Abstract
Proteins are the molecular machines of life with numerous applications in energy, health, and sustainability. However, engineering proteins with desired functions for practical applications remains slow, expensive, and specialist-dependent. Here we report a generally applicable platform for autonomous enzyme engineering that integrates machine learning and large language models with biofoundry automation to eliminate the need for human intervention, judgement, and domain expertise. Requiring only an input protein sequence and a quantifiable way to measure fitness, this automated platform can be applied to engineer a wide array of proteins. As a proof of concept, we engineer Arabidopsis thaliana halide methyltransferase (AtHMT) for a 90-fold improvement in substrate preference and 16-fold improvement in ethyltransferase activity, along with developing a Yersinia mollaretii phytase (YmPhytase) variant with 26-fold improvement in activity at neutral pH. This is accomplished in four rounds over 4 weeks, while requiring construction and characterization of fewer than 500 variants for each enzyme. This platform for autonomous experimentation paves the way for rapid advancements across diverse industries, from medicine and biotechnology to renewable energy and sustainable chemistry.
Similar content being viewed by others
Introduction
Laboratory experiments are essential to scientific research, which is traditionally driven by skilled researchers. However, many factors can slow down the conventional discovery process and limit efficiency. Manual laboratory tasks are time- and labor-intensive, prone to reproducibility issues, limited in scalability, and grow increasingly complicated when managing large datasets and high-throughput experiments. Autonomous experimentation has the potential to transform scientific research by enabling integration of artificial intelligence (AI) and robotics to iteratively propose hypotheses, design and conduct experiments, and refine models with minimal human intervention. AI-enabled systems can explore vast, multi-dimensional spaces more efficiently than traditional computational techniques while robotics and automation can perform experiments faster, more reliably and at higher throughputs with better scalability. Autonomous laboratories hold tremendous potential for diverse fields such as synthetic biology, chemical synthesis, and materials discovery by enabling more efficient and scalable research1,2,3. Early demonstration of such a system includes the Robot Scientist “Adam” that autonomously generated and tested functional genomics hypotheses for Saccharomyces cerevisiae4. Coscientist, a multi-LLMs-based intelligent agent, can autonomously perform chemical synthesis reactions5. In materials science, autonomous experimentation platform Ada optimized thin-film compositions6. The A-Lab combines robotics, machine learning (ML), and historical data to synthesize inorganic powders and successfully created 41 new compounds in 17 days of continuous operation7. A recent work developed autonomous mobile robots that can perform, analyze and decide next steps for exploratory synthetic chemistry reactions3,8.
In biology, autonomous experimentation is comparatively less mature, making even routine processes like DNA assembly, gene editing, or metabolic engineering a challenging task1,9. Moreover, integrating instruments for continuous experimentation requires skilled personnel with expertise at the intersection of biological experimentation, robotics, and programmings10,11,12. Much previous work in autonomous synthetic biology has been highly specific, targeting singular goals like engineering a single protein11, metabolic engineering for a single product13, or orphan enzyme identification4. Despite these advances, a broadly applicable autonomous system must be highly generalizable for extensive utility. Generalizable platforms are more scalable and adaptable, able to address diverse problems across different locations without the need for new workflows. Widely adopted systems enable researchers to use a common scientific framework, promoting collaboration and efficient knowledge transfer while minimizing the need to redevelop similar methods for common goals. With scalable, generalizable platforms, synthetic biology can move beyond isolated successes and drive innovation in a wide range of fields.
In this work, we use protein engineering as a case study to establish a roadmap for generalized autonomous experimentation in synthetic biology. Protein engineering provides an extensive toolkit for modifying enzymes for widespread application in fields such as medicine, biofuels, and biocatalysis14,15. By iterative design, build, test, and learn (DBTL) cycles, enzymes can be made more stable, selective, or efficient. Methods like directed evolution and computer-aided design offer diverse strategies for protein engineering and high-throughput screening strategies allow iterating over a large sample space16,17,18. Despite the wide applicability of protein engineering, there remain unmet needs, particularly in efficiently navigating vast sequence spaces and optimizing protein function in complex environments. Autonomous protein engineering represents the state-of-the-art in addressing these challenges through self-driving frameworks for executing the DBTL cycle10,19,20. Many synthetic biology applications have demonstrated decent success for automated design, such as in metabolic engineering13 and construction of plasmids21. A recent study11 demonstrated an automated protein engineering campaign that improved glycoside hydrolase thermostability using Bayesian optimization, cell-free expression and gene fragment-based variant creation11. However, reliance on an external cloud lab adds a layer of opaqueness while the high cost of gene fragments, limitations of cell free systems, and enzyme-specific generation of variant libraries reduce the workflow’s overall versatility and scalability.
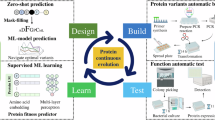
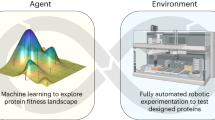
Here, we present a generalized platform for autonomous enzyme engineering enabled by the Illinois Biological Foundry for Advanced Biomanufacturing (iBioFAB) (Supplementary Fig. 1), ML, and large language models (LLMs) (Fig. 1). The process begins by designing a high quality mutant library using a protein LLM22 and epistasis model23. The library is constructed and screened by iBioFAB using optimized modular and integrated workflows. The assay data from each cycle is used for training a low-N ML model24 to predict variant fitness for subsequent iterations. Since this platform requires only a protein sequence and quantified fitness data, it can be applied to a wide array of proteins. As a proof of concept, in four rounds within 4 weeks, we engineer variants of two enzymes, Arabidopsis thaliana halide methyltransferase (AtHMT)25 and Yersinia mollaretii phytase (YmPhytase)26,27 with ~16- and 26-fold higher activity compared to the wild type enzymes, respectively.
The platform consists of three modules. First, a sequence-based unsupervised predictive model generates variants for the initial library. This variant library is created by an integrated biofoundry, a laboratory automation platform that combines various instruments for robust and continuous experimentation. Next, the biofoundry measures the fitness of the variants, and the resulting data is used to train a supervised ML model to predict subsequent variants. The seamless integration of protein language models, automated experimentation, and ML minimizes the need for human decisions and interventions during the protein engineering process. This autonomous protein engineering platform can design, test, and create improved enzymes without input from a human operator. Figure 1 was created with BioRender.
Results
Automating construction and characterization of protein variants on a biofoundry
The iBioFAB (Supplementary Fig. 1) has been used to automate various biological processes, such as pathway optimization13, yeast genome engineering28, TALEN assembly29, plasmid design and construction21, and natural product discovery30,31. In ML-guided protein engineering, sequencing and verifying the mutants generated through site-directed mutagenesis (SDM) often delays the process and increases costs. To design a robust and continuous workflow (Fig. 2a), we developed a HiFi-assembly based mutagenesis method (Fig. 2b) that eliminated the need for sequence verification in the middle of the protein engineering campaign, enabling an uninterrupted workflow. Throughout multiple rounds of protein engineering, some mutants were randomly selected, sequenced, and confirmed to have the correct targeted mutations with around 95% accuracy (Fig. 2c). All higher-order mutants are combinations of single mutants from the initial library, with one additional mutation added in each round. Variants with multiple mutations can be generated through SDM of a template plasmid containing one fewer mutation and, as such, there is no need to order new primers for each iterative cycle barring a few exceptions, thereby saving time and cost. This optimized high-fidelity approach was crucial for a reliable and continuous workflow during iterative cycles of protein engineering (Fig. 2a).
