Abstract
Brainstem gliomas (BSG) is a highly malignant central nervous system childhood tumors with 5-year survival rate <10%. Metabolism during radiotherapy is a dynamic and precisely programmed process, improving clinical outcomes and guiding therapy decisions of BSG. Here we construct diagnostic and prognostic assays of BSG via circulating metabolites based on both cross-sectional study and longitudinal cohort study with 106 BSG patients. We employ nanoparticle enhanced laser desorption/ionization mass spectrometry to characterize static and dynamic snapshots of metabolites during BSG radiotherapy. We show that this serological tool reaches the area under the curve of 0.933 for BSG diagnosis in an independent blind test and predicts risk of patients with significant differences (p < 0.05) in prognostic outcomes. We further identify eight distinct temporal patterns of metabolite regulation associated with radiotherapy responses and tracked the metabolic trajectory via dynamic metabolic snapshots throughout radiotherapy process. If further validated, this framework could be extended to derive comprehensive metabolic pictures for cancers including but not limited to BSG.
Similar content being viewed by others
Introduction
Brainstem gliomas (BSG), accounting for 10–15% of all childhood brain tumors, exhibit a peak incidence of 0.6 cases per million population annually within the age range of 6–9 years1,2,3. Patients diagnosed with BSG display a poor prognosis, with a median survival time of 10–12 months and a 5-year survival rate of less than 10%4,5. Radiotherapy is a standard BSG treatment in clinical settings, capable of alleviating symptom progression and prolonging progression-free survival by a few months6,7,8,9. Patients who undergo early and timely surgery exhibit prolonged overall survival (OS) time compared to those in late surgery group (median: 28.4 vs. 18.7 months)10,11,12. Accurate diagnosis, precise prognosis, and timely monitoring during radiotherapy are crucial in managing BSG patients. However, there are certain risks associated with biopsies, which involve sampling tissue for analysis, especially when dealing with tumors located in eloquent regions of the brain. Furthermore, biopsies hardly achieve dynamic monitoring via adequate sampling for effective therapy selection13,14.
Non-invasive radiologic imaging is the “one and only” technique to discover cancerous region before surgical intervention and evaluate tumor prognosis through timely monitoring. Nevertheless, imaging modalities afford an accuracy of only 65–73% in BSG detection due to the unsatisfactory image resolution (~1 mm), which could lead to the misdiagnosis of minimal lesions15,16. The limited precision of imaging techniques highlights an urgent need for incorporating liquid biopsy diagnostics14,17. Unlike many other solid tumors, BSG lacks clinically approved circulating biomarkers owing to the eloquent location of the brainstem within the intracranial region and the presence of the blood-brain barrier (BBB)18,19. In addition, the current collaborative technological efforts (e.g., circulating tumor DNA5 and cell-free DNA sequencing20) almost all fail in testing on peripheral blood due to the limited mutations in circulating systems.
In the context of BBB dysfunction, there may be transport-independent mechanisms for metabolites (size <1.8 nm) to cross the BBB. This is facilitated by the compatible pore size of brain microvessel endothelial cells, which typically range from about 1.4–1.8 nm, enabling metabolites to enter the bloodstream21,22,23,24. Metabolic alterations in BSG disrupt normal homeostatic processes due to tumor-related stress in the blood, leading to bioenergetic disturbances, endocrine imbalances, and oxidative stress. BSG-derived metabolism may synergistically promote adverse clinical outcomes. Our previous study has validated the function of BBB in separating blood and brain/cerebrospinal fluid (CSF) systems, as well as in regulating the permeability that allows for metabolite transport under disease status25. The feasibility of metabolic profiling has also been explored to understand the molecular pathogenesis of other tumors in the brain, such as glioblastoma, melanoma brain metastases26, and neuroblastoma27. Serum metabolite profiles, including static and dynamic metabolic snapshots, may provide an alternative for capturing critical metabolic information about BSG and enable the tracking of tumor dynamics via metabolic endpoints throughout the complete course of a patient’s radiotherapy.
Mass spectrometry (MS) is applied in the profiling and discovery of metabolic biomarkers by detecting the mass-to-charge ratio (m/z) of metabolites at a high resolution at ppm level. Recently, nanoparticle-enhanced laser desorption/ionization MS (NPELDI-MS) has been developed for capturing metabolic static and dynamic snapshots. This technique features few pretreatment operations, fast detection speed, and trace sample consumption28,29,30, making it ideal for scenarios where expeditious reporting is critical31,32,33. In particular, tailored inorganic nanoparticles have been introduced as chromatography alternatives prior to MS analysis to selectively enrich metabolites, which enhances its applicability for biofluid analysis and biomarker detection.
Molecular characteristics of BSG remain poorly characterized due to the relatively low incidence rate. Here, we constructed a cross-sectional and longitudinal cohort of pediatric BSG, ranging from 1 to 15 years old, via both static and dynamic blood sampling (Fig. 1a–c). We performed NPELDI-MS to acquire a more comprehensive metabolic picture of BSG. Based on the static metabolic snapshots, we distinguished BSG patients from healthy donors (HDs) and further predicted patient survival. Furthermore, we identified eight distinct temporal patterns of metabolite regulation corresponding to different BSG subsets associated with radiotherapy responses. We monitored the dynamic disturbances of key metabolites during the course of therapy. Our study established an approach to reveal mechanisms and heterogeneity of diseases, including but not limited to brain tumors at metabolic level.
a Serum samples were enrolled from BSG patients and healthy donors (HDs) on micro-arrayed chips, for nanoparticle-enhanced laser desorption/ionization mass spectrometry (NPELDI-MS) analysis. b Machine learning of serum metabolic profiles was conducted to obtain the biomarker panel to achieve the diagnosis, prognosis, and dynamic monitoring of BSG. c Clinical cohorts were collected to acquire the static and dynamic metabolic snapshots. d Study design of participant enrollment, including 125 HDs and 174 BSG patients. e Longitudinal sample collection of 62 BSG patients at three time points as the dynamic metabolic snapshots. Created in BioRender. Yang, Y. (2025) https://BioRender.com/uo0yzh1.
Results
Study design and patient cohort characteristics
We first applied an analytic framework with a case-control design on the cross-sectional cohort (cohort 1) to diagnose BSG from the aspects of static metabolic snapshots. We successively prognosed the radiotherapy sensitivity among BSG patients (cohort 2) via univariant and multivariant Cox regression and validated the performance by log-rank test, including 96 BSG patients with a follow-up period of more than 1 year as prognostic outcomes. Lastly, we sought progressive metabolic evolution correlated with radiotherapy responses by clustering of metabolic trajectories with dynamic metabolic snapshots on the longitudinal cohort (cohort 3). Demographic and clinical characteristics of the enrolled participants are presented in Supplementary Tables 1 and 2.
We established a Chinese BSG cohort within a duration of 6 years (from November 2017 to May 2023) at Xinhua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine. The cohort comprised 174 BSG patients and 125 HDs, from which we enrolled 106 BSG patients satisfying the following criteria. These criteria included (1) BSG diagnosed by magnetic resonance imaging (MRI) presentation and clinical symptoms, (2) completion of radiotherapy, and (3) a follow-up period of more than a year (Fig. 1d). Given the complexities and inherent risks associated with performing biopsies in brainstem tumors, particularly in cases of diffuse intrinsic pontine glioma, obtaining WHO grades for all patients present challenges, especially before 202034. To ensure the accuracy of patient enrollment and diagnosis as BSG cases, the diagnosis of each patient was collaboratively reviewed by 2 oncologists and imaging specialists. If conflicting opinions arose, senior physicians were consulted to avoid any potential biases in data evaluation. To capture the dynamic metabolic snapshots at high resolution, we performed a design of high-density blood sampling on the longitudinal cohort consisting of 62 BSG patients. Consenting participants submitted three-weekly blood draws prior to radiotherapy, until the completion of one cycle of radiotherapy (three serum samples collected for each participant, Fig. 1e).
To evaluate BSG-related metabolic snapshots, we employed a pretreatment-free MS approach that incorporated ferric nanoparticles to substitute the commonly used liquid/gas phase extraction and separation. The NPELDI-MS allowed sample multiplexing with reproducible metabolic profiles for native serum inputs as trace as 0.1 μL (Supplementary Fig. 1a, coefficient of variation <15%). In particular, a rapid analytical speed of ~2 s per sample (with 2000 laser shots at a pulse frequency of 1000 Hz) and throughput of 384 samples per batch could be accomplished by the NPELDI-MS platform (Supplementary Fig. 1b). As a result, detection process of 405 serum samples in this project was completed within ~3 h.
The raw MS spectrum recorded ~120,000 data points with a total ion count of ~7 × 107 within the mass range of 100–1000 Da. After quality control and data filtering (see details in Supporting Information), we obtained 402 metabolic features (defined as the localized highest intensity of the raw MS data) across different samples (Supplementary Fig. 1c). Among these, 287 features (71.4%) varied from HDs to BSG patients (p < 0.05, Student’s t-test). Notably, the serum metabolic snapshot database, which contains a sufficient amount of signal intensity and feature number, serves as the foundation for the downstream analysis.
