correlation coefficient, two-sided. Shaded area indicates 95% CI). K Temporal changes in expression of genes involved in iron uptake, iron storage, iron export, iron scavenging, and redox defense during RBC-based human kidney perfusion. Log2 foldchange relative to T0 is indicated by color; significance (adjusted P ≤ 0.05) by dot size and border (two-sided Wald test with Benjamini-Hochberg (BH) correction). L Immunofluorescence for DMT1 (Fe-uptake), FTH1 (Fe-storage), and Ferroportin (Fe-export) in pre- and post-perfusion biopsies (n = 7). Representative images from Kidney3 are shown. Scale bar, 200 and 50 μm, respectively. M Enhanced Perl’s Prussian Blue (EPPB) staining for iron deposition in control (n = 4) and RBC-perfused kidneys (n = 7). Representative images and quantification (Mean ± SD. Unpaired two-sided t-test). Scale bar, 50 μm. A, G Illustrated by Manon Zuurmond (scientific illustrator). Source data are provided as a Source Data file.
Hemolysis drives progressive iron accumulation
Over the course of perfusion, we observed the occurrence of hemolysis with the buildup of fHb within the perfusate (Fig. 1C) and associated hemoglobinuria (Fig. 1D), corroborating earlier preliminary data9. Distinct macroscopic differences were discernible between a control kidney, Kidney1, which exhibited minimal fHb buildup, and Kidney6, which failed after 34 h with significant fHb accumulation (Fig. 1E). Notably, both renal grafts that experienced failure during perfusion were exposed to the highest levels of fHb accumulation.
Heme oxygenase-1 (HO1), the endogenous defense mechanism against hemolysis-induced kidney injury10, was upregulated between 6 and 24 h of perfusion (Fig. 1F). This enzyme degrades heme into labile iron, carbon monoxide and biliverdin10, mitigating the deleterious effects of fHb accumulation.
Hemolysis is an inherent consequence of mechanical perfusion of (stored) RBCs10,11. Preliminary experiments, in which RBCs were perfused throughout the platform in the absence of a donor kidney, confirmed that the occurrence of hemolysis during prolonged perfusion (≥6 h) was primarily attributable to the normothermic perfusion system itself, rather than the donor graft (Figs. 1G–J; S4), corroborating previous observations reported in the context of liver perfusion12,13. Importantly, assessment of perfusate iron levels during RBC-only perfusion indicates the use of RBCs and subsequent hemolysis as a source of circulating iron, with significant iron uptake by the perfused kidney as early as 6 h into perfusion (Fig. 1I, J).
Through bulk RNA sequencing on biopsies taken during RBC-based kidney perfusion, we assessed expression dynamics of key iron uptake, iron storage, iron export, iron scavenging, and redox defense genes (Fig. 1K). The upregulation of iron uptake and storage genes, alongside the downregulation of iron export genes, strongly supports the observed uptake of circulating iron from the perfusate. This was further confirmed at the protein level for divalent metal transporter-1 (DMT1, Fe-uptake), ferritin heavy chain-1 (FTH1, Fe-storage), and ferroportin (FPN, Fe-export) (Fig. 1L). Finally, Enhanced Perl’s Prussian Blue (EPPB) staining indeed demonstrated the accumulation of tissue iron within RBC-perfused kidneys (Fig. 1M).
We conclude that the use of RBCs, and subsequent hemolysis, during machine perfusion not only results in renal exposure to high levels of fHb but also drives cellular accumulation of hemolysis-derived iron.
RBC-based perfusion triggers ferroptosis
Dysregulated cellular iron homeostasis is known to induce lipid peroxidation, the main characteristic of an iron-dependent type of cell death called ferroptosis11,14. In an untargeted approach, we sought to characterize changes in the cellular lipidome following RBC-based perfusion, using liquid chromatography tandem mass spectrometry (LC-MS/MS) with subsequent lipid identification through MS-DIAL15. Overall, 539 unique lipid species were identified, belonging to 11 lipid classes (Fig. 2B and Table S4). To depict differences between control kidneys and those subjected to RBC-based perfusion, we projected relative lipid abundance of the 539 lipid species across these 11 lipid (sub)classes on a heatmap (Fig. 2C, D). Our analysis revealed that 44 lipid species exhibited significant increases in perfused kidneys (Log2FC ≥ 1.5 and a P value < 0.05; Table S5), the majority of which were oxPL (Fig. 2E), indicating significant oxidative modifications during RBC-based kidney perfusion. Normalization to their non-oxidized PL (non-oxPL) pool confirmed significant increases in oxPL/non-oxPL abundance (Fig. 2F, G). PL peroxidation results in the formation of lipid peroxides, which can further decompose into toxic secondary products, such as malondialdehyde (MDA) and 4-hydroxynonenal (4HNE), commonly used as biomarkers for ferroptosis16. Notably, levels of MDA, assessed as thio-barbituric acid reactive substances (TBARS) in perfusate, and 4HNE in tissue increased significantly over the 48-h RBC-based perfusion period (Fig. S5).
A Experimental design comparing RBC-perfused kidneys (n = 7) to unperfused controls (n = 3) and RBC-only perfusions (n = 3). B Representation of 539 lipid species detected through untargeted LC-MS/MS lipidomics, grouped by lipid (sub-)class (also see Table S4). Heatmaps showing relative lipid abundance for individual lipid species (C) and grouped into 11 lipid (sub-)classes (D) in control and RBC-perfused kidneys (n = 3 and n = 7, respectively). Z-scores represent total area normalized abundance. TG Triradylglycerols, DG Diradylglycerols, FA Fatty acids, FE Fatty esters, PL Phospholipids, LPL Lyso-phospholipids, EtherPL Ether-linked phospholipids, oxPL Oxidized phospholipids, OtherPL Other phospholipids (BMP (4), CL (8), and MLCL (1)), SP Sphingolipids, ST Sterol lipids. E Volcano plot representing the 539 lipid species comparing RBC-perfused with control kidneys (unpaired two-sided t-test with Welch’s correction). OxPL species are highlighted in red. Significantly increased lipid species (Log2FC ≥ 1.5 and P-value < 0.05) are detailed in Table S5. Abundance of oxidized PCs/non-oxidized PCs (F) and oxidized phosphatidylinositols (PIs)/ non-oxidized PIs (G) (Mean ± SD; n = 3 and n = 7, respectively. Unpaired two-sided t-test). H Levels of arachidonic acid (AA)-derived lipid mediators in perfusate from RBC-perfused kidneys (n = 7) and RBC-only perfusions (n = 3) over time (Median with IQR). Box plot of gene set variation analysis (GSVA) enrichment score of WP_Ferroptosis (I, n = 64 genes) and FerrDb v2 driver, suppressor and marker genes ( J, n = 313 genes) (Mixed-effects ANOVA). Boxes represent median with 25th and 75th percentile and error bars minimum to maximum. K Volcano plots of differentially expressed genes over time during RBC-based kidney perfusion. Black highlights FerrDb v2 marker genes (n = 9). Red highlights validated FerrDb v2 driver or suppressor genes with a protein product (n = 312). L Dotplot illustrating temporal expression dynamics in FerrDb v2 driver, suppressor, and marker genes (|Log2FC| ≥ 2 and adjusted P ≤ 0.05 for at least one of the timepoints). Two-sided Wald test with BH correction (K, L). A Illustrated by Manon Zuurmond (scientific illustrator). Source data are provided as a Source Data file.
