Abstract
Trichomonas vaginalis, the causative agent of the venereal disease trichomoniasis, infects men and women globally and is associated with serious outcomes during pregnancy, increased risk of HIV-1 infection, and cancers of the human reproductive tract. Species of trichomonad parasitize a range of hosts in addition to humans, including birds, livestock, and pets. Genetic analysis of trichomonads recovered from columbid birds has provided evidence that they undergo frequent host-switching, and that a spillover event from columbids likely gave rise to T. vaginalis in humans. Here we describe a comparative genomics study of seven trichomonad species, generating chromosome-scale reference genomes for T. vaginalis and its avian sister species Trichomonas stableri, and assemblies of five other species that infect birds and mammals. Human-infecting trichomonad lineages have undergone recent and convergent genome size expansions compared to their avian sister species, a result of extensive repeat expansions specifically of multicopy gene families and transposable elements, with genetic drift likely a driver due to relaxed selection. Trichomonads are thought to have independently host-switched twice from birds to mammals/humans. We identify gene functions implicated in the transition, including host tissue adherence and phagocytosis, extracellular vesicle formation, and CAZyme virulence factors, which are all associated with pathogenesis phenotypes.
Similar content being viewed by others
Introduction
Trichomoniasis, the most prevalent non-viral venereal disease of humans1, is caused by the protozoan Trichomonas vaginalis that infects the lower genital tract of men (urethra and prostate) and women (vulva, vagina, and cervix). Symptoms include foul-smelling vaginal discharge and genital itching, and infections are associated with an increased risk of cervical and prostate cancer, HIV-1 infection, and complications during pregnancy2. Other human-infecting trichomonads include the oral parasite Trichomonas tenax associated with periodontal disease3, and the intestinal parasite Pentatrichomonas hominis associated with gastrointestinal distress and diarrhea4. Trichomonad species also infect a wide range of vertebrate hosts, including birds, livestock, and pets. Trichomonas gallinae5 infects the upper gastrointestinal (GI) tract of birds, including doves, pigeons, songbirds, and raptors that prey on infected birds, and is responsible for a large decline of greenfinch and chaffinch populations in Great Britain in the early 2000s6. In 2008, novel parasites with highly genetically similar markers to T. vaginalis (dubbed “T. vaginalis-like”) were reported in white-winged doves and mourning doves from Arizona and Texas, and in Pacific coast band-tailed pigeons from California7. T. vaginalis-like parasites recovered from the latter during a 2011–12 outbreak were analyzed in detail and given the species name Trichomonas stableri8. T. vaginalis may have originated as a zoonosis from American pigeons and doves during a spillover event following human colonization of the Americas9. It has been hypothesized that its ancestor moved from the upper GI tract of columbids into the human reproductive tract via barrier contraceptives or more commonly through human contact with bird-infected water9.
A draft T. vaginalis whole genome sequence generated using Sanger sequencing in 200710 revealed a highly repetitive genome comprised of thousands of highly similar transposable elements (TEs) and many multicopy gene families. The ~180 Mb genome size was unexpectedly large compared to other human mucosal parasite genomes e.g., Giardia (~11 Mb) and Entamoeba (~21 Mb)11. Extreme fragmentation of the assembly (> 64,000 scaffolds and contigs) precluded accurate counting of repetitive elements. However the sequence generated insights into (1) multicopy gene families involved in the parasite’s active endocytic and phagocytic life-style such as protein kinases and peptidases; (2) surface proteins including the highly diverse BspA-like proteins that likely mediate parasite adherence to vaginal epithelial cells required to establish and maintain an infection in the reproductive tract; and (3) novel metabolic pathways shaped by putative prokaryote-to-eukaryote lateral gene transfer (LGT) events. Other trichomonad genomes remained largely unsequenced, although in the interim, molecular phylogenies showed avian trichomonads to be the closest known relatives of T. vaginalis and T. tenax7,12, with columbids inferred to be the ancestral host of the genus, and the source of at least two independent host switches to mammals/humans9. Host switching has been posited to be a strong macroevolutionary force in genus Trichomonas9.
To expand the number of available trichomonad genome sequences, address key knowledge gaps in the evolution of the parasite, and identify genes implicated in the spillover event from avian to human host, we leveraged long-read and chromosomal conformation sequencing to generate chromosome-scale reference genome assemblies for T. vaginalis G3 and its sister species the avian parasite T. stableri. We used short-read sequencing to also assemble draft genomes of two other human-infecting species, T. tenax and P. hominis, and three other bird-infecting species. These seven assembled genomes represent an extensive whole genome sequence dataset of trichomonads, and enabled unique comparative genomics, including estimates of gene and TE content in closely-related species from different hosts, and visualization of synteny. We offer insights into trichomonad evolution, including evidence for relaxed selection accompanying the inferred host switch from birds in two human-infective Trichomonas lineages, which likely explains the striking genome size variation among these trichomonads. Finally, we identify convergently evolving genes in human-infecting species that were putatively involved in the transition from bird to human host.
Results
Comprehensive TE annotation of a new T. vaginalis chromosome-scale assembly
We generated a chromosome-scale reference assembly of T. vaginalis strain G3 using Pacific Bioscience long-read sequencing augmented with chromosome conformation capture (‘PacBio/Hi-C’). The assembly comprises six chromosome-scale scaffolds matching the published T. vaginalis karyotype number13 ranging from 20 to 40 Mb and ~177 Mb total length (in contrast to the 200710 Sanger assembly of >64,000 scaffolds [range 0.2–585 kb] and 176 Mb total length). Microsatellite and rRNA loci localized to metaphase chromosome squashes by FISH10,14 were mapped to the assembly to assign chromosome numbers I–VI to the scaffolds (Fig. 1). We improved the accuracy of the T. vaginalis predicted proteome, identifying 37,794 protein-coding genes (Table 1), with 46% annotated as ‘hypothetical’ (compared to the 59,681 genes with 75% hypotheticals predicted in 200710). Improved annotation of the 16 major multicopy gene families, many of which are associated with cell surface activity, parasite-host interactions, and the degradome, increased their copy numbers, more than quadrupling it in the case of cysteine peptidase Clan CA, family C1 (Supplementary Table 1). The >600 rDNA genes identified in 2007 collapsed to eleven 28S/5.8S/18S rRNA cassettes tandemly arrayed on chromosome II, agreeing with FISH results10 (Fig. 1). We extended a previously reported15 block of genes laterally transferred from a relative of the firmicute bacterium Peptoniphilus harei, from 37 Kb containing 27 genes to 47 Kb containing 45 genes (Supplementary Table 2). The T. vaginalis genome remains densely packed with protein-coding genes and TEs, with an average length of 1131 bp (median length 520 bp) between them.
The concentric rings, from innermost to outermost, represent: (1) chromosome size in Mb; (2) gene density (green plot) shown in 20 Kb windows; vertical blue lines represent 11 rRNA cassettes, and the vertical black line represents the 47.5 Kb block from an LGT event of the bacterium Peptoniphilus harei; (3) TE density (pink plot) shown in 20 Kb windows; (4) transcript abundance (brown plot) of all genes shown as transcripts per million (TPM) in 100 Kb windows; (5) TE transcript abundance (orange plot) of annotated TE genes shown as TPM in 100 Kb windows; and (6) dN/dS values (grey dots). The axes are shown next to chromosome I.
