Abstract
Type V CRISPR-Cas12 systems are highly diverse in their functionality and molecular compositions, including miniature Cas12f1 and Cas12n genome editors that provide advantages for efficient in vivo therapeutic delivery due to their small size. In contrast to Cas12f1 nucleases that utilize a homodimer structure for DNA targeting and cleavage with a preference for T- or C-rich PAMs, Cas12n nucleases are likely monomeric proteins and uniquely recognize rare A-rich PAMs. However, the molecular mechanisms behind RNA-guided genome targeting and cleavage by Cas12n remain unclear. Here, we present the cryo-electron microscopy (cryo-EM) structure of Rothia dentocariosa Cas12n (RdCas12n) bound to a single guide RNA (sgRNA) and target DNA, illuminating the intricate molecular architecture of Cas12n and its sgRNA, as well as PAM recognition and nucleic-acid binding mechanisms. Through structural comparisons with other Cas12 nucleases and the ancestral precursor TnpB, we provide insights into the evolutionary significance of Cas12n in the progression from TnpB to various Cas12 nucleases. Additionally, we extensively modify the sgRNA and convert RdCas12n into an effective genome editor in human cells. Our findings enhance the understanding of the evolutionary mechanisms of type V CRISPR-Cas12 systems and offer a molecular foundation for engineering Cas12n genome editors.
Similar content being viewed by others
Introduction
Numerous archaea and bacteria harness the CRISPR-Cas system as an adaptive immunity mechanism against viral and nucleic acid invasions1,2. The rich diversity in protein sequences and genomic arrangements of CRISPR-Cas systems is distinctly categorized into two main classes (class 1 and class 2), further divided into seven major types (I-VII)3 and over 30 subtypes4. Class 1 systems are characterized by their assembly into multi-subunit effector complexes, whereas class 2 systems are defined by their reliance on single, multi-domain protein effectors, rendering them advantageous for a wide array of gene editing applications. Within class 2, types II, V, and VI utilize Cas9, Cas12, and Cas13 effectors, respectively, to target and cleave DNA or RNA under RNA guidance5,6,7,8. Unique among these, the Cas12 family nucleases, distinct from Cas9, employ a single RuvC domain for DNA interference and are postulated to have originated from transposon-associated TnpB proteins4,9,10. Despite the commonality of the RuvC domain, Cas12 effector proteins display minimal sequence similarity, highlighting their varied biochemical properties. Previously reported Cas12 proteins, including Cas12a–Cas12e and Cas12g–Cas12j2,8,11,12,13,14,15,16,17,18,19,20 are larger than the typical size of TnpB proteins (∼400 a.a.)9. Conversely, the smaller Cas12 variants (V-U1 to V-U5) show a closer evolutionary connection to TnpB proteins4,9,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36.
Detailed cryo-EM and biochemical analysis of these mini-Cas12 variants and the ancestor TnpB nuclease reveals distinct working mechanisms and functional roles37. Cas12m (V-U1) lacks DNA-cleaving activity and instead prevents invasive gene expression by binding to double-stranded DNA (dsDNA)31,32,33. Cas12f1 (V-U3) functions as a dimeric dsDNA cleavage nuclease, repurposed for flexible genome editing22,23,24,25,26,27,38,39,40,41,42,43. Cas12k (V-U5) acts as a catalytically inactive CRISPR effector that facilitates programmed, site-specific RNA-directed DNA transposition by associating a Tn7-like transposition system28,29,30. In comparison, TnpB is a monomeric protein that associates with an omega RNA, exhibiting capabilities for programmed genome editing9,10,35,36.
A recent study demonstrated that type V-U4 Cas12n nucleases possess RNA-guided dsDNA-cleavage activity34. Cas12n nucleases typically harbor a disordered C-terminal loop and a trans-activating (tracr) RNA region that overlaps with its open reading frame (ORF). These features are reminiscent of TnpB proteins, suggesting possible functional similarities. Cas12n associates with CRISPR RNA (crRNA) and trans-activating RNA (tracrRNA) to target dsDNA, specifically recognizing the rare A-rich PAMs34. Remarkably, a Cas12n variant from Actinomadura craniellae displays high genome editing activity and is engineered as a multifunctional genome editor. Yet, the molecular mechanisms underpinning Cas12n activity and its evolutionary ties with TnpB remain largely unexplored, largely due to a lack of structural data.
In this study, we determine the cryo-EM structure of a Cas12n nuclease from Rothia dentocariosa (RdCas12n) in complex with a sgRNA and a target dsDNA containing an 5′-AAAC-3′ PAM. The structure highlights Cas12n as a monomeric nuclease for sgRNA association and dsDNA targeting and elucidates the mechanism by which Cas12n uniquely recognizes the rare A-rich PAM. Additionally, a comparative structural analysis between Cas12n and the ancestral TnpB protein reveals both convergences and divergences, providing insight into the evolutionary development from TnpB to mature Cas12 enzymes. Leveraging structure-guided engineering, we also successfully modify the RdCas12n sgRNA, converting RdCas12n into an effective genome editor in human cells. Our results provide a basis for Cas12n engineering and enhance the understanding of the evolutionary mechanisms of CRISPR-Cas12 family nucleases.
Results
Cryo-EM structure of the RdCas12n–sgRNA–target DNA ternary complex
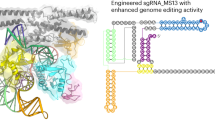
To investigate the working mechanism of Cas12n nucleases, we conducted a comprehensive phylogenetic analysis of 138 Cas12n nucleases. Based on sequence similarity, we constructed a phylogenetic tree of Cas12n (Fig. 1a, Supplementary Fig. 1a, and Supplementary Data 1) and classified Cas12n into multiple clades. Notably, the previously reported AcCas12n (Actinomadura craniellae) variant and three other orthologs—RdCas12n (Rothia dentocariosa), MlCas12n (Micrococcus luteus), and CgCas12n (Corynebacterium glutamicum)—are categorized into two distinct clades. While AcCas12n exhibits a preference for the 5′-AAG-3′ PAM sequence34, RdCas12n favor the 5′-AAC-3′ PAM sequence (Supplementary Fig. 1b), indicating the existence of different PAM recognition mechanisms across these clade-specific nucleases. It is worth noting that although 5’-AAG-3’ is the most preferred PAM for AcCas12n, AcCas12n can also cleave other 5’-AAV-3’ PAM-containing DNAs (where V = G, A, or C). Similarly, 5’-AAC-3’ is the most preferred PAM for RdCas12n, while it also retains the ability to cleave other 5’-AAH-3’ PAM-containing DNAs (where H = T, A, or C).
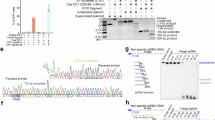
a Unrooted phylogenetic tree from representative Cas12n nucleases. AcCas12n-like and RdCas12n-like clades are shown in blue and pink, respectively. Arrows indicate Cas12n genes experimentally characterized in this study (RdCas12n, AcCas12n, CgCas12n and MlCas12n). b The tested Cas12n nucleases for purification. RdCas12n was used for final cryo-EM analysis. c Diagram of the sgRNA and target DNA used for cryo-EM analysis. d The domain structure of RdCas12n. CTD, C-terminal domain. Residues 1–10 and 268–539 (the NUC lobe) were not included in the final model. e Cryo-EM density map of the RdCas12n–sgRNA–target DNA ternary complex structure 1 (PDB ID: 9J09). f The overall structure of the RdCas12n–sgRNA–target DNA ternary complex structure 1 (PDB ID: 9J09). Disordered regions are indicated as dotted lines. The NUC region of the protein exhibits a disordered structure, and the delineation of the NUC lobe is tentative, marking a predicted domain where the NUC lobe is likely to be situated.
To isolate pure and functional Cas12n suitable for structural analysis, we evaluated four distinct Cas12n variants sourced from various bacterial species for their ability to be heterogeneously expressed in E. coli. However, when the Cas12n protein is expressed either without the accompanying sgRNA or by co-expressing both components using a single plasmid with separate T7 promoters – one for protein expression and one for RNA expression – the protein tends to aggregate into multimers and polymers. This complicates the effective isolation of individual proteins or RNPs, making it difficult to assemble or isolate active RNP monomers. To resolve this issue, we adjusted protein-RNA expression by changing plasmid copy numbers. Specifically, Cas proteins were expressed using a low-copy pCDFDuet-1-based plasmid (Novagen), while sgRNA was expressed using a high-copy pSGKP-based plasmid44. Under these conditions, we successfully purified and isolated RNP for RdCas12n. However, for several other characterized Cas12n orthologs, this purification method resulted in impurities or low yields, with either limited free RNP or low overall protein expression (Fig. 1b).
