Abstract
Hydrogel materials have emerged as versatile platforms for various biomedical applications. Notably, the engineered nanofiber-hydrogel composite (NHC) has proven effective in mimicking the soft tissue extracellular matrix, facilitating substantial recruitment of host immune cells and the formation of a local immunostimulatory microenvironment. Leveraging this feature, here we report an mRNA lipid nanoparticle (LNP)-incorporated NHC microgel matrix, termed LiNx, by incorporating LNPs loaded with mRNA encoding tumour antigens. Harnessing the high transfection efficiency of LNPs in antigen-presenting cells, LiNx demonstrates substantial levels of immune cell recruitment, antigen expression and presentation, and cellular interaction. These attributes collectively create an immunostimulating microenvironment and yield a potent immune response with a single dose at a level comparable to the conventional three-dose LNP immunization protocol. Further investigation reveals that the LiNx generates not only high levels of Th1 and Th2 responses, but also a distinct Type 17 T helper cell response critical for bolstering antitumour efficacy. Our findings elucidate the mechanism underlying LiNx’s role in potentiating antigen-specific immune responses, presenting a strategy for cancer immunotherapy.
Similar content being viewed by others
Introduction
The success of two mRNA vaccines, Spikevax® (Moderna) and Comirnaty® (BioNTech/Pfizer),1 in combating SARS-CoV-2 during the coronavirus pandemic demonstrates the safety and efficacy of the lipid nanoparticle (LNP) delivery platform for mRNA vaccines and provides impetus to the rapid expansion of mRNA LNP-based cancer vaccines and cell therapy.2 These LNP vaccines achieve protection by eliciting high antibody titre, memory B cell, and T follicular helper cell responses;3 some evidence also suggests the involvement of IFN-γ+ or IL2+CD8+ T cells and CD4+ Th1 cells.4 The collective findings from Phase 1/2 clinical studies of several mRNA vaccines for high-risk melanoma,5 non-small-cell lung cancer,6 pancreatic cancer,7 and other cancers have demonstrated the feasibility of mRNA-based cancer vaccines.8 The convergence of state-of-the-art LNP technology with personalised medicine approaches amplifies the optimism surrounding this strategy.9 Nonetheless, realising the full potential of mRNA LNP cancer vaccines requires further optimisation of LNP immune activation profiles and delivery strategies to permit more efficient programming of immune responses for potent anticancer efficacy.4
Recent literature reports that not only the choice of lipid components but also the relative proportions of the lipid ingredients in the formulation greatly influence the in vivo transfection efficiency and delivery outcomes.4,10,11 To systematically optimise LNP formulations for cell-preferential transfection, we developed a step-wise screening method that combines in vitro and in vivo assessment steps using a cohort of 1,080 LNP formulations prepared by varying molar ratios of luciferase pDNA or mRNA, ionisable lipid (DLin-MC3-DMA), cholesterol, DMG-PEG2000, and, more importantly, a helper lipid selected from those that were previously used in FDA-approved or experimental LNP formulations.10 The helper lipids were selected to represent various charge features, including cationic [1,2-dioleoyl-3-trimethylammonium-propane (DOTAP) and dimethyl dioctadecyl ammonium (DDAB)], zwitterionic [1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE) and 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC)], and anionic [1,2-dimyristoyl-sn-glycero-3-phosphate (14PA) and 1-stearoyl-2-oleoyl-sn-glycero-3-phospho-(1’-rac-glycerol) (18PG)].10 The LNPs demonstrating the highest levels of in vitro transfection were initially identified through screening tests, followed by individual formulation assessments to pinpoint the most effective LNPs. This screening identified three top-performing LNPs (C10, D6, and F5) based on transfection efficiency in primary bone marrow-derived dendritic cells (BMDCs).4 Following three doses of s.c. injections, these three LNPs induced comparably potent antigen-specific Th1 responses but substantially different Th2 responses.4 All three formulations showed significant tumour suppression and markedly prolonged survival in a prophylactic model of OVA-expressing melanoma in C57BL/6 mice.4 Compared with Th1-biased F5 and D6 LNPs, a dual Th1/Th2 activating C10 LNPs induced the highest level of potency in suppressing tumour growth and highest survival when tested in therapeutic melanoma models.4 Interestingly, the different levels of Th2 responses generated by these 3 selected LNP formulations also correlated with their distinct transfection abilities in non-APCs such as myoblasts. These data revealed that intricate tuning of the LNP composition will enable us to alter transgene expression in APCs and non-APCs when delivering antigen-encoding mRNA in vivo. Further, by biasing cell-preferential gene expression, we altered the immune activation profile elicited by these LNPs.4
Hydrogels have been increasingly proposed as delivery carriers for vaccines through various modes of action, such as sustained release of antigens and/or adjuvants, recruitment of host cells to the site of injection, enhancement of the interaction of antigens and immune cells, etc.12,13 The hydrogel-based vaccine delivery platform is capable of orchestrating an immunostimulatory niche if the key cellular components are present in the local microenvironment.14,15 Such a delivery system has a strong potential in augmenting immune responses.16,17,18 We have recently developed a biostimulatory nanofiber-hydrogel composite (NHC) that mimics the microarchitecture and mechanical properties of the extracellular matrix of soft tissues.19,20 The NHC has an integrated composite structure where electrospun poly(ε-caprolactone) (PCL) nanofiber fragments are covalently crosslinked to a hyaluronic acid (HA) network through covalent interfacial bonding. The NHC offers tunable stiffness, pore size, and degradation duration by adjusting HA structure and concentration, crosslinker concentration, and nanofiber concentration.21 A highly porous NHC with a low shear storage modulus (G’: 150–350 Pa) and 1–3 w/v% PCL nanofibers showed a high level of biostimulatory, proangiogenic, and pro-regenerative properties without using any additional exogenous biochemical cues. Using a simple mechanical fragmentation method, a particulated microgel form of the NHC can be generated for ease of injection. These NHC microgels promoted host cell recruitment, immunomodulation, angiogenesis, soft tissue remodelling, and tissue repair. Moreover, the unique property of recruiting a significant number of immune cells to form a local immunostimulatory microenvironment after subcutaneous (s.c.) or intramuscular (i.m.) injection opens up exciting possibilities. With the advent of newly developed mRNA vaccines,22 leveraging this property can potentiate efficacy by modulating the transfection of the local cell types and coordinating the recruited host immune cells.
In this work, we aim to leverage the immunostimulatory microenvironment generated by NHC4,10,11,14,19,21,23 and the potent transfection efficiency and immune-activation capability of LNPs16,20 by incorporating LNPs into the NHC composite. This mRNA LNP-incorporated NHC microgel matrix, termed LiNx, creates a conducive niche to enhance the transfection of APCs and non-APCs at the injection site, and, more importantly, facilitates the interactions and signalling of all relevant immune cell types in the local niche (Fig. 1a). In detail, we formulate three LiNx variants using top-performing LNPs from our library and show that the NHC microgel enhances immune cell recruitment and cell-type-specific transfection. LiNx induces robust antigen presentation and activation of multiple immune cell subsets, and a single dose elicits antigen-specific responses comparable to a conventional three-dose LNP regimen. Furthermore, LiNx synergises with systemic immune checkpoint blockade to improve therapeutic efficacy in cancer models. These results demonstrate that LiNx effectively amplifies mRNA vaccine-induced immunity by combining biomaterial-based immune recruitment with targeted LNP delivery.
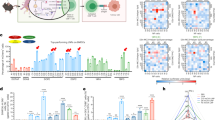
a Schematic of the LiNx vaccination platform, providing an immunostimulatory niche by recruiting host immune cells to the microgel matrix (1). LiNx-delivered LNPs can transfect non-APCs (2) and recruited macrophage-like cells, followed by antigen expression, processing, and epitope presentation on MHC-I (3) or MHC-II (4) in different pathways. b, c Kinetics of host cell recruitment following injection of three different LiNx in C57BL/6 mice (30 μg mOVA per mouse, s.c.). The number of living cells was counted within composites on both day 3 and day 7 post-injection (b). Flow cytometry was employed to determine the count of macrophage-like cells (CD3–CD11b+Ly6g–CD11c–) and DC-like cells (CD3–CD11b+ Ly6g–CD11c+) on day 3 (c). d, f Ai9 mice were administered the three LiNx loaded with mCre via s.c. injections (30 μg mCre per mouse). Transfection of cells in the composite was analysed by flow cytometry. The number of cells positive for tdTomato (d), as well as DC-like cells (CD11b+CD11c+) cells (e) and macrophage-like cells (CD11b+CD11c–) (f) positive for tdTomato on day 5 and day 10 post-injection were shown. g–i C57BL/6 mice received s.c. injections of D6 LiNx loaded with either mLuc or Cy5-labelled mRNA (30 μg per mouse). g The transfection efficiency of D6 LiNx at the injection site and in major organs was assessed using IVIS imaging. The relative percentage of transfection and biodistribution for D6 LiNx is shown in (h). IVIS images of luciferase transfection for D6 LiNx are presented in (i). Data represent the mean ± s.e.m. (n = 8 mice for (b, c), n = 4 mice for (d–f) and n = 5 mice for (g–i)). Box plots in (g) represent the median (centre line), the 25th and 75th percentiles (bounds of the box), and the minimum and maximum values (whiskers); individual data points represent values from independent biological replicates. Data were analysed using one-way ANOVA and Tukey’s multiple comparisons test. *P < 0.05, **P < 0.01, ****P < 0.0001. NS, not significant. Source data are provided as a Source Data file. Figure 1a was created in BioRender.