As hemolysis occurs already at the beginning of perfusion, we next assessed dynamics in phospholipid peroxidation during perfusion, for which oxilipidomic analysis (LC-MS/MS) was performed on perfusate taken before connection of the kidney (T0), and after 6, 24, and 48 h. This allowed differentiation between enzymatic phospholipid peroxidation—mediated by lipoxygenases (LOXs) and cyclooxygenases (COXs)—and non-enzymatic phospholipid peroxidation, which primarily occurs via Fenton reaction17. Comparison of RBC-only with RBC-based kidney perfusion revealed the presence of LOX-derived products across all timepoints in both experimental conditions (Fig. 2H). Conversely, COX- and non-enzymatic phospholipid peroxidation-derived products were absent during RBC-only perfusion, and progressively increased over the course of RBC-based kidney perfusion (Fig. 2H).
Significant enrichment of ferroptosis gene sets was observed 6 h into RBC-based perfusion (Fig. 2I-J). Next, we compared differentially expressed genes across timepoints, specifically focusing on ferroptosis marker, driver and suppressor genes (as defined by FerrDb v218) (Fig. 2K), for which the genes with the most significant dynamics (|Log2FC| ≥ 2 and padj ≤ 0.05) for at least one of the timepoints are visualized in Fig. 2L.
We conclude that the progressive accumulation of oxidized phospholipids, coupled with the enrichment of ferroptosis gene sets, points towards ferroptotic cell death as a result of hemolysis-driven iron accumulation during RBC-based kidney perfusion.
Interventions to mitigate hemolysis-driven renal injury
Having established that progressive hemolysis during machine perfusion leads to renal iron accumulation and the buildup of oxPL species, we questioned whether these effects could be mitigated. In the second phase of this study, we perfused an additional twelve human kidneys, assigning them to two intervention groups: fHb removal during RBC-based perfusion (Group2) and cell-free perfusion (Group3).
Free hemoglobin removal reduces phospholipid peroxidation
With the first intervention (Group2), we aimed to remove fHb from the perfusate by replacing the HF20 dialysis filter with the Oxiris filter, a novel blood purification filter for continuous renal replacement therapy that removes fHb (Fig. 3A). Four kidneys were perfused with the Oxiris filter up to 48-h, showing comparable perfusion dynamics and histology to the RBC-perfused kidneys (Fig. S6 and Table S3). Dialysis-based fHb removal significantly decreased perfusate fHb levels (Fig. 3B, C), with substantial clearance into the effluent (Fig. 3D). This was accompanied by a trend towards reduced tissue iron accumulation when compared to RBC-perfused kidneys (Fig. S7).
A Experimental design comparing RBC-based perfusion of deceased donor kidneys without (Group 1, n = 7) or with (Group 2, n = 4) dialysis-based free hemoglobin (fHb) removal. Perfusate fHb levels during perfusion (B), and perfusate (C) and effluent (D) fHb levels after 24 and 48 h of perfusion (Mean ± SD. Two-way ANOVA). E Total oxidized phosphatidylcholines (oxPC) in tissue as detected by targeted oxPC LC-MS/MS in control (n = 4), RBC-perfused (n = 7), and fHb removal (n = 3) groups (Mean ± SD. One-way ANOVA). F Heatmap of individual oxPC species. See Table S6 for lipid identities and their abbreviations. G Levels of arachidonic acid (AA)-derived lipid mediators in perfusate during RBC-based (n = 7) and fHb removal during RBC-based perfusion (n = 3–4) (Median with IQR). H, I Box plot of GSVA enrichment score of WP_Ferroptosis (I, n = 64 genes) and FerrDb v2 driver, suppressor and marker genes ( J, n = 313 genes) (One-way ANOVA). Boxes represent median with 25th and 75th percentile and error bars minimum to maximum. J Volcano plots of differentially expressed genes in kidneys after RBC-based perfusion or fHb removal during RBC-based perfusion, compared to control kidneys (left and middle panel). Correlation of Log2FC changes between RBC-perfused vs control kidneys and fHb removal vs control kidneys (right panel). Red highlights validated FerrDb v2 driver, suppressor, or marker genes with a protein product (n = 313). K Temporal expression dynamics in FerrDb v2 driver, suppressor, and marker genes for RBC-perfused and fHb removal groups relative to control kidneys (|Log2FC| ≥ 2 and adjusted P ≤ 0.05 for at least one of the comparisons). Two-sided Wald test with BH correction ( J, K). Association between perfusate fHb and lipid peroxidation markers after 48-h of perfusion: perfusate TBARS (MDA) (L) and tissue 4-hydroxynonenal (4HNE) (M) (Pearson’s correlation coefficient, two-sided. Shaded area indicates 95% CI). A Illustrated by Manon Zuurmond (scientific illustrator). Source data are provided as a Source Data file.
Through targeted LC-MS/MS oxidized phosphatidylcholines (oxPC) analysis on tissue at the end of preservation, we observed that the oxidative modifications during the RBC-based perfusions were reduced through fHb removal (Fig. 3E, F). However, oxilipidomic profiling revealed continued release of LOX-, COX-, and non-enzymatic oxidation products into the perfusate (Fig. 3G), indicating persistent ferroptotic activity. This was further supported by unaffected enrichment of the ferroptosis-related gene signatures following fHb removal (Fig. 3H–K).
Hence, while fHb removal attenuates phospholipid peroxidation, it does not completely prevent it, likely due to the continued presence of fHb in the perfusate, albeit at reduced levels. Notably, despite fHb removal during RBC-based perfusion, MDA (measured as TBARS) in perfusate and 4HNE in tissue still demonstrated a strong correlation with perfusate fHb and iron levels, highlighting the persistent pro-oxidative effects of residual hemolysis products (Figs. 3L, M; S7).
We conclude that, even after fHb removal, residual hemolysis continues to contribute to lipid peroxidation, albeit at lower levels when compared to RBC-based perfusion.
Cell-free perfusion negates hemolysis-driven iron accumulation and ferroptosis
The most definitive approach to avoid hemolysis, which would be the removal of RBCs from the perfusate altogether, was tested in the second intervention group (Group3) (Fig. 4A). Lowering the perfusion temperature from 37 °C to 25 °C allows for cell-free perfusion as it reduces the metabolic rate, and therewith oxygen requirements. As we previously demonstrated, this cell-free perfusion approach enables 4 day ex vivo metabolic and functional preservation of human kidneys8. Eight kidneys were preserved with a cell-free perfusate, which confirmed improved metabolic preservation during 48-h perfusion when compared to RBC-based perfusion (Fig. S8).