To elucidate the molecular architecture underpinning the functionality of RdCas12n, we employed cryo-EM to dissect the intricate complex formed by RdCas12n and a 214-nucleotide (nt) sgRNA, comprising the native 40-nt crRNA and a 170-nt tracrRNA sequences connected through a 5′-GAAA-3′ tetraloop (Fig. 1c). This assembly was further complexed with a 40-nt target DNA strand containing a 20-nt complementarity to the guide region of the crRNA (Fig. 1c) and a 14-nt non-target DNA strand containing the 5′-AAC-3′ PAM (Supplementary Fig. 2a, b and Supplementary Table 1). Our cryo-EM analysis, using two distinct box sizes, produced 3D reconstructions of the RdCas12n–sgRNA–dsDNA ternary complex, revealing two closely related structures: structure 1 (PDB ID: 9J09) at a resolution of 2.95 Å with elevated noise levels, and structure 2 (PDB ID: 9UDI) with cleaner density maps at 3.01 Å resolution. (Supplementary Fig. 2c and Supplementary Table 2). In both structures, a single RdCas12n molecule was observed to associate with one sgRNA and one dsDNA molecule, similar to the functional mechanism reported in ISDra2 TnpB35,36. While the core regions showed well-defined densities for RdCas12n residues and nucleobases at the key interaction interface (Supplementary Fig. 3a–j), the NUC domain (residues 268–539) exhibited absent densities, possibly due to its structural flexibility (Fig. 1d–f). In ternary complex structure 1, unresolved regions included sgRNA nucleotides −194 to −181, −155 to −153, −141 to −107, −90 to −77, −38 to −13, and 13 to 20, along with target strand (10–28) and non-target strand (1* and 2*) regions (Supplementary Fig. 4b). Ternary complex structure 2 displayed sgRNA gaps at nucleotides −194 to −179, −157 to −153, −143 to −107, −90 to −77, −38 to −13, and 10 to 20, with similar unresolved regions in the target and non-target strands (Supplementary Fig. 4c). Comparative analysis showed ternary complex structure 1 provides a clearer interaction interface and more complete sgRNA resolution (Supplementary Fig. 4d–g), leading us to select ternary complex structure 1 (PDB ID: 9J09) as the primary reference model. Additionally, to enhance our comprehension of the complete RdCas12n–sgRNA–dsDNA complex, we aligned the full-length RdCas12n model, as predicted by AlphaFold3, with our experimentally derived model45. This juxtaposition yielded a predicted composite structure that provides further insights into the intricate molecular interactions within the complex (Supplementary Fig. 4h).
RdCas12n domain structures
RdCas12n features a bilobed architecture made up of the recognition (REC) lobe and the nuclease (NUC) lobe, bridged by an interconnecting linker loop. The REC lobe contains the WED and REC domains, while the NUC lobe consists of the RuvC domain, the target nucleic acid-binding (TNB) domain, and a flexible C-terminal domain (residues 508–539). The DNA duplex harboring the PAM integrates within the groove formed by the REC and WED domains.
The WED domain (residues 1–20 and 182–267) is characterized by a seven-stranded β-barrel with two inserted α-helices, adopting an oligonucleotide/oligosaccharide-binding (OB) fold present in other Cas12 enzymes and the ancestor TnpB nucleases (Supplementary Fig. 5a, b). The REC domain (residues 21–181), fitted between the β1 and β2 strands of the WED domain, comprises six α-helices that are similar to counterparts in both TnpB and Cas12 family members (Supplementary Fig. 5b). Although the NUC lobe is not resolved in the density map, predictions based on AlphaFold suggest it includes the RuvC and TNB domains (Supplementary Fig. 5c). The RuvC domain displays an RNase H-like fold, with a five-stranded mixed β-sheet enveloped by four α-helices. Critical catalytic residues, D287, E386, and D484, form the enzyme’s active center, as is common across Cas12 variants (Supplementary Fig. 4d). In the AlphaFold model, the TNB domain, located next to the RuvC domain, contains a CCCC-type zinc finger motif. The structural comparison of Cas12n with TnpB and other mini Cas12 enzymes reveals that Cas12n features a compact zinc finger domain organization similar to that of TnpB (Supplementary Fig. 5f).
PAM recognition mechanism
Unlike most type V Cas12 family nucleases that recognize T- or C-rich PAMs12,13,15,22,24,25,27,46, RdCas12n uniquely identifies the rare 5′-AAC-3′ PAM sequence. This specific recognition is achieved primarily by the interaction with the complementary 3′-TTG-5′ bases (Fig. 2a, b and Supplementary Fig. 6). Within the RdCas12n complex, the nucleobases of target strand dT(−2) and dT(−3) form hydrogen bonds with R140, which is further anchored by E190 via hydrogen bonding (Fig. 2c). Moreover, the methyl group of dT(−3) engage in van der Waals interactions with P189, while dG(−1) engage in van der Waals interactions with P187 from the WED domain (Fig. 2d). Additionally, the G-C pairing of dC(−1*) and dG(−1) forms a shape-complementary interaction with V143 and Q147 (Fig. 2e). In addition to these specific interactions, RdCas12n also forms non-base-specific contacts within the complex; for instance, the side chain of R247 in the WED domain forms hydrogen bonds with target strand dG(−1) and dT(−2) (Fig. 2e). Similarly, the side chains of T95 and K102 in the REC domain create hydrogen bonds with the non-target strand nucleotides dA(−2*) and dA(−3*) (Fig. 2f).
a Recognition of the target DNA. The PAM duplex is bound to the cleft formed by the WED and REC domains. PAM upstream DNA exhibits additional interactions with the WED domain. b Detailed interactions of RdCas12n with the PAM duplex and PAM upstream DNA. c–f Major interactions between the PAM duplex and RdCas12n. Hydrogen bonding and electrostatic interactions are shown as blue dashed lines, and van der Waals interactions are shown as green dashed lines. The residues that interact with the nucleic acids through their main chains are shown in parentheses. g In vitro DNA cleavage activities of WT-RdCas12n and different PAM-recognition mutants. A linearized target DNA segment of 2.2-kb, comprising a 20-nucleotide target sequence and at a concentration of 10 nM, was subjected to incubation with the RNP complex at a concentration of 250 nM, at 37 °C across various time durations, as depicted by the gradient gray labels. Data are represented as mean ± SD (n = 3 biologically independent samples). Source data are provided as a Source Data file. h–i Major interactions between the PAM upstream DNA and RdCas12n. j Location of residues involved in PAM upstream DNA interactions. k Schematic of the molecular design for extra-WED region deletion mutants. l In vitro DNA cleavage activities of WT-RdCas12n, the PAM upstream DNA recognition mutants, and the extra-WED domain deletion mutants. Data are represented as mean ± SD (n = 3 biologically independent samples). Source data are provided as a Source Data file. m, n Guide:TS DNA recognition. Hydrogen bonding and electrostatic interactions are shown as blue dashed lines. The residues that interact with the nucleic acids through their main chains are shown in parentheses.
To further validate the roles of these residues in PAM recognition, we conducted experiments replacing them with alanine and assessed their impact on DNA cleavage activity relative to the wild-type protein. Mutations R140A, P189A, or E190A resulted in the complete abolition of the cleavage function of RdCas12n, while P187A or K102A mutations significantly diminished its DNA-cleavage capabilities, underlining the essential nature of these interactions for effective PAM recognition (Fig. 2g and Supplementary Fig. 7a, b). Collectively, our structural and functional analyses demonstrate that RdCas12n engages in both sequence-specific contacts with the target strand and non-sequence-specific interactions with the DNA backbone phosphates to achieve PAM recognition.