Results
Host cell recruitment and transfection profile of mRNA LiNx
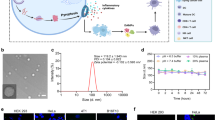
The three top-performing mRNA LNP formulations (C10, D6, and F5 LNPs, Supplementary Table 1) identified from our previous study were selected to prepare the LiNx.4 All three mRNA LNPs led to high levels of expression of the OVA-derived SIINFEKL peptide on major histocompatibility complex class I (MHC-I) on BMDCs and enhanced maturation of BMDCs, marking them as promising candidates for exploration in the context of the LiNx. The NHC microgel with a shear storage modulus G′ of 250 Pa was chosen for its previously reported effectiveness in cell recruitment, migration, and retention (Supplementary Fig. 1a). We mixed the NHC microgel particles with mRNA LNPs using the syringe extrusion method immediately before injection (Supplementary Fig. 1b). Scanning electron microscopy images revealed that, after encapsulating LNPs within the NHC, the same fibrillar microarchitecture of the NHC is observed, with nanofibers entrapped within the HA hydrogel network (Supplementary Fig. 2). The accelerated stability test of NHC at 50 °C showed that, at this temperature, NHC stiffness was reduced by ~40% after 7 days and around 70% after 14 days (Supplementary Fig. 3). Based on a scaling factor calculated empirically, the NHC scaffold used in this study can degrade over a period of 3 to 4 months at 37 °C.
To investigate host cell recruitment within various LiNx formulations loaded with C10, D6, or F5 containing ovalbumin (OVA)-encoding mRNA, or PBS, respectively, we subcutaneously injected the LiNx into the right flank of C57BL/6 mice. The composites were harvested at 3 and 7 days post-injection, and the number of viable cells within the composites was quantified using Acridine Orange/Propidium Iodide (AO/PI) staining. As depicted in Fig. 1b, a substantial number of host cells were recruited inside the scaffold for all groups at both time points. Notably, the incorporation of D6 and F5 mRNA LNPs into the composites markedly increased the number of recruited cells. In comparison to PBS LiNx, there was a 4.2-fold increase for D6-mRNA LiNx and a 2.9-fold increase for F5-mRNA LiNx in the number of cells recruited inside the composites on day 3. By day 7, there was a 2.9-fold increase for D6-mRNA LiNx and a 2.7-fold increase for F5-mRNA LiNx in the number of cells recruited inside the composites relative to the PBS LiNx group. Furthermore, a notable presence of APCs, including macrophage-like cells (CD3–CD11b+Ly6g–CD11c– cells) and DC-like cells (DCs; CD3–CD11b+Ly6g–CD11c+ cells), was identified within the composites on day 3 by flow cytometry analysis. Notably, the D6-mRNA LiNx group exhibited the highest level of recruitment of macrophage-like cells, which were 4.4-fold, 8.13-fold, and 2.0-fold higher than those observed in the PBS, C10, and F5-mRNA LiNx groups, respectively (Fig. 1c and Supplementary Fig. 4).
For an efficient vaccine, the expression profile of the antigen, along with adequate recruitment of host cells, plays a crucial role in eliciting a robust immune response.24 To assess the in vivo delivery efficacy of C10-, D6-, and F5-mRNA LiNx, we conducted s.c. injections of LiNx loaded with Cre-recombinase mRNA (mCre) in genetically engineered tdTomato (tdTom) reporter mice (Ai9 mice).17 These mice contain a LoxP-flanked stop cassette, preventing the expression of the tdTom protein until it is removed by Cre recombinase, allowing for the expression of tdTom. Our findings revealed that all three LiNx formulations resulted in notable levels of tdTom+ cells, including DC-like cells, macrophage-like cells, and CD11c-CD11b- cells in LiNx on days 5 and 10 post-injection (Fig. 1d–f and Supplementary Figs. 5–7). In comparison to C10- and F5-mRNA LiNx, D6-mRNA LiNx exhibited a substantially higher level of transfected cells in the composite. On day 10, the number of transfected cells in D6 LiNx was approximately 129-fold and 117-fold higher than those for the C10 LiNx and F5 LiNx groups, respectively. The D6-mRNA LiNx group also showed approximately a 1000-fold higher number of transfected DC-like cells compared to the other two groups. In addition, the number of transfected macrophage-like cells was 2125-fold and 1691-fold higher in D6 LiNx compared to C10 LiNx and F5 LiNx, respectively. The biodistribution of LiNx formulations was further examined using Cy5-labelled mRNA, as shown in Supplementary Fig. 8a. All three mRNA LNPs were gradually released from the LiNx injection site over two weeks post-injection. Among the formulations, D6 exhibited the most delayed release profile, with approximately 25% of mRNA LNPs still retained within the LiNx platform. In addition, at 24 h post-vaccination, D6 LiNx showed that most LNPs remained at the injection site, with no detectable signal in other major organs (Supplementary Fig. 8b, c). The decrease in signal at the injection site observed initially was likely due to the uptake and degradation of Cy5-mRNA by local or recruited cells. In terms of the transfection efficiency of D6 LiNx was further validated using mLuc mRNA. The data revealed that D6 LiNx transfection was largely confined to the injection site at 24 h post-injection, with approximately 97% of the transfection signal detected there (Fig. 1g–i) and minimal transfection observed in other organs. Taken together, these results underscore the promise of the LiNx platform, especially D6 LiNx, in facilitating in vivo host immune cell recruitment and delivering antigen-encoding mRNA.
mRNA LiNx effectively forms a local immunostimulatory niche
The cellular profiles of infiltrated cells within the composites were assessed at a later time point, specifically on day 14 post s.c. injection using mOVA LiNx. As depicted in Fig. 2a, even after 14 days, a substantially higher number of infiltrating host cells persisted in both D6 and F5 LiNx groups. In comparison to PBS LiNx (PBS + NHC) control and C10 LiNx, D6 LiNx and F5 LiNx showed approximately a 9-fold increase in terms of the number of infiltrated host cells within the composites. Moreover, a substantial increase in the presence of CD3+CD4+ T cells (Fig. 2b and Supplementary Fig. 9), CD3+CD8+ T cells (Fig. 2c and Supplementary Fig. 10), and CD3–CD11b–Npk46–CD19+ B cells (Fig. 2d and Supplementary Fig. 11) were detected in the D6 LiNx compared to other groups, showing a 73-fold higher CD4+ T cell count, a 200-fold higher CD8+ T cell count, and a 305-fold higher B cell count for the D6 LiNx, compared to the PBS LiNx control. The proportion of specific immune cell types on day 14 was assessed for various treatment groups, as illustrated in Fig. 2e (Supplementary Fig. 12). The predominant immunocytes within the composites for PBS LiNx control, C10 LiNx, and F5 LiNx were neutrophils, with approximately 60% of immunocytes within the composites were neutrophils for both PBS LiNx and C10 LiNx groups, and around 80% for the F5 LiNx. In contrast, the D6 LiNx contained substantial amounts of T cells and B cells: approximately 50% of the immunocytes within the LiNx were T cells, ~20% CD8+ T cells, and ~30% CD4+ T cells, while B cells constituted about 20% of the immunocytes. The immunostimulatory niche generated in D6 LiNx, characterised by a substantial influx of T cells and B cells, is highly advantageous for facilitating crosstalk among various immune cell types.
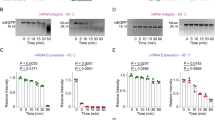
a Recruitment of host cells was assessed two weeks after s.c. administration of three different LNP/mRNA LiNx formulations in C57BL/6 mice (30 μg mOVA per mouse). The Cellaca MX High-throughput Automated Cell Counter was employed to count the living cells within the composites. b–d On day 14, flow cytometry was utilised to determine the counts of CD3+CD4+ T cells (b), CD3+CD8+ T cells (c), and CD11b–CD19+ B cells (d) within the composite. e Composition of host immune cells recruited within the composite was evaluated at two weeks after s.c. administering three different LiNx formulations in C57BL/6 mice (30 μg mOVA per mouse). This analysis included CD4 T cells, CD8 T cells, B cells, DC-like cells (CD11b+ CD11c+), macrophage-like cells (CD11b+CD11c–), neutrophils (CD11b+Ly6G+), and NK cells (CD11b–NKp46+). f–g, RT-PCR array analysis was conducted using RNA isolated from the local microenvironment generated by C10, D6, and F5 LiNx formulations. C57BL/6 mice received the three different LiNx formulations loaded with mOVA through s.c. injection (30 μg mOVA per injection). The presented heatmap shows a panel of genes relevant to immune responses (f). Subsequent RT-PCR analysis of selected genes from the panel confirmed that D6 LiNx promoted critical proinflammatory cytokine transcripts involved in the establishment of a local immunostimulatory niche. These included genes related to inflammatory cytokines and chemokines (IL-6, IL-1α, CSF-3, and CXCL10), Th1 response (TBX21, TNF, IFN-γ, GZMB, NOS-2, IL-12α, IL-15, and IL-2), and genes associated with Th2 response (IL-4, CCR-4, and IL-13). Data represent mean ± s.e.m. (n = 7 mice for a–e and n = 3 mice for f–g). Data were analysed using one-way ANOVA and Tukey’s multiple comparisons test for a–d. *P < 0.05. Source data are provided as a Source Data file.
The local microenvironments influenced by various LiNx formulations were further characterised using real-time polymerase chain reaction (PCR) arrays, analysing RNA extracted from the local injection sites on day 14 post-injection. The mRNA levels related to immune responses were detected, quantified, and analysed (Fig. 2f, g). The findings suggest that, in contrast to other groups, the local injection site for D6 mOVA LiNx displayed increased expression of genes associated with inflammatory cytokines and chemokines, including IL-6, IL-1α, CSF-3, and CXCL10. Moreover, there were elevated levels of genes linked to Th1 immune responses, such as TBX21, TNF, IFN-γ, GZMB, NOS-2, IL-12α, IL-15, and IL-2, as well as genes associated with Th2 immune responses, namely IL-4, CCR-4, and IL-13 (Supplementary Figs. 13–15).