a The protein engineering workflow was initiated by predicting 180 mutants for the first round of screening in two 96-well plates. The data from each screening cycle was used for training a low-N supervised learning model to predict variants for subsequent cycles. After the first cycle, subsequent screening cycles used a single 96-well plate, containing 90 new variants with controls. b A site-directed mutagenesis protocol based on PCR and HiFi assembly was optimized for high fidelity. The PCR using mutagenic primers generates two DNA fragments that span the ORF and contain targeted mutation, followed by ligation with the linearized vector backbone that was digested by restriction enzyme (RE) using HiFi assembly. c We created 482 mutants for AtHMT, and DNA sequencing confirmed that 70 of 74 variants had the correct mutations. For YmPhytase, we created 448 mutants and 20 of 21 were correctly mutated based on sequencing. Overall, the success rate for mutagenesis was about 95% when picking only a single colony for each variant. d Overview of the laboratory automation pipeline used for autonomous protein engineering. The workflow was divided into smaller modules, programmed into iBioFAB instruments, and optimized for robustness and continuous operation. The automated workflow for a complete round of screening, from mutagenesis PCR to functional assay, is completed within 7 days (Supplementary Fig. 2) with each module designed to automate one of the protein engineering tasks. Each module is fully automated, which is achieved by integrating various instruments on the iBioFAB using Thermo Momentum Scheduling Software and Thermo F5 robotic arm. Detailed descriptions for each automated module are provided in supplementary information (Supplementary Figs. 2–5). Source data are provided as a Source Data File. Figure 2a–c was created with BioRender.
The protein engineering workflow was divided into manageable automated modules for robustness and ease of troubleshooting (Fig. 2d, Supplementary Fig. 2), allowing recovery without restarting the entire process. Seven automated modules comprise the end-to-end workflow for each cycle of protein engineering. Experimental steps within a module are completely automated, requiring little, if any, human intervention (Supplementary Fig. 2-6). Each module was individually programmed onto the iBioFAB platform and meticulously refined to ensure robust and reliable operation during continuous automated execution. Such examples include preparation of mutagenesis PCR, DpnI digestion, 96-well microbial transformations, plating on 8-well omnitray LB plates, crude cell lysate removal from 96-well plates, and functional enzyme assays. For automated execution, the instruments were scheduled via Thermo Momentum software and fully integrated by a central robotic arm. The robotic pipeline automates all protein engineering steps including mutagenesis PCR, DNA assembly, transformation, colony picking, plasmid purification, protein expression, and enzyme assays (Supplementary Fig. 2-6).
Design of protein variants using protein LLM and ML
The design of the initial library is crucial to the protein engineering campaign as a diverse and high-quality library increases the likelihood of efficient optimization32. To create a generally applicable autonomous protein engineering platform, we employed a combination of a protein LLM and an epistasis model. This approach maximized both the diversity and quality of the initial library, enhancing the chances of identifying promising mutants early in the process. We used a state-of-the-art protein LLM, ESM-222, a transformer model trained on global protein sequences. ESM-2 predicts the likelihood of amino acids occurring at specific positions based on sequence context, and the likelihood can be interpreted as variant fitness33. Additionally, we included an epistasis model, EVmutation23 focusing on the local homologs of the target protein.
To demonstrate the generality of our autonomous enzyme engineering platform, we selected two distinct enzymes, AtHMT and YmPhytase, with a goal of improving two different enzymatic properties. Both enzymes have rarely been studied using AI/ML tools, offer potential for industrial applications, and are suitable for automation-friendly high-throughput quantification. The halide methyltransferase AtHMT offers potential for synthesizing S-adenosyl-l-methionine (SAM) analogs, which are essential for biocatalytic alkylation, a process with crucial biological applications34,35. However, the chemical synthesis and commercial availability of SAM analogs are often limited. AtHMT has shown promising promiscuous alkyltransferase activity, allowing for the synthesis of SAM analogs from more readily available and cost-effective alkyl halides and S-adenosyl-l-homocysteine (SAH)25,36. We focused here on improving the ethyltransferase activity of AtHMT, seeking to improve its preference for ethyl iodide over methyl iodide. We secondly aimed to broaden the pH range for activity of YmPhytase, a phosphate-hydrolyzing enzyme that exhibits low activity at neutral pH26. Phytases can hydrolyze phytic acid, the main storage form of phosphate in plant tissues that is indigestible by most animals, and release inorganic phosphate, an important nutrient37. However, to function in animal feed, phytase must have high activity at a broad pH range to tolerate the pH variation of the gastrointestinal tract of animals. Most phytases function optimally in acidic conditions and exhibit a sharp loss of activity at neutral pH, motivating the development of phytase enzymes with improved activity closer to neutral pH. We therefore engineered YmPhytase, which has better activity than commonly used fungal phytases and has shown promising improvements in previously reported directed evolution campaigns26,27.
By combining the two unsupervised models, we generated a list of 180 variants each of AtHMT and YmPhytase for initial screening (Fig. 3a, b). We found that 59.6% of AtHMT and 55% of YmPhytase variants performed above the wild type baseline (Fig. 3c, Supplementary Figs. 7, 10), with 50% and 23% being significantly better, respectively (two-tailed Student’s t-test p < 0.05, Supplementary Figs. 7, 10). The mutants generated by EVmutation performed better than those predicted by ESM-2, although there was significant overlap between predictions of the two models (Supplementary Fig. 27). The best mutants from the unsupervised models’ predictions were V140T for AtHMT, with a 2.1-fold improvement, and V141M for YmPhytase, with a 2.6-fold improvement. For AtHMT, both models had independently identified V140T as one of the top ranked variants, which was reported as the best mutant in a previous semi-rational design study25. Whereas, for YmPhytase, the predictions from both models did not include previously identified sites T44 and K45 in top predictions26.
a, b The individual mutated amino acid (aa) residue predicted in each round at various sites of the protein are marked and color coded by their prevalence in the screened variants. The initial sequence-based models start with a wide search, followed by narrowing down by a low-N supervised learning model to a few select mutation sites in later rounds. The active site for AtHMT (P20, V23, L27, W36, W47, Y139, V140, C143, Y172, R214) and YmPhytase (R37, R41-T47, E241, D327) are marked by red lines on x-axis. The x-axis represents the mutation positions, and the y-axis represents the mutated amino acids. The exact amino acid sequence for both is available in the supplementary materials as sequenced DNA file. c The majority of variants predicted by unsupervised models for the first round of screening show higher activity than the wild type for both AtHMT and YmPhytase with at least one variant showing remarkably improvement. The black dashed line shows wild-type activity. d None of the new single mutants predicted by the supervised learning for the second round of AtHMT engineering show better activity than the wild-type enzyme. Black dashed line indicates WT activity and orange dashed line shows V140T activity. e None of the S99T/V140T-containing triple mutants, which would typically be designed by human intuition in a traditional directed evolution approach, show better activity than the top supervised machine-learning predicted triple mutants from the third round. A comparison between S99T/V140T-containing triple mutants (the ones designed by human intuition) with all of the third-round mutants, which were predicted by the machine-learning algorithm (left). Comparison of activities for six predicted third round mutants with best prediction scores which were assayed alongside the 36 human-intuited triple mutants containing S99T/V140T mutation (right) (n = 5 biological repeats for round 3 mutants and n = 4 biological replicates for S99T/V140T triple mutants, Gray dots Q1 (25%): 2.48, Median (50%): 2.74, Q3 (75%): 3.07 IQR: 0.60 Whisker Min: 2.09 Whisker Max: 3.76; Orange dots Q1 (25%): 2.60, Median (50%): 3.08, Q3 (75%): 3.39, IQR: 0.78, Whisker Min: 1.7, Whisker Max: 4.3). f, g The predicted fitness by the low-N supervised learning model and observed fitness of mutants for AtHMT and YmPhytase. The dashed black line indicates the wild-type activity for respective proteins. Red crosses indicate mutants with the best predicted fitness according to the supervised model. The Spearman rank correlation coefficient is displayed at the bottom of the graphs. Source data underlying the figures are provided as a Source Data File.