A Experimental design comparing RBC-based perfusion (Group 1, n = 7) with cell-free perfusion (Group 3, n = 8). Perfusate iron levels during perfusion (B) and at 24 and 48 h (C) (Mean ± SD, two-way ANOVA). D Iron deposition visualized by Enhanced Perl’s Prussian Blue (EPPB) staining in control (n = 4), RBC-perfused (n = 7), and cell-free perfused (n = 5) kidneys. Representative images and quantification (Mean ± SD. One-way ANOVA). Scale bar, 200 μm. E Total oxidized phosphatidylcholines (oxPC) in tissue as detected by targeted oxPC LC-MS/MS in control (n = 4), RBC-perfused (n = 7), and cell-free perfused (n = 4) kidneys (Mean ± SD. One-way ANOVA). F Heatmap of individual oxPC species. See Table S6 for lipid identities and their abbreviations. G Perfusate TBARS (MDA) levels in RBC-perfused (n = 7) and cell-free perfused (n = 8) kidneys (Mean ± SD. Two-way ANOVA). H Levels of arachidonic acid (AA)-derived lipid mediators in perfusate during RBC-based (n = 7) and cell-free (n = 8) perfusion (Median with IQR). Box plot of GSVA enrichment score of GOBP_Iron_Ion_Transport (I, n = 58 genes, Kruskall-Wallis test), WP_Ferroptosis ( J, n = 64 genes, one-way ANOVA) and validated FerrDb v2 driver, suppressor and marker genes (K, n = 313 genes, one-way ANOVA). Boxes represent median with 25th and 75th percentile and error bars minimum to maximum. L Volcano plots of differentially expressed genes after RBC-based or cell-free perfusion, compared to control, and cell-free perfusion compared to RBC-based perfusion. Red highlights validated FerrDb v2 driver, suppressor, or marker genes with a protein product (n = 313). M Temporal expression dynamics in FerrDb v2 driver, suppressor, and marker genes after RBC-based or cell-free perfusion, compared to control kidneys, and cell-free perfusion compared to RBC-based perfusion (|Log2FC| ≥ 2 and padj ≤ 0.05 for at least one of the comparisons). Two-sided Wald test with BH correction (L, M). Perfusate LDH (N) and NGAL (O) concentration after 48 h of RBC-based (n = 7) and cell-free (n = 8) perfusion. (Mean ± SD. Mann-Whitney test, two-sided). P Association between perfusate NGAL and iron after 48 h of perfusion (Spearman’s rank correlation coefficient, two-sided. Shaded area indicates 95% CI). A Illustrated by Manon Zuurmond (scientific illustrator). Source data are provided as a Source Data file.
Perfusate iron levels did not increase during 48-h cell-free perfusion, in contrast to RBC-based perfusion. (Fig. 4B, C). Similarly, tissue iron accumulation, as observed in RBC-perfused kidneys, was absent during cell-free perfusion (Fig. 4D). This confirms that the use of RBCs, and subsequent hemolysis, drives iron accumulation during RBC-based kidney perfusion. Significantly lower perfusate bilirubin levels following cell-free perfusion further point towards a role of HO1-mediated breakdown of fHb into labile iron (Fig. S8).
Targeted oxPC analysis revealed that oxidative modifications observed after RBC-based perfusions were negated by cell-free perfusion (Fig. 4E, F). In line with this, perfusate TBARS levels did not increase during cell-free perfusion (Fig. 4G). Oxilipidomic profiling in the perfusate showed the presence of LOX- and COX-derived oxidation products during cell-free perfusion, with COX-derived products being less abundant as compared to RBC-based perfusion. Importantly, non-enzymatic oxidation products were undetectable in the perfusate during 48-h cell-free kidney perfusion (Fig. 4H), supporting the hypothesis that fHb-derived iron plays a critical role in Fenton-driven non-enzymatic lipid peroxidation during RBC-based kidney perfusion.
Although the cell-free perfusion group included both DCD and DBD donors, whereas the RBC-perfused group only contained DCD donors, donor background had a negligible impact on perfusion outcomes (Fig. S9).
The distinct differences observed in tissue iron accumulation and lipid peroxidation between RBC-based and cell-free perfusion were corroborated at the transcriptomic level, where cell-free perfusion showed markedly reduced enrichment of iron ion transport and ferroptosis gene sets (Fig. 4I–K). Next, differential gene expression analysis further highlighted differences in ferroptosis marker, driver and suppressor genes between RBC-based and cell-free perfusion (as defined by FerrDb v218) (Fig. 4L), for which the genes with the most significant changes (|Log2FC| ≥ 2 and padj ≤ 0.05) for at least one of the timepoints are visualized in Fig. 4M.
The absence of iron accumulation and subsequent lipid peroxidation during cell-free perfusion underscore that RBCs, and subsequent hemolysis, drive iron accumulation and ferroptosis during RBC-based kidney perfusion. Finally, we assessed biomarkers for AKI during perfusion, observing significantly higher levels of perfusate lactate dehydrogenase (LDH) and neutrophil gelatinase-associated lipocalin (NGAL) in RBC perfused kidneys (Fig. 4N, O). This is striking given that two donor kidneys in the cell-free perfusion group had significant dysfunction at the time of donation (as indicated by a final serum creatinine >200 µmol/L) (Fig. S10).
As NGAL is an iron-sequestering siderophore, these findings raise the possibility that NGAL is produced by the kidney during RBC-based perfusion in response to hemolysis to protect the kidney from iron-induced injury. Indeed, we did find a strong correlation between perfusate iron and NGAL (Fig. 4P), which is in line with experimental studies that have suggested that NGAL may play a protective role during AKI19. Whereas NGAL is a kidney-specific marker, LDH can also originate from hemolysis. However, supporting experiments confirmed that the observed increases are predominantly kidney-derived (Fig. S3), confirming that hemolysis during RBC-based kidney perfusion exacerbates renal injury.
Discussion
In this study, we observed that the use of RBCs and associated hemolysis during machine perfusion of diseased donor kidneys drives progressive iron accumulation and phospholipid peroxidation. The observed accumulation of fHb, iron, oxidized phospholipids, and their byproducts highlights how the use of RBCs and associated hemolysis within preservation platforms leads to ferroptosis. Finally, we demonstrate that dialysis-based fHb removal reduces, and cell-free perfusion negates, hemolysis-driven iron accumulation, phospholipid peroxidation, and AKI.
Ferroptosis is an iron-dependent form of cell death that is distinguished from other types of cell death by the occurrence of excessive phospholipid peroxidation11,20, and has been found to play a central role in renal ischemia reperfusion injury, maladaptive repair, and the development of kidney fibrosis14,21,22,23. Unlike the liver, within which resident macrophages effectively and on-demand manage the removal of damaged RBCs and fHb, and therewith provide resilience against hemolysis12,24,25, the kidney is particularly vulnerable to elevated levels of fHb and the resultant hemoglobinuria10,11,26. Preclinical studies have emphasized the importance of attenuating iron and lipid peroxide accumulation in the kidney, as this propagates ferroptotic cell death and culminates in AKI22,27. Various clinical investigations have demonstrated that hemolysis during cardiopulmonary bypass surgeries (bypass time 1–3 h) is associated with AKI, need for renal replacement therapy, and even mortality28,29.
To mitigate hemolysis-driven ferroptotic cell death during RBC-based perfusion, we performed dialysis-based fHb removal. Although this intervention allowed the removal of significant levels of fHb and reduced phospholipid peroxidation, transcriptomic analysis revealed persistent ferroptosis signature gene activity. A probable explanation is the continued exposure to residual supraphysiological levels of fHb in the absence of endogenous hemolysis scavengers. Importantly, and supporting this, we observed strong correlations between hemolysis products and lipid peroxidation biomarkers, emphasizing the importance of minimizing graft exposure to hemolysis. Despite mitigation efforts, hemolysis remains an inherent limitation to mechanical perfusion platforms. Potential strategies include platform optimization, improving fHb removal efficiency, washing stored RBCs to remove hemolytic components, or shielding RBCs through silicification30. However, in all scenarios (reduced levels of) biologically active hemolysis products will remain present. An alternative would be the addition of hemolysis-scavengers like haptoglobin, which in addition, should attenuate (or postpone) observed hemoglobinuria through size-exclusion31.