The above results reveal that, at 14 days post-injection, substantial host immune cell recruitment events were observed in the LiNx groups, particularly the D6 LiNx. The majority of these cells were associated with adaptive immune responses, specifically T cells and B cells, rather than neutrophils. The close proximity of these immune cells may enable crosstalk among them, potentially advantageous for the induction of a robust immune response. Moreover, the D6 LiNx group exhibited the formation of an immunostimulatory microenvironment characterised by elevated levels of immunostimulatory cytokines, transcripts related to both Th1 and Th2 responses, as well as an engaged innate inflammatory response.
LiNx induces a potent antigen-specific immune response
The vaccination outcomes of the three LiNx formulations were assessed in C57BL/6 mice following s.c. injections with a single dosage consisting of 30 μg OVA mRNA, with PBS LiNx serving as the control. The LiNx-induced antigen-specific CD8+ T cell response was initially examined by collecting and analysing cells from both LiNx and draining lymph nodes (dLNs) of the treated mice on day 14. As depicted in Fig. 3a, b and Supplementary Figs. 16 and 17, the D6 LiNx demonstrated a substantially higher level of antigen-specific CD8+ T cells compared to the C10 and F5 LiNx groups, with ~64-fold and ~11-fold increases detected inside the niches, respectively. Moreover, within the dLNs, the D6 LiNx exhibited a 3.7-fold and 2.9-fold higher number of antigen-specific CD8+ T cells, compared to the C10 and F5 LiNx groups, respectively. In contrast, the LiNx loaded with empty D6 LNPs did not generate a detectable immune response (Supplementary Fig. 18).
a–h C57BL/6 mice were administered three different LiNx formulations loaded with mOVA via s.c. injection (30 μg per mouse). Mice were sacrificed two weeks after the injection. a, b The number of OVA-specific CD8 T cells within the LiNx (a) and dLNs (b) was analysed by flow cytometry. c–h Their splenocytes were isolated and restimulated in vitro with OVA and SIINFEKL peptides (100 μg mL−1 OVA and 2 μg mL−1 SIINFEKL) for 12 h and assessed via flow cytometry and intracellular cytokine staining to determine the percentages of CD8+IFN-γ+ cells (c), CD8+TNFα+ cells (d), CD8+ Granzyme B+ cells (e), CD4+IFN-γ+ cells (f), CD4+TNFα+ cells (g), and CD4+IL-4+ cells (h). i Frequency of IFN-γ-producing cells among restimulated splenocytes isolated from vaccinated mice on day 30 post-vaccination, assessed via ELISPOT. j Titre of OVA-specific IgG antibodies in blood serum on day 30, determined by ELISA. k–o C57BL/6 mice were administered with three different LNP/mRNA LiNx formulations loaded with mOVA via s.c. injection (30 μg per mouse). Mice were sacrificed three months after the injection, and their splenocytes were isolated for analysis. The percentages of OVA-specific CD8 T cells (CD3+ CD8+ OVA+ cells) (k), OVA-specific effector CD8 T cells (CD3+CD8+OVA+CD44+CD62L– cells) (l), and OVA-specific central memory CD8 T cells (CD3+CD8+OVA+CD44+CD62L+ cells) (m) were determined. Splenocytes were restimulated in vitro with OVA and SIINFEKL peptide (100 μg mL−1 OVA and 2 μg mL−1 SIINFEKL) for 12 h and assessed via flow cytometry and intracellular cytokine staining to determine the percentages of CD3+CD8+TNFα+ cells (n) and CD3+CD8+IFN-γ+ cells (o). p Titre of OVA-specific IgG antibodies in blood serum on day 90, determined by ELISA. Data represent the mean ± s.e.m. (n = 7 mice (a, b), n = 4 mice (c–j), n = 8 mice (k–p)). Data were analysed using one-way ANOVA and Tukey’s multiple comparisons test for (a–p). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Source data are provided as a Source Data file.
The spleens from vaccinated mice were harvested on day 14, dissociated into a cell suspension, and subjected to ex vivo antigen restimulation. The D6 LiNx yielded increased frequencies of CD8+IFN-γ+, CD8+TNFα+, and CD8+Granzyme B+ cell populations (Fig. 3c–e and Supplementary Fig. 19). Specifically, compared to the C10 and F5 LiNx-treated groups, the D6 LiNx exhibited approximately 9.0- and 6.2-fold increases in CD8+IFN-γ+ cell frequencies, respectively. Alongside the robust CD8+ T cell response, the D6 LiNx also showed higher numbers of Th1 cells (CD4+IFN-γ+ and CD4+TNFα+), as depicted in Fig. 3f, g. On day 14 post-vaccination, there was a 16.4-fold higher level of antigen-specific CD4+IFN-γ+ Th1 cells and a 3.0-fold higher level of antigen-specific CD4+TNFα+ Th1 cells for the D6 LiNx group. In contrast, no significant increases were observed in the C10 and F5 LiNx-treated groups. Furthermore, the frequency of antigen-specific CD4+ IL-4+ Τh2 cells was examined for all groups (Fig. 3h). In comparison to the PBS LiNx control, the D6 LiNx group exhibited a higher number of antigen-specific Th2 cells (2.8-fold increase, P = 0.0156), in contrast to the C10 and F5 LiNx groups (1.8- and 1.7-fold higher, respectively; P > 0.5).
The antigen-specific response after vaccination was further evaluated on day 30 post-vaccination. As depicted in Fig. 3i, the splenocytes from vaccinated mice collected on day 30 were subjected to in vitro antigen restimulation. The D6 LiNx induced a substantially higher level of antigen-specific T-cell response (Supplementary Fig. 20). Approximately 3.7-fold and 3.8-fold higher levels of IFN-γ-secreting cells were detected within the D6 LiNx group, compared with C10 LiNx and F5 LiNx, respectively. In addition, as shown in Fig. 3j, D6 LiNx induced a higher OVA-specific IgG titre, including both IgG1 and IgG2c subclass titres, indicating a potent humoral response (Supplementary Fig. 21) as well. The antibody responses generated by the C10 and F5 LiNx groups were limited.
The memory immune responses of LiNx were evaluated at three months post-vaccination. Substantial levels of antigen-specific CD8+ T cells were still present in the spleen of mice in the D6 LiNx group (Fig. 3k and Supplementary Figs. 22 and 23). More importantly, within these T cells, there were both CD44+CD62L– effector T cells and higher levels of CD44+CD62L+ central memory T cells. Specifically, the D6 LiNx group exhibited approximately a 23.8-fold higher level of antigen-specific effector T cells and a 10.1-fold higher level of central memory T cells, underscoring the establishment of long-term memory responses (Fig. 3l, m). Splenocytes harvested from vaccinated mice on day 90 were also subjected to ex vivo antigen restimulation. As depicted in Fig. 3n, o, the D6 LiNx group exhibited 5.0-fold higher CD8+TNFα+ cells than the PBS LiNx control (P = 0.0006) and 5.3-fold higher CD8+IFN-γ+ cells than the PBS LiNx control (P = 0.0001) (Supplementary Figs. 24–26). In addition to T cell responses, a substantially higher OVA-specific IgG titre, including both IgG1 and IgG2c subclass titre, was observed in the D6 LiNx-treated group at 90 days post-vaccination, indicating a potent long-lasting humoral response (Fig. 3p and Supplementary Fig. 27). Furthermore, an examination of the biosafety profiles for C10, F5, and D6 LiNx showed no significant differences in body weight across all LiNx-treated groups throughout the vaccination schedule, and no significant changes in serum cytokine levels (IFN-γ, TNF-α, IL-4, IL-10, and IL-2) post-vaccination in the D6 LiNx treated mice (Supplementary Figs. 28 and 29).
The above results indicate that, following a single-dose vaccination, D6 LiNx effectively elicited more potent antigen-specific Th1 and Th2 responses than C10 and F5 LiNx. Moreover, D6 LiNx vaccination resulted in a strong and persistent memory immune response. In contrast, no memory immune responses were observed three months after vaccination in the C10 and F5 LiNx groups.
Anti-tumour effects induced by a single-dose vaccination of D6 mRNA LiNx
Given the potent antigen-specific immune responses induced by the D6 LiNx, we investigated its efficacy as a cancer vaccine in therapeutic and prophylactic tumour models. In the first study, C57BL/6 mice were s.c. inoculated with 3 × 105 OVA-expressing MC38 colorectal cancer cells on the right posterior side on day 0. On day 4, the mice received vaccinations with C10, D6, or F5 mRNA LiNx, each containing 30 μg of mOVA (Fig. 4a). Control groups included the NHC and OVA protein (30 μg of OVA per mouse) mixed with Alhydrogel® or OVA in NHC. In addition, C57BL/6 mice were immunised with 3 doses of D6 LNPs containing 10 μg of mOVA on days 4, 11, and 18, as a control. As illustrated in Fig. 4b–d, the single dose D6 LiNx exhibited effective tumour suppression in this treatment model, resulting in a median survival time of 75 days compared to 31 days for the negative control group (Supplementary Fig. 30). In the D6 LiNx group, 50% of the mice remained tumour-free beyond 100 days. Mice treated with the NHC, OVA in NHC, OVA in Alhydrogel®, C10 LiNx, and F5 LiNx exhibited limited levels of therapeutic efficacy, leading to a low extension of median survival time. In comparison to the group treated with the standard three-dose D6 mRNA LNP immunisation, which resulted in a median survival time of 37.5 days, a single dose of D6 LiNx produced a markedly improved antitumour effect, demonstrating the capability of LiNx in enhancing the anti-tumour effect of mRNA LNPs.