Starting from the second round, the data from all previous rounds were used to train a supervised low-N regression model24. Once trained, the model predicted variants with one additional mutation than the previous round for the subsequent screening cycle. While the unsupervised models started with a wide search space in the first round, the supervised learning model narrowed to focus on a few key mutations in the top 90 predictions for both proteins (Fig. 3a, b). We constructed the top 96 predicted mutants and 90 were used for enzyme activity screening along with controls (Supplementary Figs. 7, 10). Although there was little overall consistency between observed enzyme activities and the predictions of the low-N model, the model was nonetheless able to predict a subset of mutants with substantially improved activity in each round of engineering (Fig. 3f, g) demonstrating the observed practical effectiveness of the model. For the second round of AtHMT engineering, we predicted additional single mutations after training the low-N model using the data from the first round. The 14 new single mutant predictions performed poorly compared to the wtHMT (Fig. 3d, Supplementary Fig. 7) suggesting that the low-N model may not be well suited for finding new single mutation variants as compared to the unsupervised models used in the first round.
New insights through ML-guided protein engineering
ML-driven experimentation often leads to surprising results and unexpected directions, as shown by the model’s prediction of counter-intuitive variants during the third round of AtHMT engineering. None of the S99T/V140T-containing triple mutants were ranked in the top 100, despite S99T/V140T being the best double mutant from the second round. We tested the top 36 ranked triple mutants containing S99T/V140T and compared them against the model’s top predictions for triple mutants. Most S99T/V140T-based triple mutants failed to significantly improve activity over the V140T mutant with only 4/36 (11%) showing significantly better activity than V140T (two-tailed Student’s t-test p < 0.05, Fig. 3e, Supplementary Fig. 8), whereas the model-predicted triple mutants showed superior performance with 74/90 (82%) performing significantly better than V140T (two-tailed Student’s t-test p < 0.05, Fig. 3e, Supplementary Fig. 7). This suggests the ML model can potentially recognize synergistic effects between mutations, predicting more effective combinations than human-designed mutants.
We observed an increase in the desired activity (Fig. 4a, b) throughout all four engineering cycles (Fig. 4c, d, Supplementary Figs. 7, 10). The fitness of variants improved in each round compared to the wild type for both AtHMT and YmPhytase (Fig. 4c, d). In four rounds of protein engineering, we screened 446 variants of AtHMT and 448 variants of YmPhytase using a crude lysate assay. For AtHMT, the top performing variant in each subsequent round was 2.1-, 3-, 4.3- and 5.1-fold better than wtHMT in the crude lysate assay (Supplementary Figs. 7, 9). For YmPhytase, the top performing variant in each subsequent round was 2.6-, 7.5-, 9- and 11.1-fold better than wtPhy in the crude lysate assay (Supplementary Fig. 10). We purified some of the best performing mutants from each round and characterized the enzyme kinetics (Supplementary Figs. 11–26, Supplementary Table 1–6). The best identified AtHMT variant (round 4: V140T/L101I/E206D/S63N) showed an ~16-fold improvement in ethyltransferase activity compared to wtHMT (Fig. 4e, Supplementary Figs. 13–14, Supplementary Table 3). Another AtHMT variant (round 4: V140T/L101I/T213S/V178E) showed an ~90-fold increase in preference for ethyl iodide over methyl iodide (Supplementary Table 1–2, 4).
a Two sequential reactions were used to quantify the activity of AtHMT: (I) the HMT-catalyzed release of iodide from ethyl iodide and SAH and (II) colorimetric quantification of iodide by measuring absorbance at 570 nm. b The phytase activity assay measured 4-MUP hydrolysis to 4-MU by monitoring fluorescence (λex = 354 nm / λem = 465 nm). The wild type exhibits a sharp decrease in activity at pH 6.6. (n = 4 biological replicates) (c) The fold change in HMT ethyltransferase activity with respect to the wild type is shown in a violin plot for four rounds of crude lysate screening (n ≥ 3 biological replicates). The source data is available in Supplementary Data 3. d The fold change in phytase activity with respect to the wild type is shown in a violin plot for four rounds of crude lysate screening (n ≥ 3 biological replicates). The source data is available in Supplementary Data 4. For detailed plots for each screening round of AtHMT and YmPhytase see Supplementary Figs 7, 10. e, f A selection of the best performing mutants from each screening round were purified and specific activity was calculated at (e) 15 mM ethyl iodide for AtHMT (n = 3 biological replicates) and (f) at 0.9 mM 4-MUP for YmPhytase (n = 4 biological replicates). R1P1 and R1P2 denote plates 1 and 2 respectively from the first cycle of protein engineering. Labels with R2, R3, and R4 denote subsequent screening rounds (e.g. R2A1 denotes well A1 from the second round). The mutations present in each variant can be found in Supplementary Data 3 and 4. Full kinetic characterization of the AtHMT (Supplementary Figs. 12–14) and YmPhytase (Supplementary Figs. 20–26) mutants are provided as Supplementary Figures. Error bars (b−f) represent the standard deviation (SD) and the data are presented as mean values +/- SD. Source data underlying the figures are provided as a Source Data File.
For YmPhytase, one variant from the fourth round (round 4: V141M/K226G/I15V/Q362R) exhibited a 26.3-fold improvement in specific activity over wtPhy at pH 6.6 (Fig. 4f). Another variant (round 4: V141M/K226G/I15V/E316P) performed remarkably well at both pH 5.6 and 6.6, exhibiting 25.8- and 19.8-fold improvements in specific activity compared to wtPhy, respectively (Fig. 4f, Supplementary Fig. 18). All proteins selected for purification showed a higher specific activity (Supplementary Table 5) and improved kinetic parameters (Supplementary Table 6) compared to the wild type at pH 4.5, 5.6, and 6.6. Since the screening involved several hours to obtain complete data from multiple replicates, the cell lysates contained a protease inhibitor cocktail in the crude lysates. After purification of selected phytase variants we noticed that the addition of the protease inhibitor cocktail, and more specifically one of its components 4-(2-aminoethyl) benzenesulfonyl fluoride hydrochloride (AEBSF), was critical to observe high activities of YmPhytase enzyme variants (Supplementary Fig. 19). In a previous report, it was also noted that purified variants were less active compared to when assayed using crude lysates26. Screening conditions are therefore crucial to the design of the protein engineering campaign, as proteins become optimized for the specific assay environment—emphasizing the old adage “You get what you screen for”38.
In the previous work that identified V140T as an improved variant25, 10 residues in the active sites were screened using NNK site-saturation libraries. Further screening by generating double and triple mutant libraries combining active site residues didn’t yield a better mutant than V140T25. In the current work, V140T is present in all third and fourth round variants and is the only mutation in the active site. All other mutation sites in top performing mutants (L101, T213, V195, E206, D9, E32) from the fourth round of engineering are located relatively far from the active site and on the surface of the protein (Supplementary Movie 1). Moreover, the majority of fourth round variants show significantly improved activity over V140T alone.
In contrast, our YmPhytase engineering yielded a new mutation V141M during the first round of screening that significantly improved activity compared to the wild type and was carried over into all subsequent iterative cycles. None of the top predictions by the unsupervised models included the previously identified T44/K45 sites discovered by semi-rational design26. Since the YmPhytase crystal structure was not available, it was predicted using AlphaFold39. The majority of variants in the fourth round contain three mutations V141M, K226G, and I15V, of which V141 and K226 residues are close to the active site, while I15 is located at a more distant flexible region. Most other common mutation sites in the fourth round such as Q167, T25, L303 are distant from the active site, while A65 is located close to the active site (Supplementary Movie 2).