PAM-upstream DNA recognition
Distinct from other miniature Cas12 proteins and TnpB, RdCas12n is characterized by multiple non-sequence-specific interactions with the DNA backbone phosphates upstream of the PAM region (Fig. 2b). Here, the side chain of K191 and R220 engages in hydrogen bonding with the backbone phosphate of the non-target strand dA(−6*), while V200 creates van der Waals interactions. R201 forms a hydrogen bond with the O4’ backbone pentose of non-target strand dA(−6*) (Fig. 2h). Additionally, R211 forms a hydrogen bond with the backbone phosphate OP2 of target strand dG(−11), and R201 interacts with the nucleobases of dA(−8) and dT(−8*) (Fig. 2i). Interestingly, the multiple charged residues that interact with the PAM upstream region are clustered in the unusual sequence between β3 and β4 within the WED region. This sequence is unique to RdCas12n when compared to other Cas12n variants from different clusters such as AcCas12n or ISDra2 TnpB (Supplementary Fig. 8). In this study, we tentatively refer to this R- and K-rich expanded WED domain as extra-WED (Fig. 2j).
To experimentally validate the significance of these residues in DNA binding, we conducted alanine substitution experiments and compared the DNA cleavage efficiency of these variant proteins with the wild-type RdCas12n. Additionally, we designed three extra-WED deletion mutants (Fig. 2k). Whereas single alanine substitutions of K191, R201, or R211 did not significantly impact the DNA-cleavage activity, the R220A mutation notably diminished this activity. Concurrent alanine substitutions, either K191-and-R220 or R201-and-R211, completely abrogated or markedly reduced cleavage efficiency, respectively (Fig. 2l and Supplementary Fig. 9). Furthermore, the deletion of the extra-WED domain led to a complete loss of cleavage capacity in the modified proteins. Collectively, these findings indicate the importance of the augmented WED region for RdCas12n’s functionality.
RNA–DNA heteroduplex recognition
The guide RNA–target DNA heteroduplex is surrounded by the REC and WED domains (Fig. 2a), situated within a positively charged channel formed by the REC domain alongside the predicted RuvC domain (Supplementary Fig. 3b). This arrangement allows RdCas12n to engage with the heteroduplex through interactions with its sugar-phosphate backbone. In this context, R247 and S248 from the WED domain specifically recognize the phosphate group situated between dG(−1) and dG1 of the target strand (Fig. 2m). Within the guide RNA–target DNA heteroduplex, side chains from WED engage in hydrogen bond interactions involving the pentose backbone from G1 to U4 of the guide RNA (Fig. 2m). Side chains emanating from the REC domain primarily facilitate hydrogen bonding with the backbone from U4 to A7 of the guide RNA (Fig. 2n). However, the heteroduplex’s PAM-distal region is solvent-exposed, rendering the terminal eight base pairs (A13 to A20):(dT13 to dT20) disordered.
sgRNA architecture
The sgRNA (−194C to 20A) consists a 20 nt guide segment (1C to 20A) coupled with a 194 nt RNA scaffold (−194C to −1C), structured into five distinct stem regions (stems 1–5) and inclusive of a pseudoknot (PK) (Fig. 3a). The upper sections of stem 2 (−142G to −107A), stem 3 (−91C to −76G), stem 5 (−39U to −12A), and the 5′ region (−194C to −181A) exhibited disordered conformations in the resolved structure (Fig. 3b).
a Schematic of the sgRNA and target DNA. The upper stem regions of Stem 2 (−142G to −107A), Stem 3 (−91C to −76G), Stem 5 (−39U to −12A) and 5′ region (−194C to −181A) are disordered in the determined structure, suggesting the flexibility of these regions. These disordered regions are shown in dashed line frames. b Structure of the sgRNA scaffold. The disordered regions are denoted by dotted lines. c–j Detailed interactions and triple pairing inside the RNA scaffold. Hydrogen bonding and electrostatic interactions are shown as blue dashed lines.
The sgRNA scaffold structure presents several unanticipated features, particularly within the pseudoknot area (Fig. 3a–e). Here, −4U to −2C pairs with −168A to −170G, extending stem 1 into a seamless helix. Moreover, bases −148G to −152A engage in multicentric triple base pairings with stem 1 (−162G:−173C to −158A:−178U), contributing to the formation of a triple helix (Fig. 3a–e). Further detailed examination reveals a constellation of non-canonical pairings surrounding the pseudoknot region, which may contribute to stabilizing the RNA structure. These interactions extend from stem 1 to stem 2 (Fig. 3f), with −162G interacting with −99A and −163G with −98A, bringing stem 2 closer to stem 1. Hydrogen bonds also form between stem 3 (−69A, −98A) and the backbone phosphates of stem 1 (−171A, −172A), drawing stem 3 closer to stem 1 (Fig. 3f). Additionally, stem 3 (−67C) interacts with stem 4 (−52A) and stem 5 (−5G) (Fig. 3h), resulting in a more tightly packed overall RNA structure. The structure also unveiled unexpected interactions between stem 3 (−54U to −57U) and stem 4 (−95C, −94C, −72G, −71U) (Fig. 3i, j). This intricate web of interactions within the sgRNA likely plays a significant role in stabilizing the overall RNA scaffold structure.
sgRNA recognition
RdCas12n interacts with sgRNA majorly through interactions mediated by its WED and REC domains, which interface with the sugar-phosphate backbone of the sgRNA (Fig. 4a and Supplementary Fig. 6). Specifically, the side chain of R18 within the WED domain establishes hydrogen bonds with the phosphate backbone of −171A in stem 1 (Fig. 4b). Further interactions were observed in stem 3, where both the REC and WED structural domains formed numerous bonds with the sugar-phosphate backbone on the same side of the stem. Notably, amino acids N22 and Q23 from the REC domain engage with the phosphates of nucleotides −74G to −72G through hydrogen bonding (Fig. 4c), while the side chains of R210, H218, R228, and R217 from the WED domain form similar bonds with the phosphate backbone of nucleotides −72G to −69A (Fig. 4d). In stem 4, residue K238 anchors Y216 to facilitate interactions with the sugar-phosphate backbone at position –51U (Fig. 4e).
a Recognition of the sgRNA scaffold by RdCas12n. RdCas12n is shown as a surface model. The sgRNA scaffold is recognized mainly through the WED and RuvC domains. b Recognition between the stem 1 and WED. c Recognition between the stem 3 and REC. d Recognition between the stem 3 and WED. e Recognition between the stem 4 and WED. Hydrogen bonding and electrostatic interactions are shown as blue dashed lines and van der Waals interactions are shown as green dashed lines. f Electrophoretic mobility shift assay (EMSA) analysis of binding interactions among RdCas12n or its truncation mutants, sgRNA, and dsDNA. A schematic representation of the RdCas12n variants analyzed in this assay is depicted above the corresponding data panels. Compared to full-length RdCas12n, the NUC lobe-truncated variant displayed reduced binding affinity for dsDNA targets. Experiments were conducted in three independent biological replicates, with consistent results observed across trials. g Scheme of the design for the transcriptional activation assay in E. coli Plasmids carrying various CRISPRa effectors and target-based reporters were co-transformed into E. coli S2060 cells. Successful on-target DNA binding triggers transcription of the downstream luciferase reporter. h Transcriptional activation activities of full length RdCas12n and N-RdCas12n at 9 different target sites. Data are represented as mean ± SD (n = 3 biologically independent samples). Source data are provided as a Source Data file.
Due to the flexibility of the NUC lobe, the interactions between the sgRNA and the NUC lobe could not be observed in the present structure. This observation suggests that the NUC lobe may not be essential for the assembly of the RNP complex and its subsequent DNA targeting ability. To evaluate the influence of the NUC lobe on the DNA targeting efficiency of RdCas12n, comparative analyses were performed with both the wild-type RdCas12n and a variant devoid of the NUC lobe (N-RdCas12n) using Electrophoretic Mobility Shift Assays (EMSA) (Fig. 4f). The results revealed that the NUC lobe’s absence diminishes the RdCas12n RNP complex’s affinity for dsDNA. However, this reduced affinity can be compensated by elevating the protein concentration, enabling the NUC lobe-lacking variant of the RdCas12n RNP complex to achieve dsDNA binding effectiveness on par with that of the wild-type complex. Such findings suggest the NUC lobe, while not strictly necessary for RdCas12n RNP to associate with dsDNA, significantly contributes to enhancing the interactions between the RNP complex and dsDNA.