a–d Schematic and results of a syngeneic therapeutic vaccination model for MC38-OVA in C57BL/6 mice. Mice were inoculated s.c. with MC38-OVA and then given three s.c. injections, one week apart, of mOVA-loaded D6 LNPs (10 μg mOVA per injection) or PBS. For the LiNx treatment group, mice were administered three different LiNx formulations loaded with mOVA LNPs (30 μg per mouse) or OVA protein (10 μg protein per mouse) via a single s.c. injection. OVA protein mixed with Alhydrogel® (1:1) (10 μg protein per mouse) served as a control group. Average tumour volume (b), survival curves (c), and individual tumour volume (d) are shown. e–i Schematic and results of a therapeutic vaccine against another syngeneic model, B16F10 melanoma in C57BL/6 mice, using OVA model antigen (f, g) and melanoma-associated antigens (h, i). Mice were inoculated s.c. with B16-OVA (f, g) or B16F10 cells (h, i) and then s.c. administered with D6 LiNx loaded with mOVA, mTrp2 or mGp100 (30 μg per mouse). Three groups received the anti-CTLA-4 mAb (100 μg per i.p. injection) treatment in combination with LiNx treatment. Average tumour volume (f, h), and survival curves (g, i) are shown. j–l Schematic and results of a prophylactic vaccine against melanoma in C57BL/6 mice using OVA model antigen. Mice were s.c. administered with a single dose of D6 LiNx loaded with mOVA (30 μg per mouse) and then inoculated s.c. with B16-OVA cells on day 21. Average tumour volume (k), and survival curves (l) are shown. Data represent mean ± s.e.m. with n = 8 mice (b–d), n = 6 mice (f–i, k, l). Survival curves were compared using the log-rank Mantel–Cox test, and the stack of P-values were corrected by the Holm-Šídák method for multiple comparisons with α set to 0.05. For (b, f, h, k), data were analysed using two-way ANOVA and Tukey’s multiple comparisons test. *P < 0.05, **P < 0.01, ***P < 0.001; ****P < 0.0001; NS, not significant; i.p. intraperitoneal; \(\alpha\)CTLA-4, anti-CTLA-4 mAb. Source data are provided as a Source Data file. Figures 4a, e and j were created in BioRender.
We further evaluated the therapeutic efficacy of D6 LiNx in the B16-OVA melanoma tumour model, utilising the model OVA antigen. C57BL/6 mice were s.c. inoculated on the right posterior side with 3 × 105 B16-OVA cells on day 0. On day 4, with an average tumour size of ~25 mm³, mice were vaccinated with D6 LiNx containing 30 μg mOVA (Fig. 4e). As depicted in Fig. 4f, g, D6 LiNx demonstrated a stronger tumour suppression effect in this treatment model, resulting in a median survival time of 30.5 days compared to 17 days for the negative control (Supplementary Fig. 31). As shown in Fig. 2f, the local microenvironment at the injection site in D6 LiNx-treated group exhibited a highly elevated CTLA-4 marker, which prompted us to combine anti-CTLA-4 therapy with the LiNx vaccination to evaluate their combined effect. When D6 LiNx was administered in conjunction with an immune checkpoint inhibitor (100 μg anti-CTLA-4 monoclonal antibody, given i.p. on days 6, 13, 20, and 27), a synergistic effect was observed, extending the median survival time to 42 days. In contrast, no significant tumour suppression effect was observed in the group treated solely with anti-CTLA-4 antibody, as compared to the PBS LiNx control.
Subsequently, the D6 LiNx was evaluated in the syngeneic melanoma mouse model using two clinically relevant tumour antigens, tyrosinase-related protein 2 (Trp2) and glycoprotein 100 (Gp100) (Fig. 4e). C57BL/6 mice were s.c. inoculated on the right posterior side with 3 × 105 B16F10 cells on day 0. On day 4, with an average tumour size of ~ 25 mm³, mice were vaccinated with D6 LiNx containing 30 μg of mRNA encoding either Trp2 or Gp100. The potent anti-tumour effect was observed with these two antigens, resulting in prolonged median survival times of 22.5 and 21.5 days for D6 mTrp2 LiNx and D6 mGp100 LiNx, respectively (Fig. 4h, i and Supplementary Fig. 32). However, no significant improvement was observed when combining these two D6 LiNx formulations with anti-CTLA-4 antibody treatment.
Beyond the therapeutic cancer model, D6 LiNx was also assessed in a prophylactic B16-OVA melanoma model. C57BL/6 mice were immunised on day 0 with D6 LiNx containing 30 μg of mOVA. On day 21, animals were s.c. inoculated on the right posterior side with 1 × 106 B16-OVA cells (Fig. 4j). A single injection of D6 LiNx yielded a stronger tumour inhibition efficacy with prolonged overall survival times compared to the PBS, PBS-NHC, and OVA-NHC groups. The median survival time was 30 days for the D6 LiNx group, compared to 16–17 days for the three control groups (Fig. 4k–l and Supplementary Fig. 33). In addition, LiNx loaded with empty D6 LNPs did not generate a significant anti-tumour effect (Supplementary Fig. 34).
To further confirm the efficacy of the D6 mOVA LiNx and assess its long-term protective effect, we conducted a tumour rechallenge study with an increased sample size (Supplementary Fig. 35). C57BL/6 mice were immunised as previously described and, on day 21, were subcutaneously inoculated with 3 × 105 B16-OVA cells on the right posterior side. Survival was monitored over 100 days, and 17 of 18 mice (94.4%) remained tumour-free following the single D6 LiNx vaccination (Supplementary Fig. 35b). These mice were subsequently rechallenged with 3 × 105 B16-OVA cells on the right posterior side. As shown in Supplementary Fig. 35c, 11 of 17 (64%) mice remained tumour-free post-rechallenge, demonstrating the long-term protective effect of D6 LiNx. These findings further confirm the antitumour efficacy of the D6 LiNx platform and highlight its potential for sustained protection.
Modulation of the tumour microenvironment by LiNx
To assess the effect of LiNx treatment on the tumour microenvironment, we analysed the infiltrating immune cells in the tumour mass. As shown in Fig. 5a, C57BL/6 mice were inoculated with 3 × 105 B16-OVA cells on day 0. Starting on day 4, mice received three subcutaneous (s.c.) injections of mOVA-loaded D6 LNPs (10 μg mOVA per injection) or PBS at one-week intervals (days 4, 11, and 18). For the LiNx treatment group, mice were administered the D6 LiNx formulations (30 μg per mouse) via a single s.c. injection on day 4. On day 21, the mice were euthanized, and tumour tissues were collected, homogenised, and analysed by flow cytometry. As shown in Fig. 5b, treatment with the D6 LiNx resulted in a substantial increase in CD8+ T cells within the tumour microenvironment. Approximately 4.5% of the total cell population consisted of CD8+ T cells, representing a three-fold increase compared to treatment with 3 doses of the D6 LNPs, which only induced about 1.5% CD8+ T cells in the tumour. In addition, a 3.9-fold higher (1.3%) OVA-specific CD8+ T cells among the total cell population in the tumour were observed in the D6 LiNx group, compared to the 3 doses of D6 LNPs (Fig. 5c). Similarly, higher levels of CD4+ T cells, NK cells, macrophages, and DCs were also observed in the D6 LiNx group (Fig. 5d–g). This indicates that the D6 LiNx induced a robust antigen-specific immune response and modulated the tumour microenvironment.
a–g Schematic and analysis of the tumour microenvironment in a therapeutic vaccination model for B16-OVA in C57BL/6 mice. Mice were inoculated subcutaneously (s.c.) with B16-OVA and subsequently received three weekly s.c. injections of mOVA-loaded D6 LNPs (10 μg mOVA per injection) or PBS. On day 21, tumours were collected and analysed by flow cytometry to determine the percentages of (b) CD3+CD8+B220- T cells, (c) OVA-specific CD3+CD8+B220- T cells, (d) CD45+CD3+CD4+ T cells, (e) NKp46+CD11b-CD3- NK cells, (f) F4/80+CD11b+CD3- macrophages, and (g) MHCII+CD11c+CD11b-Ly6G-CD3- DCs. Data (b–g) represent mean ± s.e.m., and box plots display the median (centre line), the 25th and 75th percentiles (bounds of the box), and the minimum and maximum values (whiskers), n = 5 mice. Data (b–g) were analysed using one-way ANOVA and Tukey’s multiple comparisons test. Exact P-values are shown; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, NS not significant. h–k CODEX multiplex imaging analysis of tumour tissue collected on day 14 post B16-OVA inoculation. For the LiNx treatment group, mice were administered with the D6 LiNx formulation (30 μg mOVA per mouse) via a single s.c. injection on day 4. (h) Dotplot of cell types by marker expression derived from CODEX multiplexed imaging for the pooled D6 LiNx (n = 2 mice) and PBS-treated tumours (n = 3 mice) using unsupervised clustering with the percentage of cells with a marker Z-score above 0.7. (i) Percentage distribution of immune and tumour cell types within the D6 LiNx (n = 2 mice) and PBS-treated tumours (n = 3 mice) as quantified by CODEX multiplexed imaging. j Comparison of the normalised percentage of immune cell subsets within the D6 LiNx-treated tumours (n = 2 mice) and PBS-treated tumours (n = 3 mice). k Cell type map of the representative D6 LiNx tumour derived from CODEX multiplexed imaging with coloured data points representing selected immune and tumour cell types with other cell types lumped together in the same grey colour (scale bar = 500 μm). Source data are provided as a Source Data file. Figure 4a was created in BioRender.
To further validate the effect of the LiNx platform on the tumour microenvironment, we employed CODEX multiplexed fluorescence microscopy for spatial proteomics that uses iterative imaging and DNA-barcoded antibodies to enable the simultaneous imaging of multiple markers.25,26,27,28 We designed a panel to identify major adaptive and innate immune cell types (Fig. 5h and Supplementary Tables 2–4). C57BL/6 mice were inoculated with 3 × 105 B16-OVA cells on day 0 and were treated with the D6 LiNx (30 μg mOVA per mouse) via a single s.c. injection on day 4. On day 14 post-tumour inoculation, tumour tissues were collected, sectioned, and analysed. As shown in Fig. 5i–k and Supplementary Figs. 36–37, infiltration of multiple immune cell types, including CD8+ T cells, CD4+ T cells, B cells, NK cells, and CD169+ macrophages, was observed in the tumour regions for D6 LiNx formulation. These findings are consistent with the results shown above that the D6 LiNx formulation facilitates immune cell infiltration into the tumour microenvironment. Notably, recent studies have shown that a high density of CD169+ macrophages is associated with prolonged survival and favourable clinical outcomes in patients with tumours.29,30,31,32,33,34,35
Taken together, our results reveal that treatment with the D6 LiNx altered the tumour microenvironment by affecting the infiltration of T and B cells, which are involved in the adaptive immune response. These results highlight the potency of the D6 LiNx platform as a designed immunostimulatory vaccine platform.