Language model-based user interface
As mentioned above, we demonstrated that using unsupervised protein mutation prediction tools and supervised regression models can significantly speed up protein engineering without requiring domain expertise in rational design. To further simplify access, we developed a natural language-based user interface utilizing LLMs (Fig. 1). This allows users without programming skills to easily interact with our protein engineering platform. We created a user interface, leveraging OpenAI’s assistant API with customized functions, to design the initial library using simple language commands, like “help me improve phytase” or “design the initial library for AtHMT”, especially for users without required coding skills. The system responds by accessing pre-programmed functions to execute sequence-based predictions and train regression models, making the platform accessible to non-technical users.
Discussion
Through systematic development and optimization, we have demonstrated a generally applicable autonomous enzyme engineering platform which accepts a protein sequence as the sole input and requires minimal human decision-making or specialized protein knowledge. Without the need for human judgement, the automated platform is fully scalable and compatible with a wide array of proteins. Our natural language user interface further reduces barriers to entry, allowing users to operate the platform without specialized coding knowledge. We used two comprehensive case studies, AtHMT and YmPhytase, to demonstrate the automated enzyme engineering platform in action. Starting with only the wild type sequences, our approach autonomously performed successful engineering campaigns for both proteins by identifying variants with 16-fold improvements for AtHMT and 26-fold improvements for YmPhytase within four iterative cycles of protein engineering. We assayed the previously identified best mutants for AtHMT (V140T)25 and YmPhytase (T44V/K45E)26 alongside the top variants identified in this study. For both AtHMT and YmPhytase, the most active variants discovered in this study demonstrate a remarkable improvement compared to the previous engineering reports of these enzymes25,26,27.
A few previous attempts have prototyped automated experimentation using Bayesian optimization for traversing a relatively limited search space11,13,40 for protein engineering (Supplementary Table 7). With the unimaginably vast and high-dimensional sample space of proteins, state-of-the-art AI tools are better suited for efficient exploration41. A recent study11 used a cell-free expression system, which is challenging to generalize due to many issues such as limited protein yield, poor protein folding, instability and degradation. Supplementary Table 7 provides a concise comparison with prior automated protein engineering studies. In many protein engineering campaigns, initial variant libraries are tailored for proteins of interest and require deep expertise about structure and function of the target. In contrast, we present a generalizable framework that can predict variants with only the protein amino acid sequence for the initial library and fitness data for iterative engineering cycles. This coupled with our stepwise approach for mutagenesis offers a broadly applicable and cost-effective solution for autonomous enzyme engineering. Following one of the frameworks developed to evaluate levels of automation20, we assess that our current autonomous enzyme engineering platform falls between level 3 (conditional automation) and level 4 (high automation), as it can perform end-to-end protein engineering with minimal human intervention and predict variants for screening in subsequent cycles.
The modularity of our framework makes possible substantial expansions and improvements in the future. With advancements in the state-of-the-art AI models for protein engineering, we expect this framework to improve as new models can be integrated into this platform. The exploration of higher-order mutants with substantially greater numbers of mutations becomes feasible with this platform, a challenge that has proven difficult to address especially for in vitro protein engineering studies. Moreover, while we used crude lysate assays in this study, this framework is compatible with many quantified measurements of protein activity, including in vitro, in vivo, growth-coupled, or reporter-based assays. By incorporating this flexibility, we have developed a general-purpose autonomous protein engineering platform that maximizes its potential applications.
Nonetheless, our platform still contains several limitations. On the computational side, while we use ESM-2 and EVmutation for initial variant predictions, ESM-2 has performed poorly in some benchmarks and may not be suitable for all proteins42 and EVmutation depends on the availability of homologous sequences23. Applying our platform to proteins that are not well-suited for these models may result in poorly-performing initial variant libraries. Moreover, the supervised low-N model used for iterative predictions showed relatively weak correlation with experimental results, suggesting the possibility that certain proteins may perform particularly poorly using such a predictive model. Our site-directed mutagenesis method employing PCR and HiFi assembly to accumulate mutations in a stepwise manner also contains some limitations. Recycling of mutagenesis primers limits the implementation of simultaneous multiple mutations in adjacent residues. As such, implementing multiple mutations in adjacent residues may require the design and purchase of new mutagenesis primers which contain desired mutations. High GC-content sequences may also lead to difficulty with PCR amplification and HiFi assembly. We have successfully engineered two enzymes so far using the autonomous platform and extending it to additional proteins could further broaden its generalizability. Additionally, our platform requires a functional assay that is compatible with high throughput automation. Here, we focused specifically on enzymes with established high throughput quantitative assays that are functional in crude cell lysates. The suitability of our platform for transporters, chaperones, transcription factors, and other types of proteins remains unexplored at this time and would require the development of suitable functional assays.
Autonomous experimentation platforms have the potential to transform protein engineering and related fields including synthetic biology43, biocatalysis44, metabolic engineering45, and retrobiosynthesis46. The rapid progress of laboratory automation, such as liquid-handling robots47, microfluidics48, laboratory virtualization services49, and increasing global investment into biofoundries50 will make these platforms more accessible to researchers. The integration of ML tools and robotic laboratory platforms offers round-the-clock operation, improved reproducibility, and increased efficiency while minimizing the need for human decision-making. Various applications, including the design of new-to-nature enzymes51, discovery of novel natural products30,31, enhancement of drug and biomolecule efficacy, and the optimization of products for sustainable chemistry44, stand to benefit greatly from the transformative power of autonomous experimentation. The broadly applicable framework presented here for protein engineering forms a crucial component of the autonomous research toolbox, establishing a strong foundation for the future of synthetic biology.
Methods
LLM and supervised learning for prediction of protein variants
Two different zero-shot prediction models were used to guide the design of the initial variant library: EVmutation and ESM-2. EVmutation is a statistical model designed by Hopf et al.23 that captures the co-variations between pairs of residues in an amino acid sequence. This is achieved by fitting a pairwise graph model to the multiple sequence alignment (MSA) of the homologous to the target protein. The model then scores the impact of amino acid substitutions by calculating the log-ratio of sequence probabilities between the mutant and wild-type sequence. In this work, we utilized the integrated web server EVcouplings52 with the default searching parameters and highlighted bit-scores. ESM-2 is a transformer-based protein language model designed by Rives et al.53 that is trained on large and diverse protein sequence database and captured the rules governing the protein structure and functions. ESM-2 can output the probability of a certain amino acid occurring at a given position based on the up and down stream context. Thus, we can score any given mutation by using wild-type as a reference and compare the probability of the given mutation with that of the wild-type. In this work, we used the workflow implement in ESM-2 (https://github.com/facebookresearch/esm) to make the prediction with the model esm2_t36_3B_UR50D.
For each round of engineering, we trained a supervised prediction model based on all experimentally measured variant fitness data from current round and all rounds prior. We first preprocess raw data by removing all zero or negative variants and normalize the data by taking the log scale. In this work, we trained the supervised model by modifying the workflow implement by Hsu et al.24 (https://github.com/chloechsu/combining-evolutionary-and-assay-labelled-data). We initially trained the model using augmented Potts and ESM approach, but the evolutionary density score feature has negatively correlated with experimental results. Thus, we modified the original training script and used the command ‘python src/evaluate.py phytase onehot –n_seeds = 1 –n_threads = 1 –n_train = -1’ for training and prediction.
GPT-based user interface
To implement a Generative Pre-trained Transformer (GPT)-based user interface, OpenAI’s assistant Application Programming Interface (API) (https://platform.openai.com/docs/assistants/overview) was employed. The assistant was configured using the ‘gpt-4-turbo’ model. The user interface carries tools of file-search (https://platform.openai.com/docs/assistants/tools/file-search) and function-calling (https://platform.openai.com/docs/assistants/tools/function-calling). The user interface will invoke file searching to respond to user’s general questions and invoke function calling to assist the design of initial variant library.