TE sequences, difficult to identify and incompletely classified in the previous draft assembly, were meticulously annotated and found to dominate the new T. vaginalis reference genome, making up at least 46% of its length (Table 1 and Supplementary Fig. 1). While MULE TEs dominate by abundance (7322), more than 4700 Maverick (TvMav) TEs, long (~10–28 Kb) virus-like DNA transposons found in all major eukaryotic lineages except plants and mammals16, comprise >80% of the total TE length and ~40% of genome length (Table 2). We undertook extensive manual curation of TvMavs since they can contain as many as 19 TE genes, lack terminal inverted repeats (TIR), are concatenated or nested within each other, and can envelop other types of TEs16. Based on length, TIR sequence, gene repertoire, and gene order of 2788 well-defined TvMavs, we identified three classes. Class 1 (n = 902) and Class 2 (n = 181) range from 20 to 25 Kb and differ mainly in TIR sequence. The abundant and previously undescribed Class 3 (n = 1705) has a bimodal length distribution, suggesting two subclasses with peaks at 10–20 Kb and 23–26 Kb (Supplementary Fig. 2). Class 3 also has a distinct gene repertoire and order (Supplementary Tables 3–5).
Comparative genomics of trichomonads infecting humans, birds, and mammals
We chose several species within genus Trichomonas known to be closely related to T. vaginalis on the basis of single-copy gene phylogenies12 for comparative evolutionary studies, including a more distantly related trichomonad species as an evolutionary outgroup. Growing parasites in vitro proved challenging and several could not be grown continuously or in sufficient volume to generate the required quantity or quality of DNA for long-read sequencing. The final list of assembled species and their sequencing statistics is shown in Table 1: (1) the New World clade bird parasite T. stableri strain BTPI-3, the closest known relative of T. vaginalis sequenced using PacBio/HiC; (2) an Australasian bird parasite Trichomonas species genotype 1c (T. sp. 1c)9; (3) the Old World human parasite T. tenax Hs-4:NIH, (4) an Old World bird parasite Trichomonas species genotype 2a (T. sp. 2a), the closest known relative of T. tenax9, (5) the Old World bird parasite Trichomonas gallinae (TGAL)9; and (6) the human/mammal parasite P. hominis (Hs-3:NIH), used as an outgroup for our analyses. Genome size estimates calculated from short reads of the species ranged from 68.9 Mb for T. gallinae to 184.2 Mb for T. vaginalis (Table 1), the latter by far the largest genome size of the seven trichomonad species sequenced. The estimated genome size is larger for human-infecting species than bird-infecting species. It exhibits a linear relationship to estimated repeat content (multicopy genes, TE sequences, and unclassified repeats), which ranges from 21.4% in T. sp. 2a to 68.6% in T. vaginalis (Supplementary Fig. 3). Estimated repeat contents of bird-infecting species (21%–37%) are far lower than those of human-infecting species (51%–69%). Counts of predicted protein-coding genes in the assemblies ranged from 23,689 (T. sp. 1c) to 37,794 (T. vaginalis) and did not display associations with genome size or host type (Table 1).
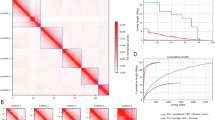
Pairwise whole genome DNA alignments of the species confirmed several previously proposed relationships7,8,9, including the presence of two lineages that exhibit close ‘sister species’ relationships between a human-infecting species and a bird-infecting species (human T. vaginalis with bird T. stableri; and human T. tenax with bird T. sp. 2a) (Supplementary Fig. 4). Whole chromosome synteny mapping of T. vaginalis with its avian sister species T. stableri showed large differences in chromosome sizes and massive genome rearrangements (Fig. 2).
Each of the six T. vaginalis chromosomes I–VI are colored uniquely, and synteny blocks between the two species are indicated by ribbons connecting the chromosomes. Chromosomes are not shown in numbered order for visualization purposes. Blue graph plots show normalized TE density (genomic sequence classified as containing TEs) in 100 Kb windows on the top and bottom of each species’ chromosomes. Hi-C interaction maps are shown as red triangles above (T. vaginalis) and below (T. stableri) the TE density plots, with lengths of contigs from the assemblies shown as white squares for T. vaginalis and green squares for T. stableri.
We identified 24,465 orthogroups (groups of evolutionarily related genes) across the seven trichomonad species, with 93.8% of all genes being assigned to an orthogroup. Of these orthogroups, 10,457 contain genes from all species (Fig. 3A), 6226 contain only single-copy genes, and 2798 orthogroups, comprising 6.6% of all genes, are species-specific. As expected, the outgroup P. hominis contained the largest number of species-specific orthogroups (1078), followed by T. vaginalis (425). We used the 6226 single-copy orthologs to infer a phylogenetic species tree (Supplementary Fig. 5). The tree strongly supports separate clades for T. vaginalis and T. tenax, in accordance with the proposal of at least two bird-to-human host switches in the evolutionary history of genus Trichomonas9. It also resolves the formerly ambiguous placement, from single-gene trees, of Australasian bird parasite T. sp. 1c among the Old or New World clades9; we find strong support for placing T. sp. 1c with the New World clade.
A An Upset plot displaying the intersection of orthogroups identified in OrthoFinder across seven trichomonad species. Each vertical bar represents the number of orthogroups shared at each species intersection, the set size indicates the number of orthogroups found in each species, and the connected dots represent the species in the intersection. B Ultrametric tree from 6226 concatenated single-copy genes. Black dots at terminal nodes are proportional to estimated genome size, and hosts are denoted by cartoons. Estimated gene family expansions/contractions from 12,345 genes are denoted as + or − values on the tree. The heat map shows the log10 transformed count of TE family members for each tree branch.
The burden of repeats/TEs in trichomonads differs by host type
The correlations between genome size, host type, and repeat content noted above are not necessarily reflected in phylogenetic proximity; for example, the human-infecting trichomonad lineages (T. vaginalis, T. tenax, P. hominis), while not closely related by evolution, all appear to have undergone recent and convergent large genome size expansions compared to their avian sister species (Fig. 3B). Kmer-based estimates of genome size and repeat content from sequencing reads clearly mark the three human-infecting species as having larger and more repetitive genomes than the bird-infecting species (Table 1). This is borne out in a comparison of the two long-read assemblies, whose lengths concur with kmer-based estimates, and whose counts of repeat sequences are the most reliable: the major contributor to the much larger genome size of T. vaginalis versus T. stableri is increased repeat content, particularly expansion of TEs (Supplementary Fig. 1).
We identified and classified 22,449 TEs in T. vaginalis, 3443 in T. stableri, and, within the limits imposed by short-read assembly, 897 in P. hominis, 459 in T. tenax, and <300 in each of the bird-infecting species (Tables 1 and 2), again showing a human/bird host disparity. The great majority of TEs identified in all species are Class II DNA transposons, with a single Class I NeSL retrotransposon family identified as particularly abundant in T. tenax (Fig. 3B). Mavericks appear to be more abundant in the three human-infecting species T. vaginalis, T. tenax, and P. hominis than the bird species, since their size often makes them the dominant TE class by length, even when it is not the most abundant class (Supplementary Fig. 6), but no pattern of abundance was seen in other TE classes.