LiNx amplifies the immune response of mRNA LNPs
To further differentiate the immune responses triggered by D6 mRNA LNPs and LiNx, we analysed the expression of 547 genes in splenocytes from vaccinated mice using the nCounter Analysis System (NanoString Technology). In detail, the D6 LiNx comprising 30 μg mOVA was s.c. injected on day 0, compared with three doses of D6 LNPs containing 10 μg mOVA each on days 0, 7, and 14. On day 21, spleens from vaccinated mice were collected, dissociated into a single-cell suspension, and then subjected to in vitro antigen restimulation. After 24 h, splenocytes were collected, and the RNA was extracted and examined using NanoString analysis. As depicted in Fig. 6a, b, the comparison between untreated mice and the D6 LiNx group revealed 376 distinctive genes, including 25 downregulated and 351 upregulated genes. When comparing the single-dose D6 LiNx with the three-dosage D6 LNPs treatment, 344 genes exhibited significant differences, consisting of 35 downregulated and 309 upregulated. Upon closer examination of various subsets of related genes, including Th1, Th2, Th17, and Treg-related genes,25 as well as B cell receptor signalling genes, the D6 LiNx-treated group exhibited upregulation in 6 out of 15 Th1-related genes, 9 out of 16 Th2-related genes, 13 out of 19 Th17-related genes, and 12 out of 30 B cell signalling-related genes, in comparison with a 3-dose free-D6 LNP control group (Fig. 6c–e). Among Treg cell-related genes, there were no significant differences, except for the downregulation of 1 out of 7 genes in the D6 LiNx group. By employing Database for Annotation, Visualisation, and Integrated Discovery (DAVID) functional annotations for Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathways to establish connections between the information and higher-order functional aspects, we discovered that the D6 LiNx platform augmented immune responses through multiple pathways (Fig. 6f). In addition to its enhancement effect on Th1 and Th2 differentiation, antigen presentation, and B cell/T cell receptor signalling pathways, a noteworthy revelation emerged: Th17 differentiation and the IL-17 signalling pathway exhibited marked enhancement following D6 LiNx treatment. This unexpected finding suggests that the increased effectiveness of D6 LiNx in anticancer efficacy, as compared to free D6 LNPs, may be attributed to the substantial enhancement of Th17 cell-mediated anti-tumour immunity, which has been recognised for its considerable therapeutic potential (Fig. 6g).26,27,28,29,30,31,32,33 To validate the functionality of the Th17 immune response induced by the D6 LiNx, we performed IL-17 depletion experiments using a prophylactic vaccination model for OVA-expressing melanoma in C57BL/6 mice. The mice received either a single s.c. injection of the D6 LiNx (30 μg mOVA per injection) or three s.c. injections of mOVA-loaded D6 LNPs (10 μg mOVA per injection) administered one week apart, followed by s.c. inoculation with B16-OVA cells. IL-17-depleting antibodies were administered i.p. every three days (200 μg per mouse). The results demonstrated that IL-17 depletion led to approximately a 40% reduction in survival rate in the D6 LiNx treatment group (Fig. 6h). In contrast, IL-17 depletion had no significant effect on the survival rate of the traditional 3-dose D6 mRNA LNP treatment group, underscoring the critical role of the Th17 immune response in the observed anti-tumour effect of the D6 LiNx formulation. Taken together, while Th17 plays an important role, the Nanostring and immune profiling data indicated that the D6 LiNx induced a more robust immune response overall, including enhanced Th1 and Th2 responses compared to the free D6 LNPs. Given this broader immune activation, we do not expect that depleting Th17 alone would fully replicate the survival pattern observed with the D6 LNP treatment. Moreover, no significant changes in lymphocyte composition were detected in the blood of α-IL-17-treated C57BL/6 control mice, indicating that antibody treatment does not substantially alter the immune system (Supplementary Fig. 38).
C57BL/6 mice were given three s.c. injections, one week apart, of mOVA-loaded D6 (10 μg mOVA per injection) or PBS. For LiNx treatment group, mice were administered D6 LiNx loaded with mOVA (30 μg per mouse) on day 0. On day 21 post-vaccination, the splenocytes were restimulated in vitro with OVA and SIINFEKL peptide (100 μg mL−1 OVA and 2 μg mL−1 SIINFEKL) for 12 h, and the RNA was extracted and assessed using a NanoString nCounter Analysis System. a, b Volcano plot illustrates the differentially expressed genes between the D6 LiNx and PBS groups (a), as well as between the D6 LiNx and D6 LNP groups (b). P < 0.05; two-sided unpaired limma-moderated t-test; absolute fold change ≥ 1.5. c, d Selected genes related to B cell signalling, Th1, Th2, Th17, and Treg differentiation are shown. e Heatmap presenting a panel of genes relevant to immune responses for D6 LNP and D6 LiNx group. f Bar plots showing differential pathways enriched after vaccination of a single-dose D6 LiNx compared with PBS or three doses of D6 LNPs using Database for Annotation, Visualisation and Integrated Discovery (DAVID) functional annotations for Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathways. Data represent mean ± s.e.m. d, e Experiments from (a–f) were conducted with n = 6 mice for the D6 LiNx group and n = 3 mice for PBS LiNx and D6 LNP groups. Pathway enrichment results in (f) were analysed using a two-sided unpaired limma-moderated t test (absolute fold change ≥ 1.5, P < 0.05). g Schematic of the immunostimulatory niche generated by LiNx through effective recruitment of host immune cells to the microgel matrix, leading to antigen presentation to CD8+ and CD4+ T cells (including Th1, Th2, and Th17 cells). h Schematic and results of IL-17 depletion experiments in the prophylactic vaccination model for OVA-expressing melanoma in C57BL/6 mice. Survival curves over time are shown (n = 6 mice for the PBS group and 8 mice for other groups). Survival curves were compared using the log-rank Mantel–Cox test, and the stack of P-values were corrected by the Holm-Šídák method for multiple comparisons with α set to 0.05. Source data are provided as a Source Data file. Figures 6g, h were created in BioRender.
Discussion
Previous reports on LNP-mediated gene delivery and mRNA vaccination revealed that both the choice of lipids34 and their molar ratios significantly impact various aspects,10 including the encapsulation efficiency35 of nucleic acid payloads36, transfection efficiency,10 targeting profiles within cells and tissues,37 as well as the immune response profile.4 Despite enormous potential, the spatial and temporal control over the immune response still persists in LNP-based engineering.38 In this study, we tested the LiNx concept, an mRNA LNP-loaded microgel matrix, which promotes endogenous immune cell recruitment and retention, and creates a programmable immunostimulating microenvironment, therefore maximising antigen expression and presentation locally. This configuration yields a more robust immune response with a single-dose vaccination in mouse colon carcinoma and melanoma models.
Here, we employed a previously developed NHC due to its ability to persistently recruit and retain host immune cells following s.c. or i.m. injection. This is crucial for facilitating effective crosstalk between relevant immune cell types. We tested three different LNPs with potent transfection capabilities of BMDCs in the LiNx context. Notably, the D6 LiNx, among the three, establishes the most effective local niche at week 2, characterised by a high abundance of both T cells and B cells, in contrast to the predominant neutrophil or macrophage composition observed in the C10 and F5 LiNx groups. The formation of a lymphoid tissue-like niche provides an immunostimulatory environment favourable for the induction of an adaptive immune response. Interestingly, substantial differences were observed regarding host cell recruitment, transfection efficiency, local microenvironment, and anti-cancer immune responses among the three LiNx formulations. These variations may be attributed to differences in LNP stability, transfection profiles, and/or immunostimulatory cues provided by the LNPs themselves. The NHC scaffold degrades gradually over 3–4 months at the physiological temperature, but the mRNA-LNP component exhibits a much shorter duration of activity. The transgene expression of D6 LiNx peaked within 24 to 48 h post-injection and diminished within 3–5 days (Supplementary Fig. 39). In addition, LNPs from LiNx were released and cleared within a similar time period (around 8 days, Supplementary Fig. 8). Collectively, the overall degradation characteristics of LiNx are primarily determined by the NHC matrix. The primary contributing factor to the enhanced immune response does not come from an extension of LNP presence or transgene expression at the immunisation site, but rather from the recruitment and retention of relevant immune cells, persistent of the cellular milieu, and facilitation of cellular communication and paracrine signalling among the transfected cells and immune cells (Fig. 2). In future applications of the LiNx platform, it is imperative to screen combinations of different LNPs and NHC. In addition, physical properties such as the stiffness and biodegradability of NHC may also influence the transfection and immune response outcomes. Therefore, optimising NHC for enhanced immune responses should involve a comprehensive examination of these physical attributes.
When we analysed the tumour microenvironment following treatment with the D6 LiNx platform using flow cytometry and CODEX multiplex imaging, we observed a marked increase in adaptive immune cells, particularly antigen-specific T cells, infiltrating the tumour. Notably, the D6 LiNx treatment group also exhibited a substantial presence of NK cells and CD169+ macrophages. The elevated NK cell levels are particularly striking, given their critical role in the innate immune response and the ability to directly target and eliminate tumour cells. Similarly, CD169+ macrophages, known for their role in antigen presentation, may facilitate T cell activation within the tumour microenvironment.29,30,31,32,33,34,35 These findings highlight the enhanced potency of the D6 LiNx platform, which strongly correlates with robust activation of both adaptive and innate immune responses. This dual activation is likely to contribute to a more sustained and effective immune response against tumours. Collectively, our results underscore the potential of the D6 LiNx platform as a highly effective immunostimulatory vaccine platform.