Expression plasmids
The HMT and CiVCPO plasmids were a gift from Uwe Bornscheuer lab. Both the HMT plasmids and empty vector (pET-28a) were transformed in Escherichia coli BL21 (Δmtn) strain, a gift from Uwe Bornscheuer lab, which is an MTA/SAH nucleosidase knockout E. coli BL21(DE3) strain25. The phytase cDNA was ordered from Twist Biosciences and cloned in pET-28a vector using HiFi assembly. These plasmids were sequence verified using Plasmidsaurus (San Francisco, CA, USA). Complete sequence of these plasmids are provided in Supplementary Data 5.
Site-directed mutagenesis
For site-directed mutagenesis, three PCR fragments were assembled using HiFi assembly (Fig. 2b). The mutagenesis primers (27 base-pairs, 12 + 3 + 12) were designed using a Python script and ordered in 96-deepwell plates from IDT. For PCR templates, the plasmids were linearized with restriction enzymes and purified using a PCR cleanup kit (Zymo #D4018). The HMT plasmid was linearized with EcoRV (for mutagenesis PCR of the ORF) and XbaI (for backbone PCR). The phytase plasmid was linearized with HpaI (for mutagenesis PCR of the ORF) and XhoI (for backbone PCR). The PCR fragments contained 27-40 overlapping base pairs. The PCR for the vector backbone was treated with DpnI (37 °C for 4 h, followed by overnight incubation at room temperature) and purified using a PCR cleanup kit (Zymo #D4018), and stored at -20 °C until used. The mutagenesis PCR for forward and reverse fragments was run in separate 96-well plates, followed by DpnI treatment. Q5 DNA polymerase (NEB #M0491) was used for mutagenesis PCR (50 µL reaction, ~200 pg template, 18 cycles, 65 °C 30 s, and 72 °C 1 min). We observed that NEB Q5 polymerase can tolerate a wide range of theoretical annealing temperatures, thus all mutagenic PCRs were annealed at 65 °C for ease of automation. Ensuring correct amount of input template DNA was critical to successful PCRs and efficient SDM, thus the DNA concentration was measured using Invitrogen Qubit Fluorometer with dsDNA Quantitation kit (Invitrogen #Q32853) and further verified by agarose gel electrophoresis. The primers were ordered from IDT in 96-well plate at 2 uM. For PCRs, 12.5 ul of primers were added to 96-well PCR plates on Tecan Fluent. After PCR, 2.5 µL of the reaction was mixed with 50 µL of 1x EvaGreen Dye (Biotinum #31000) and fluorescence (λex = 488 nm / λem = 535 nm) was measured to verify successful PCR.
HiFi assembly reactions (15 µL) were prepared using 30 ng of purified vector backbone PCR, 1.25 µL each of DpnI treated forward and reverse PCR reactions, and 7.5 µL of HiFi master mix (NEB #E2621) followed by incubation at 50 °C for 30 min. To increase the efficiency of HiFi assembly, 50 ng of single-strand binding protein (NEB #M2401) was added to 500 µL HiFi master mix54. Homemade competent DH5α cells in a 96-well plate (Biorad # HSS9641) were transformed with 5 µL of the HiFi reaction by 30 s heat shock, followed by 1 h outgrowth in 150 µL SOCS media. DH5α outgrowth cultures were spread on 8-well omnitray plates (120 µL per well, kanamycin 50 µg/mL) and incubated overnight at 37 °C. DH5α colonies were picked using Pickolo and incubated overnight at 37 °C in a 96-deepwell plate containing 1 mL of Terrific Broth (TB) and kanamycin. Minipreps were then performed using PureLink Pro Quick96 Plasmid Purification Kit (Invitrogen #K211004A). Miniprep DNA of some mutants from the 96-well plate was sent for sequencing. The miniprep DNA was used to transform competent BL21 cells in a 96-well plate for further protein expression and functional assays.
E. coli heat shock competent cell preparation in 96-well plate
DH5α, BL21, and BL21(Δmtn) competent cells were prepared in the lab and stored in a high-profile 96-well PCR plate (Bio-Rad #HSS9641) at -80 °C. Briefly, an overnight preculture (no antibiotics, 37 °C) was started from a single colony. A 250 mL LB media was inoculated with 5 mL of the starter culture and grown at 30 °C until the OD reached 0.4–0.6, and then transferred to cold room (4 °C) for 1 h. The cells were centrifuged at 1000xg for 20 min at 4 °C. The cells were then gently resuspended in Buffer RF1 (100 mM rubidium chloride, 50 mM manganese chloride, 30 mM potassium acetate, 10 mM calcium chloride, 15% w/v glycerol, pH 5.8), incubated at 4 °C for 30 min, then centrifuged. Subsequently, the cells were resuspended in 10 mL of RF2 buffer (10 mM MOPS, 10 mM rubidium chloride, 75 mM calcium chloride, 15% w/v glycerol, pH 5.8). 50 µL was aliquoted into each well of a 96-well plate in a cold room, followed by sealing and snap-freezing in liquid nitrogen. The cells were stored at -80 °C until used.
Iodide detection assay for the ethyltransferase activity of HMT
We followed the iodide quantification protocol developed by Tang et al.25 that can specifically detect iodide and is insensitive to chloride concentration as described here. For HMT screening in 96-well plates, the iodide detection reagent consisted of 1 μL purified chloroperoxidase from Curvularia inaequalis (CiVCPO, 0.75 mg/mL) and 79 μL 3,3′,5,5′-tetramethylbenzidine (TMB, Sigma #T0565), to which 20 μL of HMT reaction I (see below) was added. For enzyme kinetics of purified HMT proteins, the iodide detection reagent contained 47 μL TMB, 0.5 μL CiVCPO (0.75 mg/mL), and 2.5 μL HMT reaction I. For the standard curve (Supplementary Fig. 13), 2.5 μL of potassium iodide (KI) at varying concentrations (5 μM to 500 μM) was added to 47 μL TMB and 0.5 μL CiVCPO (0.75 mg/mL), and absorbance at 570 nm was measured (Supplementary Fig. 11) using a Tecan Infinite plate reader. Variance of iodide detection assay for AtHMT activity in 96-well plate was measured using wild-type and V140T mutant (Supplementary Fig. 6).
High throughput screening for ethyltransferase activity of HMT
Single colonies of HMT mutants in BL21 (Δmtn) cells were picked and inoculated into 800 μL LB broth supplemented with 50 μg/mL kanamycin for preculture. From preculture, 300 μL was used for glycerol stocks and stored at -80 °C. Then, 50 μL of the preculture was inoculated into 1 mL of TB (96- deepwell plate, 50 μg/mL kanamycin, 100 μM isopropyl β-D-1-thiogalactopyranoside (IPTG)) and incubated overnight at 30 °C, 900 rpm in Cytomat automated shaking incubator. Cells were pelleted at 2900xg for 15 min, then lysed in 300 μL of lysis buffer (1.5 mg/mL lysozyme, 0.1x Bugbuster reagent, 10 μg/mL DNase I, 1x Halt Protease inhibitor, 50 mM sodium phosphate buffer, pH 7.5). Lysis occurred at 30 °C for 1.5 h at 900 rpm, followed by centrifugation for 20 min at 2900xg at room temperature.
For Reaction I, 160 μL of crude cell lysate was mixed with 20 μL SAH (1 mM) and 20 μL ethyl iodide (5 mM), both freshly prepared in DMSO. After 1 h incubation at room temperature, 20 μL of Reaction I was added to 80 μL of Reaction II (1 μL CiVCPO + 79 μL TMB), and absorbance at 570 nm was measured using Tecan Infinite plate reader. A Python script calculated the slope of the increase in absorbance for each well, then calculated the fold change relative to the wild type. Each screening plate contained 90 mutants with controls (two wells each of wild type, V140T, and empty vector). For screening V140T/S99T triple mutants, each 96-well plate contained duplicates of 36 triple mutants containing V140T/S99T, six triple mutants from the third round, and the same controls as above.