Given that residual ferroptotic activity persisted despite fHb removal during RBC-based perfusion, we explored cell-free perfusion to circumvent hemolysis-driven injury. To support aerobic metabolism without an oxygen carrier, preservation temperature was reduced from 37 °C to 25 °C, thereby lowering the metabolic rate ensuring adequate oxygen delivery. Our findings indicate that cell-free perfusion prevents hemolysis-induced iron accumulation, phospholipid peroxidation, and ferroptosis gene signature enrichment, when compared to RBC-based perfusion. Hence, this approach establishes a more graft-protective environment for (prolonged) preservation of deceased donor kidneys. Although subnormothermic preservation is often considered a limitation, recent studies demonstrated it supports molecular and cellular repair processes32,33. Moreso, we previously demonstrated that it supports 4 day ex vivo metabolic and functional preservation of human kidneys8.
Machine perfusion of deceased donor organs is a rapidly developing field that holds the potential to change transplantation medicine. Despite the prevailing assumption that RBC-based NMP confers protection to deceased donor kidneys2,34, our findings challenge this notion by demonstrating that the use of RBCs as oxygen carriers, and associated hemolysis, may actually offset the anticipated benefits of normothermic perfusion. While our analysis primarily focused on prolonged preservation (≥6 h), iron uptake from the perfusate, a ferroptosis gene signature, and release of phospholipid peroxidation products were already observed after 6 h of RBC-based perfusion. This is concerning given an increasing number of clinical trials looking into normothermic kidney perfusion, and clinical implementation of kidney NMP by commercial organ procurement organizations.
Consequently, as our study underscores the pathological role of hemolysis and iron in exacerbating AKI, we urge caution in the clinical implementation of RBC-based perfusion for deceased donor kidneys. Our findings challenge the current kidney preservation dogma and necessitate reconsideration of the framework needed to establish a protective environment to facilitate ex situ reconditioning, repair, and regeneration of deceased donor kidneys.
Methods
Ethics
This research complies with all relevant ethical regulations. Leiden University Medical Center (LUMC) received authorization from the Dutch government (BWBR0008974, 2555663-CZ/IZ/2562427) for kidney transplantation and associated research. Prior to organ retrieval for all human kidneys, donor research consent was obtained by Eurotransplant, the centralized donation organization in the Netherlands (BWBR008066, Art. 13). This written consent was acquired by an independent organ donation coordinator unaffiliated with the research team.
Study design
Nineteen kidneys from deceased donors were subjected to multi-day perfusion and allocated into three experimental groups: seven kidneys were perfused with a red blood cell (RBC)-based perfusate at normothermia (37 °C) (“RBC-based”; Group1), four kidneys were perfused with a similar RBC-based perfusate at normothermia but incorporated a dialysis filter that allowed removal of fHb (“fHb removal”; Group2), and eight kidneys were perfused with a cell-free perfusate (i.e., no RBCs) at subnormothermia (25 °C) (“cell-free”; Group3). Tissue biopsies obtained at the end of perfusion were compared to biopsies taken from control deceased donor kidneys and assessed through both untargeted lipidomics and targeted oxidized phosphatidylcholines (oxPC) analysis. Additionally, three perfusions were performed using an RBC-based perfusate at normothermia in the absence of a donor kidney (“RBC-only”) to assess the interaction between the perfusion platform, stored RBCs, and deceased donor kidneys.
Perfusion platform and protocol
Adapted perfusion platform
A clinical-grade perfusion platform (CirQlife, XVIVO) was modified to support prolonged (≥24-h) preservation. This adapted platform consists of a closed-loop circuit connected to a custom-designed 3D printed organ chamber (Oceanz) that also serves as a venous reservoir. A pressure-controlled impeller pump (CirQlife, XVIVO) perfused the renal artery through silicone tubing (LS17 or LS25, Masterflex Metrohm) at a mean arterial pressure of 75 mmHg (60 BPM, 20% amplitude). Pressure in the renal artery was measured in line with a single-use pressure sensor (Edwards Lifesciences). Flow was monitored using an ultrasonic clamp-on flow probe (Transonic Systems Europe B.V.), and temperature was monitored using a temperature sensor (Heraeus Nexensos PT100 Surface Sensor [W-SZK], Heraeus Nexensos GmbH), which was positioned underneath the organ. Arterial pressure, perfusate flow and temperature were continuously recorded. The perfusate was oxygenated with a carbogen mixture of 95% O2 and 5% CO2 through a long-term membrane oxygenator (Lilliput 2 ECMO, LivaNova) that was connected to a water bath set at 37 °C. Urine was recirculated.
A Prismaflex system (Baxter) was connected in parallel to remove metabolic waste products and supply fresh nutrients whilst maintaining electrolyte homeostasis. A pediatric filter (Prismaflex HF20 set, Baxter) allowed the exchange of small molecular weight molecules. Blood flow over the filter was set at 20 mL/min. Fluid and small molecular weight molecules were removed at 80 mL/hr over the filter. Fresh substitution solution (see Table S2 for composition) was substituted post-filter at the same rate (filtration fraction of 10%). A separate infusion pump infused additional glucose (5%, WE0087G, Baxter) at a rate of 0–7 mL/h to maintain glucose above 4 mmol/L.
Human kidneys
After in situ flushing of the kidneys with cold University of Wisconsin (Belzer UW, Bridge to Life Ltd.) preservation solution, the kidneys were retrieved and transported to the LUMC by SCS or HMP. Upon arrival at the laboratory, the renal artery and ureter were cannulated using Luer lock connectors (Cole Parmer) whilst the perfusion platform was set up and primed in parallel. Shortly before the start of culture, kidneys were flushed with DMEM F12 supplemented with human serum albumin (40 g/L).
Perfusate
A detailed description of the perfusate composition is provided in Table S2. Two units of stored clinical-grade leukocyte-depleted RBCs (Sanquin) were used per perfusion. Prior to their addition, stored RBCs were washed three times with DMEM F12 through centrifugation at 1000 × g for 10 min (21 °C, acc9, dec4) to remove RBC storage solution and hemolytic RBCs9,35.
In parallel, the perfusion platform was primed with the other perfusate components, after which the washed RBCs were added. Depending on perfusate acidity, pH was adjusted with sodium bicarbonate (7.5%, Gibco) until within range (Target 7.25–7.40). Perfusion was started with a volume of ~1 L.
Intervention group: fHb removal
The pediatric HF20 filter was exchanged for an Oxiris dialysis filter (Baxter), allowing removal of fHb into the effluent (dialysis waste). fHb levels were measured in both perfusate and effluent over the course of perfusion. The substitution solution remained unchanged. Blood flow over the filter was set at 20 mL/min. Fluid was removed at 100 mL/h over the filter, with fresh substitution solution being supplemented post-filter at the same rate (filtration fraction of 13%). Of the four human kidneys perfused in this group, one was stopped after 24 h of perfusion because of logistical reasons.
Intervention group: cell-free perfusion
Kidneys were perfused using our previously described cell-free perfusion approach, which enabled 4 day ex vivo metabolic and functional preservation of human kidneys at subnormothermia (25 °C)8. Lowering perfusion temperature from 37 °C to 25 °C allows for cell-free perfusion as it reduces the metabolic rate and therewith oxygen requirements. Hence, hemolysis cannot occur and fHb is absent.