Diagnostic value of circulating metabolites in BSG patients
Given that metabolic reprogramming is crucial for cancer occurrence35,36,37, we were thus interested in delineating the distinct metabolic patterns between HDs and BSG based on the static metabolic snapshots (Supplementary Fig. 2a). We built diagnostic classification models based on machine learning algorithms on the discovery set (n = 45/54, HDs/BSG). The computational pipeline for machine learning of serum metabolic profiles from the sets included feature selection and model building (Fig. 2a, Supplementary Fig. 2b). Specifically, there were no significant differences in age and gender (p > 0.05) between BSG and HDs in the discovery set (Supplementary Table 3). The classification performance was further evaluated on an independent validation set (n = 15/19, HDs/BSG). Notably, we obtained 81.2% power at 0.05 significance level with a sample size of 80 (40/40, HDs/BSG), based on the preliminary study on 12 individuals (6/6, HDs/BSG, Supplementary Fig. 2c). The result indicated the discovery and validation sets hold adequate sample size for validating the classification models.
a Study design for BSG diagnosis using machine learning. Static metabolic snapshots from 99 participants (45/54, HDs/BSG) in the discovery set for feature selection and model building. The optimized model was tested in an independent validation set (15/19, HDs/BSG). b Coefficients of the least absolute shrinkage and selection operator (LASSO) and Ridge algorithms were shown as the evaluation index for seven metabolic features. c t-distributed stochastic neighbor embedding of serum metabolic profiles for 60 HDs and 73 BSG patients was plotted using identified metabolites. d Receiver operating characteristic (ROC) curves of the discovery set and validation set to distinguish HDs and BSG patients within the metabolic panel. The corresponding confusion matrices were shown as the inset. e ROC curves analysis was shown with each single feature (AUC of 0.662–0.818) of seven-feature diagnostic panel. f Violin-plot illustrated the differential regulation of normalized intensities for metabolic features in HDs and BSG. Statistical significance was calculated using two-sided Student’s t-test. ***p < 0.005, *p < 0.05. p = 5.224 × 10−8 (LA); 0.004 (LA’); 4.446 × 10−5 (Val); 4.944 × 10−8 (Leu); 4.422 × 10−5 (Thr); 0.0008 (2AAS); 0.025 (Orn). g Potential pathways were differentially regulated in HDs and BSG patients. Color and size of each circle were correlated to the p value and enrichment ratio. Statistical significance was calculated using enrichment analysis by MetaboAnalyst 5.0 (https://www.metaboanalyst.ca/). LA and LA’ lactic acid, Val valine, Leu leucine, Thr threonine, 2AAS 2-aminomuconic acid semialdehyde, Orn ornithine. Source data are provided as a Source Data file.
In the feature selection step, we sought to utilize dimension-reduction strategies to classify BSG versus HDs based on the outstanding features with more cost-effective combinations of the molecules (Supplementary Fig. 3a, b, Supplementary Table 4). Considering the p (number of predictors) » n (number of samples) problems, two widely used shrinkage methods, least absolute shrinkage and selection operator (LASSO) and Ridge, were employed to select the optimal biomarker combination based on the highest coefficient weights in two algorithms (Fig. 2b). As a result, seven metabolic features corresponding to six metabolites were identified as highly ranked biomarkers (mean intensity > 200, p value < 0.05, and coefficient weight > 0.18), including 2-aminomuconic acid semialdehyde, lactic acid, valine, leucine, threonine, and ornithine (Supplementary Table 5)38,39. Unsupervised clustering analysis partly intuitively distinguished BSG from HDs, including t-distributed stochastic neighbor embedding (Fig. 2c) and principal component analysis (Supplementary Fig. 2d).
We further conducted in-vitro studies to investigate the biological effects of metabolite biomarkers on SHSMU-DIPG-01 cells. Our findings revealed concentration-dependent effects of these metabolites on DIPG cell proliferation, as validated by the cell counting kit-8 assay (p < 0.05, Supplementary Fig. 4). These results suggest that these metabolites play a regulatory role in DIPG cell behavior, potentially by modulating metabolic pathways or inducing metabolic stress. This aligns with the broader impact of metabolic reprogramming on tumor development and progression37. The sensitive responses of DIPG cells to these metabolites further support their potential as biomarkers for distinguishing BSG from healthy individuals.
In the model-building step, four commonly used algorithms, including LASSO, Ridge, K-nearest neighbor (kNN), and random forest (RF), were used to construct classification models on the metabolite biomarker combinations. The optimized model by RF reached the highest area under the curve (AUC) of 0.923 (95% confidence interval (CI) = 0.870–0.976) based on the receiver operating characteristic curve (ROC) analysis in the discovery set. We then tested the model on the validation set, resulting in a promising AUC of 0.933 (95% CI = 0.853–1.000) for distinguishing BSG patients from HDs (Fig. 2d, Supplementary Fig. 5a, b, Supplementary Table 4). We recruited a new cohort (Supplementary Table 2) from Shanghai Xinhua Hospital, with 161 samples collected (96/65, BSG/HDs) as the external set. Notably, the external set was independent from the discovery and validation sets in model construction stage and blinding to the as-built model (Supplementary Fig. 6a). The situations for blood drawn were the same for all sample sets. We obtained the AUC of 0.847–0.852 (Supplementary Fig. 6b), consistent with the previous results in the methodology development. The consistent diagnostic results validated the robustness of the biomarkers, facilitating translation of the biomarkers to larger patient populations.
An evaluation of the reliability of the machine-learning strategy was conducted using confusion matrices, which demonstrated that samples were classified with a high accuracy of 86.9% in discovery set and 88.2% in validation set (Fig. 2d). Importantly, the panel constructed of seven metabolic features together outperformed the diagnostic performance by single one of these features (AUC: 0.662–0.818, Fig. 2e, Supplementary Table 5). Another major challenge in clinical practice is the differentiation of BSG from other neural diseases by MRI imaging40. We added the comparison between BSG and other neural diseases (28 encephalitis cases and 40 idiopathic epilepsy cases) through the static metabolic snapshots. We adopted the same protocol to detect the differences (Supplementary Fig. 7a). As a result, the models also achieved outstanding performance in differentiating BSG from encephalitis and idiopathic epilepsy, with AUC of 0.929 and 0.923, respectively (Supplementary Fig. 7b).
Specifically, five features were over-regulated (p value < 0.05) in BSG patients, while two features were down-regulated (p value < 0.05) with BSG diagnosis (Fig. 2f). Furthermore, metabolite set enrichment analysis showed several BSG-disturbed metabolic pathways based on Small Molecule Pathway Database (SMPDB) dataset, such as valine, leucine, and isoleucine degradation, glycine and serine metabolism, threonine and 2-oxobutanoate degradation, propanoate metabolism, as compared to HDs (Fig. 2g, Supplementary Table 6). Subsequently, we conducted metabolite set enrichment analysis using Kyoto Encyclopedia of Genes and Genomes (KEGG) dataset to identify the most strongly influenced metabolic pathways using the identified metabolites. Our findings revealed that the metabolites were notably enriched in six distinct metabolic pathways, aligning with the results obtained from the SMPDB dataset (Supplementary Fig. 8).
To further investigate the biological mechanisms underlying the differences between BSG and HDs, we performed differential expression analysis using RNAseq data obtained from orthotopically xenografted BSG specimens (brainstem tissues, n = 4) and age-matched controls (normal brainstem tissues, n = 3, Supplementary Fig. 9). We also validated the correlation based on the analysis of the GEO dataset GSE162976, comparing DIPG cells against two distinct neural progenitor controls: human neural stem cells (hNSC) and normal human astrocytes (NHA; Supplementary Fig. 10). As a result, we detected notable variances in gene expression within the perturbed metabolic pathways, which were reflected in the RNAseq expression data obtained in this study and further supported by the consistent results obtained from GEO database.
Moreover, our exploration revealed the regulation of specific genes associated with the adaptive reprogramming of tumor cells. Notably, the genes Arg1, Arg2, Nos2, and P4ha1 were found to reprogram arginine and proline metabolism. Meanwhile, Arg1, Arg2, and Nos2 were shown to disturb arginine biosynthesis. Arginine was converted into ornithine through the action of arginase. Additionally, the genes Hpgds and Nat8f6 were identified as key regulators in glutathione metabolism. Gldc and Pgam2 were also found to influence the metabolism of glycine, serine, and threonine. Furthermore, Enpp1 and Enpp3, which encode two critical enzymatic components of the pantothenate and CoA biosynthesis pathway, were shown to be epigenetically regulated in brain tumors. These modulations closely related to the Warburg effect and the TCA cycle, and regulate the expression of lactate. Lastly, Aox4 was found to regulate tyrosine metabolism as well as the degradation of valine, leucine, and isoleucine, in conjunction with Hmgcs2. Therefore, we speculated that the unique metabolic signature arises from the metabolic rewiring of brain tumors, potentially offering valuable insights for BSG.
Taken together, these findings suggest a unique metabolic signature existed in BSG and demonstrated the power of metabolic profiling for biomarker discovery. The panel constructed by circulating metabolites facilitates rapid detection of BSG more precise and accessible.