The ultra-compact size (270 amino acids) and preserved DNA targeting capability of N-RdCas12n highlight its potential utility in developing advanced genome-modification technologies, such as CRISPR activation/inhibition (CRISPRa/i), base editing, and epigenetic modification systems. These applications are particularly suited for contexts where delivery vector size constraints are a critical factor. To evaluate the transcriptional activation potential of N-RdCas12n, it was fused with the omega subunit of bacterial RNA polymerase III, employing a luxAB reporter system for assessment, and its performance was benchmarked against that of the full-length RdCas12n (Fig. 4g). Through optimization experiments involving variations in the omega subunit linker length and adjustments to the site’s proximity to the minimal lac promoter (Supplementary Fig. 10), we identified a configuration characterized by a 32 amino acid linker and variant 3 (v3) spacing. The performance of these configurations was evaluated across nine different sites, each harboring an 5′-AAC-3′ PAM. Our findings reveal that both constructs prompted a marked increase in transcriptional activity, as evidenced by enhanced luminescence per OD600 in Escherichia coli cultures (Fig. 4h). Interestingly, though the full-length RdCas12n construct generally showed superior activation capabilities across various sites, the N-RdCas12n construct demonstrated higher efficiency at specific targets (target 1 and target 8). This emphasizes the potential for fine-tuning and augmenting the N-RdCas12n’s efficacy and utility as an ultra-compact tool for genome targeting.
Structural comparison among RdCas12n, TnpB, and other miniature Cas12 effectors
Structural comparisons between RdCas12n, TnpB, and other miniature Cas12 effectors highlight both similarities and distinct variations. While Cas12n and TnpB share a bilobed architecture encompassing REC and NUC lobes, RdCas12n demonstrates a unique REC extension and an additional WED domain expansion inserted between strands β3 and β4 of the WED domains (Supplementary Fig. 11). The RuvC domain remains elusive in the RdCas12n structure, in contrast to other miniature Cas12 effectors where it is typically present. This is consistent with observations previously reported for the TnpB structure (PDB ID: 8EX9), where the RuvC structural domain is also disordered. Interestingly, the same study also reported another structure (PDB ID: 8EXA) with a resolved RuvC domain.
Further examination with another miniature Cas12 protein, Cas12m2, exhibits a pronounced REC expansion that interacts with the non-target strand DNA and the PAM-distal region of the guide RNA–target heteroduplex. Additionally, an extra WED insertion between β3–β4 and β4–β5 leads to an enlarged WED, facilitating more direct RNA interactions. Conversely, Cas12f1 variants exhibit differences in REC expansions, influencing dimerization strategies: CnCas12f1 features complex dimerization interfaces across the REC region, whereas AsCas12f1 exhibits a minor REC expansion facilitating head-to-tail dimerization, enhancing stabilizing interactions within the PAM-distal heteroduplex (Supplementary Fig. 11).
Regarding sgRNA architecture, TnpB, RdCas12n, CnCas12f1, and AsCas12f1 exhibit a conserved L-shaped core scaffold, primarily consisting of stems 1 and 2, along with the pseudoknot region. Notably, RdCas12n showcases an additional stem 4–stem 5 continuous RNA helix adjacent to the pseudoknot, akin to configurations observed in AsCas12f1/CnCas12f1. Conversely, TnpB straightforwardly bridges the pseudoknot to stem 3. Unique to RdCas12n, the bases connecting stems 3 and 4 engage with stem 4 and the WED domain, forming a stem 3–stem 4 triple base pairing through non-specific interactions with the sugar-phosphate backbone facilitated by the REC and WED domains (Supplementary Fig. 11).
Systematic sgRNA engineering improves RdCas12n editing activity
Truncating the disordered regions of sgRNA has been shown to enhance the genome editing activity of Cas12f124,25,27,43, leading us to explore this strategy for RdCas12n, which previously demonstrated minimal genome editing activity in human cells34. To improve RdCas12n’s editing efficiency, we first applied a bacterial genome targeting assay to dissect and optimize the essential structural components of its sgRNA. Utilizing a systematic truncation method focused on specific structural stems, we identified sgRNA_T6, which exhibited modifications in the 5′ region and stem 5, and selected it for detailed analysis (Fig. 5a, b). Our results showed that deletions up to sgRNA_T9 in stem 3 and sgRNA_T13 in stem 2 retained the original genome targeting efficiency. However, further deletions (sgRNA_T8 & sgRNA_T12) or the elimination of additional structural elements (sgRNA_T7) significantly reduced genome targeting activity of RdCas12n (Fig. 5a, b). Subsequently, we engineered sgRNA_T13, a 131-nt RNA integrating elements from sgRNA_T6 and stem 2 deletions, while preserving efficient bacterial genome targeting capability.
a Schematic of different sgRNA truncations for bacterial genome targeting. b Bacterial genome targeting assay using different truncated sgRNAs. +, with spacer; −, without spacer. Effective truncations are highlighted in a red box. Uncropped images are provided in the Source Data file. c Comparison of the genome editing activities of the modified sgRNAs at three different genomic loci of HEK293T cells. Data are represented as mean ± SD (n = 4 biologically independent samples). NT stands for non-targeting spacer. d Schematic of the best sgRNA version (T19) performed in HEK293T cells. e Indel efficiency comparison of the engineered version (T19) with the original version across the 19 different genomic sites (Locus_1 to 19, as specified in Supplementary Data 2) in the HEK293T cells with the 5′-VAAC-3′ PAM (where V = A, C, or G). Data are represented as mean ± SD (n = 4 biologically independent samples). NT stands for non-targeting spacer. f Impact of sgRNA-target DNA mismatch on indel activity in the HEK293T cells. Data are represented as mean ± SD (n = 4 biologically independent samples). Source data are provided as a Source Data file.
Our analysis from sgRNAs T22–T25 suggested that excessive truncation of multiple stems might interfere with the structural integrity of the stem-loop, critically impacting RNA conformation (Fig. 5a, b). To address this issue, we introduced corrective mutations in mismatch pairings of stem 2 (sgRNA_T27) and stem 3 (sgRNA_T26). These strategic mutations allowed us to optimize the design further, trimming down stem 2 and introducing a deletion in stem 3. This led to the development of sgRNA_T30, a 125-nt scaffold variant maintaining effective bacterial genome targeting (Fig. 5a, b). It’s crucial to note that overly extensive truncations could attenuate Cas12n activity, emphasizing the importance of thorough evaluation of all sgRNA versions in human cells.
To provide a precise evaluation of sgRNA optimization, 14 bacterial-optimized variants (sgRNA_WT, T6, T13–T21, and T28–T30; Supplementary Fig. 12) were tested at three target sites (VEGFA-1, HEXA-1, HEXA-2) in HEK293T cells (specified as Site_1–3; Supplementary Data 2). Remarkably, sgRNA_T19, which incorporates a stem 2 truncation from sgRNA_T6, demonstrated the highest activity across all tested human cell sites (Fig. 5c). Nonetheless, while combinational optimization of stem 2 and stem 3 (as exemplified by sgRNA_T30) was previously validated in bacterial systems, its showed reduced effectiveness in mammalian cells. Furthermore, additional efforts to optimize sgRNA through truncation and bubble complementation based on sgRNA_T19 (sgRNA_T27 to sgRNA_T37; Supplementary Fig 12) resulted in decreased editing activity.
We thereby selected sgRNA_T19 as our final optimized sgRNA for further characterizations (Fig. 5d). We evaluated its indel efficiencies against sgRNA_WT across 20 genomic sites (Locus_1–20 in Supplementary Data 2), each featuring a 5′-VAAC-3′ PAM (where V can be A, C, or G) within eight distinct genes: AAVS1, PDCD, TP53, EMX1, HEXA, KLHL, IFNG, and HBB. sgRNA_T19 demonstrated a substantial enhancement in genome-editing efficiency across the majority of sites tested (Fig. 5e). Notably, sgRNA_T19 exhibited around 40% editing efficiency at a specific site, HEXA-4 (specified as Locus_20; Supplementary Data 2), within the HEXA gene (Supplementary Fig. 13a–d). For mismatch tolerance assays, HEXA-1, a site with moderate editing efficiency, was selected to assess indel activity. The results showed that indel activity diminished in HEK293T cells as mismatches were intentionally introduced near the PAM-proximal region, showing reduced tolerance to mutations in this area (Fig. 5f).
In addition, we compared the cell gene editing efficiency of RdCas12n to that of the similar smaller Cas12 family protein Un1Cas12f1, as well as to SpCas9 and AsCas12a. Target sites were selected based on their compatibility with the PAM requirements adjacent to the target sequences. The results indicated that RdCas12n-T19 exhibits lower activity compared to the larger Cas proteins, such as SpCas9 and AsCas12a, which demonstrated an overall gene editing efficiency of over 50% at the tested loci (Supplementary Fig. 14a, b). Notably, RdCas12n achieved an approximate editing efficiency of 10% at three selected loci, surpassing Un1Cas12f1 at two of these sites, yet falling short at one (Supplementary Fig. 14c). Thus, in scenarios where the use of a compact Cas system is paramount for cellular gene editing, it is imperative to weigh both the constraints posed by PAM sequences and the actual performance of the editing system. While RdCas12n displays potential under specific conditions, it necessitates further refinement and enhancement to unlock its full gene-editing capability.