A closer inspection of the synteny between T. vaginalis with its sister species T. stableri in birds (Fig. 2) revealed the syntenic regions in T. vaginalis to be made up of almost equal numbers of TEs (47.3%) and non-TE (52.7%) protein-coding genes, whereas in T. stableri the regions are made up of 90.22% non-TE protein-coding genes (Supplementary Table 6). Analysis of the T. vaginalis protein-coding genes that are not TEs in the syntenic regions revealed many of them to be members of multicopy gene families enriched in gene ontology (GO) functions such as protein kinases, ATP/GTP binding, and protein phosphorylation. Copy numbers of several gene families are markedly higher in T. vaginalis than T. stableri, e.g., the BspA-like (73% higher), Saposin-like (SAPLIP) (65% higher), and leishmanolysin-like proteinase (64% higher) families (Supplementary Table 7), signifying that these expansions were favored in the human host. Most of the remaining gene families, e.g., membrane trafficking proteins, serine peptidases, protein kinases, vary <10% in copy number between the two species, suggesting their gene duplications largely predate the bird-human host switch.
Relaxed selection supports a neutral model for genome expansion in human-infecting trichomonads
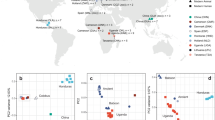
To assess levels of genetic drift (a nonadaptive possible driver of expansion of repetitive DNA when selection is relaxed) we used the hypothesis-testing framework RELAX17, which asks whether the strength of natural selection has been relaxed or “intensified’ (i.e., inferred to have undergone either purifying or positive selection) along specified test branches compared to reference branches in a phylogenetic tree. We used the 6226 single-copy orthologs occurring in all seven species as a proxy for genome-wide sampling of drift. With human-infecting branches as test (foreground) and avian branches as reference (background), we determined which genes evinced significant (p < =0.05) relaxed or intensified selection and found that human-infective branches have more genes under relaxed than intensified selection (n = 894 vs. n = 494) (Fig. 4A and Supplementary Data File 1), the converse of the bird-infecting branches.
A Graphs showing count of all single-copy orthologs (SCOs; left panel) and BUSCO genes (right panel) identified by RELAX as being under relaxed, neutral, or intensified selection in the species with expanded genomes (T. vaginalis and T. tenax) for a range of P-values. B Left panel: mean dN/dS values (plotted from 0.0 to 1.0) for SCOs under relaxed purifying selection for bird-infecting species (n = 141) and mammal-infecting species (n = 31), and right panel: dN/dS values (plotted from 1.0 to 100.0) for SCOs under relaxed positive selection for bird-infecting species (n = 401) and mammal-infecting species (n = 104). Boxplots represent the interquartile range (IQR) with the median as a horizontal line and whiskers extending to 1.5 × IQR. For relaxed purifying selection, the mammal group had a Q1 of 0.38, median of 0.52, Q3 of 0.68, and whiskers from 0.09 to 0.98; the bird group had a Q1 of 0.38, median of 0.56, Q3 of 0.83, and whiskers from 0.08 to 1.00. For relaxed positive selection, the mammal group had a Q1 of 2.36, median of 5.79, Q3 of 33.92, and whiskers from 1.02 to 80.48; the bird group had a Q1 of 2.40, median of 7.32, Q3 of 31.15, and whiskers from 1.03 to 73.13.
A gene under relaxed selection may result from an organism switching environments if the gene is obsolete in the new host or tissue, or from increased genome-wide genetic drift (due to changes in parameters such as population size and mode of reproduction)18. To rule out host environment as the driver of observed relaxed selection, we used RELAX to test the strength of selection acting on 506 genes from the seven genomes with homology to BUSCO19 genes, since the rates of evolution of conserved genes are expected to remain constant even in different environments. We found that there are more genes under relaxed selection (n = 47) than intensified selection (n = 44) in the human-infecting species relative to bird-infecting species (Fig. 4A). This suggests a role for increased genome-wide genetic drift, rather than relaxed selection targeting genes that are superfluous in the new environment. Consistent with relaxed positive selection, the distribution of average dN/dS ratios (a measure of the strength and mode of natural selection acting on protein-coding genes) for the single-copy orthologs shows a higher median in avian-infecting parasites (7.318) and lower median in human-infecting parasites (5.786). We observed a higher median in avian-infecting parasites (0.585) and lower median in human-infecting parasites (0.542) for relaxed purifying selection (Fig. 4B). In general, therefore, dN/dS in human-infecting trichomonad species has contracted towards 1, i.e., neutral evolution.
T. vaginalis has the largest net gain of expanded multicopy gene families
We previously proposed that copy number expansions in T. vaginalis multigene families may account for a significant proportion of its unexpectedly large genome size compared to other parasites10. We investigated this further by analyzing T. vaginalis gene families in the context of our other assembled trichomonad genomes. We used CAFE520, which implements a birth-death model for evolutionary inferences about gene family evolution, to identify multicopy gene families that have expanded or contracted significantly across our trichomonad phylogeny. Of the 26,244 orthogroups, 12,345 (see “Methods”) were analyzed for expansions or contractions, of which 3853 showed significant expansions or contractions in at least one extant species or inferred ancestor (Fig. 3B and Supplementary Data File 1). We found that among the trichomonad species examined, T. vaginalis had the largest net gain (n = 116) in number of expanded gene families, consistent with it having undergone the largest genome size increase. The 140 expanded T. vaginalis gene families are functionally enriched in GO terms for transmembrane transport (e.g., ABC transporters), metabolism and translation (Fig. 5). We also identified many expanded gene families (n = 61) in T. vaginalis that have published functions related to parasite pathology, such as host cell adherence21,22,23,24, phagocytosis25, and extracellular vesicles26,27. We did not find functional enrichment in the 24 multicopy gene families in T. vaginalis that have significantly contracted.
Dot size represents the number of genes with a specific GO term. Biological process (BP), cellular component (CC), and molecular function (MF) are plotted. Only significant GO enrichments after FDR correction (0.05 < ) are reported.
We found that T. vaginalis shares the largest number of expanded gene families not with its bird-infecting sister species T. stableri, but with human/mammal-infecting T. tenax (n = 33) and its sister species, bird-infecting T. sp. 2a (n = 35) (Fig. 6). These convergently expanded multicopy gene families of T. tenax and T. vaginalis are enriched for GO terms in metabolism, and include 25 genes previously reported to be associated with cell adherence21,22,28, microvesicles26,29, and putative virulence factors30. T. vaginalis shares the largest number of expanded gene families with T. sp. 2a and consists of GO terms enriched in many biological processes such as transport, telomere maintenance, signal transduction, morphogenesis, and immune response. We similarly found genes with published associations with adherence and microvesicles, but to a lesser degree (n = 13). We also identified 15 convergently contracted multicopy gene families in T. tenax and T. vaginalis species (Supplementary Fig. 7), but without any GO term enrichment.
Heatmap showing number of shared expanded orthogroups between seven trichomonad species. T. vaginalis shares its largest number of expanded orthogroups with T. tenax (n = 33) and T. sp. 2a (n = 35), two non-sister species that infect humans and birds, respectively. B GO enrichment of convergently expanded multicopy gene families in T. vaginalis and T. tenax. C GO enrichment of convergently expanded multicopy gene families in T. vaginalis and T. sp. 2a. For both (B and C): size of dot represents the number of genes with a specific GO term; biological process (BP), cellular component (CC) and molecular function (MF) are plotted; only significant GO enrichments after FDR correction (0.05 < ) are reported.