Static metabolic snapshots correlated with the prediction of BSG
We proceeded to investigate the potency of using static metabolic snapshots in cohort 2 to predict prognosis of BSG. To determine the clinical outcome of this study, we enrolled patients who had been followed up for over one year and collected survival data as well as clinical information (including gender and age). A total of 96 patients were included in the prognosis study with OS being 4 months to 51 months and a median OS being 11 months. Importantly, clinic follow-ups of these BSG patients were carried out via phone calls or health records to ensure the completeness of data (Fig. 3a). Similar to the diagnostic section, we analyzed the static metabolic snapshots from two randomly assigned sample sets, containing 58 observations with 41 deaths for training set and 38 observations with 31 deaths for a testing set (Fig. 3b).
a Clinical information of 96 BSG patients. Created in BioRender. Yang, Y.(2025) https://BioRender.com/uo0yzh1. b Workflow for building the prognostic model via seven-feature panel by calculating risk scores based on static metabolic snapshots. c Plots of overall survival (OS) duration and risk score of each patient. The vertical dash line demonstrated the median score of −2.7 and stratifies patients into high-risk group (red) and low-risk group (blue) in training set (green) and testing set (blue). d Kaplan–Meier curves of OS of patients in the low-risk group (blue line) and high-risk group (red line) with p = 0.031 by log-rank test in the training set and p = 0.044 by log-rank test in the independent testing set. Source data are provided as a Source Data file.
We conducted a variable selection on the training set and built the composite risk score on the testing set (Fig. 3b). In particular, the univariate Cox regression was applied to reduce dimensionality and select metabolic features associated with the OS of BSG. Then, the multivariant Cox regression model was constructed to predict prognosis associated with metabolic vulnerabilities. Importantly, we identified a seven-feature panel correlated with clinical outcomes, consisting of lactic acid, oxalacetic acid, niacinamide, threonic acid, succinic acid, arginine, and glucose (Supplementary Table 7). Importantly, even after adjusting for key clinical covariates such as age, gender, BMI, and Lansky score, as suggested by previous studies41, the odds ratios of risk scores remained nearly unchanged (Supplementary Table 8).
Our in vitro experiments demonstrated the specific effects of metabolic markers on cell viability in SHSMU-DIPG-01 models (Supplementary Fig. 11). In particular, we have already validated lactic acid in the diagnostic panel. Hence, its presence in the prognostic panel does not warrant further investigation. Besides, arginine, succinic acid, and lactic acid exhibited significant anti-proliferative effects on DIPG cells (p < 0.05). Glucose and niacinamide played a role in promoting cell growth (p < 0.05). These findings highlight the relevance of these metabolites in modulating tumor cell behavior.
The prognostic panel consisted of differentially regulated metabolites together identified 14 potentially altered metabolic pathways (Supplementary Fig. 12a). We thus conducted RNAseq experiments on subcutaneous xenografted BSG specimens after radiotherapy (tumor tissues, n = 5) and age-matched BSG specimens before radiotherapy (tumor tissues, n = 6). We highlighted seven differentially expressed genes (DEGs) (ALDOC, CA9, FBP1, HKDC1, LOC102724560, NDUFA4L2, and P4HA2), associated with the metabolic pathways identified in the prognostic panel and expression data (Supplementary Fig. 12b). In particular, glycolysis/gluconeogenesis appeared enriched in tumor cells. Tumor growth is characterized by the upregulation of ALDOC, FBP1, and HKDC1 correlated with increased glucose and fructose consumption and lactate production42. Similarly, amino acid metabolism, regulated by P4HA2 and LOC102724560, a significant hallmark of glioblastoma impacting patient prognosis, has been confirmed through our metabolic and sequencing results and supported by existing literature43,44.
To mechanistically characterize the pathophysiological divergence between before RT (BRT) and after RT (ART) among BSG, we identified six significantly enriched pathways correlated with the prognostic metabolic signature in BSG through gene set enrichment analysis (Supplementary Fig. 12c). Enriched pathways involve TNF signaling, NF-κB signaling, p53 signaling, HIF-1 signaling, hippo signaling, and MAPK signaling, which have been demonstrated to impact on the sensitivity of tumors to radiotherapy45,46. Intensive studies indicate that the prognostic panel of metabolites could be involved in the regulation of these signaling pathways, which have the potential to influence radiation prognosis. Specifically, lactate enhances the stability of HIF-1α, which promotes its transcriptional activity and subsequently regulates tumor growth and sensitivity to RT47. Glucose stress induces O-GlcNAcylation and further activates hippo-YAP pathways to promote tumorigenesis48. Niacinamide activates p53 activation and suppresses PARP1 to mediate DNA repair and cell proliferation in tumor49. Succinic acid promotes not only cancer cell migration and invasion but also cancer metastasis, mediated by PI3K-HIF-1α axis50. Arginine activates NF-κB signaling by regulating mTORC1-mediated lysosomal release of essential amino acids. It also induces the generation of intracellular nitric oxide, which stimulates HIF-1α signaling and further promotes tumor proliferation51. Threonine metabolism influences tumor cell proliferation by driving abnormal tRNA modifications and protein translation within tumors52. Thus, we conclude that the prognostic panel reflects the rewired metabolism of tumors.
The risk score for every patient in both the training set and the testing set was estimated based on the formula, by summing up the product of each feature intensity and coefficient weight. Based on the dichotomized label on the median risk score, the BSG patients were stratified into two groups: the high-risk group and the low-risk group. Notably, the high-risk group (risk score > −2.7) contained 29 observations with 24 events in the training set and 14 observations with 14 events in the testing set. Meanwhile, there were 29 observations with 17 events in the training set and 24 observations with 16 events in the testing set among the low-risk group (risk score < −2.7, Fig. 3c). Multivariate Cox regression analysis demonstrated that the risk score tightly correlated with mortality risk and became an important factor of survival in both the training and testing sets.
We compared the OS duration of BSG patients. Kaplan-Meier curves were applied to evaluate the prognostic prediction efficiency of the risk score. Accordingly, the median OS duration in the low-risk group was significantly more prolonged than the OS duration in the high-risk group (p = 0.031 in the training set and p = 0.044 in the testing set, by log-rank test, Fig. 3d). In this study, we also collected the information of molecular subtypes and MRI imaging, and further explored the prognostic effectiveness of H3K27M mutation (p = 0.004, log-rank test, Supplementary Fig. 13a), TP53 mutation (p = 0.554, log-rank test, Supplementary Fig. 13b), and ring enhancement (p = 0.106, log-rank test, Supplementary Fig. 13c). These results suggested higher levels of risk score acquired from static metabolic snapshots are correlated with significant or borderline significant worse patient outcomes. Notably, we validated the prognosis efficiency of static metabolic snapshots in predicting prognostic risks of BSG patients.
Dynamic metabolic snapshots during radiotherapy
Circulating metabolites undergo dynamic changes during radiotherapy and are closely associated with the therapeutic efficacy of this treatment modality53,54. We examined the dynamic metabolic snapshots with a time series to evaluate the connection with the radiotherapy prognosis. Specifically, the metabolic profiles of 43 BSG patients were investigated at a total of three time points before, during, and after radiotherapy (BRT/DRT/ART, Fig. 4a). Corresponding MRI images were also collected for all enrolled patients to facilitate comprehensive analysis and interpretation of the data.
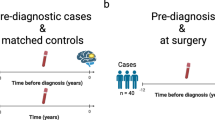
a Sample information collected for 43 BSG participants. Created in BioRender. Yang, Y.(2025) https://BioRender.com/uo0yzh1. b Calculation of the change in tumor size based on magnetic resonance imaging (MRI) images. c ROC curves to distinguish volume-decreasing participants from volume-increasing participants using the static metabolic snapshots collected BRT. d Clustering of metabolic trajectories using dynamic metabolic snapshots along HDs, BRT, DRT, and ART. e Probabilities of six biomarkers tracing with potential clustering tendency. f Scatter plots of six metabolites constructed the diagnostic panel among HDs (black), BRT (red), DRT (blue), and ART (green). Orange lines referred to the average intensity of the metabolites in each group. Statistical significance was calculated using pairwise comparison via multiple two-sided t test. g Dynamic alterations of lactic acid in low-risk (blue) and high-risk (red) groups. Statistical significance was calculated using two-sided Student’s t-test. *p < 0.05: p = 3.686 × 10−7. n.s. p > 0.05. Source data are provided as a Source Data file.
Clinically, difference in tumor volume size measured by MRI is an evaluation criterion of radiotherapy55,56. Based on the tendency of tumor change shown from MRI images (Fig. 4b), we achieved a good predictive capability in distinguishing volume-decreasing participants from volume-increasing participants based on the BRT metabolic profiles, with an AUC of 0.818 by 5-fold cross validation (95% CI: 0.690–0.946, Fig. 4c). The results demonstrated consistency between metabolic profiles and actual tumor volume changes.
Our samples were mainly from the HDs and BSG (as cohort 3) in the different periods of radiotherapy, including 117 HDs, 101 BRT, 85 DRT, and 85 ART, revealed detailed temporal patterns of metabolic changes. Interestingly, we found that metabolites with uni-directional behaviors dominated the features, and over half of them were significantly different between HDs and disease groups, as well as made a change in the development of radiotherapy (Supplementary Fig. 14a). These findings highlighted that even though the amount of each compound dynamically changes between HDs and BSG during radiotherapy, a highly coordinated metabolite regulatory network underlies the therapy process.