Discussion
In this study, we determined the cryo-EM structure of the RdCas12n–sgRNA–dsDNA ternary complex, revealing the detailed molecular mechanisms underlying A-rich PAM recognition and the interactions between RdCas12n and the nucleic acids. Structural comparisons across RdCas12n, TnpB, and other small Cas12 effectors emphasize the evolutionary transition of Cas12n nucleases from the ancestral TnpB to the fully functional Cas12 effectors. Utilizing structure-guided sgRNA engineering, we successfully enhanced the initially modest editing capabilities of RdCas12n, transforming it into a potent genome-editing tool in human cells.
Phylogenetic analysis reveals that RdCas12n and AcCas12n belong to two distinct clades (Supplementary Fig. 1a). Sequence alignments and structural analyses demonstrate that proteins similar to RdCas12n engage in PAM recognition by interacting with dT(−2) and dT(−3) of the target strand through an arginine (R) residue within the REC domain (Fig. 2c), while AcCas12n-like proteins feature an enrichment of tyrosine (Y) residues in the corresponding region (Supplementary Fig. 8). Additionally, proteins akin to RdCas12n apply PAPEK(V) residues for recognizing PAM-complementary bases (Fig. 2d), whereas AcCas12n-like proteins adapt HHDV(K)-like residues (Supplementary Fig. 1a and 8). These observations highlight the presence of two distinct mechanisms for recognizing 5′-AAN-3′ PAMs.
The ancestral TnpB nuclease, a monomeric protein with the capacity for dsDNA cleavage, stands in contrast to the distinct features exhibited by currently characterized miniature Cas12 nucleases35,36. For instance, Cas12m and Cas12k have lost their dsDNA cleavage capability, whereas Cas12f1 nucleases operate as dimeric proteins for DNA cleavage30,32. These differences hint at earlier evolutionary intermediates in the Cas12 family lineage. The Cas12n nuclease serves as such an intermediate, displaying monomeric nuclease characteristics similar to TnpB, alongside the retention of dsDNA cleavage capabilities. Through detailed structural comparisons among Cas12n, TnpB, and other Cas12 family members, Cas12n emerges as a crucial evolutionary bridge from the ancestral TnpB to the fully developed Cas12 nucleases (Fig. 6). Our analysis supports the conclusion that Cas12n is the nuclease within the Cas12 family that bears a close resemblance to the ancestral TnpB, notwithstanding the similarity of its sgRNA to exhibit stem-loop structures comparable to those observed in Cas12f1.
Structural comparison of RdCas12n (PDB ID: 9J09) with TnpB (PDB ID: 8H1J), Un1Cas12f1 (PDB ID: 7C7L), AsCas12f1 (PDB ID: 8J12), Cas12m (PDB ID: 8HHL), Cas12e (PDB ID: 6NY2), and Cas12a (PDB ID: 6I1K). The second Cas12f1 molecule (Mol. 2), as well as the specific REC expansions in Cas12n and Cas12m and REC2 insertions in Cas12e and Cas12a, are highlighted in blue. The region corresponding to the REC expansion in Cas12m that is already present in Cas12n, as well as the REC expansion in Cas12a that is already present in Cas12m, are highlighted in gray. REC1 expansion in Cas12f1, Cas12e and Cas12a are highlighted in green. The WED domain expansions from AsCas12f1 to Cas12a are highlighted in orange, the PI domain in Cas12a is highlighted in red. Other major differences are indicated by dashed line frames.
A comparative structural analysis of TnpB35, miniature Cas12 effectors (including Cas12n, Cas12f122,26,27, and Cas12m232), and larger Cas12 proteins (like Cas12e15 and Cas12a46) elucidates the evolutionary trajectory of Cas12 enzymes. Throughout their evolution, Cas12 family proteins consistently exhibit structural expansions, particularly notable in the REC domain, manifesting through two primary mechanisms: direct expansion and expansion via dimerization (Fig. 6). The miniature enzymes—TnpB, Cas12n, and Cas12m—show a consistent pattern of direct REC domain expansions extending toward the PAM-distal end (Fig. 6 and Supplementary Fig. 15a, b). Specific amino acids, particularly R and K, within these expanded regions, appear to encircle the NTS DNA, enhancing NTS DNA binding and stabilizing R-loop formation. Larger Cas12 enzymes harbor an additional REC domain (REC2) dedicated to RNA–DNA heteroduplex binding (Fig. 6 and Supplementary Fig. 15c, d). Unique among the Cas12 family, the Cas12f1 enzymes engage in REC domain expansion by forming an asymmetric homodimer that interacts with RNA–DNA heteroduplex, as highlighted in Mol. 2 (Fig. 6). Additionally, direct REC domain expansions are also observed in Cas12f1 enzymes. Major structural expansions also occur in other domains, such as the WED domain, which frequently exhibits R- and K-rich expansions between beta strands, extending toward the sgRNA or the PAM duplex (Fig. 6 and Supplementary Fig. 15c, d). These expansions likely enhance interactions between the effector proteins and nucleic acids or serve as structural surrogates for RNA segments, potentially altering their genome editing capabilities.
Our research has shown that N-RdCas12n preserves its RNA-guided DNA targeting function in E. coli, highlighting its potential as a programmable molecule for developing advanced genome manipulation technologies. Notably, its compact structure, comprising merely 270 amino acids, facilitates the integration with additional effectors. Even after coupling with larger effectors such as TETv1, the cumulative construct size remains under 4.7 kb, rendering it compatible with AAV vector encapsulation (Supplementary Fig. 16a, b). Furthermore, the discovery of RdCas12n’s RuvC-independent target recognition mechanism suggests that other miniature Cas12 nucleases, which share sgRNA structural motifs similar to that of RdCas12n, are likely to retain their targeting abilities through truncations akin to those observed in RdCas12n. In all, this structural, evolutionary, and engineering analysis lays a foundation for further development and optimization of the CRISPR-Cas12n system and potential derivative genome editing technologies.
Methods
Plasmid construction
For the heterologous co-expression and purification of RdCas12n, an E. coli-codon-optimized RdCas12n gene or its variants were cloned into a pCDFDuet-based plasmid. This construct included the fusion of an N-terminal His6-SUMO tag and a human rhinovirus 3C (HRV3C) protease cleavage site. Additionally, the wild-type sgRNA was cloned into a pSGKP-based plasmid using a T7 promoter for transcription. For in vitro dsDNA cleavage assays, substrate plasmids with various PAMs were generated using circular polymerase extension cloning, with a pUC19-based plasmid serving as the template. This method utilizes a single polymerase to seamlessly assemble and clone multiple inserts with any vector in a one-step in vitro reaction47. Template plasmids were subsequently removed using DpnI (ABclonal). In bacterial genome targeting assays, a p15a-based vector was used to constitutively express RdCas12n, under the control of an rpsL promoter. The corresponding sgRNAs or variants were expressed by a pSGKP-based plasmid with a J23119 promoter to facilitate transcription. For CRISPRa assays, a p15a-based vector was used to express the ω-fused RdCas12n or its variants under the control of an rpsL promoter. The cognate sgRNAs were separately expressed by a pET175-based plasmid with a J23119 promoter that contains a reversed PAM-Target-plac-LuxA/B reporter. Various PAM-Target combinations and gap lengths between the Target-plac of the pET175 reporter system were constructed using Golden Gate assembly48. For gene disruption assays in human cells, an all-in-one plasmid was created to contain a cytomegalovirus (CMV)-driven nuclear localization signal (NLS)-tagged human-codon-optimized RdCas12n-expressing cassette and a U6-driven sgRNA transcription cassette. All plasmids were extracted using the MolPure® Plasmid Mini Kit (Yeasen). A list of all plasmids utilized in this study can be found in Supplementary Data 3.