Control group: RBC-only perfusions
For the three perfusions performed without a kidney graft, we followed the same procedure as for RBC-based kidney perfusion, except that the donor kidney was absent. The resistance normally provided by the kidney was simulated by partially clamping the tubing at the place where the renal artery is normally connected, thereby creating a flow rate and vascular resistance similar to those observed during RBC-based kidney perfusions.
Assessment of kidney preservation and injury
Perfusate was sampled at least three times per day throughout the perfusion period. Arterial, venous and urine samples were measured directly for blood-gas analyzes and monitoring of electrolytes and metabolites using an iSTAT1 blood analyzer (Abbott) and CG4+ (pH, pO2, pCO2, HCO3−, lactate, and saturation) and Chem8 iSTAT cartridges (sodium, potassium, chloride, glucose, and Hb) (Abott). Oxygen uptake (mL O2/min/100 g) was calculated as:
Perfusate samples were centrifuged at 1000 × g for 7 min (4 °C, acc9, dec4), whereafter the supernatant was aliquoted and frozen at −80 °C for later analysis. Effluent samples (i.e., fluid removed through continuous hemofiltration) were sampled after 24 and 48 h.
The Clinical Chemistry Laboratory within the LUMC measured perfusate, urine, and effluent fHb, albumin, total bilirubin (tBil), total iron, LDH, and ferritin according to standard operating procedures, which had previously been stored at −80 °C.
NGAL and kidney injury molecule-1 (KIM1) levels in the perfusate were measured using a quantitative sandwich enzyme immunoassay technique with NGAL Quantikine ELISA kit (DLCN20, R&D systems) and KIM-1 DuoSet ELISA kit (DY1750B, R&D systems) according to manufacturer’s instruction in samples that had previously been frozen and stored at −80 °C. A thiobarbituric acid reactive substances (TBARS) assay kit (CaymanChemical, 10009055) was used to measure levels of MDA and related lipid oxidation end products in the perfusate according to manufacturer’s instructions in samples that had previously been frozen and stored at −80 °C.
Tissue sampling and processing
Wedge biopsies were taken at the end of perfusion and from control deceased donor kidneys and either fixed in 4% formaldehyde (VWRK4187F, VWR) or snap frozen in liquid nitrogen and stored at −80 °C for further analysis. Punch biopsies (6 mm) were taken at four timepoints during perfusion (Hour 0, Hour 6, Hour 24, and Hour 48). At each timepoint biopsies were cut longitudinally into three pieces of which one was fixed in formaldehyde, one frozen in RNAlater (AM7020, ThermoFisher), and one snap frozen in liquid nitrogen. Paraffin-embedded tissue sections were sectioned at 5 µm for histochemical and immunohistochemistry staining.
Histochemical staining
For EPPB, paraffin-embedded tissue sections were dewaxed and stained with an iron stain kit (ab150674, Abcam). The human spleen was stained as a positive control. To enhance iron detection, we adapted the EPPB staining protocol as described previously36, by adding a blocking step (Protein block, s1699, Dako) for 60 min followed with 30 min of peroxidase blocking solution (ab64218, Abcam) and a 30 min incubation with 3,3-diaminobenzidine (DAB) substrate (ab64238, Abcam) on sections that had been previously stained with the iron stain kit. A counterstain with nuclear fast red solution was performed for 2 min. Samples were dehydrated and mounted with Entellan (1079610100, Merck). Whole slides were digitized using a 3D Histech Pannoramic MIDI Scanner (Sysmex) and viewed with CaseViewer software. A minimum of 7 Fields of View (3870 × 2050 µm) were selected per sample. Prussian blue positive area was quantified in ImageJ using the H DAB plugin after correction for background staining. PAS staining was performed on paraffin-embedded tissue sections by the Pathology department using an automatic slide stainer. Slides were digitized and scored by a blinded renal pathologist on acute tubular injury, tubular casts, tubular vacuolization, tubular dilation, and interstitial edema.
Immunohistochemistry
For immunohistochemistry, paraffin-embedded tissue sections were dewaxed. Antigen retrieval was performed using antigen retrieval buffer (s1699, Dako) in an autoclave, after which a 60 min protein block (X0909, Dako) and 15 min peroxidase block (ab64218, Abcam) were performed. Primary anti-HO1 (MA1-112, ThermoFisher) and anti-4HNE (MA5-27570, ThermoFisher) and isotype mouse IgG1 (14-4714-82, Invitrogen) were used. All primary antibodies and isotypes were incubated at 4 µg/mL overnight at 4 °C in 1%NDS/1% NGS/1%BSA in PBS. Thereafter, samples were incubated for 1–2 h at RT with goat anti-Mouse HRP (2 µg/mL, P0447, Dako), followed by incubation with DAB substrate (ab64238, Abcam) for ~10 min. A counterstain with Mayer’s hematoxylin was performed for 40 s. Samples were dehydrated and mounted with Entellan (1079610100, Merck). Whole slides were digitized using a 3D Histech Pannoramic MIDI Scanner (Sysmex) and viewed with CaseViewer software.
Immunofluorescence
For immunofluorescence, paraffin-embedded tissue sections were dewaxed. Antigen retrieval was performed using antigen retrieval buffer (s1699, Dako) in an autoclave, followed by a 60 min protein block (X0909, Dako). Primary anti-DMT1 (Ab55735, Abcam), anti-FTH1 (701934, Invitrogen), and anti-ferroportin (PA5-2293, ThermoFisher) and isotype mouse IgG2a (X0943, DAKO) and rabbit IgG (X0936, DAKO) were used. All primary antibodies and isotypes were incubated at 5 µg/mL overnight at 4 °C in PBS. Thereafter, samples were incubated for 1–2 h at RT with goat anti-Mouse IgG2a A647 (4 µg/mL, A-21241, ThermoFisher) or goat anti-rabbit IgG A568 (4 µg/mL, A-11011, ThermoFisher). A counterstain with Hoechst 33258 (1:10.000 in MQ, H3569, Life Technologies) was performed for 5 min. Samples were mounted with Prolong Gold (P36930, Invitrogen). Whole slides were digitized using a Zeiss Axioscan 7 microscope slide scanner (Zeiss) and viewed with Zen3.4.
Western blotting
Frozen kidney tissues were sectioned on a Cryostar NX70 cryostat (ThermoFisher) at 20 µm, with ~30 sections per kidney being collected in Eppendorf tubes, and dissolved in RIPA buffer (89900, ThermoFisher) freshly supplemented with protease and phosphatase inhibitors (78440, ThermoFisher) by pipetting. Protein content was determined through Pierce BCA assay kit (23227, ThermoFisher), and lysates were diluted to a concentration of 2222 µg/mL. Kidney lysate (27 µL) and Laemmli sample buffer (9 µL; 1610747, BioRAD) were combined, and boiled for 5 min at 100 °C. Then, 30 µL of lysates per lane (equal to 50 µg protein) and a ladder (1610374, BioRAD) were resolved on 10% Mini-PROTEAN TGX Precast Protein gels 4569034, BioRAD) and transferred onto 0.2 µm PVDF membranes (1704156, BioRAD), using Trans-Blot Turbo Transfer System (BioRAD). For blocking and antibody incubation steps, 5% skim milk (70166, Merck) in PBS was used. All washing steps were performed with 0.05% Tween 20 (P1379, Merck) in PBS. Blots were blocked overnight at 4 °C, incubated overnight at 4 °C with anti-4HNE (1 µg/ mL, MA5-27570, ThermoFisher) or for 1–2 h at room temperature (RT) with anti-ACTB (0.4 µg/mL, MA5-15739, ThermoFisher), and subsequently incubated with anti-Mouse HRP antibodies (0.02 µg/mL, P0447, DAKO) for 1–2 h at RT. For substrate visualization, SuperSignalTM West Femto (34095, ThermoFisher) and SuperSignalTM West Pico PLUS (34580, ThermoFisher) were used, respectively. Stripping buffer (21059, ThermoFisher) was used between 4HNE and ACTB. Blots were imaged with a ChemiDocTM Touch imaging system (BioRAD) and analyzed using Image Lab (v5.2). For densitometry, OD of the protein of interest relative to the housekeeping gene was normalized to the averages obtained in control samples.