Trichomonas genes under positive selection and involved in bird-to-mammal/human host switch
We used a branch-site model implemented in aBRASEL31 to test the 6226 single-copy orthologs across our trichomonad species for evidence of positive selection (Supplementary Fig. 8 and Supplementary Data File 1). Approximately 18% of the 1201 genes with evidence of positive selection were shared between two or more species, the rest being specific to a single species. The shared genes were enriched in GO terms for translation, intracellular transport, and cytoskeleton/motility, most likely reflecting functions essential to trichomonads generally (Supplementary Fig. 9). A relatively large number of these shared genes have been previously associated with phagocytosis25 (n = 44) and include proteases, cytoskeleton genes, transmembrane and transporter genes, vesicular trafficking, and metabolism-related genes, and a similar number were associated with microvesicles26 (n = 40), including a number of tRNA synthetases and peptidases, regulatory and binding proteins. Smaller numbers of shared genes were associated with adherence21,23,28,32 (n = 21) and included transporters and membrane proteins; exosomes29 (n = 9), including one core exosomal protein; and proteins of the secretome27 (n = 4), and carbohydrate-active enzymes (CAZymes, n = 1) implicated as virulence factors30.
A total of 138 of the 1201 genes with evidence of positive selection were found in T. vaginalis, and 69 of them are unique to the T. vaginalis lineage. While no GO terms were found to be enriched among them, ten genes (TVAGG3_0302500, TVAGG3_1001150, TVAGG3_1088290, TVAG_005750, TVAG_062520, TVAG_117090, TVAG_152520, TVAG_313880, TVAG_437950, TVAG_453350; Supplementary Data File 1) are specific to T. vaginalis and have experimentally verified functions associated with adherence21,22,24, microvesicles26, the secretome27, phagocytosis25, and CAZymes30. T. tenax shows 45 genes with evidence of positive selection, 26 of which are unique to the lineage. We did not find GO enrichment among these 45 genes. However, six genes (TVAG_097660, TVAG_127300, TVAG_137880, TVAG_237760, TVAG_270770, TVAG_459530) under positive selection and shared between other trichomonad species have been associated with adherence23, microvesicles26, exosomes29, and phagocytosis25; all of these genes are shared with T. vaginalis.
Assuming that trichomonads have independently host-switched twice, from birds to humans to generate the T. vaginalis lineage, and from birds to mammals and humans to generate the T. tenax lineage9, and that selection will act on similar genes when different lineages independently adapt to similar environments, we applied the convergent evolution model RERconverge33 to identify single-copy genes putatively involved in the transition to a mammalian/human host (Supplementary Data File 1). Of 6226 single-copy orthologs, 320 showed evidence of convergent purifying evolution in the human-infecting branches T. vaginalis and T. tenax. Several of these genes are reported to be associated with phenotypes of phagocytosis25 and adherence21,28, as well as microvesicle26 and exosome29 structures, and CAZymes30. A total of 93 single-copy orthologs showed evidence of convergent positive evolution in T. vaginalis and T. tenax; several of these have been reported to be involved in adherence28, phagocytosis25, and microvesicle-like structures26.
Discussion
We present here a comparative analysis of chromosome-scale genomes of the human sexually transmitted parasite T. vaginalis with its sister species in birds, T. stableri. These are compared with genomes from five other species of human- and bird-infecting trichomonads. These comparisons illuminate differences in protein-coding gene and TE content, genomic architecture, and gene evolution across the trichomonad phylogeny, and identify genes implicated in the inferred spillover event from avian to human host.
All of the trichomonads we sequenced have much larger genomes than other orders of single-celled parasites that cause important human diseases11,34. The major contributor to increased genome size is increased repeat content, in particular TE expansion, which has been proposed to be triggered by major environmental changes35. TEs constitute the bulk of repetitive DNA in T. vaginalis, and likely the other genomes presented here as well. The presence of the same classes of TEs in all of the trichomonad genomes points to either multiple invasions of an ancient common ancestor or multiple invasions and expansions after divergence. For example, the very high sequence similarity of the hundreds of Mariners we recently reported in T. vaginalis points to their recent expansion in that genome36, and high polymorphism in Mariner insertion sites across different T. vaginalis strains also suggests recent active transposition of this TE class in the species36. At least 45% of the T. vaginalis genome length is made up of three classes of an ancient DNA transposon lineage, Maverick DNA transposons (TvMavs). A greater abundance of TvMavs in the human-infective species T. tenax, T. vaginalis, and P. hominis appears to be the main contributor to their genome size increases relative to bird-infecting species. A ‘transposome’ analysis across species is needed to clarify the likely complex evolutionary history of trichomonad TEs, and to extend the previous studies on the TE transcriptional silencing mechanisms elucidated in some T. vaginalis TE families37.
Another constituent of genome repeat content is paralogous gene families. Paralogs originate from gene duplication, and while most duplicates are deleted, gene duplication is the primary origin of new gene functions38. The high copy numbers of T. vaginalis gene families found in the draft genome sequence10 raised the question of whether they persist due to being adaptive or due to other evolutionary mechanisms such as genetic drift. Selection for a paralog with a new function (neofunctionalization) or to maintain gene dosage balance can contribute to long-term preservation of gene duplicates38. And differential expression of paralogs in different environments, e.g., different host species or different host tissues, has been hypothesized to infer adaptive new roles for some gene duplicates. Evidence of this has been reported in T. vaginalis for paralogs in a limited number of multicopy gene families, such as cysteine proteases (see ref. 11), but such evidence exists for only a small fraction of paralogs in T. vaginalis gene families overall. In addition, adaptive processes are also unlikely to explain the high burden of T. vaginalis TEs, which are assumed to be deleterious and of exogenous origin. Alternatively, when selection is relaxed—as when functional constraints on a gene are removed, or effective population size is reduced, both of which can occur when an organism switches host environments—genetic drift comes to the fore, randomly fixing or deleting alleles. Elevated levels of drift are predicted by the ‘mutation hazard hypothesis’39 to increase sequence copy number, including of genes and TEs. Our analyses identified a neutral process of evolution driving the expansion of repeat content in T. vaginalis, congruent with the mutation hazard hypothesis, alongside a subset of candidate multicopy gene families under selection. Future studies should prioritize categorizing the various paralogs within these gene families (as under neofunctionalization, subfunctionalization, dosage balance, etc.) and assessing their functional roles in the parasite.