HMT kinetics assay
The Michaelis-Menten kinetics for HMT and its mutants were determined for ethyl iodide and methyl iodide. A 50 μL reaction was prepared with 1 mM SAH, 10 μL haloalkane, and purified HMT in 50 mM sodium phosphate buffer (pH 7.5). To prevent hydrolysis, 5x haloalkane stocks were freshly prepared in DMSO. For ethyl iodide, purified wild-type HMT (wtHMT) was used at a final concentration of 87 μM (2.4 mg/mL), V140T, round 2, and round 3 mutants at 29 μM, and round 4 mutants at 8.7 μM. For methyl iodide, wtHMT was used at 0.145 nM, and V140T, round 2, round 3, and round 4 mutants at 0.29 nM. After a 10 min incubation, 2.5 μL of the above reaction was added to the iodide assay reagent (0.5 μL CiVCPO + 47 μL TMB), and absorbance at 570 nm was measured to determine iodide concentration. For the kinetic assay, the concentration of haloalkanes was varied while keeping HMT and SAH concentration constant. The specific activity was calculated at 15 mM ethyl iodide for all purified HMT proteins. Autohydrolysis rates for each haloalkane concentration were simultaneously measured and subtracted from enzymatic reaction rates. The amount of iodide produced per minute was calculated using the standard curve (Supplementary Fig. 9) and fit to the Michaelis-Menten model using Origin software. All reactions were performed in triplicate.
Purification of HMT proteins
We followed the published protocol25 for purifying chloroperoxidase from Curvularia inaequalis (CiVCPO) and stored in 50 mM sodium phosphate buffer (pH 8.0) with 100 µM sodium orthovanadate. For the iodide detection assay, 0.375 mg CiVCPO was added per 50 μL reaction.
For HMT purification, starter cultures were prepared from glycerol stocks of mutant libraries in BL21 Δmtn (DE3) strain. For protein expression, a 1:100 preculture was added to TB (50 μg/mL kanamycin) and grown at 37 °C until the OD600 reached ~0.6. Cells were cooled and incubated overnight at 20 °C with 200 μM IPTG. The cells were harvested by centrifugation at 10,000 x g at 4 °C for 10 min, and the pellet was resuspended in lysis buffer (20 mM sodium phosphate, 0.5 M sodium chloride, 20 mM imidazole, 1× PIC, pH 7.5). Cells were lysed by sonication (10 min on ice, 20% amplitude), followed by centrifugation (12,000xg, 20 min at 4 °C). His6-tagged proteins were purified using immobilized metal-affinity chromatography (IMAC). Lysates were clarified through a 0.45 μm filter, and 2 mL of pre-washed nickel beads (Nuvia™ IMAC Resin, Bio-Rad #7800800) were added. After 45 min of incubation and mixing at 4 °C, the lysates were centrifuged at 2900xg, and the supernatant was discarded. The affinity beads were transferred to a column and washed with 10x volume of lysis buffer. Proteins were eluted with elution buffer (20 mM sodium phosphate, 0.5 M sodium chloride, and 0.2 M imidazole, pH 7.5), then desalted using PD-10 desalting columns (Cytiva #17085101) and Amicon Ultra 10 kDa centrifugal filters. The proteins were stored in 50 mM sodium phosphate (pH 7.5), and concentrations were determined using absorbance at 280 nm with a Nanodrop, using the HMT extinction coefficient (42,065 M−1cm−1). Purity was analyzed via SDS-PAGE (Supplementary Fig. 15).
High throughput screening for phytase activity using 4-MUP assay
Single colonies of phytase plasmids in BL21 cells were picked and inoculated into 800 μL LB broth supplemented with 50 μg/mL kanamycin for preculture in a 96-deepwell plate. 1 mL of autoinduction media (15 g/L peptone, 30 g/L yeast extract, 6.25 mL/L glycerol, 90 mL 1 M potassium phosphate buffer pH 7, 10 mL glucose (50 g/L), 100 mL lactose (20 g/L), and 50 μg/mL kanamycin) was inoculated with 5 μL aliquot of the preculture in a 96-deepwell plate. Phytase was expressed overnight (16 h, 37 °C, 900 rpm) in a Cytomat automated shaking incubator. Cells were pelleted by centrifugation (2900xg, 15 min, room temperature) and resuspended in 200 μL lysis buffer (1 mg/mL lysozyme, 50 mM Tris-HCl buffer, pH 7.5). Lysis was carried out by incubating at 37 °C, 900 rpm for 1 h, followed by centrifugation to remove cell debris.
Phytase activity was measured using a fluorescence-based 4-MUP assay26,27. Cleared cell lysate (10 μL) was added to 90 μL of 1.11 mM 4-MUP (4-methylumbelliferyl phosphate, Sigma #M8168) in Tris-maleate buffer (0.2 M, pH 6.6) in black, clear-bottom plates. The increase in fluorescence (λex = 354 nm / λem = 465 nm) was measured over time using a Tecan Infinite M1000 plate reader. Each screening plate contained 90 mutants and six control wells (wild type, M16 (T44V/K45E) mutant, empty vector, and the best mutant from the previous round). The slope of the fluorescence increase was calculated for each well using a Python script, with empty vector values subtracted at each time point for normalization.
Standard curve for 4-MUP assay
4-Methylumbelliferone (4-MU, Sigma #M1381) was dissolved into methanol and 50 µL of dissolved 4-MU solution was mixed with 50 µL of appropriate pH buffer. For pH 4.5, the buffer used was 0.25 M sodium acetate, 1 mM calcium chloride, 0.01% Tween-20. For pH 5.6 and 6.6, 0.2 M tris maleate buffer was used. Fluorescence (λex = 354 nm / λem = 465 nm) was measured using Tecan Infinite plate reader and plotted against 4-MU concentrations to generate standard curves for each pH (Supplementary Fig. 17).
Phytase kinetics assay
Kinetics of phytase variants was determined using a 4-methylumbelliferyl phosphate (4-MUP) assay as previously described with some modifications5 as described here. Assays at pH 5.6 and 6.6 were performed in 0.2 M tris maleate buffer while assays at pH 4.5 were performed in 0.25 M sodium acetate, 1 mM calcium chloride, 0.01% Tween-20 buffer. Standard curves for pH 4.5, 5.6 and 6.6 were prepared for different concentrations of 4-MU (4-methylumbelliferone), the fluorescent product of 4-MUP (pH 4.5: 2.5-640 μM; pH 5.6: 2.5-320 μM; pH 6.6: 3.125-40 μM) and can be viewed in Supplementary Fig. 15. 50 μL of 4-MU dissolved in methanol was mixed with 50 μL of the specified buffer followed by fluorescent measurement with λex 354 nm and λem 465 nm on a Tecan INFINITE plate reader. To prepare enzymes for kinetic assays, proteins were first mixed with 1 mM AEBSF (4-benzenesulfonyl fluoride hydrochloride) and incubated at 37 °C for 30 min. 90 μL of 4-MUP substrate in respective buffer was then mixed with 10 μL of the enzyme solution followed by kinetic measurement on the TECAN infinite. Fluorescence data was converted to mM 4-MU using the appropriate standard curves then divided by the enzyme concentration and used to calculate initial reaction rates. As Yersinia mollaretii phytase is a tetrameric enzyme with allosteric interactions, initial reaction rates were fit using OriginLab graphing software to the Hill equation y = Vmax * x / (Khalf + x) where y is the initial reaction rate, Vmax is the maximum reaction rate, X is the substrate concentration, and Khalf is the substrate concentration that enables half the maximum reaction rate.