Expression and purification of RdCas12n
For the purification of RdCas12n wild-type, RdCas12n alanine replacement mutant and extra-WED deletion mutant proteins, the RdCas12n-expressing plasmid and sgRNA-expressing plasmid were transformed into E. coli BL21(AI) for heterologous protein expression. The E. coli cells were cultured in fresh LB supplemented with 50 μg·mL−1 spectinomycin and 50 μg·mL−1 kanamycin at 37 °C until the OD600 reached to 0.6–0.7. IPTG was subsequently added to a final concentration of 0.25 mM, along with 0.16% L-arabinose, to induce protein expression at 16 °C for 16–20h. Harvested cells should be used immediately or stored on ice for no longer than 12h. It is important to note that any RNP-containing products should not be frozen in liquid nitrogen or stored at −80 °C for later use.
For every 1 L of E. coli culture, cells were pelleted by centrifugation, resuspended in buffer A (10 mM MgCl2, 20 mM Tris-HCl, pH 7.5, 500 mM NaCl, 10% glycerol v/v, 1 mM DTT), and then disrupted using a high-pressure cell disruptor at 800 bar for 15min at 4 °C. The clarified supernatant was loaded onto a buffer A pre-equilibrated 5-mL HisTrap HP column (Cytiva). 30 mL volume of buffer A containing 50 mM imidazole was used to wash unbound proteins, followed by elution of the RdCas12n ribonucleoprotein (RNP) complex using buffer B (10 mM MgCl2, 20 mM Tris-HCl, pH 7.5, 500 mM NaCl, 250 mM imidazole, 10% v/v glycerol and 1 mM DTT). The eluent was concentrated to 1 mL and further purified by using a HiLoad 16/600 Superdex 200 pg column (GE healthcare/Cytiva) with buffer A. RdCas12n RNP peak elution around 77 mL was collected and diluted into a final buffer C (10 mM MgCl2, 20 mM Tris-HCl, pH 7.5, 300 mM NaCl and 1 mM DTT) before being loaded onto a buffer C pre-equilibrated 5-mL HiTrap Capto Q column (Cytiva). 30 mL of buffer C was used to wash unbound proteins, and then buffer D (10 mM MgCl2, 20 mM Tris-HCl, pH 7.5, 1 M NaCl and 1 mM DTT) was used for gradient elution to elute RdCas12n RNP (prefer 1 mL·min−1 flow rate, 30 mL to 90% Buffer D). The purified RdCas12n RNP protein was exchanged into buffer E (10 mM MgCl2, 20 mM Tris-HCl, pH 7.5, 500 nM NaCl and 1 mM DTT), concentrated to ~0.3 mg·mL−1, and then used for structure determination and in vitro DNA cleavage assay.
For the purification of N-RdCas12n, the RdCas12n-expressing plasmid were transformed alone into E. coli BL21(AI) for heterologous protein expression. The E. coli cells were cultured in fresh LB supplemented with 50 μg·mL−1 spectinomycin at 37 °C until the OD600 reached to 0.6–0.7. IPTG was subsequently added to a final concentration of 0.25 mM, along with 0.16% L-arabinose, to induce protein expression at 16 °C for 16–20h. Harvested cells were purified through 5-mL HisTrap HP column and HiLoad 16/600 Superdex 200 pg column as mentioned above. N-RdCas12n peak elution around 85 mL was collected for EMSA assay.
In vitro DNA cleavage assay
For the in vitro cleavage assays, complexes of RdCas12n and sgRNA were purified employing the identical protocol established for the preparation of samples in prior cryo-EM analyses, ensuring purification within 24 h prior to the DNA cleavage activity. Quantification of protein concentrations was performed using a Bradford Protein Assay Kit (TransGen Biotech, China) to ensure accuracy in the experimental setup. In the in vitro DNA cleavage assay, the standard reaction mixture comprises 10 nM of a PCR-linearized substrate plasmid (2.0 kb), along with 250 nM of RdCas12n in a 1× reaction buffer condition containing 10 mM MgCl2, 20 mM Tris-HCl, pH 7.5, 100 mM NaCl and 1 mM DTT. The reaction is prepared on ice, incubated at 37 °C and halted at various time points by adding stop buffer (50 mM EDTA and 50 μg·mL−1 proteinase K (Solarbio)) at 50 °C for 10 min. The resulting products are analyzed using a 1% agarose gel in 1× TAE running buffer at 180 V for 20 min. The gel is stained with 4S Red Plus dye (Sangon Biotech) and visualized with the ChemiDoc MP System (Bio-Rad), unless otherwise specified. In vitro cleavage experiments were performed at least three times.
For 16-PAM in vitro DNA cleavage assay, the DNA substrate (500 bp) was PCR-amplified with 5′-end FAM-labeled primers and purified by using TIANquick Midi Purification Kit (Tiangen Biotech, Beijing, China). 30 nM DNA substrate (500 bp) was incubated with 400 nM RdCas12n RNP in the 1× reaction buffer at 37 °C for 1 h and stopped by adding stop buffer at 50 °C for 10 min. The sample was mixed with 6×gel loading dye (NEB) and analyzed through 6% TBE-PAGE gel electrophoresis, and visualized using the ChemiDoc MP System (Bio-Rad). In vitro cleavage experiments were performed at least three times.
Double-stranded DNA preparation
For EMSA and RdCas12n–sgRNA–dsDNA complex preparation assay, various 5′-end FAM (6-carboxyfluorescein)-labeled or unlabeled ssDNA molecules were synthesized from Sangon and were annealed with their complementary unlabeled ssDNA with a molar ratio of 1:1 to obtain dsDNA.
Target DNA strand (40-nt; unlabeled):
5′-GGCAGTGTTTTCACTTTCACCTGAACCGGTTTCTCACAGC-3′
Non-target DNA strand (14-nt; 5′-FAM fluorescein labeled or unlabeled):
5′-GCTGTGAGAAACCG-3′
sgRNA preparation
Single guide RNAs (sgRNAs) were generated by in vitro transcription (IVT) using the HiScribe T7 High Yield RNA Synthesis Kit (NEB), following the manufacturer’s instructions. After transcription, sgRNAs were treated with DNase I to remove residual DNA templates, then purified by phenol-chloroform extraction and isopropanol precipitation. Purified sgRNAs were stored at –80 °C until use. The transcription templates utilized in this study are detailed in Supplementary Table 1.
RdCas12n–sgRNA–dsDNA complex preparation
2.5 nmol of purified RdCas12n RNP was mixed with 5 nmol pre-annealed target DNA (40-nt TS and 14-nt NTS) in 600 μL of buffer E at 37 °C for 30 min. 500 uL of the RdCas12n–sgRNA–dsDNA complex was purified by size-exclusion chromatography with a Superose 6 10/300 GL column (Cytiva) pre-equilibrated with buffer E. The purified complex was concentrated to a concentration of ∼0.277 mg·mL−1 of RdCas12n protein. The protein concentration was determined using SDS-PAGE, where the concentrated protein complex was diluted and loaded onto the gel for relative quantification. This process included utilizing various concentrations of BSA standard protein as references to create a calibration curve. 3 uL complex, at a concentration of 0.277 mg·mL−1, was applied onto the H2/O2 glow-discharged holey carbon grids (CryoMatrix Amorphous alloy film R1.2/1.3, 300 mesh) in a Vitrobot Mark IV (Thermo Fisher Scientific) with a waiting time of 4 s and a blotting time of 2 s under 100% humidity conditions. The grids were plunge frozen in liquid ethane cooled in liquid nitrogen.
Electron microscopy data collection
The cryo-EM data were collected on a Krios G4 cryo-TEM (Thermo Fisher Scientific) equipped with a cold-FEG operating at 300 kV acceleration voltage, a Selectris X energy filter, and a EF-Falcon 4 direct electron detector. Data were acquired using EPU at 0.96 Å pixel size (at a nominal magnification of 130,000×), 60 electrons·Å−2 total exposure at a 9–10 electron·pixel−1·sec−1 in EER format, with a defocus range of −1.2 μm to −1.8 μm. A total of 5396 images were gathered.
Image processing
All movies were motion corrected and dose weighted by MotionCor2 software49, and their contrast transfer functions (CTF) were estimated by cryoSPARC patch CTF estimation50. For patch CTF estimation, manually curated exposures, blob picking, particle extraction and futher refinement of the RdCas12n–sgRNA–dsDNA complex, specifically ternary complex structure 1 (PDB ID: 9J09), were carried out using cryoSPARC (v3.2.0). Further data processing for ternary complex structure 2 of the RdCas12n–sgRNA–dsDNA complex (PDB ID: 9UDI) was performed utilizing cryoSPARC (v4.6.2). Out of the 5396 collected images, 5296 were accepted based on stringent criteria, including an average defocus value of less than 50,000, a CTF under 5.00, a defocus range within 3000, relative ice thickness below 1.6, and astigmatism lower than 400. And a total of 4,617,590 raw particles were examined through blob picking, focusing on particles with a diameter range of 90–140 Å.