We further performed c-means clustering using the differential metabolites among four phases (HDs, BRT, DRT, and ART) and identified eight distinct clusters (Fig. 4d and Supplementary Fig. 14b). Notably, 23.1% of all metabolites in cluster 1 and cluster 2 showed a marked alteration in BSG patients from HDs and a stepwise increase or decrease during radiotherapy. Metabolites in cluster 3 showed an increase and achieved the top in DRT, then a large decrease afterward, whereas metabolites in cluster 4 had a marked increase in disease groups and a large decrease with radiotherapy, suggesting that unique metabolic vulnerabilities may appear in disease groups which could be exploited to monitoring the therapeutic effects of radiotherapy.
According to the diagnostic biomarker panel, we collected the probabilities of these biomarkers tracing in the potential cluster tendency (Fig. 4e). In particular, the 2-aminomuconic acid semialdehyde approached cluster 1 and gradually increased deviated the normal levels. Valine and threonine from cluster 2 were opposite of the tendency in cluster 1. Lactic acid and ornithine were included in cluster 3 and largely influenced by the radiotherapy process. Leucine from cluster 4 had no difference during the radiotherapy process (Fig. 4f). Importantly, we found lactic acid involved in pyruvate metabolism had different tendencies along with radiotherapy progression in low-risk and high-risk groups within the longitudinal cohorts of 62 BSG participants (Fig. 4g).
We further performed preclinical animal study using mouse tumor models to acquire paired samples (i.e., mice brain tumor and mice serum before and after radiation) with the same analysis from discovery set. The preclinical animal study also indicated significant alterations in metabolite levels before and after radiotherapy treatment in BSG, aligning with observations in paired serum samples (Supplementary Fig. 15).
Taken together, these findings depicted a programmed metabolic trajectory over the course of radiotherapy, providing a systematic value for treatment response and prognostic implications.
Discussion
Precise diagnosis and prognosis contribute to improving clinical outcomes and guiding therapy decisions of BSG patients, highlighting an urgent need to gain deep mechanistic insights into the comprehensive alterations associated with tumor occurrence and responses. Metabolic phenotypes have been reported to be involved in metabolism regulation within cancer progression. Stem-like glioma cells alter their metabolic process in order to sustain self-renewal and provide energy for the tumor hierarchy, uncovering potential cancer vulnerabilities that can be targeted with personalized therapy. In this study, we captured the static metabolic snapshots in BSG from HDs, as well as tracked the metabolic trajectory via dynamic metabolic snapshots throughout radiotherapy process.
A highly detailed metabolic picture of a cross-sectional cohort with specific outcomes at a given point in time can provide a static view of that sample and aid in disease diagnosis and prognosis. We first built a diagnostic model using six selected metabolic biomarkers. We found that this model could accurately and invasively discriminate BSG patients from HDs with higher accuracy of 88.2% with AUC of 0.933, sensitivity of 94.7%, and specificity of 80%, compared with the accuracy of 82.5% of CSF ctDNA samples and 38% of plasma ctDNA5. The accuracy of this model for BSG diagnosis was also superior to those acquired from the traditional MRI information (accuracy of ~76%)57. While it is acknowledged that the presence of BSG would typically be confirmed by imaging modalities, such as MRI, it is essential to consider the global healthcare landscape, particularly in low-income countries where imaging facilities may not be uniformly available. The World Health Organization reports that less than 30% of low-income countries have adequate imaging facilities, leading to a substantial number of undiagnosed cases and delayed interventions58. BSG, especially in pediatric cases, exhibits atypical clinical presentations that include subtle mental signs like emotional and behavioral changes or acquired torticollis59,60. These symptoms can be easily missed or misinterpreted, resulting in delays in diagnosis and treatment initiation61. Given the aggressive nature of BSG and its rapid progression, delays in the commencement of essential treatments, such as radiotherapy, can significantly impact patient outcomes and reduce survival periods.
Subsequently, a prognostic prediction model was developed by employing a seven-feature panel. With this model, the calculated risk score successfully differentiated patients of BSG based on their respective prognoses, and the validity as an independent prognostic risk factor was confirmed through log-rank test. When compared to other prognostic risk factors (MRI imaging ring enhancement62 and genotyping15,63,64), the discrimination potential of the score was found to be superior (Supplementary Fig. 13a–c). Significantly, the H3K27M mutation demonstrated superior distinguishing ability between high- and low-risk groups compared to the risk scores via static metabolic snapshots (p = 0.031, log-rank test, Fig. 3d). Unlike molecular testing on tissue biopsies, risk scores from static metabolic snapshots were derived from serum samples, facilitating treatment monitoring through serial liquid biopsy. Furthermore, metabolic profiling using NPELDI MS technology offers rapid and cost-effective analysis compared to sequencing methodologies. Overall, regular monitoring of metabolic snapshots allows for prompt assessment of the patients’ treatment effectiveness and provides valuable support in clinical decision-making processes.
Notably, both diagnostic and prognostic assays were built on a NPELDI-MS platform (see workflow in Supplementary Fig. 16), featuring little sample pretreatment (~min) and fast analytical speed (2 s per sample) for metabolic profiling of tracing serum sample (down to 100 nL). The typical platforms require complex derivatization or chromatographic separation prior to MS detection, to address the sample complexity and enrich the target metabolites from clinical biofluids (usually consuming 50–200 μL). The key is to introduce nanoparticles to selectively enhance the desorption and ionization efficiency of metabolites. Notably, coupled with throughput of 384 samples per chip can be achieved by microarray design automatically, indicating application potency for biomarker discovery especially suitable for high-throughput cohorts.
While radiotherapy is the standard treatment for BSG, chemotherapy is often utilized in combination with radiotherapy for other brain tumors like GBMs. The metabolism of tumors is expected to undergo changes based on the chemoradiation regimens employed65. Importantly, we have previously observed significant alterations in carbohydrates and amino acids during the chemotherapy process for various cancer types such as gastric cancer, pancreatic cancer, and breast cancer29,66. Thus, it is feasible to extend this framework to incorporate the combined use of chemotherapy and radiation in brain tumor treatment.
The alteration in tumor volume directly reflects the growth and progression of the tumor. Metabolic reprogramming of tumor cells plays a critical role in the initiation, proliferation, and advancement of cancer. The behavior of cancer cells is regulated by both intrinsic cellular metabolism and the availability of metabolites in the tumor microenvironment67. Therefore, BRT metabolic profiles can serve as indicators of the growth and development status of tumors, enabling the prediction of tumor volume changes (Fig. 4c), a concept that is further supported by existing literature68,69.
Static snapshots of the blood provide accurate diagnosis and prognosis, monitoring measurements longitudinally enables the most precise evaluation of metabolic transitions throughout therapy. Although previous molecular profiling studies adopted a cross-sectional approach, the challenge remains in understanding how the circulating metabolites as a whole change, given the metabolic heterogeneity among different individuals70,71,72. Relatively few studies have assessed the whole metabolites at many time points, hardly predicting the dynamic alteration through snapshots of its current state. Several studies have combined untargeted metabolomics and high-density sampling to develop prediction models for specific activities within sampling time points, such as peak oxygen consumption during acute physical activity or delivery time73,74. Dynamic metabolic profiling for predicting follow-up activity is essential for better healthcare, but remains unexplored, for instance in predicting regrowth rhythmicity of remaining cancer cells after therapy75,76.
The use of serum samples as a less invasive alternative to traditional pathological molecular testing offers promising implications for the diagnosis and prognosis of tumors, particularly in challenging-to-biopsy brain tumors such as BSG. The non-invasive nature of serum metabolic testing yields benefits such as reduced patient anxiety, fewer postoperative complications, and decreased physical and mental burdens, fostering a positive doctor-patient relationship. Regular monitoring of metabolite levels enables a timely assessment of the patient’s condition and treatment efficacy. This approach allows for personalized treatment adjustments and the development of individualized dietary plans, leading to improved patient care and outcomes.
Thanks to the non-invasive sampling and facile operation, our assay was further expanded to determine if serial sampling of serum could be used to monitor tumor metabolic dynamics by tracking response to therapy. Observational studies utilizing serology tests require a well-designed blood sampling interval that bridges the appropriate molecular assays with the outcomes of interest. For instance, weekly blood sampling during pregnancy reflects rapid metabolic changes in pregnant women74. Our designed scheme entailed a dense sequential blood sampling interval of three weeks, which is different from previous cohorts that had annual blood sampling for mortality evaluation. Eight metabolic clusters were determined with different metabolism patterns leading to distinct survival outcomes. As a proof-of-concept demonstration in clinical use, the actual OS can be predicted via static metabolic profiles, allowing the regimen to be adjusted timely via dynamic metabolic monitoring. This longitudinal cohort study depicted a programmed metabolic trajectory over the course of radiotherapy, filling the clinical gap related to cancer’s poor prognosis, including but not limited to BSG.
Strikingly, we also performed further comparisons between diagnostic and prognostic panels to demonstrate the metabolic differences and similarities among HDs and BSG groups. Diagnostic biomarkers here were singular parameters that cannot be used to directly predict the probability associated with tumor development, due to the metabolic preferences with a tumor change during cancer progression77,78,79. For metabolic differences, we identified two new metabolic panels, one for distinguishing the BSG from HDs and another for classifying the high-risk and low-risk groups by predicting the OS duration. The altered valine, leucine, and isoleucine degradation, and glycine and serine metabolism agree with previous reports in BSG diagnosis80,81. In the development of peripheral neuropathy, changes in valine, serine, and glycine amounts are associated with occurrence of brain tumors82,83,84. For metabolic similarities, we identified one overlapping metabolite that was differentially regulated. Lactic acid, as the product of the Warburg effect, was considered as the diagnostic biomarker reflecting tissue necrosis in high-grade brain tumors85,86,87, and was also evaluated as a poor prognostic factor in brain tumor progression88.