When comparing a single dose of D6 LiNx to three doses of conventional D6 LNPs, we observed enhanced immune responses and therapeutic efficacy with D6 LiNx. This phenomenon may be attributed to the initial successful transfection of host cells by D6 LNPs in the LiNx, leading to continuous host cell recruitment and effective crosstalk among recruited cells in LiNx, creating a local environment conducive to the potentiation of the immune response. In this study, we selected a 30 µg OVA mRNA dose for the LiNx groups to match the total mRNA dose used in the free OVA mRNA-LNP conditions. In the recombinant antigen control group, a 30 µg OVA protein dose was used. Nonetheless, we did not perform a dosage titration study for OVA protein antigen, and it remains possible that other OVA doses could elicit a stronger response. Dose responses for all the test groups discussed here should be explored more thoroughly in future studies.
In this study, we did not directly compare the D6 LiNx, which showed superior efficacy than three doses of free D6 LNPs, with the FDA-approved mRNA-LNP vaccines such as SM102 and ALC-0315. Previous studies from our group have shown that the D6 free LNPs exhibit a comparable level of anti-tumour efficacy as the SM102 LNPs under equivalent dosing and scheduling conditions.4 Importantly, the LiNx represents a complementary strategy that could further enhance the performance of other LNP formulations, even though the exact incorporation process and composite features will need to be fine-tuned. Under the optimised conditions, integrating LNPs into the LiNx system can potentially enhance antigen expression, immune cell recruitment, retention, and cell-cell signalling and create a more immunostimulatory microenvironment, thereby amplifying vaccine efficacy. The mRNA LNP vaccines have already been used in clinical settings, and extensive R&D efforts have been devoted to the development of mRNA LNP cancer immunotherapies. The NHC matrix has successfully completed a Phase 1 clinical trial as a biostimulatory dermal filler, demonstrating its safety as a standalone biodegradable matrix. These safety track records will accelerate the translational process from bench to bedside. In addition, the LiNx represents a complementary strategy that could further enhance the performance of LNP formulations. It is possible to mix LNPs into NHC right before injection to preserve the stability features of both LNPs and NHC, thus forming the LiNx in situ, to create an immunostimulatory niche, thereby amplifying vaccine efficacy. More interestingly, our analysis revealed that the LiNx elicited a more robust Th17 response, in addition to Th1 and Th2 responses, suggesting the critical role of Th17 cells in generating potent antitumour efficacy in this context (Fig. 5g).
While our current LiNx platform primarily functions as an immune activation niche, its future iterations could be applied directly in or near the tumour microenvironment to mimic the functions of a tertiary lymphoid tissue. Such a design may incorporate tumour-responsive elements to further enhance efficacy. Dynamic biomaterials capable of responding to specific tumour microenvironmental cues—such as enzymatic degradation, pH sensitivity, or reactive oxygen species-triggered release—could enable more precise and controlled immune modulation.39,40 This approach could facilitate the timely release of immunostimulatory factors, improve immune cell infiltration into tumours, and enhance antigen presentation. Exploring these biomaterial-based design principles within the LiNx system represents an exciting future direction for improving cancer immunotherapy outcomes.
In summary, we present an effective approach to potentiate mRNA LNP cancer vaccine by creating an immune cell-recruiting and retention microgel matrix as a local immunostimulating niche. The LiNx matrix does not rely on lymph node trafficking of LNPs, but rather leverages its capability to locally enhance antigen expression and presentation, as well as immune cell crosstalk, resulting in potent antigen-specific responses. Among the three tested LiNx formulations, D6 LiNx exhibited continuous host cell recruitment and differentiation, as well as Th1, Th2, and Th17 responses that correlate with the most potent antitumour efficacy among the three LiNx formulations and when compared to the traditional three-dose regimen for D6 LNP vaccination. The results showcase a potent strategy to augment immune responses elicited by mRNA LNPs through a biomaterials-enabled lymphoid niche. This presents a versatile vaccine platform applicable to a variety of disease treatment and prevention strategies, thereby expanding the utility of mRNA LNP-based immunotherapies.
Methods
Ethical statement
All animal experiments were conducted in compliance with relevant ethical regulations and approved by the Johns Hopkins University Institutional Animal Care and Use Committee (IACUC) under protocol number MO22E117. Male and female C57BL/6 mice (6–8 weeks old) were obtained from The Jackson Laboratory and housed under standard conditions in accordance with institutional guidelines.
The maximum tumour size permitted by the IACUC was 2000 mm³ or a maximum tumour diameter of 20 mm in any direction. Mice were euthanized when either threshold was reached. We confirm that these limits were not exceeded in any of the experiments reported in this study.
Materials
DLin-MC3-DMA was purchased from MedKoo Biosciences. DOPE, DSPC, 18PG, and DMG-PEG-2000 were obtained from Avanti Polar Lipids. Cholesterol was from Sigma-Aldrich. B16F10 (CRL-6475) were purchased from ATCC (American Type Culture Collection, USA). B16-OVA cells were kindly provided by the lab of Prof. Jonathan Schneck. MC38 OVA (KC-2370) was purchased from KYINNO Biotechnology. All eukaryotic cell lines used in this study were authenticated by short tandem repeat profiling and routinely tested for mycoplasma contamination using the Universal Mycoplasma Detection Kit (ATCC 30-1012 K) every two months. No contamination was detected during the study period. All mRNA (Cre mRNA, OVA mRNA, Trp2 mRNA, or Gp100 mRNA) constructs were purchased from TriLink BioTechnologies. The anti-CTLA-4 (α-CTLA-4) monoclonal antibody (mctla4-mab10-10) was purchased from InvivoGen. CODEX antibody information was compiled in Supplementary Table 4.
Sodium hyaluronate with a molecular weight of 1.5 × 106 Da (HA, research grade) was purchased from LifeCore Biomedical Inc. (Chaska, MN, USA). Glycidyl acrylate was obtained from TCI America Inc. (Portland, OR, USA). Poly(ethylene glycol) dithiol with an average molecular weight of 5 kDa (PEG-SH, MW 5 kDa) was from JenKem Technology (Plano, TX, USA). All other chemical reagents were purchased from Sigma Aldrich (St. Louis, MO, USA) unless otherwise noted.
LNP synthesis and characterisation
LNPs were synthesised by directly adding an organic phase containing the lipids to an aqueous phase containing mRNA in 1.5 mL microcentrifuge tubes. To prepare the organic phase, a mixture of DLin-MC3 DMA, cholesterol, DMG-PEG2000, and a helper lipid selected from a group consisting of DOPE, DSPC, and 18PG were dissolved in ethanol. The mRNA (Cre mRNA, OVA mRNA, Trp2 mRNA, or Gp100 mRNA) was dissolved in 25 mM magnesium acetate buffer (pH 4.0). For larger-scale LNP production, the aqueous and ethanol phases prepared were mixed at a 3:1 ratio in a flash complexation (FNC) device using syringe pumps and purified by dialysis against DI water using a 100 kDa MWCO cassette at 4 °C for 24 h and were stored at 4 °C before injection. The size, polydispersity index, and zeta potentials of LNPs were measured using dynamic light scattering (ZetaPALS, Brookhaven Instruments). Diameters are reported as the intensity average.
Animals and primary cells
All animal procedures were performed under an animal protocol approved by the Johns Hopkins Institutional Animal Care and Use Committee (protocol #MO22E117). Male and female C57BL/6 mice, 6–8 weeks of age, were purchased from the Jackson Laboratory. Male Ai9 mice, 6–8 weeks of age, were bred in Johns Hopkins Animal Facilities and randomly grouped. The mice were supplied with free access to pelleted feed and water. The pelleted feed generally contained 5% fibre, 20% protein, and 5–10% fat. The mice usually ate 4–5 g of pelleted feed (120 g per kg body weight) and drank 3–5 mL of water (150 mL per kg body weight) per day. The temperature of the mouse rooms was maintained at 18–26 °C (64–79 °F) at 30–70% relative humidity with a minimum of 10 room air changes per hour. Standard shoebox cages with corncob as bedding were used to house the mice. The LNPs or LiNx were given through s.c. (right flank) injection at a predetermined dose per mouse.
Antibodies, cell isolation, and staining for flow cytometry
Antibodies used in this study are FITC, APC, Brilliant Violet 750 anti-mouse CD11c (BioLegend #117306, 117310, 117357); Brilliant Violet 421 anti-mouse CD86 (BioLegend #105032); PE anti-mouse SIINFEKL H-2KB (ThermoFisher # 12574382); FITC, Brilliant Violet 605, Brilliant Violet 421 anti-mouse CD45 (BioLegend # 103108, 103140, 103134); APC anti-mouse CD3 (BioLegend # 100236); FITC, APC, Brilliant Violet 750 anti-mouse CD8 (BioLegend # 100706, 100712, BD Biosciences # 747502); PerCP-Cyanine 5.5 anti-mouse CD4 (BioLegend # 100540); PE anti-mouse IFN-γ (BioLegend # 505808); Brilliant Violet 421 anti-mouse IL-4 (BioLegend # 504120); PE-Cyanine 7 anti-mouse TNFα (BioLegend # 506324); and APC anti-mouse Granzyme B (BioLegend # 396408). All antibodies were diluted at a ratio of 1:100 before use.