We previously hypothesized that the genome size expansion of T. vaginalis reflected a relaxation of selection when the parasite underwent a population size bottleneck during its transition from a GI environment to the urogenital tract10. In the present study, we found an overall trend of relaxed selection amongst human-infecting compared to bird-infecting trichomonad species, suggesting higher levels of genetic drift as a factor in their genome expansion. An intriguing question is whether host switch bottlenecks alone account for the relaxation of selection. Peters et al.9, estimated the co-divergence between columbid host and Trichomonas and observed relatively shallow branches in the parasite tree, indicating recent divergence in the parasites but not the hosts. Additionally, T. gallinae, and T. sp. 2a have been identified across bird orders, not just genera9. These observations suggest that recent host shifting, including across fairly large evolutionary distance, is a general phenomenon amongst columbid Trichomonas, and that we would therefore expect bottlenecks (and relaxed selection) in these species as well, if the hypothesis is true. But the relative lack of relaxed selection we observed in columbid Trichomonas overall (Fig. 4) suggests that factors other than bottlenecks contributed to the host-associated difference in selection strength. Mode of parasite reproduction is one such possible contributor. Asexual reproduction can lower the effective population size through decreased genetic variation and global reduction of variation due to background selection and genetic hitchhiking18. The last common ancestor to eukaryotes is thought to have reproduced sexually, and among extant eukaryotes sexual reproduction is generally the norm. We previously accumulated evidence that T. vaginalis may have undergone sexual recombination in its evolutionary past40; a putative hybridization event has also been described in T. gallinae41, raising the possibility that sex occurs in other Trichomonas species. Thus, it could be that a shift from sexual to asexual reproduction, in addition to a bottleneck, accompanied host switching, and facilitated relaxed selection, enabling large-scale structural changes to the genomes. Further investigation of reproduction in genus Trichomonas is needed to confirm this hypothesis.
Among the trichomonads in our study, we found the largest net gain in number of expanded multicopy gene families in T. vaginalis, and the highest number of gene family expansions shared with T. vaginalis in the T. tenax/T. sp. 2a clade, indicating that the latter similarity results from convergent evolution. The set of expanded families shared by T. vaginalis and human-infecting T. tenax is different from that shared by T. vaginalis and bird-infecting T. sp. 2a. Families that expanded in T. vaginalis and T. tenax feature more genes involved specifically in metabolism, cell adherence, microvesicles, and virulence, than those expanded/shared in T. vaginalis and T. sp. 2a, which could be evidence for human- (or at least mammal-) specific adaptations. Indeed, the diverse array of glycoside hydrolases, Carbohydrate Active enZymes (CAZymes), and carbohydrate-binding modules, identified through a recent comparative analysis of T. vaginalis and T. tenax30 are likely shared virulence factors that potentially target host or bacterial glycans, and induce and/or amplify damaging inflammation and bacterial dysbiosis, known to exacerbate periodontitis and vaginitis. The functions associated with the shared T. vaginalis/T. sp. 2a families, on the other hand, could be those useful to parasitic trichomonads with bird-host ancestors. The recent reports of T. tenax in birds3 complicates this hypothesis. However, that report is based upon genotyping of the multicopy ITS1/5.8S/ITS2 rRNA small subunit gene, where discrimination between species can be based on as little as <=1% difference in sequence identity. Moreover, our T. tenax Hs-4:NIH genome sequence is of a strain isolated from a human subject and presumably adapted to that host. T. tenax has also been isolated from a range of birds and mammals such as cats, dogs, and horses3, raising the possibility that the ancestor of T. tenax was transmitted from birds to mammals before jumping to humans. The recently identified Trichomonas brixi42 appears to have undergone a similar bird-to-mammal transmission event, an interesting parallel. More sequence data of T. tenax isolates from humans, other mammals, and birds are needed to clarify this. The columbid upper GI tract and the oral and vaginal cavities of humans are lined with stratified, non-cornified epithelia43,44, a histological similarity that conceivably enabled the ancestral colonization of a human tissue by a bird trichomonad. At the same time, convergent changes in the human-infective species suggest there was enough microscale difference in the host environments to drive adaptation. Convergently evolving multicopy gene families in T. vaginalis and T. tenax included some associated with cell adherence, suggesting specifically that differences in surface membrane proteins in bird versus human mucosal epithelium could foster selection for differential adherence to host tissues.
Multicopy genes are challenging to use in some evolutionary analyses because it is difficult to identify orthologues between them. But evidence from analysis of single-copy gene evolution can illuminate phenomena such as host-switching or spillover. For this analysis we looked at single-copy orthologues two ways: (1) specifically for positive selection, and (2) more generally for rates of evolution, since convergent evolutionary rate shifts can indicate whether changes in selection in a gene cohort are due to purifying selection versus relaxed or positive selection compared to the average rate across the phylogeny. Most of the single-copy genes we found with evidence for positive selection were species-specific, suggesting fine-tuning of the parasite to particular environments. However, the single-copy orthologues in T. vaginalis and T. tenax we identified as displaying convergent purifying or positive evolution were often related to the endo- and cell membrane systems, and also to adherence, phagocytosis, and mitosis. The endomembrane system generates extracellular vesicles, e.g., exosomes and microvesicles, which have recently been identified in T. vaginalis and shown to prime host cells for adherence, modulate the host’s immune response, facilitate cell-to-cell communication, and promote host cell colonization22,24,26,29. Convergent selection for endomembrane system genes could reflect adaptation of the parasite to new host cell surface membranes and immune system; vesicles can carry cargo that affect host gene expression, and the removal of these vesicles from the extracellular milieu reduces the adherence of the parasite to host cells22. Parasite adherence to host mucosal cells is essential in establishing an infection, and parasite phagocytosis is involved in nutrient acquisition45 and immune cell evasion46. Autophagy is associated with the pathogenicity of several protozoan parasites and has been demonstrated to increase the survivability of T. vaginalis under nutrient starvation47 as well as participate in proteolysis48. Both phagocytosis and autophagy also involve the endomembrane system. Peculiarly among eukaryotes, T. vaginalis mitosis can occur during phagocytosis, which has been hypothesized to be advantageous for a parasite in a hostile environment with scarce nutrients49. In conclusion, our results implicate several genes and gene families involved in parasite adherence, phagocytosis, and microvesicle-like structures in the spillover of parasites from the upper GI tract of columbids into the human reproductive tract. This analysis provides candidates for further investigation into trichomonad evolution and adaptation to human hosts.
Methods
Generation of a T. vaginalis chromosome-scale assembly and annotation
DNA was extracted from T. vaginalis strain G3 parasites cultured in modified Diamond’s media and sequenced using Pacific Biosciences Inc. sequencing chemistry on 56 SMRT cells on a PacBio RSII instrument, generating 2,043,705,869 reads that were initially assembled using FALCON50. The initial assembly had a total span of 173 Mb across 1194 contigs with a contig N50 size of 321 Kb. Hi-C library preparation and sequencing were performed as described51, and PBJelly52 run to close any scaffold gaps. In total, this yielded six chromosome-scale scaffolds containing 97.4% of the original assembly with a scaffold N50 size of 27.3 Mb, scaffold N90 size of 20.0 Mb, and improved the contig N50 size to 444 Kb. Pilon53 was used for assembly polishing two times using published G3 Illumina reads (SRA# SRR4734558), Sanger reads10, and RNA-seq reads37 mapped to the assembly using BWA54.
Structural annotation used BRAKER255, STAR56-mapped RNA-seq reads, and a training set of 539 high-confidence T. vaginalis protein sequences. De novo structural annotation was augmented by gene model transfer from the 2007 T. vaginalis assembly (TrichDB release 52), using Liftoff57 with parameters -s 0.9 and -a 0.9. Functional annotation used one of six criteria: (1) identity to proteins with previously experimentally characterized function; (2) identity to proteins previously inferred as horizontally transferred from firmicute bacterium Peptoniphilus harei15; (3) strong similarity (90% identity over 90% length) to UniProtKB/Swissprot entries; (4) orthology group membership and function using eggnog-mapper58 and the eggNOG database of orthology groups59; (5) protein domains returned by Interproscan (v 5.52.86)60; (6) DeepFRI function prediction from predicted protein structure61; and for the remainder (6) DeepGOPlus62 version 1.0.20 function prediction. Proteins that could not be assigned a function by these means were called ‘conserved hypothetical’. GO enrichment analysis was undertaken using the hypergeometric distribution incorporated into an inhouse Python script.