Purification of phytase proteins
For phytase protein purification, BL21 glycerol stock from each round was used to inoculate a starter culture in LB medium followed by growth at 37 °C for 12–16 h. 5 mL of starter cultures were added to flasks containing 500 mL Terrific Broth (6 g tryptone, 12 g yeast extract, 2 mL glycerol, 0.085 mol potassium dihydrogen phosphate, 0.36 mol dipotassium hydrogen phosphate) supplemented with 50 μg/mL kanamycin. Flasks were then shaken at 200 rpm and 37 °C for 12–16 h. Following growth, protein expression was induced by adding 100 µM IPTG and culturing for an additional 4 h at 200 rpm and 37 °C. The cells were harvested by centrifugation at 10,000xg at 4 °C for 10 min, and the pellet was resuspended in lysis buffer (20 mM sodium phosphate, 0.5 M sodium chloride, 30 mM imidazole, 1× Protease Inhibitor Cocktail (Sigma #P8849), pH 7.5) supplemented with 1 mg/mL lysozyme. Cells were then lysed for 1 h at 37 °C followed by centrifugation (12,000xg, 20 min at 4 °C). His6-tagged proteins were purified using immobilized metal-affinity chromatography (IMAC). Lysates were clarified through a 0.45 μm filter, and 3 mL of pre-washed nickel beads (Nuvia™ IMAC Resin, Bio-Rad #7800800) were added. After 45 min of incubation and mixing at room temperature, the lysates were centrifuged at 2900xg, and the supernatant was discarded. The affinity beads were transferred to a column and washed with 10x volume of lysis buffer. Protein was eluted by adding 10 mL elution buffer (20 mM sodium phosphate, 0.5 M sodium chloride, and 0.2 M imidazole, pH 7.5). Protein was concentrated and the buffer was exchanged to 0.25 M sodium acetate, pH 5.5, 1 mM calcium chloride, 0.01% Tween-20 using Amicon Ultra 10 kDa centrifugal filters. Concentrations were determined using absorbance at 280 nm with a Nanodrop and enzymes were stored at 4 °C. Purity was analyzed via SDS-PAGE (Supplementary Fig. 16).
PAGE gels for purified proteins
For SDS-PAGE, the purified proteins were mixed with 1x Laemmli sample buffer (Biorad #1610737) with 5% 2-mercaptoethanol (Sigma #M6250) and boiled at 95 °C for 5 min. An equal amount of each sample was then loaded onto a 10% precast gel (Biorad #4561034) with protein ladder (Biorad # 1610374) and run using Tris/Glycine/SDS running buffer (Biorad # 1610732). After Coomassie staining, the gels were thoroughly washed before imaging. For native gel for phytase proteins, the purified proteins were mixed with native sample buffer (Biorad #1610738) and equal amount was loaded onto 7.5% gel (Biorad #4568024) and run with Tris/glycine running buffer (Biorad # 1610771). NativeMark unstained protein standard (Invitrogen # LC0725) ladder was used for native gel.
Worklist generation and laboratory automation
The seamless integration of various modules required generating worklists for PCR, which were created using Python scripts. Each round of mutagenesis added one additional mutation to plasmids generated in the previous round, such that the variants in the third round of evolution contained three total mutations. Thus, linearized plasmids containing mutations from one round of screening serve as templates for the next round. A Python script efficiently selected the minimal number of PCR templates needed for the next cycle. The linearized PCR templates were transferred to a 384-well plate, and a worklist was generated to distribute them into a 96-well PCR plate. Another worklist was created to distribute the primers using Tecan Fluent in 96-well PCR plates. Primers were ordered in a 96-well format from IDT and diluted to 2 μM stocks with 12.5 μL being added to each PCR reaction. Worklists were also used for spreading the transformed E. coli cells in 8-well omnitray LB plates. Laboratory automation was facilitated using the iBioFAB facility, which employs a Thermo F5 robotic arm to connect instruments and Thermo Momentum Scheduling software to execute integrated and automated experimental workflows. The comprehensive descriptions about instrumentation and automation are provided in Supplementary Fig. 2-5.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data for all figures and tables are included in the Source Data File. The AtHMT and YmPhytase mutant functional assay screening data generated in this study are provided in Supplementary Data 3 and 4, respectively. The plasmids created during this research are available upon request. Correspondence and requests for materials should be addressed to Huimin Zhao. The corresponding author will respond within a month for any plasmid requests. Source data are provided with this paper.
Code availability
The source code for zero-shot predictions and supervised learning model is available at https://github.com/Zhao-Group/closed-loop. The python scripts used for primer design and worklist generation are available at https://github.com/Zhao-Group/Primer_Design_and_Worklists. Both repositories can also be found at https://zenodo.org/records/15243671 and https://zenodo.org/records/15233300.
References
Martin, H. G. et al. Perspectives for self-driving labs in synthetic biology. Curr. Opin. Biotechnol. 79, 102881 (2023).
Tom, G. et al. Self-driving laboratories for chemistry and materials science. Chem. Rev. 124, 9633–9732 (2024).
Dai, T. et al. Autonomous mobile robots for exploratory synthetic chemistry. Nature 635, 890–897 (2024).
King, R. D. et al. The automation of science. Science 324, 85–89 (2009).
Boiko, D. A., MacKnight, R., Kline, B. & Gomes, G. Autonomous chemical research with large language models. Nature 624, 570–578 (2023).
MacLeod, B. P. et al. Self-driving laboratory for accelerated discovery of thin-film materials. Sci. Adv. 6, eaaz8867 (2020).
Szymanski, N. J. et al. An autonomous laboratory for the accelerated synthesis of novel materials. Nature 624, 86–91 (2023).
Burger, B. et al. A mobile robotic chemist. Nature 583, 237–241 (2020).
Carbonell, P., Radivojevic, T. & García Martín, H. Opportunities at the intersection of synthetic biology, machine learning, and automation. ACS Synth. Biol. 8, 1474–1477 (2019).
Yu, T., Boob, A. G., Singh, N., Su, Y. & Zhao, H. In vitro continuous protein evolution empowered by machine learning and automation. Cell Syst. 14, 633–644 (2023).
Rapp, J. T., Bremer, B. J. & Romero, P. A. Self-driving laboratories to autonomously navigate the protein fitness landscape. Nat. Chem. Eng. 1, 97–107 (2024).
Thurow, K. Automation for life science laboratories. Adv. Biochem Eng. Biotechnol. 182, 3–22 (2022).
HamediRad, M. et al. Towards a fully automated algorithm driven platform for biosystems design. Nat. Commun. 10, 5150 (2019).
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. Int. Ed. 57, 4143–4148 (2018).
Brannigan, J. A. & Wilkinson, A. J. Protein engineering 20 years on. Nat. Rev. Mol. Cell Biol. 3, 964–970 (2002).
Romero, P. A. & Arnold, F. H. Exploring protein fitness landscapes by directed evolution. Nat. Rev. Mol. Cell Biol. 10, 866–876 (2009).
Wang, Y. et al. Directed evolution: methodologies and applications. Chem. Rev. 121, 12384–12444 (2021).
Lutz, S. & Iamurri, S. M. Protein engineering: past, present, and future. In Protein Engineering: Methods and Protocols (eds. Bornscheuer, U. T. & Höhne, M.) 1–12 (Springer, 2018).
Yang, J., Li, F.-Z. & Arnold, F. H. Opportunities and challenges for machine learning-assisted enzyme engineering. ACS Cent. Sci. 10, 226–241 (2024).
Stephenson, A. et al. Physical laboratory automation in synthetic biology. ACS Synth. Biol. 12, 3156–3169 (2023).