Nevertheless, this study still has its limitations. Firstly, the enrolled cohort consisted of the Chinese population only, with small variations in clinical characteristics. In particular, the clinical follow-up is relatively short, with a median follow-up duration of 11 months. In the future, we need to explore the generalizability of assays in a larger cohort with diverse ethnicities and complex complications, as well as longer clinical surveillance. Although this work delineates the concentration-dependent effects of specific metabolites on tumor cell proliferation dynamics, additional biological validation is essential to confirm the metabolic regulatory networks involved in cellular nutrient uptake or receptor-mediated signaling pathways. Meanwhile, we only performed untargeted metabolic profiling to discover biomarkers related to BSG occurrence and therapy response. Targeted analytical assays need to be developed on the selected biomarker panels to measure the absolute metabolite abundance independent of batches, toward clinical translation and reduced costs. When implementing NPELDI-MS for routine BSG diagnosis and monitoring, size- and cost-effective mass spectrometers with adequate analytical performance for metabolic profiling are needed in dedicated clinical environments, particularly in remote locations. The exploration of miniaturization within mass spectrometers presents an opportunity for further advancements in this area. Additionally, simplifying the sample collection process, such as utilizing dried blood spots for patient blood samples, could facilitate the transition toward point-of-care testing in laboratory settings.
In summary, we demonstrated the potency of metabolic snapshots for diagnosis, prognosis, and surveillance of BSG by launching both cross-sectional studies and longitudinal cohort studies and employing NPELDI-MS to record static metabolic snapshots and dynamic metabolic trajectories. The results of this study offer support for setting up large-scale randomized clinical trials to prospectively validate its clinical applicability.
Methods
Ethics statement
This retrospective cohort study was approved by the institutional ethics committees of Xinhua Hospital, Shanghai Jiao Tong University (#XHEC-C-2019-101). All participants in this study provided written consent and approved the use of serum samples for analysis. All animal care and experiments were approved (#XHEC-F-2023-031) by the Animal Ethics Committee of Xinhua Hospital, School of Medicine, Shanghai Jiao Tong University.
Chemicals and reagents
All chemicals and reagents used in this study are listed in Supplementary Tables 9–11 with all corresponding details. Ferric nanoparticles were synthesized with these reagents via a revised solvo-thermal method32,89.
Subject enrollment and serum sample collection
A total of 299 participants, comprising 125 HDs and 174 pediatric brainstem glioma patients (BSG) aged 1–15 years, were initially recruited at Xinhua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine from November 2017 to May 2023. Among these 299 subjects, 68 patients were excluded for various reasons, including: (1) failure to complete the radiation therapy; (2) receipt of other treatments; and (3) improper serum preservation. BSG patients were diagnosed based on abnormalities observed in MRI images of the brainstem and confirmed by two independent oncologists and imaging specialists. All individuals with BSG were confirmed through the use of radiological imaging and clinical symptoms.
We designed a cross-sectional study for BSG diagnosis and prognosis, as well as a longitudinal study to monitor responses to radiotherapy. We first collected 244 serum samples from 60 HDs and 73 BSG patients (HDs/BRT/DRT/ART = 60/73/59/52, Supplementary Table 1), as the discovery and validation cohort. Following the same process, we newly collected 161 serum samples, including 65 HDs, 33 BRT, 28 DRT, and 35 ART (Supplementary Table 2), as the external cohort. Finally, A total of 186 serum samples were collected longitudinally from 62 patients at three time points: before radiotherapy (BRT), during radiotherapy (DRT), and after radiotherapy (ART)2,64,74.
All subjects underwent a complete clinical examination, including the physical signs (gender and age), radiological imaging, and follow-up investigation. All patients underwent radiotherapy according to guidelines7. During the treatment, all patients maintain the same hospital-provided diet without additional nutritional food or drugs. All subjects kept an emotional balance without heavy physical exercise for two days before serum collection. Blood samples were collected from participants who had fasted for at least 8 h. All serum samples were stored in microtubes at −80 °C until analysis. Such strict enrollment criteria were implemented during sample collection to mitigate potential influences from external factors, such as medication and diet according to existing literature90.
Cohort design
All serum samples were analyzed by nanoparticle-enhanced laser desorption/ionization mass spectrometry (NPELDI-MS) platform to obtain the serum metabolic profiles.
For BSG diagnosis (cohort 1), we randomly assigned 294 serum subjects to (1) a discovery set of 54 samples from BSG patients and 45 samples from HDs, (2) a validation set of 19 samples from BSG patients and 15 samples from HDs, and (3) an independent external set of 96 samples from BSGs and 65 samples from HDs. For BSG prognosis (cohort 2), we randomly assigned 96 BSG subjects to (1) a training set of 58 observations with 41 deaths and (2) a testing set of 38 observations with 31 deaths. For dynamic metabolic snapshots during radiotherapy (cohort 3), we enrolled 388 samples, including 117 HDs, 101 BRT, 85 DRT, and 85 ART.
In studies based on static metabolic snapshots, the age and gender of BSG patients and HDs were matched with no significant difference for the discovery set by two-sided Student’s t-test and Chi-Squared test in SPSS platform (version 19.0, SPSS Inc., USA), respectively.
LDI-MS detection
All 405 serum samples (including 125 HDs, 106 BRT, 87 DRT, and 87 ART) were used for MS analysis without any complex pre-treatment91,92. In a typical metabolic profiling process, 100 nL of naive serum for each sample was dropped onto a substrate chip to form a high-throughput MS microarray. Nanoparticle suspension was further distributed over the dried serum sample spot. The microarray chip was stored under constant temperature and humidity conditions for nanoparticles to in situ enrich the metabolites and achieve a uniform co-crystallization93.
Sample targets were further loaded into a Bruker Autoflex III MALDI TOF/TOF mass spectrometer (Brucker Daltonics, Bremen, Germany). The MS instrument is equipped with a Nd:YAG laser source with a wavelength of 355 nm. Mass spectra were collected in the positive ion mode, using a pulse laser at a frequency of 1000 Hz with 2000 shots. Full scan was carried out at the m/z range of 100–1000. The acceleration voltage and delay time were optimized as 20 kV and 150 ns, respectively. Five technical replicates were implemented.
Current metabolic profiling techniques mainly include nuclear magnetic resonance and MS. LDI-MS stands out for its ability to rapidly detect metabolites in a high-throughput manner. Throughout the analysis process, the abundance of metabolites and the complexity of samples significantly affect MS analysis, necessitating rigorous pretreatment procedures to separate and enrich metabolites from complex biological mixtures. Utilizing nanomaterials as substrates and fine-tuning the interaction interfaces between molecules and substrate materials by controlling the physicochemical properties of the material interface can significantly enhance the detection efficiency of LDI-MS.
Ferric nanoparticle as an alternative of chromatograph in LDI-MS possesses three advantages: (1) nano-scale surface roughness and large-scale uniformity that enable the rapid and selective laser desorption/ionization of metabolites; (2) optimized surface charge that promotes ion formation and facilitates electron transfer; and (3) ease of mass-production with cost-effectiveness that are suitable for extensive clinical applications28. Based on these advantages, NPELDI-MS platform avoids the necessity for sample pretreatment, leading to faster analysis speed (~seconds) and a significant reduction in sample consumption (100 nL) by ~2–3 orders of magnitude.
Metabolite identification
The verification of the metabolites in this study was achieved by comparing the m/z features with the human metabolome database (HMDB, http://www.hmdb.ca/). Further verification was conducted by comparing the exact mass in serum to the obtained mass of standard molecules, acquired by high-resolution MALDI-Fourier transform ion cyclotron resonance (FTICR) MS (Supplementary Tables 5 and 7). The elemental composition of MS peak was determined by the exact accurate mass at an m/z tolerance of 10 ppm.
Authentic standards were used in this study. Following the guidelines of the Metabolomics Standards Initiative94, the level of metabolite IDs should be stated as Level 1 (Annotated compounds: accurate mass confirmation and real-case serum sample validation relative to an authentic compound analyzed under identical experimental conditions). This level of identification covered all identified metabolite biomarkers except for 2-AAS, oxalacetic acid, and threonic acid, for which we failed to obtain the chemical standards. For 2-AAS, oxalacetic acid, and threonic acid, theoretical mass of adducts was compared within an m/z tolerance of 10 ppm.
Pathway analysis
We conducted overrepresentation analysis using MetaboAnalyst 5.0 (https://www.metaboanalyst.ca/) to perform metabolite set enrichment analysis (MSEA) on the identified metabolite biomarkers from both diagnostic and prognostic panels. The enrichment analysis was based on the metabolic network in the Small Molecule Pathway Database (SMPDB) dataset, comprising 99 metabolite sets representing normal human metabolic pathways. Pathways with an enrichment ratio greater than 5 (calculated by observed hits/expected hits) and a significance level of p < 0.05 were considered relevant, aligning with established practices in the field95,96,97. Importantly, MSEA enables the investigation of functionally related metabolite groups rather than individual metabolites, helping to identify pathways that may be involved in the observed metabolic changes.