Sample preparation of kidney tissue samples for untargeted lipidomics analysis
Frozen kidneys were homogenized in chilled LC-MS-grade water to obtain a final concentration of 100 mg/mL by means of a Bullet BlenderTM 24 (NY, USA, Next Advance Inc.) and 0.9–2.0 mm diameter stainless steel beads. To 25 µL of kidney tissue homogenate (equal to 2.5 mg of frozen tissue), 600 µL of methyl-tert-butyl ether (MTBE) and 150 µL LC-MS grade methanol were added. Then samples were vortexed and centrifuged at 18,000 × g for 5 min and supernatants were transferred to new Eppendorf tubes. The extraction was repeated with 300 µL MTBE and 100 µL methanol and after centrifugation, supernatants were combined. 300 µL of LC-MS grade water were added and after centrifugation at 18,000 × g for 5 min, the upper organic layer were transferred to a glass vial and evaporated under a gentle stream of N2. Dried samples were stored at −80 °C until the day of analysis. On the day of analysis, samples were reconstituted in 100 µL 2-propanol, vortexed and sonicated for 5 min. 100 µL of LC-MS grade water was added, and samples were vortexed and sonicated for 5 min again. The samples were transferred to inserts and were ready to be analyzed by the LC-MS/MS untargeted lipidomics platform. A quality control (QC) was made by pooling equal aliquots of every single sample and was analyzed periodically during the analytical run.
Untargeted lipidomic analysis by LC-MS/MS
10 µL of the lipid extracts were injected in a LC-MS/MS equipment composed of a Shimadzu Nexera X2 (Shimadzu) LC system using as eluents water:acetonitrile 80:20 (mobile phase A) and water:2-propanol:acetonitrile 1:90:9 (mobile phase B), both containing 5 mM ammonium formate and 0.05% formic acid. The applied gradient, at a 300 µL/min flow rate was: 0 min 40% B, 10 min 100% B, 12 min 100% B and then a 3.6 min re-equilibration at 40% B, using a Kinetex C18, 2.7 µm particle, 50 × 2.1 mm (Phenomenex) column and a SecurityGuard Ultra C8, 2.7 µm, 5 × 2.1 mm cartridge (Phenomenex) guard column, both kept at 50 °C The MS system was a Sciex TripleTOF 6600 (AB Sciex Netherlands B.V.) operated both in positive and negative electrospray (ESI) mode, with the following conditions: ion source gas 1 45 psi, ion source gas 2 50 psi, curtain gas 35 psi, source temperature 350 °C, acquisition range m/z 100–1800, ion spray voltage 5500 and −4500 V for ESI+ and ESI–, respectively, declustering potential 80 V (ESI+) and −80 V (ESI–). An information-dependent acquisition (IDA) strategy was used to identify lipids. The following parameters were used for MS: collision energy ±10 V, acquisition time 250 ms and for MS/MS, these were: collision energy ±45 V, collision energy spread 25 V, ion release delay 30, ion release width 14 and acquisition time 40 ms. The IDA switching criteria were set so that only a maximum number of 20 ions were taken into account, these being greater than m/z 300, exceeding 200 cps and excluding the former target for 2 s and the isotopes within 1.5 Da.
MS-DIAL (v5.2.240424.3) with the associated standard lipid library belonging to this version15,37 was used to align the data and identify the different lipids based on their MS/MS fragmentation pattern. The following parameters were applied: peaks with an intensity of at least 500 ions, eluting from 0.5 to 10 min in the chromatogram, were considered. Alignment within samples was performed using a retention time tolerance of 0.15 min and a MS tolerance of 0.025 Da.
Lipid identification was enhanced by the decision tree-based algorithm of MS-DIAL which integrates exact mass, adduct recognition and MS/MS spectral matching to improve annotation accuracy15. The following settings were applied: accurate mass tolerance of 0.01 Da for MS1 and 0.025 Da for MS2, including as adducts [M + H]+, [M+Na]+, [M + K]+, [M + NH4]+ and [M-H2O + H]+ in ESI+ and [M–H]–, [M+Cl]–, [M + HCOO]– and [M–H2O–H]– in ESI–. The following default identification thresholds were used: simple dot product score ≥600, weighted dot product score ≥600, reverse dot product score ≥600, matched spectrum percentage ≥0%, and a minimum number of one matched spectra. After obtaining a list of annotated lipids we performed three-step filtering. First, the lipid list was filtered to make sure that all identified lipids have at least a weighted dot product and a reversed dot product ≥600. Next, we kept those lipids having a signal at least 2 times greater than the blank signal in a minimum of 80% of the samples. Finally, we kept those lipids that had a relative standard deviation lower than 30% in the pooled sample (measured at regular intervals through the analytical run).
After automated identification and filtering, we manually curated all MS/MS spectra by retaining only lipids with matching experimental and reference spectra while considering lipid class retention time window and elution patterns. The resulting annotation level corresponds to the molecular lipid species level since the lipid subclass and fatty acyl composition are known but the sn-positions and double bond positions are unknown, e.g., PC 18:1_16:0 (following the guidelines of Liebisch on lipid shorthand notation). Finally, total area normalization of samples was performed prior to subsequent statistical analysis.
Materials for targeted oxidized PCs analysis
All reagents used were of LC-MS grade or higher. Non-oxidized PC standards included 1-palmitoyl-2-linoleoyl-sn-glycero-3-phosphocholine, 1-palmitoyl-2-arachidonoyl-sn-glycero-3-phosphocholine, 1-stearoyl-2-linoleoyl-sn-glycero-3-phosphocholine, 1-stearoyl-2-arachidonoyl-sn-glycero-3-phosphocholine, 1-palmitoyl-2-docosahexaenoyl-sn-glycero-3-phosphocholine, and 1-stearoyl-2-docosahexaenoyl-sn-glycero-3-phosphocholine were purchased from Avanti Polar Lipids. Commercially available oxidized phospholipid standards including 1-palmitoyl-2-(5-oxovaleroyl)-sn-glycero-3-phosphocholine, 1-palmitoyl-2-azelaoyl-sn-glycero-3-phosphocholine, 1-palmitoyl-2-(9-oxo)nonanoyl-sn-glycero-3-phosphocholine, and 1-palmitoyl-2-glutaroyl-sn-glycero-3-phosphocholine were from Avanti Polar Lipids while 1-(palmitoyl)−2-(5-keto-6-octene-dioyl)phosphatidylcholine and 1-palmitoyl-2-(4-keto-dodec-3-ene-dioyl)phosphatidylcholine were from Cayman Chemicals. Internal standards (ISs) 2-dinonanoyl-sn-glycero-3-phosphocholine (DNPC) and 1-heptadecanoyl-2-docosatetraenoyl-sn-glycero(d5)−3-phosphocholine (17:0-22:4 PC-d5) were from Avanti Polar Lipids.