Enghiad, B. et al. PlasmidMaker is a versatile, automated, and high throughput end-to-end platform for plasmid construction. Nat. Commun. 13, 2697 (2022).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Hopf, T. A. et al. Mutation effects predicted from sequence co-variation. Nat. Biotechnol. 35, 128–135 (2017).
Hsu, C., Nisonoff, H., Fannjiang, C. & Listgarten, J. Learning protein fitness models from evolutionary and assay-labeled data. Nat. Biotechnol. 40, 1114–1122 (2022).
Tang, Q. et al. Directed evolution of a halide methyltransferase enables biocatalytic synthesis of diverse sam analogs. Angew. Chem. Int. Ed. 60, 1524–1527 (2021).
Korfer, G. et al. Directed evolution of an acid Yersinia mollaretii phytase for broadened activity at neutral pH. Appl. Microbiol. Biotechnol. 102, 9607–9620 (2018).
Shivange, A. V. et al. Directed evolution of a highly active Yersinia mollaretii phytase. Appl Microbiol. Biotechnol. 95, 405–418 (2012).
Si, T. et al. Automated multiplex genome-scale engineering in yeast. Nat. Commun. 8, 15187 (2017).
Chao, R. et al. Fully automated one-step synthesis of single-transcript TALEN pairs using a biological foundry. ACS Synth. Biol. 6, 678–685 (2017).
Ayikpoe, R. S. et al. A scalable platform to discover antimicrobials of ribosomal origin. Nat. Commun. 13, 6135 (2022).
Yuan, Y., Huang, C., Singh, N., Xun, G. & Zhao, H. Self-resistance-gene-guided, high-throughput automated genome mining of bioactive natural products from Streptomyces. Cell Syst. 16, 101237 (2025).
Wittmann, B. J., Yue, Y. & Arnold, F. H. Informed training set design enables efficient machine learning-assisted directed protein evolution. Cell Syst. 12, 1026–1045.e7 (2021).
Meier, J. et al. Language models enable zero-shot prediction of the effects of mutations on protein function. In Advances in Neural Information Processing Systems 34 (NeurIPS 2021).
Schönherr, H. & Cernak, T. Profound Methyl Effects in Drug Discovery and a Call for New C−H Methylation Reactions. Angew. Chem. Int. Ed. 52, 12256–12267 (2013).
Tang, Q., Pavlidis, I. V., Badenhorst, C. P. S. & Bornscheuer, U. T. From natural methylation to versatile alkylations using halide methyltransferases. ChemBioChem 22, 2584–2590 (2021).
Yang, G.-Y., Zheng, G.-W., Zeng, B.-B., Xu, J.-H. & Chen, Q. Engineering of halide methyltransferases for synthesis of SAE and its application in biosynthesis of ethyl vanillin. Mol. Catal. 550, 113533 (2023).
Haefner, S. et al. Biotechnological production and applications of phytases. Appl. Microbiol. Biotechnol. 68, 588–597 (2005).
Schmidt-Dannert, C. & Arnold, F. H. Directed evolution of industrial enzymes. Trends Biotechnol. 17, 135–136 (1999).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Hu, R. et al. Protein engineering via Bayesian optimization-guided evolutionary algorithm and robotic experiments. Brief. Bioinform. 24, bbac570 (2023).
Santoni, M. L., Raponi, E., Leone, R. D. & Doerr, C. Comparison of high-dimensional bayesian optimization algorithms on BBOB. ACM Trans. Evol. Learn Optim. 4, 17:1–17:33 (2024).
Marquet, C., Schlensok, J., Abakarova, M., Rost, B. & Laine, E. Expert-guided protein language models enable accurate and blazingly fast fitness prediction. Bioinformatics 40, btae621 (2024).
Radivojević, T., Costello, Z., Workman, K. & Garcia Martin, H. A machine learning automated recommendation tool for synthetic biology. Nat. Commun. 11, 4879 (2020).
Reisenbauer, J. C., Sicinski, K. M. & Arnold, F. H. Catalyzing the future: recent advances in chemical synthesis using enzymes. Curr. Opin. Chem. Biol. 83, 102536 (2024).
Lawson, C. E. et al. Machine learning for metabolic engineering: a review. Metab. Eng. 63, 34–60 (2021).
Gricourt, G., Meyer, P., Duigou, T. & Faulon, J.-L. Artificial intelligence methods and models for retro-biosynthesis: a scoping review. ACS Synth. Biol. 13, 2276–2294 (2024).
Norton-Baker, B. et al. Enabling high-throughput enzyme discovery and engineering with a low-cost, robot-assisted pipeline. Sci. Rep. 14, 14449 (2024).
Diefenbach, X. W. et al. Enabling biocatalysis by high-throughput protein engineering using droplet microfluidics coupled to mass spectrometry. ACS Omega 3, 1498–1508 (2018).
Segal, M. An operating system for the biology lab. Nature 573, S112–S113 (2019).
Global Centers (GC). NSF—National Science Foundation. https://new.nsf.gov/funding/opportunities/gc-global-centers (2024).
Bell, E. L. et al. Biocatalysis. Nat. Rev. Methods Prim. 1, 1–21 (2021).
Hopf, T. A. et al. The EVcouplings python framework for coevolutionary sequence analysis. Bioinformatics 35, 1582–1584 (2019).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl Acad. Sci. 118, e2016239118 (2021).
Rabe, B. A. & Cepko, C. A simple enhancement for gibson isothermal assembly. bioRxiv https://doi.org/10.1101/2020.06.14.150979 (2020).
Acknowledgements
This work was supported by the Molecule Maker Lab Institute: An AI Research Institute program supported by U.S. National Science Foundation (NSF) under grant no. 2019897 (H.Z.), the NSF iBioFoundry (DBI- 2400058 to H.Z.), the NSF Global Center for Reliable and Scalable Biofoundries (OISE-2435374 to H.Z.), and U.S. Department of Energy (DOE) Center for Advanced Bioenergy and Bioproducts Innovation (award no. DE-SC0018420 to H.Z.). Any opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect those of the funding agencies. We thank members of the Biosystems Design theme at the Carl R. Woese Institute for Genomic Biology for their discussions and feedback. We thank Prof. Uwe Bornscheuer (University of Greifswald) for sharing research materials necessary for the AtHMT enzyme and Prof. Ulrich Schwaneberg and Dr. Anna Joelle Ruff for helpful discussions related to the YmPhytase enzyme. We also thank Nicole Setiawan for general laboratory duties that helped this project. All or parts of Fig. 1, Fig. 2 and Supplementary Fig. 2 has been created in Biorender (https://www.biorender.com/).
Author information
Authors and Affiliations
Contributions
T.Y. predicted enzyme mutations using the unsupervised and supervised models and designed the natural language agent. N.S. developed the site-directed mutagenesis protocol and designed the automated workflow while N.S. and S.L. performed automation implementation and operation. N.S. and S.L performed the four cycles of engineering of the AtHMT and YmPhytase enzymes. H.C. validated the phytase 4-MUP assay. A.R. helped with performing protein engineering experiments and SDM. N.S., S.L., and J.L. purified the AtHMT and YmPhytase protein variants and performed kinetic characterization. N.S., S.L., T.Y., and H.Z. wrote the manuscript. All authors contributed to editing the manuscript. H.Z. directed research activities and procured the funding to support this study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tong Si, Yongcan Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, N., Lane, S., Yu, T. et al. A generalized platform for artificial intelligence-powered autonomous enzyme engineering. Nat Commun 16, 5648 (2025). https://doi.org/10.1038/s41467-025-61209-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61209-y
This article is cited by
-
Green bioconversion of insoluble chitin: chitinase development pathways via multi-strategy synergy
Bioresources and Bioprocessing (2026)
-
Will self-driving ‘robot labs’ replace biologists? Paper sparks debate
Nature (2026)
-
Protein foundation models: a comprehensive survey
Science China Life Sciences (2026)