This approach is advantageous as it avoids bias that could arise from pre-selecting metabolites based on an arbitrary threshold98. By employing pathway analyses such as MSEA, we were able to gain insights into the significant metabolic pathways influenced by the identified metabolites. This approach served as a valuable tool for knowledge-based dimensionality reduction, allowing us to focus on essential pathways while excluding redundant or less influential ones. Through this analysis, we aimed to provide a more streamlined interpretation of the metabolic pathways and their associations with the identified metabolites, without oversimplifying the dataset.
Cell culture
SHSMU-DIPG-01 cells were derived from surgically resected diffuse intrinsic pontine glioma (DIPG) tissue samples obtained during pediatric neurosurgery from a two-year-old boy at Shanghai Jiao Tong University School of Medicine-affiliated Xinhua Hospital. Fresh tumor specimens were aseptically collected, pathologically confirmed as DIPG, and subjected to short tandem repeat profiling for cell line authentication. The DIPG cells were cultured in tumor stem medium supplied with fetal bovine serum, penicillin, streptomycin, amphotericin B, and a series of growing factors (Supplementary Tables 10–11).
In vitro cell experiments
SHSMU-DIPG-01 cells were seeded in a 96-well plate at the density of 1 × 104 cells per well and incubated at 37 °C for 24 h. To determine the effect of lactic acid, ornithine, and succinic acid, the cells were cultured in complete medium and treated with each metabolite at varying concentrations (0, 0.25, 0.5, 1, 2, 4, 8, and 16 mM) in triplicate for 72 h. For the assessment of valine, leucine, threonine, arginine, niacinamide, and glucose, the cells were cultured in medium devoid of these metabolites and subsequently treated with each metabolite at the same concentrations in triplicate for 72 h. The cell viability was determined using an enhanced cell counting kit-8 (CCK8, Beyotime). Specifically, 10 μL of CCK8 solution was added to each well. After 1 h of cell culture, the optical density (OD) value at 450 nm was detected via a plate reader (BioTek Synergy H1). The metabolites originally present in the culture medium had the control group that was cultured with a medium lacking these metabolites during the co-culture treatment.
In vivo mouse experiments
SPF-grade female BALB/c nude mice, aged 4–6 weeks and weighing 18–20 g, were used as experimental subjects in this study99. Xinhua Hospital Animal Ethics Committee mandates that tumor-bearing murine models must adhere to strict ethical standards, with tumors not exceeding 15 mm in maximum diameter or 2000 mm³ in total volume. Throughout this study, no tumors surpassed these predefined limits. Euthanasia was required when exhibited significant body weight loss (loss greater than 20% of initial weight or no weight gain for 7 consecutive days) along with severe neurobehavioral deficits, including unsteady gait, circular walking, and frequent falls. Assessments and euthanasia procedures were performed in compliance with established laboratory animal euthanasia guidelines.
The SPF-grade animal facility maintains stringent environmental parameters: temperature regulation at 20–26 °C, relative humidity of 40–70%, and a positive pressure differential of 10–15 Pa to mitigate cross-contamination. Operational standards include a 12-h photoperiod with UV-free illumination.
To establish the orthotopic intracranial xenograft model, SHSMU-DIPG-01 cells were transduced with lentiviral vectors encoding enhanced green fluorescent protein and firefly luciferase. Positively transduced cells were subsequently sorted by fluorescence-activated cell sorting to achieve a purity greater than 95% prior to implantation. Under deep anesthesia, mice were secured on a stereotactic frame, and a cranial burr hole was drilled at stereotactic coordinates relative to the lambda suture: 0.8 mm posterior, 1 mm lateral to the midline, and 4.5 mm depth. A total of 1 × 106 cells suspended in 3 μL phosphate-buffered saline (PBS) were infused via Hamilton syringe at a controlled rate of 0.3 μL/min. To minimize reflux, the injection needle was retained for 5 min post-infusion. Tumor engraftment was validated 72 h post-surgery by intraperitoneal administration of D-luciferin potassium salt (75 mg/kg; Promega) followed by bioluminescent signal acquisition using the IVIS Spectrum In Vivo Imaging System (PerkinElmer). At the humane endpoint, mice were euthanized. Brain tumor tissue and normal mice brainstem tissues were collected and subjected to RNA sequencing.
To establish the subcutaneous tumor model, 3 × 106 SHSMU-DIPG-01 cells were resuspended in a 1:1 mixture with chilled Matrigel matrix (Corning), maintained on ice throughout preparation. A 100 μL aliquot of the cell-Matrigel suspension was injected subcutaneously into the left dorsal flank of mice using a sterile 25-gauge needle. Tumor volume was measured using a vernier caliper. Radiotherapy was initiated in subcutaneous tumor-bearing mice when the tumor size exceeded 100 mm3. Mice in the radiotherapy group were administered a daily radiation dose of 2 Gy at the tumor site for five consecutive days.
RNA sequencing and analyses
Total RNA was extracted using the TRIzol reagent (Invitrogen, CA, USA) according to the manufacturer’s protocol. RNA purity and quantification were evaluated using the NanoDrop 2000 spectrophotometer (Thermo Scientific, USA). All samples were analyzed in triplicate. RNA integrity was assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Then the libraries were constructed using VAHTS Universal V6 RNA-seq Library Prep Kit.
The libraries were sequenced on an Illumina Novaseq 6000 platform and 150 bp paired-end reads were generated. About 21.39–30.61 M raw reads for each sample were generated. Raw reads of FASTQ format were firstly processed using fastp and the low-quality reads were removed to obtain the clean reads. Then, about 19.79-26.21 M clean reads for each sample were retained for subsequent analyses. The clean reads were mapped to the reference genome using HISAT2. FPKM of each gene was calculated and the read counts of each gene were obtained by HTSeq-count.
Differential expression analysis was performed using the DESeq2. p value < 0.05 and fold change >2 or fold change <0.5 was set as the threshold for significantly DEGs. Based on the hypergeometric distribution, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of DEGs was performed to screen the significantly enriched term using R (v 3.2.0).
Statistics and reproducibility
The average of five mass spectra obtained during each 20-s NPELDI-MS analysis was utilized to construct databases for serum metabolic profiles. To ensure MS data quality, each run was evaluated by assessing the coefficients of variation (CVs) for 70% of features from quality-control samples and was considered failed and discarded from analysis when CV > 30%. Raw MS data was transformed into a Microsoft Excel spreadsheet format. After quality control and data filtering, we obtained 402 metabolic features (defined as the localized highest intensity of the raw MS data) across different samples. It should be noted that the metabolic features here refer to MS peaks that have not been identified. Models were generated using four common algorithms, including the least absolute shrinkage and selection operator (LASSO), Ridge, K-Nearest Neighbor (kNN), and random forest (RF).
A mathematical coefficient weight was calculated for each statistically informative feature using LASSO and Ridge, depending on the significance of the mass spectral feature in characterizing a specific class (BSG vs. HDs). Classification was conducted using a 20-fold cross-validation approach to evaluate the predictive accuracy within the discovery set via Orange (version 3.34.0). The difference of metabolites among HDs, BRT, DRT, and ART were evaluated using pairwise comparison via multiple t test on SPSS software. Delong tests were calculated based on “DeLong UI” codes by MATLAB (version R2024b, The MathWorks, USA)100. Power analysis was conducted to predetermine sample size on MetaboAnalyst 5.0101. Unsupervised analyses, such as principal component analysis and t-stochastic neighbor embedding, were performed by Python (version 3.7). RseQC (RNA-Seq Quality Control, version 4.0.0) was employed to evaluate the quality of RNA-seq data. All significance levels for p value were set as 0.05 throughout the experiments. The c-means clustering was performed by “mfuzz” package in R (version 3.2.2). One-way ANOVA was used to perform statistical analyses in cellular experiments. Stars represented p value as follows: * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001. Randomization and blinding were used in this study.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNAseq data have been deposited in the public GEO database under accession code GSE 293092. Metabolic profiles by laser desorption/ionization MS are provided in the Supplementary Data 1. Other data generated in this study are provided in the Supplementary Information and Source Data file. Source data are provided with this paper.
References
Cohen, A. R. Brain tumors in children. N. Engl. J. Med. 386, 1922–1931 (2022).
Larson, J. D. et al. Histone H3.3 K27M accelerates spontaneous brainstem glioma and drives restricted changes in bivalent gene expression. Cancer Cell 35, 140–155 (2019).
Khuong-Quang, D.-A. et al. K27M mutation in histone H3.3 defines clinically and biologically distinct subgroups of pediatric diffuse intrinsic pontine gliomas. Acta Neuropathol. 124, 439–447 (2012).
Wu, G. et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet. 44, 251–253 (2012).
Pan, C. et al. Molecular profiling of tumors of the brainstem by sequencing of CSF-derived circulating tumor DNA. Acta Neuropathol. 137, 297–306 (2019).
Ayoub, R. et al. Assessment of cognitive and neural recovery in survivors of pediatric brain tumors in a pilot clinical trial using metformin. Nat. Med. 26, 1285–1294 (2020).
Varlet, P. et al. WHO grade has no prognostic value in the pediatric high-grade glioma included in the HERBY trial. Neuro Oncol. 22, 116–127 (2019).