Maverick TEs were identified by BLAST using ORFs from 11 ‘canonical’ Mavericks identified previously16. Ordered blocks of Maverick ORFs were marked in the polished assembly. Canonical and novel TIRs flanking the blocs were identified with BLAST and Inverted Repeats Finder63. Other TE families were identified through BLASTn queries using consensus TE sequences from Repbase64, GyDB65, and a custom database of previously identified TEs (Supplementary Data File 2); RepeatModeler266 was used to identify novel potential TEs. We used phylogenetic analysis, motif identification, and Interproscan to validate the classification of RepeatModeler TE consensuses, and einverted (EMBOSS67) and GenericRepeatFinder68 to annotate TIRs and target site duplications followed by manual inspection of a multiple sequence alignment of the TE family.
Sequencing, assembly, and annotation of six additional trichomonad species
T. gallinae strain TGAL (see Supplementary Methods), Trichomonas species genotype 1c, Trichomonas species genotype 2a, T. tenax strain Hs-4:NIH (ATCC 30207), P. hominis strain Hs-3:NIH (ATCC 30000), T. stableri strains CA015840 (ATCC PRA-430) and BTPI-3 (ATCC PRA-412), and two strains of T. vaginalis CDC 085 (ATCC 50143) and NYH 286 (ATCC 50148) were grown axenically in vitro under standard conditions, DNA extracted, libraries generated, and sequenced on an Illumina HiSeq platform. Barcode sequences were trimmed using the fastx toolkit (https://github.com/agordon/fastx_toolkit), sequencing errors were corrected using Quake69 and then assembled using SOAPdenovo270, yielding contig N50 sizes of 10 Kb to 25 Kb (Table 1). Genome sizes were estimated from Illumina reads using GenomeScope71. T. stableri strains BTPI-3 and CA015840 were sequenced using Pacific Biosciences Sequel II SMRT technology, assembled using hierarchical genome-assembly HGAP (Pacific Biosciences, SMRT Link V11.1) and Canu72, and the resulting assemblies scaffolded using Hi-C data51. RNA-seq data were generated in triplicate for T. stableri strains BTPI-3 and CA015840, using total RNA extracted from three biological replicate cultures for each strain, stranded mRNA preparation, and the resulting libraries run in HighOutput mode on a NextSeq 500 sequencer to produce 2 × 75 bp paired-end reads. TE expression was estimated in T. stableri as for T. vaginalis G3 above.
De novo gene finding and annotation of the remaining five assemblies was performed using AUGUSTUS73 with ab initio training using the standard translation code. RNA-seq data were used for transcript assembly and annotation, where available for each species, and annotation was manually curated when possible. For annotation of TEs, we used BLASTn74 with conserved sequence motifs of all TE consensus sequences identified in T. stableri and T. vaginalis as queries. For the other species, their lower assembly quality precluded annotation of TE sequences. To quantify them, TE queries based on T. vaginalis/T. stableri consensus sequences were used in BLASTn to find matches in each species, which were used to generate per-species consensus sequences. Raw reads were mapped to these consensus sequences using deviaTE75, to estimate the true insertion frequency of each TE family except Mavericks, where raw reads mapping to the integrase ORF were used to estimate TE frequency.
Comparative genomics
DNA sequence similarity across the seven Trichomonas species at the whole genome level was calculated using MUMmer v.3.23 dnadiff algorithm76 using default parameters, and Bray-Curtis dissimilarity statistics. Synteny analysis was determined using MCScanX77 using default parameters (Maximum Evalue: 1e-10, Num. of BlastHits: 5 [minimum collinearity length]), which identified collinear blocks consists of ≥ five genes conserved between the two species. OrthoFinder78 was used to identify 6,226 single-copy orthologs (SCOs) across the seven trichomonad species, and GO terms assigned to them using embedding similarity79. Other analyses used custom in-house Python Scripts and packages in R, such as UpSetR in version 1.3.3; 2017.
Phylogenetic and evolutionary analyses
A species tree of the seven trichomonad species was generated from 6226 genes present in one copy in each genome (‘single-copy orthologues’). Orthologues were aligned using PRANK80 with default parameters and concatenated to generate a supergene matrix for phylogenetic inference with Phangorn81, estimating the best evolution model as GTR + G + I using AICc and executing 1000 bootstraps for analysis. To test if expanded genomes experienced genome-wide relaxed selection, we used RELAX17 on the 6226 single-copy orthologs. We tested human-infecting Trichomonas species (T. vaginalis and T. tenax) with the four avian-infecting species set as background, and the outgroup P. hominis excluded. We tested the avian sister species (T. stableri and T. sp. genotype 2a) of T. vaginalis and T. tenax against all Trichomonas species (i.e., excluding P. hominis). BUSCO19 was used to identify single-copy orthologs that are near-universal across eukaryotes. Significant genes in RELAX results were searched against a curated database of published papers associated with specific phenotypes such as virulence (Supplementary Data File 3). CAFE520 was used to implement a birth-death model for evolutionary inferences about gene family evolution. A total of 12,345 of 26,244 orthogroups were tested that met the CAFE requirement that each orthogroup include the outgroup species P. hominis. The R package RERConverge33 was used to test for association between relative evolutionary rates of genes and the evolution of traits across the phylogeny. We performed the association by designating human-infecting Trichomonas species (excluding P. hominis) as the foreground against all bird-infecting species for all 6226 single-copy genes. We assessed significance using permutations, phylogenetic simulations and trait permutation, with RERconverge. This enabled the generation of a list of candidate genes associated with host type. aBRASEL31 was used to test for positive selection in all 6226 single-copy genes across all Trichomonas species using default parameters without a priori selection of foreground branches.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data supporting the findings of this work are available within the paper and its Supplementary Information files. The raw sequence data have been deposited in GenBank under the following accession numbers: T. vaginalis strain G3: Bio-Project PRJNA885811, Genome Accession SAMN31107788, RNA-Seq SUB12510628; T. vaginalis strain CDC 085: SRA SRX1017343, SRX1017342; T. vaginalis strain NYH 286: SRA SRX1017345, SRX1017344; T. stableri strain BTPI-3: Bio-Project PRJNA816543, Genome SUB11194301, RNA-seq SUB12511058; T. stableri strain CA015840: Bio-Project PRJNA828130, Genome SUB11350628, RNA-seq SUB12511233, Illumina DNA-seq reads SRA SRR30350957; T. gallinae strain TGAL: Bio-Project PRJNA885811, Genome SUB14002162; Trichomonas species genotype 1c: Bio-Project PRJNA885811, Genome: SUB14002711; Trichomonas species genotype 2a: Bio-Project PRJNA885811, Genome SUB14002751; T. tenax strain Hs-4:NIH: Bio-Project PRJNA885811, Genome: SUB14002737; P. hominis strain Hs-3:NIH: Bio-Project PRJNA885811, Genome SUB14002786. The source data underlying the figures and tables are provided as a Source Data File. Icons used in figures and tables were from Adobe Stock and Microsoft Stock which are copyright-free. Source data are provided with this paper.