For isolation, re-stimulation, and staining of splenocytes, the spleen was removed and minced using a sterile blade and dissociated in 250 μL of digestion medium (45 units µL−1 collagenase I, 25 units µL−1 DNase I and 30 units µL−1 hyaluronidase). The suspension was transferred into a 15 mL tube containing 5–10 mL of digestion medium and then filtered through a 70 μm filter and washed once with PBS. Cells were pelleted at 300 × g for 5 min at 4 °C resuspended in 5 mL of red blood cell lysis buffer (BioLegend), and then incubated on ice for 5 min. Cells were then pelleted at 300 × g for 5 min at 4 °C and washed twice with PBS. Isolated splenocytes were counted using the Cellometer cell counter and ViaStain AOPI staining solution (CS2-0106-25 mL, Nexcelom) and diluted in PBS to be used for restimulation. Splenocytes were re-stimulated in vitro with OVA (InvivoGen Cat. vac-pova) and SIINFEKL peptide (InvivoGen Cat. vac-sin) (10 μg mL−1 OVA and 2 μg mL−1 SIINFEKL) for 12 h. After re-stimulation, cells were collected and centrifuged at 300 × g for 5 min. The cell pellet was washed with staining buffer 3 times and stained with antibodies against surface markers (total volume 100 μL) for 30 min in the dark at 4 °C. The stained cells were washed twice with 1 mL of PBS and then fixed and permeabilized using the fixation/permeabilization solution kit (BD Cat# 555028). Then, cells were stained with anti-IFN-γ or other antibodies against intracellular cytokines. Flow data were acquired on Attune (ThermoFisher) and analysed using FlowJo software.
For isolation and staining of cells from lymph nodes or LiNx, isolated lymph nodes or LiNx were mechanically digested through 70 μm nylon cell strainers to prepare single-cell suspensions. The cell suspension was washed once with PBS via centrifugation (300 × g) for 5 min. Then, the cells were resuspended in 100 µL of staining buffer and stained with antibodies (total volume 100 μL) for 20 min in the dark at 4 °C. The stained cells were washed twice with 1 mL of PBS and resuspended in 300 µL of staining buffer for flow cytometry analysis. Flow data were acquired on Attune (ThermoFisher) and analysed using FlowJo software.
ELISpot assay
Multiscreen filter plates (Millipore-Sigma #S2EM004M99) were coated with antibodies specific for IFN-γ (BD Biosciences #551881) and blocked following the manufacturer’s protocols. Then, 1 × 105 isolated splenocytes were plated per well and stimulated with SIINFEKL peptide (2 μg mL−1 SIINFEKL) for 24 h. All tests were performed in duplicate or triplicate and included assay-positive controls as well as cells from a reference donor with known reactivity. Spots were visualised with mouse IFN-γ detection antibody (BD Biosciences #551881) followed by incubation with Streptavidin-HRP (BD Biosciences #557630) and AEC Substrate (BD Biosciences #551951). Plates were then sent to the SKCCC Immune Monitoring Core for analysis.
Enzyme-linked immunosorbent assay (ELISA)
For antibody detection, groups of C57BL/6 mice were immunised with different vaccines on days 0, 7, and 14. On day 21, 100 μL of blood sample was drawn from the tail vein, and levels of antigen-specific IgG in the serum were measured by ELISA. Flat-bottomed 96-well plates (Nunc) were precoated with OVA protein at a concentration of 2 μg protein per well in 100 mM carbonate buffer (pH 9.6) at 4 °C overnight, which were then blocked with 10% foetal bovine serum (FBS) in PBS-Tween (PBS-T). Sera obtained from immunised animals were diluted 100 times in PBS-T (PBS-0.05% Tween), pH 7.4, and then in a 4-fold serial dilution. The undiluted and diluted serum was added to the wells and incubated at 37 °C for 2 h. Horseradish peroxidase-conjugated goat anti-mouse IgG (Southern Biotech Associates, #1013-05) was used at a dilution of 1:5,000 in PBS-T-10% FBS for labelling. After adding the horseradish peroxidase substrates, optical densities were determined at a wavelength of 450 nm in an ELISA plate reader (Bio-Rad). A sample is considered positive if its absorbance is twice as much as or higher than the absorbance of the negative control.
Immunisation and tumour therapy experiments
Mice aged 6–8 weeks were injected subcutaneously with B16-OVA, MC38-OVA cells (1 × 106 in the prophylactic model and 3 × 105 in the therapeutic model) or 3 × 105 B16F10 melanoma cells into the right flank. In therapeutic studies, vaccinations began when tumour sizes were less than 50 mm3 (on day 4 after tumour inoculation). Animals were immunised by subcutaneous injection of different LNP or LiNx formulations containing OVA mRNA, mTrp2, or m Gp100 as described in the main text. One dosage of LiNx was given, and a total of three doses were given for the LNP group. For combinatorial immunotherapy, at days 6, 13, and 20 and an additional at day 27 for OVA-expressing melanoma after inoculation, some groups were intraperitoneally injected with 100 μg checkpoint inhibitor (α-CTLA-4 mAb). Tumour growth was measured three times a week using a digital caliper and calculated as 0.5 × length × width × width. Mice were euthanized when the tumour volumes reached 2000 mm3, and mice were also euthanized if any single tumour dimension reached 20 mm.
IL-17 depletion study
Depletions of IL-17 were done using α-IL-17A (clone 17F3, BioXCell) at 200 μg i.p. every 3 d. The dosing was initiated at 3 d after the first vaccination and continued every 3 d until day 60. On day 21 post-vaccination, 3 × 105 B16-OVA melanoma cells were injected into the right flank. Tumour growth was measured three times a week using a digital caliper and calculated as 0.5 × length × width × width. Mice were euthanized when the tumour volumes reached 2000 mm3, and mice were also euthanized if any single tumor dimension reached 20 mm.
Subsequent RT-PCR analysis
Real-time polymerase chain reaction (RT-PCR) was carried out using TaqMan™ Array Mouse Immune Response (Applied Biosystems, Cat#4414079). The results were analysed according to the 2 − ΔΔCT method and normalised to the housekeeping gene GAPDH.
nCounter Analysis system (NanoString Technology)
Total RNA was extracted from stimulated spleen cells using an RNA extraction kit (Zymo Research, Cat# R2062). Quality and concentration were evaluated using NanoDrop. Predesigned NanoString nCounter CodeSets targeting mouse immune responses-related genes were employed (NanoString, Cat#115000052). Hybridisation of RNA samples to these CodeSets was performed following the manufacturer’s protocol. Post-hybridisation, samples were processed on the nCounter Analysis System, and data normalisation was conducted using nSolver Analysis Software with reference to housekeeping genes.
CODEX multiplexed imaging
Tumour tissues were collected, embedded in optimal cutting temperature (OCT) compound for CODEX, and immediately frozen. After freezing, explants were sectioned to a thickness of 7 µm using an EprediaTM HM525 NX cryostat and arranged on slides. The tissue arrays were stained with the validated panels of CODEX antibodies and imaged in accordance with a previously established protocol.41 Briefly, this entailed cyclic stripping, annealing, and imaging of fluorescently labelled oligonucleotides complementary to the oligonucleotide conjugated to the antibody. Each array underwent CODEX multiplexed imaging; metadata from each CODEX run can be found in Supplementary Table 2. Raw imaging data were processed using the Akoya PhenoCylcer Fusion 2.2.0 software for image stitching, drift compensation, deconvolution, and cycle concatenation. After the raw imaging data were processed, they were evaluated for specific signals. Any markers that produced an untenable pattern or a low signal-to-noise ratio were excluded from the ensuing analysis. Uploaded images were visualised in ImageJ (https://imagej.net/software/fiji/).
CODEX single-cell segmentation
To obtain quantitative single-cell information, individual cells and extracted single-cell protein expression were segmented. Processed data were segmented using the SPACEc package, which can also be downloaded here (https://github.com/yuqiyuqitan/SPACEc/tree/master).42 SPACEc incorporates Mesmer and Cellpose, which are both deep learning-based segmentation methods.43,44 Mesmer was used for our segmentation, with DAPI (nuclei) along with CD45 and CD90 (surface membrane) as reference channels.
Cell-type analysis
Across the LiNx and PBS-treated tumours, 208,486 cells were identified and classified into 19 cell types and states (197652 cells) with noise excluded based on marker expression. Cell type identification was done following the strategies that were developed.45,46 Briefly, segmented cells of appropriate sizes were selected by gating DAPI-positive cells, followed by Z-normalisation of protein markers used for clustering (some phenotypic markers were not used in the unsupervised clustering). The data were overclustered with Leiden-based clustering with the scanpy Python package. Clusters were assigned a cell type based on average cluster protein expression and location within the image. Impure clusters were split or reclustered with K-means clustering with the sklearn Python package, following mapping back to the original fluorescent images.
Statistics & reproducibility
Statistical analyses were performed using GraphPad Prism 8.0 and Microsoft Excel. Comparisons between two groups were conducted using two-tailed unpaired Student’s t tests. Comparisons among more than two groups were conducted using one-way analysis of variance (ANOVA) followed by Tukey’s multiple comparisons test, with alpha set at 0.05. Survival data were analysed using the log-rank (Mantel–Cox) test, and multiple comparisons were adjusted using the Holm–Šídák method.
No statistical method was used to predetermine sample size. Sample sizes were based on prior studies, expected effect sizes, and resource availability. For in vivo vaccination studies, a minimum of n = 4 mice per group was used to permit statistical evaluation, and tumour growth studies were conducted with n = 7–8 mice per group, as stated in the figure legends. The number of biologically independent replicates is provided in each figure or figure legend.
No data were excluded from the analyses. Animal group allocation was randomised while ensuring consistent baseline characteristics (body weight: 17–28 grams; age: 6–8 weeks). Investigators were blinded to group allocation during tumour experiments and outcome assessment. All attempts at replication were successful, and each key experiment was independently repeated at least twice with consistent results.
Some figures in this manuscript were created using BioRender.com, including Fig. 1a (Created in BioRender. Mao, H. (2025) https://BioRender.com/xqqu4bg), Fig. 4a (Created in BioRender. Mao, H. (2025) https://BioRender.com/orhacbc), Fig. 4e (Created in BioRender. Mao, H. (2025) https://BioRender.com/orhacbc), Fig. 4j (Created in BioRender. Mao, H. (2025) https://BioRender.com/pdelh7y), Fig. 5a (Created in BioRender. Mao, H. (2025) https://BioRender.com/580lwzu), Fig. 6g (Created in BioRender. Mao, H. (2025) https://BioRender.com/po3cpbg) and Fig. 6h (Created in BioRender. Mao, H. (2025) https://BioRender.com/tsl22jd).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analysed during this study are provided in the paper and the Supplementary Information. Source data supporting the findings of this study are available in the accompanying Source Data file. No additional datasets requiring deposition in external repositories were generated. Source data are provided in this paper.