For the RdCas12n–sgRNA–dsDNA ternary complex structure 1 (PDB ID: 9J09), 4,617,590 particles were extracted (box size 384 pixels) and subjected to multiple rounds of reference-free two-dimensional (2D) classification with a circular mask of 130 Å in diameter to generate particle sets. From this process, 2,175,481 particles, each sized 384 pixels by 384 pixels, were selected for further analysis. Further curation was performed through cryoSPARC’s heterogeneous refinement (n = 4), employing the map obtained from the cryoSPARC ab-initio reconstruction as a template. Subsequently, 879,820 particle projections of the best class (Class_0) were further applied for homogenous refinement and non-uniform refinement, generating the best density map with a global resolution of 2.95 Å estimated by cryoSPARC’s validation (FSC) job.
For the RdCas12n–sgRNA–dsDNA ternary complex structure 2 (PDB ID: 9UDI), 4,617,590 particles were extracted (box size 384 pixels) and subjected to multiple rounds of reference-free (2D) classification with a circular mask of 130 Å in diameter to generate particle sets. From these, 2,047,648 selected particles, each sized 256 pixels by 256 pixels, were extracted for further processing. Further curation was performed through cryoSPARC heterogeneous refinement (n = 4), employing the map obtained from the cryoSPARC ab-initio reconstruction as a template. Then 786,732 particle projections of the best class (Class_1) were further applied for homogenous refinement and non-uniform refinement, generating the best density map with a global resolution of 3.01 Å estimated by cryoSPARC’ s validation (FSC) job.
Model building and validation
The structure model of RdCas12n–sgRNA–dsDNA complex was manually constructed in Coot (v.0.9.4.1)51 and iteratively refined against the sharpened cryo-EM density map using the phenix.real_space_refine tool52 within the Phenix (v.1.20.1-4487) software suite. For visualization, molecular figures were rendered in Cuemol2 (v.2.2.3.443), and the local-resolution analysis of the cryo-EM density map was performed using ChimeraX (v.1.6.1).
Electrophoretic mobility shift assay (EMSA)
Duplexed DNA was prepared as above, with the 5′-FAM label at the non-target strand (40-nt TS and 14-nt 5′-FAM-NTS). For dRdCas12n, reactions were performed by incubating 0 to 25 nM dCas12n RNP to 10 nM FAM-label dsDNA for 30 min at 37 °C in a 20-μL binding buffer system containing 10 mM MgCl2, 20 mM HEPES pH 7.5, 100 mM NaCl and 1 mM DTT. For N-RdCas12n, the N-RdCas12n protein and IVT RNA were pre-mixed at a molar ratio of 1:1 ranging from 0 to 400 nM, or a combination of 0 to 400 nM N-RdCas12n with 25 nM RNA, in a 19.5-μL binding buffer system. The mixture was then pre-incubated at 37 °C for 15 min. Subsequently, a final 10 nM concentration of FAM-labeled dsDNA was introduced into the system, resulting in a 20-μL binding buffer setup, and incubated at 37 °C for 30 min. Samples were then mixed with 6×gel loading (10 mM MgCl2, 20 mM HEPES pH 7.5, 0.1% bromophenol blue, glycerol 40%(v/v) and 1 mM DTT) and separated using native polyacrylamide gels (6%, 20 mM HEPES) and visualized by fluorescence imaging.
Bacterial luciferase assays
To assess the transcriptional activation activity in E. coli, plasmids of interest (including the p15a-based RdCas12n-ω plasmid and the pET175-based sgRNA-target-reporter plasmid) were introduced into E. coli S2060 cells53 and cultured overnight at 37 °C on LB agar plates supplemented with 50 μg·mL−1 carbenicillin and 75 μg·mL−1 apramycin. For each biological replicate, single colonies were selected and cultured overnight at 37 °C. These cultures were then diluted 1:50 into 1 ml of fresh medium. The cells were grown until they reached to an OD600 of 0.3–0.4, at which point they were transferred to assay plates (150 µl per well, 96 well, Black Plate, Clear Bottom with Lid (Corning® REF 3603)). The SpectraMax i3 was utilized to monitor OD600 and luminescence (with an integration time of 500 ms). OD600-normalized luminescence values were calculated by dividing the raw luminescence by the background-subtracted OD600. The background value was established as the absorbance at 600 nm in wells containing only ddH2O.
Bacterial genome targeting
The bacterial genome targeting assay was carried out as previously described27, a 20-nucleotide sequence of the arcB gene was selected as the target. 200 ng of pSGKP-sgRNA plasmid was transformed into 50 μL of E. coli electrocompetent cells containing the p15a-RdCas12n plasmid with an electroporation parameter as 0.1-cm cuvette, 1.8 kV, 200 Ω and 25 μF. After recovering in 500 μL of SOC medium for 1.0 h at 37 °C, the cultures were diluted with six consecutive 10-fold dilutions. Then, 5 μL of each dilution was spotted onto LB agar plates (75 μg·mL−1 apramycin, 50 μg·mL−1 kanamycin) for incubation at 37 °C for 16–24 h. All target sequences used in this study are listed in Supplementary Data 2.
Cell culture and plasmid transfection
The HEK293 cells were cultured in DMEM medium (Corning) supplemented with 10% FBS (Excell) at 37 °C in 5% CO2. For plasmid transfection, the HEK293T cells were seeded into a 24-well plate with a density of 0.8E5 cells per well. The next day, each well was transfected with 1 μg editing plasmids using lipofectamine 3000 according to the manufacturer’s protocol (Life Technologies). After 24 h post-transfection, successfully transfected cells were selected by the addition of 2 μg·mL−1 puromycin to the cell culture, and the cells were harvested for genome extraction 48 h later.
Assessment of indel efficiencies
The cells transfected with editing plasmid were collected for genomic DNA extraction using the Ezup Column Animal Genomic DNA Purification Kit (Sangon Biotech). Approximately 200 bp regions covering the target loci were amplified with primers containing dual 8-nt barcodes using Phanta Max Super-Fidelity DNA Polymerase Mastermix (Vazyme Biotech Co., Ltd). The DNA products were merged and purified by using TIANquick Midi Purification Kit (Tiangen Biotech, Beijing, China). Subsequently, an amplicon-seq library was prepared with the VAHTS® Universal Plus DNA Library Prep Kit for Illumina® (Vazyme Biotech Co., Ltd) following the manufacturer’s instructions, and subjected to Illumina NovaSeq sequencing (PE150) at the HaploX Genomics Center (Jiangxi, China). The sequencing data were demultiplexed based on the 8-nt barcodes, and the efficiency of indels and substitutions was analyzed using CRISPResso2 (ref. 54). The target sites utilized in this study are detailed in Supplementary Data 2.
Phylogenetic analysis of Cas12n
The 138 Cas12n homologous sequences were gathered using PSI-Blast from NCBI and JGI based on AcCas12n sequence34. Sequences were aligned using Clustal Omega55, and a tree was computed from the final alignment using IQtree56. The best model made on the final set was WAG + F + R6, and bootstrap values were estimated with ultrafast bootstrap with 1000 iterations. Final phylogenetic trees were visualized and annotated using the Interactive Tree Of Life (iTOL) platform57. All compared sequences utilized in this study are detailed in Supplementary Data 1.
Statistics and reproducibility
Statistical analyses used GraphPad Prism (version 8.0.2), and replication numbers are specified in the figure legends and all numerical values are presented as mean ± SD. The experimental findings reported in this study were independently replicated at least three times.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The structure of the RdCas12n–sgRNA–dsDNA complex have been deposited in the Protein Data Bank under the accession code 9J09 (ternary complex structure 1) and 9UDI (ternary complex structure 2). The cryo-EM density map have been deposited in the Electron Microscopy Data Bank under the accession code EMD-61051 (ternary complex structure 1) and EMD-64070 (ternary complex structure 2). The sequencing data generated in this study are available in the NCBI database under accession code PRJNA1261697. The uncropped gel images are included in the Source Data file. Source data are provided with this paper.