Sample preparation of kidney tissue samples for targeted oxidized PCs analysis
Frozen kidneys were homogenized as described in “Sample preparation of kidney tissue samples for untargeted lipidomics analysis”. To 25 µL of kidney tissue homogenate (equal to 2.5 mg of frozen tissue), 20 µL of the IS mix (containing 100 ng/mL DNPC and 17:0-22:4 PC-d5 each in methanol), 500 µL LC-MS grade methanol and 500 µL chloroform were added. Samples were vortexed and 400 µL LC-MS grade water was added to the kidney tissue samples. The samples were then vortexed and centrifuged at 4000 × g for 10 min and 400 µL of the lower chloroform layer were transferred to a glass vial and evaporated under a gentle stream of N2. Dried samples were stored at −80 °C until the day of analysis. On the day of analysis, samples were reconstituted in 50 µL 2-propanol, vortexed and sonicated for 5 min. 50 µL of LC-MS grade water was added and samples were vortexed and sonicated for 5 min again. The samples were transferred to inserts and were ready to be analyzed by the LC-MS/MS targeted oxidized PCs platform. Same as in the untargeted lipidomics analysis, a QC was made by pooling equal aliquots of every single sample and was analyzed periodically during the analytical run.
Targeted LC-MS/MS method for oxidized PCs
10 µL of the lipid extracts were injected in a LC-MS/MS system composed of a Shimadzu LC-40 HPLC system coupled to a Sciex QTRAP 6500 + MS system in ESI+ multiple reaction monitoring (MRM) mode. Lipid separation was achieved using a Kinetex® C18 100 Å LC Column (50 × 2.1 mm, 1.7 μm) from Phenomenex, kept at 50 °C. Mobile phase A consisted of acetonitrile:water (50:50, v/v), and mobile phase B was 2-propanol:acetonitrile:water (85:10:5, v/v/v), both containing 5 mM ammonium formate and 0.1% formic acid delivered at a flow rate of 400 µL/min. The gradient elution was 10% B at 0.0 min, 25% B at 2.0 min, 30% B at 3.5 min, 75% B at 5.0 min, 77% B at 8.5 min, 100% B at 9.0 min, 100% B at 10.0 min, reverting to 10% B at 10.5 min and then allowed to re-equilibrate for 2 min. Data acquisition and integration were both performed using Sciex Analyst and Sciex OS software, respectively. The peak area ratios of lipids to the ISs were used for further analysis. The MRM transitions are detailed in Table S6. Other MS parameters include a curtain gas of 26 psi, ion source gas 1 and 2 set to 40 psi and 30 psi, respectively, a “low” collision gas, an ion spray voltage of 4500 V, a source temperature of 450 °C, and an entrance, collision cell exit declustering potential of 10 V, 15 V and 100 V, respectively.
Oxilipidomic analysis by LC-MS/MS on perfusate
Oxilipids were analyzed from 100 µL perfusate samples according to published protocols38. Briefly, to 100 µL perfusate was added 100 µL water, 600 µL methanol and an internal standard mix. Following acidification with formic acid, samples were cleaned up by solid phase extraction using C18 cartridges before being analyzed by LC-MS/MS in MRM mode on a Shimadzu Nexera series UHPLC system coupled to a Sciex 6500 QTrap.
Bulk RNA-sequencing
RNA was extracted from cryopreserved kidney biopsies using RNeasy Kits for RNA purification (Qiagen, 74106), following manufacturer’s instructions. The library was constructed by BGI Laboratories using the DNBSEQ Eukaryotic Strand-specific Transcriptome Resequencing protocol. Sequencing was performed on the DNBSEQ platform with PE150 or PE100. Trimmed FastQ files were provided by BGI. After which, sequence reads were mapped to the reference genome (GRCh38/hg38) and a count matrix was generated using Rsubread (v2.20.0). Count matrices were read into DEseq2, and size factors were estimated. Read counts were analyzed using DEseq2 (v1.46.0)39.
All bulk RNA sequencing analysis were performed using R (version 4.4.0). Gene set variation analysis (GSVA, v2.0.2) was performed for GOBP_Iron_Ion_Transport (n = 58 genes; GO:0006826, 2024.1.Hs), WP_Ferroptosis (n = 64 genes; WP4313, 2024.1.Hs), and FerrDb v2 validated driver, suppressor and marker genes with a protein product (n = 313 genes)18,40.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 9. We tested for Gaussian distribution by performing the Shapiro-Wilk normality test. When normal distribution was confirmed, we used an unpaired two-tailed t-test to compare two datasets and a one-way ANOVA to compare more than two datasets. In cases where multiple comparisons involved paired observations, we used a two-way ANOVA. Bonferroni’s correction was used when multiple comparisons were made. If the normality test indicated a non-Gaussian distribution, we used the Mann-Whitney U-test to compare two datasets and a Kruskal-Wallis test to compare more than two datasets. Dunn’s correction was used when multiple comparisons were made. For the untargeted lipidomics analysis, we performed unpaired t-tests with Welch’s correction. Correlation coefficients, where reported, are Pearson’s Rho for normally distributed or Spearman’s rank correlation for non-normally distributed variables. A p-value of 0.05 was used as threshold for statistical significance. Data are presented as mean ± SD, unless indicated otherwise.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All the raw untargeted lipidomics, targeted oxPC, and oxilipins data generated in this study have been deposited in Figshare at https://doi.org/10.6084/m9.figshare.29113910, https://doi.org/10.6084/m9.figshare.29113907, and https://doi.org/10.6084/m9.figshare.29113874. The processed untargeted lipidomics, targeted oxPC, and oxilipins data are provided as supplementary data files. The raw and normalized RNA-seq data are available from GEO under accession number GSE289688. Source data are provided with this paper.
Code availability
The code used for analysis is available via GitHub at https://github.com/RabelinkLab/deHaan_2025.
References
Kidney disease: a global health priority. Nat. Revi. Nephrol. 20, 421–423 (2024).
Elliott, T. R., Nicholson, M. L. & Hosgood, S. A. Normothermic kidney perfusion: an overview of protocols and strategies. Am. J. Transpl. 21, 1382–1390 (2021).
Parente, A. et al. Machine perfusion techniques for liver transplantation—a meta-analysis of the first seven randomized-controlled trials. J. Hepatol. 79, 1201–1213 (2023).
Croome, K. P. Should advanced perfusion be the standard of care for donation after circulatory death liver transplant?. Am. J. Transpl. 24, 1127–1131 (2024).
Lau, N.-S. et al. Long-term ex situ normothermic perfusion of human split livers for more than 1 week. Nat. Commun. 14, 4755 (2023).
Clavien, P. A. et al. Transplantation of a human liver following 3 days of ex situ normothermic preservation. Nat. Biotechnol. 40, 1610–1616 (2022).
Hosgood, S. A. et al. Normothermic machine perfusion versus static cold storage in donation after circulatory death kidney transplantation: a randomized controlled trial. Nat. Med 29, 1511–1519 (2023).
de Haan, M. J. A. et al. A cell-free nutrient-supplemented perfusate allows four-day ex vivo metabolic preservation of human kidneys. Nat. Commun. 15, 3818 (2024).
Arykbaeva, A. S. et al. Red blood cells as oxygen carrier during normothermic machine perfusion of kidney grafts: friend or foe?. Am. J. Transpl. 24, 1172–1179 (2024).
Van Avondt, K., Nur, E. & Zeerleder, S. Mechanisms of haemolysis-induced kidney injury. Nat. Rev. Nephrol. 15, 671–692 (2019).