Code availability
Code for T. vaginalis GO enrichment is available at https://github.com/biofcallejas/TriGO and is permanently referenced with the link https://doi.org/10.5281/zenodo.15002692.
References
Van Gerwen, O. T. & Muzny, C. A. Recent advances in the epidemiology, diagnosis, and management of Trichomonas vaginalis infection. F1000Res. 8, F1000 (2019).
Van Gerwen, O. T., Muzny, C. A. & Marrazzo, J. M. Sexually transmitted infections and female reproductive health. Nat. Microbiol. 7, 1116–1126 (2022).
Matthew, M. A., Yang, N., Ketzis, J., Mukaratirwa, S. & Yao, C. Trichomonas tenax: a neglected protozoan infection in the oral cavities of humans and dogs-A scoping review. Trop. Med. Infect. Dis. 8, 60 (2023).
Zhang, N. et al. High prevalence of Pentatrichomonas hominis infection in gastrointestinal cancer patients. Parasit. Vectors 12, 423 (2019).
Stabler, R. M. Trichomonas gallinae: a review. Exp. Parasitol. 3, 368–402 (1954).
Lawson, B. et al. A clonal strain of Trichomonas gallinae is the aetiologic agent of an emerging avian epidemic disease. Infect. Genet. Evol. 11, 1638–1645 (2011).
Gerhold, R. W. et al. Molecular characterization of the Trichomonas gallinae morphologic complex in the United States. J. Parasitol. 94, 1335–1341 (2008).
Girard, Y. A. et al. Trichomonas stableri n. sp., an agent of trichomonosis in Pacific Coast band-tailed pigeons (Patagioenas fasciata monilis). Int. J. Parasitol. Parasites Wildl. 3, 32–40 (2014).
Peters, A., Das, S. & Raidal, S. R. Diverse Trichomonas lineages in Australasian pigeons and doves support a columbid origin for the genus Trichomonas. Mol. Phylogenet. Evol. 143, 106674 (2020).
Carlton, J. M. et al. Draft genome sequence of the sexually transmitted pathogen Trichomonas vaginalis. Science 315, 207–212 (2007).
Barratt, J., Gough, R., Stark, D. & Ellis, J. Bulky trichomonad genomes: encoding a swiss army knife. Trends Parasitol. 32, 783–797 (2016).
Malik, S. B. et al. Phylogeny of parasitic parabasalia and free-living relatives inferred from conventional markers vs. Rpb1, a single-copy gene. PLoS ONE 6, e20774 (2011).
Yuh, Y. S., Liu, J. Y. & Shaio, M. F. Chromosome number of Trichomonas vaginalis. J. Parasitol. 83, 551–553 (1997).
Conrad, M. et al. Microsatellite polymorphism in the sexually transmitted human pathogen Trichomonas vaginalis indicates a genetically diverse parasite. Mol. Biochem. Parasitol. 175, 30–38 (2011).
Strese, A., Backlund, A. & Alsmark, C. A recently transferred cluster of bacterial genes in Trichomonas vaginalis-lateral gene transfer and the fate of acquired genes. BMC Evol. Biol. 14, 119 (2014).
Pritham, E. J., Putliwala, T. & Feschotte, C. Mavericks, a novel class of giant transposable elements widespread in eukaryotes and related to DNA viruses. Gene 390, 3–17 (2007).
Wertheim, J. O., Murrell, B., Smith, M. D., Kosakovsky Pond, S. L. & Scheffler, K. RELAX: detecting relaxed selection in a phylogenetic framework. Mol. Biol. Evol. 32, 820–832 (2015).
Glemin, S., Francois, C. M. & Galtier, N. Genome evolution in outcrossing vs. selfing vs. asexual species. Methods Mol. Biol. 1910, 331–369 (2019).
Manni, M., Berkeley, M. R., Seppey, M., Simao, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Mendes, F. K., Vanderpool, D., Fulton, B. & Hahn, M. W. CAFE 5 models variation in evolutionary rates among gene families. Bioinformatics 36, 5516–5518 (2021).
de Miguel, N. et al. Proteome analysis of the surface of Trichomonas vaginalis reveals novel proteins and strain-dependent differential expression. Mol. Cell Proteom. 9, 1554–1566 (2010).
Kochanowsky, J. A. et al. Trichomonas vaginalis extracellular vesicles up-regulate and directly transfer adherence factors promoting host cell colonization. Proc. Natl. Acad. Sci. USA 121, e2401159121 (2024).
Molgora, B. M. et al. A novel trichomonas vaginalis surface protein modulates parasite attachment via protein: host cell proteoglycan interaction. mBio 12, e03374–20 (2021).
Nievas, Y. R. et al. Extracellular vesicles released by anaerobic protozoan parasites: current situation. Cell Microbiol. 22, e13257 (2020).
Zimmann, N. et al. Proteomic analysis of Trichomonas vaginalis phagolysosome, lysosomal targeting, and unconventional secretion of cysteine peptidases. Mol. Cell Proteom. 21, 100174 (2022).
Nievas, Y. R. et al. Membrane-shed vesicles from the parasite Trichomonas vaginalis: characterization and their association with cell interaction. Cell Mol. Life Sci. 75, 2211–2226 (2018).
Stafkova, J. et al. Dynamic secretome of Trichomonas vaginalis: case study of beta-amylases. Mol. Cell Proteom. 17, 304–320 (2018).
Nievas, Y. R. et al. Protein palmitoylation plays an important role in Trichomonas vaginalis adherence. Mol. Cell Proteom. 17, 2229–2241 (2018).
Twu, O. et al. Trichomonas vaginalis exosomes deliver cargo to host cells and mediate hostratioparasite interactions. PLoS Pathog. 9, e1003482 (2013).
Mpeyako, L. A. et al. Comparative genomics between Trichomonas tenax and Trichomonas vaginalis: CAZymes and candidate virulence factors. Front. Microbiol. 15, 1437572 (2024).
Smith, M. D. et al. Less is more: an adaptive branch-site random effects model for efficient detection of episodic diversifying selection. Mol. Biol. Evol. 32, 1342–1353 (2015).
Coceres, V. M. et al. The C-terminal tail of tetraspanin proteins regulates their intracellular distribution in the parasite Trichomonas vaginalis. Cell Microbiol. 17, 1217–1229 (2015).
Kowalczyk, A. et al. RERconverge: an R package for associating evolutionary rates with convergent traits. Bioinformatics 35, 4815–4817 (2019).
Zubacova, Z., Cimburek, Z. & Tachezy, J. Comparative analysis of trichomonad genome sizes and karyotypes. Mol. Biochem. Parasitol. 161, 49–54 (2008).
Piacentini, L. et al. Transposons, environmental changes, and heritable induced phenotypic variability. Chromosoma 123, 345–354 (2014).
Bradic, M., Warring, S. D., Low, V. & Carlton, J. M. The Tc1/mariner transposable element family shapes genetic variation and gene expression in the protist Trichomonas vaginalis. Mob. DNA 5, 12 (2014).
Warring, S. D. et al. Small RNAs are implicated in regulation of gene and transposable element expression in the protist Trichomonas vaginalis. mSphere 6, e01061–20 (2021).