References
Ishino, Y., Shinagawa, H., Makino, K., Amemura, M. & Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 169, 5429–5433 (1987).
Yan, W. X. et al. Functionally diverse type V CRISPR-Cas systems. Science 363, 88–91 (2019).
Altae-Tran, H. et al. Uncovering the functional diversity of rare CRISPR-Cas systems with deep terascale clustering. Science 382, eadi1910 (2023).
Makarova, K. S. et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Karvelis, T. et al. Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease. Nature 599, 692–696 (2021).
Altae-Tran, H. et al. The widespread IS200/IS605 transposon family encodes diverse programmable RNA-guided endonucleases. Science 374, 57–65 (2021).
Shmakov, S. et al. Discovery and Functional Characterization of Diverse Class 2 CRISPR-Cas Systems. Mol. Cell 60, 385–397 (2015).
Dong, D. et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Nature 532, 522–526 (2016).
Yamano, T. et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165, 949–962 (2016).
Burstein, D. et al. New CRISPR-Cas systems from uncultivated microbes. Nature 542, 237–241 (2017).
Liu, J. J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Pausch, P. et al. CRISPR-CasPhi from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Carabias, A. et al. Structure of the mini-RNA-guided endonuclease CRISPR-Cas12j3. Nat. Commun. 12, 4476 (2021).
Urbaitis, T. et al. A new family of CRISPR-type V nucleases with C-rich PAM recognition. EMBO Rep. 23, e55481 (2022).
Sun, A. et al. The compact Caspi (Cas12l) ‘bracelet’ provides a unique structural platform for DNA manipulation. Cell Res. 33, 229–244 (2023).
Al-Shayeb, B. et al. Diverse virus-encoded CRISPR-Cas systems include streamlined genome editors. Cell 185, 4574–4586.e4516 (2022).
Karvelis, T. et al. PAM recognition by miniature CRISPR-Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res. 48, 5016–5023 (2020).
Takeda, S. N. et al. Structure of the miniature type V-F CRISPR-Cas effector enzyme. Mol. Cell 81, 558–570.e553 (2021).
Xiao, R., Li, Z., Wang, S., Han, R. & Chang, L. Structural basis for substrate recognition and cleavage by the dimerization-dependent CRISPR-Cas12f nuclease. Nucleic Acids Res. 49, 4120–4128 (2021).
Wu, Z. et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat. Chem. Biol. 17, 1132–1138 (2021).
Wang, Y. et al. Guide RNA engineering enables efficient CRISPR editing with a miniature Syntrophomonas palmitatica Cas12f1 nuclease. Cell Rep. 40, 111418 (2022).
Hino, T. et al. An AsCas12f-based compact genome-editing tool derived by deep mutational scanning and structural analysis. Cell 186, 4920–4935.e4923 (2023).
Su, M. et al. Molecular basis and engineering of miniature Cas12f with C-rich PAM specificity. Nat. Chem. Biol. 20, 180–189 (2024).
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science 365, 48–53 (2019).
Chen, W. et al. Targeted genetic screening in bacteria with a Cas12k-guided transposase. Cell Rep. 36, 109635 (2021).
Xiao, R. et al. Structural basis of target DNA recognition by CRISPR-Cas12k for RNA-guided DNA transposition. Mol. Cell 81, 4457–4466.e4455 (2021).
Wu, W. Y. et al. The miniature CRISPR-Cas12m effector binds DNA to block transcription. Mol. Cell 82, 4487–4502 e4487 (2022).
Omura, S. N. et al. Mechanistic and evolutionary insights into a type V-M CRISPR-Cas effector enzyme. Nat. Struct. Mol. Biol. 30, 1172–1182 (2023).
Bigelyte, G. et al. Innate programmable DNA binding by CRISPR-Cas12m effectors enable efficient base editing. Nucleic Acids Res. 52, 3234–3248 (2024).
Chen, W. et al. Cas12n nucleases, early evolutionary intermediates of type V CRISPR, comprise a distinct family of miniature genome editors. Mol. Cell 83, 2768–2780 e2766 (2023).
Nakagawa, R. et al. Cryo-EM structure of the transposon-associated TnpB enzyme. Nature 616, 390–397 (2023).
Sasnauskas, G. et al. TnpB structure reveals minimal functional core of Cas12 nuclease family. Nature 616, 384–389 (2023).
Tang, N. & Ji, Q. Miniature CRISPR-Cas12 systems: mechanisms, engineering, and genome editing applications. ACS Chem. Biol. 19, 1399–1408 (2024).
Wu, Z. W. et al. Structure and engineering of miniature Acidibacillus sulfuroxidans Cas12f1. Nat Catal 6, 695–709 (2023).
Wang, Y. et al. Characterization and engineering of a novel miniature Eubacterium siraeum CRISPR-Cas12f system. ACS Synth. Biol. 13, 2115–2127 (2024).
Xu, X. et al. Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing. Mol. Cell 81, 4333–4345.e4334 (2021).
Wu, T. et al. An engineered hypercompact CRISPR-Cas12f system with boosted gene-editing activity. Nat. Chem. Biol. 19, 1384–1393 (2023).
Bigelyte, G. et al. Miniature type V-F CRISPR-Cas nucleases enable targeted DNA modification in cells. Nat. Commun. 12, 6191 (2021).
Kim, D. Y. et al. Efficient CRISPR editing with a hypercompact Cas12f1 and engineered guide RNAs delivered by adeno-associated virus. Nat. Biotechnol. 40, 94–102 (2022).
Wang, Y. et al. CRISPR-Cas9 and CRISPR-assisted cytidine deaminase enable precise and efficient genome editing in Klebsiella pneumoniae. Appl. Environ. Microbiol. 84, e01834-18 (2018).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Swarts, D. C. & Jinek, M. Mechanistic insights into the cis- and trans-acting DNase activities of Cas12a. Mol. Cell 73, 589–600.e584 (2019).
Quan, J. & Tian, J. Circular polymerase extension cloning of complex gene libraries and pathways. PLoS ONE 4, e6441 (2009).
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PLoS ONE 3, e3647 (2008).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D. Struct. Biol. 74, 531–544 (2018).
Hubbard, B. P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Madeira, F. et al. The EMBL-EBI Job Dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res. 52, W521–W525 (2024).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Acknowledgements
This work was financially supported by grants 22277078 (Q.J.) and 22207074 (Zhaowei Wu) from the National Natural Science Foundation of China, 2023ZD0500500 (Zhaowei Wu) from Noncommunicable Chronic Diseases-National Science and Technology Major Project, 23HC1400800 (Q.J.) from the Shanghai Committee of Science and Technology, China, 2022YFC3403400 (Q.J.) and 2023YFC3400200 (Q.J.) from the National Key R&D Program of China, SPST-YSFZ-2024-01 (Q.J.) from School of Physical Science and Technology, ShanghaiTech University, and 2025AI4S04002 (Q.J.) from ShanghaiTech AI4S Initiative, ShanghaiTech University. The authors thank the Bio-EM facility at ShanghaiTech University for cryo-EM data collections. Additionally, the authors also thank the staff members of the Electron Microscopy System (https://cstr.cn/31129.02.NFPS.EMAS) at the National Facility for Protein Science in Shanghai (https://cstr.cn/31129.02.NFPS), for providing technical support and assistance in data collection and analysis. The work was supported by HPC Platform of ShanghaiTech University. ChatGPT-4 was used for language editing.
Author information
Authors and Affiliations
Contributions
W.F. and J.M. contributed equally to this work. Q.J. conceived the initial study. W.F. and Q.J. designed the research. W.F., J.G. determined the structure of the RdCas12n effector complex. W.F., J.M. performed plasmid construction, protein purification and biochemical experiments. W.F. performed the genome editing in E. coli and in vitro experiments. J.M. performed the genome editing in human cells. Zhipeng Wang searched all Cas12n homologous sequences and W.F. performed the phylogenetic tree analysis. N.T., D.P., M.S., Zhaowei Wu helped with other experiments. W.F., J.M., Q.J. analyzed and discussed the experimental data. W.F., Q.J. prepared the figures and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
Zhaowei Wu and Q.J. are cofounders of Castalysis Bioscience. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jun-Jie (Gogo) Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fu, W., Ma, J., Wang, Z. et al. Mechanisms and engineering of a miniature type V-N CRISPR-Cas12 effector enzyme. Nat Commun 16, 5667 (2025). https://doi.org/10.1038/s41467-025-61290-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61290-3