Code availability
No custom code was developed for this study. CODEX image segmentation was performed using the publicly available SPACEc package, which is accessible at https://github.com/yuqiyuqitan/SPACEc. SPACEc incorporates the open-source Mesmer and Cellpose segmentation algorithms.
References
Noor, R. Developmental status of the potential vaccines for the mitigation of the COVID-19 pandemic and a focus on the effectiveness of the Pfizer-BioNTech and Moderna mRNA vaccines. Curr. Clin. Micro Rep. 8, 178–185 (2021).
Han, J. et al. Lipid nanoparticle-based mRNA delivery systems for cancer immunotherapy. Nano Converg. 10, 36 (2023).
Pardi, N. et al. Nucleoside-modified mRNA vaccines induce potent T follicular helper and germinal center B cell responses. J. Exp. Med. 215, 1571–1588 (2018).
Zhu, Y. et al. Screening for lipid nanoparticles that modulate the immune activity of helper T cells towards enhanced antitumour activity. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-023-01131-0 (2023).
Xu, Z. & Fisher, D. E. mRNA melanoma vaccine revolution spurred by the COVID-19 pandemic. Front. Immunol. 14, https://doi.org/10.3389/fimmu.2023.1155728 (2023).
García-Pardo, M. et al. Vaccine therapy in non-small cell lung cancer. Vaccines 10, https://doi.org/10.3390/vaccines10050740 (2022).
Rojas, L. A. et al. Personalized RNA neoantigen vaccines stimulate T cells in pancreatic cancer. Nature 618, 144–150 (2023).
Wang, B., Pei, J., Xu, S., Liu, J. & Yu, J. Recent advances in mRNA cancer vaccines: meeting challenges and embracing opportunities. Front. Immunol. 14, 1246682 (2023).
Mukai, H., Ogawa, K., Kato, N. & Kawakami, S. Recent advances in lipid nanoparticles for delivery of nucleic acid, mRNA, and gene editing-based therapeutics. Drug Metab. Pharmacokinet. 44, 100450 (2022).
Zhu, Y. et al. Multi-step screening of DNA/lipid nanoparticles and co-delivery with siRNA to enhance and prolong gene expression. Nat. Commun. 13, 4282 (2022).
Zhu, Y. et al. Optimization of lipid nanoparticles for gene editing of the liver via intraduodenal delivery. Biomaterials 308, 122559 (2024).
Pacifici, N., Bolandparvaz, A. & Lewis, J. S. Stimuli-responsive biomaterials for vaccines and immunotherapeutic applications. Adv. Therap. 3, 2000129 (2020).
Chen, W. et al. Combination of bacterial-photothermal therapy with an Anti-PD-1 peptide depot for enhanced immunity against advanced cancer. Adv. Funct. Mater. 30, 1906623 (2020).
Pal, S. et al. Extracellular matrix scaffold-assisted tumor vaccines induce tumor regression and long-term immune memory. Adv. Mater. 36, e2309843 (2024).
Hao, H. et al. Immunization against Zika by entrapping live virus in a subcutaneous self-adjuvanting hydrogel. Nat. Biomed. Eng. 7, 928–942 (2023).
Huang, P. et al. Nano-, micro-, and macroscale drug delivery systems for cancer immunotherapy. Acta Biomater. 85, 1–26 (2019).
Kerr, M. D., Johnson, W. T., McBride, D. A., Chumber, A. K. & Shah, N. J. Biodegradable scaffolds for enhancing vaccine delivery. Bioeng. Transl. Med. 8, e10591 (2023).
Chua, C. Y. X., Viswanath, D. I., Huston, D. P. & Grattoni, A. Engineering platforms for localized long-acting immune modulation. J. Allergy Clin. Immunol. 153, 572–575 (2024).
Li, X. et al. The effect of a nanofiber-hydrogel composite on neural tissue repair and regeneration in the contused spinal cord. Biomaterials 245, 119978 (2020).
Yao, Z.-C. et al. Biostimulatory micro-fragmented nanofiber-hydrogel composite improves mesenchymal stem cell delivery and soft tissue remodeling. Small 18, e2202309 (2022).
Li, X. et al. Nanofiber-hydrogel composite–mediated angiogenesis for soft tissue reconstruction. Sci. Transl. Med. 11, eaau6210 (2019).
Chen, W., Zhu, Y., He, J. & Sun, X. Path towards mRNA delivery for cancer immunotherapy from bench to bedside. Theranostics 14, 96–115 (2024).
Li, L. et al. Extracellular vesicles delivered by a nanofiber-hydrogel composite enhance healing in vivo in a model of crohn’s disease perianal fistula. Adv. Healthc. Mater. 14, 2402292 (2024).
Kaczmarek, M. et al. Cancer vaccine therapeutics: limitations and effectiveness—a literature review. Cells 12, 2159 (2023).
Zou, W. & Restifo, N. P. TH17 cells in tumour immunity and immunotherapy. Nat. Rev. Immunol. 10, 248–256 (2010).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 Cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Zhang, J.-P. et al. Increased intratumoral IL-17-producing cells correlate with poor survival in hepatocellular carcinoma patients. J. Hepatol. 50, 980–989 (2009).
Charles, K. A. et al. The tumor-promoting actions of TNF-alpha involve TNFR1 and IL-17 in ovarian cancer in mice and humans. J. Clin. Invest. 119, 3011–3023 (2009).
Dubin, P. J. & Kolls, J. K. Th17 cytokines and mucosal immunity. Immunol. Rev. 226, 160–171 (2008).
Hinrichs, C. S. et al. Type 17 CD8+ T cells display enhanced antitumor immunity. Blood 114, 596–599 (2009).
Kryczek, I. et al. Phenotype, distribution, generation, and functional and clinical relevance of Th17 cells in the human tumor environments. Blood 114, 1141–1149 (2009).
Martin-Orozco, N. et al. T helper 17 cells promote cytotoxic T cell activation in tumor immunity. Immunity 31, 787–798 (2009).
Bronte, V. Th17 and cancer: friends or foes?. Blood 112, 214 (2008).
Hald Albertsen, C. et al. The role of lipid components in lipid nanoparticles for vaccines and gene therapy. Adv. Drug Deliv. Rev. 188, 114416 (2022).
Zhang, L. et al. Effect of mRNA-LNP components of two globally-marketed COVID-19 vaccines on efficacy and stability. NPJ Vaccines 8, 156 (2023).
Li, S. et al. Payload distribution and capacity of mRNA lipid nanoparticles. Nat. Commun. 13, 5561 (2022).
Dilliard, S. A., Cheng, Q. & Siegwart, D. J. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc. Natl. Acad. Sci. USA 118, e2109256118 (2021).
Chen, J. et al. Combinatorial design of ionizable lipid nanoparticles for muscle-selective mRNA delivery with minimized off-target effects. Proc. Natl. Acad. Sci. 120, e2309472120 (2023).
Li, X. & Gong, J. P. Design principles for strong and tough hydrogels. Nat. Rev. Mater. 9, 380–398 (2024).
Lu, P. et al. Harnessing the potential of hydrogels for advanced therapeutic applications: current achievements and future directions. Sig. Transduct. Target Ther. 9, 1–66 (2024).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Tan, Y. et al. SPACEc: A streamlined, interactive Python workflow for multiplexed image processing and analysis. 2024.06.29.601349 Preprint at https://doi.org/10.1101/2024.06.29.601349 (2024).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Greenwald, N. F. et al. Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning. Nat. Biotechnol. 40, 555–565 (2022).
Hickey, J. W. et al. Organization of the human intestine at single-cell resolution. Nature 619, 572–584 (2023).
Hickey, J. W., Tan, Y., Nolan, G. P. & Goltsev, Y. Strategies for accurate cell type identification in CODEX multiplexed imaging data. Front. Immunol. 12, https://doi.org/10.3389/fimmu.2021.727626 (2021).
Acknowledgements
Funding: This study is partially supported by National Institutes of Health grants U01AI155313 (H.-Q.M.), P41EB028239 (J.P.S. and H.Q.M.), and R01CA293906-01A1 (H.Q.M., J.W.H., and J.P.S.).
Author information
Authors and Affiliations
Contributions
Y.Z. and H.-Q.M. conceived of and designed this study. H.-Q.M., J.P.S. and J.W.H. secured the funding for this study. Y.Z., R.S., J.M., Z.-C.Y., S.L., I.V., J.L., X.L., K.D.G., X.L., D.Y., C.W., J.L.S., B.Y.X.N., Y.M., K.V.B., J.W.H. and J.K. performed the experiments. Y.Z., R.S., Y.D., J.M., I.V., N.K.L., S.L., G.P.H., S.K.R., J.C., J.P.S., J.C.D., B.Y.X.N., Y.M., J.W.H. and H.-Q.M. participated in data analysis and interpretation. The manuscript was written by Y.Z. and H.-Q.M., with revisions by J.W.H., B.Y.X.N., C.W., J.M., J.P.S., J.C.D., L.C., and inputs from all the other authors.
Corresponding author
Ethics declarations
Competing interests
H.-Q.M., Y.Z., J.M., Z.C.Y. and C.W. are co-inventors of a pending patent application covering the LiNx formulation described in this paper, filed in April 2024 through and managed by Johns Hopkins Technology Ventures. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Wen Jiang, Lulu Xue, Helena Florindo, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.”
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, Y., Yao, ZC., Li, S. et al. An mRNA lipid nanoparticle-incorporated nanofiber-hydrogel composite for cancer immunotherapy. Nat Commun 16, 5707 (2025). https://doi.org/10.1038/s41467-025-61299-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61299-8