Birchler, J. A. & Yang, H. The multiple fates of gene duplications: deletion, hypofunctionalization, subfunctionalization, neofunctionalization, dosage balance constraints, and neutral variation. Plant Cell 34, 2466–2474 (2022).
Lynch, M. The Origins of Genome Architecture (Sunderland, Sinauer Associates, Inc., 2007).
Bradic, M. & Carlton, J. M. Does the common sexually transmitted parasite Trichomonas vaginalis have sex?. PLoS Pathog. 14, e1006831 (2018).
Alrefaei, A. F. et al. Multilocus analysis resolves the European finch epidemic strain of Trichomonas gallinae and suggests introgression from divergent trichomonads. Genome Biol. Evol. 11, 2391–2402 (2019).
Kellerova, P. & Tachezy, J. Zoonotic Trichomonas tenax and a new trichomonad species, Trichomonas brixi n. sp., from the oral cavities of dogs and cats. Int. J. Parasitol. 47, 247–255 (2017).
Bragulla, H. H. & Homberger, D. G. Structure and functions of keratin proteins in simple, stratified, keratinized and cornified epithelia. J. Anat. 214, 516–559 (2009).
Mahdy, M. A. A. & Mohammed, E. S. I. Anatomical, histological, and scanning electron microscopic features of the esophagus and crop in young and adult domestic pigeons (Columba livia Domestica). BMC Vet. Res. 20, 428 (2024).
Midlej, V. & Benchimol, M. Trichomonas vaginalis kills and eats-evidence for phagocytic activity as a cytopathic effect. Parasitology 137, 65–76 (2010).
Mercer, F. et al. Leukocyte lysis and cytokine induction by the human sexually transmitted parasite Trichomonas vaginalis. PLoS Negl. Trop. Dis. 10, e0004913 (2016).
Huang, K. Y. et al. Adaptive responses to glucose restriction enhance cell survival, antioxidant capability, and autophagy of the protozoan parasite Trichomonas vaginalis. Biochim. Biophys. Acta 1840, 53–64 (2014).
Huang, K. Y. et al. Potential role of autophagy in proteolysis in Trichomonas vaginalis. J. Microbiol. Immunol. Infect. 52, 336–344 (2019).
Pereira-Neves, A. & Benchimol, M. Phagocytosis by Trichomonas vaginalis: new insights. Biol. Cell 99, 87–101 (2007).
Chin, C. S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
English, A. C. et al. Mind the gap: upgrading genomes with Pacific Biosciences RS long-read sequencing technology. PLoS ONE 7, e47768 (2012).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Bruna, T., Hoff, K. J., Lomsadze, A., Stanke, M. & Borodovsky, M. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genom. Bioinform. 3, lqaa108 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Paysan-Lafosse, T. et al. InterPro in 2022. Nucleic Acids Res. 51, D418–D427 (2023).
Gligorijevic, V. et al. Structure-based protein function prediction using graph convolutional networks. Nat. Commun. 12, 3168 (2021).
Kulmanov, M., Zhapa-Camacho, F. & Hoehndorf, R. DeepGOWeb: fast and accurate protein function prediction on the (Semantic) Web. Nucleic Acids Res. 49, W140–W146 (2021).
Warburton, P. E., Giordano, J., Cheung, F., Gelfand, Y. & Benson, G. Inverted repeat structure of the human genome: the X-chromosome contains a preponderance of large, highly homologous inverted repeats that contain testes genes. Genome Res. 14, 1861–1869 (2004).
Bao, W., Kojima, K. K. & Kohany, O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Hafez, A. I. et al. Client applications and server-side Docker for management of RNASeq and/or VariantSeq workflows and pipelines of the GPRO suite. Genes (Basel) 14, 267 (2023).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117, 9451–9457 (2020).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Shi, J. & Liang, C. Generic repeat finder: a high-sensitivity tool for genome-wide de novo repeat detection. Plant Physiol. 180, 1803–1815 (2019).
Kelley, D. R., Schatz, M. C. & Salzberg, S. L. Quake: quality-aware detection and correction of sequencing errors. Genome Biol. 11, R116 (2010).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 11, 1432 (2020).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19, ii215–ii225 (2003).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Weilguny, L. & Kofler, R. DeviaTE: assembly-free analysis and visualization of mobile genetic element composition. Mol. Ecol. Resour. 19, 1346–1354 (2019).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Littmann, M., Heinzinger, M., Dallago, C., Olenyi, T. & Rost, B. Embeddings from deep learning transfer GO annotations beyond homology. Sci. Rep. 11, 1160 (2021).
Loytynoja, A. Phylogeny-aware alignment with PRANK. Methods Mol. Biol. 1079, 155–170 (2014).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Acknowledgements
We thank Mari Shiratori, Sally D. Warring, Martina Bradic, Akash Sookdeo, Charlotte Darby, and Srividya Ramakrishnan for initial wet lab and genome analyses. We thank Ellen Pritham for supplying canonical Maverick sequences. Research reported in this publication was partially supported by: the NYU IT High Performance Computing resources, services, and staff expertise; CMRPD1M0571-2 from Chang Gung Memorial Hospital and NSTC–110–2320B–182–016–MY3 from National Science and Technology Council, Taiwan; Australian Government Wildlife Exotic Disease Preparedness Program; and the National Institute Of Allergy And Infectious Diseases of the National Institutes of Health under Award Number R21AI149449 and U24AI183870. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
J.M.C. and M.C.S. designed the study, provided the funding for resequencing T. vaginalis, supervised the work, and interpreted the analyses. Y.A.G., C.K.J., K.H.R. and R.G. provided Trichomonas isolates. P.-J.H., Y.-M.Y., C.-C.L., H.L, T.-W.C., P.T. and C.-H.C. provided the T. tenax, P. hominis, T. stableri CA015840, T. vaginalis strain CDC 085, and T. vaginalis strain NYH 286 Illumina whole genome sequence data, and A.P. and S.R.R. provided the T. gallinae TGAL, T. sp. genotype 1c, and T. sp. genotype 2a Illumina whole genome sequence data. S.A.S., J.C.O., F.C.-H., F.B., T.R.R.-B. and H.L. undertook genome annotation and analysis, and additionally F.B. and F.C.-H. generated the T. stableri BTPI-3 genome sequence and undertook wet lab work. H.M. and I.L. undertook Hi.-C. H.S., C.C., D.B., V.G. and R.A.B. helped with T. vaginalis functional annotation. All authors contributed to the writing of the manuscript and approved the final version before submission.
Corresponding author
Ethics declarations
Competing interests
I.L. and H.M. are employees of Phase Genomics, a company that provides Hi-C services and products. The following author wishes to disclose his industry relations although there are no competing interests to the work published here: R.A.B. is an employee of Genentech/Roche. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Neil Young and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sullivan, S.A., Orosco, J.C., Callejas-Hernández, F. et al. Comparative genomics of the parasite Trichomonas vaginalis reveals genes involved in spillover from birds to humans. Nat Commun 16, 6487 (2025). https://doi.org/10.1038/s41467-025-61483-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61483-w
This article is cited by
-
Chromosome-level genome assembly of Trichomonas vaginalis strain IR-78 (ATCC 50138)
Scientific Data (2025)