Deland, K. et al. Tumor genotype dictates radiosensitization after Atm deletion in primary brainstem glioma models. J. Clin. Investig. 131, e142158 (2021).
Wang, K. & Tepper, J. E. Radiation therapy-associated toxicity: etiology, management, and prevention. CA Cancer J. Clin. 71, 437–454 (2021).
Mouliere, F. et al. Fragmentation patterns and personalized sequencing of cell-free DNA in urine and plasma of glioma patients. EMBO Mol. Med. 13, e12881 (2021).
Kawauchi, D. et al. Early diagnosis and surgical intervention within 3 weeks from symptom onset are associated with prolonged survival of patients with glioblastoma. Neurosurgery 91, 741–748 (2022).
Hashizume, R. & Gupta, N. Patient-derived tumor models for diffuse intrinsic pontine gliomas. Curr. Neuropharmacol. 15, 98–103 (2017).
Hollon, T. C. et al. Near real-time intraoperative brain tumor diagnosis using stimulated Raman histology and deep neural networks. Nat. Med. 26, 52–58 (2020).
Miller, A. M. et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 565, 654–658 (2019).
Indira Chandran, V. et al. Ultrasensitive immunoprofiling of plasma extracellular vesicles identifies syndecan-1 as a potential tool for minimally invasive diagnosis of glioma. Clin. Cancer Res. 25, 3115–3127 (2019).
Regazzo, G. et al. A restricted signature of serum miRNAs distinguishes glioblastoma from lower grade gliomas. J. Exp. Clin. Cancer Res. 35, 124 (2016).
Nassiri, F. et al. Detection and discrimination of intracranial tumors using plasma cell-free DNA methylomes. Nat. Med. 26, 1044–1047 (2020).
Han, L. et al. Systemic delivery of monoclonal antibodies to the central nervous system for brain tumor therapy. Adv. Mater. 31, 1805697 (2019).
Tian, D.-S. et al. FSAP aggravated endothelial dysfunction and neurological deficits in acute ischemic stroke due to large vessel occlusion. Signal Transduct. Target. Ther. 7, 6 (2022).
Zhang, Y. et al. Pan-cancer circulating tumor DNA detection in over 10,000 Chinese patients. Nat. Commun. 12, 11 (2021).
Duperron, M.-G. et al. Genomics of perivascular space burden unravels early mechanisms of cerebral small vessel disease. Nat. Med. 29, 950–962 (2023).
Abbasi, J. Guided ultrasound opens blood-brain barrier to cancer drugs. JAMA 326, 1785 (2021).
Zheng, F. et al. Applications of nanobodies in brain diseases. Front. Immunol. 13, 978513 (2022).
Song, Y., Hu, C., Fu, Y. & Gao, H. Modulating the blood–brain tumor barrier for improving drug delivery efficiency and efficacy. VIEW 3, 20200129 (2022).
Huang, L. et al. Plasmonic silver nanoshells for drug and metabolite detection. Nat. Commun. 8, 220 (2017).
Fischer, G. M. et al. Molecular profiling reveals unique immune and metabolic features of melanoma brain metastases. Cancer Discov. 9, 628–645 (2019).
Tao, L. et al. MYCN-driven fatty acid uptake is a metabolic vulnerability in neuroblastoma. Nat. Commun. 13, 3728 (2022).
Huang, L. et al. Machine learning of serum metabolic patterns encodes early-stage lung adenocarcinoma. Nat. Commun. 11, 3556 (2020).
Xu, Z. et al. Efficient plasma metabolic fingerprinting as a novel tool for diagnosis and prognosis of gastric cancer: a large-scale, multicentre study. Gut 72, 2051 (2023).
Sun, L.-Y. et al. Nuclear receptor NR1D1 regulates abdominal aortic aneurysm development by targeting the mitochondrial tricarboxylic acid cycle enzyme Aconitase-2. Circulation 146, 1591–1609 (2022).
Chen, Y. et al. Plasma metabolic fingerprints for large-scale screening and personalized risk stratification of metabolic syndrome. Cell Rep. Med. 4, 101109 (2023).
Huang, Y. et al. Diagnosis and prognosis of breast cancer by high-performance serum metabolic fingerprints. Proc. Natl. Acad. Sci. USA 119, e2122245119 (2022).
Ding, C. et al. Pt/NiFe-LDH hybrids for quantification and qualification of polyphenols. Mater. Today Bio 26, 101047 (2024).
Grimm, S. A. & Chamberlain, M. C. Brainstem glioma: a review. Curr. Neurol. Neurosci. Rep. 13, 346 (2013).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
Hoxhaj, G. & Manning, B. D. The PI3K-AKT network at the interface of oncogenic signalling and cancer metabolism. Nat. Rev. Cancer 20, 74–88 (2020).
Faubert, B., Solmonson, A. & Deberardinis, R. J. Metabolic reprogramming and cancer progression. Science 368, 152 (2020).
Li, X. et al. Plasmonic array assisted mass spectrometry for preferential metabolite detection. Chem. Eng. J. 486, 150224 (2024).
Chen, X., Shu, W., Zhao, L. & Wan, J. Advanced mass spectrometric and spectroscopic methods coupled with machine learning for in vitro diagnosis. VIEW 4, 20220038 (2023).
Maldonado, M. D. et al. Features of diffuse gliomas that are misdiagnosed on initial neuroimaging: a case control study. J. Neuro Oncol. 140, 107–113 (2018).
Arrillaga-Romany, I. L. A. M. ACTION: a randomized phase 3 study of ONC201 (dordaviprone) in patients with newly diagnosed H3K27M-mutant diffuse glioma. Neuro Oncol. 26, 173–181 (2024).
De Vitis, C. et al. ALDOC- and ENO2-driven glucose metabolism sustains 3D tumor spheroids growth regardless of nutrient environmental conditions: a multi-omics analysis. J. Exp. Clin. Cancer Res. 42, 69 (2023).
Chinnaiyan, P. et al. The metabolomic signature of malignant glioma reflects accelerated anabolic metabolism. Cancer Res. 72, 5878–5888 (2012).
Jin, E. et al. P4HA2 activates mTOR via hydroxylation and targeting P4HA2-mTOR inhibits lung adenocarcinoma cell growth. Oncogene 43, 1813–1823 (2024).
Wang, Y. et al. Mechanisms of radioresistance and radiosensitization strategies for triple negative breast cancer. Transl. Oncol. 55, 102351 (2025).
Ding, M., Zhang, E., He, R. & Wang, X. Newly developed strategies for improving sensitivity to radiation by targeting signal pathways in cancer therapy. Cancer Sci. 104, 1401–1410 (2013).
Taneja, N., Chauhan, A., Kulshreshtha, R. & Singh, S. HIF-1 mediated metabolic reprogramming in cancer: mechanisms and therapeutic implications. Life Sci. 352, 122890 (2024).
Peng, C. et al. Regulation of the hippo-YAP pathway by glucose sensor O-glcNAcylation. Mol. Cell 68, 591–604.e595 (2017).
Nikas, I. P., Paschou, S. A. & Ryu, H. S. The role of nicotinamide in cancer chemoprevention and therapy. Biomolecules 10, 477 (2020).
Xu, J. & Richard, S. Cellular pathways influenced by protein arginine methylation: implications for cancer. Mol. Cell 81, 4357–4368(2021).
Yang, J. S., Wang, C. C., Qiu, J. D., Ren, B. & You, L. Arginine metabolism: a potential target in pancreatic cancer therapy. Chin. Med. J. 134, 28–37 (2020).
Wu, X. et al. Threonine fuels glioblastoma through YRDC-mediated codon-biased translational reprogramming. Nat. Cancer 5, 1024–1044 (2024).
Lewis, J. E. & Kemp, M. L. Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance. Nat. Commun. 12, 2700 (2021).
Martínez-Reyes, I. & Chandel, N. S. Cancer metabolism: looking forward. Nat. Rev. Cancer 21, 669–680 (2021).
Wang, Z. et al. Silk microneedle patch capable of on-demand multidrug delivery to the brain for glioblastoma treatment. Adv. Mater. 34, 2106606 (2022).
Xue, J. et al. Deep learning-based detection and segmentation-assisted management of brain metastases. Neuro Oncol. 22, 505–514 (2019).
Marner, L. et al. Diagnostic accuracy and clinical impact of [18F]FET PET in childhood CNS tumors. Neuro Oncol. 23, 2107–2116(2021).
Wang, R. et al. A sustainable approach to universal metabolic cancer diagnosis. Nat. Sustain. https://doi.org/10.1038/s41893-41024-01323-41899 (2024).
Milanaccio, C. DIPG-27. Behavioral disturbances as underestimated presenting symptoms in children with Diffuse Intrinsic Pontine Glioma (DIPG). Neuro Oncol. 24, i24 (2022).
Baklizi, N., Raswoli, M., Burges, M., Moreira, D. C. & Qaddoumi, I. Torticollis as a presenting symptom of pediatric CNS tumors: a systematic review. Semin. Oncol. 49, 419–425 (2022).
Patel, V., McNinch, N. L. & Rush, S. Diagnostic delay and morbidity of central nervous system tumors in children and young adults: a pediatric hospital experience. J. Neuro Oncol. 143, 297–304(2019).
Hoffman, L. M. et al. Clinical, radiologic, pathologic, and molecular characteristics of long-term survivors of diffuse intrinsic pontine glioma (DIPG): a collaborative report from the international and European society for pediatric oncology DIPG registries. J. Clin. Oncol. 36, 1963–1972 (2018).