Bayır, H., Dixon, S. J., Tyurina, Y. Y., Kellum, J. A. & Kagan, V. E. Ferroptotic mechanisms and therapeutic targeting of iron metabolism and lipid peroxidation in the kidney. Nat. Rev. Nephrol. 19, 315–336 (2023).
Eshmuminov, D. et al. An integrated perfusion machine preserves injured human livers for 1 week. Nat. Biotechnol. 38, 189–198 (2020).
Schuler, M. J. et al. Observations and findings during the development of a subnormothermic/normothermic long-term ex vivo liver perfusion machine. Artif. Organs 47, 317–329 (2023).
Zheng, J. & Conrad, M. Ferroptosis: when metabolism meets cell death. Physiol. Rev. 105, 651–706 (2025).
Tsugawa, H. et al. A lipidome atlas in MS-DIAL 4. Nat. Biotechnol. 38, 1159–1163 (2020).
Zhang, D. D. Ironing out the details of ferroptosis. Nat. Cell Biol. 26, 1386–1393(2024).
O’Donnell, V. B., Aldrovandi, M., Murphy, R. C. & Krönke, G. Enzymatically oxidized phospholipids assume center stage as essential regulators of innate immunity and cell death. Sci. Signal. 12, eaau2293 (2019).
Zhou, N. et al. FerrDb V2: update of the manually curated database of ferroptosis regulators and ferroptosis-disease associations. Nucleic Acids Res. 51, D571–d82 (2023).
Mori, K. et al. Endocytic delivery of lipocalin-siderophore-iron complex rescues the kidney from ischemia-reperfusion injury. J. Clin. Investig. 115, 610–621 (2005).
Wiernicki, B. et al. Excessive phospholipid peroxidation distinguishes ferroptosis from other cell death modes including pyroptosis. Cell Death Dis. 11, 922 (2020).
Balzer, M. S. et al. Single-cell analysis highlights differences in druggable pathways underlying adaptive or fibrotic kidney regeneration. Nat. Commun. 13, 4018 (2022).
Friedmann Angeli, J. P. et al. Inactivation of the ferroptosis regulator Gpx4 triggers acute renal failure in mice. Nat. Cell Biol. 16, 1180–1191 (2014).
Ide, S. et al. Ferroptotic stress promotes the accumulation of pro-inflammatory proximal tubular cells in maladaptive renal repair. Elife 10, e68603 (2021).
Nasralla, D. et al. A randomized trial of normothermic preservation in liver transplantation. Nature 557, 50–56 (2018).
Theurl, I. et al. On-demand erythrocyte disposal and iron recycling requires transient macrophages in the liver. Nat. Med. 22, 945–951 (2016).
Deuel, J. W. et al. Hemoglobinuria-related acute kidney injury is driven by intrarenal oxidative reactions triggering a heme toxicity response. Cell Death Dis. 7, e2064 (2016).
Zarjou, A. et al. Proximal tubule H-ferritin mediates iron trafficking in acute kidney injury. J. Clin. Investig. 123, 4423–4434 (2013).
Vermeulen Windsant, I. C. et al. Hemolysis is associated with acute kidney injury during major aortic surgery. Kidney Int. 77, 913–920 (2010).
Leaf, D. E. et al. Increased plasma catalytic iron in patients may mediate acute kidney injury and death following cardiac surgery. Kidney Int. 87, 1046–1054 (2015).
Lei, C. et al. Improving normothermic machine perfusion and blood transfusion through biocompatible blood silicification. Proc. Natl. Acad. Sci. USA 121, e2322418121 (2024).
Buehler, P. W., Humar, R. & Schaer, D. J. Haptoglobin Therapeutics and Compartmentalization of Cell-Free Hemoglobin Toxicity. Trends Mol. Med. 26, 683–697 (2020).
Andrijevic, D. et al. Cellular recovery after prolonged warm ischaemia of the whole body. Nature 608, 405–412 (2022).
Andrijevic, D. et al. Mechanisms and strategies for organ recovery. Nature Reviews Bioengineering. Nat. Rev. Bioeng. https://doi.org/10.1038/s44222-025-00293-7 (2025).
Ogurlu, B. et al. Utilizing pathophysiological concepts of ischemia-reperfusion injury to design renoprotective strategies and therapeutic interventions for normothermic ex vivo kidney perfusion. Am. J. Transpl. 24, 1110–1126 (2024).
Callaghan, C. J., Phillips, B. L., Foukaneli, T., Robinson, S. & Watson, C. J. E. The use of third-party packed red blood cells during ex situ normothermic machine perfusion of organs for transplantation: underappreciated complexities? Am. J. Transpl. 21, 1376–1381 (2021).
Maus, M. et al. Iron accumulation drives fibrosis, senescence and the senescence-associated secretory phenotype. Nat. Metab. 5, 2111–2130 (2023).
Tsugawa, H. et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Körner, A. et al. Inhibition of Δ24-dehydrocholesterol reductase activates pro-resolving lipid mediator biosynthesis and inflammation resolution. Proc. Natl. Acad. Sci. USA 116, 20623–20634 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Acknowledgments
We gratefully acknowledge the support provided by Rutger van Rooden (Transplant Center, LUMC, Leiden, the Netherlands) in making the 3D renderings of the organ chambers used for perfusion experiments, Mohan Ghorasaini and Kevin Brewster (Center of Proteomics and Metabolomics, LUMC, Leiden, the Netherlands) for their technical assistance with lipidomic analysis, and Manon Zuurmond (Department of Internal Medicine, LUMC, Leiden, the Netherlands) for making the illustrations. This work is supported by the Dutch Kidney Foundation through the Participants of the Friends Lottery (20INI011). The Novo Nordisk Foundation Center for Stem Cell Medicine (reNEW) is supported by Novo Nordisk Foundation grants (NNF21CC0073729). Ton J. Rabelink is funded by the European Union through ERC grant (SPARK 101140863).
Author information
Authors and Affiliations
Contributions
Conceptualization: M.dH., M.J., M.E., T.R.; Methodology: M.dH., M.J., E.S.L., M.G., M.E., T.R.; Software: R.S., R.D., E.S.L.; Validation: M.dH., M.J., A.dG, R.S., R.D., E.S.L., M.G., M.E., T.R.; Formal analysis: M.dH., M.J., R.S., R.D., J.K., E.S.L.; Investigation: M.dH., M.J., A.dG., R.D., E.S.L.; Resources: D.dV., J.K., I.A., C.vK., E.S.L., M.G., M.E., T.R.; Data curation: M.dH, M.J., R.S., R.D., E.S.L., M.G.; Writing—Original draft: M.dH., T.R.; Writing—Review and editing: M.dH., M.J., A.dG, R.S., R.D., D.dV., J.K., I.A., C.vK., E.S.L., M.G., M.E., T.R.; Visualization: M.dH., M.J., R.S.; Supervision: M.G., M.E., T.R.; Project administration: M.dH.; Funding acquisition: T.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tong Shen and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
de Haan, M.J.A., Jacobs, M.E., de Graaf, A.M.A. et al. Normothermic human kidney preservation drives iron accumulation and ferroptosis. Nat Commun 16, 5420 (2025). https://doi.org/10.1038/s41467-025-61058-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61058-9
This article is cited by
-
Opportunities and challenges with the implementation of normothermic machine perfusion in kidney transplantation
Nature Communications (2025)