Chen, J. et al. APOLLO: an accurate and independently validated prediction model of lower-grade gliomas overall survival and a comparative study of model performance. EBio Med. 79, 104007 (2022).
Wu, G. et al. The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat. Genet. 46, 444–450 (2014).
Xiao, Y. et al. Emerging therapies in cancer metabolism. Cell Metab. 35, 1283–1303 (2023).
Vernieri, C. et al. Targeting cancer metabolism: dietary and pharmacologic interventions. Cancer Discov. 6, 1315–1333 (2016).
Elia, I. & Haigis, M. C. Metabolites and the tumour microenvironment: from cellular mechanisms to systemic metabolism. Nat. Metab. 3, 21–32 (2021).
Qu, N. et al. Integrated proteogenomic and metabolomic characterization of papillary thyroid cancer with different recurrence risks. Nat. Commun. 15, 3175 (2024).
Chen, Y. et al. Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer. Nat. Commun. 15, 1657 (2024).
Christopher et al. Metabolic heterogeneity in human lung tumors. Cell 164, 681–694 (2016).
Ohno, M. et al. Assessment of the diagnostic utility of serum microRNA classification in patients with diffuse glioma. JAMA Netw. Open 2, e1916953 (2019).
Nikbakht, H. et al. Spatial and temporal homogeneity of driver mutations in diffuse intrinsic pontine glioma. Nat. Commun. 7, 11185 (2016).
Contrepois, K. et al. Molecular choreography of acute exercise. Cell 181, 1112–1130 (2020).
Liang, L. et al. Metabolic dynamics and prediction of gestational age and time to delivery in pregnant women. Cell 181, 1680–1692 (2020).
Buergel, T. et al. Metabolomic profiles predict individual multidisease outcomes. Nat. Med. 28, 2309–2320 (2022).
Ben-Hamo, R. et al. Predicting and affecting response to cancer therapy based on pathway-level biomarkers. Nat. Commun. 11, 3296 (2020).
Shu, T. et al. Plasma proteomics identify biomarkers and pathogenesis of COVID-19. Immunity 53, 1108–1122 (2020).
Boehm, K. M., Khosravi, P., Vanguri, R., Gao, J. & Shah, S. P. Harnessing multimodal data integration to advance precision oncology. Nat. Rev. Cancer 22, 114–126 (2022).
Luo, H. et al. Circulating tumor DNA methylation profiles enable early diagnosis, prognosis prediction, and screening for colorectal cancer. Sci. Transl. Med. 12, eaax7533 (2020).
Funato, K., Major, T., Lewis, P. W., Allis, C. D. & Tabar, V. Use of human embryonic stem cells to model pediatric gliomas with H3.3K27M histone mutation. Science 346, 1529–1533 (2014).
Jones, C. & Baker, S. J. Unique genetic and epigenetic mechanisms driving paediatric diffuse high-grade glioma. Nat. Rev. Cancer 14, 651–661 (2014).
Peyre, M. et al. Somatic PIK3CA mutations in sporadic cerebral cavernous malformations. N. Engl. J. Med. 385, 996–1004 (2021).
Handzlik, M. K. et al. Insulin-regulated serine and lipid metabolism drive peripheral neuropathy. Nature 614, 118–124 (2023).
Subbiah, V. et al. Dabrafenib plus trametinib in BRAFV600E-mutated rare cancers: the phase 2 ROAR trial. Nat. Med. 29, 1103–1112 (2023).
Zhang, X. & Dang, C. V. Time to hit pause on mitochondria-targeting cancer therapies. Nat. Med. 29, 29–30 (2023).
Ribas, A. et al. Association of pembrolizumab with tumor response and survival among patients with advanced melanoma. JAMA 315, 1600 (2016).
Rizk, N., Kamiya-Matsuoka, C. & Tremont-Lukats, I. W. Lower extremity edema, anxiety, and cyanosis during chemoradiation therapy for glioblastoma. JAMA Oncol. 2, 265 (2016).
Yamasaki, F. et al. Magnetic resonance spectroscopic detection of lactate is predictive of a poor prognosis in patients with diffuse intrinsic pontine glioma. Neuro Oncol. 13, 791–801 (2011).
Liu, W. et al. Monitoring retinoblastoma by machine learning of aqueous humor metabolic fingerprinting. Small Methods 6, 2101220 (2022).
Simundic, A. M. et al. Joint EFLM-COLABIOCLI recommendation for venous blood sampling. Clin. Chem. Lab. Med. 56, 2015–2038(2018).
Cao, J. et al. Metabolic fingerprinting on synthetic alloys for medulloblastoma diagnosis and radiotherapy evaluation. Adv. Mater. 32, 2000906 (2020).
Wang, R. et al. A “one-stop shop” decision tree for diagnosing and phenotyping polycystic ovarian syndrome on serum metabolic fingerprints. Adv. Funct. Mater. 32, 2206670 (2022).
Wu, J. et al. Glaucoma characterization by machine learning of tear metabolic fingerprinting. Small Methods 6, 2200264 (2022).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3, 211–221 (2007).
Wang, X. et al. Targeting pyrimidine synthesis accentuates molecular therapy response in glioblastoma stem cells. Sci. Transl. Med. 11, eaau4972 (2019).
Deng, L. et al. Sparse PLS-based method for overlapping metabolite set enrichment analysis. J. Proteome Res. 20, 3204–3213 (2021).
Morel, J. D. et al. The mouse metallomic landscape of aging and metabolism. Nat. Commun. 13, 607 (2022).
Xia, J. & Wishart, D. S. MSEA: a web-based tool to identify biologically meaningful patterns in quantitative metabolomic data. Nucleic Acids Res. 38, W71–W77 (2010).
Grasso, C. S. et al. Functionally defined therapeutic targets in diffuse intrinsic pontine glioma. Nat. Med. 21, 555–559 (2015).
Sun, X. & Xu, W. Fast implementation of DeLong’s algorithm for comparing the areas under correlated receiver operating characteristic curves. IEEE Signal Process. Lett. 21, 1389–1393 (2014).
Pang, Z. et al. Using MetaboAnalyst 5.0 for LC-HRMS spectra processing, multi-omics integration and covariate adjustment of global metabolomics data. Nat. Protoc. 17, 1735–1761 (2022).
Acknowledgements
We thank the financial support from the National Key Research and Development Program of China (2022YFC2502800 (J.Wan)), National Natural Science Funds (Grant Nos. 22474040 (J.Wan), 82422042 (L.H.), 82372148 (L.H.)), Medical-Engineering Joint Funds of Shanghai Jiao Tong University (Nos. YG2024ZD07 (K.Q.), YG2023ZD08 (K.Q.)), and Shanghai Institutions of Higher Learning (No. 2021-01-07-00-02-E00083 (K.Q.)). This work was also sponsored by the Innovative Research Team of High-Level Local Universities in Shanghai (SHSMU-ZDCX20210700 (K.Q.)), Innovation Group Project of Shanghai Municipal Health Commission (2019CXJQ03 (K.Q.)), Innovation Research Plan by the Shanghai Municipal Education Commission (ZXWF082101 (K.Q.)), National Research Center for Translational Medicine Shanghai (Nos. TMSK-2021-124 (K.Q.), NRCTM(SH)-2021-06 (K.Q.)), Science and Technology Commission of Shanghai Municipality (20DZ2220400 (K.Q.)), Hospital Funded Clinical Research, Xin Hua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine (24XHCR09B (M.J.)), and Fundamental Research Funds for the Central Universities. We thank Prof. Yujie Tang from Key Laboratory of Cell Differentiation and Apoptosis of National Ministry of Education in Shanghai Jiao Tong University School of Medicine for providing us the DIPG cell lines.
Author information
Authors and Affiliations
Contributions
K.L., R.W., Z.G., and W.W. contributed equally to this work. K.Q., M.J., L.H., and J.Wan conceived the study. K.L., R.W., and Z.G. carried out the study and wrote the initial article. K.L. and J.S. contributed to the collection of serum samples. R.W. completed all the MS experiments and wrote the manuscript with W.L., Y.H., and J.Wu., R.W., Z.Z., and S.Y. contributed to the MS data analysis. R.W., Z.G., W.W., and Y.T. designed and completed the cellular and animal experiments. All authors joined in the critical discussion and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.W., L.H., and K.Q. have filed patents (CN117191934A, China, 2023) for both the technology and the use of the technology to detect bio-samples. These patents are owned and managed by Shanghai Jiao Tong University. The other authors declare no competing interests. No patient titles or affiliations influenced the design, analysis, or interpretation of this study. This study employed 367 anonymized residual clinical participants collected under observational protocols at Xin Hua Hospital, Shanghai Jiao Tong University School of Medicine (#XHEC-C-2019-101). The cohort included 174 pediatric brainstem glioma patients, 125 HDs, and 68 other neural diseases; no protected health information was disclosed.
Peer review
Peer review information
Nature Communications thanks Rintaro Hashizume and the other anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, K., Wang, R., Gu, Z. et al. Serum metabolic profiling enables diagnosis, prognosis, and monitoring for brainstem gliomas. Nat Commun 16, 6108 (2025). https://doi.org/10.1038/s41467-025-61163-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61163-9